Abstract
Secreted from intestine, human fibroblast growth factor 19 (hFGF19) is an endocrine metabolic regulator that controls bile acid synthesis in the liver. Earlier studies have suggested that hFGF19 at 10–100 nm levels signals through FGF receptor 4 (FGFR4) in the presence of a co-receptor, betaKlotho, but its activity and receptor specificity at physiological concentrations (picomolar levels) remain unclear. Here we report that hFGF19 at picomolar levels require sulfated glycosaminoglycans (sGAGs), such as heparan sulfate, heparin, and chondroitin sulfates, for its signaling via human FGFR4 in the presence of human betaKlotho. Importantly, sGAGs isolated from liver are highly active in enhancing the picomolar hFGF19 signaling. At nanomolar levels, in contrast, hFGF19 activates all types of human FGFRs, i.e. FGFR1c, FGFR2c, FGFR3c, and FGFR4 in the co-presence of betaKlotho and heparin and activates FGFR4 even in the absence of betaKlotho. These results show that sGAGs play crucial roles in specific and sensitive hFGF19 signaling via FGF receptors and suggest that hepatic sGAGs are involved in the highly potent and specific signaling of picomolar hFGF19 through FGFR4 and betaKlotho. The results further suggest that hFGF19 at pathological concentrations may evoke aberrant signaling through various FGF receptors.
Keywords: Glycosaminoglycan, Growth factors, Liver, Receptors, Signal Transduction, FGF, Co-receptor
Introduction
In both humans and mice, the fibroblast growth factor (FGF) family is composed of 22 structurally related proteins that can be divided into several subfamilies (1). The endocrine FGF subfamily, which function as metabolic regulators (2–5), is composed of FGF19, FGF21, and FGF23 in humans and FGF15, FGF21, and FGF23 in mice. Mouse (m)Fgf15 is an ortholog for human (h)FGF19 (6). Both mFGF15 and hFGF19 are expressed in the distal small intestine. Their secretion into the circulation is induced by bile acid, and after reaching the liver via the portal vein they suppress a key liver enzyme involved in bile acid biosynthesis, thereby closing a feedback loop regulating bile acid homeostasis (7). hFGF19/mFGF15 also reportedly contributes to the regulation of blood glucose levels, along with other metabolic regulators, including FGF21 and insulin (8–11).
Endocrine FGFs act via receptor tyrosine kinases but also require the presence of a co-receptor, alphaKlotho (KLA)4 or betaKlotho (KLB) (12). In humans, FGF23 signaling via several FGF receptor (FGFR) subtypes is thought to specifically require KLA, whereas hFGF19 and FGF21 reportedly require KLB (12–14). But other studies have shown that KLA can also act as a co-receptor for hFGF19 (15). It has thus been unclear how the receptor specificity of hFGF19 is achieved, or even what its precise receptor specificity is. Results from Fgf15 knock-out mice suggest that mFGFR4 is the sole functional receptor for mFGF15 (7).
Moreover, the potential applicability of hFGF19 to the treatment of metabolic diseases such as diabetes (8–11) has prompted attempts to develop protocols for its clinical application, but these have been discouraged by the finding that transgenic mice expressing high levels of hFGF19 develop liver tumors (16, 17). The results presented here shed new light on the mechanisms underlying the physiological and pathological activities of hFGF19.
EXPERIMENTAL PROCEDURES
Reagents
Human FGF19 was purchased from R&D Systems. Heparan sulfate (HS), chondroitin sulfate (CS)-A, CS-B, CS-C, CS-D, and CS-E were from Seikagaku Biobusiness Corp. (Tokyo, Japan). Heparin was from Sigma.
Preparation of Hepatic Sulfated Glycosaminoglycans (sGAGs)
Hepatic sGAGs were isolated essentially as described previously (18, 19) with some modification. Briefly, bovine liver was boiled and homogenized, and acetone precipitate was prepared. It was delipidated by chloroform-methanol and subjected to extensive actinase digestion in 50 mm Tris-HCl, pH 8.0, containing 2 mm CaCl2 at 50 °C for 24 h, then the enzyme was denatured by heat. After centrifugation, the cleared supernatant was dialyzed against water and subjected to DEAE-Sepharose Fast Flow column chromatography run by stepwise gradient of NaCl. The sGAG pool was treated with sodium borohydride to remove the GAG chain from its core protein by beta elimination. After neutralization with HCl, perchloric acid was added to obtain a final concentration of 5% (v/v) to allow protein precipitation. Then the cleared supernatant was dialyzed extensively against water, filtered through a 0.22-μm membrane filter, and subjected to biological experiments. Concentration of the sGAGs was determined by a carbazole sulfuric acid method.
Culture of BaF3 Cells and DNA Synthesis Assay
BaF3 cells were obtained from RIKEN BioResource Center. Construction of stable BaF3 cell transfectants and analysis of hFGF19-induced DNA synthesis were performed as described previously (14).
Plasmid Preparation
cDNAs encoding hFGFR1c (NM_015850.3) and hFGFR2c (NM_000141.3) were cloned from human fetal brain RNA (Clontech), hFGFR3c (NM_000142.2) was from human brain RNA (Clontech), hFGFR4 (NM_002011.3) and hKLB (NM_175737.2) were from human liver RNA (Clontech). The nucleotide sequences of these clones were confirmed to be coding the correct amino acid sequences as appear in the database, including one single-nucleotide polymorphism (rs376618).
Western Blotting
Western blotting of FGFRs, KLB, and p-ERK1/2 was performed as described previously (14).
RESULTS
A limiting feature of the various cell and animal systems used to study the mechanisms involved in hFGF19 recognition and signaling is the endogenous expression of one or more components involved in those processes. These components include FGFRs, KLA/KLB, and/or sGAGs. To investigate more precisely the molecular mechanism by which hFGF19 induces cellular signaling, we used a modified BaF3 cell system that was initially established to determine the receptor specificity of canonical FGF signaling. BaF3 cells are a pro-B lymphoma line that does not endogenously express detectable levels of any of the aforementioned components of hFGF19 recognition/signaling, but does harbor the intracellular mediators necessary to transduce a mitogenic signal from FGFRs. We prepared BaF3 transfectants stably expressing one of the subtypes of four human tyrosine kinase FGFR genes (hFGFR1c, hFGFR2c, hFGFR3c, or hFGFR4) with or without hKLB (Fig. 1A). In addition, for hFGFR2c, hFGFR3c, and hFGFR4, the intracellular kinase domains were swapped with that of hFGFR1c, as described previously (20), which enabled more direct comparison of the receptor activation induced by ligand binding to the respective ectodomains. These cells were then examined for their ability to respond to hFGF19 in the presence or absence of sGAGs and hKLB. When evaluating responses evoked by canonical FGFs, inclusion of heparin in the assay system has become a standard procedure (20–23). It is now widely accepted that, for canonical FGFs but not endocrine FGFs, HS, a heparin-like sGAG found on the surfaces of cells, forms a signaling complex with the corresponding FGFR (24, 25). Heparin is a fully sulfated form of HS found in a few mammalian cells/tissues, including mast cells and intestinal mucosa.
FIGURE 1.
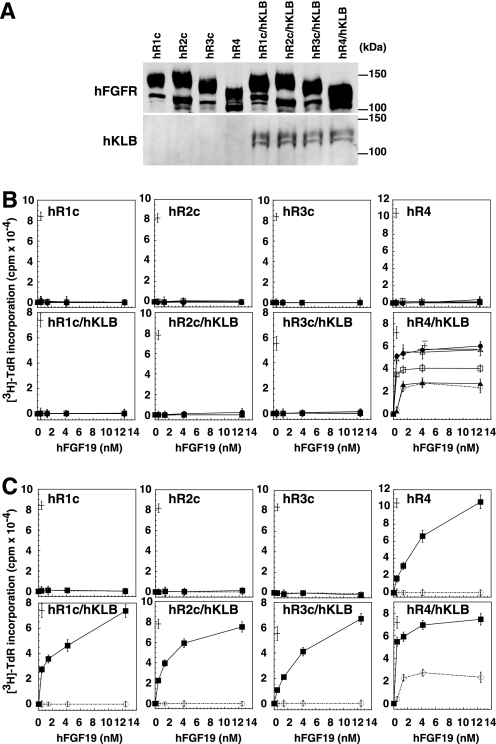
Activation of hFGFRs by hFGF19 at nanomolar concentrations in the presence of sGAGs and hKLB. A, hFGFR and hKLB are expressed in the indicated BaF3 transfectants. The expression levels of hFGFR and hKLB were detected using antibodies against the C terminus of hFGFR1 and hKLB, respectively. B, HS and CS enhance specific activation of hFGFR4 by hFGF19 in the presence of co-expressed hKLB. BaF3 cells, which do not endogenously express FGFRs, KLA/KLB, or sGAGs, were stably transfected with an expression vector encoding the indicated hFGFR subtype (hR1c, hR2c, hR3c, and hR4), without/with an expression vector encoding hKLB. Each FGFR construct used in this figure harbors the intracellular kinase domain of hFGFR1, as described previously (20), and receptor-mediated signaling was measured based on evoked DNA synthesis. The cells were stimulated with hFGF19 at the indicated concentrations in the absence (open circles, dotted line) or presence (5 μg/ml) of HS (filled circles, solid line), CS-B (open triangles, solid line), CS-D (filled triangles, solid line), or CS-E (open squares, solid line). After 42 h of culture, [3H]thymidine was added, and its incorporation into the cellular DNA during the subsequent 6 h was analyzed using a scintillation counter. To confirm the signaling capability of the respective FGFRs, their activation by 460 pm FGF1 was examined in the presence of 5 μg/ml heparin (a cross in each panel); FGF1 is known to activate every FGFR subtype. C, heparin enables activation of all four hFGFR subtypes in the presence of co-expressed hKLB. Receptors were analyzed as in B in the absence (open circles, dotted line) or presence of heparin (5 μg/ml; filled squares, solid line). Note that heparin also enables activation of hFGFR4 in the absence of co-expressed hKLB. In B and C, each symbol represents the mean ± S.D. of triplicate samples. All experiments were performed at least three times and yielded essentially the same results. In addition, at least two independent clones of each transfectant were examined in each combination, yielding essentially the same results.
We initially examined the ability of hFGF19 at nanomolar levels, the concentration range that has been studied in earlier studies, to induce DNA synthesis in the presence of HS or the indicated classes of CS (Fig. 1B). We found that hFGF19 at 1.4, 4, and 12 nm was able to induce DNA synthesis via hFGFR4 only when the receptor was co-expressed with hKLB (Fig. 1B, hR4/hKLB, dotted line). Moreover, although the combination of hFGFR4 and hKLB was sufficient to elicit a partial response to hFGF19, the response was enhanced by HS, CS-B, or CS-E, but not CS-D, and the enhancement elicited by CS-B was comparable with that of HS (Fig. 1B, hR4/hKLB, solid lines). The three other receptors tested, hFGFR1c, hFGFR2c, and hFGFR3c, were not activated by hFGF19, even when co-expressed with hKLB. Thus, HS and CS enhance specific hFGF19 signaling via the combination of hFGFR4 and hKLB.
In contrast to HS and CS, heparin enabled hFGF19 (at 0.46 nm and higher) to signal through hFGFR1c, hFGFR2c, and hFGFR3c when co-expressed with hKLB (Fig. 1C). In addition, heparin also enabled hFGF19 (at 0.46 nm and higher) to signal via hFGFR4 in the absence of hKLB (Fig. 1C, hR4). Such KLB-independent signaling was not observed in the presence of HS or CS (Fig. 1B).
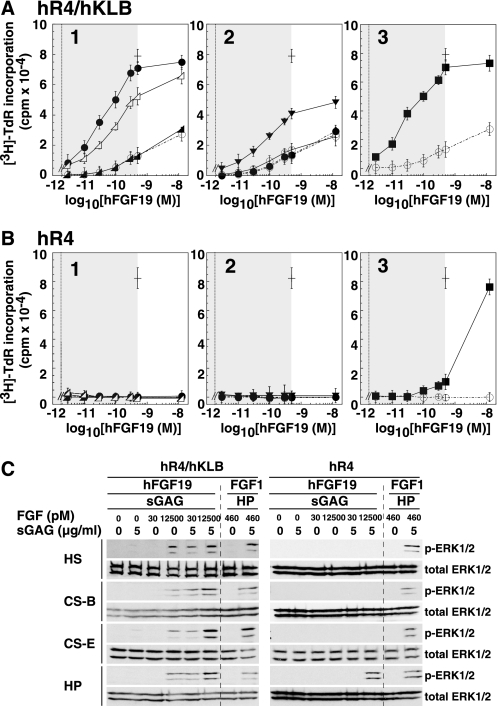
Fig. 1 summarizes the effects of hFGF19 at nanomolar concentrations. Our findings are consistent with those of earlier studies, which examined the activity of hFGF19 at concentrations ranging from 10 to 100 nm (15, 26, 27). However, hFGF19 concentrations in the human circulation are reportedly in the subnanomolar range, most likely around 30 pm, which is 3–4 orders of magnitude lower than the concentrations tested previously (28–30). We therefore also examined the receptor specificity of hFGF19 at more physiological concentrations (Fig. 2). We found that activation of hFGFR4 by hFGF19 at concentrations ranging from 3 to 500 pm was strongly dependent on the presence of sGAGs and on the co-expression of hKLB (Fig. 2, A and B). In the presence of HS or heparin, 500 pm hFGF19 alone did not activate hFGFR4, but the receptor was maximally activated when co-expressed with hKLB (Fig. 2A). With this system, evoked DNA synthesis could be detected at hFGF19 concentrations as low as 3 pm in the presence of heparin, HS, CS-B, or CS-E; in contrast, CS-A, CS-C, and CS-D were not sufficient to mediate the effect (Fig. 2A). These findings were confirmed by assessing the activation (phosphorylation) of ERK1/2 MAP kinase (p-ERK, Fig. 2C). sGAG-dependent activation of ERK by 30 pm hFGF19 in cells co-expressing hFGFR4 and hKLB was clearly observed in the presence of HS, CS-B, CS-E, or heparin (Fig. 2C), and the response-enhancing effects of the sGAGs were detected at their concentrations of 0.3 μg/ml and higher (Fig. 3).
FIGURE 2.
Sulfated GAGs enable potent and specific hFGF19 signaling via hFGFR4 co-expressed with hKLB. Signaling was evaluated based on evoked DNA synthesis. A and B, stable hFGFR4/hKLB/BaF3 transfectants (A) or stable hFGFR4/BaF3 transfectants (B) were stimulated with hFGF19 as in Fig. 1 at the indicated concentrations in the absence (open circles, dotted line) or presence (5 μg/ml) of various GAGs: 1, HS (filled circles, solid line), CS-A (filled triangles, solid line), CS-B (open triangles, solid line); 2, CS-C (filled circles, dotted line), CS-D (open circles, solid line), CS-E (inverted filled triangles, solid line); 3, heparin (filled squares, solid line). As a positive control, activation of each receptor by 460 pm FGF1 (cross) was examined in the presence of 5 μg/ml heparin. The reported physiological concentration range of hFGF19 (less than 500 pm) is shaded in each panel. Note that in B1 and B2, none of the sGAGs enabled hFGF19 signaling. All experiments were performed at least three times and yielded essentially the same results. C, BaF3 transfectants like those in A and B were treated for 10 min with the indicated concentrations of hFGF19 or FGF1 in the absence and presence of various sGAGs, after which signaling was evaluated based on phosphorylation of ERK1/2 detected by Western blotting.
FIGURE 3.
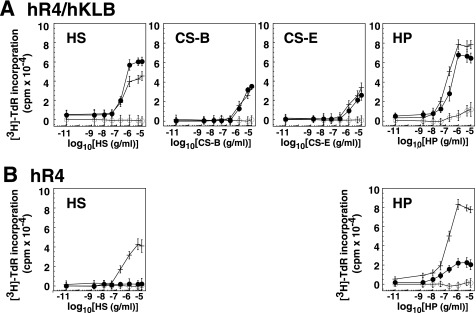
Sulfated GAG dose response for hFGF19-induced signaling via hFGFR4/hKLB. Stable hFGFR4/hKLB/BaF3 transfectants (A) or stable hFGFR4/BaF3 transfectants (B) were stimulated with the absence (open circles) or presence of FGF (500 pm hFGF19 (filled circles) or 460 pm FGF1 (crosses)) together with the indicated concentrations of HS, CS-B, CS-E, or heparin (HP). Signaling was evaluated based on evoked DNA synthesis. All experiments were performed at least three times and yielded essentially the same results.
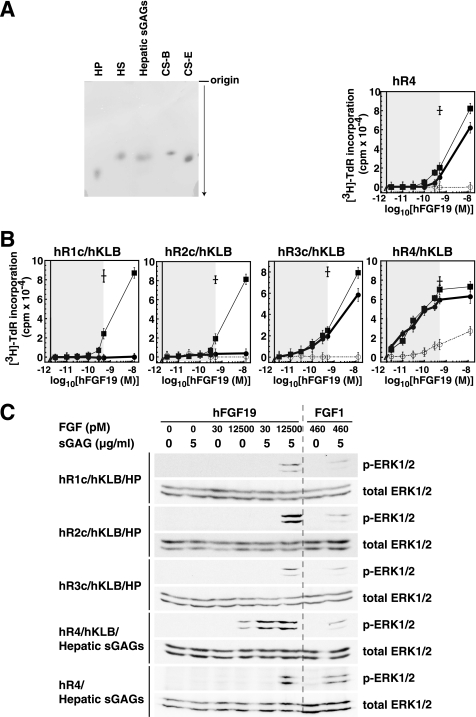
Because liver is the target organ of hFGF19, we then examined whether the sGAGs of the liver had potential to assist hFGF19 signaling. Thus, we prepared sGAGs from liver, i.e. hepatic sGAGs, and compared the effects of hepatic sGAGs and heparin on the signaling of picomolar hFGF19 through FGFRs (Fig. 4). In cellulose acetate electrophoresis a preparation of the hepatic sGAGs migrated close to the authentic HS, CS-B, and CS-E, but clearly differently from heparin (Fig. 4A). The disaccharide compositions of the sGAGs used in this study are shown in Table 1. Importantly, this hepatic sGAGs preparation and heparin were equally potent enhancers of picomolar hFGF19 signaling toward hFGFR4 when it is co-expressed with hKLB (Fig. 4B). The hepatic sGAGs also enhanced signaling through hFGFR3c/hKLB weakly, similar to heparin. However, unlike heparin, the hepatic sGAGs did not enhance signaling through hFGFR1c/hKLB or hFGFR2c/hKLB (Fig. 4B). These results were confirmed by the activation of ERK1/2 MAP kinase (p-ERK, Fig. 4C).
FIGURE 4.
Hepatic sGAGs enable potent and specific hFGF19 signaling via hFGFR4 co-expressed with hKLB. A, preparation of hepatic sGAGs used in this study is shown. The preparation of hepatic sGAGs was resolved by cellulose acetate electrophoresis and visualized by Alcian blue staining. Authentic samples of heparin (HP), HS, CS-B, and CS-E as used in this study were run on the same membrane. Repeated experiments yielded identical results. B, hepatic sGAGs and heparin enhance specific activation of hFGFR4 by hFGF19 in the presence of co-expressed hKLB. Signaling was evaluated by BaF3 transfectants as in Fig. 1 at the indicated concentrations of hFGF19 in the absence (open circles, dotted line) or presence (5 μg/ml) of hepatic sGAGs (filled circles, solid bold line) or heparin (filled squares, solid thin line). As a positive control, activation of each receptor by 460 pm FGF1 (cross) was examined in the presence of 5 μg/ml heparin. The reported physiological concentration range of hFGF19 (less than 500 pm) is shaded. All experiments were performed at least three times and yielded essentially the same results. C, BaF3 transfectants like those in B were treated for 10 min with the indicated concentrations of hFGF19 or FGF1 in the presence of heparin (HP) or hepatic sGAGs, after which signaling was evaluated based on phosphorylation of ERK1/2 detected by Western blotting. Note that the results of hR4/hKLB/HP are shown in Fig. 2B.
TABLE 1.
Disaccharide composition of the sulfated glycosaminoglycans used in this study
| Heparan sulfate/heparina | |||||||
|---|---|---|---|---|---|---|---|
| Disaccharide | 0S | NS | 6S | US | (6,N)S | (U,N)S | (U,6,N)S |
| Hepatic sGAG (HS) | 41.4 | 14.9 | 12.3 | —b | 8.0 | 5.5 | 17.9 |
| HS | 51.7 | 16.9 | 11.5 | — | 8.5 | 5.7 | 5.6 |
| Heparin | 6.4 | 5.5 | 3.5 | — | 29.8 | 7.3 | 47.5 |
| Chondroitin sulfatec | |||||||
|---|---|---|---|---|---|---|---|
| Disaccharide | 0S | 6S | 4S | (U,6)S | (U,4)S | (4,6)S | (U,4,6)S |
| CS-A | 1.6 | 19.3 | 76.1 | 2.7 | — | 0.3 | — |
| CS-B | 0.7 | 1.9 | 90.3 | 0.6 | 6.5 | — | — |
| CS-C | 1.7 | 73.2 | 15.5 | 9.3 | — | 0.3 | — |
| CS-D | 0.6 | 43.9 | 26.9 | 21.3 | — | 7.0 | 0.3 |
| CS-E | 5.9 | 9.6 | 22.9 | — | — | 61.6 | — |
a Disaccharide unit compositions of the HS fraction of the hepatic sGAGs, HS, and heparin used in this study were analyzed, and their molar ratios (%) are shown.
b —, Not detected.
c These data were provided by Seikagaku Biobusiness Corp. as typical compositions of their products used in this study.
DISCUSSION
hFGF19 (and mFGF15) is expressed in the intestine in response to bile acid and then secreted into the blood. After reaching the liver via the portal vein, it suppresses transcription of Cyp7A1, a key enzyme involved in bile acid biosynthesis. This endocrine mode of action would require hFGF19 to possess a low affinity for extracellular sGAGs to avoid being trapped within blood vessel lumens and would require hepatocytes to possess a highly sensitive and specific ability to recognize hFGF19. Consistent with those ideas, hFGF19 shows very low affinity to heparin (31, 32), and the requirement for hKLB as a co-receptor and the strong positive regulation of hFGF19/hFGFR4 signaling by HS/CS establishes a highly specific and sensitive recognition system. Several earlier studies have shown that the hFGF19 concentration in the circulation of healthy volunteers is around 30 pm (28–30). We found that at these low levels, hFGF19 signaling is only induced in the presence of HS/CS/heparin/hepatic sGAGs (Figs. 2–4). Thus HS, CS-B, and CS-E, together with hKLB, are crucial for hFGF19/hFGFR4 signaling at physiological levels of hFGF19. Although the hepatic sGAGs are likely to be composed of HS and CSs, their detailed structures responsible for this activity, such as sulfate modification patterns, await future studies. In addition, the structure and composition of hepatic sGAGs may vary by species, and the relevant human or mouse sGAGs might be different. The enhancement of hFGF19 activity by HS/CS also implies that formation of a complex composed of sGAGs, hFGFR4, and hKLB is required for optimal signaling in hepatocytes. Indeed, both KLB and FGFRs exhibit affinity for heparin (data not shown). We suggest that this mechanism underlies normal hFGF19 activity in situ.
In contrast to the picomolar range, the results obtained with nanomolar hFGF19 in the presence of heparin indicate these conditions severely reduce the target specificity of hFGF19. At concentrations higher than 1 nm, hFGF19 signaling via hFGFR4 was observed in the absence of sGAGs and, in the presence of heparin or hepatic sGAGs, hFGFR4 mediated hFGF19 signaling in the absence of hKLB (Figs. 1, 2B, and 4). The presence of heparin also enabled hFGF19 to stimulate all of the hFGFR subtypes in the presence of hKLB (Fig. 1). These results likely reflect pathological hFGF19 activity or perhaps normal physiological activity found in only a few selected tissues. Although HS and CSs are the dominant sGAGs found on cell surfaces and within the extracellular matrix, some tissues and cells do harbor heparin. For instance, extracts of mammalian lung and intestine are sources of commercially available heparin. In addition, there may be tissues other than liver that harbor HS/CSs with biological activity similar to that of hepatic sGAGs we showed in this paper. In that context, it is intriguing that hFGF19 expression is elevated in the cancerous tissues of some patients with lung or colon cancer (17), and some cancers also show up-regulation of FGFR4 (17). Whether hFGF19 signaling via hFGFR4 in the presence of endogenous heparin/sGAGs contributes to the pathology of these diseases remains unknown, but our findings shed new light on the mechanisms underlying the physiological and pathological activities of hFGF19.
Acknowledgments
We thank Profs. Yukio Ishimi and Zheng Yu Wang for tutorial assistance; Dr. Yukishige Ito for kindly providing access to his laboratory facilities in RIKEN; Dr. Yasushi Kamisaka, Miho Kimura-Ueki, Akiko Komi-Kuramochi, and Junko Oki at National Institute of Advanced Industrial Science and Technology and Yuniko Shibata and Takeshi Ishimaru at Seikagaku Biobusiness Corp. for providing technical help and information.
This work was supported in part by the National Institute of Advanced Industrial Science and Technology and by a grant-in-aid from the Ministry of Education, Culture, Sports, Science, and Technology, Japan.
- KLA
- alphaKlotho
- KLB
- betaKlotho
- CS
- chondroitin sulfate
- GAG
- glycosaminoglycan
- HS
- heparan sulfate
- R
- receptor
- sGAG
- sulfated GAG.
REFERENCES
- 1. Itoh N., Ornitz D. M. (2004) Trends Genet. 20, 563–569 [DOI] [PubMed] [Google Scholar]
- 2. Jones S. (2008) Mol. Pharm. 5, 42–48 [DOI] [PubMed] [Google Scholar]
- 3. Fukumoto S. (2008) Endocr. J. 55, 23–31 [DOI] [PubMed] [Google Scholar]
- 4. Kuro-o M. (2008) Trends Endocrinol. Metab. 19, 239–245 [DOI] [PubMed] [Google Scholar]
- 5. Wu X., Li Y. (2009) Aging 1, 1023–1027 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6. Wright T. J., Ladher R., McWhirter J., Murre C., Schoenwolf G. C., Mansour S. L. (2004) Dev. Biol. 269, 264–275 [DOI] [PubMed] [Google Scholar]
- 7. Inagaki T., Choi M., Moschetta A., Peng L., Cummins C. L., McDonald J. G., Luo G., Jones S. A., Goodwin B., Richardson J. A., Gerard R. D., Repa J. J., Mangelsdorf D. J., Kliewer S. A. (2005) Cell Metab. 2, 217–225 [DOI] [PubMed] [Google Scholar]
- 8. Tomlinson E., Fu L., John L., Hultgren B., Huang X., Renz M., Stephan J. P., Tsai S. P., Powell-Braxton L., French D., Stewart T. A. (2002) Endocrinology 143, 1741–1747 [DOI] [PubMed] [Google Scholar]
- 9. Fu L., John L. M., Adams S. H., Yu X. X., Tomlinson E., Renz M., Williams P. M., Soriano R., Corpuz R., Moffat B., Vandlen R., Simmons L., Foster J., Stephan J. P., Tsai S. P., Stewart T. A. (2004) Endocrinology 145, 2594–2603 [DOI] [PubMed] [Google Scholar]
- 10. Strack A. M., Myers R. W. (2004) Endocrinology 145, 2591–2593 [DOI] [PubMed] [Google Scholar]
- 11. Kharitonenkov A. (2009) Curr. Opin. Pharmacol. 9, 805–810 [DOI] [PubMed] [Google Scholar]
- 12. Tomiyama K., Maeda R., Urakawa I., Yamazaki Y., Tanaka T., Ito S., Nabeshima Y., Tomita T., Odori S., Hosoda K., Nakao K., Imura A., Nabeshima Y. (2010) Proc. Natl. Acad. Sci. U.S.A. 107, 1666–1671 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13. Lin B. C., Wang M., Blackmore C., Desnoyers L. R. (2007) J. Biol. Chem. 282, 27277–27284 [DOI] [PubMed] [Google Scholar]
- 14. Suzuki M., Uehara Y., Motomura-Matsuzaka K., Oki J., Koyama Y., Kimura M., Asada M., Komi-Kuramochi A., Oka S., Imamura T. (2008) Mol. Endocrinol. 22, 1006–1014 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15. Wu X., Lemon B., Li X., Gupte J., Weiszmann J., Stevens J., Hawkins N., Shen W., Lindberg R., Chen JL., Tian H., Li Y. (2008) J. Biol. Chem. 283, 33304–33309 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16. Nicholes K., Guillet S., Tomlinson E., Hillan K., Wright B., Frantz G. D., Pham T. A., Dillard-Telm L., Tsai S. P., Stephan J. P., Stinson J., Stewart T., French D. M. (2002) Am. J. Pathol. 160, 2295–2307 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17. Desnoyers L. R., Pai R., Ferrando R. E., Hötzel K., Le T., Ross J., Carano R., D'Souza A., Qing J., Mohtashemi I., Ashkenazi A., French D. M. (2008) Oncogene 27, 85–97 [DOI] [PubMed] [Google Scholar]
- 18. Vongchan P., Warda M., Toyoda H., Toida T., Marks R. M., Linhardt R. J. (2005) Biochim. Biophys. Acta 1721, 1–8 [DOI] [PubMed] [Google Scholar]
- 19. Kimura A., Toyoki Y., Hakamada K., Yoshihara S., Sasaki M. (2008) J. Hepatobiliary Pancreat. Surg. 15, 608–614 [DOI] [PubMed] [Google Scholar]
- 20. Ornitz D. M., Xu J., Colvin J. S., McEwen D. G., MacArthur C. A., Coulier F., Gao G., Goldfarb M. (1996) J. Biol. Chem. 271, 15292–15297 [DOI] [PubMed] [Google Scholar]
- 21. Yoneda A., Asada M., Oda Y., Suzuki M., Imamura T. (2000) Nat. Biotechnol. 18, 641–644 [DOI] [PubMed] [Google Scholar]
- 22. Allen B. L., Rapraeger A. C. (2003) J. Cell Biol. 163, 637–648 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23. Smith S. M., West L. A., Govindraj P., Zhang X., Ornitz D. M., Hassell J. R. (2007) Matrix Biol. 26, 175–184 [DOI] [PubMed] [Google Scholar]
- 24. Pellegrini L., Burke D. F., von Delft F., Mulloy B., Blundell T. L. (2000) Nature 407, 1029–1034 [DOI] [PubMed] [Google Scholar]
- 25. Kan M., Wu X., Wang F., McKeehan W. L. (1999) J. Biol. Chem. 274, 15947–15952 [DOI] [PubMed] [Google Scholar]
- 26. Wu X., Ge H., Lemon B., Weiszmann J., Gupte J., Hawkins N., Li X., Tang J., Lindberg R., Li Y. (2009) Proc. Natl. Acad. Sci. U.S.A. 106, 14379–14384 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27. Wu X., Ge H., Lemon B., Vonderfecht S., Baribault H., Weiszmann J., Gupte J., Gardner J., Lindberg R., Wang Z., Li Y. (2010) Proc. Natl. Acad. Sci. U.S.A. 107, 14158–14163 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28. Lundåsen T., Gälman C., Angelin B., Rudling M. (2006) J. Intern. Med. 260, 530–536 [DOI] [PubMed] [Google Scholar]
- 29. Dostálová I., Kaválková P., Haluzíková D., Lacinová Z., Mráz M., Papezová H., Haluzík M. (2008) J. Clin. Endocrinol. Metab. 93, 3627–3632 [DOI] [PubMed] [Google Scholar]
- 30. Schaap F. G., van der Gaag N. A., Gouma D. J., Jansen P. L. (2009) Hepatology 49, 1228–1235 [DOI] [PubMed] [Google Scholar]
- 31. Goetz R., Beenken A., Ibrahimi O. A., Kalinina J., Olsen S. K., Eliseenkova A. V., Xu C., Neubert T. A., Zhang F., Linhardt R. J., Yu X., White K. E., Inagaki T., Kliewer S. A., Yamamoto M., Kurosu H., Ogawa Y., Kuro-o M., Lanske B., Razzaque M. S., Mohammadi M. (2007) Mol. Cell. Biol. 27, 3417–3428 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32. Asada M., Shinomiya M., Suzuki M., Honda E., Sugimoto R., Ikekita M., Imamura T. (2009) Biochim. Biophys. Acta 1790, 40–48 [DOI] [PubMed] [Google Scholar]