Abstract
Lactic acid bacteria (LAB) have been used in fermentation processes for centuries. More recent applications including the use of LAB as probiotics have significantly increased industrial interest. Here we present a comparative genomic analysis of four completely sequenced Lactobacillus strains, isolated from the human gastrointestinal tract, versus 25 lactic acid bacterial genomes present in the public database at the time of analysis. Lactobacillus acidophilus NCFM, Lactobacillus johnsonii NCC533, Lactobacillus gasseri ATCC33323, and Lactobacillus plantarum WCFS1are all considered probiotic and widely used in industrial applications. Using Differential Blast Analysis (DBA), each genome was compared to the respective remaining three other Lactobacillus and 25 other LAB genomes. DBA highlighted strain-specific genes that were not represented in any other LAB used in this analysis and also identified group-specific genes shared within lactobacilli. Initial comparative analyses highlighted a significant number of genes involved in cell adhesion, stress responses, DNA repair and modification, and metabolic capabilities. Furthermore, the range of the recently identified potential autonomous units (PAUs) was broadened significantly, indicating the possibility of distinct families within this genetic element. Based on in silico results obtained for the model organism L. acidophilus NCFM, DBA proved to be a valuable tool to identify new key genetic regions for functional genomics and also suggested re-classification of previously annotated genes.
Electronic supplementary material
The online version of this article (doi:10.1007/s12263-010-0191-9) contains supplementary material, which is available to authorized users.
Keywords: In silico analysis, PAU, Adhesion, Stress response, Bacteriophage
Introduction
Historically lactic acid bacteria (LAB) have been used in many industrial fermentation processes. Only recently have some members of this group emerged as probiotics [36, 37, 41] and several strains are also under consideration for delivery of biotherapeutics [19, 27, 47]. LAB represent a remarkably heterogeneous group, with members assigned by their ability to produce lactic acid either by homo- or heterofermentative metabolism. The subgroup of the Lactobacillus complex [38] is of particular interest due to the fact that many members occupy important ecological niches in the gastrointestinal tracts of humans and animals such as mice, piglets, dogs, cats, cattle, and others [12, 45, 51, 54, 56]. Four complete Lactobacillus genomes were available at the time of analysis: Lactobacillus acidophilus NCFM, Lactobacillus gasseri ATCC33323, Lactobacillus johnsonii NCC533, and Lactobacillus plantarum WCFS1. Three more are completed: Lactobacillus brevis ATCC367 [44], Lactobacillus bulgaricus [66], and Lactobacillus casei ATCC334 [44], and were pending public release.
Often, closely related strains display unique features and metabolic capabilities. However, the genetic background facilitating these attributes remained largely unknown. With the recent availability of complete genome sequences, a plethora of new information became available, albeit that a significant proportion of predicted genes remain functionally unclassified. Furthermore, problems in the reliability of automated and manually verified annotation efforts aggravate functional predictions and the precision of targeted functional genomic efforts.
Previous genome analyses focused mainly on predicted metabolic features, relying on functional annotation, and genome composition and synteny [6, 18, 39, 55].
However, these types of analyses seldom answer some of the most interesting questions. What enables related organisms to fill different niches and what genes are involved in their unique metabolic capabilities? What genes do single organisms or groups of organisms share with each other? Which genes are unique within these groups? What are the differences and similarities between different (related) groups of organisms? How can new genetic targets be selected that investigate the unique features found for smaller group-subsets or single organisms? Another challenge is to identify new genetic targets independent from functional classifications and presumed context, thus becoming more independent from mis-annotated and unidentified ORFs. Discovery of these genetic regions might prove to be a key to targeting functional genomic research in the future.
Here we present a comparison of four Lactobacillus genomes of human origin. In contrast to previous analyses, we primarily focused on unique or group-specific genome regions using Differential Blast Analysis (DBA), identifying new genetic targets with a high likelihood of importance to functional genomics. We attempt to analyze these genetic regions in context and propose possible mechanistic models based on gene synteny and function.
Materials and methods
In silico solutions
Genome visualizations were realized using Artemis v7 [57], Artemis Comparison Tool (ACT) [17], and Genewiz [52]. Data generation and processing for Genewiz were performed using in-house developed software (unpublished). Creation of customized databases and localized Blast analyses were realized using the stand alone Blast distribution from NCBI (ftp://ftp.ncbi.nih.gov/blast/executables/LATEST/).
Database creation
Four complete Lactobacillus genomes from human origins, L. acidophilus NCFM (ACC:CP000033) [6], L. gasseri ATCC33323 (ACC:CP000413) [8], L. johnsonii NCC533 (ACC:AE017198) [55], and L. plantarum WCFS1 (ACC:AL935263) [39] were analyzed and compared using BlastP and Differential Blast Analysis (DBA). Existing gene models (manually verified and published GAMOLA [4] model for L. acidophilus NCFM, the verified gene model for L. gasseri ATCC33323 [44], and published models for L. johnsonii NCC533 and L. plantarum WCFS1) were used to deduce amino acid sequences for each predicted open reading frame (ORF). These amino acid sequences were used to compile the custom Lactobacillus BlastP-database (dbLB), utilizing formatdb from NCBI’s Blast distribution. Similarly, a second custom database (dbLAB), excluding the aforementioned four organisms, was created, consisting of 25 lactic acid bacteria genomes and 104 plasmids, present at the time of analysis. A summary of the database content is shown in Table 1 and a detailed description can be found in Supplemental Table 1.
Table 1.
Firmicutes and Actinobacteria used to generate the custom LAB database
| Organisms | Genomes | Plasmids |
|---|---|---|
| Bacilli | 1 | 5 |
| Bifidobacteriae | 2 | 11 |
| Brevibacteriae | 1 | 0 |
| Enterococci | 1 | 9 |
| Lactobacilli | 3 | 31 |
| Lactococci | 2 | 23 |
| Leuconostoc sp | 1 | 5 |
| Oenococci | 1 | 0 |
| Pediococci | 1 | 2 |
| Streptococci | 12 | 18 |
BlastP and differential blast analyses
Each ORFeome of the four genomes was analyzed using BlastP and both custom databases. BlastP hits to the respective query organisms were ignored. The resulting best BlastP hit was used as basis for the respective trust level assignment. A trust level represents an empirically deduced range of probabilities, permitting a clustering of e-value ranges of similar significance. Trust level ranges and values are shown in Table 2. The trust levels for each predicted ORF and each database are then directly displayed (BlastP results) and differentially compared and represented (DBA results) on the respective genome atlas circles (Fig. 1a–d).
Table 2.
Trust level assignments used for in silico analyses
| e-value range | Trust level |
|---|---|
| >1e-10 | 1 |
| 1e-10 to 1e-40 | 2 |
| 1e-40 to 1e-70 | 3 |
| 1e-70 to 1e-100 | 4 |
| 1e-100 to 1e-130 | 5 |
| 1e-130 to 1e-160 | 6 |
| 1e-160 to 0.0 | 7 |
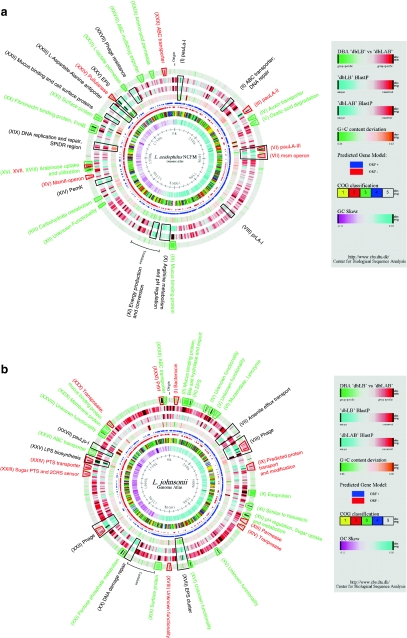
Fig. 1.
Comparative DBA analysis of four Lactobacillus genomes. The atlases each represent a circular view of the complete genome sequences of L. acidophilus NCFM (a), L. johnsonii NCC533 (b), L. gasseri ATCC33323 (c), and L. plantarum WCFS1 (d), respectively. The right-hand legend describes the single circles in the top-down- outermost-innermost direction. The circle was created using Genewiz and in-house developed software. Innermost circle 1, GC-Skew. Circle 2, COG classification. Predicted ORFs were analyzed using the COG database and grouped into the 4 major categories. 1, Information storage and processing; 2, Cellular processes and signaling; 3, Metabolism; 4, Poorly characterized; and 5, ORFs with uncharacterized COGs or no COG assignment. Circle 3, ORF orientation. ORFs in sense orientation (ORF +) are shown in blue; ORFs oriented in anti-sense direction (ORF −) in red. Circle 4, G+C content deviation. Deviations from the average GC content are shown in either green (low GC spike) or orange (high GC spike). A boxfilter was applied to visualize contiguous regions of low or high deviations. Circles 5 and 6, Blast similarities, using LAB and Lactobacillus custom databases, respectively. Deduced amino acid sequences compared against the respective database using gapped BlastP [7]. Regions in blue represent unique proteins in NCFM, whereas highly conserved features are shown in red. The degree of color saturation corresponds to the level of similarity. Circle 7, DBA analysis. Predicted ORFs present in at least one member of the Lactobacillus database (dbLB), but not equally conserved in any member of the LAB database (dbLAB) are highlighted in green. Geneproducts present in at least one member of the LAB database, but not equally conserved in any member of the Lactobacillus database are highlighted in red. Features described in more detail are indicated by trapezoids in the corresponded color and roman numbers. Black trapezoids represent unique genome regions, either not found in any custom database or otherwise outstanding within the genome (color figure online)
Differential Blast Analysis is both independent from ORF position on the respective reference and subject genomes, and from given ORF annotations. Thereby, impacts caused by genome rearrangements or mis-annotations are significantly reduced. Trust levels (tl) found for each investigated ORF of the reference genome were differentially analyzed against both databases (ΔDBA = tldbLB − tldbLAB). Extreme values range from −6 to +6, indicating the complete lack of similarity of a deduced amino acid sequence to entries in one of the two databases. Ranges in between describe the relative difference between trust levels with respect to the chosen databases. DBA in combination with genome atlases created by Genewiz indicates group-specific genes by color shades in relationship to a reference genome (either red or blue in this study). The intensity of the respective color is a direct visual indicator of the relative affiliation of the subject ORF to a given group. In this study, regions marked in red indicate ORFs conserved within at least one member of dbLAB and the reference genome, but not within the other three lactobacilli (dbLB) and vice versa for regions shown in green. Because DBA does not rely on any annotation, ORFs without a functional classification can be taken into account, thus allowing for a more complete analysis of the ORFeome. Selected homologous genome regions highlighted on the genome circles of more than one organism are described once in detail and referenced thereafter.
Protein ACT
Genome similarity and synteny on amino acid level was realized using an in-house developed duplex BlastP algorithm that compares the complete ORFeome of two organisms (GAMOLA [4] annotated) and assigns up to 20 similarity hits per ORF based on a trust interval assignment. Raw data are then converted into an ACT-compatible format. Sequential analyses of several genomes allowed the direct comparison of multiple complete genomes, simultaneously. By changing the identity threshold in ACT, displayed hits can be varied from relaxed to stringent.
Software availability
All software scripts for in-house solutions were developed in PERL and are available from the authors upon request.
Results and discussion
Lactobacillus acidophilus NCFM
L. acidophilus NCFM was originally isolated from humans and has been used extensively in industrial applications as a probiotic culture. Its complete genome sequence and content have been described [6], highlighting various genome regions harboring potentially significant genes and gene clusters. These analyzes were performed using regular similarity and gene synteny searches using standard databases. To investigate strain-specific traits within lactobacilli of human origin and other LAB, custom databases dbLB and dbLAB were utilized and revealed the presence of several unique regions, including three previously described genetic elements designated as pauLA-I through pauLA-III (potential autonomous unit; PAU). These elements resemble features from both plasmids and phages and were not found in other lactic acid bacteria to date. Because the core region of each PAU is mostly unique with regard to amino acid sequence similarities to LAB, use of the custom databases dbLB and dbLAB highlighted all three regions on the respective genome atlas (Fig. 1a, I, III, and VI, blue shading). In contrast, the genome regions adjacent to the core for pauLA-II and pauLA-III were identified by DBA due to the presence of a Type II and a Type III restriction endonuclease (LBA332 and 475), respectively, that was not present in the other three lactobacilli. However, homologs to these nucleases can be found in Streptococcus thermophilus, Bacillus subtilis and to a lesser degree, in the draft phase sequence of Lactobacillus bulgaricus. The Doc-Phd system featured by pauLA-I remained unique for L. acidophilus NCFM, with no close homologs among LAB. Strikingly, despite the lack of apparent core sequence similarities, more PAUs were identified by functional classifications and synteny analyses in other LAB and a detailed analysis was performed in reference to L. johnsonii NCC533.
Another L. acidophilus NCFM-specific locus at ~250 kb features a potential DNA repair system (Fig. 1a, II). The gene cluster describes a multidrug resistance protein that may resemble a permease (Lba251 to 253) and a 6-O-methylguanine DNA methyltransferase (Lba255), likely to be arranged in an operon-like structure. The formation of O-6-alkylguanine represents one of the major mutagens for DNA [48]. The repair of such DNA is facilitated by DNA S-methyltransferases by transferring the alkyl group at the O-6 position to a cysteine residue, consequently inactivating the enzyme in a suicide reaction. These enzymes are featuring two distinct domains: an N-terminal methyltransferase and a C-terminal DNA-binding domain. Lba255 revealed a well-conserved DNA-binding domain and although the methyltransferase domain was only weakly conserved, most of the absolutely conserved residues were maintained [48], indicating a preservation of the methylase function. An alternative function of the methyltransferase is that of a transcriptional activator which is mediated by self-methylation at Cys-38 [60]. The immediate presence of a predicted multidrug efflux system might point to a potential export mechanism for (conjugated) alkylating agents [63, 72]. The genetic arrangement of this locus may suggest a dual functionality of the methyltransferase, acting both as a DNA repair system and a chemosensor for adaptive response. Although L. johnsonii NCC533 features a related methyltransferase (Ljo252) at a similar genome position, no transport mechanism could be identified upstream of this gene, implicating this as a unique DNA repair mechanism in L. acidophilus NCFM. However, the transport system may be inactivated by a premature stop-codon and its functionality remains to be verified.
Located upstream of the terminus of replication, two large genome regions were identified (1.012–1.055 Mb and 1.073–1.092 Mb), predominantly specific to L. acidophilus NCFM (Fig. 1a, X and XI). Within the first region, a unique arginase (Lba1022, EC3.5.3.1) was identified, preceded by a transcriptional regulator of the merR family (Lba1021). This enzyme might catalyze the hydrolysis of arginine into l-ornithine and urea. As reported earlier, L. acidophilus NCFM encodes an ornithine decarboxylase (Lba996) that decarboxylates ornithine to putrescine and carbon dioxide [9], consuming a proton and potentially raising the cytoplasmic pH. The presence of a putrescine export system (Lba709 to 712) further strengthens the model of a specific internal mechanism to compensate for lowering pH during acidification. The presence of a periplasmatic-binding protein for putrescine (Lba713) points to the fact that under normal circumstances this ABC transporter is responsible for the uptake of this metabolite. However, it is noteworthy that under certain conditions ABC transporters may also act bi-directionally and are capable of generating ATP in the process [13].
Within the second L. acidophilus NCFM–specific region (Fig. 1a, XI), a gene cluster was identified, consisting of 16 genes (LBA1100 to 1115). This cluster might be divided into three groups, each consisting of a flavodoxin, an associated oxidoreductase, and some accessory proteins. Although no immediate functional assignment within the metabolic network could be performed, the close proximity of these genes to each other could indicate an intimate cooperation between the single gene sets. Furthermore, the COG classification of all three groups referred to energy production and conversion and the presence of two membrane transporters further suggests a strain-specific mechanism for metabolite uptake and conversion. Despite the presence of several predicted premature stop-codons and frameshifts within the first set of genes, it certainly would be intriguing to further characterize this unusual genome region.
A ppGpp regulated growth inhibitor protein of the PemK family (Lba1405) has been described for L. acidophilus NCFM (Fig. 1a, XIV). PemK is part of a plasmid maintenance system in E. coli, whereby PemK inhibits growth of host cells. PemK has also been reported to show DNA-binding capabilities, autoregulating its own synthesis (PFam2452). The antagonist, PemI, binds to PemK thus inactivating the protein [46]. However, the corresponding suppressor PemI was not identified in the L. acidophilus NCFM genome. Alternatively, the presence of a predicted bacterial transcription activator (LBA1408) upstream of LBA1405 could indicate a possible interaction of LBA1405 and LBA1408 in transcriptional regulation of this genome region. While this system is unique to L. acidophilus NCFM, a toxin–antitoxin system was recently described for the L. johnsonii NCC2761 (closely related to L. johnsonii NCC533) prophage LJ771. There is evidence that the mazE-pemK system contributes to prophage stability within L. johnsonii NCC2761, although the specific role of this system for both the phage and its host remains to be elucidated [20].
A putative l-aspartate-l-alanine conversion system was identified in L. acidophilus NCFM that was unique among the LAB examined (LBA1695 and 1696) (Fig. 1a, XXIII). LBA1695 has been annotated as aspartate aminotransferase, converting l-aspartate into oxaloacetate (EC2.6.1.1). However, metabolic pathway mapping using PathwayVoyager [5] and the KEGG database [33] also revealed a second highly conserved hit to an l-aspartate beta-decarboxylase (EC 4.1.1.12) that converts l-aspartate into l-alanine while releasing carbon dioxide. A membrane-bound antiporter (LBA1696) then mediates the exchange of l-alanine and l-aspartate, generating a physiological membrane potential. This system might be involved in yielding metabolic energy by generating a proton motif force and regulating internal pH. Similar systems have been described in Tetragenococcus halophila D10 and are generally referred to as proton motif metabolic cycles [1]. It is noteworthy that related systems were described for other lactobacilli (i.e. L. sakei ssp. sakei) that are not considered commensals.
The above-described genome regions highlighted strain-specific traits. DBA, however, identifies areas that are group-specific and may represent features shared by all or some members of this group, whereas the same regions are absent in another set of organisms. Among the most prominent regions identified by DBA are those harboring surface, mucus-, and fibrinogen-binding proteins (LBA1019, 1020, 1611, 1612, and 1634) (Fig. 1a, IX and XXI). All of these regions (green) are generally conserved between at least one of the three lactobacilli, but not found in other LAB, indicating their importance in strain/group-specific cell–cell interactions. Functional analyses of selected surface genes confirmed their role in cell adhesion and potentially cell–host interactions, in vitro [16]. It might be noteworthy that besides these group-specific genes, a number of strain-specific genes, potentially involved in cell adhesion (LBA1495, 1496, and 1654) or located on the cell surface (LBA1654), were also identified (Fig. 1a, XXII). Strain specificity of cell surface proteins appears to be a widespread and important factor mediating particular cell–cell interactions. Recently, a novel genetic locus, spaABCDEF coding for LPXTG-like pilins and a pilin-dedicated sortase in Lactobacillus rhamnosus GG, was identified [34]. The functional characterization of this locus revealed a mucus adhesion mechanism not previously described in lactobacilli. Specifically, presence of the cell wall–bound SpaC pilin was of critical importance to mucus binding, directly affecting retention time of L. rhamnosus GG in the intestinal tract [68]. Because of the biological significance, all four genomes analyzed here were screened for the presence of such a pilus cluster. A PSI-Blast over three iterations against the non-redundant database provided by NCBI and a BlastP analysis against the ORFeomes of L. acidophilus NCFM, Lactobacillus gasseri ATCC33323, Lactobacillus johnsonii NCC533, and Lactobacillus plantarum WCFS1 were carried out. While hits against other L. rhamnsosus and Lactobacillus casei strains were identified for the spaABCDEF cluster, no significant matches were detected against L. acidophilus NCFM, L. johnsonii NCC533, L. gasseri ATCC33323 or L. plantarum WCFS1—with the exception of hits against L. gasseri strain 224-1 (spaA, spaB, and spaE) and one weak hit to spaF in L. plantarum WCFS1. Similar results were found when the spaABCDEF cluster was compared to the four genomes directly. Only L. plantarum WCFS1 showed some weak matches, most of them unspecific hits to surface proteins. It may be interesting to point out that PSI-Blast analyses revealed an extensive number of matches throughout the gene cluster to numerous enterococci, supporting the proposed model of acquisition of this gene cluster via horizontal gene transfer [34].
Directly adjacent to the strain-specific gene cluster LBA251 to LBA255 (Fig. 1a, II), a three gene operon was identified potentially involved in drug resistance (Lba246 to Lba248). The operon consists of an ABC transport system (ATPase and permease component) and a LytR-type response regulator. Although the ATPase component was annotated as a Daunorubicin resistance protein, alignments of the Daunorubicin (DNR) resistance genes DrrA and DrrB from Streptomyces peucetius [29] did not reveal a high degree of similarity between the respective protein sequences (DrrA: 32% identity, 53% positive; DrrB: no similarities detected) and it appears to be more likely for this operon to be involved in other types of drug transport. Only L. johnsonii NCC533 showed significant homologies to these genes, whereas other LAB merely displayed limited levels of conservation for certain conserved domains.
Oxalic acid is a strong dicarboxylic acid that can cause pathological disorders [49]. At this point, degradation of oxalic acid in the gastrointestinal tract has been reported to be linked exclusively to the bacterial commensal Oxalobacter formigenes [21]. The reaction is carried out by two enzymes, a formyl CoA-transferase (frc) and an oxalyl CoA-decarboxylase (oxc). L. acidophilus NCFM and L. gasseri ATCC33323, but not the closely related L. johnsonii NCC533, show highly conserved homologs to frc (LBA395; LGA130244) and oxc (LBA396; LGA130245). Flanking genes in L. acidophilus NCFM constitute a second acyl CoA-transferase (LBA394) and the ATPase component of an ABC transporter (LBA397) (Fig. 1a, V). Although the genetic organization might indicate an operon-like structure spanning from LBA397 to LBA394, only LBA395 and LBA396 were inducible and expressed upon acid stress [10]. Functional characterization and gene expression analyses in L. acidophilus NCFM demonstrated the capability to degrade oxalate, potentially identifying one probiotic role [10]. Interestingly, both CoA-transferases were grouped into the PFam CoA-transferase family 3, a novel family distinctively different from the two other, previously described families [30]. Based on gene synteny, functional classification, and the slightly higher GC content of 38.64 and 39.53% in L. acidophilus NCFM and L. gasseri ATCC33323 respectively, one might speculate that this region was acquired via horizontal gene transfer (HGT) and then subjected to subsequent deletion events that left only frc and oxc functional. To further investigate the possibility of HGT, we have analyzed dinucleotide frequencies and codon usage for both genes and compared them to the average found in the L. acidophilus NCFM genome. A chi-squared test was used to determine whether the observed forward strand dinucleotide counts for the genes of interest were similar to the expected counts under the assumption that the genes came from the genome of interest. Results clearly indicate a different dinucleotide frequency when compared to the average genome (chi-squared value: 100.6, degrees of freedom = 15, p-value = 1.00E-14). Similarly, a chi-squared test was used to compare the codon usage frequencies for the two genes to the overall genome frequencies. Rare codons (expected values <5) were pooled to create a single category before the test was done. Again, results indicate a different codon usage of both genes (chi-squared value of 224.8, degrees of freedom = 46, and p-value = 2.14E-25). Both the dinucleotide frequencies and codon usage differences support the hypothesis of HGT acquisition.
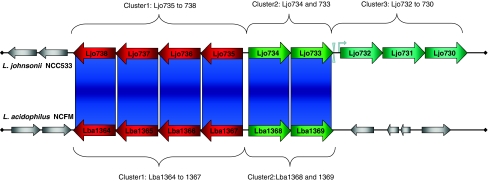
Carbohydrate metabolism is a central anchor of energy generation with living organisms. Transport and metabolism systems are often specific to certain sugars and rely on dedicated enzymes for processing [35]. DBA partially highlighted two divergently oriented gene clusters in L. acidophilus NCFM, LBA1364 to 1367 and LBA1368 to 1369, respectively, likely to be involved in group-specific carbohydrate metabolism (Fig. 1a, XIII and Fig. 2, cluster 1 and 2). The first gene cluster consists of three alpha- and beta-galactosidases and a transcriptional regulator of the ROK family, predicted to be involved in galactose metabolism. Although the three structural genes feature many homologs throughout LAB, it is interesting to note that the transcriptional regulator LBA1367 is much less conserved and only L. johnsonii NCC533 revealed a close homolog (LJO735) within the same gene synteny. The second gene cluster consisted of the cellobiose-specific phosphotransferase component EIIC, celB, (LBA1369) and a transcriptional regulator of the ROK family (LBA1368), similar to LBA1367. CelB is one of eight cellobiose-specific PTS EIIC components identified in L. acidophilus NCFM, responsible for the phosphorylation during translocation of the sugar. Both genes of this cluster show only very limited similarities throughout LAB, again with the exception of L. johnsonii NCC533 (Ljo733 and 734). It is tempting to speculate that the specific transcriptional regulators found for these two gene clusters are involved in modulating strain-specific degradation of complex sugars and moiety uptake. Based on gene synteny at this locus, it might be noteworthy to highlight a third gene cluster found exclusively in L. johnsonii NCC533 (Fig. 2, cluster 3). This L. johnsonii NCC533–specific cluster is located immediately downstream of Ljo733 and consists of three ORFs; two phosphatidyl-serine decarboxylases (Ljo730 and 732) and a predicted permease/antiporter (Ljo731). Detailed analysis of the intergenic region between Ljo732 and 733 revealed the presence of a predicted rho-independent terminator (−16.09 kcal/mol) and a highly conserved promoter-like structure, likely to control expression of the gene cluster independently from the upstream transcriptional regulator Ljo734. In silico analyses using PathwayVoyager [5] and the KEGG database revealed a partial metabolic pathway for aminophospholipids, originating from serine and terminated at phosphatidylethanolamine. Phosphatidylserine decarboxylases (EC4.1.1.65) mediate the conversion of an activated serine (phosphatidyl-l-serine) into phosphatidylethanolamine and CO2. Both decarboxylases identified in this cluster are highly conserved to each other at their respective N and C-termini (75% identity, 86% similarity) and show similarities only to LBA1607 in L. acidophilus NCFM. However, neither gene synteny nor gene content is conserved in L. acidophilus NCFM and LBA1607 could not be placed into the same context. No genes could be identified in L. johnsonii NCC533 that might further convert phosphatidylethanolamine. However, the presence of the predicted permease/antiporter (Ljo731) might point to a different functionality than aminophospholipid metabolism, thus employing an otherwise dead-end metabolic reaction. Decarboxylation reactions have the general potential of raising the internal pH. Hence, the decarboxylation of the activated serine might change the pH, and ethanolamine could subsequently be exported from the cell to avoid accumulation of a dead-end product in the organism. Based on this model, the gene cluster shows a possible involvement in internal pH regulation, unique to L. johnsonii NCC533. The presence of two decarboxylase genes has been reported for other organisms as well and both enzymes have the capability to complement each other [65]. Interestingly, the KEGG database does not currently list the required serine-activating phosphatidyltransferase (EC2.7.8.5) for L. johnsonii NCC533. However, reverse BlastP using a customized database featuring prokaryotic orthologs to the phosphatidyltransferase listed in the KEGG database and the complete proteome of L. johnsonii NCC533 as queries revealed the presence of a predicted glycerophosphate phosphatidyltransferases (Ljo838). Further analyses strengthened the functional classification of Ljo838 by disclosing the presence of CDP-alcohol phosphatidyltransferase domain (PFam 1066) and showing similarities to COG class 558, harboring phosphatidylglycerophosphate synthases. Therefore, Ljo838 might complete this strain-specific pathway, possibly giving L. johnsonii NCC533 the ability to utilize more substrates for internal pH regulation.
Fig. 2.
Schematic representation of a pACT analysis of two homologous genome regions in L. johnsonii NCC533 (upper panel) and L. acidophilus NCFM (lower panel). Genomes are indicated by the black horizontal lines (not to scale). Predicted ORFs are shown as red (cluster 1), green (cluster 2), and cyan (cluster 3) arrows. ORF numbers are indicated within the arrows. Gray arrows represent ORFs with no similarity to the corresponding ORFs at the respective genome location of the other genome. Alignments were created and similarity scores were calculated as described. Blue vertical bars indicate a level of similarity below an e-value of 1e-80. Predicted terminator and promoter structures between ORFs Ljo733 and Ljo732 are indicated by gray inverted triangles and a gray arrow, respectively (color figure online)
Other prominently highlighted group-regions represent specific transport systems for other carbohydrates and amino acids (i.e. an auxin transporter LBA367, an l-lactate permease LBA1768, an ABC-type multidrug exporter LBA1859, and an amino acid permease LBA1902 (Fig. 1a, IV, XXVI, XXVIII, and XXIX)). Considering their similar ecological niches, the intestinal lactobacilli may share a large array of similar transport systems, not found in other LAB, reflecting their adaptation to the intestine.
A region likely involved in pentose uptake and conversion shows an intriguing mosaic structure of a gene cluster conserved in L. acidophilus NCFM and other LAB (red), flanked by two loci conserved only in at least one of the other three lactobacilli of human origin (green) (Fig. 1a, XVI, XVII, and XVIII). The central region (LBA1481 to 1485) encodes for an ABC-type ribose transporter and a ribose kinase, likely activating the sugar moiety for further conversion after uptake [62]. Immediately downstream of this cluster, an ORF was identified likely to encode for a cellulase (LBA1480). Only L. johnsonii NCC533 showed a closely related homolog (LJO1263). This particular cellulase was classified within the glycosyl hydrolase family 5, a widespread group of enzymes that hydrolyze the glycosidic bond between two or more carbohydrates and are known to act on both hexoses (cellulose) and pentoses (xylans) [31]. While it has been shown that L. acidophilus NCFM is able to grow on cellobiose and weakly on xylobiose (van Santen and Lahtinen, personal communication), growth on cellulose has not yet been experimentally validated. Upstream of the central cluster, a predicted l-fucose isomerase was identified (LBA1486) and only L. johnsonii NCC533 revealed a closely related homolog (LJO1264). However, the gene synteny between the two strains is not preserved. Whereas in L. johnsonii NCC533 the cellulase and the isomerase genes are located adjacent to each other and appear to be organized in one operon, the two genes are flanking the sugar ABC transporter (LBA1481–LBA 1485) in L. acidophilus NCFM. More interestingly, l-arabinose isomerase revealed a very similar domain structure to l-fucose isomerases. In conjunction with the other genes present in this cluster, a partial pathway for arabinose uptake and utilization could be proposed. The predicted cellulase might be involved in xylan or arabinan degradation, both molecules are an essential part of plant and mycobacterial cell walls. Arabinose molecules are then transported into the cell by the ABC-type transporter, predicted to be specific for pentoses. l-arabinose can then be converted into l-ribulose by the l-arabinose isomerase and further activated by the ribose kinase. Sa-Nogueira et al. [58] reported the utilization of arabinose by Bacillus subtilis to depend on three enzymes, namely an l-arabinose isomerase, an l-ribulokinase, and an l-ribulose-5-phosphate 4-epimerase. Although the first two enzyme activities were identified in L. acidophilus NCFM, no 4-epimerase was predicted. However, the presence of a transposase (LBA1487) directly adjacent to the isomerase could indicate a previous genome rearrangement resulting in the loss of the epimerase gene. It might be interesting to investigate this predicted partial pathway by providing the missing enzyme activity in trans and analyze the organisms potential capability to utilize xylans or arabinans.
On the other hand, there are a number of genome regions conserved between L. acidophilus NCFM and at least one other member of the LAB database dbLAB that are not, or only weakly conserved in the other three lactobacilli (red). The multiple sugar metabolism (msm) operon (LBA500 to 507) (Fig. 1a, VII) has been extensively analyzed and described previously [14]. A similar operon also involved in sugar uptake and metabolism has been identified and was designated msmII (LBA1437 to 1443) [6] (Fig. 1a, XV). The enzymatic part (sucrose phosphorylase, LBA1437 and alpha galactosidase, LBA1438) and the energy conversion unit of the ABC transporter (ATPase, LBA1439) of the msmII operon are highly conserved in both the lactobacilli of human origin and many other LAB. However, the substrate-specific genes (permease subunits LBA1440 and 1441 and the sugar-binding component LBA1442) are unique to L. acidophilus NCFM and two Streptococcus strains, namely S. mutans and S. pneumoniae. The unique uptake and binding proteins may indicate the possibility of varying substrate specificities for these systems.
Pullulanases are involved in the hydrolysis of (1 − >6)-alpha-D-glucosidic linkages in pullulan, amylopectin, and glycogen. The smallest sugar released by its enzymatic activity is maltose. This enzyme (Lba1710) appears to be unique for L. acidophilus NCFM among the other lactobacilli (Fig. 1a, XXIV). A BlastP search against the non-redundant database revealed further similarities to firmicutes, in particular to bacilli. Based on these results, a phylogenetic tree was constructed using the Cobalt multiple sequence alignment tool and the Fast Minimum Evolution algorithm [50] (data not shown). The closest homologs to Lba1710 were found in Lactobacillus amylolyticus DSM11664 (beer malt and beer wort), Lactobacillus crispatus JV-V01 and Lactobacillus iners DSM 13335 (both are considered part of the normal human microbial flora), whereas bacilli and bifidobacteria exhibited weaker similarities. In particular, pullulanases from bifidobaceria were nearly twice the size than Lba1710–extending the N-terminal part of the protein while maintaining a more conserved C-terminus. Further detailed analyses will be required to examine the differences in functionality between those enzyme groups. A SignalP analysis [22] revealed the presence of a signal peptide with a predicted cleavage site between position 35 and 36 of the deduced amino acid sequence, indicating an extracellular location and thus further supporting the proposed enzymatic activity. The presence of this unique enzyme in L. acidophilus NCFM could reflect an adaptation to the nutritional content of the hosts GI-tract. Amylopectin is a highly branched polymer of glucose found in plants and is one of the two components of starch. Glycogen is a polysaccharide and the principal storage form of glucose in animal cells. Pullulan is a polysaccharide polymer consisting of maltotriose units and is produced from starch by the yeast Aureobasidium pullulans. Because of its physical and biochemical properties, it is increasingly used in food industry as a non-digestible carbohydrate [70]. Hence, all three components are likely to be present in the GI tract as integral parts of a diet. Accordingly, L. acidophilus NCFM might be able to utilize some or all of these complex carbohydrates. However, in vitro growth experiments have failed to show that NCFM can grow on pullulan (O’Flaherty & Klaenhammer, unpublished data).
As reported earlier, L. acidophilus NCFM is auxotrophic for many amino acids, including the branched chain amino acids leucine, isoleucine, and valine [6]. An ABC transport system was identified (Lba1943 to 1946), likely to be specific for uptake of these amino acids (Fig. 1a, XXX). Interestingly, no other Lactobacillus of human origin revealed a system with significant amino acid similarities. Only weak similarities were identified to the periplasmic and ATPase components. The substrate-specific permeases, however, were unique to L. acidophilus NCFM. In contrast, the branched chain amino acid ABC transporter was well conserved throughout LAB, most notably in Lactobacillus delbrueckii ssp. bulgaricus. Both gene synteny and content are highly conserved on amino acid level. Interestingly, L. delbrueckii ssp. bulgaricus features a high overall GC content of 51.37%, and the ABC transporter exhibits an even higher GC content of 52.26%. L. acidophilus NCFM on the other hand belongs to the low GC branch of LAB and has an overall GC content of only 34.71%, with the ABC transporter featuring an only slightly higher GC content of 35.44%. This is also reflected by a DNA alignment of both regions that shows only a few regions to be conserved (data not shown). The observation that this system is, apart from L. delbrueckii, almost exclusively conserved in low to medium GC content LAB, such as E. faecalis, S. pyogenes, S. thermophilus, S. agalactiae, O. oeni, or L. mesenteroides, might indicate an ancient genetic transfer to L. delbrueckii, with a subsequent GC and codon adaptation, while preserving the secondary structure.
Lactobacillus johnsonii NCC533
As described previously, L. johnsonii NCC533 and L. gasseri ATCC33323 share a significant amount of genetic information [55] and are considered closely related [24]. Despite this high level of similarity, a number of regions were identified unique to L. johnsonii NCC533 when compared to dbLB and dbLAB. Most prominent were the two prophages of L. johnsonii NCC533 (Lj965 ranging from Ljo288 to Ljo330 and Lj928, ranging from Ljo1418 to Ljo1465 [67]) (Fig. 1b, VIII and XXII). Both elements were recognized by BlastP and DBA analyses, since some phage genes displayed similarities to LAB phages unrelated to those of lactobacilli (data not shown).
Furthermore, a putative arsenite-efflux transport system was identified to be unique to L. johnsonii NCC533 among LAB (Fig. 1b, VII). This system is likely to be organized in an operon and consists of an operon repressor ArsR (Ljo230), an arsenite efflux transporter ArsB (Ljo231), and an arsenate reductase ArsC (Ljo232). This operon might provide an efficient detoxification system for arsenate by forming arsenite which, subsequently, is exported from the cell. This might reflect an interesting lifestyle adaption to the increasing concentrations of these substances in the environment (i.e. use of metal-containing pesticides) which have been shown to accumulate in higher organisms.
An L. johnsonii NCC533 unique region predicted to be involved in exopolysaccharide biosynthesis was also identified (Ljo1707 to Ljo1711) (Fig. 1b, XXV). In this small gene cluster, four genes (Ljo1707 to Ljo1710) showed similarities to glycosyl transferases of family 8. This family includes enzymes that transfer sugar residues to donor molecules. Based on genome context analysis, the gene product of Ljo1711, a 3039 aa protein with an LPXTG membrane anchor and a 10 amino acid repeat structure, might act as donor protein.
A region with strain-specific characteristics, analyzed previously [36], encodes an EPS cluster. Highly conserved genes (epsA, B, C, D, E, J, and I) flank a unique core consisting of sugar transferases and polymerases (Fig. 1b, XVII).
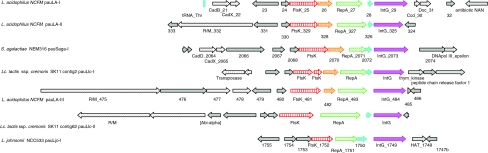
Lastly, a genome region (Ljo1748 to Ljo1755) encoding a potential autonomous unit (PAU) has been identified (Fig. 1b, XXVII), previously believed to be unique to L. acidophilus NCFM [6]. PAUs in L. acidophilus NCFM resembled elements from both bacteriophage and plasmids. Analysis of pauLjo-I region in L. johnsonii NCC533 revealed a striking similarity in both functional classification and gene synteny to pauLa-I, pauLA-II, and pauLa-III of L. acidophilus NCFM (Fig. 3). Consequently, this region was designated as pauLjo-I. Interestingly, the amino acid similarities of the conserved core genes (ftsK, repA and intG) were very low to the respective L. acidophilus NCFM elements. Further analyses (gene ortholog neighborhoods) using the Integrated Microbial Genomes system (IMG) provided by the DOE Joint Genome Institute (http://img.jgi.doe.gov/pub/main.cgi) revealed the presence of three more genetic elements with significant similarities to the core region of PAUs. These elements comprise of pauSaga-I (ORFs2064 to 2075) found in Streptococcus agalactiae NEM316, and pauLlc-I (scaffold18, ORFs1220 to 1228, of the current draft phase genome) and pauLlc-II (scaffold6, ORFs 3359 to 3364, of the current draft phase genome) found in Lactococcus lactis ssp. cremoris SK11 (Fig. 3). Gene synteny and functional classification remain highly conserved and little variation was observed in the presence or absence of the two smaller hypothetical core genes. Like L. johnsonii NCC533, the amino acid similarities of the classified core elements remain very weak and do not cluster consistently among the three proteins (Supplemental Fig. 1). This suggests either a differentiation of a common ancestor or distinctive different roots of these elements. Adjacent of the core, PAUs displayed genes that might stabilize or maintain the respective PAU in the chromosome. In case of L. acidophilus NCFM, the death on curing (doc) system and several restriction/modification (R/M) systems were identified in close proximity to the core genes [6]. Similarly, pauLlc-II exhibited the presence of an R/M system, an adenine specific DNA methylase, and a putative abortive infection (abi) system alpha, upstream of FtsK. Interestingly, pauSaga-I exhibited a cadmium export system (CadD and CadX), similar to the one determined in pauLa-I (CadB and CadX). This system consists of a cadmium exporter (CadD family) and a regulatory protein (CadX). Amino acid alignments of CadD and CadB revealed a 60% identity (75% similarity), indicating similar transporter functionalities for both genes. Similarly, both regulatory proteins exhibited 44% sequence identity (67% similarity).
Fig. 3.
Potential autonomous units identified in LAB. Alignment of previously described and newly identified PAUs in L. acidophilus NCFM, L. johnsonii NCC533, Lactococcus lactis ssp. cremoris SK11, and Streptococcus agalactiae NEM316. Elements of the core region are shown as colored arrows: DNA segregation ATPase FtsK, red arrow; replication protein RepA, green arrow; Integrase IntG, pink arrow; conserved hypothetical ORFs, orange and cyan arrows, respectively. Flanking ORFs with predicted functions are displayed in light gray arrows. Different functional classes are indicated by distinct pattern. Adjacent ORFs with no functional characterization are indicated by solid gray arrows. Predicted tRNAs are displayed as vertical cyan boxes. Alignments are centered on the start position of IntG and are drawn to scale (color figure online)
With the exceptions of pauLjo-I and pauLlc-I, every investigated PAU harbored genes adjacent to its core that might promote stabilization of the PAU in the chromosome. The discovery of seven different PAUs in four different organisms clearly suggests that PAUs are not a curiosity, but represent a new class of potentially mobile elements that might be further classified into different distinct families. However, no functional analyses have been performed to date and the in vivo activities remain to be investigated.
DBA analysis highlighted a large number of genes found to be shared between L. acidophilus NCFM, L. gasseri ATCC33323, and L. johnsonii NCC533 but which are not highly conserved within other LAB (green color shading). As described earlier, a significant portion of these regions are comprised of cell surface proteins mediating host–cell interactions (Ljo46, 47, 48, 484, and 1128), transport systems (Ljo733 and Ljo734), and metabolism islands (Ljo730, 731, 732 and Ljo1263 to Ljo1268) (Fig. 1b, II, X, XII, XIX, and XXI).
A genome region located at ~60 kbp (Ljo59 to Ljo63) was identified to harbor a partial exopolysaccharide (EPS) biosynthesis and transport cluster (Fig. 1b, III). This cluster was highly conserved to L. gasseri ATCC33323 and L. acidophilus NCFM and to a significantly lower degree in similarity and synteny to Lactobacillus bulgaricus. Interestingly, Ljo60 to Ljo62 show similarities to glycosyl transferases of family 8. Despite their chromosomal distance, perhaps this gene cluster and the L. johnsonii NCC533–specific cluster (Ljo1707 to Ljo1711) described earlier may act synergistically by providing different moieties to the LPS structure.
Directly adjacent to the LPS cluster, a smaller set of genes was identified, coding for a bile salt hydrolase (Ljo56) and two bile salt transporters (Ljo57 and Ljo58) (Fig. 1b, II). Not surprisingly, the bile salt hydrolase is highly conserved in L. acidophilus NCFM,L. gasseri ATCC33323, and L. plantarum WCFS1, likely reflecting their common adaptation to the intestinal environment. Other LAB, in particular bifidobacteriae, show only a limited degree of similarity to Ljo56. In contrast, both bile salt transporters are only conserved in L. gasseri ATCC33323 and L. johnsonii NCC533. It is not clear if bile salt hydrolases have a wide substrate spectrum, allowing the deconjugation of many different bile salts. In contrast, it is more likely that bile salt transporters might be dedicated to certain classes of bile salts.
Despite the exceptional degree of similarity to L. gasseri ATCC33323, several genome regions were identified that were uniquely shared between L. johnsonii NCC533 and other LAB, but not present in either three of the other Lactobacilli (red color shading).
Interestingly, a genome region close to the origin of replication was originally annotated to harbor an enzyme involved in thiamine biosynthesis (Ljo20, 21, 22) (Fig. 1b, I). However, when analyzed in context, these ORFs are likely to encode for different functionalities. Ljo20 was annotated as a transcriptional regulator and is likely to control expression of Ljo21 and Ljo22. The gene product of Ljo21 shows significant similarities to COG0476. This cluster represents dinucleotide-utilizing enzymes involved in molybdopterin and thiamine biosynthesis, as indicated by the original annotation. The presence of Ljo22, a predicted efflux transporter, rendered the initial functional classification of Ljo21 questionable. Further analyses of Ljo 21 and Ljo22 revealed similarities to the microcin C51 production gene mccB in Escherichia coli [23] and the microcin C7 secretion protein mccC in Heliobacter pylori [32], respectively. Interestingly, the identified similarity of MccB to COG0476 has been described for both MccB of microcin C7 [26] and C51 [23]. It has been proposed that the adenylation conferred by MccB plays a role in the substitution of the C-terminus of the heptapetide by AMP [23] and is not involved in molybdopterin or thiamine biosynthesis. MccC has been shown to provide partial immunity, complemented by MccE. However, in L. johnsonii NCC533, the microcin operon appears to be only partially conserved, as mccA, the gene coding for the heptapeptide moiety, mccD, and mccE (immunity) are absent from the operon. Furthermore, the presence of the divergently oriented transcriptional regulator Ljo20 does not comply with the reported operon structure of microcins. The low GC content of 25.78% found for this region might indicate horizontal gene transfer (HGT) as source of acquisition. This is further strengthened by the absence of this gene cluster in all other lactobacilli present in dbLB. Analysis of dbLAB revealed homologous proteins only in Streptococcus thermophilus CNRZ1066 (CP000024: ORF 1944, mccB and 1943, pmrB). A similarly low GC content also indicates HGT as possible source of acquisition; however, the predicted transcriptional regulator (ORF1947) is separated from proposed mccB and pmrB by two predicted transposase genes (ORFs1945 and 1946). It might be speculated that this genome region represents an acquired resistance mechanism to microcin-like substances giving it a strain-specific competitive advantage. Alternatively, the gene cluster could resemble the remnants of a bacteriocin-producing operon, similar in structure to microcins.
An uptake system found in L. johnsonii NCC533, but not any other Lactobacillus, consisted of a PTS sugar transporter (predicted mannose specificity) and a 2-CRS sensor (Ljo1652 to 1660) (Fig. 1b, XXIII). Ljo1653 to Ljo1656 represent PTS components A to D. These genes are relatively highly conserved in Enterococcus faecalis, Lactobacillus casei, Streptococcus mutans, and Leuconostoc mesenteroides, with decreasing levels of similarity from PTS EIID to EIIA. Located in the same operon-like gene cluster, Ljo1652 was found to share similarities to membrane proteins and permeases, potentially acting as an accessory protein to the PTS system. Homologous genes were identified in L. casei, S. mutans, and L. mesenteroides. Of the other lactobacilli in dbLB, only L. acidophilus NCFM showed very weak levels of similarity to this gene cluster (except for Ljo1652), indicating only a general functional similarity that is unlikely to cover the same substrate spectrum. This uptake system might give L. johnsonii NCC533 the advantage of using additional sugars as carbon and energy sources in shared ecosystems. Genes likely to be involved in gene regulation were oriented in the opposite direction (Ljo1657 to Ljo1660) (Fig. 1b, XXIII). Ljo1658 and 1659 encoded for a predicted 2 CRS, with similarities of the histidine kinase indicating sugar specificity (COG2851). Interestingly, the flanking genes Ljo1657 and 1660 revealed similarities to periplasmatic sugar-binding protein. In addition, Ljo1657 revealed conserved domains indicating a possible role as primary receptor for sugar transport and transcriptional repressor (PFam00532 and COG1609). These four genes do not feature homologs in the other three lactobacilli and the closest relatives were found in streptococci.
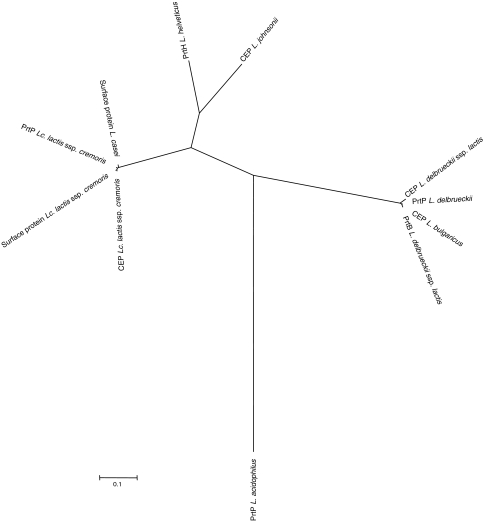
Lastly, a gene likely to encode a cell-envelop-associated proteinase was identified in L. johnsonii NCC533 (Ljo1840) (Fig. 1b, XXXI). Cell-envelop proteinases (CEP) facilitate the proteolysis of casein in lactic acid bacteria [53, 61] and are often essential for the growth in milk [25]. Furthermore, CEPs were also associated with periodontal disease in Bacteroides forsythes [64]. Most CEPs require the presence of chaperones for maturation (lactococcal PrtP [69] and PrtH from Lactobacillus helveticus [53]), whereas others are capable of an autocatalytic process that substitutes the chaperone protein PrtM with an intramolecular chaperone (PrtB of L. delbrueckii spp. bulgaricus [25]). The identified CEP of L. johnsonii NCC533 is a unique gene among lactobacilli of human origins. Only L. acidophilus NCFM harbored a weakly conserved homolog (Lba1512). However, as described earlier, the homologs are present and conserved in other LAB, further implicating a key role in extracellular protein processing. Most notably, several lactobacilli (L. casei, L. bulgaricus) and L. lactis ssp. cremoris showed conserved homologs of PrtH. A phylogenetic tree featuring previously described and predicted cell-envelop proteinases from several lactobacilli and lactococci revealed a clear clustering of PrtH from L. helveticus and CEP of L. johnsonii NCC533 (Fig. 4) likely positioning Ljo1840 into the same functional group (Group I) as PrtH, as was noted previously [55]. This is further supported by the presence of the divergently oriented maturation protein PrtM (Ljo1841). However, two of the five reported substrate-binding regions differ significantly in amino acid composition in Ljo1840, indicating a different specificity than previously reported CEPs. Interestingly, PrtP of L. acidophilus NCFM is the phylogenetically most different protein included in the analysis, although many conserved regions remain intact. In contrast to Ljo1840, no adjacent maturation protein could be identified; the closest chaperone-like protein was found 70 kb downstream (PrtM, Lba1588) and it is unknown whether PrtM is involved in PrtH processing.
Fig. 4.
Phylogenetic tree of cell-envelop proteinases (CEP) of selected LAB. Deduced amino acid sequences of CEPs identified in other LAB were aligned using ClustalW [42], and the phylogenetic tree was calculated and visualized using Mega3.1 [40]. The evolutionary history was inferred using the neighbor-joining method [59]. The evolutionary distances were computed using the Poisson correction method [73] and are in the units of the number of amino acid substitutions per site. Protein names and respective organisms are indicated on the phylogenetic tree (color figure online)
Lactobacillus gasseri ATCC33323
Originally isolated from the human GI tract, L. gasseri ATCC33323 occupies a similar ecological niche as L. acidophilus NCFM and L. plantarum WCFS1 [8, 43]. However, overall genome synteny and similarity were found to be significantly more conserved between L. gasseri ATCC33323 and L. johnsonii NCC533, than between L. gasseri ATCC33323 and the other two lactobacilli (Fig. 5). More than 50% of the predicted ORFs in L. gasseri ATCC33323 share similarities to L. johnsonii NCC533 ORFs at a level of 1e-100 and below (Fig. 6a). This likeness is not surprising as phylogenetically these two species are the most closely related of the members of the L. acidophilus NCFM complex. However, it is noted that the two species are often isolated from distinctly different hosts: L. gasseri ATCC33323 from humans and L. johnsonii NCC533 from the crop of chickens [2, 28, 41]. These are distinctive environments with unique metabolic challenges. The L. johnsonii NCC533 strain was sequenced and used in this analysis, however, was isolated from a human [55].
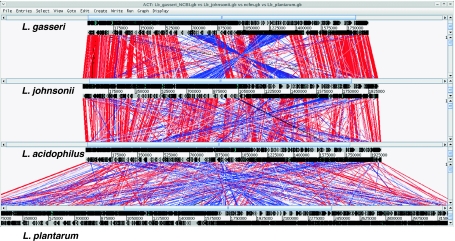
Fig. 5.
Protein similarity and genome synteny of four Lactobacillus genomes pACT were used to simultaneously analyze four complete genomes on amino acid level. Double lines featuring pointed boxes indicate predicted ORFs in their respective orientation. Red and blue lines in between the genomes represent amino acid similarities between ORFs below the selected threshold. Red lines show direct alignments, whereas blues lines depict inversions. The threshold was set to 1e-70 and only hits with a more significant e-value are indicated (reprinted from [37]) (color figure online)
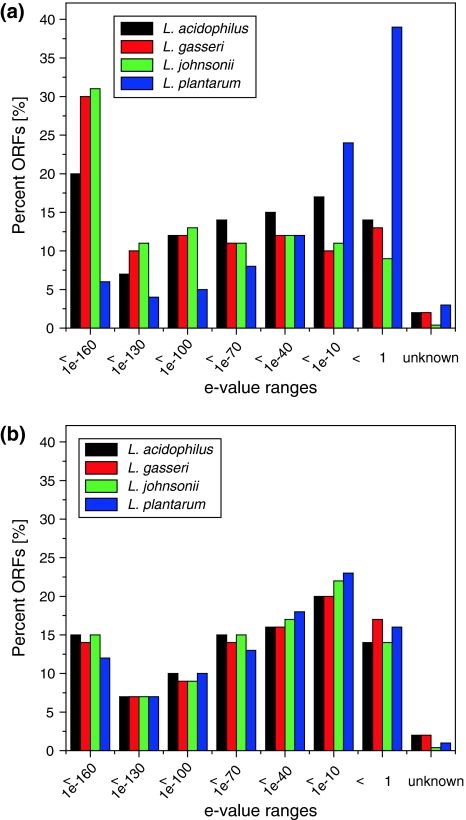
Fig. 6.
Differences in e-value distribution between the four lactobacilli of human origin when compared to the two custom databases dbLB and dbLAB. Based on the best BlastP hit, similarities were grouped within the assigned e-value ranges. Solid bars represent results for the custom Lactobacillus database dbLB (a) and the custom LAB database dbLAB (b). E-value ranges correspond to trust level intervals described in Table 2 (color figure online)
A significant percentage of ORFs are very similar between L. gasseri ATCC33323, L. johnsonii NCC533, and L. acidophilus NCFM and only to a lesser degree conserved to other LAB (green). Among the most dominant features identified are exoproteins like mucus- and fibrinogen-binding proteins (Lga130040, 130041, 130042, 130139, 130140, 131623, and 131641), as described earlier (Fig. 1c, I, III, XXII, and XXIII). One of the largest proteins in the genome, a specific mucus-binding protein (Lga130041), was found in L. johnsonii NCC533 (Ljo0047) and L. acidophilus NCFM (Lba1019 and 1020), but was absent in L. plantarum WCFS1. Homologs in L. gasseri ATCC33323 and L. johnsonii NCC533 were located close to the origin of replication, whereas the corresponding ORF in L. acidophilus NCFM was found close to the terminus of replication. No evidence of apparent genome rearrangements was observed. Quite contrary, the region close to the origin shows a relatively high degree of synteny between all three genomes (Fig. 5).
Adjacent to this surface protein, a bile salt export system was identified (Fig. 1 C, II). This system consists of a bile salt hydrolase and two permeases. As described for the corresponding system in L. johnsonii NCC533, closely related genes to this export system can only be found in these two strains. Interestingly, one of the permease genes (Lga130054) appears to be truncated by a premature stop-codon and its integrity remains to be verified.
Furthermore, a tagatose-diphosphate aldolase was identified (Lga130142), present only in L. johnsonii NCC533, L. gasseri ATCC33323 and to a lesser degree, in certain LAB (Fig. 1c, III). This enzyme is able to convert tagatose-1,6-diphospate into glycerone phosphate and glyceraldehyde-3-phosphate. Both metabolites can be directly fed into the glycolysis for further conversions. Although all genes necessary for lactose uptake and conversion are present in the L. gasseri ATCC33323 genome (data not shown), the presence of a sugar phosphate permease (Lga130141) directly adjacent to the aldolase could indicate an additional tagatose-specific sugar uptake and metabolism system, not present in L. acidophilus NCFM and L. plantarum WCFS1. Tagatose is widely used as a commercial low-calorie sweetener in soft drinks and fruit juices but is otherwise present in only small amounts in dairy products. The increasing consumption of low-calorie foods and the resulting rising presence of tagatose in the GIT might represent a new source of energy, promoting the growth of L. gasseri ATCC33323 and L. johnsonii NCC533.
A gene-pair which is conserved in L. gasseri ATCC33323 and L. acidophilus NCFM, but not in any other LAB consists of an oxalyl CoA-decarboxylase (Lga130245) and a formyl CoA-transferase (Lga130244) (Fig. 1c, IV). As described for L. acidophilus NCFM above, this system is likely involved in oxalic acid degradation.
The synthesis and modification of bacterial cell walls and the incorporated teichoic acids require the coordinated interaction of many different enzyme groups. The most predominant group is composed of glycosyltransferases, catalyzing the transfer of sugar moieties from activated donor molecules to specific acceptor molecules. A group of three genes (Lga131543 to 131545), presumably organized in an operon-like structure, has been identified in L. gasseri ATCC33323 that is likely involved in teichoic acid biosynthesis (Fig. 1c, XIX). All three genes were predicted to be glycosyltransferases and classified into separate families. Lga131544 represents a glycosyltransferase of group 1 which is highly conserved in L. acidophilus NCFM (Lba520), L. johnsonii NCC533 (Ljo1736), and many other LAB. However, close homologs cannot be found in L. plantarum WCFS1. A glycosyl transferase of the WecB/TagA/CpsF family (Lga131543) might be involved in the synthesis of cell wall polymers and was highly conserved in many LAB, including the four lactobacilli. Notably, the last gene of this operon-like gene cluster, Lga131543, a glycosyltransferase of group 2 that transfers sugar moieties to teichoic acid, is unique to L. gasseri ATCC33323 and L. johnsonii NCC533. A comparison to other LAB revealed that this region appears to be highly variable. L. acidophilus NCFM only harbors two genes (Lba 519 and 520), L. johnsonii NCC533 and L. gasseri ATCC33323 each feature a three gene cluster, whereas the region in L. bulgaricus consists of five genes (ORFs 111644 to 111648). One could speculate that the components of the cell wall synthesized by the proteins encoded in this operon harbor a conserved core that then features strain-specific entities, possibly altering cell surface properties and consequently, immunological responses.
Despite the predominant similarity to L. johnsonii NCC533, several genome regions were revealed in L. gasseri ATCC33323 that exhibited more significant similarities to other LAB (red).
Two loci, separated by ~40 kb, each harbored a carbohydrate transport system (Fig. 1c, V and VI). The first region (Lga130338 to Lga130341) consisted of four genes, composing a lactose-specific ABC transporter. LacE (Lga130340), representing the sugar-specific permease component, featured only very weak homologs in the other three lactobacilli. Similarly, LacG (Lga130341), a 6-phospho-galactosidase, was only weakly conserved in this dataset, with homologs displaying only certain conserved domains. However, highly conserved homologs were identified in several streptococcal strains. The second locus (Lga130393 to 130395) also represents an ABC transporter. However, the permease (Lga130395) and sugar-specific hydrolase (Lga130396) feature relatively close homologs in both databases. Interestingly, BglX (Lga130394) a beta-glucosidase and the first structural gene of this operon-like structure, cannot be found in the three other lactobacilli but features close homologs in several E. faecalis and other LAB strains. Glycoside hydrolases of this specific family are known to accept larger carbohydrates as substrates and release beta-D-glucosides. In E. coli BglX homologs were found to be located in the periplasm or the cytoplasm [71], and the absence of a signal peptide sequence in Lga130394 could indicate a cytoplasmic location. Substrate specificities for this predicted carbohydrate transporter remain unresolved, and the presence of a BglX homolog might point to a different, more complex substrate for Lga130395 than glucose or maltose.
Another sugar metabolism and uptake system is located at ~500 kb (Fig. 1c, VII). It mainly consists of two PTS systems, likely to be specific for galactitol and cellobiose, respectively and an adjacent operon featuring both subunits of a galactose isomerase. A PTS system consisting of the three genes gatA, gatB, and gatC (Lga130491 to 130494) is likely to be specific for galactose uptake. In succession, the adjacent isomerase operon (Lga130487 to 130489), induced by the presence of galactose, may catalyze the conversion of D-galactose 6-phosphate to D-tagatose and 6-phosphate in the tagatose 6-phosphate pathway of galactose catabolism. Both systems are weakly conserved or not present in the other three Lactobacillus strains, but can be readily found in many other LAB. The second PTS system identified in this genome region (Lga130496 to 130499) is likely to be specific for lactose or cellobiose. Similar to the other PTS system, significant similarities can be found almost exclusively outside the Lactobacillus group.
Centered on a phage-related integrase (Lga130904), a type I restriction/modification (R/M) system (Lga130902 to 130906) was identified via DBA, not found in this form in the other three lactobacilli of human origin (Fig. 1c, X). The main function of R/M system lies in the protection of the bacterial organism against introduced foreign DNA. The foreign DNA will be degraded by endonucleolytic cleavage, whereas the hosts’ DNA is protected by a system-specific pattern of modifications. The R/M system consists of three different complexes: a DNA methylase (family M), a target recognition domain (family S), and a restriction unit (R), present in varying quantities. Here, one N-6 adenine-specific DNA methylase (Lga130902) mediated the methylation of specific DNA sequences. Two target recognition peptides (Lga130903 and 130905) will then interact with either the DNA methylase to protect the host DNA or with the restriction enzyme (Lga130906) to degrade unprotected DNA. Interestingly, a DEAD/H box helicase (Lga130900) was also present in the gene cluster and showed close homologs only within the LAB group. This helicase might play a role in unwinding the DNA prior to its methylation. Interestingly, this gene complex features a sharp decline in GC content (29.9%), possibly indicating an acquisition via horizontal gene transfer. Alternatively, the region could represent mobile DNA or a remnant thereof, as indicated by the presence of the integrase gene. It might be interesting to investigate the range of protection this system may provide against foreign DNA, including DNA introduced by bacteriophage or electroporation.
A 5′-nucleotidase (Lga131078) was identified that is highly conserved in LAB, but not found in L. acidophilus NCFM, L. johnsonii NCC533, or L. plantarum WCFS1 (Fig. 1c, XIII). This gene is partly annotated as a secreted or peptidoglycan-bound enzyme. There it might function by degrading free DNA molecules and providing nucleotides for readily uptake into the cell.
Threonine dehydratase (Lga131436) usually mediates the conversion of l-threonine to 2-oxobutanoate and NH3 (Fig. 1c, XVI). Although L. gasseri ATCC33323 was predicted to harbor the complete pathway to convert l-aspartate into l-threonine (data not shown), the subsequent pathway to further utilize 2-oxobutanoate in the valine, leucine, and isoleucine metabolism appears to be absent. However, this enzyme is also able to act as an l-serine ammonia-lyase, mediating the conversion from l-serine to pyruvate and NH3. Lga131436 is not conserved in any of the remaining Lactobacillus strains of human origin and only E. faecalis strain V583 revealed a protein with some degree of similarity. Interestingly, Lactobacillus strains do harbor an l-serine dehydratase, mediating the same reaction. This enzyme consists of two subunits and does not show any sequence similarities to Lga131436. Furthermore, the presence of an adjacent permease (Lga131437) and a transcriptional regulator (Lga31438) could indicate a different metabolic role for this operon-like structure for L. gasseri ATCC33323. One could speculate that the permease upon induced expression would transport L-serine into the cell, where it is converted into pyruvate by the serine lyase. Pyruvate is then further metabolized to create ATP. This model would suggest a unique energy gaining mechanism for L. gasseri ATCC33323 dependent on the presence of free extracellular serine.
A second restriction/modification system (Lga133002 to 131482) largely unique to L. gasseri ATCC33323 was identified at ~1.45 Mbp (Fig. 1c, XVII). In contrast to the previously described type I system, this one is likely to represent a type III R/M system, consisting of a chromosome aggregation ATPase (Lga131480), two DNA methylases (Lga131477 and 131478), a type III restriction endonuclease (Lga131476), and a DNA helicase (Lga131474). This system does not feature any homologs in the other three lactobacilli, and it might provide a unique protection for L. gasseri ATCC33323 on the nucleotide level. Although single components are conserved in other LAB, most notably Pediococcus pentosaceus and Lactobacillus brevis, no organism represented in either database appears to harbor a complete homologous protection system.
Adjacent to the unique R/M system, a high-GC region harbors a phage remnant (the terminase, structural module, and lysis module are partially present), oriented against the main coding direction (Fig. 1c, XVII). Interestingly, this phage remnant also appears to feature a DNA helicase which additionally exhibits a type III endonuclease domain (Lga131490) and a DNA methylase (Lga131492). It is unknown whether this system might interact with other R/M systems or if it represents a phage-specific system.
One of the most unique features of L. gasseri ATCC33323 is the presence of a tandem phage, exactly duplicated in the genome. These two phages (Lga130573 to 130635 and Lga130636 to 130698), located at ~600 kbp, appear to be genetically complete (Fig. 1c, IX). Both phages are identical on nucleotide level and are directly adjacent to each other, with no other intermediate genes present (publication pending). At this point, the occurrence of a tandem phage represents a novel genome structure, and further experiments are required to investigate the functionality of these phages and the impact of the tandem organization on their life cycle.
A second L. gasseri ATCC33323-specific genome region was identified close to the terminus of DNA replication. A set of ORFs (Lga130942 to 130947) is comprised mostly of mucus-binding proteins, which could indicate strain-specific cell-binding or adhesion properties (Fig. 1c, XI).
Lactobacillus plantarum WCFS1
L. plantarum WCFS1 is by far the largest Lactobacillus genome. With more than 3050 predicted ORFs, it is approximately 50% larger than the other three genomes. This is, in part, reflected by the large number of L. plantarum WCFS1-specific genes (Fig. 6a), when compared to genes similar to other LAB in dbLAB (Fig. 6b). More than 60% of the ORFeome showed BlastP hits at 1e-10 and above. However, less than 15% of its genome shares similarities to the other three lactobacilli at a level of 1e-100 or below. In contrast, approximately 40% of the predicted ORFs of L. acidophilus NCFM reside within this similarity range. As previously pointed out by Boekhorst et al. [15], L. plantarum WCFS1 does not appear to be significantly related to either one of the other lactobacilli and in particular to L. johnsonii NCC533 [15]. Figure 5 further illustrates these differences by highlighting the complete lack of genome synteny found between L. plantarum WCFS1 and the other three genomes.
Not surprisingly, most of the results obtained through DBA analysis represent genome regions conserved in L. plantarum WCFS1 and other LAB, but which are not present in the other three lactobacilli analyzed (red). These include the previously described nonribosomal peptide synthesis module (Lp0578 to 584) [39] which could not be identified within the other three lactobacilli but reveals close homologs in Bacillus subtilis and other LAB (Fig. 1d, IV). Interestingly, this gene cluster displays a significantly lower average GC content (35.9%) than the overall genome (45.2%), possibly indicating gene acquisition by horizontal gene transfer.
The respiratory nitrate reductase (Lp1498 to 1501) consists of three subunits (NarG, NarH, and NarI) and reduces nitrate to nitrite under anaerobic conditions [39] (Fig. 1d, VIII). This forms a redox loop that in turn aids in generating the proton motive force of the organism. Additionally, a chaperone (NarJ) might be present to aid in protein maturing and assembly. This complex was not found in the other lactobacilli and comparison to other LAB revealed an interesting pattern of similarities for the different subunits. The alpha and beta chains NarG (Lp1498) and NarH (Lp1499), respectively, are highly conserved in Bacillus subtilis ssp. subtilis 168, but do not share homologs in other LAB strains. In contrast, the gamma chain NarI (Lp1502) and the chaperone NarH (Lp1501) are only weakly conserved in B. subtilis 168, whereas the gene synteny is still maintained. Although no significant changes in GC content could be observed for this gene cluster (47.6%), two low GC spikes upstream of Lp1482 and downstream of Lp1503 identify potential hotspots for genome rearrangements. Interestingly, the genes enclosed by the two spikes comprise not only those of the narGHJI cluster, but also those of the molybdopterin biosynthesis cluster moeB (Lp1496), moaB (Lp1495), moeA (Lp1494), mobA (Lp1493), moaC (Lp1492), and mobA (Lp1491). This cluster, divergently oriented to the narGHJI operon, could provide the co-factor required for the nitrate reductase alpha chain, NarG. Notably, adjacent to the proximal low GC spike, the nitrite extrusion protein NarK (Lp1481) and the molybdopterin biosynthesis proteins MoaA (Lp1480), MoaD (Lp1479), and MoaE (Lp1478) were identified, further strengthening the hypothesis of an ancient genome insertion (see also http://www.cmbi.ru.nl/plantarum/supplementary/supplementary_text.html#molyb for more information on this genome region).
Another low-GC region (Lp3129 to 3138) featuring a sugar uptake (specific for glucose or maltose) and metabolism system was identified, which was not present in the other three lactobacillus strains (Fig. 1d, XIV). However, it is likely that similar systems for these essential sugars are widespread throughout all LABs. The previously described low-GC sugar metabolism island [39] (genome position ~3.1 to ~3.25 Mbp) also shares more similarities to other LAB than to the other three lactobacilli (Fig. 1d, XV).
Despite the low level of protein similarities to the other lactobacilli of human origin, some genome regions were identified that are shared between the four lactobacilli (green). A fumarate reductase (Lp55), predicted to mediate the conversion from fumarate to succinate in the citrate cycles, was highly conserved in L. gasseri ATCC33323 (Lga 130908) (Fig. 1d, I). Other LAB only share the N-terminal part of this enzyme. FMN reductases are members of the flavoprotein clan, whose protein families have arisen from a single evolutionary origin. Interestingly, a second predicted fumarate reductase (Lp952) was also identified, sharing extensive similarities with the previously described ORF Lp55 in L. plantarum WCFS1, FrdA (Lga130908) from L. gasseri ATCC33323, and other homologs within the Lactobacillus group (Fig. 1d, VI). Because different functionalities are so closely related, the exact substrate specificity might be difficult to predict, and further in vitro characterization is required to determine the true function of these proteins.
A predicted ATPase (Lp1520) that might be related to the helicase subunit of the Holliday junction resolvase is highly conserved in all four lactobacilli, but only to a much lesser extent in other LAB (Fig. 1d, IX). Whether this enzyme will act as a resolvase on unresolved Holliday junctions or is otherwise involved in DNA repair or modification remains to be verified.
Finally, a cation antiporter (Lp2674) was found to be highly conserved only within the four lactobacilli (Fig. 1d, XIII). Na+/H+ antiporters of the NhaP-type are important for maintaining the pH of metabolizing cells and may promote survival during gastro-intestinal passage. Interestingly, an efflux permease likely to be specific for arabinose (Lp2675) was identified immediately upstream of the antiporter. These proteins are also partially annotated as an H + antiporter, possibly indicating a functional link between the two proteins and providing an alternative model. One could speculate that the import of arabinose, at the cost of exporting protons, might lead to changes in the internal pH. These would then be subsequently compensated by the cation antiporter, thus maintaining a physiological state within the cell.
Conclusions
Analyzing genome contents of complete bacterial genomes and identifying key genetic elements for further in vitro characterization represents one of the more challenging tasks of genome research today. Here we employ a new approach of comparing genomes and their predicted ORFeomes to each other, and to groups of related organisms, using differential Blast analysis. Due to the nature of DBA, careful selection of the content of both customized databases is vital for meaningful results. This also implies that DBA should not be used to randomly generate data, but to answer specific questions of interest. These specific questions will then in turn determine the content of each database. The model organism L. acidophilus NCFM was initially used to evaluate the significance of DBA results. L. acidophilus NCFM has been extensively analyzed using more traditional methods and many genetic elements have been identified and subsequently selected for in vitro characterization [3, 9–11, 14, 16]. Many of these genetic elements were chosen to facilitate the understanding of the genetic basis of bacteria considered probiotic and promote their subsequent optimization for industrial use. The significance of DBA was clearly demonstrated by the fact that the majority of these genome regions was independently re-discovered by DBA. This might also imply that other genetic targets tagged by DBA are likely to be of considerable interest for further in vitro characterization. Because DBA is not dependent on existing genome annotation, highlighted targets comprise both classified and uncharacterized ORFs. Since genome annotation still considers up to 40% of the ORFeome as functionally unclassified or conserved hypothetical genes, DBA provides a valuable tool to distinguish among a plethora of options and rationally select unique uncharted regions for future research (Fig. 1a–d, regions highlighted as “Unknown functionality”).
Furthermore, the analytical combination of BlastP, DBA, and genome synteny analyses led to the discovery of novel genetic elements. In particular, the distribution of the previously described potential autonomous regions has been significantly increased. Apart from the suggestion of distinct PAU families, their presence has been substantiated and they are likely to represent a new group of mobile genetic elements.
The four lactobacilli analyzed here clearly share different degrees of similarity and genome synteny in the order L. gasseri ATCC33323 ≤ L. johnsonii NCC533 < L. acidophilus NCFM ≪ L. plantarum WCFS1. DBA was most successful on the more closely related strains, whereas L. plantarum WCFS1 revealed a predominantly unique genome content. Genetic regions shared exclusively within the Lactobacillus group were identified and indicated that the common ecological niche led to the acquisition of similar functions (i.e. surface, mucus-, and fibrinogen-binding proteins, drug resistance systems, degradation of oxalic acid, transport and metabolism systems for simple and complex carbohydrates, amino acids, and cations, bile salt hydrolases, and proteins involved in group-specific cell wall synthesis/modification). Whereas, on the other hand, many specific traits were also identified suggesting different requirements based on other, previously occupied niches. In particular, the analyses revealed group-specific genome regions in all four genomes that may reflect lifestyle adaptations to ecological niches and provide distinct defense and protection mechanisms (i.e. in L. acidophilus NCFM: an arginase as part of a pH mediating mechanism, a potential proton motif force generator based on l-alanine, the substrate-specific components of the msmII operon, a pullulanase mediating the degradation of amylopectin or glycogen, and specific amino acid uptake systems; in L. johnsonii NCC533: an arsenite efflux system, a predicted LPS biosynthesis gene cluster, a potential microcin synthesis or resistance system, and a cell-envelop proteinase potentially facilitating proteolysis of macromolecules; in L. gasseri ATCC33323: strain-specific mucus-binding proteins, a galactose uptake system, DNA restriction/modification systems, a nucleotidase degrading extracellular DNA molecules, and an energy-generating system based on l-serine; in L. plantarum WCFS1: the non-ribosomal peptide synthesis module, a respiratory nitrate reductase complex, and the sugar metabolism island). These traits may also provide distinct functional advantages and could potentially be exploited for future strain improvements.
Traditional genome analyses have indicated that L. johnsonii NCC533 and L. gasseri ATCC33323 feature extensive differences in their genomes and, contrary to earlier beliefs, represent two distinct species [15]. Based on the identified strain-specific differences, this observation was further strengthened. Indeed, DBA might provide a valuable tool to assess the similarity of sequenced bacterial strains and aid in their respective phylogenetic classification.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Acknowledgments
This research was supported by The Southeast Dairy Foods Research Center, Dairy Management Inc., Danisco USA Inc., and the North Carolina Dairy Foundation. The authors wish to thank Sonja Lick, Mick Callanan, Mike Russel (Rhodia), Olivia McAuliffe, Andrea Azcarate, B. Logan Buck, Alleson Dobson, Mike Miller, Evelyn Durmaz, Erika Pfeiler, Jun Goh, Tri Duong, and Rodolphe Barrangou for their contributions to the manual annotation of Lactobacillus acidophilus NCFM and Lactobacillus gasseri ATCC33323 genomes. In particular, we thank Andrea Azcarate-Peril, Evelyn Durmaz, and B. Logan Buck for their help and critical review of the manuscript and Zaneta Park-Ng for the statistical data evaluation. We are grateful to Gariella van Zanten and Sampo Lahtinen for the communication of L. acidophilus NCFM growth characteristics. We would also like to thank Hans-Henrik Staerfeldt for kindly providing the “Genewiz” software.
Contributor Information
Eric Altermann, Email: Eric.Altermann@agresearch.co.nz.
Todd R. Klaenhammer, Phone: +1-919-5152972, Phone: +1-919-5154197, FAX: +1-919-5157124, Email: Klaenhammer@ncsu.edu
References
- 1.Abe K, Ohnishi F, Yagi K, Nakajima T, Higuchi T, Sano M, Machida M, Sarker RI, Maloney PC. Plasmid-encoded asp operon confers a proton motive metabolic cycle catalyzed by an aspartate-alanine exchange reaction. J Bacteriol. 2002;184(11):2906–2913. doi: 10.1128/JB.184.11.2906-2913.2002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Ahrne S, Lonnermark E, Wold AE, Aberg N, Hesselmar B, Saalman R, Strannegard IL, Molin G, Adlerberth I. Lactobacilli in the intestinal microbiota of Swedish infants. Microbes Infect. 2005;7(11–12):1256–1262. doi: 10.1016/j.micinf.2005.04.011. [DOI] [PubMed] [Google Scholar]
- 3.Altermann E, Buck LB, Cano R, Klaenhammer TR. Identification and phenotypic characterization of the cell-division protein CdpA. Gene. 2004;342(1):189–197. doi: 10.1016/j.gene.2004.08.004. [DOI] [PubMed] [Google Scholar]
- 4.Altermann E, Klaenhammer TR. GAMOLA: a new local solution for sequence annotation and analyzing draft and finished prokaryotic genomes. Omics. 2003;7(2):161–169. doi: 10.1089/153623103322246557. [DOI] [PubMed] [Google Scholar]
- 5.Altermann E, Klaenhammer TR. PathwayVoyager: pathway mapping using the Kyoto encyclopedia of genes and genomes (KEGG) database. BMC Genomics. 2005;6(1):60. doi: 10.1186/1471-2164-6-60. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Altermann E, Russell WM, Azcarate-Peril MA, Barrangou R, Buck BL, McAuliffe O, Souther N, Dobson A, Duong T, Callanan M, Lick S, Hamrick A, Cano R, Klaenhammer TR. Complete genome sequence of the probiotic lactic acid bacterium Lactobacillus acidophilus NCFM. Proc Natl Acad Sci USA. 2005;102(11):3906–3912. doi: 10.1073/pnas.0409188102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Altschul SF, Madden TL, Schaffer AA, Zhang J, Zhang Z, Miller W, Lipman DJ. Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res. 1997;25(17):3389–3402. doi: 10.1093/nar/25.17.3389. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Azcarate-Peril MA, Altermann E, Goh YJ, Tallon R, Sanozky-Dawes RB, Pfeiler EA, O’Flaherty S, Buck BL, Dobson A, Duong T, Miller MJ, Barrangou R, Klaenhammer TR. Analysis of the genome sequence of Lactobacillus gasseri ATCC 33323 reveals the molecular basis of an autochthonous intestinal organism. Appl Environ Microbiol. 2008;74(15):4610–4625. doi: 10.1128/AEM.00054-08. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Azcarate-Peril MA, Altermann E, Hoover-Fitzula RL, Cano RJ, Klaenhammer TR. Identification and inactivation of genetic loci involved with Lactobacillus acidophilus acid tolerance. Appl Environ Microbiol. 2004;70(9):5315–5322. doi: 10.1128/AEM.70.9.5315-5322.2004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Azcarate-Peril MA, Bruno-Barcena JM, Hassan HM, Klaenhammer TR. Transcriptional and functional analysis of oxalyl-coenzyme A (CoA) decarboxylase and formyl-CoA transferase genes from Lactobacillus acidophilus. Appl Environ Microbiol. 2006;72(3):1891–1899. doi: 10.1128/AEM.72.3.1891-1899.2006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Azcarate-Peril MA, McAuliffe O, Altermann E, Lick S, Russell WM, Klaenhammer TR. Microarray analysis of a two-component regulatory system involved in acid resistance and proteolytic activity in Lactobacillus acidophilus. Appl Environ Microbiol. 2005;71(10):5794–5804. doi: 10.1128/AEM.71.10.5794-5804.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Baillon ML, Marshall-Jones ZV, Butterwick RF. Effects of probiotic Lactobacillus acidophilus strain DSM13241 in healthy adult dogs. Am J Vet Res. 2004;65(3):338–343. doi: 10.2460/ajvr.2004.65.338. [DOI] [PubMed] [Google Scholar]
- 13.Balakrishnan L, Venter H, Shilling RA, Veen HW. Reversible transport by the ATP-binding cassette multidrug export pump LmrA: ATP synthesis at the expense of downhill ethidium uptake. J Biol Chem. 2004;279(12):11273–11280. doi: 10.1074/jbc.M308494200. [DOI] [PubMed] [Google Scholar]
- 14.Barrangou R, Altermann E, Hutkins R, Cano R, Klaenhammer TR. Functional and comparative genomic analyses of an operon involved in fructooligosaccharide utilization by Lactobacillus acidophilus. Proc Natl Acad Sci USA. 2003;100(15):8957–8962. doi: 10.1073/pnas.1332765100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Boekhorst J, Siezen RJ, Zwahlen MC, Vilanova D, Pridmore RD, Mercenier A, Kleerebezem M, Vos WM, Brussow H, Desiere F. The complete genomes of Lactobacillus plantarum and Lactobacillus johnsonii reveal extensive differences in chromosome organization and gene content. Microbiology. 2004;150(Pt 11):3601–3611. doi: 10.1099/mic.0.27392-0. [DOI] [PubMed] [Google Scholar]
- 16.Buck BL, Altermann E, Svingerud T, Klaenhammer TR. Functional analysis of putative adhesion factors in Lactobacillus acidophilus NCFM. Appl Environ Microbiol. 2005;71(12):8344–8351. doi: 10.1128/AEM.71.12.8344-8351.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Carver TJ, Rutherford KM, Berriman M, Rajandream MA, Barrell BG, Parkhill J. ACT: the Artemis comparison tool. Bioinformatics. 2005;21(16):3422–3423. doi: 10.1093/bioinformatics/bti553. [DOI] [PubMed] [Google Scholar]
- 18.Claesson MJ, Li Y, Leahy S, Canchaya C, Pijkeren JP, Cerdeno-Tarraga AM, Parkhill J, Flynn S, O’Sullivan GC, Collins JK, Higgins D, Shanahan F, Fitzgerald GF, Sinderen D, O’Toole PW. Multireplicon genome architecture of Lactobacillus salivarius. PNAS. 2006;103(17):6718–6723. doi: 10.1073/pnas.0511060103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Collado MC, Isolauri E, Salminen S, Sanz Y. The impact of probiotic on gut health. Curr Drug Metab. 2009;10(1):68–78. doi: 10.2174/138920009787048437. [DOI] [PubMed] [Google Scholar]
- 20.Denou E, Pridmore RD, Ventura M, Pittet A-C, Zwahlen M-C, Berger B, Barretto C, Panoff J-M, Brussow H. The role of prophage for genome diversification within a clonal lineage of Lactobacillus johnsonii: characterization of the defective prophage LJ771. J Bacteriol. 2008;190(17):5806–5813. doi: 10.1128/JB.01802-07. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Duncan SH, Richardson AJ, Kaul P, Holmes RP, Allison MJ, Stewart CS. Oxalobacter formigenes and its potential role in human health. Appl Environ Microbiol. 2002;68(8):3841–3847. doi: 10.1128/AEM.68.8.3841-3847.2002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Dyrlov Bendtsen J, Nielsen H, Heijne G, Brunak S. Improved prediction of signal peptides: SignalP 3.0. J Mol Biol. 2004;340(4):783–795. doi: 10.1016/j.jmb.2004.05.028. [DOI] [PubMed] [Google Scholar]
- 23.Fomenko DE, Metlitskaya AZ, Peduzzi J, Goulard C, Katrukha GS, Gening LV, Rebuffat S, Khmel IA. Microcin C51 plasmid genes: possible source of horizontal gene transfer. Antimicrob Agents Chemother. 2003;47(9):2868–2874. doi: 10.1128/AAC.47.9.2868-2874.2003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Fujisawa T, Benno Y, Yaeshima T, Mitsuoka T. Taxonomic study of the Lactobacillus acidophilus group, with recognition of Lactobacillus gallinarum sp. nov. and Lactobacillus johnsonii sp. nov. and synonymy of Lactobacillus acidophilus group A3 (Johnson et al. 1980) with the type strain of Lactobacillus amylovorus (Nakamura 1981) Int J Syst Bacteriol. 1992;42(3):487–491. doi: 10.1099/00207713-42-3-487. [DOI] [PubMed] [Google Scholar]
- 25.Germond JE, Delley M, Gilbert C, Atlan D. Determination of the domain of the Lactobacillus delbrueckii subsp. bulgaricus cell surface proteinase PrtB involved in attachment to the cell wall after heterologous expression of the prtB gene in Lactococcus lactis. Appl Environ Microbiol. 2003;69(6):3377–3384. doi: 10.1128/AEM.69.6.3377-3384.2003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Gonzalez-Pastor JE, San Millan JL, Castilla MA, Moreno F. Structure and organization of plasmid genes required to produce the translation inhibitor microcin C7. J Bacteriol. 1995;177(24):7131–7140. doi: 10.1128/jb.177.24.7131-7140.1995. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Grangette C, Muller-Alouf H, Geoffroy M, Goudercourt D, Turneer M, Mercenier A. Protection against tetanus toxin after intragastric administration of two recombinant lactic acid bacteria: impact of strain viability and in vivo persistence. Vaccine. 2002;20(27–28):3304–3309. doi: 10.1016/S0264-410X(02)00301-8. [DOI] [PubMed] [Google Scholar]
- 28.Guan LL, Hagen KE, Tannock GW, Korver DR, Fasenko GM, Allison GE. Detection and identification of Lactobacillus species in crops of broilers of different ages by using PCR-denaturing gradient gel electrophoresis and amplified ribosomal DNA restriction analysis. Appl Environ Microbiol. 2003;69(11):6750–6757. doi: 10.1128/AEM.69.11.6750-6757.2003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Guilfoile PG, Hutchinson CR. A bacterial analog of the mdr gene of mammalian tumor cells is present in Streptomyces peucetius, the producer of daunorubicin and doxorubicin. Proc Natl Acad Sci USA. 1991;88(19):8553–8557. doi: 10.1073/pnas.88.19.8553. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Heider J. A new family of CoA-transferases. FEBS Lett. 2001;509(3):345–349. doi: 10.1016/S0014-5793(01)03178-7. [DOI] [PubMed] [Google Scholar]
- 31.Henrissat B. A classification of glycosyl hydrolases based on amino acid sequence similarities. Biochem J. 1991;280(Pt 2):309–316. doi: 10.1042/bj2800309. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Hofreuter D, Haas R. Characterization of two cryptic Helicobacter pylori plasmids: a putative source for horizontal gene transfer and gene shuffling. J Bacteriol. 2002;184(10):2755–2766. doi: 10.1128/JB.184.10.2755-2766.2002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Kanehisa M. The KEGG database. Novartis Found Symp. 2002;247:91–101. doi: 10.1002/0470857897.ch8. [DOI] [PubMed] [Google Scholar]
- 34.Kankainen M, Paulin L, Tynkkynen S, Ossowski I, Reunanen J, Partanen P, Satokari R, Vesterlund S, Hendrickx AP, Lebeer S, Keersmaecker SC, Vanderleyden J, Hamalainen T, Laukkanen S, Salovuori N, Ritari J, Alatalo E, Korpela R, Mattila-Sandholm T, Lassig A, Hatakka K, Kinnunen KT, Karjalainen H, Saxelin M, Laakso K, Surakka A, Palva A, Salusjarvi T, Auvinen P, Vos WM. Comparative genomic analysis of Lactobacillus rhamnosus GG reveals pili containing a human- mucus binding protein. Proc Natl Acad Sci USA. 2009;106(40):17193–17198. doi: 10.1073/pnas.0908876106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Klaenhammer T, Altermann E, Arigoni F, Bolotin A, Breidt F, Broadbent J, Cano R, Chaillou S, Deutscher J, Gasson M, Guchte M, Guzzo J, Hartke A, Hawkins T, Hols P, Hutkins R, Kleerebezem M, Kok J, Kuipers O, Lubbers M, Maguin E, McKay L, Mills D, Nauta A, Overbeek R, Pel H, Pridmore D, Saier M, Sinderen D, Sorokin A, Steele J, O’Sullivan D, Vos W, Weimer B, Zagorec M, Siezen R. Discovering lactic acid bacteria by genomics. Antonie Van Leeuwenhoek. 2002;82(1–4):29–58. doi: 10.1007/978-94-017-2029-8_3. [DOI] [PubMed] [Google Scholar]
- 36.Klaenhammer TR, Azcarate-Peril MA, Barrangou R, Duong T, Altermann E. Genomic perspectives on probiotic lactic acid bacteria. Biosci Microflora. 2005;24:31–33. [Google Scholar]
- 37.Klaenhammer TR, Barrangou R, Buck BL, Azcarate-Peril MA, Altermann E. Genomic features of lactic acid bacteria effecting bioprocessing and health. FEMS Microbiol Rev. 2005;29(3):393–409. doi: 10.1016/j.fmrre.2005.04.007. [DOI] [PubMed] [Google Scholar]
- 38.Klaenhammer TR, Russell WM. Species of the Lactobacillus acidophilus complex, vol 2. Encyclopedia of food microbiology. San Diego: Academic Press; 2000. [Google Scholar]
- 39.Kleerebezem M, Boekhorst J, Kranenburg R, Molenaar D, Kuipers OP, Leer R, Tarchini R, Peters SA, Sandbrink HM, Fiers MW, Stiekema W, Lankhorst RM, Bron PA, Hoffer SM, Groot MN, Kerkhoven R, Vries M, Ursing B, Vos WM, Siezen RJ. Complete genome sequence of Lactobacillus plantarum WCFS1. Proc Natl Acad Sci USA. 2003;100(4):1990–1995. doi: 10.1073/pnas.0337704100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Kumar S, Tamura K, Nei M. MEGA3: integrated software for molecular evolutionary genetics analysis and sequence alignment. Briefings Bioinform. 2004;5(2):150–163. doi: 10.1093/bib/5.2.150. [DOI] [PubMed] [Google Scholar]
- 41.La Ragione RM, Narbad A, Gasson MJ, Woodward MJ. In vivo characterization of Lactobacillus johnsonii FI9785 for use as a defined competitive exclusion agent against bacterial pathogens in poultry. Lett Appl Microbiol. 2004;38(3):197–205. doi: 10.1111/j.1472-765X.2004.01474.x. [DOI] [PubMed] [Google Scholar]
- 42.Larkin MA, Blackshields G, Brown NP, Chenna R, McGettigan PA, McWilliam H, Valentin F, Wallace IM, Wilm A, Lopez R, Thompson JD, Gibson TJ, Higgins DG. Clustal W and clustal X version 2.0. Bioinformatics. 2007;23(21):2947–2948. doi: 10.1093/bioinformatics/btm404. [DOI] [PubMed] [Google Scholar]
- 43.Lauer E, Kandler O. Lactobacillus gasseri sp. nov., a new species of the subgenus Thermobacterium. Zentralbl Bakteriol Mikrobiol Hyg Abt. 1980;1(C1):75–78. [Google Scholar]
- 44.Makarova K, Slesarev A, Wolf Y, Sorokin A, Mirkin B, Koonin E, Pavlov A, Pavlova N, Karamychev V, Polouchine N, Shakhova V, Grigoriev I, Lou Y, Rohksar D, Lucas S, Huang K, Goodstein DM, Hawkins T, Plengvidhya V, Welker D, Hughes J, Goh Y, Benson A, Baldwin K, Lee JH, Diaz-Muniz I, Dosti B, Smeianov V, Wechter W, Barabote R, Lorca G, Altermann E, Barrangou R, Ganesan B, Xie Y, Rawsthorne H, Tamir D, Parker C, Breidt F, Broadbent J, Hutkins R, O’Sullivan D, Steele J, Unlu G, Saier M, Klaenhammer T, Richardson P, Kozyavkin S, Weimer B, Mills D. Comparative genomics of the lactic acid bacteria. PNAS. 2006;103(42):15611–15616. doi: 10.1073/pnas.0607117103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Marshall-Jones ZV, Baillon ML, Croft JM, Butterwick RF. Effects of Lactobacillus acidophilus DSM13241 as a probiotic in healthy adult cats. Am J Vet Res. 2006;67(6):1005–1012. doi: 10.2460/ajvr.67.6.1005. [DOI] [PubMed] [Google Scholar]
- 46.Masuda Y, Miyakawa K, Nishimura Y, Ohtsubo E. chpA and chpB, Escherichia coli chromosomal homologs of the pem locus responsible for stable maintenance of plasmid R100. J Bacteriol. 1993;175(21):6850–6856. doi: 10.1128/jb.175.21.6850-6856.1993. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Mercenier A, Pavan S, Pot B. Probiotics as biotherapeutic agents: present knowledge and future prospects. Curr Pharm Des. 2003;9(2):175–191. doi: 10.2174/1381612033392224. [DOI] [PubMed] [Google Scholar]
- 48.Moore MH, Gulbis JM, Dodson EJ, Demple B, Moody PC. Crystal structure of a suicidal DNA repair protein: the Ada O6-methylguanine-DNA methyltransferase from E. coli. EMBO J. 1994;13(7):1495–1501. doi: 10.1002/j.1460-2075.1994.tb06410.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Ogawa Y, Miyazato T, Hatano T. Oxalate and urinary stones. World J Surg. 2000;24(10):1154–1159. doi: 10.1007/s002680010193. [DOI] [PubMed] [Google Scholar]
- 50.Papadopoulos JS, Agarwala R. COBALT: constraint-based alignment tool for multiple protein sequences. Bioinformatics. 2007;23(9):1073–1079. doi: 10.1093/bioinformatics/btm076. [DOI] [PubMed] [Google Scholar]
- 51.Pascher M, Hellweg P, Khol-Parisini A, Zentek J. Effects of a probiotic Lactobacillus acidophilus strain on feed tolerance in dogs with non-specific dietary sensitivity. Arch Anim Nutr. 2008;62(2):107–116. doi: 10.1080/17450390801892583. [DOI] [PubMed] [Google Scholar]
- 52.Pedersen AG, Jensen LJ, Brunak S, Staerfeldt H-H, Ussery DW. A DNA structural atlas for Escherichia coli. J Mol Biol. 2000;299(4):907–930. doi: 10.1006/jmbi.2000.3787. [DOI] [PubMed] [Google Scholar]
- 53.Pederson JA, Mileski GJ, Weimer BC, Steele JL. Genetic characterization of a cell envelope-associated proteinase from Lactobacillus helveticus CNRZ32. J Bacteriol. 1999;181(15):4592–4597. doi: 10.1128/jb.181.15.4592-4597.1999. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 54.Peterson RE, Klopfenstein TJ, Erickson GE, Folmer J, Hinkley S, Moxley RA, Smith DR. Effect of Lactobacillus acidophilus strain NP51 on Escherichia coil O157:H7 fecal shedding and finishing performance in beef feedlot cattle. J Food Prot. 2007;70(2):287–291. doi: 10.4315/0362-028x-70.2.287. [DOI] [PubMed] [Google Scholar]
- 55.Pridmore RD, Berger B, Desiere F, Vilanova D, Barretto C, Pittet AC, Zwahlen MC, Rouvet M, Altermann E, Barrangou R, Mollet B, Mercenier A, Klaenhammer T, Arigoni F, Schell MA. The genome sequence of the probiotic intestinal bacterium Lactobacillus johnsonii NCC 533. Proc Natl Acad Sci USA. 2004;101(8):2512–2517. doi: 10.1073/pnas.0307327101. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Rogelj I, Bogovic Matijasic B, Canzek Majhenic A, Stojkovic S. The survival and persistence of Lactobacillus acidophilus LF221 in different ecosystems. Int J Food Microbiol. 2002;76(1–2):83–91. doi: 10.1016/S0168-1605(02)00019-3. [DOI] [PubMed] [Google Scholar]
- 57.Rutherford K, Parkhill J, Crook J, Horsnell T, Rice P, Rajandream MA, Barrell B. Artemis: sequence visualization and annotation. Bioinformatics. 2000;16(10):944–945. doi: 10.1093/bioinformatics/16.10.944. [DOI] [PubMed] [Google Scholar]
- 58.Sa-Nogueira I, Nogueira TV, Soares S, Lencastre H. The Bacillus subtilisl-arabinose (ara) operon: nucleotide sequence, genetic organization and expression. Microbiology. 1997;143(3):957–969. doi: 10.1099/00221287-143-3-957. [DOI] [PubMed] [Google Scholar]
- 59.Saitou N, Nei M. The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol. 1987;4(4):406–425. doi: 10.1093/oxfordjournals.molbev.a040454. [DOI] [PubMed] [Google Scholar]
- 60.Sedgwick B, Lindahl T. Recent progress on the Ada response for inducible repair of DNA alkylation damage. Oncogene. 2002;21(58):8886–8894. doi: 10.1038/sj.onc.1205998. [DOI] [PubMed] [Google Scholar]
- 61.Siezen RJ. Multi-domain, cell-envelope proteinases of lactic acid bacteria. Antonie Van Leeuwenhoek. 1999;76(1–4):139–155. doi: 10.1023/A:1002036906922. [DOI] [PubMed] [Google Scholar]
- 62.Sigrell JA, Cameron AD, Jones TA, Mowbray SL. Purification, characterization, and crystallization of Escherichia coli ribokinase. Protein Sci. 1997;6(11):2474–2476. doi: 10.1002/pro.5560061124. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63.Smitherman PK, Townsend AJ, Kute TE, Morrow CS. Role of multidrug resistance protein 2 (MRP2, ABCC2) in alkylating agent detoxification: MRP2 potentiates glutathione S-transferase A1–1-mediated resistance to chlorambucil cytotoxicity. J Pharmacol Exp Ther. 2004;308(1):260–267. doi: 10.1124/jpet.103.057729. [DOI] [PubMed] [Google Scholar]
- 64.Tan KS, Song KP, Ong G. Bacteroides forsythus prtH genotype in periodontitis patients: occurrence and association with periodontal disease. J Periodontal Res. 2001;36(6):398–403. doi: 10.1034/j.1600-0765.2001.360608.x. [DOI] [PubMed] [Google Scholar]
- 65.Trotter PJ, Pedretti J, Yates R, Voelker DR. Phosphatidylserine decarboxylase 2 of Saccharomyces cerevisiae. Cloning and mapping of the gene, heterologous expression, and creation of the null allele. J Biol Chem. 1995;270(11):6071–6080. doi: 10.1074/jbc.270.11.6071. [DOI] [PubMed] [Google Scholar]
- 66.van de Guchte M, Penaud S, Grimaldi C, Barbe V, Bryson K, Nicolas P, Robert C, Oztas S, Mangenot S, Couloux A, Loux V, Dervyn R, Bossy R, Bolotin A, Batto JM, Walunas T, Gibrat JF, Bessieres P, Weissenbach J, Ehrlich SD, Maguin E (2006) The complete genome sequence of Lactobacillus bulgaricus reveals extensive and ongoing reductive evolution. PNAS 103:9274–9279 [DOI] [PMC free article] [PubMed]
- 67.Ventura M, Canchaya C, Pridmore RD, Brussow H. The prophages of Lactobacillus johnsonii NCC 533: comparative genomics and transcription analysis. Virology. 2004;320(2):229–242. doi: 10.1016/j.virol.2003.11.034. [DOI] [PubMed] [Google Scholar]
- 68.Ossowski I, Reunanen J, Satokari R, Vesterlund S, Kankainen M, Huhtinen H, Tynkkynen S, Salminen S, Vos WM, Palva A. Mucosal adhesion properties of the probiotic Lactobacillus rhamnosus GG SpaCBA and SpaFED Pilin subunits. Appl Environ Microbiol. 2010;76(7):2049–2057. doi: 10.1128/AEM.01958-09. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69.Vos P, Asseldonk M, Jeveren F, Siezen R, Simons G, Vos WM. A maturation protein is essential for production of active forms of Lactococcus lactis SK11 serine proteinase located in or secreted from the cell envelope. J Bacteriol. 1989;171(5):2795–2802. doi: 10.1128/jb.171.5.2795-2802.1989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 70.Wolf BW, Garleb KA, Choe YS, Humphrey PM, Maki KC. Pullulan is a slowly digested carbohydrate in humans. J Nutr. 2003;133(4):1051–1055. doi: 10.1093/jn/133.4.1051. [DOI] [PubMed] [Google Scholar]
- 71.Yang M, Luoh SM, Goddard A, Reilly D, Henzel W, Bass S. The bglX gene located at 47.8 min on the Escherichia coli chromosome encodes a periplasmic beta-glucosidase. Microbiology. 1996;142(Pt 7):1659–1665. doi: 10.1099/13500872-142-7-1659. [DOI] [PubMed] [Google Scholar]
- 72.Zhou S-F, Wang L-L, Di YM, CChangli Xue, Duan W, CGuang Li, Li Y. Substrates and inhibitors of human multidrug resistance associated proteins and the implications in drug development. Curr Med Chem. 2008;15:1981–2039. doi: 10.2174/092986708785132870. [DOI] [PubMed] [Google Scholar]
- 73.Zuckerkandl E, Pauling L. Evolutionary divergence and convergence in proteins. In: Bryson V, Vogel HJ, editors. Evolving genes and proteins. New York: Academic Press; 1965. pp. 97–166. [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Below is the link to the electronic supplementary material.