Abstract
The copper-transporting ATPase ATP7A has an essential role in human physiology. ATP7A transfers the copper cofactor to metalloenzymes within the secretory pathway; inactivation of ATP7A results in an untreatable neurodegenerative disorder, Menkes disease. Presently, the mechanism of ATP7A-mediated copper release into the secretory pathway is not understood. We demonstrate that the characteristic His/Met-rich segment Met672–Pro707 (HM-loop) that connects the first two transmembrane segments of ATP7A is important for copper release. Mutations within this loop do not prevent the ability of ATP7A to form a phosphorylated intermediate during ATP hydrolysis but inhibit subsequent dephosphorylation, a step associated with copper release. The HM-loop inserted into a scaffold protein forms two structurally distinct binding sites and coordinates copper in a mixed His-Met environment with an ∼2:1 stoichiometry. Binding of either copper or silver, a Cu(I) analog, induces structural changes in the loop. Mutations of 4 Met residues to Ile or two His-His pairs to Ala-Gly decrease affinity for copper. Altogether, the data suggest a two-step process, where copper released from the transport sites binds to the first His(Met)2 site, triggering a structural change and binding to a second 2-coordinate His-His or His-Met site. We also show that copper binding within the HM-loop stabilizes Cu(I) and protects it from oxidation, which may further aid the transfer of copper from ATP7A to acceptor proteins. The mechanism of copper entry into the secretory pathway is discussed.
Keywords: ATPases, Copper, Membrane Proteins, Metalloproteins, Transport Metals, EXAFS, Secretory Pathway
Introduction
Copper is required for normal cell homeostasis and serves as an essential cofactor for a variety of metalloenzymes involved in neurotransmitter biosynthesis, amidation of neuroendocrine peptides, vasculature and connective tissue formation, and many other processes. Copper exists in two oxidation states: reduced and unstable Cu(I) and a more stable oxidized Cu(II). The secreted and plasma membrane-bound enzymes acquire their copper cofactor within the secretory pathway (trans-Golgi network and vesicles) in a process that requires the activity of copper-transporting ATPases, ATP7A and ATP7B (1).
ATP7A is ubiquitously expressed in human tissues and is thought to play a housekeeping role in cellular copper metabolism (2), whereas the physiological function of ATP7B is likely to be tissue-specific (3). Genetic defects in ATP7A are associated with a lethal disorder, Menkes disease, caused by insufficient copper delivery into the brain and other tissues and the malfunction of numerous copper-dependent enzymes. Mutations in ATP7B result in Wilson disease, a severe hepato-neurologic disorder. The genetic lesions in both disorders are firmly established, and significant progress has been made in dissecting the structure and function of the transporters. However, the mechanisms of copper release from Cu-ATPases and its transfer to cuproenzymes in the secretory pathways remain largely unknown.
The transport of copper by Cu-ATPases requires the hydrolysis of ATP and involves the formation of a transient phosphorylated intermediate (catalytic phosphorylation, see Fig. 1A). This step depends on Cu(I) binding to the intramembrane transport sites (4, 5). The release of copper from the intramembrane sites into the lumen of the trans-Golgi or vesicles triggers protein dephosphorylation (2, 4, 5). ATP7A and ATP7B, which are highly homologous, share this general mechanism. In in vitro assays, these two Cu-ATPases work interchangeably, as evidenced by their ability to deliver copper to the same enzymes when overexpressed in either human fibroblasts or yeast cells (6, 7). However, in the organism, the functions of the two Cu-ATPases are not equivalent. Severe disorders due to ATP7A or ATP7B inactivation indicate that these Cu-ATPases do not fully complement each other's activity. This is in part due to different levels of expression of ATP7A and ATP7B in tissues and their distinct trafficking behavior in polarized epithelia (8). In addition, kinetic studies of catalytic phosphorylation and dephosphorylation show that ATP7A performs each of these steps faster than ATP7B (2). These functional differences may become critical in Menkes and Wilson diseases, when the function of one of the Cu-ATPases is lost and cells have to rely solely on activity of the remaining Cu-ATPase.
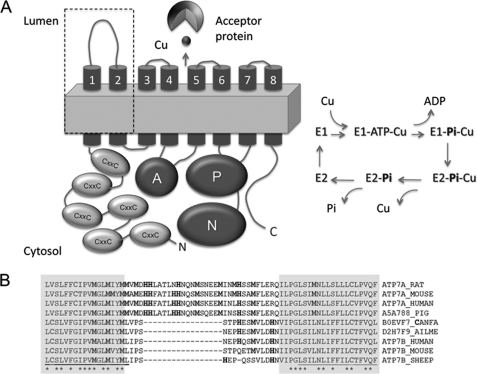
FIGURE 1.
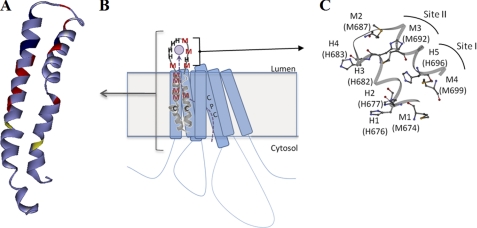
ATP7A contains a characteristic sequence insert in the first luminal loop. A, graphic structure of ATP7A depicting functional domains (left) and the schematic of catalytic cycle (right). The N terminus of ATP7A contains six metal-binding domains with CXXC copper-binding sites. ATP binds to the ATP-binding domain consisting of the N- and P-domains, and during hydrolysis, the γ-phosphate of ATP is transferred to the P-domain forming transient phosphorylated intermediate. This reaction requires copper binding within the TM domain. Copper release at the luminal side is accompanied by dephosphorylation. The released copper binds to acceptor proteins (cuproenzymes). The dashed square indicates the TM1,2 hairpin with the insert, HM-loop. B, sequence alignment of the TM1,2 hairpin of ATP7A and ATP7B. The shaded sequence indicates residues present in the TM; Met and His residues are in bold.
To better understand the mechanism of copper entry to the secretory pathway and the molecular determinants that contribute to a faster copper release by ATP7A, we searched for structural differences between ATP7A and ATP7B. Human Cu-ATPases are large membrane proteins, consisting of an N-terminal domain with six Cu(I)-binding sites (CXXC; MBS1–6), a nucleotide-binding domain, an actuator domain, a phosphorylation domain, eight transmembrane (TM)4 domains, and a C-terminal tail (see Fig. 1A). We hypothesized that the cytosolic domains of Cu-ATPases regulate copper delivery to the membrane from the cytosol, whereas subsequent copper release from the intramembrane sites may be controlled by structural elements at the lumenal side of the transporters. Our results suggest that a sequence insert in the lumenal portion of ATP7A, which is absent in ATP7B, binds copper and influences the rate of copper release from ATP7A. In addition, this region appears to play a unique role in stabilizing reduced copper prior to its further transfer to cuproenzymes. These results provide the first information about events that facilitate copper entry into the secretory pathway.
EXPERIMENTAL PROCEDURES
Generation and Characterization of the Full-length ATP7A with Mutations in the HM-loop
Mutations were introduced into the full-length ATP7A by site-directed mutagenesis using the previously generated pFBD-MNK(BamHI) plasmid (9) as a template and primers Mut1fwd, Mut1rv, Mut2fwd, and Mut2rv (supplemental Table 1): ATP7A-Mut1 has 6 residues (in bold) mutated to either Ala or Gly: 672M(G)VM(A)DH(A)H(G)FATLH(A)H(G)NQNMSKEEM692, whereas ATP7A-Mut2 has 2 Met residues mutated, 672MVMDHHFATLHHNQNM(A)SKEEM(A)692. Attempts to replace all Met/His were unsuccessful. A list of all constructs in this study can be found in supplemental Table 2. Baculoviruses expressing ATP7A-Mut1 and ATP7A-Mut2 were produced as described previously (2). For protein expression, 2 × 106 Sf9 insect cells growing in 50–100 ml of Sf9 cell medium containing 1% ethanol and 1% FBS were infected with recombinant baculovirus. Cells were collected after 3 days by centrifugation, and total membrane protein was isolated as in Ref. 10, and the protein concentration was determined by Lowry et al. (11).
Phosphorylation and Dephosphorylation of ATP7A Variants
Phosphorylation and dephosphorylation of ATP7A variants were performed as in Ref. 2. Briefly, microsomal membranes (50 μg of protein) in 100 μl of cold reaction buffer (20 mm Bis-Tris-propane-HCl, pH 6.0, 200 mm KCl, and 5 mm MgCl2) were incubated with 1 μm [γ-32P]ATP (12.5 mCi/μmol in each reaction) at room temperature for 0, 15, 30, 60, and 120 s and immediately precipitated with 50 μl of ice-cold 1 mm NaH2PO4 in 50% TCA to measure phosphorylation. To examine the effect of copper chelation on protein phosphorylation, one aliquot was treated with 200 μm bathocuproine disulphonate (BCS) for 30 min before [γ-32P]ATP addition. For dephosphorylation, after 2 min of phosphorylation, cold ATP was added to a final concentration of 25 μm (to facilitate dephosphorylation), and aliquots were removed at 0, 10, 20, and 30 s after ATP addition and immediately precipitated with 50 μl of ice-cold 1 mm NaH2PO4 in 50% TCA to measure phosphorylation. Precipitates were collected by centrifugation, washed with H2O, and resuspended in SDS-PAGE acidic gel sample buffer (2). The disappearance of the phosphorylated band was monitored following acidic SDS-PAGE on a 7.5% acrylamide gel and exposure to x-ray film overnight at −80 °C. Intensities were quantified by densitometry using the Spot Densitometry Analysis software (AlphaView, Alpha Innotech), with the level of phosphorylation after 2 min set as 100%.
Constructs for the Analysis of Metal Binding
Initially, a construct containing the HM-loop alone in the intein-containing vector, pTXB1 (New England Biolabs), was generated. The sequence 671MMVMDHHFATLHHNQNMSKEEMINLHSSMFLERQILP707 was amplified from the ATP7A-encoding pFBD-MNK(BamHI) plasmid (a kind gift from Michael Petris, University of Missouri) using primers HM-fw and HM-rv (supplemental Table 1) that contained restriction sites NdeI at the 5′ end and SapI at the 3′ end. After amplification, insert and plasmid were digested with corresponding restriction enzymes and then ligated overnight at 16 °C.
Metal-binding domain 2 (MBS2) with the HM-loop (MBS2-HM) was designed by replacing the native copper-binding loop MTCQSC with the sequence 674MDHHFATLHHNQNMSKEEMINLHSSM699 (“breast-tumor” and “breast-cancer” 1). To do this, the MBS2 region of human ATP7B (residues 141–212) was amplified with T7-rv and intein-fw primers (supplemental Table 1) from a plasmid containing the corresponding sequence in pTYB12 (New England Biolabs) vector (12). The PCR product was digested with BslI and BsaWI restriction enzymes to cut out the cDNA region encoding the MTCQSC sequence. In parallel, the HM-loop region was amplified from the pFBD-MNK plasmid using the primers MBSHM-fw, MBSHM-rv1, MBSHM-rv2, and MBSHM-rv3 (supplemental Table 1). This procedure added the BslI restriction site and a flanking sequence of MBS2 at the N terminus and inserted a Gly-Ser-Ala and flanking sequence containing the BsaWI restriction site at the C terminus of the HM insert. After digestion with BslI and BsaWI, the insert was ligated into cut MBS2 at 16 °C overnight. MBS2 containing the HM sequence was digested with NdeI and SalI and ligated into cut pTYB12.
ScoHM Chimera
ScoHM chimera was constructed by replacing the 45CETIC49 copper-binding motif of Bacillus subtilis Sco (Sco) protein (13) with the sequence 674MDHHFATLHHNQNMSKEEMINLHSSM699 (supplemental Fig. 1). Native His and Met residues in Sco were mutated to Ala and Ile, respectively, to generate the M52I/M56I/H55A/H135A variant of Sco. This construct was used to replace the native 45CETIC49 loop with the HM-loop using overlap extension PCR with T7 forward primer, HMSco-rv (supplemental Table 1), MXE intein reverse primer (New England Biolabs), and HMSco-fw (supplemental Table 1). The PCR product was digested with NcoI and BseRI and ligated into the NcoI-BseRI-digested Sco (13), generating the M52I/M56I-ScoHM. The final ScoHM construct was generated by introducing the H55A/H135A substitution into the M52I/M56I-ScoHM using overlap PCR and the T7 fwd, M52I/M56I/H55A/H135A-rv, M52I/M56I/H55A/H135A-fw, and MXE intein primers (supplemental Table 1). The M52I/M56I/H55A/H135A-ScoHM protein was termed ScoHM. The control M52I/M56I/H55A/H135A-Sco without the HM-loop was also made using the same primers.
ScoHM Mutants
ScoHM mutants ScoHis MDH(A)H(G)FATLH(A)-H(G)NQNMSKEEMINLHSSM, ScoM12 Mdhhfa-TLHHNQNMSKEEM(I)INLHSSM(I), ScoM0 M(I)DHHFATLHHNQNM(I)SKEEM(I)INLHSSM(I), and ScoM1 MDHHFATLHHNQNM(I)SKEEM(I)INLHSSM(I) were constructed by PCR amplification of the ScoHM construct with primers containing desired mutations and intein-rv as the reverse primer (supplemental Table 1). After digestion, inserts were ligated into pTXB3 plasmid at 16 °C overnight. To introduce a Trp residue into ScoHM, the mutagenesis primer WSco-fw was used to replace the only Phe residue in the HM-loop with Trp. The sequence was extended to the NcoI restriction site with primers ScoHMmut-fw1, ScoHMmut-fw2, and ScoHMmut-fw3 and intein-rv. PCR products were digested with NcoI and SpeI and ligated into cut pTXB3. Plasmids were transformed into Escherichia coli BL21 for expression.
Expression and Purification of Recombinant Proteins
Protein expression was induced with 0.5 mm isopropyl β-d-1-thiogalactopyranoside, added at A600 = 0.6–0.8 of E. coli cell density at 16 °C overnight. Cells were then pelleted by centrifugation, resuspended in 50 mm sodium phosphate and 500 mm NaCl buffer, pH 7.2 (15 ml/liter cell culture), containing Complete-EDTA protease inhibitor (Roche Applied Science). Proteins were purified on chitin resin with subsequent intein cleavage/elution with 50 mm DTT for 48 h (for pTYB12-derived proteins) or 50 mm 2-mercapto-ethane sulfonate sodium for 24 h for pTXB3-derived proteins, in 50 mm sodium phosphate and 500 mm NaCl, pH 7.2. Protein purity was assessed by 10% SDS-PAGE, typically ∼90%, and protein concentration was determined by Bradford assay and/or by absorption at 280 nm using an extinction coefficient, calculated for each protein sequence at the ExPASy Proteomics Server. Circular dichroism analyses were performed at 0.5 mg/ml protein concentration on an AVIV 215 spectrometer at 25 °C in 50 mm sodium phosphate buffer, pH 7.2 using a 1.0-mm path length cuvette and averaged from three independent scans.
Copper Binding
Excess CuCl2 (typically 3–5 copper atoms per protein) was added aerobically after reduction with glutathione (1 copper:10 glutathiones) or ascorbate (1 copper:2 or 4 ascorbates) in water to purified MBS2-HM, ScoHM, and ScoHM mutants, in 50 mm sodium phosphate buffer, pH 7.2, at a protein concentration of 5–25 μm. Excess copper was removed by dialysis in 50 mm sodium phosphate buffer, pH 7.2. Anaerobically, Cu(I) was added as a 3-fold excess of (Cu(CH3CN)4)+ followed by dialysis against sodium phosphate buffer, pH 7.2 (14). For copper retention experiments, proteins were present at 15 μm in 50 mm sodium phosphate buffer, pH 7.2, in a 500-μl volume in a 1-ml quartz cuvette. Copper was added at 2:1 or 3:1 per protein with ascorbate present at 2:1 or 4:1 per copper. After copper addition, 1-μl aliquots of 5 mm BCS were titrated into the solution, and the formation of Cu(I)(BCS)2 complex was monitored at 483 nm. The amount of complex was calculated using ϵ of 13,500 m−1·cm−1 and then subtracted from total copper in solution to calculate uncomplexed copper.
For Cu(I) stabilization studies, 200 μm BCS was added to a CuCl2 solution in 50 mm sodium phosphate buffer, pH 7.2, containing ascorbate (2-fold to copper) with or without 15 μm ScoHM protein at 0, 15, 30, and 60 min after mixing. CuCl2 solutions were incubated at room temperature for 30 min before BCS addition. The amount of Cu(I)-BCS complex was calculated as above. To monitor metal binding using intrinsic Trp fluorescence, a 1-μl solution of 1 mm AgNO3 in 20 mm HEPES-KOH, pH 7.0, was titrated into the wells of a 96-well plate (Reacti-Bind white opaque or Falcon) containing 100 μl of 15 μm protein (ScoHM, ScoHis, or ScoM1 containing the Phe/Trp mutation) in 20 mm HEPES-KOH, pH 7.0. Fluorescence (excitation at 280 nm, emission at 350 nm) was measured in a FLUOstar OPTIMA FL (BMG Labtech). Silver nitrate in buffer alone was used for background subtraction. To monitor the binding of Cu(II), 1-μl aliquots of 1 mm CuCl2 in 50 mm sodium phosphate, pH 7.2, were titrated into protein in the same buffer.
Ab Initio Modeling of the TM1,2-HM Region
Ab initio modeling of the TM1,2-HM region was performed on a locally installed version 3.0 of Rosetta ab initio modeling software. Briefly, the protocol used transmembrane region prediction from the OCTOPUS server to set initial membrane normal and membrane center vectors, which subsequently help define the membrane-specific environment (hydrophobic core, interface, polar, and water layers). Using limited constraints based on residue-residue interactions obtained from the transmembrane regions of the ϵ subunit of F1F0-ATPase (2E5T) (a typical transmembrane hairpin), the protocol constrains helix-helix packing arrangements at particular positions. The predicted membrane-spanning regions of ATP7A are consistent with those obtained using TMPred and MEMSAT2. Structure fragments were generated using the standard web-based Robetta fragment server and the secondary structure prediction method during the fragment selection procedure. 5000 independent models were generated. The resulting structures were clustered using the clustering program in the Rosetta ab initio software. The centers of the five biggest clusters were chosen as our best models, defined as having the lowest standard mean deviation value (between positions of carbon atoms of all residues) to all other models in a cluster. Amino acid side chains were added to the structures using a locally installed version of the program SCWRL (15).
X-ray Absorption Spectroscopy
To generate samples for x-ray absorption spectroscopy, copper was added to proteins as (Cu(CH3CN4)+ described in Ref. 14. Excess copper and acetonitrile were removed by two cycles of desalting using spin columns (Pierce). After concentration to 300–500 μm in copper, 20% ethylene glycol was added as cryoprotectant, and the samples were flash-frozen in liquid nitrogen. All steps were carried out at room temperature in an anaerobic chamber. Copper concentration was analyzed by atomic absorption spectroscopy (AA-6650G, Shimadzu, Columbia, MD) or by inductively coupled plasma optical emission spectrometry on a PerkinElmer Life Sciences Optima 2000 DV spectrometer.
Copper K-edge Data
Copper K-edge data (8.980 keV) were collected at the Stanford Synchrotron Radiation Lightsource. The extended x-ray absorption fine structure (EXAFS) and the x-ray absorption near edge structure data were measured on beam line 9-3, operating at 3 GeV with beam currents either between 200 and 160 mA or between 100 and 80 mA. The beam line was configured with a silicon[220] monochromator (crystal orientation ϕ = 90°) and a rhodium-coated mirror upstream of the monochromator with a 13-keV energy cutoff to reject harmonics. A second rhodium mirror downstream of the monochromator was used to focus the beam. Data were collected in fluorescence mode using a liquid nitrogen-cooled, high count rate Canberra 100-element or 30-element germanium array detector with maximum count rates below 120 kHz. Soller slits with a Z-1 metal oxide (nickel) filter were placed in front of the detector to selectively attenuate the elastic scatter peak. Under these conditions, no dead time corrections were necessary. Energy calibration was achieved by reference to the first inflection point of a copper foil (8980.3 eV) placed between the second and third ionization chamber. 4–6 scans of a sample containing only sample buffer (50 mm NaPO4, pH 7.2) were collected, averaged, and subtracted from the averaged data of the protein samples to remove Z-1 Kβ fluorescence and produce a flat pre-edge baseline. Protein samples (80 μl) were measured as aqueous glasses (containing ≥ 20% ethylene glycol) at 10–15 K in a liquid helium cryostat. The number of scans collected for each sample varied from 6 to 10, depending on the concentration of copper in the samples. The scans were collected to k = 12.8 Å−1 at the copper K-edge to avoid possible interference by traces of zinc in the samples. Data reduction and background subtraction were performed with the program modules of EXAFSPAK (16). Spectra simulation was carried out by least-squares curve fitting using the program EXCURVE 9.2 as described previously (14, 17, 18). The quality of the fits was evaluated by the goodness-of-fit parameter, F, also referred to as the fit index.
 |
RESULTS
The First Lumenal Loop of ATP7A Contains a Unique Sequence Insert
To identify structural element(s) that may contribute to the faster rate of copper release by ATP7A, we carried out sequence alignments of human Cu-ATPases and their orthologues. We found that the TMs and the lumenal loops were well conserved between ATP7A and ATP7B with one notable exception; ATP7A had a sequence insert in the first lumenal loop that was enriched with His and Met residues (Fig. 1B). The insert is present in all mammalian orthologues of ATP7A, and the motif 674MDHHXXXXHHXXXMSXEEMXXXHSxM699 is conserved. The lumenal loop of ATP7B is much shorter and has fewer His/Met residues (Fig. 1B). His and Met are commonly found in copper-binding sites. Because during the Cu-ATPase transport cycle copper is released at the luminal side and the rate of copper release is faster for ATP7A, the following questions were raised. Are the His and Met residues in the insert involved in copper binding? Is this insert responsible for the faster kinetics of copper release from ATP7A? Does the insert play any role in subsequent transfer to cuproenzymes? We addressed these questions by functional analysis of ATP7A mutants, by copper binding measurements after introduction of the His/Met rich loop (HM-loop) into scaffold proteins, and by spectroscopic studies on the Cu(I) complexes of the HM-loop.
Mutations in the HM-loop Inhibit ATP7A Dephosphorylation
To test whether the HM-loop influences copper release from the TM sites, we mutated several potential copper ligands in the HM-loop (Fig. 2A). In ATP7A-Mut1, 2 Met and 4 His residues in the first half of the loop were changed to Ala or Gly to preserve presumed flexibility of the loop. In ATP7A-Mut2, 2 Met residues in the second half of the HM-loop were replaced, both to Ala (Fig. 2A). The mutants and wild-type (WT) ATP7A were expressed in Sf9 cells, and the presence of full-length protein in membrane preparations was verified by Western blotting (supplemental Fig. 2A). WT ATP7A and mutants were then auto-phosphorylated by incubating with [γ-32P]ATP. Both mutants were active and had similar kinetics of phosphorylation (Fig. 2B). (The time dependence of phosphorylation was also comparable with that of WT ATP7A (supplemental Fig. 2B).)
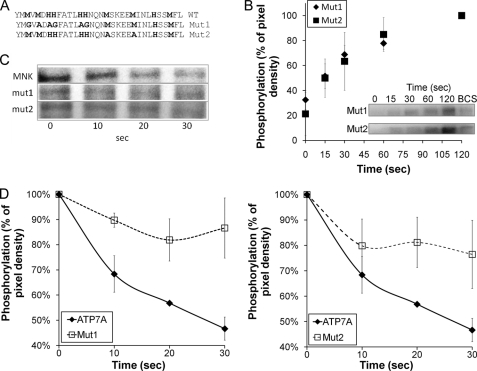
FIGURE 2.
Mutations within the HM-loop inhibit the ATP7A dephosphorylation without affecting phosphorylation. A, the region of the HM-loop mutated in full-length ATP7A; the residues replaced in ATP7A-Mut1 and Mut2 are shown in bold. B, phosphorylation of Mut1 and Mut2 over time where densitometry data for three independent experiments were averaged; phosphorylation at 2 min is taken as 100%. For copper depletion, protein was pretreated with 200 μm BCS before phosphorylation. C, autoradiography of gels showing changes in protein phosphorylation for WT ATP7A (MNK) and both mutants; Mut1 and Mut2 dephosphorylate more slowly than wild-type ATP7A. D, densitometry data for three independent experiments; the initial level of phosphorylation is taken as 100%. Error bars indicate S.E.
In contrast, dephosphorylation (the reaction coupled to copper release) for both mutants was significantly slower when compared with that of WT ATP7A (Fig. 2, C and D). ATP7A-Mut1 was dephosphorylated by about 16% after 20 s when compared with 57 ± 1% of WT (Fig. 2C). ATP7A-Mut2 also inhibited dephosphorylation, but not as significantly as Mut1 (Fig. 2D). Overall, both mutants never dephosphorylated to the level of the WT within the 30-s experiment.
The HM-loop of ATP7A Binds Copper
Slow dephosphorylation suggested that copper release was retarded in the ATP7A variants with mutations within the HM-loop. We hypothesized that normally the HM-loop facilitates copper release by binding copper and that mutations disrupted metal binding. Examining copper binding to the HM-loop in the context of the full-length ATP7A is difficult because ATP7A has multiple copper-binding sites: six in the N terminus and possibly two within the TM domain (Fig. 1A). Consequently, to examine copper binding, we expressed the HM-loop along with several flanking residues as a recombinant protein. The purified peptide MMVMDHHFATLHHNQNMSKEEMINLHSSM-FLERQILP was unstable, precipitated over time, and/or was absorbed on dialysis membranes. Therefore, we decided to anchor the HM-loop with a flanking scaffold protein.
First, we used the recombinant metal-binding domain 2 of ATP7B, MBS2, and replaced its native copper-binding loop, MTCQSC (Fig. 3A), with the sequence MDHHFATLHHNQNMSKEEMINLHSSM. To improve solubility, this sequence was shortened when compared with the recombinant peptide but still contained all residues unique for ATP7A. As a control, we used MBS2 in which copper binding was inactivated by replacing both Cys residues with Ala (MBS2-C>A). MBS2-HM was expressed as a soluble protein and was folded, as illustrated by CD spectroscopy (Fig. 3B). The presence of the HM insert altered the spectra when compared with the MBS2-C>A scaffold; particularly noticeable was a larger negative signal at 208 nm (Fig. 3B). The addition of Cu(I) to MBS2-HM induced structural changes that led to protein unfolding (supplemental Fig. 3), whereas no structural change was seen with the MBS2-C>A mutant. We speculated that the small size of MBS2 and the proximity of the insert to the N terminus (only 9 residues from the Met start site) made MBS2-HM vulnerable to copper-induced structural changes within the relatively large HM insert. To avoid complications associated with instability of MBS2-HM, we inserted the HM-loop into a larger scaffold, the B. subtilis Sco protein (Sco), replacing its copper-binding site 45CETIC49 (Fig. 4A). Endogenous His and Met residues in the Sco scaffold were mutated (to Ala and Ile, respectively, generating M52I/M56I/H55A/H135A-Sco). This Sco protein with the inserted HM-loop was termed ScoHM.
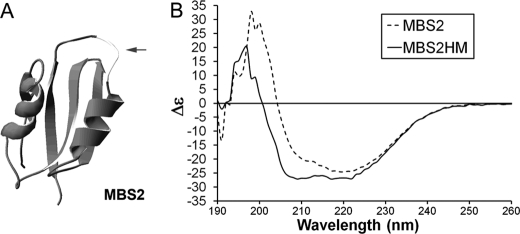
FIGURE 3.
Properties of MBS2-HM. A, structure of the MBS2 scaffold (dark gray) with the location of the CXXC loop replaced with the HM insert shown in light gray. B, CD spectra of MBS2 with and without the loop HM.
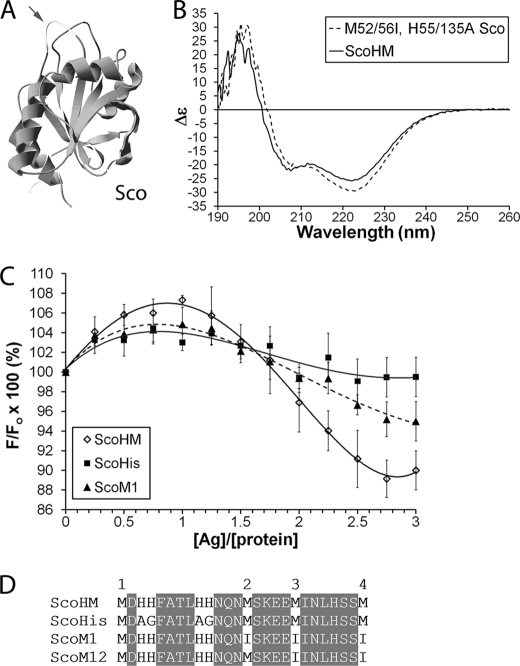
FIGURE 4.
Characterization of the ScoHM chimera. A, the structure of B. subtilis Sco (dark gray) with the CETIC sequence was replaced with the HM-loop shown in light gray with an arrow. B, CD spectra of Sco with (ScoHM) and without the HM-loop. C, sequence of ScoHM mutants; the position of mutants is indicated by the unshaded letters. Met residues are numbered 1–4 (see under “Results”). D, a single Phe residue in the HM-loop was replaced with Trp, and changes in Trp fluorescence were monitored following titration with Ag(I): ScoHM (diamonds, solid black line); ScoHis (squares, solid gray line); and ScoM1 (triangles, dashed line). The experiment was repeated in triplicate, and data were averaged. The data points were fit with a fourth polynomial curve. Error bars indicate S.E.
ScoHM was expressed as a soluble protein of the expected size, ∼22 kDa. The CD spectroscopy of ScoHM showed a more negative signal at 208 nm when compared with control M52I/M56I/H55A/H135A-Sco without insert (Fig. 4B); this change was similar to what was seen for MBS2-HM. Unlike MBS2-HM, ScoHM was stable as judged by the lack of precipitation, consistent CD spectroscopy, and a single band on SDS-PAGE. To evaluate metal binding to ScoHM, Cu(I) was added both aerobically and anaerobically (to better control for the oxidation state). After aerobic addition of 3 copper equivalents in the presence of 30-fold glutathione and extensive dialysis, ScoHM bound 2.0 ± 0.75 copper atoms/protein and remained soluble. Anaerobic addition of Cu(I) followed by desalting led to a slightly higher ratio of 2.7 ± 0.3 Cu(I) ions/protein.
To examine copper binding to ScoHM in more detail, we replaced the non-conserved Phe in the HFATL sequence of the HM-loop (corresponding to Phe678 of ATP7A) with Trp to use as a fluorescent probe. Titration with Cu(I) in the presence of glutathione produced a response that varied depending on incubation time prior to measurements (not shown). Because rapid Cu(I) oxidation to Cu(II) may cause this variability, we utilized Ag+ as a Cu(I) analog. This substitution is also physiologically relevant because ATP7A can facilitate Ag+ efflux (19). Silver titration into ScoHM was accompanied by reproducible changes of Trp fluorescence (Fig. 4C). Initially, fluorescence increased, with a maximum signal seen at 1:1 Ag+ ions per protein, whereas at 2–3 Ag+ ions per protein, fluorescence was quenched to ∼90% of initial level and did not change further. The apparent two-step change of Trp fluorescence was consistent with the presence of more than one metal-binding site in the HM-loop.
Copper Binds to His and Met Residues in the HM-loop
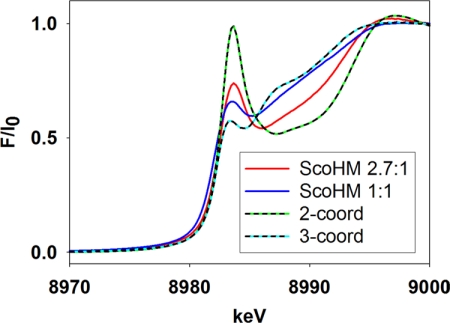
To identify structural components of these binding sites, we examined Sco-HM that was anaerobically loaded with Cu(I) at low (1:1) and high (2.7:1) stoichiometries by absorption edge and EXAFS spectroscopy. The intensity of the edge feature at 8983 eV is characteristic of coordination number; it increases in proportion to decreasing number of coordinating ligands (20, 21). At either high or low copper, the 8983 eV intensity was in between values for the 2- and 3-coordinate states, although high copper samples had a more intense feature, indicative of changes in coordination environment upon binding of additional copper (Fig. 5).
FIGURE 5.
Comparison of the intensities of the 8983 eV feature for Cu(I) complexes of the ScoHM chimera at Cu:P ratios of 2.7:1 (red trace) and 1:1 (blue trace) with those of genuine 2-coordinate (2-coord) (Cu(I)(His)2 peptide (22)) and 3-coordinate (3-coord) (Cu(I)(Met)2(His) in CusF (37)) systems. For example, the 2-coordinate bis-histidyl sites such as those present in the Cu(I) complex of the amyloid-β peptide (22, 38), and the bis-cysteinyl Cu(I)-binding sites of HAH1 (14) and CueR (39) exhibit intense 8983 eV transitions in normalized absorption edge data with intensities close to 1.
These conclusions were confirmed by EXAFS (Fig. 6). The Fourier transform of data for samples with 2.7 copper atoms yielded a spectrum typical of Cu(I)-His coordination. An intense first shell peak around 1.9 Å was assigned to Cu(I)-N single scattering, and peaks at 3 and 4 Å were assigned to single and multiple scattering interactions of the imidazole ring. Simulation of the EXAFS data produced a best fit corresponding to two Cu(I)-His interactions with a small component (∼0.5 ± 0.2) of Cu(I)-S scattering. In contrast, quantitative analysis of the EXAFS data for the sample with 1 bound copper atom showed 1.5 sulfur atom at 2.23 Å and 1 N(His) molecule at 1.88 Å, suggesting that at low copper, the binding site has a higher degree of Met coordination. Given that EXAFS fitting cannot determine shell occupancies to better than ∼25% precision, we also analyzed bond lengths, which are good indicators of coordination number. For both copper binding stoichiometries, the Cu(I)-N(His) bond length was 1.88 ± 0.2 Å, a value typical of linear 2-coordination in either Cu(I)N2 (22, 23) or mixed Cu(I)NS environments (24). The Cu(I)-S distance was 2.23 ± 0.2 Å, a value closer to the typical 3-coordinate distance of 2.25–2.30 Å (17, 18, 25–27). Altogether, the EXAFS data confirmed the presence of at least two structurally non-equivalent Cu(I)-binding sites. The site occupied at low copper is most likely a pseudo 3-coordinate formed by two sulfur atoms from Met and one oxygen/nitrogen atom, which could be either a His residue or another donor from a protein side chain, main chain amide, or solvent. The lower affinity site is most likely a 2-coordinate with a Cu(I)(His)2 or Cu(I)(His)(Met) configuration.
FIGURE 6.
Experimental (black) and simulated (red) Fourier transforms and EXAFS (insets) for the ScoHM Cu(I) complexes at a Cu:P ratio of 2.7:1 (bottom) and 1:1 (top). Spectra were nitrogen/oxygen phase shift-corrected. The blue dashed line in each spectrum corresponds to the Cu-N scattering, whereas the green dashed line corresponds to the Cu-S scattering.
His Pairs Are Important for Copper Binding
To clarify the role of His and Met in each site, we generated two ScoHM mutants. In ScoM12, 2 Met residues (corresponding to Met692 and Met699 of ATP7A) were mutated to Ile (Fig. 7A); in ScoHis, both His-His pairs (corresponding to His676/677 and His682/683 of ATP7A) were changed to Ala-Gly. ScoM12 bound 2 copper atoms and behaved similarly to ScoHM. In contrast, ScoHis had a decreased Cu(I) binding stoichiometry of 1.2, suggesting the loss of coordinating ability due to His mutation.
FIGURE 7.
Stronger copper retention by the HM-loop is associated with Cu(I) stabilization. A, BCS titration to 45 μm copper in the absence of protein (ascorbate alone) or in the presence of 15 μm ScoHM variants (in addition to ascorbate). The formation of Cu(I)(BCS)2 was monitored at 483 nm; the amount of complexed (BCS-chelated) copper was calculated using ϵ = 13,500 m−1·cm−1. ScoHM (dark blue) is right-shifted when compared with the ascorbate alone or mutants, indicating that copper is retained more strongly and is less available to BCS. Inset: at saturating concentration of BCS, sample containing ScoHM yields more Cu(I)(BCS)2 when compared with ascorbate alone (light blue) or the ScoHis mutant (green), suggesting stabilization of Cu(I) against oxidation by ScoHM. B, time-dependent oxidation of Cu(I) in the presence (light bars) and absence (dark bars) of 15 μm ScoHM monitored by the decrease in Cu(I)-BCS complex (calculated as above). The experiments in A and B were repeated in triplicate. Error bars indicate S.E.
The retention of 1 bound copper atom in the ScoHis mutant raised the possibility that multiple metal-coordinating residues in the HM-loop may substitute for each other (albeit not completely). This conclusion was further supported by measuring Trp fluorescence in ScoHis with a Phe/Trp replacement and the analysis of an additional mutant, ScoM1, in which 3 Met residues (corresponding to Mer687, Met692, and Met699 of ATP7A) were replaced with Ile (Fig. 4C). Similarly to ScoHM, Trp fluorescence was increased for both ScoHis and ScoM1 at low Ag+ stoichiometry, although increase was lower (Fig. 4D). At 2–3 silver atoms per protein, the ScoM1 fluorescence was quenched; however, the quenching (∼95% of initial) was less than that of ScoHM (∼90% of initial). For ScoHis, fluorescence only returned to the initial level. Thus, the mutation of either His or Met residues altered the structure of the binding sites without eliminating the ability of ScoHM to bind metals.
Cu(I) Retention and Stabilization
To test whether the mutations simply rearrange the binding sites or also diminish metal affinity, we compared copper retention by the ScoHM variants using BCS. (To prevent rapid copper oxidation, ascorbate was included in the buffer; the ascorbate-to-copper ratio was optimized prior to the experiment (supplemental Fig. 4).) As expected, the presence of ScoHM diminished the formation of the BCS-copper complex when compared with buffer alone, indicating competition between the HM insert and BCS for copper (Fig. 7A). ScoM1, ScoM12, and ScoHis also competed with BCS for copper, but overall showed lower copper retention when compared with ScoHM (Fig. 7A).
These experiments also yielded an unanticipated and interesting result. It should be noted that unlike Cu(I), the oxidized Cu(II) does not form a spectroscopically detectable complex with BCS. We noticed that upon titrations, the final amount of Cu(I)-BCS complex was higher in the presence of ScoHM when compared with buffer alone or when compared with ScoHis mutant (Fig. 7B, inset). This result was reproducible and suggested that during titration, the ScoHM stabilized copper in the reduced form better than ascorbate, releasing it eventually to BCS. The effect was especially noticeable in low ascorbate when copper oxidizes more rapidly (data not shown). To verify this conclusion, we assessed the amount of Cu(I) in the low ascorbate buffer in the absence and presence of ScoHM at several time points (Fig. 7B). Over time, Cu(I) was oxidized to Cu(II) and thus became unavailable for Cu(I)-BCS complex formation. However, when ScoHM was present, Cu(I) did not oxidize as quickly; 2-fold as much copper was available for BCS even after a 1-h incubation (Fig. 7B). Thus, the HM-loop stabilizes Cu(I) and protects it from oxidation. Finally, we tested whether after the oxidation copper could rebind to the HM-loop. The addition of CuCl2 and dialysis demonstrated that the HM-loop binds Cu(II) at 1.9 ± 0.2 ions per protein (supplemental Table 2).
DISCUSSION
Despite significant progress in characterization of copper transporters (1, 2, 4, 5, 28), the mechanisms of copper transfer into intracellular compartments remain poorly understood. Unlike many other members of the P-type ATPase family, Cu-ATPases do not generate ion gradients; instead, transported copper is incorporated into acceptor molecules. The apparent high affinity of intramembrane binding sites (shown for archaeal orthologues CopA (28)) and a small number of potential copper-binding ligands raise a question about driving force and the mechanics of copper release from the transporters, as well as the role lumenal acceptors. Direct interaction between Cu-ATPase ATP7B and its target ceruloplasmin has been demonstrated (29, 30); whether this interaction is important for copper release remains to be tested. ATP7A has multiple targets (dopamine-β-hydroxylase (31), lysyl oxidase (32), tyrosinase (6), and peptidylglycine-α-amidating mono-oxygenase (33)), and tight interaction with them is not required for copper transport (34). This latter observation suggested that specific structural elements within ATP7A might contribute to copper release, making it faster than in ATP7B. We demonstrate that the unique TMS1,2 loop of ATP7A (HM-loop) may play such a regulatory role. Mutation of Met/His residues in this loop inhibit the dephosphorylation of ATP7A, consistent with impaired copper release. When inserted into scaffold proteins, the HM-loop binds 2–3 copper atoms and undergoes conformational transitions, which in full-length ATP7A can be transmitted to the membrane domain. Furthermore, the HM-loop stabilizes Cu(I), which may facilitate transfer of copper from ATP7A to acceptor proteins. Altogether, our results suggest that the HM-loop serves as a molecular device to guide transiently bound copper away from the transmembrane domain.
How does copper released from the transport site reach the HM-loop? To address this issue, we modeled the TM1,2 hairpin structure using the Rosetta software. The model revealed a string of Met residues (a “Met-wire”) in TM1 (Fig. 8A). A similar Met-wire is also present in the TM1 of ATP7B (not shown), suggesting the existence of a common copper pathway to the surface for Cu-ATPases (Fig. 8, B and C). The presence of Met wires in copper transporters was previously noticed in mammalian copper uptake transporter Ctr1 (35) and in the bacterial copper efflux protein CusA (36). As in Ctr1 and CusA, Met residues appear to provide transient sites that assist Cu(I) passage through and exit from the membrane. The 3-coordinate environment of the first Cu(I)-binding site in the HM-loop (Fig. 8C) adds to the emerging coordination chemistry of copper transporters, whereas the 2-coordinate (Met)(His) pair(s) may represent the first stable homologues of the transient Met pairs observed in Ctr1 and CusA.
FIGURE 8.
Suggested spatial relationship of the metal-coordinating residues in the TM1,2 hairpin. A and B, Rosetta model of the TM1,2 hairpin (A) is overlaid in a graphic of ATP7A (B) to illustrate a possible Cu(I) path from the intramembrane copper-binding site CPC to the luminal surface. Met residues (shown in red) are clustered along the same surface of the TM1. C, potential arrangement of Cu(I)-binding sites (Site I and Site II) in the HM-loop. In the first cluster (closer to the membrane), the first and fourth Met residues (Met674 and Met699 of the full-length ATP7A, respectively) are in the vicinity of the first His pair. The residues are all within ∼6 Å of one another so that small movements of the loop backbone could result in binding to Cu(I). In the second cluster, one of the His residues from the His pair together with either the second or the third Met residue (Met687 or Met692) could form a mixed 2-coordinate site, and H4 (His683) and M2 (Met687) are within 4 Å of each other.
Our CD, Trp fluorescence, and EXAFS data suggest that binding of copper to the HM-loop induces significant conformational changes. At first, copper binds to a 3-coordinate site. This binding results in structural changes and, at higher copper, the second state becomes populated. The binding of a second copper triggers further change in the HM structure, as demonstrated by fluorescence quenching and modification of the coordination environment to (His)2 and/or (His)(Met). It is interesting that even at high copper levels, only a fraction of the His and Met residues in the loop is involved in Cu(I) binding, although different residues may be utilized in different conformational states. This suggests a highly dynamic environment for Cu(I) binding, where the coordination and conformation of the loop can be significantly influenced by external factors such as pH and redox potential. The HM-loop is also able to bind Cu(II). These properties of the HM-loop may play a role in the copper transfer to the enzymes within the secretory pathway. The target enzymes to which ATP7A transfers copper differ in their chemical properties as well as intracellular location (Golgi, secretory vesicles, melanosomes). The conditions of pH, oxygen tension, and redox potential vary significantly within different cellular compartments. Therefore, the ability of the lumenal loop to adopt different structures and conformations as a function of pH and/or redox potential may be the controlling factor in ensuring rapid and efficient copper release to cuproenzymes.
Supplementary Material
This work was supported, in whole or in part, by National Institutes of Health Grant P01GM067166 (to S. L. and N. J. B.).

The on-line version of this article (available at http://www.jbc.org) contains supplemental Tables 1 and 2 and Figs. 1–4.
- TM
- transmembrane
- HM-loop
- the loop Met672–Pro707 of ATP7A
- BCS
- bathocuproine disulfonate
- EXAFS
- extended X-ray absorption fine structure
- Bis-Tris
- 2-(bis(2-hydroxyethyl)amino)-2-(hydroxymethyl)propane-1,3-diol.
REFERENCES
- 1. Lutsenko S., Barnes N. L., Bartee M. Y., Dmitriev O. Y. (2007) Physiol. Rev. 87, 1011–1046 [DOI] [PubMed] [Google Scholar]
- 2. Tsivkovskii R., Eisses J. F., Kaplan J. H., Lutsenko S. (2002) J. Biol. Chem. 277, 976–983 [DOI] [PubMed] [Google Scholar]
- 3. Kuo Y. M., Gitschier J., Packman S. (1997) Hum. Mol. Genet. 6, 1043–1049 [DOI] [PubMed] [Google Scholar]
- 4. Pilankatta R., Lewis D., Adams C. M., Inesi G. (2009) J. Biol. Chem. 284, 21307–21316 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5. Voskoboinik I., Mar J., Strausak D., Camakaris J. (2001) J. Biol. Chem. 276, 28620–28627 [DOI] [PubMed] [Google Scholar]
- 6. Petris M. J., Strausak D., Mercer J. F. (2000) Hum. Mol. Genet. 9, 2845–2851 [DOI] [PubMed] [Google Scholar]
- 7. Forbes J. R., Cox D. W. (1998) Am. J. Hum. Genet. 63, 1663–1674 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8. Michalczyk A., Bastow E., Greenough M., Camakaris J., Freestone D., Taylor P., Linder M., Mercer J., Ackland M. L. (2008) J. Histochem. Cytochem. 56, 389–399 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9. Barnes N., Tsivkovskii R., Tsivkovskaia N., Lutsenko S. (2005) J. Biol. Chem. 280, 9640–9645 [DOI] [PubMed] [Google Scholar]
- 10. Leonhardt K., Gebhardt R., Mössner J., Lutsenko S., Huster D. (2009) J. Biol. Chem. 284, 7793–7802 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11. Lowry O. H., Rosebrough N. J., Farr A. L., Randall R. J. (1951) J. Biol. Chem. 193, 265–275 [PubMed] [Google Scholar]
- 12. Walker J. M., Huster D., Ralle M., Morgan C. T., Blackburn N. J., Lutsenko S. (2004) J. Biol. Chem. 279, 15376–15384 [DOI] [PubMed] [Google Scholar]
- 13. Andruzzi L., Nakano M., Nilges M. J., Blackburn N. J. (2005) J. Am. Chem. Soc. 127, 16548–16558 [DOI] [PubMed] [Google Scholar]
- 14. Ralle M., Lutsenko S., Blackburn N. J. (2003) J. Biol. Chem. 278, 23163–23170 [DOI] [PubMed] [Google Scholar]
- 15. Canutescu A. A., Shelenkov A. A., Dunbrack R. L., Jr. (2003) Protein Sci. 12, 2001–2014 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16. George G. N. (1995) EXAFSPAK, Stanford Synchrotron Radiation Laboratory, Menlo Park, CA [Google Scholar]
- 17. Blackburn N. J., Rhames F. C., Ralle M., Jaron S. (2000) J. Biol. Inorg. Chem. 5, 341–353 [DOI] [PubMed] [Google Scholar]
- 18. Bagai I., Rensing C., Blackburn N. J., McEvoy M. M. (2008) Biochemistry 47, 11408–11414 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19. Ibricevic A., Brody S. L., Youngs W. J., Cannon C. L. (2010) Toxicol. Appl. Pharmacol. 243, 315–322 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20. Kau L. S., Spira-Solomon D. J., Penner-Hahn J. E., Hodgson K. O., Solomon E. I. (1987) J. Am. Chem. Soc. 109, 6433–6442 [Google Scholar]
- 21. Pickering I. J., George G. N., Dameron C. T., Kurz B., Winge D. R., Dance I. G. (1993) J. Am. Chem. Soc. 115, 9498–9505 [Google Scholar]
- 22. Himes R. A., Park G. Y., Barry A. N., Blackburn N. J., Karlin K. D. (2007) J. Am. Chem. Soc. 129, 5352–5353 [DOI] [PubMed] [Google Scholar]
- 23. Sanyal I., Karlin K. D., Strange R. W., Blackburn N. J. (1993) J. Am. Chem. Soc. 115, 11259–11270 [Google Scholar]
- 24. Siluvai G. S., Mayfield M., Nilges M. J., Debeer George S., Blackburn N. J. (2010) J. Am. Chem. Soc. 132, 5215–5226 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25. Chong L. X., Ash M. R., Maher M. J., Hinds M. G., Xiao Z., Wedd A. G. (2009) J. Am. Chem. Soc. 131, 3549–3564 [DOI] [PubMed] [Google Scholar]
- 26. Sarret G., Favier A., Covès J., Hazemann J. L., Mergeay M., Bersch B. (2010) J. Am. Chem. Soc. 132, 3770–3777 [DOI] [PubMed] [Google Scholar]
- 27. Bagai I., Liu W., Rensing C., Blackburn N. J., McEvoy M. M. (2007) J. Biol. Chem. 282, 35695–35702 [DOI] [PubMed] [Google Scholar]
- 28. González-Guerrero M., Eren E., Rawat S., Stemmler T. L., Argüello J. M. (2008) J. Biol. Chem. 283, 29753–29759 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29. di Patti M. C., Maio N., Rizzo G., De Francesco G., Persichini T., Colasanti M., Polticelli F., Musci G. (2009) J. Biol. Chem. 284, 4545–4554 [DOI] [PubMed] [Google Scholar]
- 30. Maio N., Polticelli F., De Francesco G., Rizzo G., Bonaccorsi di Patti M. C., Musci G. (2010) J. Biol. Chem. 285, 20507–20513 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31. Kaler S. G., Holmes C. S., Goldstein D. S. (1998) Adv. Pharmacol. 42, 66–68 [DOI] [PubMed] [Google Scholar]
- 32. Tchaparian E. H., Uriu-Adams J. Y., Keen C. L., Mitchell A. E., Rucker R. B. (2000) Arch. Biochem. Biophys. 379, 71–77 [DOI] [PubMed] [Google Scholar]
- 33. Steveson T. C., Ciccotosto G. D., Ma X. M., Mueller G. P., Mains R. E., Eipper B. A. (2003) Endocrinology 144, 188–200 [DOI] [PubMed] [Google Scholar]
- 34. El Meskini R., Culotta V. C., Mains R. E., Eipper B. A. (2003) J. Biol. Chem. 278, 12278–12284 [DOI] [PubMed] [Google Scholar]
- 35. De Feo C. J., Aller S. G., Siluvai G. S., Blackburn N. J., Unger V. M. (2009) Proc. Natl. Acad. Sci. U.S.A. 106, 4237–4242 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36. Long F., Su C. C., Zimmermann M. T., Boyken S. E., Rajashankar K. R., Jernigan R. L., Yu E. W. (2010) Nature 467, 484–488 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37. Loftin I. R., Franke S., Blackburn N. J., McEvoy M. M. (2007) Protein Sci. 16, 2287–2293 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38. Himes R. A., Park G. Y., Siluvai G. S., Blackburn N. J., Karlin K. D. (2008) Angew. Chem. Int. Ed. Engl. 47, 9084–9087 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39. Chen K., Yuldasheva S., Penner-Hahn J. E., O'Halloran T. V. (2003) J. Am. Chem. Soc. 125, 12088–12089 [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.