Abstract
KLF2 is a Krüppel-like zinc-finger transcription factor required for blood vessel, lung, T-cell and erythroid development. KLF2-/- mice die by embryonic day 14.5 (E14.5), due to hemorrhaging and heart failure. In KLF2-/- embryos, β-like globin gene expression is reduced, and E10.5 erythroid cells exhibit abnormal morphology. In this study, other genes regulated by KLF2 were identified by comparing E9.5 KLF2-/- and wild-type (WT) yolk sac erythroid precursor cells, using laser capture microdissection and microarray assays. One hundred and ninety-six genes exhibited significant differences in expression between KLF2-/- and WT; eighty-nine of these are downregulated in KLF2-/-. Genes involved in cell migration, differentiation and development are over-represented in the KLF2-regulated gene list. The SOX2 gene, encoding a pluripotency factor, is regulated by KLF2 in both ES and embryonic erythroid cells. Previous work had identified genes with erythroid-enriched expression in the yolk sac. The erythroid-enriched genes reelin, adenylate cyclase 7, cytotoxic T lymphocyte-associated protein 2 alpha, and CD24a antigen are downregulated in KLF2-/- compared to WT, and are therefore candidates for controlling primitive erythropoiesis. Each of these genes contains a putative KLF2 binding site(s) in its promoter and/or an intron. Reelin has an established role in neuronal development. Luciferase reporter assays demonstrated that KLF2 directly transactivates the reelin promoter in erythroid cells, validating this approach to identify KLF2 target genes.
Keywords: yolk sac, embryonic erythropoiesis, KLF2, expression profiling, laser capture microdissection
Introduction
Mammalian erythropoiesis occurs in successive primitive and definitive phases. Primitive or embryonic erythropoiesis begins in the mouse yolk sac at approximately embryonic day 7.5 (E7.5) and results in large nucleated erythroblasts that produce the embryonic globins. Definitive or adult erythropoiesis begins in the fetal liver at approximately E10.5. Definitive erythroid cells are small, enucleated and synthesize the adult globins. Less is known about the genes that regulate primitive, as opposed to definitive, erythropoiesis. Identifying genes that are important for embryonic erythroid development could provide new therapeutic approaches for sickle-cell anemia and β-thalassemia. Gene ablation studies in the mouse have established the essential roles of transcription factors (GATA-1, GATA-2, LMO2, EKLF, KLF2), signaling and growth factors (BMP4, VEGFA, FLK1), and proto-oncogenes/leukemia genes (such as TAL1) in primitive erythropoiesis [1-7].
The Krüppel-like factor (KLF) family has 17 members, each with 3 Cys2/His2 zinc finger DNA-binding domains (reviewed in [8-10]). The KLF transcription factors control cellular proliferation, differentiation, apoptosis and embryonic development. The CACCC elements in the human and mouse β-like globin promoters are important regulators of globin gene expression, and consensus binding sites for KLF factors. Erythroid Krüppel-like factor (EKLF or KLF1) positively regulates adult β-globin gene expression. More recently, it has become clear that EKLF also plays an essential role in embryonic erythropoiesis and globin gene expression [11-14].
Phylogenetic analyses indicate that EKLF and KLF2 belong to a subfamily of KLFs with 90% similarity in their zinc finger DNA binding domains [8,9,15]. KLF2 is expressed in a variety of tissues and is necessary for blood vessel, lung, T-cell and embryonic erythroid development [16-20]. KLF2-/- mice die between E12.5 and E14.5 due to hemorrhaging and cardiac failure [16,17,21]. KLF2 positively regulates the expression of the mouse and human embryonic globin genes [20]. In addition, KLF2-/- E10.5 primitive yolk sac erythroid precursor cells are abnormally shaped, displaying pseudopodia-like appendages. Recent evidence indicates that the EKLF and KLF2 genes interact in embryonic erythropoiesis, resulting in a more severe phenotype in double than in single null embryos [14].
Because morphological changes are observed in KLF2-/- embryonic erythroid cells, it seems likely that genes other than the globin genes are normally regulated by KLF2 in these cells. Previously, we used laser capture microdissection (LCM) to isolate E9.5 erythroid and epithelial cells from frozen embryonic yolk sac sections and generated high quality expression profiles [22]. Sixty-seven unique erythroid-enriched genes were identified, and are candidates for positively regulating primitive erythropoiesis. Because KLF2 is essential for primitive erythropoiesis, in this study, we determined which genes are differentially expressed between wildtype (WT) and KLF2-/- embryonic erythroid cells. A biological filter was then applied to determine which of the genes that are downregulated in KLF2-/- embryonic erythroid cells also have an erythroid-enriched expression pattern, as determined in the previous work [22]. Using erythroid-enriched genes as a biological filter, the aim was to focus on genes more likely to control primitive erythropoiesis. A limited number of genes are erythroid-enriched and downregulated in KLF2-/- compared to wildtype (WT) embryonic erythroid cells. The microarray expression assays showed that KLF2 acts directly or indirectly to positively regulate 89 genes. In addition to the globin genes, four of the 89 KLF2-regulated genes are also erythroid-enriched [22], and these genes encode the cell signaling factors CD24a antigen, cytotoxic T-lymphocyte associated protein 2 alpha (Ctla2a), adenylate cyclase 7 and reelin. These genes are candidates for controlling embryonic erythropoiesis. We also show that KLF2 directly regulates the reelin gene, which has an established role in nervous system development.
Material and Methods
Matings, sample preparation and microarray analyses
KLF2+/- 129/Bl6/Swiss mice [17] were bred with female FVB/N inbred mice for at least 5 generations to generate KLF2+/- offspring with at least 97% FVB/N character. These KLF2+/-mice were used in timed matings to obtain E9.5 KLF2-/- embryonic yolk sacs for freezing, cryosectioning and laser capture microdissection [22]. A small portion of the embryo tail was used for PCR genotyping. For each of the 4 erythroid KLF2-/- microarrays, approximately 4,000 erythroid cells were collected from 30 to 80 microscope slides, using 2 to 4 embryonic yolk sacs obtained from 2 different pregnant females. RNA extractions from the 4 KLF2-/- erythroid samples yielded 10 to 20 ng of RNA, and quality was assessed using capillary electrophoresis, microarray hybridizations and RNA digestion plots as previously described [23]. Fifteen μg of labeled erythroid cRNA was fragmented and 10 μg was hybridized to Affymetrix GeneChip® Mouse Genome 430A 2.0 arrays for 18 hours (Affymetrix Inc.). The Robust Multi-average (RMA) algorithm [24] was used to obtain probe set expression summaries which were calculated using the rma function available in Bioconductor in the affy package, within the R programming environment [25-27].
Statistical analysis, Ingenuity pathway analysis and gene classification
An extension to the probe-level S-score method for performing probe set level hypothesis tests when comparing two GeneChips [28], which is capable of comparing two independent conditions where more than one GeneChip is available for each condition, is called the Multi-Chip S-score [29]. The Multi-Chip Significance score method is available in the publicly available sscore package [30] and was used to compare the 4 wildtype and KLF2-/- GeneChips to identify significant differences in gene expression between these types of erythroid cells. P-values were calculated from absolute S-score values and then used to obtain q-values. A q-value <0.05 indicates a false discovery rate (FDR) <5%.
Ingenuity Pathway Analysis (IPA, Ingenuity Systems, Mountain View, CA; (http://www.ingenuity.com) was used to determine how differentially expressed genes interact in erythroid cells. The KLF2-regulated genes were mapped to their corresponding gene object/symbol in the IPA biological knowledge database. These focus genes were used as the basis to generate significant biological networks.
Functional gene categories were determined with GO (Gene Ontology) using DAVID (Database Annotation Visualization and Integrated Discovery). In addition, a gene annotation enrichment analysis was performed using DAVID, to determine whether any of the gene categories were over-represented in the gene list compared to their representation in the genome.
cDNA synthesis and quantitative real-time RT-PCR (qRT-PCR)
For the qRT-PCR verification of microarray data, cDNA was prepared from total RNA from between approximately 1,500 and 2,900 microdissected erythroid cells, as previously described [23]. Oligonucleotide primers were designed using PrimerExpress (Applied Biosystems), and these sequences are indicated in Table 1. The NCBI database (http://www.ncbi.nih.gov) was used to establish that the primers are gene specific. qRT-PCR experiments were performed using the ABI-Prism 7300 system (Applied Biosystems, Foster City, CA, USA). A pre-designed Taqman probe and primer set was used for qRT-PCR for cyclophilin A mRNA (Applied Biosystems, Foster City, CA, USA), which was the internal standard to normalize the expression data. All other reactions were performed with SYBR Green chemistry, and a dissociation curve was used to verify that only a single product was amplified. qRT-PCR was performed in duplicate using three biological replicates.
Table 1.
Primer sequences for real-rime PCR.
| Gene symbol | Forward primer | Reverse primer |
|---|---|---|
| CD24a | 5′TCAGGCCAGGAAACGTCTCTA3′ | 5′TCTTTCTTCTGATCACATTGGA′ |
| Ctla2a | 5′TCTGTCTCCTGGTATGAGAGGAATG3′ | 5′CAAAGCAGGTGCTGGAAGCT3′ |
| Reln | 5′CAAGAACAATACCGCTGATTGG3′ | 5′GATGTGGATGACTGTGCTCACA 3′ |
Transient co-transfections and reporter gene assays
The human reelin wildtype and mutant promoter/luciferase constructs were previously described [31,32]. The reelin –514 minimal promoter (-514 to +76 bp) was subcloned into a PGL3-basic luciferase construct. This region contains three putative Sp1/KLF consensus binding sites located at -230, -180 and -150. These binding sites were individually mutated to create three mutant constructs (mSp1-230, mSp1-180, and mSp1-150). The KLF2 cDNA expression construct driven by the CMV (cytomegalovirus) immediate–early promoter was described by Anderson et al. [33].
Transfection assays were performed with 1×106 log-phase K562 cells and Effectene reagent (Qiagen), according to the manufacturer's protocol with slight modifications. Briefly, 0.05 μg of a reelin promoter-luciferase construct (wildtype, mSp1-230, mSp1-180, mSp1-150) and 0.35 μg of the KLF2 expression construct or PMT2-empty vector (control) was transfected. The promoter-less PGL3 basic-luciferase construct was transfected as a negative control. Two different plasmid preparations per construct were transfected and the transfection assays were repeated at least 4 times and the mean for the trials was determined. Luciferase activities were determined for equal amounts of protein from cell lysates. The amount of luciferase activity for transfections of the wildtype construct was set at 1, and three different data comparisons were performed. For the initial comparison, transfections with the reelin wildtype promoter-luciferase construct were compared to wildtype plus the KLF2 expression construct. In the second, wildtype was compared to each of the mutant promoter-luciferase constructs. An additional comparison was conducted between wildtype plus KLF2 and each of the three mutant constructs plus KLF2. The statistical analyses were performed using a two-sample Student's t test, and all findings were judged to be significant using an α=0.05 level of significance.
Results and Discussion
Because morphological changes are observed in KLF2-/- embryonic erythroid cells, it seems likely that genes other than the globin genes are normally regulated by KLF2 in these cells. The objective of this study was to identify KLF2-regulated genes that may be essential for embryonic erythropoiesis and globin gene regulation in E9.5 erythroid precursor cells, using microarray assays. The LCM approach was used to isolate erythroid precursor cells from the embryonic yolk sac. The E9.5 developmental time point was selected because by E10.5 KLF2-/- cells appear dysmorphic and apoptotic, and this could confound the results. Also, at E10.5 some adult erythroid cells are present.
KLF2-regulated genes in E9.5 yolk sac erythroid cells
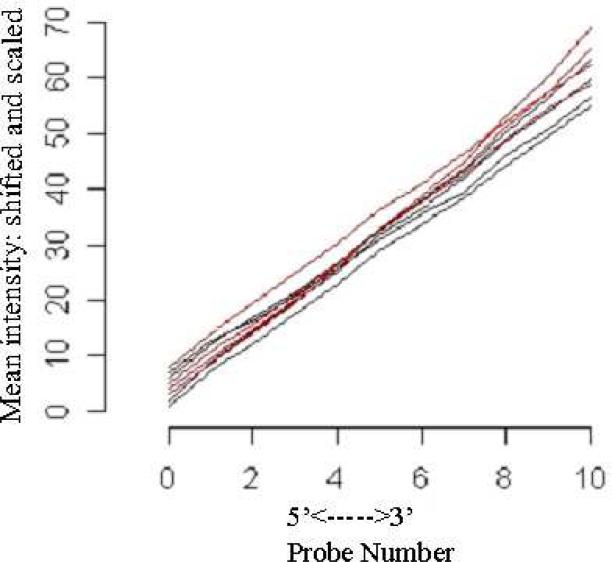
LCM was performed to collect erythroid cells from E9.5 KLF2-/- yolk sacs. E9.5 yolk sacs were chosen for the study to enrich for precursor cells in the primitive erythroid population. High quality RNA was isolated, as determined by the presence of strong rRNA peaks, and greater than 15% of the RNA was rRNA (Table 1). The RNA was linearly amplified, labeled, and hybridized to Affymetrix GeneChip® Mouse Genome 430A 2.0 arrays. For all hybridizations, greater than 40% of genes were present as determined by the Affymetrix detection call algorithm, confirming successful amplification and hybridization (Table 2). RNA digestion plots show that the KLF2-/- and previously performed WT E9.5 erythroid microarrays [22] have similar linear relationships, indicating equivalent RNA quality (Figure 1). Therefore, it is valid to compare the WT and KLF2-/- E9.5 erythroid microarrays.
Table 2.
Quality assessment parameters for KLF2–/– erythroid microarrays.
| Total RNA |
Labeled cRNA |
Microarray hybridizations |
||||
|---|---|---|---|---|---|---|
| Classification | Number of captured cells | % rRNA | cRNA bp median size | Average background | Scaling Factor (SF) | % genes present |
| 1-KLF2–/– | 4137 | 23.5 | 1000 | 60.2 | 0.58 | 53.0 |
| 2-KLF2–/– | 4200 | 29.0 | 1000 | 37.2 | 1.24 | 49.2 |
| 3-KLF2–/– | 4438 | 17.2 | ND | 54.5 | 0.84 | 40.2 |
| 4-KLF2–/– | 3625 | 15.9 | ND | 52.1 | 1.05 | 46.0 |
ND= not determined.
Figure 1.
Quality control assessment of wildtype and KLF2-/- E9.5 erythroid microarray assays. Shown is the RNA digestion plot for the 4 wildtype and 4 KLF2-/- Affymetrix 430A 2.0 Mouse Genome GeneChip hybridizations. The simpleaffy package in the Bioconductor R program (http://www.bioconductor.org) was used to assess RNA quality. The x-axis shows the 5’ to 3’ probe number for the position of the 22,690 probes. The y-axis indicates the mean intensity of the perfect-matched probes. The red lines are WT and black lines are the KLF2-/- samples.
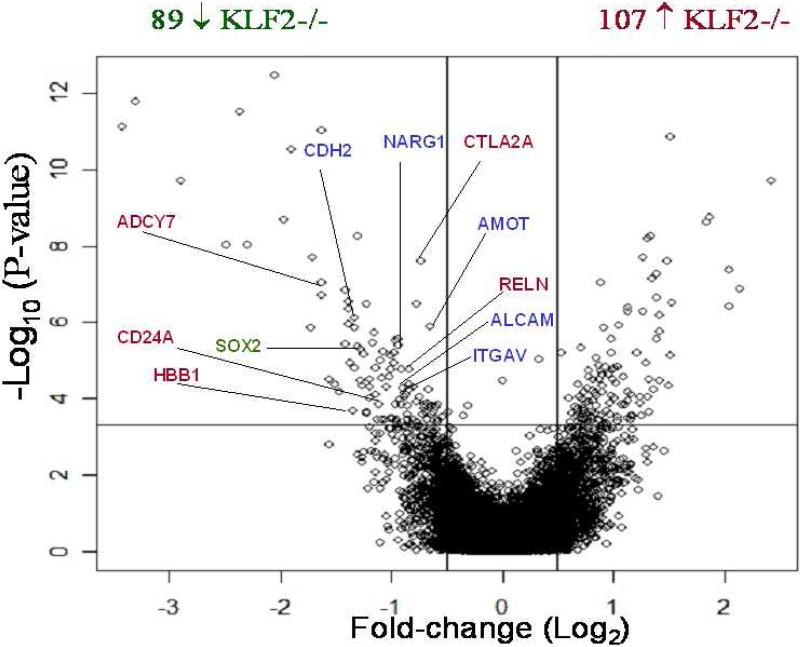
After applying the Robust multi-array (RMA) average algorithm to calculate probe set expression summaries, the multi-chip S-score statistical method was used to compare the wildtype and KLF2-/- data sets and p-values were obtained. These raw p-values were used to estimate the q-value, which is a quantifiable measure of the false discovery rate (FDR). Figure 2 is a volcano plot, which is a scatter-plot of the negative log10-transformed p-values from the S-score versus the log2-fold change between KLF2-/- and WT groups. The log2-fold change was estimated by first estimating the mean log2 expression in each group and then taking the difference. The q-value estimates were used to identify genes that are differentially expressed between the KLF2-/- and WT microarrays. At a q-value of 5%, 196 genes exhibited significant differences in expression. In the scatter plot, genes to the left of the leftmost vertical line, and above the horizontal line are expressed significantly lower in KLF2-/- compared to WT samples. The genes to the right of the rightmost vertical line and above the horizontal line are expressed significantly higher in KLF2-/- compared to WT samples. The genes labeled in red are known to be erythroid-specific genes. The genes labeled in blue are involved in cardiovascular development, perhaps reflecting a close relationship between KLF2 regulation in erythropoiesis and in vascular development. Eighty-nine unique genes are expressed lower in KLF2-/- than in wildtype cells, suggesting that they are directly or indirectly positively regulated by KLF2 (Table 3). One hundred-seven unique genes are expressed higher in KLF2-/- than in wildtype erythroid cells, and these genes are negatively regulated by KLF2 (Table 4). Most of the subsequent studies were focused on genes that are positively regulated by KLF2.
Figure 2.
Volcano plot showing differentially expressed genes between KLF2-/- and WT E9.5 erythroid cells using both biological and statistical dimensions. The x-axis is the fold change on a log scale. The y-axis represents the negative log10-transformed p-values of differences between the samples calculated using the S-score method. Above the horizontal line, there is a p-value <0.0004 and a q-value<0.05. Genes located between the two vertical lines had no change in expression. The genes in red are erythroid-enriched genes and those in blue are essential for cardiovascular development. The single gene in green (SOX2) is important for stem cell renewal.
Table 3.
Genes with decreased expression in KLF2–/– compared to WT erythroid cells.
| Symbol | Gene Description | S-score | Q-value | Classification | Probe ID |
|---|---|---|---|---|---|
| Prss11 | Protease, serine, 11 (Igf binding) | –7.27472 | 3.94E-09 | CS | 1416749_at |
| Plf2 | Proliferin 2 | –7.05638 | 1.29E-08 | CS | 1427760_s_at |
| Dtprp | Decidual/trophoblast prolactin-related protein | –6.97692 | 1.71E-08 | CS | 1448608_at |
| Csh1 | Chorionic somatomammotropin hormone 1 | –6.85038 | 3.34E-08 | CS | 1439002_s_at |
| Prlpe | Prolactin-like protein E | –6.35877 | 4.61E-07 | CS | 1449529_s_at |
| Fas | Fas (TNF receptor superfamily member 6) | –5.99074 | 3.95E-06 | CS/DEV/DIF/M | 1434279_at |
| Tra1 | Tumor rejection antigen gp96 | –5.83118 | 8.61E-06 | CS | 1438040_a_at |
| Eif2s3y | Eukaryotic translation initiation factor 2, subunit 3, structural gene Y-linked | –5.7371 | 1.21E-05 | OF | 1417210_at |
| Ddx3y | DEAD (Asp-Glu-Ala-Asp) box polypeptide 3, Y-linked | –5.74578 | 1.21E-05 | OF | 1426438_at |
| S100a6 | S100 calcium binding protein A6 (calcyclin) | –5.61071 | 2.36E-05 | CCR/DIF | 1421375_a_at |
| Ctla2a | Cytotoxic T lymphocyte-associated protein 2 alpha | – 5.57544 | 2.67E-05 | KES | 1448471_a_at |
| Cald1 | Caldesmon 1 | –5.34214 | 7.76E-05 | CS | 1424768_at |
| Gpr155 | G protein-coupled receptor 155 | –5.2657 | 0.000109 | CSR | 1452353_at |
| Prap1 | Proline-rich acidic protein 1 | –5.19941 | 0.000151 | UF | 1455996_x_at |
| Adcy7 | Adenylate cyclase 7 | –5.13724 | 0.000198 | KES/CS | 1456307_s_at |
| Csnk2a1 | Casein kinase II, alpha 1 polypeptide | –5.10952 | 0.000215 | CS | 1419038_a_at |
| Crabp2 | Cellular retinoic acid binding protein II | –5.0471 | 0.000268 | DEV | 1451191_at |
| Prlpa | Prolactin-like protein A | –4.94436 | 0.000413 | CS | 1448572_at |
| Cdh2 | Cadherin 2 | –4.87305 | 0.00058 | CA/DEV/DIF/M | 1418815_at |
| Amot | Angiomotin | –4.8393 | 0.000657 | DEV/DEV/M | 1454890_at |
| Ramp2 | Receptor (calcitonin) activity modifying protein 2 | –4.82988 | 0.000672 | CS | 1438403_s_at |
| Sgk | Serum/glucocorticoid regulated kinase | –4.76111 | 0.000874 | OF | 1416041_at |
| Wasl | Wiskott-Aldrich syndrome-like (human) | –4.68802 | 0.001206 | TF | 1452193_a_at |
| Adm | Adrenomedullin | –4.67536 | 0.001233 | CS/DEV | 1416077_at |
| Serpinb9e | Serine (or cysteine) proteinase inhibitor, clade B, member 9e | –4.63238 | 0.001491 | OF | 1418423_s_at |
| Tcfap2c | Transcription factor AP-2, gamma | –4.61909 | 0.001562 | TF | 1436392_s_at |
| Narg1 | NMDA receptor-regulated gene 1 | –4.60903 | 0.00161 | TF/.DEV | 1418024_at |
| Star | Steroidogenic acute regulatory protein | –4.59187 | 0.001719 | OF | 1418728_at |
| Sox2 | SRY-box containing gene 2 | –4.55385 | 0.001992 | TF/DEV | 1416967_at |
| Pycs | Pyrroline-5-carboxylate synthetase (glutamate gamma-semialdehyde synthetase) | –4.52626 | 0.002233 | OF | 1437325_x_at |
| Dnmt1 | DNA methyltransferase (cytosine-5) 1 | –4.51464 | 0.002284 | TF | 1435122_x_at |
| Hnrpu | Heterogeneous nuclear ribonucleoprotein U | –4.50275 | 0.002306 | OF | 1450849_at |
| Nap1l1 | Nucleosome assembly protein 1-like 1 | –4.47708 | 0.002551 | CM | 1420479_a_at |
| Ifi16 | Interferon, gamma-inducible protein 16 | –4.41792 | 0.003185 | TF/DEV/DIF | 1419603_at |
| Reln | Reelin | – 4.38168 | 0.003661 | KES/CS/DEV/CA | 1449465_at |
| 1600014E20Rik | RIKEN cDNA 1600014E20 gene | –4.37727 | 0.003685 | UF | 1460480_at |
| Syncrip | Synaptotagmin binding, cytoplasmic RNA interacting protein | –4.31957 | 0.00449 | RP | 1450743_s_at |
| Fabp7 | Fatty acid binding protein 7, brain | –4.31291 | 0.00457 | OF | 1450779_at |
| A130034M23Rik | RIKEN cDNA A130034M23 gene | –4.29795 | 0.004819 | UF | 1434513_at |
| Cnn3 | Calponin 3, acidic | –4.29331 | 0.004819 | OF | 1436759_x_at |
| Ddx6 | DEAD (Asp-Glu-Ala-Asp) box polypeptide 6 | –4.19517 | 0.006874 | OF | 1424598_at |
| Lgals3 | Lectin, galactose binding, soluble 3 | –4.17113 | 0.00741 | OF/DEV | 1426808_at |
| Slc2a2 | Solute carrier family 2 (facilitated glucose transporter), member 2 | –4.15784 | 0.007438 | OF | 1449067_at |
| Mt4 | Metallothionein 4 | –4.16199 | 0.007438 | OF | 1450645_at |
| 1600015I10Rik | RIKEN cDNA 1600015I10 gene | –4.14718 | 0.007713 | UF | 1449327_at |
| Fabp1 | Fatty acid binding protein 1, liver | –4.09344 | 0.009185 | OF | 1448764_a_at |
| A230046K03Rik | RIKEN cDNA A230046K03 gene | –4.07107 | 0.009924 | UF | 1439450_x_at |
| Nr2f1 | Nuclear receptor subfamily 2, group F, member 1 | –4.0561 | 0.010387 | TF/DEV | 1418157_at |
| Cd24a | CD24a antigen | – 4.04998 | 0.010471 | KES/DEV/CA/ CS/M | 1437502_x_at |
| Tpbpa | Trophoblast specific protein alpha | –4.05086 | 0.010471 | UF | 1438190_x_at |
| Alcam | Activated leukocyte cell adhesion molecule | –4.04102 | 0.010686 | CA/DIF/M | 1426301_at |
| Itgav | Integrin alpha V | –4.01708 | 0.011626 | CA/DEV/CS/DIF | 1452784_at |
| Gja1 | Gap junction membrane channel protein alpha 1 | –4.01239 | 0.011698 | CS/DEV/DIF | 1437992_x_at |
| Slc2a3 | Solute carrier family 2 (facilitated glucose transporter), member 3 | –3.95713 | 0.013993 | OF | 1455898_x_at |
| Lrrc58 | Leucine rich repeat containing 58 | –3.9114 | 0.016016 | UF | 1427131_s_at |
| Crabp1 | Cellular retinoic acid binding protein I | –3.90156 | 0.016289 | OF | 1448326_a_at |
| Fip1l1 | FIP1 like 1 (S. cerevisiae) | –3.88261 | 0.017241 | RP | 1428280_at |
| 2010109N14Rik | RIKEN cDNA 2010109N14 gene | –3.83141 | 0.020503 | UF | 1435524_at |
| Sdpr | Serum deprivation response | –3.8077 | 0.021223 | OF | 1416779_at |
| Hmgcs1 | 3-hydroxy-3-methylglutaryl-Coenzyme A synthase 1 | –3.80763 | 0.021223 | OF | 1433443_a_at |
| Hmgb1 | High mobility group box 1 | –3.80804 | 0.021223 | CM | 1448235_s_at |
| Snx6 | Sorting nexin 6 | –3.80385 | 0.021372 | CS/ CS | 1425148_a_at |
| Qk | Quaking | –3.77601 | 0.02309 | DEV/CS/RP/DIF | 1417073_a_at |
| Elavl4 | ELAV (embryonic lethal, abnormal vision, Drosophila)-like 4 (Hu antigen D) | –3.77538 | 0.02309 | RP | 1428741_at |
| Edg2 | Endothelial differentiation, lysophosphatidic acid G-protein-coupled receptor, 2 | –3.77616 | 0.02309 | CSR | 1448606_at |
| Acly | ATP citrate lyase | –3.75657 | 0.024661 | OF | 1439459_x_at |
| Rai14 | Retinoic acid induced 14 | –3.73454 | 0.026339 | TF | 1417400_at |
| Hes5 | Hairy and enhancer of split 5 (Drosophila) | –3.70697 | 0.02902 | TF/DEV/CA | 1456010_x_at |
| 5730427N09Rik | RIKEN cDNA 5730427N09 gene | –3.68658 | 0.030697 | UF | 1423132_a_at |
| Aass | Aminoadipate-semialdehyde synthase | –3.67887 | 0.030997 | OF | 1423523_at |
| Eif2s2 | Eukaryotic translation initiation factor 2, subunit 2 (beta) | –3.67696 | 0.030997 | OF | 1448819_at |
| Actb | Actin, beta, cytoplasmic | –3.65973 | 0.03255 | CS | 1419734_at |
| Sfrs6 | Splicing factor, arginine/serine-rich 6 | –3.65179 | 0.033196 | RP | 1416721_s_at |
| PDZ | PDZ and LIM domain 5 | –3.641 | 0.034046 | UF | 1450786_x_at |
| Eif3s8 | Eukaryotic translation initiation factor 3, subunit 8 | –3.63838 | 0.034086 | OF | 1415859_at |
| Sec61g | SEC61, gamma subunit | –3.63786 | 0.034086 | OF | 1423090_x_at |
| Elovl5 | ELOVL family member 5, elongation of long chain fatty acids (yeast) | –3.60471 | 0.037124 | OF | 1437211_x_at |
| Plac8 | Placenta-specific 8 | –3.60094 | 0.037276 | UF | 1451335_at |
| Sh3md2 | SH3 multiple domains 2 | –3.59081 | 0.038358 | CS | 1455149_at |
| Pten | Phosphatase and tensin homolog | –3.57595 | 0.039647 | CCR/DEV/CA | 1422553_at |
| Tfpi | Tissue factor pathway inhibitor | –3.57416 | 0.039663 | CS | 1451791_at |
| Cxcl7 | Chemokine (C-X-C motif) ligand 7 | –3.57233 | 0.039743 | CS | 1418480_at |
| Rgs2 | Regulator of G-protein signaling 2 | –3.56819 | 0.040003 | CS | 1419248_at |
| Ttc13 | Tetratricopeptide repeat domain 13 | –3.56212 | 0.040129 | UF | 1438631_x_at |
| Fabp4 | Fatty acid binding protein 4, adipocyte | –3.56462 | 0.040129 | TF | 1451263_a_at |
| Hbb-b1 | Hemoglobin, beta adult major chain | –3.53607 | 0.043467 | KES/DEV | 1417184_s_at |
| Snrpg | Small nuclear ribonucleoprotein polypeptide G | –3.53714 | 0.043467 | RP | 1448357_at |
| Tiparp | TCDD-inducible poly(ADP-ribose) polymerase | –3.52588 | 0.044545 | OF | 1452161_at |
| Xlr5c | X-linked lymphocyte-regulated 5C | –3.49823 | 0.048086 | UF | 1422933_at |
Classifications: OF = other factors; ES = known hematopoietic/erythroid specific; UF = unknown factors; CS = cell signaling; TF = transcription; CCF = cell cycle factors; CSF = cell surface receptors; CM = chromatin remodeling/assembly; RP = RNA processing; DIF = differentiation; DEV = development; M = migration.
Bold font indicates genes verified by qRT-PCR.
E10.5 KLF2-/- erythroid cells are abnormally shaped, suggesting that KLF2 regulates genes that are essential for erythroid cell morphology [20]. Circulating E9.5 EKLF-/- erythroid cells also have abnormal morphological characteristics [14]. EKLF-/- erythroid cells have reduced amounts of mRNA for major red cell membrane proteins and heme synthesis enzymes [11,12,34]. The erythroid membrane genes and heme synthesis genes that are controlled by EKLF do not appear to be regulated by KLF2. However, there are some similarities in the genes regulated by EKLF and by KLF2 in erythroid cells. For example, members of the solute carrier family 2, which are facilitated glucose transporters, are positively regulated by KLF2 (Slc2a2/GLUT2 and Slc2a3/GLUT3) and by EKLF (Slc2a1/GLUT1 and Slc2a4/GLUT4) [11,12]. Interestingly, GLUT1 has recently been shown to bind directly to dematin and adducin, and thus may play a role in attachment of the spectrin-actin junction to the lipid bilayer in erythrocyte membranes [35]. It is possible that dysregulated glucose transporter expression contributes to the abnormal shape of KLF1-/- and KLF2 -/- erythrocytes [36], but this has not yet been directly tested. In addition, the reelin and CD24a genes are positively regulated by KLF2 in primitive and by EKLF in definitive erythroid cells [11,12].
A reduction in Ey- and βh1-globin gene expression was detected by qRT-PCR in KLF2-/- compared to wild-type erythroid cells [20]. Unexpectedly, differential expression of these genes was not detected on the microarrays. This may be due to the fact that these genes are expressed in very high amounts, which may not be within the linear range of the microarray analyses. In support of this theory, the signals for Ey- and βh1-globin expression are higher than any other genes on the array, except for 2 of the ribosomal protein genes. The mouse adult β-globin gene, which is expressed at levels much lower than Ey- and βh1-globin at E9.5, was expressed in lower amounts in KLF2-/- than in normal erythroid cells in the microarray analyses. This was unexpected, because we had previously shown by qRT-PCR that KLF2 does not regulate the adult gene at a later time point, E12.5. This may indicate that KLF2 has a different role in adult β-globin gene expression at E9.5 than it has at E12.5.
Functional classification of E9.5 KLF2-regulated genes and identification of biological networks
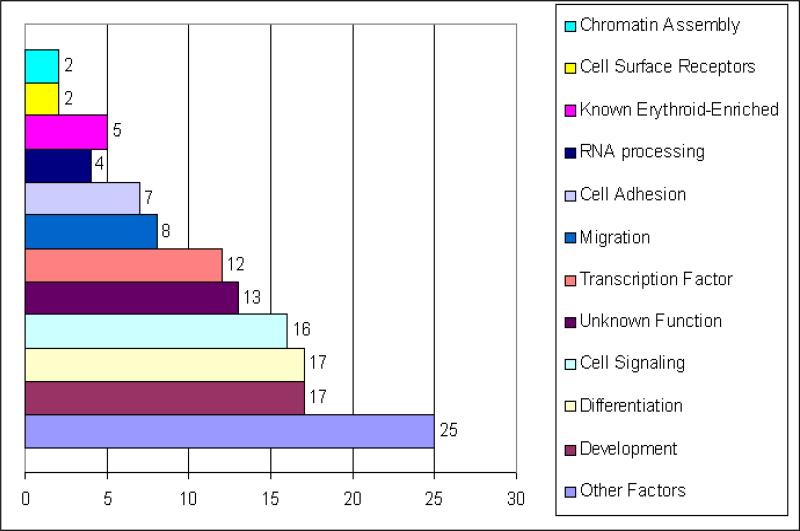
The list of genes positively regulated by KLF2 was arranged into functional categories using GO through DAVID and the available literature. The functional categories identified are in the bar graph in Figure 3. They include transcription factor, cell signaling, chromatin assembly, cell surface receptor, development, differentiation, erythroid-enriched, migration/motility, cell adhesion and RNA processing genes, genes of unknown function, and genes in other categories. The number of genes in specific GO categories in the differentially expressed gene list was compared to the number of genes in that category in the genome, to determine if certain gene categories are over-represented in the KLF2-regulated gene list. Interestingly, genes in 3 GO categories, cell migration, development and differentiation are over-represented in the KLF2-regulated gene list using a p-value <0.05. These over-represented categories indicate important biological themes in the gene list.
Figure 3.
Functional Classifications of genes exhibiting decreased expression in KLF2-/- compared to wildtype embryonic erythroid cells. Functional gene categories were determined with GO (Gene Ontology) using DAVID (Database Annotation Visualization and Integrated Discovery, http://david.abcc.ncifcrf.gov/). Other Factors are genes of known function that do not fit in the other categories. The bar chart represents the number of genes assigned per category. Some genes are in more than one category so the total is greater than 89.
Erythroid-enriched genes were used as a biological filter, and four of the genes positively regulated by KLF2 are erythroid-enriched in E9.5 yolk sac cells [22]. The genes which meet these conditions encode the cell signaling proteins reelin, adenylate cyclase 7, cytotoxic T lymphocyte-associated protein 2 α (Ctla2a), and CD24a antigen. The cellular roles of these proteins are discussed in more depth below. These genes are all potential direct targets of KLF2 (http://www.ncbi.nlm.nih.gov/). The reelin, Adcy7 and CD24a genes have at least one consensus binding site for KLF2 (CCACCC or CCGCCC, [37]) in their promoters, within 400 base pairs upstream of their transcription start sites. The Ctla2a gene has a CCACCC site in the first intron. Transcription factors such as SRY-box containing gene 2 (SOX2) and Hairy and Enhancer of Split 5 are also downregulated in KLF2-/- compared to wildtype erythroid cells, and these genes are essential for development/differentiation.
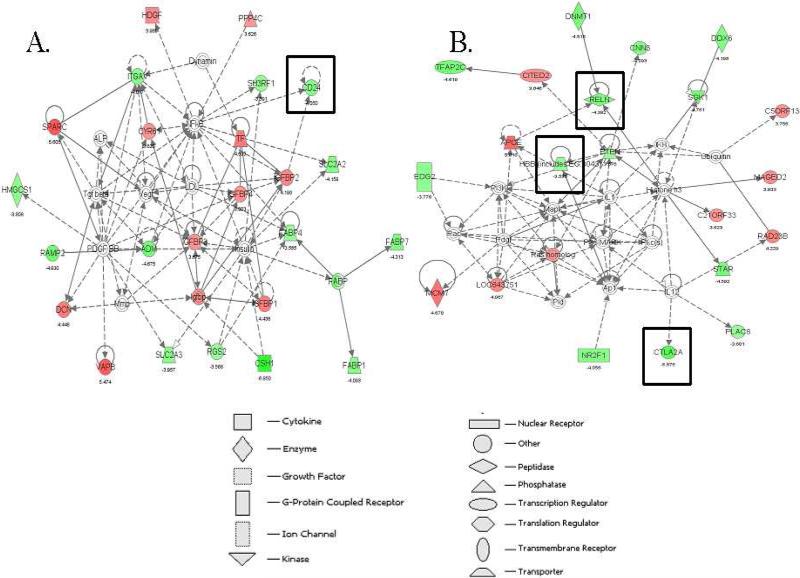
To further define how KLF2-regulated genes might interact in gene networks, an Ingenuity Pathway Analysis (IPA) was performed with the 196 genes that are either up- or down-regulated in KLF2-/- compared to WT erythroid cells. IPA uses a knowledge base program to generate relevant biological networks. These IPA networks visually describe direct or indirect functional relationships between genes, based on known interactions from the literature. One hundred and thirty-eight of the 196 differentially expressed genes were mapped to significant genetic networks using the IPA tool. Thirteen major networks were discovered in the gene list and nine of these networks have a p-value of 1.0E-21 or less, indicating that it is highly unlikely that they were detected by chance. The top ranked network had a score of 48 (p-value of 1.0E-48), and contains Cadherin 2 (Cdh2) and quaking (Qk), which are downregulated in KLF2-/- compared to wildtype erythroid cells (data not shown). These two KLF2-regulated genes are important for blood vessel development [38,39], and in so far as the two processes are related, may potentially be important for primitive erythropoiesis. The second network includes activated leukocyte cell adhesion molecule (ALCAM), lectin, galactoside-binding soluble 3, and S100 calcium binding protein A6 (S100A6), and is necessary for cardiovascular and nervous system development and function and cell-to-cell signaling (score of 43, data not shown). The third network (Figure 4A; score 43) is required for cellular proliferation, cancer and cell-to-cell signaling and includes cadherin 2 (CDH2), alpha integrin V (ITGAV), and CD24a antigen (CD24a), which the microarray assays indicate are downregulated in KLF2-/- compared to wildtype erythroid cells. Alpha integrin V is involved in blood vessel development [40]. The fourth network (Figure 4B; score 36) includes globin, reelin and cytotoxic T lymphocyte-associated protein 2 alpha (Ctla2a), and is important for cellular movement, molecular transport, and lipid metabolism. Interestingly, CD24a antigen, reelin and cytotoxic T lymphocyte-associated protein 2 alpha are erythroid-enriched [22] and are KLF2-regulated genes, and therefore are prime candidates for regulating primitive erythropoiesis. Overall, the data indicate that KLF2-regulated genes cluster in highly significant biological networks.
Figure 4.
Representative gene networks identified in KLF2-regulated gene list. Gene networks were explored using Ingenuity Pathway Analysis. Networks A and B are representative of the 11 significant networks identified. Network A scored 43 and B scored 36, indicating that their p-values are 1E-43 and 1E-36, respectively, and that they are highly significant. The genes shaded in red are upregulated and those shaded in green are downregulated in KLF2-/- compared to WT erythroid cells. Genes in white are not differentially expressed or not detected. Solid lines indicate direct relationships and dotted lines are indirect relationships. The genes surrounded by boxes are erythroid-enriched according to previous data.
Reelin is a large glycoprotein expressed in the Cajal-Retzius cells of the cerebral cortex. During development, reelin signaling is required for normal cortical neuronal migration (reviewed in [41]). Human reelin mutations can cause an autosomal recessive form of lissencephaly [42]. These clinical findings predict a role for reelin in cells other than neural cells, as the patients can have persistent lymphoedema neonatally. The role of reelin in adult neural cells is less clear, but it may play a role in synaptic transmission, and some evidence suggests that reelin could be involved in neurological conditions, such as epilepsy, autism and schizophrenia (reviewed in [43]). Reelin is frequently silenced in pancreatic cancers, and this correlates with increased cellular motility and invasiveness [44]. So, reelin is involved in the migration/mobility of both neural and pancreatic tumor cells. Interestingly, in erythroid cells lacking KLF2, there is reduced expression of the cell adhesion molecules N-cadherin (Cdh2) and integrin αV, suggesting that motility of the cells could be altered.
CD24a antigen is a cell surface glycoprotein expressed on immature cells of most major hematopoietic lineages [45]. Although CD24a null adult mice are viable, among other defects, hematopoiesis is compromised. Erythrocytes tend to aggregate, are more susceptible to hypotonic lysis, and have a shorter half-life in CD24a-/- than in WT mice [46]. The potential role of CD24a in regulating globin gene expression has not been explored. As a membrane protein, CD24a may play a role in establishing or maintaining normal erythroid morphology or cell-cell interactions.
Ctla2a is a cysteine proteinase inhibitor that was first identified in activated T lymphocytes and mast cells [47]. Although Ctla4 is well-characterized, the function of other Ctla family members, such as Ctla2a, remains unclear (reviewed in [48]). Two studies have reported that Ctla2a is expressed in both hematopoietic and germline stem cell niches [49,50]. This suggests that Ctla2a could be a regulator for stemness. In several recent studies, KLF2 and KLF4 have been implicated in the regulation of embryonic stem cell pluripotency and renewal [37,51]. It is possible that Ctla2a is required for hematopoietic stem cell renewal. Further work would need to be done to determine whether the Ctla2a gene is a direct KLF2 target and is essential for embryonic erythropoiesis.
Adenylate cyclase type VII (Adcy7) belongs to the family of enzymes that convert ATP to the intracellular second messenger cAMP. Forskolin-activation of the cAMP pathway increases expression of the fetally expressed β-like globin gene, γ-globin, in primary adult erythroblasts derived from BFU-E progenitors, but in contrast the cAMP pathway blocks -globin gene expression in the K562 erythroid cell line [52]. In human CD34+ cell cultures, cAMP production is required for full induction of fetal hemoglobin by hydroxyurea, sodium butyrate and 5-azacytidine [53]. It is possible that regulation of the Adcy7 gene by KLF2 may have a role in embryonic and fetal globin gene regulation.
Interestingly, our microarray assays indicate that Sox2, a transcription factor essential for self-renewal of undifferentiated embryonic stem cells, is regulated by KLF2. A cocktail of the Sox2, KLF2 and Oct3/4 genes has been used to generate induced pluripotent stem cells (iPS) from mouse fibroblasts [51]. KLF2 binds directly to the Sox2 promoter in ES cells, in chromatin immunoprecipitation and microarray assays (ChIP-on-chip) [37]. It is interesting that our microarray assays indicate that the Sox2 gene is also regulated by KLF2 in primitive erythroid cells. Because Sox2 null mice die early in embryogenesis [54], the role of SOX2 in erythropoiesis has not been fully explored. However, SOX2 may have a role in globin gene regulation. In fact, SOX6 is known to down-regulate Ey-globin expression during definitive erythropoiesis [55].
qRT-PCR validation of KLF2-regulated genes identified in microarray analyses
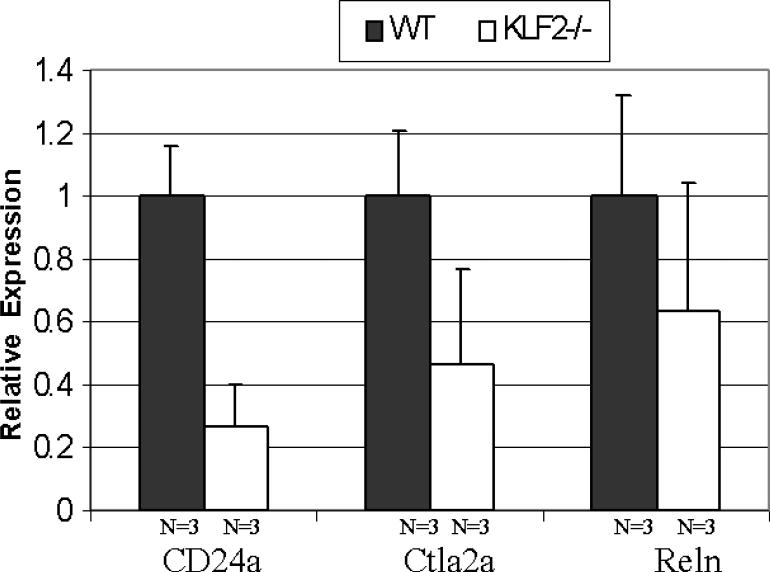
Three genes were of particular interest because they are KLF2-regulated, erythroid-enriched and found in high-scoring IPA networks. Therefore, the Reln, Ctla2a, and CD24a genes were selected for quantitative real-time PCR verification of differential expression. Independent LCM experiments were performed to collect additional samples for qRT-PCR, to avoid any possible bias in the microarray data. Between approximately 1,500 and 2,900 erythroid cells were collected from each of three individual E9.5 wildtype and from three KLF2-/- yolk sacs. qRT-PCR was performed using SYBR green chemistry on 3 samples of each genotype, to verify the microarray expression data. Cyclophilin A was used as an internal standard to normalize the expression data. As shown in Figure 5, CD24a mRNA is 3-fold, Ctla2a mRNA is 2-fold, and Reln mRNA is 1.5-fold lower in KLF2-/- than in WT E9.5 erythroid cells. The fold change for these three genes is higher in the qRT-PCR analysis than estimated from the microarray assays (at least 1.2-fold). Importantly, there is a positive association between the microarray and qRT-PCR data for all three genes. This confirms the accuracy of the microarray assays in identifying genes differentially expressed between the wildtype and KLF2-/- erythroid samples.
Figure 5.
qRT-PCR verification of KLF2-regulated genes identified by microarray analyses. Laser capture microdissection was performed to collect independent E9.5 wildtype and KLF2-/- erythroid cells. qRT-PCR was performed using between 70 and 150 erythroid cells per assay. Cyclophilin A mRNA was used to normalize the data. The normalized amount of mRNA in WT erythroid cells was set as 1. The relative fold expression in KLF2-/- cells is indicated on the y-axis. The error bars indicate the data mean ± standard error.
KLF2 upregulates reelin promoter activity
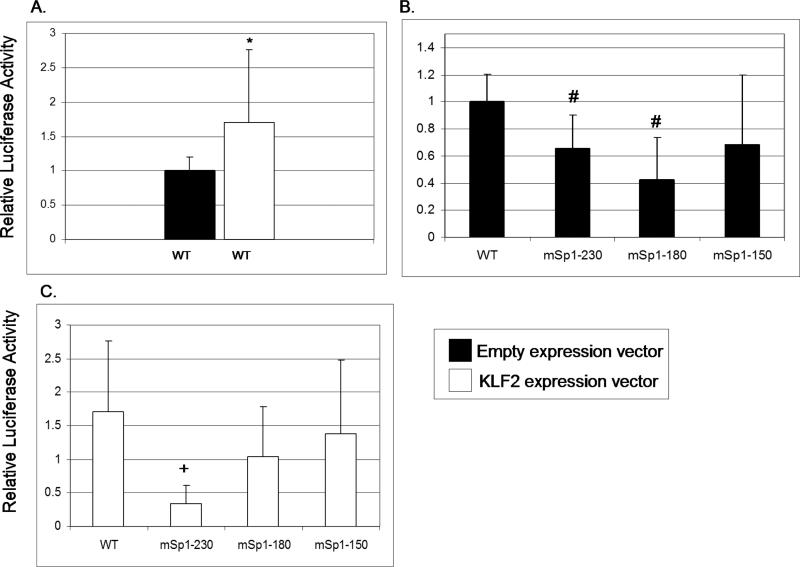
Microarray assays indicate that reelin is regulated by KLF2 and is erythroid-enriched in the E9.5 yolk sac [22], making it a strong candidate for regulating primitive erythropoiesis. In addition, reelin is involved in development and cellular migration, categories of genes that are over-represented in the KLF2-regulated gene list. To determine whether KLF2 has a direct effect on reelin promoter activity, transient co-transfection assays were performed in a human erythroleukemia cell line, K562. A reelin promoter-luciferase reporter construct was transfected into K562 cells in the presence or absence of a KLF2 cDNA expression construct. KLF2 activated the human reelin promoter-luciferase reporter by 1.7-fold, which was a statistically significant effect (Figure 6A). This modest effect may reflect the fact that some KLF2 is already present in K562 cells [15]. The -514 reelin promoter contains three GC rich (Sp1/KLF) binding sites located at -230, -180 and -150 within its enhancer region [31]. To determine whether any of these sites are KLF2 responsive elements, reelin promoter-luciferase constructs mutated at these three sites were transfected into K562 cells. There is a small but significant reduction (p<0.05) in luciferase activity for the K562 cells transfected with the constructs with the mutant -230 and -180 sites, compared to wildtype (Figure 6B). This data suggests that the -230 and -180 reelin promoter sites are necessary for positive regulation of the reelin gene in erythroid cells. Interestingly, in K562 cells co-transfected with the KLF2 expression construct, there is a significant 4-fold reduction in luciferase activity in cells with the -230 mutant site compared to the wildtype reelin promoter-luciferase construct (Figure 6C). This indicates that KLF2 regulates the reelin promoter via the -230 site (CCCCGCCC) in K562 cells. In NT2 neural progenitor cells, the -150 element (GCCCGCCC) is the main retinoic acid responsive site in the reelin promoter, mediating an Sp1-dependent response [32]. The -230 site is identical to the KLF2 consensus binding site [37]. It appears likely that KLF2 regulates the reelin gene through the -230 site in erythroid cells, whereas Sp1 regulates the gene through the -150 element in neural cells.
Figure 6.
KLF2 transactivates the reelin promoter in K562 cells. Data shown are the mean values from at least 4 independent transfection experiments. Error bars indicate standard deviation. (A) K562 cells were co-transfected with the wildtype reelin promoter-luciferase construct, and either an empty vector plasmid or the KLF2 expression vector. The “*” indicates significantly different luciferase activity at P<0.05 when cells with the WT luciferase construct are compared to WT plus KLF2 expression vector. (B) K562 cells were co-transfected with the wildtype reelin promoter-luciferase construct, or the mutant reelin promoter-luciferase constructs mSp1(-230)-Luc, mSp1(-180)-Luc or mSp1(-150)-Luc, plus the empty expression vector. The “#” indicates significantly different luciferase activity at P<0.05 for cells transfected with the mutant compared to WT reelin promoter-luciferase constructs. (C) The same luciferase constructs were transfected as in (B), but with the KLF2 expression construct. The “+” indicates significantly different luciferase activity in cells with mutant compared to WT luciferase constructs.
Future studies will be performed to determine the specific roles of CD24a, Ctla2a, Adcy7 and Sox2 in embryonic erythroid development. In addition, other biological filters will be used to identify candidate genes likely to have a role in primitive erythropoiesis. The EKLF and KLF2 genes interact to produce a more severe anemia phenotype in embryos, so microarray assays will be used to identify genes are regulated by both KLF2 and EKLF in E9.5 yolk sac erythroid cells. This approach should pinpoint key players in embryonic erythropoiesis.
Acknowledgments
We thank Dr. Jerry B Lingrel for his kind gifts of the KLF2 knockout mice and KLF2 cDNA expression construct, and Dr. David C. Williams Jr. for his critical evaluation of the mouse cytospin data. This work was supported by NIH U54 HL090516 to JAL and NIH F31 DK072548 to LCR. The GEO Series record for the microarray data in this article is GSE2760.
Footnotes
Publisher's Disclaimer: This is a PDF file of an unedited manuscript that has been accepted for publication. As a service to our customers we are providing this early version of the manuscript. The manuscript will undergo copyediting, typesetting, and review of the resulting proof before it is published in its final citable form. Please note that during the production process errors may be discovered which could affect the content, and all legal disclaimers that apply to the journal pertain.
References
- 1.Saleque S, Cameron S, Orkin SH. The zinc-finger proto-oncogene Gfi-1b is essential for development of the erythroid and megakaryocytic lineages. Genes Dev. 2002;16:301–306. doi: 10.1101/gad.959102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Fujiwara Y, Chang AN, Williams AM, Orkin SH. Functional overlap of GATA-1 and GATA-2 in primitive hematopoietic development. Blood. 2004;103:583–585. doi: 10.1182/blood-2003-08-2870. [DOI] [PubMed] [Google Scholar]
- 3.Warren AJ, Colledge WH, Carlton MB, Evans MJ, Smith AJ, Rabbitts TH. The oncogenic cysteine-rich LIM domain protein rbtn2 is essential for erythroid development. Cell. 1994;78:45–57. doi: 10.1016/0092-8674(94)90571-1. [DOI] [PubMed] [Google Scholar]
- 4.Winnier G, Blessing M, Labosky PA, Hogan BL. Bone morphogenetic protein-4 is required for mesoderm formation and patterning in the mouse. Genes Dev. 1995;9:2105–2116. doi: 10.1101/gad.9.17.2105. [DOI] [PubMed] [Google Scholar]
- 5.Martin R, Lahlil R, Damert A, Miquerol L, Nagy A, Keller G, Hoang T. SCL interacts with VEGF to suppress apoptosis at the onset of hematopoiesis. Development. 2004;131:693–702. doi: 10.1242/dev.00968. [DOI] [PubMed] [Google Scholar]
- 6.Baron MH. Embryonic origins of mammalian hematopoiesis. Exp. Hematol. 2003;31:1160–1169. doi: 10.1016/j.exphem.2003.08.019. [DOI] [PubMed] [Google Scholar]
- 7.Sanchez MJ, Bockamp EO, Miller J, Gambardella L, Green AR. Selective rescue of early haematopoietic progenitors in Scl(-/-) mice by expressing Scl under the control of a stem cell enhancer. Development. 2001;128:4815–4827. doi: 10.1242/dev.128.23.4815. [DOI] [PubMed] [Google Scholar]
- 8.Philipsen S, Suske G. A tale of three fingers: the family of mammalian Sp/XKLF transcription factors. Nucleic Acids Res. 1999;27:2991–3000. doi: 10.1093/nar/27.15.2991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Bieker JJ. Kruppel-like factors: three fingers in many pies. J Biol Chem. 2001;276:34355–34358. doi: 10.1074/jbc.R100043200. [DOI] [PubMed] [Google Scholar]
- 10.McConnell BB, Yang VW. Mammalian Kruppel-like factors in health and diseases. Physiol Rev. 2010;90:1337–1381. doi: 10.1152/physrev.00058.2009. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Drissen R, von LM, Kolbus A, Driegen S, Steinlein P, Beug H, Grosveld F, Philipsen S. The erythroid phenotype of EKLF-null mice: defects in hemoglobin metabolism and membrane stability. Mol. Cell Biol. 2005;25:5205–5214. doi: 10.1128/MCB.25.12.5205-5214.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Hodge D, Coghill E, Keys J, Maguire T, Hartmann B, McDowall A, Weiss M, Grimmond S, Perkins A. A global role for EKLF in definitive and primitive erythropoiesis. Blood. 2006;107:3359–3370. doi: 10.1182/blood-2005-07-2888. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Zhou D, Pawlik KM, Ren J, Sun CW, Townes TM. Differential binding of erythroid Kruppel-like factor to embryonic/fetal globin gene promoters during development. J. Biol. Chem. 2006;281:16052–16057. doi: 10.1074/jbc.M601182200. [DOI] [PubMed] [Google Scholar]
- 14.Basu P, Lung TK, Lemsaddek W, Sargent TG, Williams DC, Jr., Basu M, Redmond LC, Lingrel JB, Haar JL, Lloyd JA. EKLF and KLF2 have compensatory roles in embryonic β-globin gene expression and primitive erythropoiesis. Blood. 2007;110:3417–3425. doi: 10.1182/blood-2006-11-057307. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Zhang P, Basu P, Redmond LC, Morris PE, Rupon JW, Ginder GD, Lloyd JA. A functional screen for Krüppel-like factors that regulate the human γ-globin gene through the CACCC promoter element. Blood Cells, Molecules, and Diseases. 2005;35:227–235. doi: 10.1016/j.bcmd.2005.04.009. [DOI] [PubMed] [Google Scholar]
- 16.Kuo CT, Veselits ML, Barton KP, Lu MM, Clendenin C, Leiden JM. The LKLF transcription factor is required for normal tunica media formation and blood vessel stabilization during murine embryogenesis. Genes Dev. 1997;11:2996–3006. doi: 10.1101/gad.11.22.2996. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Wani MA, Means RTJ, Lingrel JB. Loss of LKLF function results in embryonic lethality in mice. Transgenic Res. 1998;7:229–238. doi: 10.1023/a:1008809809843. [DOI] [PubMed] [Google Scholar]
- 18.Wani MA, Wert SE, Lingrel JB. Lung Kruppel-like factor, a zinc finger transcription factor, is essential for normal lung development. J. Biol. Chem. 1999;274:21180–21185. doi: 10.1074/jbc.274.30.21180. [DOI] [PubMed] [Google Scholar]
- 19.Kuo CT, Veselits ML, Leiden JM. LKLF: A transcriptional regulator of single-positive T cell quiescence and survival. Science. 1997;277:1986–1990. doi: 10.1126/science.277.5334.1986. [DOI] [PubMed] [Google Scholar]
- 20.Basu P, Morris PE, Haar JL, Wani MA, Lingrel JB, Gaensler KML, Lloyd JA. KLF2 is essential for primitive erythropoiesis and regulates the human and murine embryonic beta-like globin genes. Blood. 2005;106:2566–2571. doi: 10.1182/blood-2005-02-0674. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Lee JS, Yu Q, Shin JT, Sebzda E, Bertozzi C, Chen M, Mericko P, Stadtfeld M, Zhou D, Cheng L, Graf T, MacRae CA, Lepore JJ, Lo CW, Kahn ML. Klf2 is an essential regulator of vascular hemodynamic forces in vivo. Dev. Cell. 2006;11:845–857. doi: 10.1016/j.devcel.2006.09.006. [DOI] [PubMed] [Google Scholar]
- 22.Redmond LC, Dumur CI, Archer KJ, Haar JL, Lloyd JA. Identification of erythroid-enriched gene expression in the mouse embryonic yolk sac using microdissected cells. Dev. Dyn. 2008;237:436–446. doi: 10.1002/dvdy.21426. [DOI] [PubMed] [Google Scholar]
- 23.Redmond LC, Haar JL, Giebel ML, Dumur CI, Basu P, Ware JL, Lloyd JA. Isolation of erythroid cells from the mouse embryonic yolk sac by laser capture microdissection and subsequent microarray hybridization. Blood Cells, Molecules, and Diseases. 2006;37:27–32. doi: 10.1016/j.bcmd.2006.02.006. [DOI] [PubMed] [Google Scholar]
- 24.Irizarry RA, Bolstad BM, Collin F, Cope LM, Hobbs B, Speed TP. Summaries of Affymetrix GeneChip probe level data. Nucleic Acids Res. 2003;31:e15. doi: 10.1093/nar/gng015. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Gentleman RC, Carey VJ, Bates DM, Bolstad B, Dettling M, Dudoit S, Ellis B, Gautier L, Ge Y, Gentry J, Hornik K, Hothorn T, Huber W, Iacus S, Irizarry R, Leisch F, Li C, Maechler M, Rossini AJ, Sawitzki G, Smith C, Smyth G, Tierney L, Yang JY, Zhang J. Bioconductor: open software development for computational biology and bioinformatics. Genome Biol. 2004;5:R80. doi: 10.1186/gb-2004-5-10-r80. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Gautier L, Cope L, Bolstad BM, Irizarry RA. affy--analysis of Affymetrix GeneChip data at the probe level. Bioinformatics. 2004;20:307–315. doi: 10.1093/bioinformatics/btg405. [DOI] [PubMed] [Google Scholar]
- 27.R Core Development Team . R: A Language and Environment for Statistical Computing. R Foundation for Statistical Computing; Vienna, Austria: 2007. [Google Scholar]
- 28.Zhang L, Wang L, Ravindranathan A, Miles MF. A new algorithm for analysis of oligonucleotide arrays: application to expression profiling in mouse brain regions. J. Mol. Biol. 2002;317:225–235. doi: 10.1006/jmbi.2001.5350. [DOI] [PubMed] [Google Scholar]
- 29.Kennedy RE. Probe level analysis of Affymetrix microarray data. Virginia Commonwealth University Thesis; 2008. [Google Scholar]
- 30.Kennedy RE, Kerns RT, Kong X, Archer KJ, Miles MF. SScore: an R package for detecting differential gene expression without gene expression summaries. Bioinformatics. 2006;22:1272–1274. doi: 10.1093/bioinformatics/btl108. [DOI] [PubMed] [Google Scholar]
- 31.Chen Y, Sharma RP, Costa RH, Costa E, Grayson DR. On the epigenetic regulation of the human reelin promoter. Nucleic Acids Res. 2002;30:2930–2939. doi: 10.1093/nar/gkf401. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Chen Y, Kundakovic M, gis-Balboa RC, Pinna G, Grayson DR. Induction of the reelin promoter by retinoic acid is mediated by Sp1. J. Neurochem. 2007;103:650–665. doi: 10.1111/j.1471-4159.2007.04797.x. [DOI] [PubMed] [Google Scholar]
- 33.Anderson KP, Kern CB, Crable SC, Lingrel JB. Isolation of a gene encoding a functional zinc finger protein homologous to erythroid Kruppel-like factor: identification of a new multigene family. Mol Cell Biol. 1995;15:5957–5965. doi: 10.1128/mcb.15.11.5957. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Nilson DG, Sabatino DE, Bodine DM, Gallagher PG. Major erythrocyte membrane protein genes in EKLF-deficient mice. Exp. Hematol. 2006;34:705–712. doi: 10.1016/j.exphem.2006.02.018. [DOI] [PubMed] [Google Scholar]
- 35.Khan AA, Hanada T, Mohseni M, Jeong JJ, Zeng L, Gaetani M, Li D, Reed BC, Speicher DW, Chishti AH. Dematin and adducin provide a novel link between the spectrin cytoskeleton and human erythrocyte membrane by directly interacting with glucose transporter-1. J. Biol. Chem. 2008;283:14600–14609. doi: 10.1074/jbc.M707818200. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Weber YG, Storch A, Wuttke TV, Brockmann K, Kempfle J, Maljevic S, Margari L, Kamm C, Schneider SA, Huber SM, Pekrun A, Roebling R, Seebohm G, Koka S, Lang C, Kraft E, Blazevic D, Salvo-Vargas A, Fauler M, Mottaghy FM, Munchau A, Edwards MJ, Presicci A, Margari F, Gasser T, Lang F, Bhatia KP, Lehmann-Horn F, Lerche H. GLUT1 mutations are a cause of paroxysmal exertion-induced dyskinesias and induce hemolytic anemia by a cation leak. J. Clin. Invest. 2008;118:2157–2168. doi: 10.1172/JCI34438. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Jiang J, Chan YS, Loh YH, Cai J, Tong GQ, Lim CA, Robson P, Zhong S, Ng HH. A core Klf circuitry regulates self-renewal of embryonic stem cells. Nat. Cell Biol. 2008;10:353–360. doi: 10.1038/ncb1698. [DOI] [PubMed] [Google Scholar]
- 38.Noveroske JK, Lai L, Gaussin V, Northrop JL, Nakamura H, Hirschi KK, Justice MJ. Quaking is essential for blood vessel development. Genesis. 2002;32:218–230. doi: 10.1002/gene.10060. [DOI] [PubMed] [Google Scholar]
- 39.Luo Y, Radice GL. N-cadherin acts upstream of VE-cadherin in controlling vascular morphogenesis. J. Cell Biol. 2005;169:29–34. doi: 10.1083/jcb.200411127. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.McCarty JH, Monahan-Earley RA, Brown LF, Keller M, Gerhardt H, Rubin K, Shani M, Dvorak HF, Wolburg H, Bader BL, Dvorak AM, Hynes RO. Defective associations between blood vessels and brain parenchyma lead to cerebral hemorrhage in mice lacking alphav integrins. Mol. Cell Biol. 2002;22:7667–7677. doi: 10.1128/MCB.22.21.7667-7677.2002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Kerjan G, Gleeson JG. A missed exit: Reelin sets in motion Dab1 polyubiquitination to put the break on neuronal migration. Genes Dev. 2007;21:2850–2854. doi: 10.1101/gad.1622907. [DOI] [PubMed] [Google Scholar]
- 42.Hong SE, Shugart YY, Huang DT, Shahwan SA, Grant PE, Hourihane JO, Martin ND, Walsh CA. Autosomal recessive lissencephaly with cerebellar hypoplasia is associated with human RELN mutations. Nat. Genet. 2000;26:93–96. doi: 10.1038/79246. [DOI] [PubMed] [Google Scholar]
- 43.Herz J, Chen Y. Reelin, lipoprotein receptors and synaptic plasticity. Nat. Rev. Neurosci. 2006;7:850–859. doi: 10.1038/nrn2009. [DOI] [PubMed] [Google Scholar]
- 44.Sato N, Fukushima N, Chang R, Matsubayashi H, Goggins M. Differential and epigenetic gene expression profiling identifies frequent disruption of the RELN pathway in pancreatic cancers. Gastroenterology. 2006;130:548–565. doi: 10.1053/j.gastro.2005.11.008. [DOI] [PubMed] [Google Scholar]
- 45.Pierres M, Naquet P, Barbet J, Marchetto S, Marics I, Devaux C, Barad M, Hyman R, Rougon G. Evidence that murine hematopoietic cell subset marker J11d is attached to a glycosyl-phosphatidylinositol membrane anchor. Eur. J. Immunol. 1987;17:1781–1785. doi: 10.1002/eji.1830171216. [DOI] [PubMed] [Google Scholar]
- 46.Nielsen PJ, Lorenz B, Muller AM, Wenger RH, Brombacher F, Simon M, von der WT, Langhorne WJ, Mossmann H, Kohler G. Altered erythrocytes and a leaky block in B-cell development in CD24/HSA-deficient mice. Blood. 1997;89:1058–1067. [PubMed] [Google Scholar]
- 47.Denizot F, Brunet JF, Roustan P, Harper K, Suzan M, Luciani MF, Mattei MG, Golstein P. Novel structures CTLA-2 alpha and CTLA-2 beta expressed in mouse activated T cells and mast cells and homologous to cysteine proteinase proregions. Eur. J. Immunol. 1989;19:631–635. doi: 10.1002/eji.1830190409. [DOI] [PubMed] [Google Scholar]
- 48.Teft WA, Kirchhof MG, Madrenas J. A molecular perspective of CTLA-4 function. Annu. Rev. Immunol. 2006;24:65–97. doi: 10.1146/annurev.immunol.24.021605.090535. [DOI] [PubMed] [Google Scholar]
- 49.Ivanova NB, Dimos JT, Schaniel C, Hackney JA, Moore KA, Lemischka IR. A stem cell molecular signature. Science. 2002;298:601–604. doi: 10.1126/science.1073823. [DOI] [PubMed] [Google Scholar]
- 50.Mizukami T, Kuramitsu M, Takizawa K, Momose H, Masumi A, Naito S, Iwama A, Ogawa T, Noce T, Hamaguchi I, Yamaguchi K. Identification of transcripts commonly expressed in both hematopoietic and germ-line stem cells. Stem Cells Dev. 2008;17:67–80. doi: 10.1089/scd.2007.0077. [DOI] [PubMed] [Google Scholar]
- 51.Nakagawa M, Koyanagi M, Tanabe K, Takahashi K, Ichisaka T, Aoi T, Okita K, Mochiduki Y, Takizawa N, Yamanaka S. Generation of induced pluripotent stem cells without Myc from mouse and human fibroblasts. Nat. Biotechnol. 2008;26:101–106. doi: 10.1038/nbt1374. [DOI] [PubMed] [Google Scholar]
- 52.Kuroyanagi Y, Kaneko Y, Muta K, Park BS, Moi P, Ausenda S, Cappellini MD, Ikuta T. cAMP differentially regulates gamma-globin gene expression in erythroleukemic cells and primary erythroblasts through c-Myb expression. Biochem. Biophys. Res. Commun. 2006;344:1038–1047. doi: 10.1016/j.bbrc.2006.03.203. [DOI] [PubMed] [Google Scholar]
- 53.Keefer JR, Schneidereith TA, Mays A, Purvis SH, Dover GJ, Smith KD. Role of cyclic nucleotides in fetal hemoglobin induction in cultured CD34+ cells. Exp. Hematol. 2006;34:1151–1161. doi: 10.1016/j.exphem.2006.03.018. [DOI] [PubMed] [Google Scholar]
- 54.Avilion AA, Nicolis SK, Pevny LH, Perez L, Vivian N, Lovell-Badge R. Multipotent cell lineages in early mouse development depend on SOX2 function. Genes Dev. 2003;17:126–140. doi: 10.1101/gad.224503. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Yi Z, Cohen-Barak O, Hagiwara N, Kingsley PD, Fuchs DA, Erickson DT, Epner EM, Palis J, Brilliant MH. Sox6 directly silences epsilon globin expression in definitive erythropoiesis. PLoS. Genet. 2006;2:e14. doi: 10.1371/journal.pgen.0020014. [DOI] [PMC free article] [PubMed] [Google Scholar]