Abstract
The mitochondrial genome of trypanosomes is composed of ~50 maxicircles and thousands of minicircles. Maxi- (~25 kb) and mini-(~1 kb)circles are catenated and packed into a dense structure called a kinetoplast. Both types of circular DNA are transcribed by a phage-like RNA polymerase: maxicircles yield multicistronic rRNA and mRNA precursors, while guide RNA (gRNA) precursors are produced from minicircles. To function in mitochondrial translation, pre-mRNAs must undergo a nucleolytic processing and 3′ modifications, and often uridine insertion/deletion editing. gRNAs, which represent short (50-60 nt) RNAs directing editing reactions, are produced by 3′ nucleolytic processing of a much longer precursor followed by 3′ uridylation. Ribosomal RNAs are excised from precursors and their 3′ ends are also trimmed and uridylated. All tRNAs are imported from the cytoplasm and some are further modified and edited in the mitochondrial matrix. Historically, the fascinating phenomenon of RNA editing has been extensively studied as an isolated pathway in which nuclear-encoded proteins mediate interactions of maxi- and minicircle transcripts to create open reading frames. However, recent studies unraveled a highly integrated network of mitochondrial genome expression including critical pre- and postediting 3′ mRNA processing, and gRNA and rRNA maturation steps. Here we focus on RNA 3′ adenylation and uridylation as processes essential for biogenesis, stability and functioning of mitochondrial RNAs.
Keywords: Trypanosoma, Mitochondria, RNA editing, RNA stability, Polyadenylation, Uridylation, mRNA, gRNA, rRNA, PPR proteins, Translation
1. Introduction
Notwithstanding a common eubacterial origin, mitochondria of different phyla display a variety of post-transcriptional processes including endo- and exonucleolytic degradation, splicing, 3′ modification, nucleoside modifications and RNA editing. The processing pathways are often evolutionarily unrelated, even though their outcomes may ostensibly serve the same purpose. For example, trypanosomal U-insertion/deletion editing and C to U editing in plants (Stern et al., 2010) change the mRNA sequence to create open reading frames or alter amino acid identities. Conversely, similar modifications performed by homologous enzymes may carry out opposing functions: 3′ addition of 20-30 adenosines stimulates degradation of mitochondrial mRNAs in algae and higher plants (Lange et al., 2009), while similar A-tails are required for mRNA maintenance in trypanosomes (Ryan et al., 2003;Kao and Read, 2005;Etheridge et al., 2008). The RNA degradation apparatus may involve endonucleolytic cuts in combination with 5′-3′ and/or 3′-5′ exonucleolytic digestion or rely predominantly on exonucleases. Overall, the only recurring theme among different systems is that the organellar genome expression is largely controlled post-transcriptionally; specific mechanisms typically invoke a phylogenetically and functionally diverse collage of prototypic bacterial and acquired eukaryotic components.
The biomedical significance of Kinetoplastids, a group of flagellated protozoans including the species Trypanosoma and Leishmania, has continuously stimulated research of their unusual mitochondrial gene expression as a premise for drug discovery (Stuart et al., 2008). From a basic science perspective, the bizarre mechanisms of kinetoplast DNA (kDNA) replication (Liu et al., 2005) and U-insertion/deletion mRNA editing remain captivating after decades of research. Somewhere between these trends, the molecular details of mitochondrial transcription, pre- and postediting mRNA processing, ribosomal RNA and guide RNA biogenesis and translation remain very sketchy. However, the ‘fringe benefits’ of editing research, such as identification of RNA uridylation and adenylation enzymes, and recent advances in genetic and biochemical approaches position the ultimate goal of understanding the developmentally regulated mitochondrial gene expression within experimental reach.
2. Transcription
The catalytic C-terminal domain of a single-subunit mitochondrial RNA polymerase(mtRNAP) resembles a prototypic T7 phage RNA polymerase-like module (Wang et al., 2000; Clement and Koslowsky, 2001) which is found in mitochondria of most organisms. Although neither transcription factors nor mitochondrial promoters have been identified, available evidence implicates the 144 kDa mtRNAP in transcription of both maxicircle- and minicircle genomes and in kDNA replication (Grams et al., 2002;Hashimi et al., 2009). In this context, the key question is whether maxicircles and minicircles share cis-elements and transcriptional complexes, or whether different promoters recruit diverse transcription factors to engage the RNA polymerase. In the maxicircle, rRNA and protein coding genes are located on both strands of the ~17 kb-long conservative region, while the remaining part, termed the divergent region, is composed of repeated sequences (Sloof et al., 1992). Transcription of the major strand is initiated ~1,200 nt upstream of the 12S rRNA gene (Michelotti et al., 1992), while origins of minor strand transcription have not been determined (Fig. 1). The minicircle genome is more complex. With an estimated 200-400 minicircle sequence classes, each encoding 2–5 gRNAs, the number of potential guide RNAs greatly exceeds the requirement for known editing sites, although gRNAs of slightly different sequences may edit the same sites (Hong and Simpson, 2003;Ochsenreiter et al., 2007). In addition to gRNAs, minicircles encode similarly-sized RNAs without apparent function (Madej et al., 2007;Madej et al., 2008). Unlike maxicircle-encoded RNAs, guide RNAs and gRNA-like molecules maintain 5′ triphosphates, which indicates a lack of 5′ processing of their primary transcripts (Blum and Simpson, 1990). Although transcription start sites have been mapped in several cases (Blum et al., 1990), minicircle promoters remain elusive. Guide RNA coding sequences are typically flanked by short 18 bp inverted repeats (Jasmer and Stuart, 1986;Pollard et al., 1990), but participation of these sequences in transcription initiation or any other function is unclear. For example, in T. brucei, the maxicircle encodes a single trans-acting gRNA (gMURF2-II) which is located within the 5′ region of the ND4 gene. Although surrounding sequences lack apparent repeats, the gMURF2-II transcription pattern closely resembles that of minicircle-encoded molecules: gRNA is produced as a ~800 nt precursor (Clement et al., 2004;Aphasizheva and Aphasizhev, 2010;Grams et al., 2000). Given the minicircle's length of ~1 kb and the presence of more than one gRNA gene per minicircle, the bulk of gRNA-containing primary transcripts are also likely to be polycistronic thus requiring the nucleolytic and 3′ modification processing to achieve a functional state of 50-60 nt molecules terminating with 15-20-nt oligo(U) tails.
Fig. 1.
General outline of mitochondrial RNA processing in trypanosomes. Polycistronic transcripts are produced by mitochondrial RNA polymerase from maxicircle and minicircle components of the kinetoplast DNA. ND and NADH dehydrogenase. CO, cytochrome c oxidase. MURF, mitochondrial unidentified frame. GR, G-rich region. A single trans-acting gRNA (ND2, gMurf1-[II]) is transcribed from the maxicircle. Irrespective of genomic location, pre-gRNAs undergo 3′ nucleolytic processing by a cryptic nuclease via a pathway requiring RET1 activity (Aphasizheva and Aphasizhev, 2010). Processed guide RNAs are stabilized by binding to the gRNA binding complex (GRBC, (Weng et al., 2008;Aphasizheva and Aphasizhev, 2010)) and 3′ uridylated by RET1 TUTase (Aphasizhev et al., 2002;Aphasizhev et al., 2003b), but the order of events is unclear. Pre-rRNAs are also trimmed at the 3′ end and uridylated by RET1 (Aphasizheva and Aphasizhev, 2010). Guide RNA hybridization with pre-edited mRNA activates the editing process, but may also be responsible for transient association of RET1 with 3′ adenylation (KPAP1) complex. Editing events in the 3′ region confer a requirement for the short A-tail as cis-element necessary (Etheridge et al., 2008) and sufficient for mRNA stability (Aphasizheva et al., 2011). The completion of editing, typically at the 5′ region, triggers extension of the pre-existing A-tail into A/U-heteropolymer by KPAF1-2-coordinated, KPAP1/RET1-catalyzed reaction. This final processing event (green arrows) constitutes a rate-limiting step in mitochondrial mRNA processing which facilitates mRNA binding to the small ribosomal subunit (Aphasizheva et al., 2011).
3. Nucleolytic processing
It is generally presumed that both endo- and exonuclease activities are required to form mature mRNAs from polycistronic RNAs. In T. brucei, this process is further complicated by ubiquitous intersections between adjacent transcripts within a precursor. In several cases, the mature 3′-end of the upstream transcript is produced at the expense of the 5′-untranslated region (UTR) in the downstream unit, and vice-versa (e.g. ND7 and CO3 pre-mRNAs (Koslowsky and Yahampath, 1997), 9S ribosomal RNA and ND8 pre-mRNA (Aphasizheva and Aphasizhev, 2010)). Because mature RNAs with both correctly processed ends are present in the steady-state population, alternative cleavage events may occur stochastically or in a regulated fashion. Several nucleases have been investigated, but none could be unambiguously assigned to a specific processing function. A mitochondrial-associated (endo)ribonuclease (MAR1) has been purified and the gene cloned from L. tarentolae (Alfonzo et al., 1998). In addition, three distinct 3′-5′ exonuclease activities were characterized in L. tarentolae: a U-specific exonuclease, a processive non-specific enzyme and an exonuclease with preference for 3′ phosphorylated RNAs (Aphasizhev and Simpson, 2001). Protein identities of these three exonucleases remain to be determined and MAR1's function needs further investigation. In T. brucei, the RNA editing core complex (RECC, also referred to as ~20S editosome) is associated with three RNase III-type endonucleases and two 3′-5′ U-specific exonucleases typified by the presence of endonuclease/exonuclease/phosphatase (EEP) catalytic domains (reviewed in (Aphasizhev and Aphasizheva, 2011)). Although a “moonlighting” function of editing endo- and/or exonucleases in processing cannot be excluded, available evidence indicates their confinement to the U-insertion/deletion RNA editing. A mitochondrial RNase P activity has been reported in T. brucei, but the protein responsible for cleavage of model RNA substrate in vitro remains to be identified (Salavati et al., 2001). Since the entire set of tRNAs acting in mitochondrial translation is imported from the cytosol as processed molecules (reviewed in (Alfonzo and Soll, 2009)), tRNA maturation is unlikely to be the major function of mitochondrial RNase P. The trypanosomal homolog of yeast DSS-1 exonuclease participates in RNA surveillance with one documented example being 3′-5′ degradation of the non-coding fragment removed from the 12S pre-rRNA (Mattiacio and Read, 2008). Finally, the existence of 5′-3′ RNA degradation activities in mitochondria of trypanosomes is not supported by the available data. Although identification of processing nucleases remains a major challenge, recent work (Aphasizheva and Aphasizhev, 2010) indicates that short anti-sense RNAs may direct these cleavage reactions similarly to the gRNA-dependent cleavage during mRNA editing (Figs. 2 and 4).
Fig. 2.
A. Schematic representation of U-deletion (left) and U-insertion (right) enzymatic cascades catalyzed by RECC1 and RECC2, respectively. RECC – RNA editing core complex. U-insertion and U-deletion subcomplexes are depicted in blue and green, respectively. MP: mitochondrial protein; REX: RNA editing exonuclease; REN: RNA editing endonuclease; REL: RNA editing ligase; RET: RNA editing TUTase; gRNA: guide RNA; anchor: 5-15-nt long double-stranded region formed by the 5′-portion of the gRNA and pre-edited mRNA. Initial endonucleolytic mRNA cleavage occurs at the first unpaired nucleotide upstream of the anchor. B. Polarity of mRNA editing. The 3′-5′ (mRNA) polarity of RNA editing is determined by hierarchical gRNA binding in which editing at the 3′ region often creates a binding site for the next gRNA.
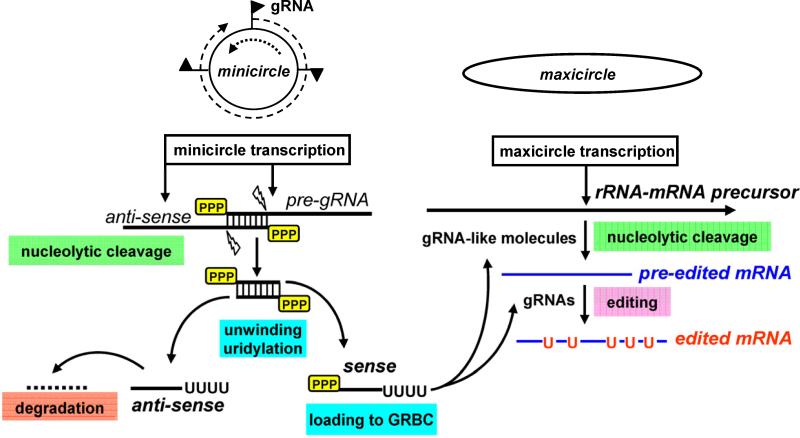
Fig. 4.
A model for generation of gRNAs and gRNA-like molecules involved in RNA editing and maxicircle precursor processing, respectively. The double-stranded substrate for the processing endonuclease may be generated by overlapping transcripts synthesized from opposite strands. Uridylation likely takes place after unwinding of the gRNA-sized duplex. It is also possible that the otherwise processive RET1 adds only 15-20 Us to GRBC-bound gRNAs and gRNA like molecules. We propose that the first editing step (gRNA-directed mRNA cleavage) and multicistronic precursor cleavage are directed by trans-acting short RNA molecules bound to the gRNA biding complex (GRBC). The editing endonucleases have been identified as RNase III-type enzymes (reviewed in (Aphasizhev and Aphasizheva, 2011), suggesting that members of the same family may participate in RNA-directed pre-mRNA cleavage.
4. Polyadenylation of messenger RNAs
Following excision from the multicistronic precursor, mitochondrial pre-mRNAs typically possess short 5′ monophosphorylated UTRs without discernable ribosome binding sites and short 3′ UTRs. In addition, 12 out of 18 annotated mitochondrial pre-mRNAs require U-insertion/ deletion editing to create open reading frames. Therefore, the ultimate task of producing translation-competent mRNAs can be tentatively divided into: 1) pre-mRNA stabilization; 2) reconstruction of the protein coding sequence by U-insertion/deletion RNA editing, if required; and 3) adding structural features that enable translation. The editing process (Fig. 2A) has been extensively described elsewhere (Aphasizhev and Aphasizheva, 2011;Hajduk and Ochsenreiter, 2010), but for the purposes of this review, we will keep in mind that editing of most mRNAs is initiated in the 3′ region and proceeds toward the 5′ end (Maslov and Simpson, 1992). Such 3′-5′ (mRNA) polarity is determined by sequential gRNA binding followed by editing, which generates a binding site for the next gRNA (Fig. 2B).
The insect (procyclic, PF) developmental form of African trypanosomes produces energy by mitochondrial cytochrome-mediated respiration using amino acids as major carbon source, while the mammalian (bloodstream form, BF) parasites rely on glycolysis to synthesize ATP (reviewed in (Van Hellemond et al., 2005)). It is thought that the mitochondrion of the bloodstream T. brucei consumes ATP to maintain transmembrane potential (Schnaufer et al., 2005;Brown et al., 2006). Importantly, mRNA editing was shown to be essential for the viability of both forms (Schnaufer et al., 2001), but the role of polyadenylation in producing translation-competent mRNAs and regulating their abundance remained unclear. Early comparative analysis of mRNA abundance and size distribution in insect and bloodstream forms left behind a rather convoluted picture which may be summarized as follows: 1) mitochondrial mRNAs possess short (20-50) or long (200-300) poly(A) tails; 2) mRNA editing and polyadenylation processes are developmentally regulated in a transcript-specific manner; 3) the editing status and the length of the poly(A) tail do not correlate with mRNA abundance and 4) pre-edited mRNAs tend to have short tails, while edited and never-edited molecules typically possess either short or long tails (Bhat et al., 1991;Bhat et al., 1992;Read et al., 1994;Feagin et al., 1987).
The ensuing studies of mRNA stability, in organello (Militello and Read, 2000;Ryan and Read, 2005), and RNA degradation activities in submitochondrial fractions (Ryan et al., 2003;Kao and Read, 2005) showed that the short poly(A) tail protects edited mRNAs against 3′-5′ degradation. Conversely, the same short tail reportedly stimulates degradation of pre-edited mRNAs and non-mitochondrial adenylated mRNAs in vitro. Furthermore, in organello, the mRNA degradation could be stimulated by exogenous UTP and this effect was dependent on RET1 TUTase activity. Although somewhat conflicting, these reports provided the first indications that the short poly(A) tail's function may “switch” from destabilizing in pre-edited mRNAs to stabilizing in edited transcripts. Indeed, very few editing events at the 3′ region are sufficient to induce this switch. The reconciliation of RNA analysis and in vitro data, however, was further complicated by reports of cleavage/polyadenylation and editing being independent events (Koslowsky and Yahampath, 1997;Militello and Read, 1999). Indeed, the functional significance of degrading adenylated pre-edited mRNAs and maintaining short and long-tailed mRNA populations was puzzling.
Search of the T. brucei genome (El-Sayed et al., 2005) for homologs of mitochondrial RET1 (Aphasizhev et al., 2002) and RET2 TUTases (Aphasizhev et al., 2003a;Panigrahi et al., 2003) defined an apparently monophyletic family (Aphasizhev, 2005;Aphasizhev and Aphasizheva, 2008) which included a third mitochondrial TUTase TbMEAT1 (Aphasizheva et al., 2009;Stagno et al., 2010), cytosolic TUTases TbTUT3 (Aphasizhev et al., 2004), TbTUT4 (Stagno et al., 2007b;Stagno et al., 2007a), TbncPAP1 and TbncPAP2 nuclear non-canonical poly(A) polymerases (Etheridge et al., 2009). Most importantly for our subject, mitochondrial (kinetoplast) poly(A) polymerases (KPAP) 1 (Etheridge et al., 2008) and 2 (Kao and Read, 2007) were among the nucleotidyl transferases identified in a biochemical/genetic pursuit of data mining predictions. The biological function and the nucleotide specificity of KPAP2 remain unclear; its RNAi knockdowns affected neither cell viability nor polyadenylation of mitochondrial mRNAs (Aphasizheva, unpublished observations and (Kao and Read, 2007)). On the other hand, KPAP1 repression led to a rapid decline in both short- and long-tailed forms of never-edited and fully-edited mRNAs followed by the loss of mitochondrial function and cell death (Etheridge et al., 2008). In contrast to in vitro mRNA decay results (Ryan et al., 2003;Kao and Read, 2005), the steady-state levels of pre-edited mRNAs in vivo were not affected by the loss of short A-tails, indicating their neutral role in stabilizing pre-edited mRNAs. In agreement with in vitro data, however, the short A-tail is required for maintaining RPS12 transcripts edited beyond the few sites at the 3′ region, thereby confirming a “switch” of its function into a stabilizing mode upon initiation of editing. Still more surprising, the long tail is added to completely or nearly-completely edited messenger RNA in the form of a 200-300 nt-long A/U heteropolymer. These findings clearly implicated the pre-editing addition of the short A-tail in stabilization of partially and fully edited mRNAs, but left open a number of questions: 1) what is the mechanism of the short A-tail's “switching” from neutral to stabilizing function; 2) how does completion of U-insertion/deletion editing at the 5′ region trigger random addition of As and Us, in ~7:3 ratio, to the 3′ end; 3) given that KPAP1 is a poly(A) polymerase, the enzyme(s) of which contribute uridines to the A/U-tail; and 4) what is the function of this bizarre A/U structure?
5. mRNA polyadenylation/uridylation factors: the emerging role of PPR proteins
Transient association of RET1 TUTase with KPAP1 (Etheridge et al., 2008), partial collapse of long tails in CO1 mRNA (Aphasizheva and Aphasizhev, 2010) when RET1 is repressed by RNAi and RET1's requirement for UTP-stimulated mRNA degradation in organello (Militello and Read, 2000;Ryan and Read, 2005) implicated this enzyme as a possible KPAP1 counterpart in the mRNA adenylation/uridylation process. However, RET1-catalyzed uridylation also affects nucleolytic processing and 3′ uridylation of gRNA and rRNAs (Aphasizheva and Aphasizhev, 2010), while KPAP1 is responsible for short A-tail addition to mRNAs (Etheridge et al., 2008). Proteomic analysis of the KPAP1 complex identified five pentatricopeptide repeat-containing (PPR) proteins, thus providing initial clues regarding the identity of mRNA-specific 3′ modification factors. PPR-containing RNA binding proteins are defined by the presence of 35-amino acid tandem repeats (Small and Peeters, 2000). In terrestrial plants, hundreds of non-redundant PPR factors control various aspects of organellar RNA processing, including editing, splicing, processing and stability (Schmitz-Linneweber and Small, 2008). In T. brucei, 28 PPR proteins, including those associated with the KPAP1 complex, were predicted by data mining (Pusnik et al., 2007) and 16 were detected experimentally in purified mitochondrial ribosomes (Zikova et al., 2008). At least some ribosome-associated PPRs are essential for rRNA biogenesis or stability, as evidenced by the loss of either 9S or 12S rRNAs in respective RNAi knockdowns (Pusnik et al., 2007). By functional characterization of KPAP1-associated PPR factors, Aphasizheva et al. identified a heterodimer of PPR proteins, termed kinetoplast polyadenylation/ uridylation factors (KPAFs) 1 and 2, as an inducer of the post-editing A/U-tail addition by KPAP1 and RET1 (Aphasizheva et al., 2011). Furthermore, edited transcripts bearing 200-300 nucleotide-long A/U-tails, but not short A-tails, were found in translating ribosomal complexes. Finally, KPAF1 repression was shown to inhibit A/U-addition, but not the short A-tail formation, and to block mitochondrial translation. To conclude, these studies established the KPAP1-catalyzed pre-editing short A-tail addition as required and sufficient for mRNA stabilization during and after the editing process. Conversely, the KPAF1-2 coordinated post-editing A/U-addition by RET1 and KPAP1 does not significantly affect mRNA stability; instead this unique structure enables mRNA binding to the ribosome, thus representing a defining cis-elements of translation-competent mRNAs in both procyclic and bloodstream developmental forms of T. brucei (Aphasizheva et al., 2011).
Considering the multitude of mitochondrial PPRs, which in plants often act as sequence-specific RNA binding proteins (Delannoy et al., 2007), one may envisage these factors acting in processes that remain poorly understood. For example, the “stability switch” phenomenon for the short A-tail functioning as a neutral or stabilizing cis-element depending on the mRNA's editing status may be explained in terms of PPR factor binding to the 3′ region. By analogy to PPR10 in maize chloroplasts (Pfalz et al., 2009;Prikryl et al., 2010), high-affinity binding of a PPR protein (Factor X, Fig. 3) at a specific site may define the 3′ terminus by protecting the RNA from exonucleolytic degradation and possibly recruiting the polyadenylation complex for a short A-tail addition. RNA editing then may remove this stabilizing PPR factor either because of sequence changes or by RNA helicase-driven displacement. In any event, in the absence of a trans-acting factor occupying the 3′ region, mRNA stability becomes dependent on a cis-element (short A-tail) as RNA editing proceeds toward the 5′ end (Fig. 3). Along the same lines, signaling between the completion of editing at the 5′ end and A/U-addition to the short A-tail at the 3′ end may be mediated by a PPR factor that recognizes a sequence created de novo by editing (Factor Y, Fig. 3). In this scenario, mRNA circularization is likely to be required for recruitment of RET1 and KPAFs at the 3′ end.
Fig. 3.
A model for the editing status-dependent ‘switch’ in short A-tail function. After precursor cleavage, pre-edited mRNA is protected against 3′-5′ degradation by virtue of hypothetical protein factor (X) binding to the 3′ region. Hence, the short A-tail is dispensable for pre-edited mRNA stabilization, and KPAP1 knockdown has no effect on mRNA abundance (Etheridge et al., 2008). Progression of editing through the 3′ region displaces factor X; therefore, mRNA stability becomes dependent on the short A-tail and, most likely, protein factors bound to the short tail, e.g., KPAP1. Thus, KPAP1 knockdown abolishes edited mRNAs. Generation of a new sequence by RNA editing leads to binding of a sequence-specific factor Y, mRNA circularization and recruitment of KPAF1 and 2, and RET1 to the 3′ end. A combination of KPAP1, RET1 and KPAFs is sufficient to induce A/U-tail formation in vitro (Aphasizheva et al., 2011).
6. RET1-catalyzed RNA uridylation is a major transcriptome-shaping force
The discovery of 3′ oligo (U)-tails in gRNAs (Blum and Simpson, 1990) and rRNAs (Adler et al., 1991) led to detection (Bakalara et al., 1989), purification and gene cloning (Aphasizhev et al., 2002) of the first RNA uridylyl transferase termed RNA editing TUTase 1 (RET1). Because RET1 RNAi knockdown triggered shortening of the gRNA population, this enzyme's function was initially associated with gRNA uridylation (Aphasizhev et al., 2003b). However, the mechanism by which RET1 knockdown blocked editing was unclear: the U-tail's participation in the editing process (direct effect) and/or U-tail requirement for gRNA maintenance in mitochondria (indirect effect) were among the most obvious culprits. The direct participation in editing may involve interaction with purine-rich pre-edited mRNA (Blum and Simpson, 1990). Alternatively, the U-tail may function as a landing pad for protein factors (Blom et al., 2001;Hayman and Read, 1999;Pelletier et al., 2000;Vanhamme et al., 1998). Although its role in editing remains debatable, the U-tail appears to be dispensable for gRNA stability in vivo. A combination of RNAi and dominant negative approaches with in vivo stability assays demonstrated that RET1 repression inhibits gRNA precursor processing but does not affect decay rates of mature gRNAs (Aphasizheva and Aphasizhev, 2010). Apparently, the U-tail is rapidly degraded unless it is continuously rebuilt by RET1, while the non-uridylated gRNAs remain relatively stable by virtue of binding to the gRNA binding complex (GRBC) (Weng et al., 2008;Aphasizheva and Aphasizhev, 2010;Ryan et al., 2006). Unlike maxicircle-encoded precursors, which span thousands of nucleotides, gRNAs are transcribed as uniformly sized (~800 nt) precursors. Mature gRNAs maintain 5′ triphosphates, indicating a lack of 5′ end processing while the 3′ end is formed by removal of a 3′ trailer and uridylation. Although the exact mechanism of RET1-controlled pre-gRNA processing remains to be elucidated, it can be concluded that RET1 TUTase activity and not the cryptic nuclease activity of this protein is required for nucleolytic processing of gRNA precursors (Aphasizheva and Aphasizhev, 2010) .
Most unexpectedly, precursors for 9S rRNA and some mRNAs also accumulated in RET1 RNAi cell lines (Aphasizheva and Aphasizhev, 2010). Thus, it appears that processing of both mini- and maxicircle-encoded RNA precursors is affected by RET1 repression. These effects are clearly indirect and likely to be caused by inhibited uridylation of RNAs that are required for precursor cleavage. To that end, a model was proposed in which the nucleolytic processing of maxicircle-encoded RNA precursors is directed (guided) by short, gRNA-like anti-sense molecules encoded elsewhere in kDNA. This would explain the alternative cleavage of precursors by stochastic binding of different anti-sense RNAs. In minicircles, anti-sense transcripts may originate via transcription of the opposite strand; indeed, gRNA- and gRNA precursor-sized anti-sense molecules have been detected (Aphasizheva and Aphasizhev, 2010). It is plausible that anti-sense RNAs are stabilized by RET1-catalyzed 3′- uridylation and are rapidly degraded in RET1-depleted cells. Although the mechanistic link between the anti-sense transcripts and pre-gRNA processing remains to be firmly established, we propose that the double-stranded region formed by overlapping 5′-ends of the opposing 600-800 nt transcription units may constitute a substrate for a specific processing nuclease (Fig. 4). Considering the length of gRNA precursors, these transcription units traverse almost the entire minicircle which in T. brucei may encode up to five gRNAs. This means that a primary transcript must contain more than one gRNA sequence, of which only the most 5′ end unit would be triphosphorylated. It remains to be established whether the 5′-triphosphate is necessary for gRNA function and/or stability. If this is indeed the case, then only the most 5′-end gRNA can be processed into a functional molecule. Clarification of the 3′ trailer's metabolic fate will be critical to understanding the entire gRNA processing pathway.
Interestingly, gRNAs are present at much higher steady-state levels than the corresponding anti-sense RNAs. Assuming that gRNAs enter the editing cascade as single-stranded molecules, the double-stranded RNA unwinding and asymmetric degradation of anti-sense molecules would be required to liberate functional gRNA from a 50-60 nt-long duplex. In sum, the current hypothesis suggests that following the cleavage reaction and duplex unwinding, gRNA and gRNA-sized anti-sense transcripts enter different pathways: gRNAs are loaded onto the gRNA binding complex and channeled to the editing process, while anti-sense molecules are targeted for degradation. It is unclear at which point uridylation takes place but given RET1's strong preference for single-stranded RNA, this process may be expected to take place after duplex unwinding. It is entirely possible that not only gRNAs, but also gRNA-like molecules directing nucleolytic processing of maxicircle transcripts, are loaded onto GRBC (Weng et al., 2008), which is likely to be the same particle as the putative mitochondrial RNA binding complex 1 (MRB1) identified by the Stuart laboratory (Panigrahi et al., 2007)). This would explain processing defects observed in RNAi knockdown of MRB1 subunit (Acestor et al., 2009).
Generation of short effector RNA molecules by cleaving double-stranded substrates and selective loading of one strand onto a protein complex is remarkably similar to RNAi/miRNA pathways. Along these lines, a prediction can be made that RNase III-like enzymes recognize a double-stranded region produced by overlapping 5′ ends of opposite transcription units and then cleave precursors at or near the ends of the duplex region. RNA helicase-driven duplex unwinding would be among the most obvious next step in this model (Fig. 4). While multiple representatives of DEA(H/D) RHA helicases are localized in the mitochondrion, reports of gRNA loss in knockdown of the GRBC-associated RNA editing helicase 2 (REH2) are consistent with its possible role in gRNA – anti-gRNA duplex unwinding (Hashimi et al., 2009;Hernandez et al., 2010).
7. Interplay of mRNA adenylation and uridylation
The first indications that uridylation may promote mRNA decay came from reports of UTP-stimulated degradation of adenylated RNAs in organello (Militello and Read, 2000;Ryan et al., 2003). This phenomenon was no longer observed in the mitochondrial fraction isolated from RET1-depleted parasites (Ryan and Read, 2005). Identification of mitochondrial poly(A) polymerase KPAP1 (Etheridge et al., 2008) and polyadenylation/uridylation factors KPAF1 and KPAF2 (Aphasizheva et al., 2011) demonstrated that short A-tails are required and sufficient for maintenance of most mRNAs, while long A/U-heteropolymers enable mRNA translation. However, recent work indicated that RET1 participates in two distinct modes of mRNA 3′-uridylation: formation of long A/U-tails and transcript-specific addition of continuous short U-tails (Aphasizheva et al., 2011).
Because RET1 RNAi ablates editing via blockade of gRNA biogenesis, the most convincing data suggesting that addition of continuous U-tails accelerates mRNA decay have been obtained for never-edited ND1 and MURF1 transcripts (Aphasizheva and Aphasizhev, 2010). Both RET1 repression and overexpression of a dominant negative mutant triggered a significant increase in steady-state levels of ND1 and MURF1 transcripts and stabilized MURF5 mRNA. Cloning and sequencing of ND1 mRNA 3′ ends from RET1 RNAi cells directly demonstrated that 3′ UTRs were generally longer and lacking short stretches of U otherwise present in the control cell line. This finding indicates that a transcript-specific addition of continuous short U-tails may act as a mechanism of selective post-transcriptional gene repression. In this context, stimulating RET1 activity by increasing UTP level in organello may have led to non-specific mRNA 3′-uridylation followed by rapid degradation. It is not clear whether UTP-stimulated mRNA decay represents a biologically relevant regulatory pathway (Militello and Read, 2000;Ryan and Read, 2005), but it is evident that the nature of transcript-specific RET1 activators or inhibitors needs to be further investigated.
8. What's next?
Post-transcriptional RNA processing is critical for organellar genome expression in all systems studied to date, but its intricacy in trypanosomes appears gratuitous by any account. While the evolutionary basis of such complexity remains contentious (Gray et al., 2010), biochemistry and genetics only recently began expanding beyond RNA editing. Discoveries of RNA adenylation (KPAP1) and RNA uridylation (RET1) enzymes underscored the key roles and tightly regulated nature of 3′ modifications for generation and stabilization of functional RNAs. However, few KPAP1 and RET1 functions could be readily attributed to their intrinsic RNA substrate binding properties. While the length of mRNA short A-tail is consistent with KPAP1's in vitro processivity (Etheridge et al., 2008), RET1 TUTase is capable of adding several hundred instead of 15-20 uridines to single-stranded RNAs (Aphasizhev et al., 2002). Identification of KPAF1 and KPAF2 as mRNA polyadenylation/ uridylation factors points to the expanded family of pentatricopeptide repeat proteins as a rich source of candidates for “missing” regulatory functions (Pusnik et al., 2007;Aphasizheva et al., 2011). Binding of PPR proteins to specific RNA sequences may define mRNA 3′ ends by protecting against nuclease degradation or trigger 3′ A/U addition upon completion of RNA editing. Given the uridylation-induced degradation of select mRNAs, the existence of factors that prevent fortuitous mRNA U-tailing may be anticipated. Finally, the deep sequencing of short mitochondrial RNAs is bound to clarify the possible role of anti-sense transcription and gRNA-like molecule in the processing of maxiand minicircle-encoded multicistronic precursors.
Acknowledgements
We thank Marian Waterman and Yongsheng Shi for discussions. The work in authors’ laboratory is supported by NIH grants AI091914 and AI083863.
Footnotes
Publisher's Disclaimer: This is a PDF file of an unedited manuscript that has been accepted for publication. As a service to our customers we are providing this early version of the manuscript. The manuscript will undergo copyediting, typesetting, and review of the resulting proof before it is published in its final citable form. Please note that during the production process errors may be discovered which could affect the content, and all legal disclaimers that apply to the journal pertain.
References
- Acestor N, Panigrahi AK, Carnes J, Zikova A, Stuart KD. The MRB1 complex functions in kinetoplastid RNA processing. RNA. 2009;15:277–286. doi: 10.1261/rna.1353209. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Adler BK, Harris ME, Bertrand KI, Hajduk SL. Modification of Trypanosoma brucei mitochondrial rRNA by posttranscriptional 3' polyuridine tail formation. Mol. Cell. Biol. 1991;11:5878–5884. doi: 10.1128/mcb.11.12.5878. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Alfonzo JD, Soll D. Mitochondrial tRNA import--the challenge to understand has just begun. Biol Chem. 2009;390:717–722. doi: 10.1515/BC.2009.101. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Alfonzo JD, Thiemann OH, Simpson L. Purification and characterization of MAR1 - A mitochondrial associated ribonuclease from Leishmania tarentolae. J. Biol. Chem. 1998;273:30003–30011. doi: 10.1074/jbc.273.45.30003. [DOI] [PubMed] [Google Scholar]
- Aphasizhev R. RNA uridylyltransferases. Cell. Mol. Life Sci. 2005;62:2194–2203. doi: 10.1007/s00018-005-5198-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Aphasizhev R, Aphasizheva I. Terminal RNA uridylyltransferases of trypanosomes. Biochim. Biophys. Acta. 2008;1779:270–280. doi: 10.1016/j.bbagrm.2007.12.007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Aphasizhev R, Aphasizheva I. Uridine Insertion/Deletion mRNA Editing In Trypanosomes: A Playground for Non-templated Information Transfer. Wiley Interdisciplinary Reviews: RNA . 2011. [DOI] [PMC free article] [PubMed]
- Aphasizhev R, Aphasizheva I, Nelson RE, Gao G, Simpson AM, Kang X, Falick AM, Sbicego S, Simpson L. Isolation of a U-insertion/deletion editing complex from Leishmania tarentolae mitochondria. EMBO J. 2003a;22:913–924. doi: 10.1093/emboj/cdg083. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Aphasizhev R, Aphasizheva I, Simpson L. A tale of two TUTases. Proc. Natl. Acad. Sci. U.S.A. 2003b;100:10617–10622. doi: 10.1073/pnas.1833120100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Aphasizhev R, Aphasizheva I, Simpson L. Multiple terminal uridylyltransferases of trypanosomes. FEBS Lett. 2004;572:15–18. doi: 10.1016/j.febslet.2004.07.004. [DOI] [PubMed] [Google Scholar]
- Aphasizhev R, Sbicego S, Peris M, Jang SH, Aphasizheva I, Simpson AM, Rivlin A, Simpson L. Trypanosome Mitochondrial 3' Terminal Uridylyl Transferase (TUTase): The Key Enzyme in U-insertion/deletion RNA Editing. Cell. 2002;108:637–648. doi: 10.1016/s0092-8674(02)00647-5. [DOI] [PubMed] [Google Scholar]
- Aphasizhev R, Simpson L. Isolation and characterization of a U-specific 3'-5' exonuclease from mitochondria of Leishmania tarentolae. J. Biol. Chem. 2001;276:21280–21284. doi: 10.1074/jbc.M100297200. [DOI] [PubMed] [Google Scholar]
- Aphasizheva I, Aphasizhev R. RET1-catalyzed Uridylylation Shapes the Mitochondrial Transcriptome in Trypanosoma brucei. Mol. Cell Biol. 2010;30:1555–1567. doi: 10.1128/MCB.01281-09. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Aphasizheva I, Maslov D, Wang X, Huang L, Aphasizhev R. Pentatricopeptide Repeat Proteins Stimulate mRNA Adenylation/Uridylation to Activate Mitochondrial Translation in Trypanosomes. Mol. Cell. 2011 doi: 10.1016/j.molcel.2011.02.021. in press. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Aphasizheva I, Ringpis GE, Weng J, Gershon PD, Lathrop RH, Aphasizhev R. Novel TUTase associates with an editosome-like complex in mitochondria of Trypanosoma brucei. RNA. 2009;15:1322–1337. doi: 10.1261/rna.1538809. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bakalara N, Simpson AM, Simpson L. The Leishmania kinetoplast-mitochondrion contains terminal uridylyltransferase and RNA ligase activities. J. Biol. Chem. 1989;264:18679–18686. [PubMed] [Google Scholar]
- Bhat GJ, Myler PJ, Stuart K. The two ATPase 6 mRNAs of Leishmania tarentolae differ at their 3' ends. Mol. Biochem. Parasitol. 1991;48:139–150. doi: 10.1016/0166-6851(91)90110-r. [DOI] [PubMed] [Google Scholar]
- Bhat GJ, Souza AE, Feagin JE, Stuart K. Transcript-specific developmental regulation of polyadenylation in Trypanosoma brucei mitochondria. Mol. Biochem. Parasitol. 1992;52:231–240. doi: 10.1016/0166-6851(92)90055-o. [DOI] [PubMed] [Google Scholar]
- Blom D, Burg J, Breek CK, Speijer D, Muijsers AO, Benne R. Cloning and characterization of two guide RNA-binding proteins from mitochondria of Crithidia fasciculata: gBP27, a novel protein, and gBP29, the orthologue of Trypanosoma brucei gBP21. Nucleic Acids Res. 2001;29:2950–2962. doi: 10.1093/nar/29.14.2950. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Blum B, Bakalara N, Simpson L. A model for RNA editing in kinetoplastid mitochondria: “Guide” RNA molecules transcribed from maxicircle DNA provide the edited information. Cell. 1990;60:189–198. doi: 10.1016/0092-8674(90)90735-w. [DOI] [PubMed] [Google Scholar]
- Blum B, Simpson L. Guide RNAs in kinetoplastid mitochondria have a nonencoded 3' oligo-(U) tail involved in recognition of the pre-edited region. Cell. 1990;62:391–397. doi: 10.1016/0092-8674(90)90375-o. [DOI] [PubMed] [Google Scholar]
- Brown SV, Hosking P, Li J, Williams N. ATP synthase is responsible for maintaining mitochondrial membrane potential in bloodstream form Trypanosoma brucei. Eukaryot Cell. 2006;5:45–53. doi: 10.1128/EC.5.1.45-53.2006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Clement SL, Koslowsky DJ. Unusual organization of a developmentally regulated mitochondrial RNA polymerase (TBMTRNAP) gene in Trypanosoma brucei. Gene. 2001;272:209–218. doi: 10.1016/s0378-1119(01)00538-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Clement SL, Mingler MK, Koslowsky DJ. An intragenic guide RNA location suggests a complex mechanism for mitochondrial gene expression in Trypanosoma brucei. Eukaryot. Cell. 2004;3:862–869. doi: 10.1128/EC.3.4.862-869.2004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Delannoy E, Stanley WA, Bond CS, Small ID. Pentatricopeptide repeat (PPR) proteins as sequence-specificity factors in post-transcriptional processes in organelles. Biochem. Soc. Trans. 2007;35:1643–1647. doi: 10.1042/BST0351643. [DOI] [PubMed] [Google Scholar]
- El-Sayed NM, Myler PJ, Blandin G, Berriman M, Crabtree J, Aggarwal G, Caler E, Renauld H, et al. Comparative genomics of trypanosomatid parasitic protozoa. Science. 2005;309:404–409. doi: 10.1126/science.1112181. [DOI] [PubMed] [Google Scholar]
- Etheridge RD, Aphasizheva I, Gershon PD, Aphasizhev R. 3' adenylation determines mRNA abundance and monitors completion of RNA editing in T. brucei mitochondria. EMBO J. 2008;27:1596–1608. doi: 10.1038/emboj.2008.87. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Etheridge RD, Clemens DM, Aphasizhev R. Identification and characterization of nuclear non-canonical poly(A) polymerases from Trypanosoma brucei. Mol. Biochem. Parasitol. 2009;164:66–73. doi: 10.1016/j.molbiopara.2008.11.004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Feagin J, Jasmer D, Stuart K. Developmentally regulated addition of nucleotides within apocytochrome b transcripts in Trypanosoma brucei. Cell. 1987;49:337–345. doi: 10.1016/0092-8674(87)90286-8. [DOI] [PubMed] [Google Scholar]
- Grams J, McManus MT, Hajduk SL. Processing of polycistronic guide RNAs is associated with RNA editing complexes in Trypanosoma brucei. EMBO J. 2000;19:5525–5532. doi: 10.1093/emboj/19.20.5525. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Grams J, Morris JC, Drew ME, Wang Z, Englund PT, Hajduk SL. A trypanosome mitochondrial RNA polymerase is required for transcription and replication. J. Biol. Chem. 2002;277:16952–16959. doi: 10.1074/jbc.M200662200. [DOI] [PubMed] [Google Scholar]
- Gray MW, Lukes J, Archibald JM, Keeling PJ, Doolittle WF. Cell biology. Irremediable complexity? Science. 2010;330:920–921. doi: 10.1126/science.1198594. [DOI] [PubMed] [Google Scholar]
- Hajduk S, Ochsenreiter T. RNA editing in kinetoplastids. RNA Biol. 2010;7:229–236. doi: 10.4161/rna.7.2.11393. [DOI] [PubMed] [Google Scholar]
- Hashimi H, Cicova Z, Novotna L, Wen YZ, Lukes J. Kinetoplastid guide RNA biogenesis is dependent on subunits of the mitochondrial RNA binding complex 1 and mitochondrial RNA polymerase. RNA. 2009;15:588–599. doi: 10.1261/rna.1411809. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hayman ML, Read LK. Trypanosoma brucei RBP16 is a mitochondrial Y-box family protein with guide RNA binding activity. J. Biol. Chem. 1999;274:12067–12074. doi: 10.1074/jbc.274.17.12067. [DOI] [PubMed] [Google Scholar]
- Hernandez A, Madina BR, Ro K, Wohlschlegel JA, Willard B, Kinter MT, Cruz-Reyes J. REH2 RNA helicase in kinetoplastid mitochondria: ribonucleoprotein complexes and essential motifs for unwinding and guide RNA (gRNA) binding. J. Biol. Chem. 2010;285:1220–1228. doi: 10.1074/jbc.M109.051862. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hong M, Simpson L. Genomic organization of Trypanosoma brucei kinetoplast DNA minicircles. Protist. 2003;154:265–279. doi: 10.1078/143446103322166554. [DOI] [PubMed] [Google Scholar]
- Jasmer D, Stuart K. Sequence organization in African trypanosome minicircles is defined by 18 base pair inverted repeats. Mol. Biochem. Parasitol. 1986;18:321–332. doi: 10.1016/0166-6851(86)90089-7. [DOI] [PubMed] [Google Scholar]
- Kao CY, Read LK. Opposing effects of polyadenylation on the stability of edited and unedited mitochondrial RNAs in Trypanosoma brucei. Mol. Cell Biol. 2005;25:1634–1644. doi: 10.1128/MCB.25.5.1634-1644.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kao CY, Read LK. Targeted depletion of a mitochondrial nucleotidyltransferase suggests the presence of multiple enzymes that polymerize mRNA 3' tails in Trypanosoma brucei mitochondria. Mol. Biochem. Parasitol. 2007;154:158–169. doi: 10.1016/j.molbiopara.2007.04.014. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Koslowsky DJ, Yahampath G. Mitochondrial mRNA 3' cleavage polyadenylation and RNA editing in Trypanosoma brucei are independent events. Mol. Biochem. Parasitol. 1997;90:81–94. doi: 10.1016/s0166-6851(97)00133-3. [DOI] [PubMed] [Google Scholar]
- Lange H, Sement FM, Canaday J, Gagliardi D. Polyadenylation-assisted RNA degradation processes in plants. Trends Plant. Sci. 2009;14:497–504. doi: 10.1016/j.tplants.2009.06.007. [DOI] [PubMed] [Google Scholar]
- Liu B, Liu Y, Motyka SA, Agbo EE, Englund PT. Fellowship of the rings: the replication of kinetoplast DNA. Trends Parasitol. 2005;21:363–369. doi: 10.1016/j.pt.2005.06.008. [DOI] [PubMed] [Google Scholar]
- Madej MJ, Alfonzo JD, Huttenhofer A. Small ncRNA transcriptome analysis from kinetoplast mitochondria of Leishmania tarentolae. Nucleic Acids Res. 2007;35:1544–1554. doi: 10.1093/nar/gkm004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Madej MJ, Niemann M, Huttenhofer A, Goringer HU. Identification of novel guide RNAs from the mitochondria of Trypanosoma brucei. RNA Biol. 2008;5:84–91. doi: 10.4161/rna.5.2.6043. [DOI] [PubMed] [Google Scholar]
- Maslov DA, Simpson L. The polarity of editing within a multiple gRNA-mediated domain is due to formation of anchors for upstream gRNAs by downstream editing. Cell. 1992;70:459–467. doi: 10.1016/0092-8674(92)90170-h. [DOI] [PubMed] [Google Scholar]
- Mattiacio JL, Read LK. Roles for TbDSS-1 in RNA surveillance and decay of maturation by-products from the 12S rRNA locus. Nucleic Acids Res. 2008;36:319–329. doi: 10.1093/nar/gkm690. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Michelotti EF, Harris ME, Adler B, Torri AF, Hajduk SL. Trypanosoma brucei mitochondrial ribosomal RNA synthesis, processing and developmentally regulated expression. Mol. Biochem. Parasitol. 1992;54:31–42. doi: 10.1016/0166-6851(92)90092-x. [DOI] [PubMed] [Google Scholar]
- Militello KT, Read LK. Coordination of kRNA editing and polyadenylation in Trypanosoma brucei mitochondria: complete editing is not required for long poly(A) tract addition. Nucleic Acids Res. 1999;27:1377–1385. doi: 10.1093/nar/27.5.1377. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Militello KT, Read LK. UTP-dependent and -independent pathways of mRNA turnover in Trypanosoma brucei mitochondria. Mol. Cell. Biol. 2000;20:2308–2316. doi: 10.1128/mcb.20.7.2308-2316.2000. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ochsenreiter T, Cipriano M, Hajduk SL. KISS: the kinetoplastid RNA editing sequence search tool. RNA. 2007;13:1–4. doi: 10.1261/rna.232907. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Panigrahi AK, Schnaufer A, Ernst NL, Wang B, Carmean N, Salavati R, Stuart K. Identification of novel components of Trypanosoma brucei editosomes. RNA. 2003;9:484–492. doi: 10.1261/rna.2194603. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Panigrahi AK, Zikova A, Dalley RA, Acestor N, Ogata Y, Anupama A, Myler PJ, Stuart KD. Mitochondrial complexes in trypanosoma brucei: a novel complex and a unique oxidoreductase complex. Mol. Cell. Proteomics. 2007;7:534–545. doi: 10.1074/mcp.M700430-MCP200. [DOI] [PubMed] [Google Scholar]
- Pelletier M, Miller MM, Read LK. RNA-binding properties of the mitochondrial Y-box protein RBP16. Nucleic Acids Res. 2000;28:1266–1275. doi: 10.1093/nar/28.5.1266. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pfalz J, Bayraktar OA, Prikryl J, Barkan A. Site-specific binding of a PPR protein defines and stabilizes 5' and 3' mRNA termini in chloroplasts. EMBO J. 2009;28:2042–2052. doi: 10.1038/emboj.2009.121. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pollard VW, Rohrer SP, Michelotti EF, Hancock K, Hajduk SL. Organization of minicircle genes for guide RNAs in Trypanosoma brucei. Cell. 1990;63:783–790. doi: 10.1016/0092-8674(90)90144-4. [DOI] [PubMed] [Google Scholar]
- Prikryl J, Rojas M, Schuster G, Barkan A. Mechanism of RNA stabilization and translational activation by a pentatricopeptide repeat protein. Proc. Natl. Acad. Sci. U.S. A. 2010;108:415–420. doi: 10.1073/pnas.1012076108. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pusnik M, Small I, Read LK, Fabbro T, Schneider A. Pentatricopeptide repeat proteins in Trypanosoma brucei function in mitochondrial ribosomes. Mol. Cell. Biol. 2007;27:6876–6888. doi: 10.1128/MCB.00708-07. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Read LK, Stankey KA, Fish WR, Muthiani AM, Stuart K. Developmental regulation of RNA editing and polyadenylation in four life cycle stages of Trypanosoma congolense. Mol. Biochem. Parasitol. 1994;68:297–306. doi: 10.1016/0166-6851(94)90174-0. [DOI] [PubMed] [Google Scholar]
- Ryan CM, Kao CY, Sleve DA, Read LK. Biphasic decay of guide RNAs in Trypanosoma brucei. Mol. Biochem. Parasitol. 2006;146:68–77. doi: 10.1016/j.molbiopara.2005.10.019. [DOI] [PubMed] [Google Scholar]
- Ryan CM, Militello KT, Read LK. Polyadenylation regulates the stability of Trypanosoma brucei mitochondrial RNAs. J. Biol. Chem. 2003;278:32753–32762. doi: 10.1074/jbc.M303552200. [DOI] [PubMed] [Google Scholar]
- Ryan CM, Read LK. UTP-dependent turnover of Trypanosoma brucei mitochondrial mRNA requires UTP polymerization and involves the RET1 TUTase. RNA. 2005;11:763–773. doi: 10.1261/rna.7248605. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Salavati R, Panigrahi AK, Stuart KD. Mitochondrial ribonuclease P activity of Trypanosoma brucei. Mol. Biochem. Parasitol. 2001;115:109–117. doi: 10.1016/s0166-6851(01)00273-0. [DOI] [PubMed] [Google Scholar]
- Schmitz-Linneweber C, Small I. Pentatricopeptide repeat proteins: a socket set for organelle gene expression. Trends Plant Sci. 2008;13:663–670. doi: 10.1016/j.tplants.2008.10.001. [DOI] [PubMed] [Google Scholar]
- Schnaufer A, Panigrahi AK, Panicucci B, Igo RP, Salavati R, Stuart K. An RNA ligase essential for RNA editing and survival of the bloodstream form of Trypanosoma brucei. Science. 2001;291:2159–2161. doi: 10.1126/science.1058955. [DOI] [PubMed] [Google Scholar]
- Schnaufer A, Clark-Walker GD, Steinberg AG, Stuart K. The F1-ATP synthase complex in bloodstream stage trypanosomes has an unusual and essential function. EMBO J. 2005;24:4029–4040. doi: 10.1038/sj.emboj.7600862. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sloof P, De Haan A, Eier W, Van Iersel M, Boel E, Van Steeg H, Benne R. The nucleotide sequence of the variable region in Trypanosoma brucei completes the sequence analysis of the maxicircle component of mitochondrial kinetoplast DNA. Mol. Biochem. Parasitol. 1992;56:289–300. doi: 10.1016/0166-6851(92)90178-m. [DOI] [PubMed] [Google Scholar]
- Small ID, Peeters N. The PPR motif - a TPR-related motif prevalent in plant organellar proteins. Trends Biochem. Sci. 2000;25:46–47. doi: 10.1016/s0968-0004(99)01520-0. [DOI] [PubMed] [Google Scholar]
- Stagno J, Aphasizheva I, Aphasizhev R, Luecke H. Dual Role of the RNA Substrate in Selectivity and Catalysis by Terminal Uridylyl Transferases. Proc. Natl. Acad. Sci. U.S. A. 2007a;104:14634–14639. doi: 10.1073/pnas.0704259104. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Stagno J, Aphasizheva I, Bruystens J, Luecke H, Aphasizhev R. Structure of the mitochondrial editosome-like complex associated TUTase 1 reveals divergent mechanisms of UTP selection and domain organization. J. Mol. Biol. 2010;399:464–475. doi: 10.1016/j.jmb.2010.04.021. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Stagno J, Aphasizheva I, Rosengarth A, Luecke H, Aphasizhev R. UTP-bound and Apo structures of a minimal RNA uridylyltransferase. J. Mol. Biol. 2007b;366:882–899. doi: 10.1016/j.jmb.2006.11.065. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Stern DB, Goldschmidt-Clermont M, Hanson MR. Chloroplast RNA metabolism. Annu. Rev. Plant. Biol. 2010;61:125–155. doi: 10.1146/annurev-arplant-042809-112242. [DOI] [PubMed] [Google Scholar]
- Stuart K, Brun R, Croft S, Fairlamb A, Gurtler RE, McKerrow J, Reed S, Tarleton R. Kinetoplastids: related protozoan pathogens, different diseases. J. Clin. Invest. 2008;118:1301–1310. doi: 10.1172/JCI33945. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Van Hellemond JJ, Opperdoes FR, Tielens AG. The extraordinary mitochondrion and unusual citric acid cycle in Trypanosoma brucei. Biochem. Soc. Trans. 2005;33:967–971. doi: 10.1042/BST20050967. [DOI] [PubMed] [Google Scholar]
- Vanhamme L, Perez-Morga D, Marchal C, Speijer D, Lambert L, Geuskens M, Alexandre S, Ismaïli N, Göringer U, Benne R, Pays E. Trypanosoma brucei TBRGG1, a mitochondrial oligo(U)-binding protein that co-localizes with an in vitro RNA editing activity. J Biol Chem. 1998;273:21825–21833. doi: 10.1074/jbc.273.34.21825. [DOI] [PubMed] [Google Scholar]
- Wang Z, Morris JC, Drew ME, Englund PT. Inhibition of Trypanosoma brucei gene expression by RNA interference using an integratable vector with opposing T7 promoters. J. Biol. Chem. 2000;275:40174–40179. doi: 10.1074/jbc.M008405200. [DOI] [PubMed] [Google Scholar]
- Weng J, Aphasizheva I, Etheridge RD, Huang L, Wang X, Falick AM, Aphasizhev R. Guide RNA-Binding Complex from Mitochondria of Trypanosomatids. Mol. Cell. 2008;32:198–209. doi: 10.1016/j.molcel.2008.08.023. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zikova A, Panigrahi AK, Dalley RA, Acestor N, Anupama A, Ogata Y, Myler PJ, Stuart K. Trypanosoma brucei mitochondrial ribosomes: affinity purification and component identification by mass spectrometry. Mol. Cell. Proteomics. 2008;7:1286–1296. doi: 10.1074/mcp.M700490-MCP200. [DOI] [PMC free article] [PubMed] [Google Scholar]