SUMMARY
Background
Cell size control systems, coupled with apoptotic and cell proliferation regulatory mechanisms, determine the overall dimensions of organs and organisms, and their dysregulation can lead to tumor formation. The interrelationship between regulatory mechanisms for cell growth and apoptosis during normal development and cancer is not understood. The TRK-fused gene (TFG) promotes tumorigenesis when present in chromosomal rearrangements from a variety of human cancer types by unknown mechanisms. Apaf1/CED-4 is essential for apoptosis but has not been shown to function in cell growth control.
Results
We found that loss of TFG-1, the TFG homologue in Caenorhabditis elegans, results in supernumerary apoptotic corpses, while its overexpression is sufficient to inhibit developmentally programmed cell death. TFG-1 is also required for both cells and nuclei to grow to normal size. Further, we found that CED-4 is required to inhibit cell growth in animals lacking all TFG-1. However, caspases, the downstream effectors of CED-4-mediated apoptosis, are not required in TFG-1/CED-4-regulated cell size control. CED-4 acts broadly to inhibit cell growth by antagonizing other conserved cell size-regulating proteins, including cAMP response element-binding (CREB) protein, translation initiation factor eIF2B, and the nucleolar p53-interacting protein nucleostemin.
Conclusions
These findings show that TFG-1 is both a suppressor of apoptosis and is essential for normal cell size control, suggesting that abnormalities in the cell growth-promoting and apoptosis-inhibiting functions of TFG may be responsible for its action in tumorigenesis. Further, they reveal that CED-4 plays a pivotal role both in activating apoptosis and restricting cell and nuclear size, thereby determining the appropriate overall size of an animal. Thus, these findings reveal links between the control mechanisms for apoptosis and cell growth.
INTRODUCTION
Cells actively modulate growth parameters to achieve reproducible dimensions appropriate to their functions. Cell size control systems, coupled with regulatory mechanisms for apoptotic cell death and cell proliferation, determine the overall dimensions of organs and organisms, and dysregulation of these processes can lead to cancer. Apoptosis and cell size are controlled by largely distinct molecular regulatory events.
The pathways regulating apoptosis are highly conserved among animals. The core apoptotic regulatory pathway in C. elegans consists of four proteins (CED-3, CED-4, CED-9, and EGL-1) that are critical for the switch between cell survival and apoptosis [1]. CED-3, a member of the caspase family of cysteine proteases and the main executor of apoptosis, is triggered to undergo autoproteolytic activation into its pro-apoptotic form by CED-4, a homologue of mammalian Apaf-1. CED-9, the worm equivalent of mammalian proto-oncoprotein Bcl-2, promotes cell survival by binding to CED-4 and preventing it from activating CED-3. In cells fated to undergo apoptosis, the anti-apoptotic activity of CED-9 is blocked by EGL-1, a BH3-only domain protein.
In contrast to apoptosis, several independent mechanisms appear to regulate cell size. The best-understood mechanism controlling cell and body size in C. elegans is the Sma/Mab pathway, involving members of the TGF-β superfamily and their transducers [2]. Smads encoded by sma-2, sma-3, and sma-4 transduce TGF-β signals into the nucleus and regulate cell and body size through transcriptional regulation of lon-1 and other downstream factors. The Sma/Mab pathway is negatively regulated by LON-2, a member of the glypican family [3]. Epistasis studies have shown that LON-2 functions upstream of the Sma/Mab pathway, as mutations in lon-2 produce long animals only when the TGF-β ligand DBL-1 and the TGF-β receptor SMA-6 are intact [3]. A number of other cell size regulatory mechanisms exist, including the insulin/insulin-like growth factor (IGF) and target of Rapamycin (TOR) signaling pathways [4]; however, their roles in cell size regulation in C. elegans are less defined.
Though the control of cell growth and apoptosis must be coordinated, little is known about how this is achieved. We have found that the essential apoptotic regulator, ced-4, and the C. elegans homologue of the human TFG proto-oncogene participate in both cell size control and apoptosis, suggesting that they link these two processes. The human TFG gene was identified from chromosomal rearrangements in a variety of malignant tumor types, including papillary thyroid carcinomas, anaplastic large cell lymphomas, and extraskeletal myxoid chondrosarcomas [5–8] (Fig. 1A). All resulting protein chimeras contain an in-frame fusion of TFG at the amino-terminus and some have been shown to possess oncogenic activity [5, 7, 9]. The TFG portion of the oncogenic fusion form is essential for this transforming activity as well as for interactions with other proteins [10–13]. TFG is upregulated in response to TALL-1 [14], a member of the tumor necrosis factor family implicated in B cell proliferation and autoimmunity. TFG was recently reported to interact with PTEN [15], a key tumor suppressor that modulates cell growth, division, and death. These findings raise the possibility that TFG might contribute to cancer by regulating such cellular activities.
Figure 1. Participation of tfg-1 in cell death regulation.
(A) Schematic representation of human and C. elegans TFG homologues and human TFG protein chimeras in various cancers. Red box, SH2 binding motif; cc, coiled-coil domain; black box, SH3 binding motif. (B) Histogram of cell corpse numbers in comma stage embryos of the given genotype. (C, D) Histogram of cell corpse numbers in comma stage embryos without (black bars) and with (yellow bars) heat-shock. Heat-shock leads to reduced number of cell corpses specifically in hs-tfg-1 transgenic animals (C) but not in control hs-lacZ animals (D).
We report here that the single TFG orthologue in C. elegans, tfg-1 [16] (Fig. 1A), both suppresses apoptotic cell death and activates cell and nuclear growth by a pathway that is apparently independent of TGF-β and insulin signaling pathways. The reduced cell and nuclear size seen in the absence of TFG requires CED-4 function, implicating CED-4 in cell growth inhibition in addition to its pro-apoptotic action. Moreover, we found that three other proteins whose function is required for normal cell and body size, are also antagonized by CED-4, suggesting that CED-4 performs a general role in restricting cell growth and body size. Thus, CED-4 may provide a mechanism for coordinating the processes of cell death and cell growth/cell size control.
RESULTS
TFG-1 regulates apoptotic cell death
To evaluate its possible role in growth, proliferation, and apoptosis, we investigated tfg-1 gene function in C. elegans. We found that inactivation of tfg-1 by RNAi results in a significant increase in the number of apoptotic corpses during embryogenesis (Fig. 1B). At the “comma” stage of embryogenesis, when much programmed cell death occurs, tfg-1(RNAi) embryos contain an average of 12.6 ± 2.8 corpses, significantly higher than the number in control wild-type embryos (8.5 ± 2.7 corpses; Wilcoxon rank sum, p = 2.98 × 10−6). A similar elevation in cell corpse number is also seen in a cell corpse engulfment-defective ced-1(e1735) mutant (Supplemental Figure S1A). This elevated cell death was blocked by mutations in the core cell death pathway (data not shown), indicating that most cell death in tfg-1(RNAi) worms relies on the canonical CED-3, CED-4-dependent cell death pathway. However, later in development, some cells undergo CED-3-independent death in tfg-1(RNAi) animals (Supplemental Table S1). Such CED-4-dependent, CED-3-independent apoptosis has also been reported with the ICD-1 cell death suppressor [17].
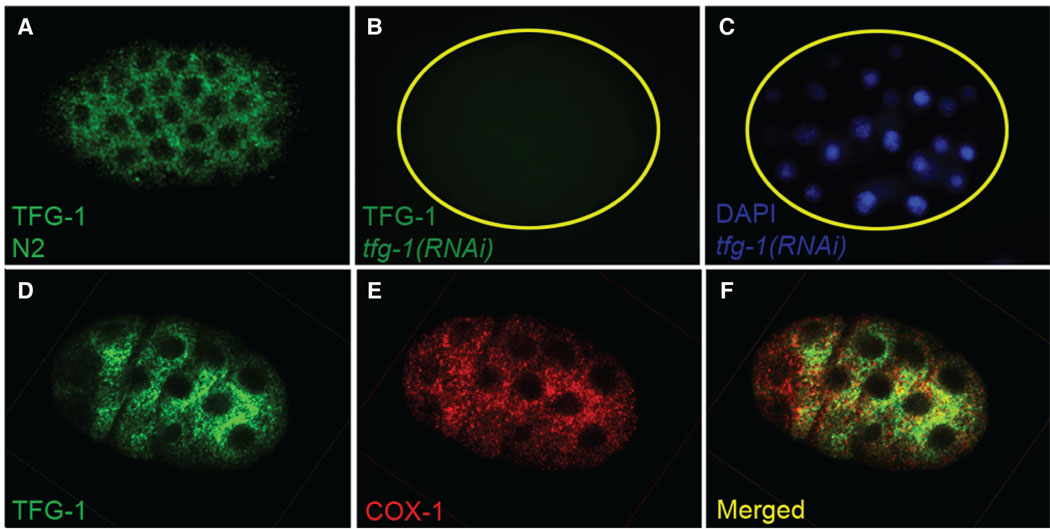
Localization of TFG-1 is consistent with a role in apoptosis: immunoreactive TFG-1 is detectable as early as the two-cell stage of embryogenesis and its expression continues throughout embryogenesis (e.g., Fig. 2A, D). Closer examination of the subcellular localization of TFG-1 revealed a punctuate staining pattern in the cytoplasm, which partially colocalizes with mitochondrial COX1 (Fig. 2D, E, F). As mitochondria are key sites for apoptotic regulators [1], the partial mitochondrial localization of TFG-1 and the tfg-1(RNAi) phenotype are suggestive of a possible anti-apoptotic action in this organelle.
Figure 2. Intracellular localization of TFG-1.
(A) An example of the TFG-1 staining in the embryo. (B) TFG-1 is undetectable in tfg-1(RNAi) embryos. (C) The DAPI image of the embryo in panel B. (D, E, F) TFG-1 partially colocalizes with mitochondria, marked with anti-COX-1 antibody. D, an embryo stained with anti-TFG-1 antibody; E, the same embryo stained with anti-COX-1 antibody; F, the merged image of panels D and E.
The elevation in cell corpse number in tfg-1(RNAi) animals results either from inappropriate cell death or reduced engulfment of cell corpses. To further assess the role of tfg-1 in apoptotic regulation, we asked whether its overexpression is sufficient to suppress developmentally programmed cell death by driving it ubiquitously under control of a heat-shock promoter. Embryos overexpressing TFG-1 contain significantly fewer cell corpses at comma stage (average of 4.1 ± 1.9) than in the non-heat shocked controls (6.9 ± 1.9, Wilcoxon rank sum, p = 1 × 10−6, Fig. 1C). This decrease in cell corpse number reflects bona fide suppression of the apoptotic pathway, as we observed extra surviving cells in the anterior pharynx of the resulting larvae in the same positions as the extra survivors in mutants defective for essential pro-apoptotic genes (Supplemental Figure S1B). Thus, TFG-1 is sufficient, and may be necessary, to inhibit the normal apoptotic pathway.
TFG-1 is required to regulate normal nuclear, cell, and body size
The temporal flexibility of gene inactivation by RNAi provides an opportunity to study functions of essential genes that might otherwise be masked by lethality. We found that while elimination of TFG-1 from embryos results in lethality prior to hatching, removal of all detectable TFG-1 by RNAi from developing larvae that had expressed it at normal levels during embryogenesis does not prevent growth to adulthood. However, such worms are much smaller than their wild-type counterparts (Fig. 3A, B; average length 0.73 ± 0.13 mm vs. 1.34 ± 0.1 mm respectively), revealing that TFG-1 is required during postembryonic development to regulate body size. Body and organ size in animals is a function of total cell number and individual cell size. As TFG-1 suppresses cell death, the reduced size of tfg-1(RNAi) worms might be a consequence of decreased cell number owing to cell loss through increased apoptosis. This was not the case: we found that the total number of DAPI-stained somatic nuclei quantifiable in tfg-1(RNAi) worms is not significantly different from that in wild-type (938.7±44.7; n=27, and 941.3±43.6; n=25, respectively; see Table 1). In contrast, we found that the average size of both locomotory muscle cells and epidermal cells, as well as their respective nuclei, is dramatically reduced in tfg-1(RNAi) adult worms than in wildtype (Fig. 3C, Table 1, and not shown). The reduction in cell size accounts for the reduced body size in these animals. Thus, TFG-1 influences body size primarily by controlling the overall dimensions of individual cells.
Figure 3. Cell size regulation by TFG-1.
(A) Adult tfg-1(RNAi) worms are much smaller than the same stage wild-type adult worms. (B) The body length distribution of control (black bars) and tfg-1(RNAi) (white bars) worms. Body length data were pooled from three independent RNAi experiments. (C) Muscle cells and their nuclei are smaller in adult tfg-1(RNAi) worms. Scale bars: 20µm. (D) Seam cell size during larval stages. ** Student t test, p<0.01.
Table 1.
ced-4-dependent reduction of cell size in tfg-1(RNAi) mutants.
| Genotype | L4440(RNAi) | tfg-1(RNAi) | |||
|---|---|---|---|---|---|
| Number of muscle | WT | 94.6 ± 0.9 | (n=15) | 94.2 ± 1.9 | (n=15) |
| cells | |||||
| Muscle cell size | WT | 1709.2 ± 224.9 | (n=27) | 679 ± 137.9 | (n=42) ** |
| (µm2) * | ced-4(n1162) | 1322.5 ± 282 | (n=50) | 1254.4 ± 237.5 | (n=56) |
| Muscle nucleus size | WT | 79.7 ± 15.8 | (n=27) | 46.3 ± 9.7 | (n=42) ** |
| (µm2) * | ced-4(n1162) | 71.2 ± 14.9 | (n=50) | 69.1 ± 14.3 | (n=56) |
| Total cell number | WT | 941.3 ± 43.6 | (n=27) | 938.7 ± 44.7 | (n=25) |
RNAi was performed by feeding with either a control feeder (L4440, carrying an uninserted plasmid) or a tfg-1 RNAi feeder. Muscle cell and nucleus size were determined as the area of the largest section through the cell.
Student t test, p<0.01.
Cell size is determined by the combined action of two opposing processes, cell growth and cell division. The number of muscle cells in adult tfg-1(RNAi) worms is normal (Table 1), suggesting that TFG-1 does not affect cell division. During postembryonic development, all but one of the lateral epidermal (seam) cells divide during each larval stage [18]. This invariant cell division pattern provides a useful tool to further distinguish the influence of TFG-1 on cell division vs. growth. By examining a marker of apical junctions surrounding epithelial cells [19], we found that seam cells undergo a similar division pattern in tfg-1(RNAi) larvae as in wildtype, through the molt to adulthood, confirming that TFG-1 does not alter the number of cell divisions. In contrast, we found that seam cells are substantially growth-inhibited in tfg-1(RNAi) worms starting at the L3 stage, demonstrating that TFG-1 is a novel regulator of cell growth and size. This is apparent from a decrease in the longitudinal dimension (Fig. 3D). This effect is particularly dramatic by the end of development: the body volume and volume of the diminutive cells in tfg-1(RNAi) adults are only ~30% those of wildtype cells (Fig. 5D). Thus, TFG-1 is largely responsible for determining the size of adult worms.
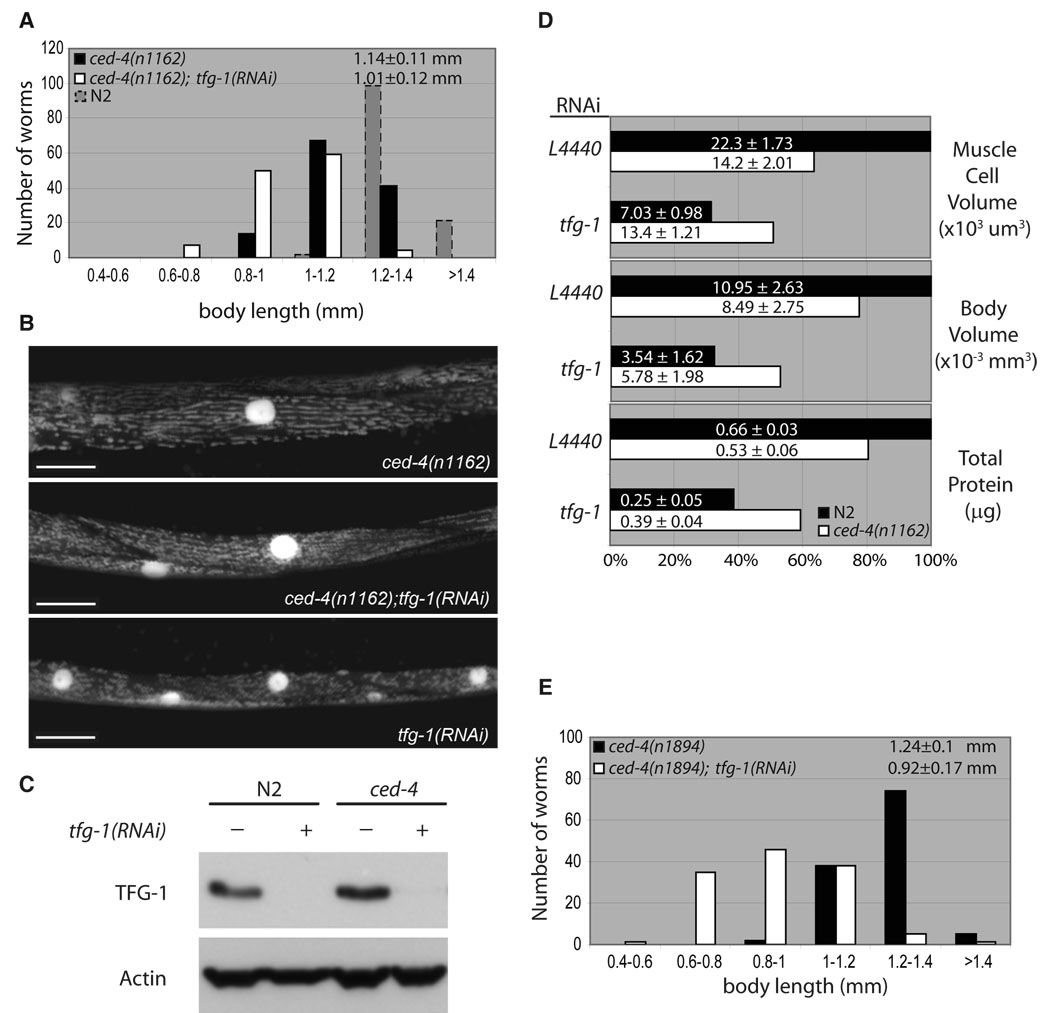
Figure 5. CED-4 is required for TFG-1-dependent body and cell size regulation.
(A) The adult body length distribution of control ced-4(n1162); L4440(RNAi) (black bars) and ced-4(n1162);tfg-1(RNAi) (white bars) worms. The n1162 allele is the reference allele that results in a stop codon at amino acid 80. Body length data were pooled from three independent RNAi experiments. (B) Inactivation of ced-4 restores the normal muscle cell and nucleus size of adult tfg-1(RNAi) worms. Scale bars: 20 µm. (C) Western blot of extracts from wildtype and ced-4(−) worms without (−) and with (+) treatment of tfg-1 RNAi. tfg-(RNAi) eliminates all detectable TFG-1. (D) CED-4-dependent cell and body volume changes are proportional to total protein content. More than 30 worms per strain were used to measure body volume. Total protein content is expressed as the mean ± s.d. from three independent experiments. (E) Rescue of the Sma phenotype of tfg-1(RNAi) by ced-4(n1894). The n1894 allele introduces a stop codon at amino acid 401, resulting in a truncated protein.
The small locomotory muscle cells in tfg-1(RNAi) worms also contain smaller nuclei (Table 1), prompting us to examine whether DNA content is reduced in nuclei of TFG-1-depleted worms. A change in DNA content is often associated with an alteration in nuclear volume and cell size: for example, the large size of intestinal cells in worms is accompanied by increased DNA ploidy of up to 32N as a result of endoreduplication [20]. Moreover, somatic ploidy has recently been shown to be causally linked to body size in C. elegans [21]. Nevertheless, ploidy is only one factor underlying cell and nuclear size changes: many cells of dramatically different size remain diploid throughout life and mutants with smaller body size but the same ploidy as wildtype have been reported [21]. We found no significant difference in the average ploidy of intestinal cells between dwarf tfg-1(RNAi) and control animals (30.4N ± 6.6, n=26 vs. 31.7N ± 3.7, n=22, respectively, Student t test, p=0.41; Fig. 4A). In addition, we observed that epidermal nuclei, which also undergo larval endoreduplication, contained a similar DNA content in tfg-1(RNAi) and wildtype adults (7.4N ± 2.3, n=26 vs. 8.4N ± 1.6, n=22, respectively, Student t test, p=0.1). Thus, differences in nuclear DNA content cannot explain the dramatic effect of TFG-1 on the dimensions of nuclei and cells.
Figure 4. TFG-1 regulates body and cell size independent of ploidy, the Sma/Mab pathway, and the caspase activity.
(A) Ploidy vs. body size in diminutive tfg-1(RNAi) and control L4440(RNAi) worms. (B) tfg-1(RNAi) and mutations of the Sma/Mab pathway show additivity in body size reduction. (C) A ced-3(−) mutation does not alter the small body size of tfg-1(RNAi) worms. (D) Removal of all known caspase activity does not restore the small body size of tfg-1(RNAi) worms.
We next tested for possible genetic interactions between TFG-1 and other cell size regulatory pathways. We found that inactivation of TFG-1 in sma-3 and sma-4 mutant animals additively reduced body size (Fig. 4B), implying that the TGF-β pathway and TFG-1 act in parallel to control cell size. Similarly, the elongated body phenotype resulting from a mutation in lon-2 is suppressed by tfg-1(RNAi) in a simple additive relationship (Fig. 4B), implying that TFG-1 and LON-2 act independently to regulate body size. This contrasts with the strong epistatic relationship between SMA-4 and LON-2, which are believed to act in the same pathway: sma-4(−); lon-2 double mutants are as small as sma-4(−) single mutant animals [22]. Mutations that disrupt IGF or TOR signaling pathways also showed no conspicuous interactions with TFG-1 in cell size phenotype (not shown). Human tumor suppressor PTEN [15], which is known to regulate cell size, interacts with TFG; however, we found that mutations that eliminate the function of worm PTEN, DAF-18, do not alter the cell size phenotype of tfg-1 (not shown). Thus, TFG-1 apparently functions in a novel pathway independent of TGF-β and insulin signaling to control cell growth and size in C. elegans.
CED-4 reduces cell size in mutants lacking TFG-1
Our discovery that TFG-1 functions both to repress apoptosis and regulate cell size prompted us to ask whether proteins known for their roles in apoptosis might also act in cell size regulation. We found that key apoptotic factors, CED-3, CED-9, and EGL-1, do not significantly affect the diminutive cell size in tfg-1(RNAi) animals (Fig. 4C, Supplemental Fig. S2). In striking contrast, we observed that two strong loss-of-function alleles of ced-4 strongly suppress the cell size phenotypes of adult tfg-1(RNAi) worms (Fig. 5A, E). For example, ced-4(n1162); tfg-1(RNAi) worms (average length 1.01 ± 0.12 mm, n = 120) are similar in size to ced-4(n1162) single mutant animals (average length 1.14 ± 0.11 mm, n = 122). Consistent with the restored body size, we found that muscle cells in ced-4(n1162); tfg-1(RNAi) worms are much larger than those in tfg-1(RNAi) worms and are similar in size to muscles in ced-4(n1162) single mutants (Fig. 5B, Table 1). ced-4 mutants do not appear to show reduced susceptibility to RNAi [17]. We further confirmed the efficacy of tfg-1 RNAi in the ced-4 mutants: TFG-1 was undetectable in extracts from both tfg-1(RNAi) and ced-4(n1162); tfg-1(RNAi) animals (Fig. 5C).
The restoration of body and cell dimensions by ced-4 mutations demonstrates that CED-4 is not only necessary to activate the apoptotic program, but also functions to limit cellular dimensions. In apoptosis, the ced-4 locus possesses dual functions as both a pro-and anti-apoptotic regulator [23]. A similar dual role in both restricting and promoting cell growth might also exist in CED-4-dependent cell size regulation, as we observed that ced-4(−) animals have a slightly smaller body and cell size than wildtype worms (Figure 5A, D).
Caspases are the well-documented downstream effectors of CED-4 and Apaf-1 in apoptosis and the only proteins in C. elegans known to possess caspase activity are CED-3 and CSP-1 [24]. We found that removal of either one or both of these caspases does not alter the small body size of tfg-1(RNAi) animals (Fig. 4C, D). For example, the average body length of adult ced-3(n717); csp-1(tm917); tfg-1(RNAi) worms (0.74 ± 0.13 mm; n = 108) is virtually identical to that of tfg-1(RNAi) worms (0.73 ± 0.13 mm; n = 137). Further, we found that ubiquitous high level expression of baculovirus p35, a potent pan-caspase inhibitor that effectively blocks normal apoptotic death in C. elegans [25], does not detectably alter the size of tfg-1(RNAi) animals (not shown). These findings indicate that while caspases are critical for CED-4-mediated apoptosis, CED-4 apparently mediates cell size control independent of caspase function.
Regulation of ribosomal biogenesis and protein synthesis is known to govern cell growth control. However, the protein content of some C. elegans mutants with altered body size, including the larger-than-normal egl-4 mutants and diminutive sma-1 mutants, does not change in proportion to their respective body sizes. In these cases, the alteration in body size has been attributed to changes in DNA content or fluid content [26]. In contrast, we observed CED-4-dependent reduction in total protein content of tfg-1(RNAi) worms that parallels the CED-4-dependent decrease in body and cell size (Fig. 5D). The strong correlation between body size and protein content suggests that CED-4/TFG-1 likely controls cell and body size through modulation of the protein production and accumulation machinery.
CED-4 antagonizes the function of CREB, eIF2B, and nucleostemin in cell size regulation
Suppression of the diminutive cell size phenotype of tfg-1(RNAi) by ced-4 mutations might indicate that CED-4 antagonizes normal cell size regulatory mechanisms, or may instead reflect a specific regulatory relationship between TFG-1 and CED-4. To address this issue, we investigated whether mutants of other cell size regulators show a CED-4-dependent decrease in cell size. The CREB/ATF family of transcription factors is required for cell growth and survival [27] and CREB null mice are ~30% smaller than their wild-type counterparts [28]. Furthermore, cells expressing a dominant-negative form of CREB are reduced in size [29]. We found that RNAi knockdown of the gene encoding CREB in C. elegans similarly results in a profound decrease in body size (Fig. 6A) that is even more severe than that seen with tfg-1 mutants. The volume of cells in these mutant adults is remarkably only ~20% that of wildtype cells. This profound cell size phenotype was strongly suppressed by a ced-4(n1162) mutation (Fig. 6A), underscoring the broad role of CED-4 in cell size control.
Figure 6. CED-4 acts broadly to regulate body size.
(A) Inactivation of ced-4 suppresses the Sma phenotype of animals depleted for the function of the C. elegans CREB gene (F57B10.1) by RNAi. (B) Inactivation of ced-4 suppresses the Sma phenotype of iftb-1(RNAi) animals. (C) Inactivation of ced-4 suppresses the Sma phenotype of nst-1(RNAi) animals. Student t test, p<0.01. (D) Model for the action of TFG-1 and three other cell size regulators with CED-4 in cell size/growth control and apoptosis.
Factors that mediate translation affect cell size; for example, eIF2B mediates hypertrophic growth of cardiac myocytes [30]. We found that RNAi of the eIF2B-encoding gene, iftb-1, in C. elegans results in a small body size phenotype, which is also suppressed by a ced-4 mutation (Fig. 6B). Thus, the action of CED-4 in limiting cell size extends to regulation of cell growth through a translational regulator. Nucleostemin, a nucleolar GTPase enriched in proliferating stem cells and cancer cells, has been implicated in regulating ribosomal assembly [31]. Nucleostemin can physically interact with, and modulate the activity of, tumor suppressor p53. Loss of nucleostemin results in p53-dependent apoptosis and cell cycle arrest [32, 33] and its overexpression results in cell cycle arrest and formation of giant cells [34]. We found that knocking down expression of the worm homolog of nucleostemin results in a reduced body size phenotype that is also suppressed by a ced-4(n1162) mutation (Fig. 6C).
These observations demonstrate that CED-4 represses cell growth as a result of mutations in several apparently distinct pathways, revealing that it acts broadly to regulate cell and body size.
DISCUSSION
Human TFG is involved in several different cancer types; however, the mechanism of its action in tumorigenesis is unclear. We have found that C. elegans TFG-1 acts both as a novel apoptotic suppressor and an activator of cell growth. These observations help to illuminate the role of human TFG in cancer: the inappropriate activity of oncogenic forms of TFG-1 might promote tumor development by inhibiting the normal cell death program, which prevents tumor development, while also activating the cell growth regulatory machinery. In both of these processes, TFG-1 appears to act antagonistically with CED-4, which is essential to activate the apoptotic program and to dramatically reduce the size of cells when TFG-1 is absent.
CED-4 and its mammalian homologue, Apaf-1, have been well studied for their critical roles in apoptotic regulation. Both act as adapter proteins that trigger the activation of downstream caspases, thereby executing the cell death program. However, several recent studies have demonstrated that CED-4/Apaf-1 has functions in other processes in addition to apoptosis. Apaf-1 mediates DNA-damage-induced cell cycle arrest at the S phase [35, 36]. This DNA damage checkpoint function is conserved between human and worms: loss of CED-4 function leads to a similar cell-cycle arrest defect in the animal’s germline after exposure to γ or UVC irradiation [36]. CED-4 has also been found to be required for hypoxic and volatile anesthetic preconditioning in C. elegans [37, 38].
Our findings that CED-4 opposes the cell size regulatory action of TFG-1 reveal a new aspect of the non-apoptotic vital functions of CED-4. CED-4 likely performs a general role in body and cell size control as it also limits the size of cells in animals depleted for three other cell growth-regulating genes. It is interesting to note that cells developmentally programmed to die in C. elegans are much smaller than their surviving sisters at the time they are born [39]. The CED-4 dependence of cell size control in the adult soma and in cells doomed to die through the developmental program may reflect a related process mediated by this key apoptotic regulator. Thus, some of the cell death activating machinery, particularly that involving CED-4, may actively promote this diminutive cell size at the time a cell has committed to the cell death pathway. Our results imply that CED-4 is required in many, if not all, cells to regulate the size of surviving cells that never trigger the apoptotic program.
As with human TFG, CREB appears to be a proto-oncoprotein [40]. Both CREB and PTEN, which interacts with TFG, regulate glucose metabolism and functionally interact: for example, PTEN suppresses cyclin D2 expression and therefore cell proliferation in a CREB-dependent manner [41]. We found that RNAi knockdown of either gene in C. elegans leads to a diminutive body size phenotype that is suppressed by inactivation of ced-4. Moreover, similar to tfg-1, RNAi knockdown of CREB also leads to an increase in embryonic cell death (Supplemental Fig. S3). These observations suggest that the cell size regulatory mechanisms of TFG and the CREB transcription factor might act through related pathways.
It has been suggested that C. elegans cell size is influenced by changes in DNA content, protein synthesis, cell cycle, and/or metabolism [42]. We found that TFG-1/CED-4-regulated body and cell size control is largely independent of nuclear DNA content but highly correlated with total protein level. This suggests that a possible mechanism for TFG-1/CED-4 function in cell and body size regulation is to modulate protein production and accumulation. In support of this notion, we found two additional factors associated with protein synthesis, eIF2B and nst-1, that show CED-4-dependent body size regulation.
In summary, our findings on the function of the proto-oncogene TFG-1 and the pro-apoptotic factor CED-4 in apoptosis and cell size regulation suggest a possible link between these two key cellular processes (Fig. 6D). That CED-4 antagonizes the action of at least four genes that function in apparently distinct processes to regulate body, and presumably cell, size, indicates a prominent role of CED-4 in establishing appropriate cell and body size during development.
Supplementary Material
ACKNOWLEDGMENTS
We thank A. M. van der Bliek for the pmyo-3::mt-GFP strain, A. Fire for heat shock vectors, and members of the Rothman laboratory for discussions. Some nematode strains used in this work were provided by the Caenorhabditis Genetics Center, which is funded by the National Institutes of Health National Center for Research Resources. This work was supported by a Cancer Center of Santa Barbara postdoctoral fellowship to L.C. and by grants from the National Institutes of Health to J.H.R.
Footnotes
Publisher's Disclaimer: This is a PDF file of an unedited manuscript that has been accepted for publication. As a service to our customers we are providing this early version of the manuscript. The manuscript will undergo copyediting, typesetting, and review of the resulting proof before it is published in its final citable form. Please note that during the production process errors may be discovered which could affect the content, and all legal disclaimers that apply to the journal pertain.
REFERENCES
- 1.Lettre G, Hengartner MO. Developmental apoptosis in C. elegans: a complex CEDnario. Nat Rev Mol Cell Biol. 2006;7:97–108. doi: 10.1038/nrm1836. [DOI] [PubMed] [Google Scholar]
- 2.Patterson GI, Padgett RW. TGF beta-related pathways. Roles in Caenorhabditis elegans development. Trends Genet. 2000;16:27–33. doi: 10.1016/s0168-9525(99)01916-2. [DOI] [PubMed] [Google Scholar]
- 3.Gumienny TL, Macneil LT, Wang H, de Bono M, Wrana JL, Padgett RW. Glypican LON-2 is a conserved negative regulator of BMP-like signaling in Caenorhabditis elegans. Curr Biol. 2007;17:159–164. doi: 10.1016/j.cub.2006.11.065. [DOI] [PubMed] [Google Scholar]
- 4.Stocker H, Hafen E. Genetic control of cell size. Curr Opin Genet Dev. 2000;10:529–535. doi: 10.1016/s0959-437x(00)00123-4. [DOI] [PubMed] [Google Scholar]
- 5.Greco A, Mariani C, Miranda C, Lupas A, Pagliardini S, Pomati M, Pierotti MA. The DNA rearrangement that generates the TRK-T3 oncogene involves a novel gene on chromosome 3 whose product has a potential coiled-coil domain. Mol Cell Biol. 1995;15:6118–6127. doi: 10.1128/mcb.15.11.6118. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Hernandez L, Pinyol M, Hernandez S, Bea S, Pulford K, Rosenwald A, Lamant L, Falini B, Ott G, Mason DY, Delsol G, Campo E. TRK-fused gene (TFG) is a new partner of ALK in anaplastic large cell lymphoma producing two structurally different TFG-ALK translocations. Blood. 1999;94:3265–3268. [PubMed] [Google Scholar]
- 7.Hernandez L, Bea S, Bellosillo B, Pinyol M, Falini B, Carbone A, Ott G, Rosenwald A, Fernandez A, Pulford K, Mason D, Morris SW, Santos E, Campo E. Diversity of genomic breakpoints in TFG-ALK translocations in anaplastic large cell lymphomas: identification of a new TFG-ALK(XL) chimeric gene with transforming activity. Am J Pathol. 2002;160:1487–1494. doi: 10.1016/S0002-9440(10)62574-6. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Hisaoka M, Ishida T, Imamura T, Hashimoto H. TFG is a novel fusion partner of NOR1 in extraskeletal myxoid chondrosarcoma. Genes Chromosomes Cancer. 2004;40:325–328. doi: 10.1002/gcc.20044. [DOI] [PubMed] [Google Scholar]
- 9.Armstrong F, Duplantier MM, Trempat P, Hieblot C, Lamant L, Espinos E, Racaud-Sultan C, Allouche M, Campo E, Delsol G, Touriol C. Differential effects of X-ALK fusion proteins on proliferation, transformation, and invasion properties of NIH3T3 cells. Oncogene. 2004;23:6071–6082. doi: 10.1038/sj.onc.1207813. [DOI] [PubMed] [Google Scholar]
- 10.Greco A, Fusetti L, Miranda C, Villa R, Zanotti S, Pagliardini S, Pierotti MA. Role of the TFG N-terminus and coiled-coil domain in the transforming activity of the thyroid TRK-T3 oncogene. Oncogene. 1998;16:809–816. doi: 10.1038/sj.onc.1201596. [DOI] [PubMed] [Google Scholar]
- 11.Roccato E, Pagliardini S, Cleris L, Canevari S, Formelli F, Pierotti MA, Greco A. Role of TFG sequences outside the coiled-coil domain in TRK-T3 oncogenic activation. Oncogene. 2003;22:807–818. doi: 10.1038/sj.onc.1206189. [DOI] [PubMed] [Google Scholar]
- 12.Roccato E, Miranda C, Raho G, Pagliardini S, Pierotti MA, Greco A. Analysis of SHP-1-mediated down-regulation of the TRK-T3 oncoprotein identifies Trk-fused gene (TFG) as a novel SHP-1-interacting protein. J Biol Chem. 2005;280:3382–3389. doi: 10.1074/jbc.M407522200. [DOI] [PubMed] [Google Scholar]
- 13.Miranda C, Roccato E, Raho G, Pagliardini S, Pierotti MA, Greco A. The TFG protein, involved in oncogenic rearrangements, interacts with TANK and NEMO, two proteins involved in the NF-kappaB pathway. J Cell Physiol. 2006;208:154–160. doi: 10.1002/jcp.20644. [DOI] [PubMed] [Google Scholar]
- 14.Xu LG, Wu M, Hu J, Zhai Z, Shu HB. Identification of downstream genes up-regulated by the tumor necrosis factor family member TALL-1. J Leukoc Biol. 2002;72:410–416. [PubMed] [Google Scholar]
- 15.Herlevsen M, Oxford G, Ptak C, Shabanowitz J, Hunt DF, Conaway M, Theodorescu D. A novel model to identify interaction partners of the PTEN tumor suppressor gene in human bladder cancer. Biochem Biophys Res Commun. 2007;352:549–555. doi: 10.1016/j.bbrc.2006.11.067. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Mencinger M, Aman P. Characterization of TFG in mus musculus and Caenorhabditis elegans. Biochem Biophys Res Commun. 1999;257:67–73. doi: 10.1006/bbrc.1999.0417. [DOI] [PubMed] [Google Scholar]
- 17.Bloss TA, Witze ES, Rothman JH. Suppression of CED-3-independent apoptosis by mitochondrial betaNAC in Caenorhabditis elegans. Nature. 2003;424:1066–1071. doi: 10.1038/nature01920. [DOI] [PubMed] [Google Scholar]
- 18.Sulston JE, Horvitz HR. Post-embryonic cell lineages of the nematode, Caenorhabditis elegans. Dev Biol. 1977;56:110–156. doi: 10.1016/0012-1606(77)90158-0. [DOI] [PubMed] [Google Scholar]
- 19.Koppen M, Simske JS, Sims PA, Firestein BL, Hall DH, Radice AD, Rongo C, Hardin JD. Cooperative regulation of AJM-1 controls junctional integrity in Caenorhabditis elegans epithelia. Nat Cell Biol. 2001;3:983–991. doi: 10.1038/ncb1101-983. [DOI] [PubMed] [Google Scholar]
- 20.Hedgecock EM, White JG. Polyploid tissues in the nematode Caenorhabditis elegans. Dev Biol. 1985;107:128–133. doi: 10.1016/0012-1606(85)90381-1. [DOI] [PubMed] [Google Scholar]
- 21.Lozano E, Saez AG, Flemming AJ, Cunha A, Leroi AM. Regulation of growth by ploidy in Caenorhabditis elegans. Curr Biol. 2006;16:493–498. doi: 10.1016/j.cub.2006.01.048. [DOI] [PubMed] [Google Scholar]
- 22.Tischler J, Lehner B, Chen N, Fraser AG. Combinatorial RNA interference in Caenorhabditis elegans reveals that redundancy between gene duplicates can be maintained for more than 80 million years of evolution. Genome Biol. 2006;7:R69. doi: 10.1186/gb-2006-7-8-r69. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Shaham S, Horvitz HR. An alternatively spliced C. elegans ced-4 RNA encodes a novel cell death inhibitor. Cell. 1996;86:201–208. doi: 10.1016/s0092-8674(00)80092-6. [DOI] [PubMed] [Google Scholar]
- 24.Shaham S. Identification of multiple Caenorhabditis elegans caspases and their potential roles in proteolytic cascades. J Biol Chem. 1998;273:35109–35117. doi: 10.1074/jbc.273.52.35109. [DOI] [PubMed] [Google Scholar]
- 25.Sugimoto A, Friesen PD, Rothman JH. Baculovirus p35 prevents developmentally programmed cell death and rescues a ced-9 mutant in the nematode Caenorhabditis elegans. Embo J. 1994;13:2023–2028. doi: 10.1002/j.1460-2075.1994.tb06475.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Nagamatsu Y, Ohshima Y. Mechanisms for the control of body size by a G-kinase and a downstream TGFbeta signal pathway in Caenorhabditis elegans. Genes Cells. 2004;9:39–47. doi: 10.1111/j.1356-9597.2004.00700.x. [DOI] [PubMed] [Google Scholar]
- 27.Persengiev SP, Green MR. The role of ATF/CREB family members in cell growth, survival and apoptosis. Apoptosis. 2003;8:225–228. doi: 10.1023/a:1023633704132. [DOI] [PubMed] [Google Scholar]
- 28.Rudolph D, Tafuri A, Gass P, Hammerling GJ, Arnold B, Schutz G. Impaired fetal T cell development and perinatal lethality in mice lacking the cAMP response element binding protein. Proc Natl Acad Sci U S A. 1998;95:4481–4486. doi: 10.1073/pnas.95.8.4481. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Sordella R, Classon M, Hu KQ, Matheson SF, Brouns MR, Fine B, Zhang L, Takami H, Yamada Y, Settleman J. Modulation of CREB activity by the Rho GTPase regulates cell and organism size during mouse embryonic development. Dev Cell. 2002;2:553–565. doi: 10.1016/s1534-5807(02)00162-4. [DOI] [PubMed] [Google Scholar]
- 30.Hardt SE, Tomita H, Katus HA, Sadoshima J. Phosphorylation of eukaryotic translation initiation factor 2Bepsilon by glycogen synthase kinase-3beta regulates beta-adrenergic cardiac myocyte hypertrophy. Circ Res. 2004;94:926–935. doi: 10.1161/01.RES.0000124977.59827.80. [DOI] [PubMed] [Google Scholar]
- 31.Beekman C, Nichane M, De Clercq S, Maetens M, Floss T, Wurst W, Bellefroid E, Marine JC. Evolutionarily conserved role of nucleostemin: controlling proliferation of stem/progenitor cells during early vertebrate development. Mol Cell Biol. 2006;26:9291–9301. doi: 10.1128/MCB.01183-06. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Tsai RY, McKay RD. A nucleolar mechanism controlling cell proliferation in stem cells and cancer cells. Genes Dev. 2002;16:2991–3003. doi: 10.1101/gad.55671. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Ma H, Pederson T. Depletion of the nucleolar protein Nucleostemin causes G1 cell cycle arrest via the p53 pathway. Mol Biol Cell. 2007 doi: 10.1091/mbc.E07-03-0244. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Han C, Zhang X, Xu W, Wang W, Qian H, Chen Y. Cloning of the nucleostemin gene and its function in transforming human embryonic bone marrow mesenchymal stem cells into F6 tumor cells. Int J Mol Med. 2005;16:205–213. [PubMed] [Google Scholar]
- 35.Mouhamad S, Galluzzi L, Zermati Y, Castedo M, Kroemer G. Apaf-1 Deficiency Causes Chromosomal Instability. Cell Cycle. 2007;6:3103–3107. doi: 10.4161/cc.6.24.5046. [DOI] [PubMed] [Google Scholar]
- 36.Zermati Y, Mouhamad S, Stergiou L, Besse B, Galluzzi L, Boehrer S, Pauleau AL, Rosselli F, D'Amelio M, Amendola R, Castedo M, Hengartner M, Soria JC, Cecconi F, Kroemer G. Nonapoptotic role for Apaf-1 in the DNA damage checkpoint. Mol Cell. 2007;28:624–637. doi: 10.1016/j.molcel.2007.09.030. [DOI] [PubMed] [Google Scholar]
- 37.Dasgupta N, Patel AM, Scott BA, Crowder CM. Hypoxic preconditioning requires the apoptosis protein CED-4 in C. elegans. Curr Biol. 2007;17:1954–1959. doi: 10.1016/j.cub.2007.10.017. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Jia B, Crowder CM. Volatile anesthetic preconditioning present in the invertebrate Caenorhabditis elegans. Anesthesiology. 2008;108:426–433. doi: 10.1097/ALN.0b013e318164d013. [DOI] [PubMed] [Google Scholar]
- 39.Sulston JE, Schierenberg E, White JG, Thomson JN. The embryonic cell lineage of the nematode Caenorhabditis elegans. Dev Biol. 1983;100:64–119. doi: 10.1016/0012-1606(83)90201-4. [DOI] [PubMed] [Google Scholar]
- 40.Shankar DB, Cheng JC, Kinjo K, Federman N, Moore TB, Gill A, Rao NP, Landaw EM, Sakamoto KM. The role of CREB as a proto-oncogene in hematopoiesis and in acute myeloid leukemia. Cancer Cell. 2005;7:351–362. doi: 10.1016/j.ccr.2005.02.018. [DOI] [PubMed] [Google Scholar]
- 41.Huang W, Chang HY, Fei T, Wu H, Chen YG. GSK3beta mediates suppression of cyclin D2 expression by tumor suppressor PTEN. Oncogene. 2007;26:2471–2482. doi: 10.1038/sj.onc.1210033. [DOI] [PubMed] [Google Scholar]
- 42.Wang J, Tokarz R, Savage-Dunn C. The expression of TGFbeta signal transducers in the hypodermis regulates body size in C. elegans. Development. 2002;129:4989–4998. doi: 10.1242/dev.129.21.4989. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.