Abstract
Regulation of histone methylation levels has long been implicated in multiple cellular processes, many of which involve transcription. Here, however, we report a unique role for the Caenorhabditis elegans histone demethylase SPR-5 in meiotic DNA double-strand break repair (DSBR). SPR-5 shows enzymatic activity toward H3K4me2 both in vitro and in the nematode germline, and spr-5 mutants show several phenotypes indicating a perturbation of DSBR, including increased p53-dependent germ cell apoptosis, increased levels of the DSBR marker RAD-51, and sensitivity toward DSB-inducing treatments. spr-5 mutants show no transcriptional misregulation of known DSBR involved genes. Instead, SPR-5 shows a rapid subcellular relocalization upon DSB-inducing treatment, which suggests that SPR-5 may function directly in DSBR.
Keywords: lysine specific histone demethylase, meiosis
Dynamic regulation of histone methylation status has been linked to multiple biological processes, including transcriptional activation and repression, sex chromosome silencing, and DNA damage response (reviewed in refs. 1 and 2). The recent discovery of histone demethylase enzymes has offered opportunities to further understand the function of these modifications. Histone H3 methylation occurs at multiple lysine (K) residues (H3K4, K9, K27, K36, and K79), and each of these lysine moieties can be methylated to mono-, di-, and trimethylated states. Both the location and the degree of methylation are believed to play a role in the determination of the distinct function a specific methylation event plays in the regulation of chromatin structure and function. Histone H3 methylation at lysine 4 (H3K4me) is a particularly intriguing modification: it has not only been strongly linked to transcriptional activation and enhancer function but more recently has been shown to be enriched at meiotic recombination hotspots (3–7), indicating a potential role for epigenetic control of hotspot localization in regulating meiotic double-strand break (DSB) initiation.
The mammalian histone demethylase LSD1 was originally identified as an H3K4me2/1-specific demethylase (8), and studies of LSD1 in various species have presented several common phenotypes suggestive of a role for LSD1 in meiosis. For example, fission yeast lacking LSD1 have a sporulation defect, and flies show a female sterility phenotype (9–11). More recently, a mutation in spr-5, a Caenorhabditis elegans ortholog of LSD1, was found to show a phenotype of progressive sterility (12). The authors used whole-animal samples to identify spr-5 target genes and showed a progressive increase of H3K4me2 for a subset of genes, including spermatogenesis genes (12). The sterility phenotypes detected in several organisms are suggestive of a role for LSD1 orthologs in meiosis; however, LSD1 function in the germline remains largely unknown.
Here we examined the germline function of spr-5, using tissue-specific techniques in C. elegans. These experiments uncovered an unexpected set of phenotypes indicating perturbed DSB repair (DSBR), linking H3K4me2 modulation via SPR-5 to efficient repair of meiotic DSBs. Specifically, SPR-5 acts in vitro to demethylate H3K4me2, and spr-5 mutants show increased H3K4me2 levels in the germline, indicating that spr-5 is responsible for germline H3K4me2 modulation. In addition, spr-5 mutants show an increase in p53-dependent germ cell death, along with increased meiotic RAD-51 foci, a DSBR marker, suggesting a disruption of meiotic DSBR progression. These findings are further supported by the specific sensitivity of spr-5 mutants to DSB-inducing γ-irradiation and the observation of a rapid relocalization of germline SPR-5 in response to irradiation. Importantly, these phenotypes are immediately apparent in early generation (F2–F4) spr-5 animals, and they become progressively more severe over multiple generations, suggesting an epigenetic nature for this regulation. However, this regulation may not be transcriptional in nature: germline-specific microarray experiments were carried out to assess whether spr-5 target genes may be involved in these DSBR phenotypes, and no known DSBR genes were identified. Instead, spr-5 may be acting in parallel to repress various classes of somatic genes within the germline. These findings therefore represent evidence for dual roles for the LSD1 family of demethylases in both transcriptional control and meiotic DSBR progression, further linking epigenetic regulation to these biological processes.
Results and Discussion
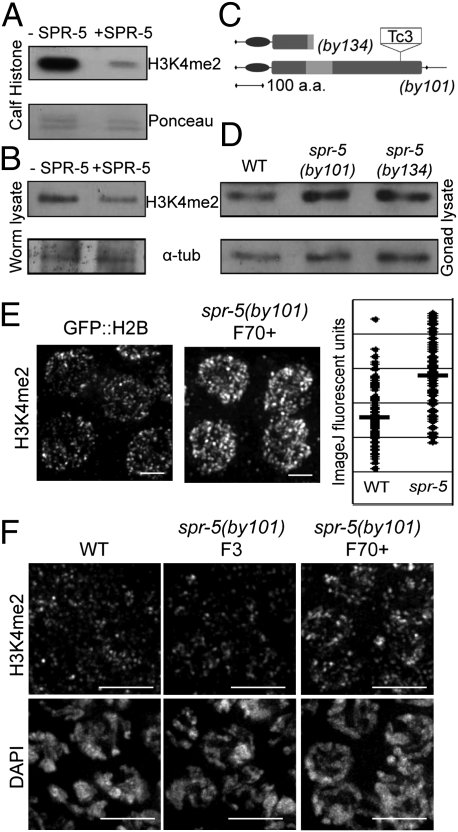
Orthologs of LSD1 in various other species are confirmed histone demethylases both in vitro and in vivo (reviewed in ref. 13). SPR-5, however, has not had its demethylase activity confirmed in vitro, although in vivo studies support its activity as an H3K4me2 demethylase in C. elegans (12). To address this issue, we expressed and purified full-length FLAG-tagged SPR-5 from Sf9 insect cells. Purified SPR-5 was then tested in an in vitro demethylase assay against histones extracted from calf thymus containing various post-translational modifications. Histones incubated with and without SPR-5 were separated by Western blot and probed using antibodies against H3K4me2 (Fig. 1A). Histones incubated with SPR-5 showed a noticeable decrease in H3K4me2 level compared with a Ponceau dye loading control (only 29% of control level was observed), indicating in vitro demethylase activity for this protein. To confirm activity on a C. elegans substrate, we incubated full-length SPR-5 with whole-worm protein lysate and observed a 50% reduction in H3K4me2 levels by Western blot as well (Fig. 1B). Taken together, these results indicate that SPR-5 is an H3K4me2 demethylase.
Fig. 1.
SPR-5 is an H3K4me2 demethylase that functions in the C. elegans germline. (A and B) SPR-5 demethylates H3K4me2. Full-length FLAG-SPR-5 was incubated with purified calf thymus histones (A) or whole-worm protein lysate (B). Western blots were used to separate samples, then probed using anti-H3K4me2 antibodies and Ponceau dye (Upper) or anti-tubulin antibody (Lower) for loading controls. (C) Schematic of spr-5 mutant alleles. spr-5(by134) contains an early nonsense mutation within the TOWER domain (light gray). spr-5(by101) contains a transposon insertion (Tc3) adjacent to predicted catalytic residues in the amine oxidase domain (medium gray). The SWIRM domain (dark gray), predicted for protein–protein interactions, is unaffected. (D) spr-5 mutants show increased germline H3K4me2. Gonads were dissected from 50 animals per genotype. After separation by Western blot, samples were probed with anti-H3K4me2 and anti-tubulin (loading control) antibodies. (E) Immunostaining reveals that spr-5 mutant germ cells show increased H3K4me2. Gonads from either GFP::H2B-expressing animals (control) or spr-5 (F70+) worms were immunostained for H3K4me2 (Left and Center). (Scale bars, 2 μm.) Individual nuclei were sampled for fluorescent intensity using ImageJ (Right). The bars represent the mean fluorescent intensity. (F) Germ cell H3K4me2 is progressively misregulated in spr-5 mutants. Germline H3K4me2 immunostaining was compared between wild-type and both late-generation (F70+) and early-generation (F3, created from a single outcross of the F70+ strain) spr-5 mutants. (Scale bars, 5 μm.)
Given that SPR-5 acts in vitro to demethylate H3K4me2 and that previous studies have shown by chromatin immunoprecipitation on whole-worm extracts that H3K4me2 levels are increased at target gene promoters in spr-5 mutant animals (12), we next examined whether SPR-5 acts in the germline to modulate H3K4me2. We first confirmed that SPR-5 is expressed in the germline using a C-terminal specific antibody, which detected SPR-5 expression in the soma as well (Fig. S1). This finding confirms the importance of using tissue-specific methods to assess SPR-5 germline function (see below).
We focused on addressing the importance of SPR-5 in germline histone methylation control by comparing germline-specific levels of H3K4me2 in wild-type worms with those in two spr-5 mutants: spr-5(by101) and spr-5(by134) (Fig. 1 C and D). A transposon insertion is adjacent to a predicted catalytic residue in the by101 allele, and a nonsense mutation eliminates most of the enzymatic domain in the by134 allele, suggesting that both may be functionally null alleles. Consistent with this, both spr-5 mutants showed equivalent phenotypes in all further experiments. We assessed germline H3K4me2 levels both by Western blots and immunostaining of whole mounted gonads. Specifically, dissected gonads from age-matched adult hermaphrodites were probed for H3K4me2 levels by Western blot (Fig. 1D). Both spr-5(by101) and spr-5(by134) mutants have higher levels of H3K4me2 in their germline tissue compared with wild-type (23% and 37%, respectively) as measured by Western blot intensity, indicating a role for spr-5 in down-regulating this modification. To obtain comparable immunofluorescent images, we used a transgenic GFP::H2B, and otherwise wild-type, strain as our control strain. These animals were dissected on the same slide as spr-5(by101) homozygous animals, and therefore both control and mutant germlines were processed and imaged under identical conditions. The control gonads were identified by the GFP-positive signal on chromosomes, and the H3K4me2 was imaged using a Cy3 secondary antibody, allowing for comparison of H3K4me2 intensity between genotypes (Fig. 1E, Left). Fluorescent intensity was sampled from multiple nuclei from different gonads and quantitated using ImageJ software (Fig. 1E, Right). Although there is overlap between the H3K4me2 intensities in control and spr-5 mutant gonads, there is a statistically significant increase in H3K4me2 in spr-5 mutant germlines (P < 0.0001, two-tailed Mann-Whitney test, 95% confidence interval), further supporting a role for spr-5 in germline H3K4me2 regulation.
On the basis of previous reports of a progressive sterility phenotype and a progressive increase of H3K4me2 on target gene promoters from whole-worm samples (12), we set out to examine whether the H3K4me2 misregulation in the germline was progressive as well. Immunostaining of wild-type and spr-5 mutant germlines indicates that early-generation animals do not show a detectable change in H3K4 dimethylation, whereas late-generation animals (animals maintained as homozygous mutants for more than 70 generations) show an increase in H3K4me2 (Fig. 1F). These results suggest that the misregulation in germline H3K4me2 is progressive in spr-5 mutants.
Misregulation of various methylation modifications has been shown to affect transcription, but several recent studies have implicated histone methylation state in DNA damage repair as well (reviewed in ref. 14). We therefore examined spr-5 mutant germlines for defects in DNA damage repair. Because germ cell apoptosis can be triggered by unresolved DNA damage (15), spr-5 mutants were analyzed for possible increase in germline apoptosis using acridine orange staining. Both spr-5 mutants show a two- to threefold increase in apoptosis across multiple experiments (Fig. 2 A and B and Fig. S2). This two- to threefold increase is apparent in second-generation (F2) spr-5 mutant animals but becomes progressively more severe in subsequent generations before leveling off by generation F30 (Fig. 2 A and B).
Fig. 2.
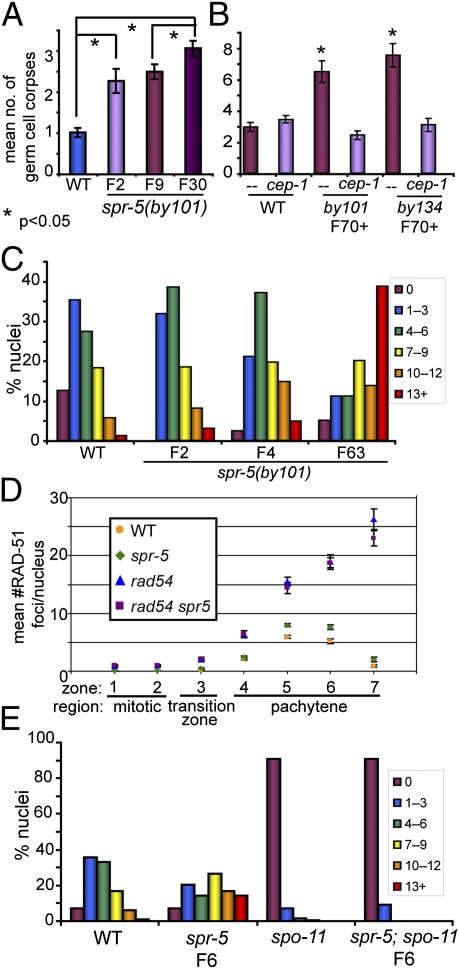
spr-5 mutants show a progressive defect in DNA double-strand break repair. (A) spr-5 mutants show a progressive increase in germ cell apoptosis. Error bars represent the SEM. P values were determined using the two-tailed Mann-Whitney test, 95% confidence interval. (B) Increased apoptosis in late generation spr-5 mutants is cep-1-dependent. Wild-type as well as late-generation spr-5(by101) and spr-5(by134) animals were examined after growth in either control(RNAi) or cep-1(RNAi) plates. As reported previously, RNAi treatment (both control and target gene-specific) results in higher levels of germ cell apoptosis (approximately three apoptotic germ cells in the control) (31), but the relative increase in spr-5 mutants remains. Error bars represent the SEM; P values were determined by the two-tailed Mann-Whitney test, 95% confidence interval. (C) spr-5 mutants show a progressive increase in mid-pachytene RAD-51 foci. The number of RAD-51 foci per nucleus was quantified in the mid-pachytene zone (zone 5 as in ref. 18 and 20) for at least three gonads per strain. Ranges (number of foci detected per nucleus) and color codes are indicated in the key. The rightward shift of the histograms indicates a higher percentage of nuclei with more RAD-51 foci. (D) Analysis of repair-defective mutants of the indicated genotypes reveals no difference in the number of DSBs formed. The total number of RAD-51 foci in each nucleus was quantified as in C and Fig. S3. However, here the data are displayed as the mean number of RAD-51 foci per nucleus (y axis) in each zone or region of the germline (x axis) for at least three gonads per genotype. There is no statistically significant difference between rad-54 and rad-54 spr-5 mutants at any stage. Experiments were done on generation F3–F5 mutants. (E) The increase in RAD-51 foci is dependent on the meiotic endonuclease spo-11. Histograms depict the quantitation of RAD-51 foci in midpachytene (zone 5) nuclei for at least three germlines for each indicated genotype.
Increased germ cell apoptosis can be due to triggering of a DNA damage checkpoint mediated by the p53 homolog CEP-1 (16). For both spr-5(by101) and spr-5(by134) mutants (≥F70), RNAi depletion of cep-1 reduces apoptosis to wild-type levels (Fig. 2B), indicating that spr-5 mutants are experiencing DNA damage-induced apoptosis. At generation F2–F3, cep-1(lg12501) spr-5(by101) double mutants show wild-type levels of apoptosis compared with the twofold increase seen in an spr-5(by101) single mutant (Fig. S2), further confirming that the increased apoptosis is dependent on CEP-1 and therefore DNA damage-induced, at both early and late generations. Supporting this connection between meiotic DNA damage and spr-5 function, the increase in germ cell apoptosis observed in spr-5 mutants is also dependent on spo-11, the endonuclease responsible for meiotic DSB formation (17) (Fig. S2).
Cytological examination of the nuclei within the germline can provide evidence of perturbations to the progression of DSBR through quantitation of DNA damage repair markers, such as RAD-51. In wild-type germlines, few RAD-51 foci are observed in the premeiotic region. Levels then begin to increase in late transition zone (leptotene/zygotene stages), peak in early- to mid-pachytene, and are resolved by the end of pachytene (18). spr-5 mutants show an increase in the number of RAD-51 foci per nucleus in early- to mid-pachytene (Fig. 2 C–E and Fig. S3). The persistence of high levels of RAD-51 foci throughout the late pachytene region suggests that either more DSBs are being formed or that there is a delay in DSBR. To distinguish between these possibilities, we examined the levels of RAD-51 foci in a rad-54 mutant background, where DSBs and RAD-51 foci remain colocalized, and where DSBR is blocked and RAD-51 foci are not removed, allowing for quantitation of the total number of breaks formed (19). This analysis revealed no difference between rad-54 single and rad-54 spr-5 double mutant animals, supporting a primary role for spr-5 in DSBR, rather than in DSB formation (Fig. 2D). In addition, the lack of RAD-51 foci later at diplotene, accompanied by the lack of chromosome fragments at diakinesis (Fig. S4), suggests that DSBR is eventually successfully completed in spr-5 mutants. However, in the case of a spr-5;syp-1 double mutant, in which homologous chromosomes are unsynapsed and therefore lack the ability to repair damage off the homologous template (20), altered chromosome morphogenesis indicative of unresolved DNA damage is observed at diakinesis. This further supports a role for spr-5 in promoting both interhomolog and intersister recombination (Fig. S4).
As with germ cell apoptosis, the increase in RAD-51 foci is apparent in early-generation spr-5 mutants but becomes more severe over multiple generations (Fig. 2C), indicating a progressive defect. Importantly, the increase in RAD-51 foci is dependent on the meiosis-specific endonuclease SPO-11 (Fig. 2E), emphasizing the meiotic-specific nature of this DSBR phenotype.
SPR-5 may function to actively promote repair after DNA damage, therefore explaining the DSBR defect present in early-generation spr-5 mutants. H3K4 methylation has been reported at meiotic recombination hotspots in budding yeast and in mouse (4, 7). SPR-5 may act to remove the H3K4me2 at the sites of DSBs in a manner that promotes repair. If H3K4me2 removal by SPR-5 is important for efficient progression of DSBR, we might expect some observable effect on SPR-5 activity or localization upon induction of DSBs.
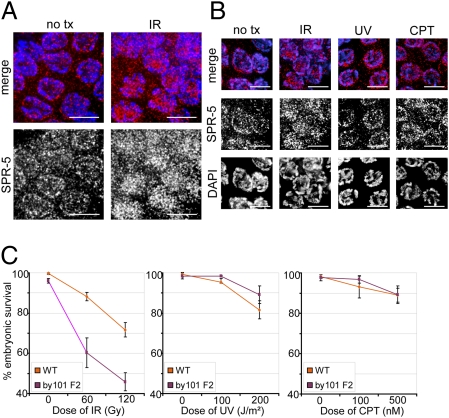
Exposure of adult hermaphrodite animals to γ-irradiation (IR) introduces DSBs in the germline, providing a model to study SPR-5 activity in the wild-type gonad and determine whether SPR-5 levels or subcellular distributions change upon DSB induction. In the absence of IR, SPR-5 shows a nuclear-associated pattern (Fig. 3 A and B). However, upon IR, SPR-5 shows a rapid (within 15 min) relocalization resulting in an increase in perichromosomal intensity (Fig. 3 A and B). Two additional types of DNA-damaging agents were tested: UV-C, which largely leads to pyrimidine dimers, and camptothecin (CPT), which leads to single-strand breaks. Importantly, SPR-5 does not seem to relocate after exposure to UV or CPT, only to DSB-inducing IR (Fig. 3B and Fig. S5). SPR-5 is therefore engaging in a rapid and dynamic relocalization specifically after the induction of DSBs in the germline. In a spo-11 mutant background where DSBs are not being formed, SPR-5 displays a localization pattern that is indistinguishable from that observed in non-irradiated germlines, supporting a function for SPR-5 downstream of break formation (Fig. S6). However, given that an average of only 2.1 DSBs are made per meiotic chromosome pair in C. elegans (19), we cannot exclude the possibility that it may be too difficult to detect a SPO-11–dependent change of SPR-5 localization by immunofluorescence.
Fig. 3.
SPR-5 relocalizes in response to DNA double-strand breaks and is required for IR resistance. (A) SPR-5 relocalizes rapidly upon irradiation. Gonads were dissected from 24 h post-L4 wild-type worms 15–30 min after exposure to 60 Gy of IR and compared with untreated controls (no tx). Images are from pachytene nuclei. (Scale bars, 5 um.) (B) SPR-5 relocalizes upon DSB-inducing IR but not in response to other DNA damage-inducing reagents. Animals were treated as in A with 60 Gy for the IR, with 200 J/m2 for the UV and 500 nM for the CPT treatments. (C) Early-generation (F2) spr-5(by101) mutants show decreased embryonic viability compared with wild-type after treatment with the indicated doses of IR, but not UV-C or CPT. At least 30 animals were assayed per genotype and condition. Points represent the mean embryonic survival, error bars represent the SEM.
On the basis of the relocalization of SPR-5 upon exogenous DSB formation, we next examined spr-5 mutants for sensitivity to IR as well as other sources of DNA damage. Adult hermaphrodites were exposed to different types of DNA damage, and the embryonic survival of their offspring, judged by egg-hatching frequencies, was quantified over the next 24 h, providing a read-out of germ cell fate after damage. spr-5 mutants show IR sensitivity compared with wild-type animals but no statistically significant difference when treated with UV-C or CPT (Fig. 3C). These data, along with the relocalization results, indicate a specific sensitivity to DSBs, the predominant type of damage occurring during meiosis.
The rapid relocalization of SPR-5 upon DSB-inducing IR treatment suggests that SPR-5 may have an important role in DSB response and repair. The in vitro activity of SPR-5 and in vivo regulation of H3K4me2 observed in the analysis of spr-5 mutants indicates that H3K4me2 is the major substrate for SPR-5. Perhaps this enzymatic activity is important for SPR-5 activity in DSBR as well. However, one alternative explanation for the spr-5 mutant phenotypes of perturbed DSBR and increased sensitivity to IR could be a misregulation of genes involved in DSBR. To address this issue, as well as to identify other spr-5 transcriptional targets, we carried out a germline-specific microarray comparing wild-type with spr-5 mutant samples. This microarray was done on RNA isolated from dissected gonads from age-matched young adults, compared with a previous microarray done on mixed-stage whole animals (12). Our microarray experiment identified 126 genes consistently up-regulated in late-generation (F70+) spr-5(by101) mutants and 4 genes down-regulated in quadruplicate samples, suggesting that SPR-5 functions predominantly as a repressor, not an activator of transcription, in C. elegans germ cells (Table S1).
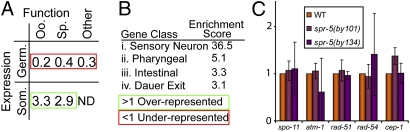
In contrast to results reported in ref. 12, this gene list does not show an enrichment for germline-expressed spermatogenesis genes, and in fact genes up-regulated in spr-5 mutant germlines are statistically significantly under-enriched for germline-expressed genes (Fig. 4A). However, spr-5 target genes are over-enriched in the somatically expressed sets of genes shown to be functionally important for spermatogenesis or oogenesis (Fig. 4A). These data suggest that spr-5 represses somatically expressed genes important for germline function, so that in its absence they are inappropriately up-regulated in the germline. The previous experiments were done using mixed-stage intact animals and identified a cohort of coregulated genes, including germline-expressed spermatogenesis genes (12). Taken together, these data sets suggest two potential models. One is that loss of spr-5 transcriptionally derepresses germline genes at early to middle generations but not at later generations, as supported by the lack of derepression seen at generation F28 by Katz et al. (12) and at much later generations in our work. A second potential model is that spr-5 may repress somatic genes in the germline and repress germline genes in the soma, leading to the observable deregulation of germline genes when whole animals are used for isolating RNA. Comparison of our spr-5 gene target list with other published microarray datasets shows that spr-5 targets are over-enriched for genes identified as specifically enriched in several classes of somatic tissues or during specific developmental processes, including sensory neurons, pharyngeal, intestinal, and dauer (Fig. 4B), supporting a role for spr-5 in repressing somatic genes in the germline. Given the different generations and tissue sources used for RNA isolation and our observation of spr-5 expression in the soma, both models are formally possible. Future tissue-specific work is therefore needed to confirm whether spr-5 mutants show a deregulation of germline genes in the soma, which will have implications for our understanding of the mechanism behind the progressive sterility phenotype.
Fig. 4.
spr-5 germline gene targets are under-enriched for germline-expressed genes and do not include known DSBR genes. (A) Comparison of genes up-regulated in spr-5 mutants with published germline function and expression data (32). Germline expression refers to genes enriched in wild-type hermaphrodites compared with germline-deficient glp-4 mutants. Somatic expression refers to genes of each functional class that are not germline-enriched. Genes important for either oogenesis (Oo.) or spermatogenesis function (Sp.) were determined by comparing oocyte-only producing mutants [fem-1(lf)] to sperm-only producing mutants [fem-3(gf)] or vice versa (32). Other genes with germline expression are those genes that are germline-enriched but with neither oogenesis nor spermatogenesis known functions. Enrichment scores are statistically significant (P < 0.05). (B) Comparison of spr-5 gene targets with other published datasets. Four tissue-specific datasets showed statistically significant overrepresentation of spr-5 gene targets within the datasets. i. (33); ii. (34); iii. (35); iv. (36). Enrichment scores are statistically significant (P < 0.05). (C) Examination of DSBR repair gene expression levels in spr-5 mutants. qRT-PCR was performed on samples confirmed in Fig. S5 for candidate DSBR genes.
To explore a potentially transcriptional role for spr-5 in DSBR, we analyzed our gene target list for genes with known or predicted roles in DSBR. Each of the 130 target genes was examined in the literature and through online database searches (WormBase) for any reported role in DSBR, and none was found to be implicated in this process. We next set out to directly test DSBR candidate genes for misexpression in spr-5 mutant germlines using germline-specific quantitative RT-PCR (qRT-PCR). Between 5 and 20 gonads were dissected per genotype for these experiments and the RNA then used for standard RT followed by quantitative PCR. The samples were first assayed for known spr-5 targets based on our microarrays, including the LSD2 ortholog amx-1, which had been previously reported to be up-regulated in spr-5 mutants (12). The qRT-PCR shows similar levels of up-regulation as reported by the microarrays in late-generation animals with either spr-5 allele (Fig. S7), supporting the comparable sensitivity of each assay. The same samples show no discernable difference in the expression of a selection of DSBR-related genes, including the DNA damage checkpoint regulators atm-1 and cep-1, the meiotic endonuclease spo-11, and the DSBR genes rad-51 and rad-54 (Fig. 4C). These data do not exclude the possibility that DSBR genes or proteins could be misregulated in other ways in spr-5 mutants, but their transcriptional profile as assessed by microarray and qRT-PCR seems normal. This, along with the rapid relocalization of SPR-5 upon damage, suggests a potentially direct role for SPR-5 in meiotic DSBR. Over multiple generations, the build-up of H3K4me2 in spr-5 mutants could result in both transcriptional misregulation of somatic genes in the germline and a worsening of the DSBR phenotypes (in the absence of DSBR gene misregulation).
Recent studies have emphasized the connection between H3K4 methylation and DSBR. For example, H3K4me3 is enriched at recombination hotspots and strongly implicated in meiotic DSB formation and crossover formation, respectively, in yeast and mice (7, 21). Moreover, Prdm9, the meiotic H3K4me3 methyltransferase, has been recently implicated as a possible regulator of hotspot specification in mice and humans (21). H3K4me2 has also been identified to be enriched, albeit it to a lesser degree than H3K4me3, at the sites of active hotspots (4). Given that H3K4 methylation is an early marker of sites of recombination, perhaps the regulated removal of this modification is also important for processing of meiotic DSBs. Several lines of evidence support a role for SPR-5 in the efficient processing of DSBs, including the increase in p53/CEP-1–dependent apoptosis, the increase in RAD-51 foci in pachytene, and the IR sensitivity of spr-5 mutants. In C. elegans, DNA damage-triggered apoptosis occurs in a CEP-1–dependent manner near the end of pachytene (22). An increase in CEP-1–dependent apoptosis therefore indicates that more nuclei are reaching this stage of pachytene with unresolved DNA damage. Similarly, the elevated levels of RAD-51 foci observed during pachytene in spr-5 mutants and our analysis of rad-54 spr-5 double mutants, which ruled out an increase in DSB formation, suggest instead impaired DSBR progression. The fact that spr-5 mutant germlines are specifically sensitive to IR, which causes DSBs, further supports a defect in DSBR in the absence of spr-5.
However, DSBR is not completely abrogated in spr-5 mutants. Specifically, RAD-51 foci do not persist beyond late pachytene. Moreover, spr-5 mutants do not display entangled or frayed chromosomes at diakinesis, as is seen in rad-51 mutants (23, 24). Furthermore, late-generation spr-5 mutant populations do not contain higher frequencies of males or the appearance of pleiotropic phenotypes (i.e., Dpy, Unc, Pvul) indicative of errors in meiotic chromosome segregation or a mutator phenotype, respectively, due to unresolved DSBs. These findings suggest that animals lacking spr-5 experience a defect or delay in DSBR but are still able to repair DSBs. The finding of an increase in frayed or fragmented chromatin in spr-5;syp-1 mutants, which are reliant on intersister-based repair, also suggests that this defect affects both interhomolog and intersister-based pathways of DSBR. Furthermore, the role of SPR-5 may be partially redundant with other histone demethylases, explaining the ability of spr-5 mutant animals to eventually complete DSBR.
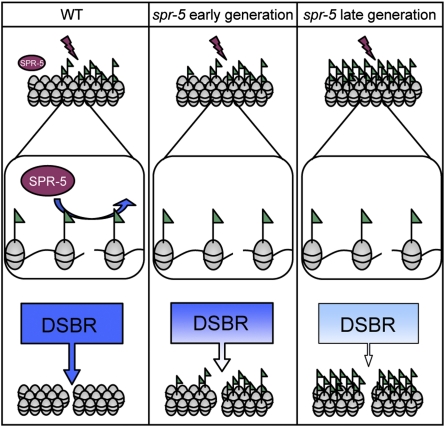
Taken together, the observation of rapid relocalization by SPR-5 upon DSB induction, along with the spr-5 mutant DSBR phenotypes, suggest a model whereby an early response by SPR-5 upon DSB induction is required for efficient DSBR (Fig. 5). In spr-5 mutants, DSBR is impaired, possibly owing to failure of SPR-5 to remove H3K4me2. If H3K4me2 does impair repair in some manner, the global and progressive increase of H3K4me2 in spr-5 mutants over multiple generations could therefore contribute to the progressive nature of the DSBR phenotypes as well (Fig. 5, Right). We propose that SPR-5 may act as an H3K4me2 demethylase at the sites of DNA damage during meiosis to promote DSBR, so that in its absence spr-5 mutant animals display a meiotic-specific phenotype of perturbed or delayed DSBR. SPR-5 does not seem to colocalize with either RPA-1 or RAD-51, which act at early to mid stages in DSBR (Fig. S8). However, these findings do not exclude an earlier interaction for SPR-5 with DSB sites, as might be predicted by the rapid observed kinetics of its relocalization.
Fig. 5.
Model for DSBR progression defect phenotype in both early- and late-generation spr-5 mutants. In wild-type animals, we propose that SPR-5 acts after DSBs occur to remove H3K4me2 (green flags), allowing DSBR factors to complete repair efficiently (Left). In early-generation spr-5 mutants, the global levels of H3K4me2 are minimally affected, but after DSBs occur H3K4me2 is not removed by SPR-5, leading to inefficient DSBR (Center). In late-generation spr-5 mutants, the lack of SPR-5 to act after damage and the general increase in H3K4me2 combine to further impair DSBR progression (Right).
The proposed removal of H3K4me2 by SPR-5 at the sites of DSBR may support efficient DSBR by several potential mechanisms. For example, recruitment of DSBR proteins may require SPR-5 directly or indirectly via the removal of hotspot-directed H3K4 methylation. In addition, H3K4me2 demethylation may be important for regulating chromatin structure in a manner that prevents additional DSB formation and/or transcription from occurring at or around the sites of DSBR. Future studies will differentiate between these nonmutually exclusive possibilities.
In conclusion, we have shown that SPR-5 is an H3K4me2 demethylase that is active in the C. elegans germline. In its absence, H3K4me2 builds up progressively in the germline. In addition, F2 generation spr-5 mutants show phenotypes indicative of impaired meiotic DSBR, such as increased DNA damage-induced germ cell apoptosis, increased RAD-51 foci at early- to mid-pachytene, and a specific sensitivity to DSB-inducing treatments. These data, along with the relocalization of SPR-5 protein upon DSB-inducing treatment and the absence of DSBR genes among spr-5 transcriptional targets, support an active and likely a direct role for SPR-5 in meiotic DSBR. The subsequent progressivity of the DSBR phenotypes along with the progressive increase in H3K4me2 is suggestive of a connection between H3K4me2 regulation and efficient DSBR. Furthermore, the microarray data suggest an additional role for spr-5 in the transcriptional repression of somatic genes in the germline. These studies both support previously reported transcriptional roles for histone demethylases as identified in various model systems, and introduce a unique role for a histone demethylase in meiotic DSBR through the use of the C. elegans germline.
Materials and Methods
Worm Culture and Strains.
The N2 Bristol strain was used as the wild-type background in these studies. A strain containing a transgene expressing RPA-1::YFP was used for colocalization studies (25). C. elegans strains were cultured at 20 °C under standard conditions as described in ref. 26. The following mutations and chromosome rearrangements were used: LGI: cep-1(lg12501), rad-54(ok615), hT2[bli-4(e937) let-?(q782) qIs48] (I; III), spr-5(by101), spr-5(by134); LGIV: spo-11(ok79), nT1 [unc-?(n754) let-?(m435)] (IV; V); LGV: syp-1(me17) (17, 19, 27–30). Further details can be found in SI Materials and Methods.
Supplementary Material
Acknowledgments
We thank Fei Lan, Nima Mosammaparast, Sarit Smolikov, and Kate Meyer for their assistance and helpful discussions. Some strains were provided by the Caenorhabditis Genetics Center. This work was supported by National Institutes of Health Grant R01GM058012 (to Y.S. and M.P.C.).
Footnotes
The authors declare no conflict of interest.
*This Direct Submission article had a prearranged editor.
This article contains supporting information online at www.pnas.org/lookup/suppl/doi:10.1073/pnas.1102298108/-/DCSupplemental.
References
- 1.Martin C, Zhang Y. The diverse functions of histone lysine methylation. Nat Rev Mol Cell Biol. 2005;6:838–849. doi: 10.1038/nrm1761. [DOI] [PubMed] [Google Scholar]
- 2.Ng SS, Yue WW, Oppermann U, Klose RJ. Dynamic protein methylation in chromatin biology. Cell Mol Life Sci. 2009;66:407–422. doi: 10.1007/s00018-008-8303-z. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Myers S, et al. Drive against hotspot motifs in primates implicates the PRDM9 gene in meiotic recombination. Science. 2010;327:876–879. doi: 10.1126/science.1182363. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Buard J, Barthès P, Grey C, de Massy B. Distinct histone modifications define initiation and repair of meiotic recombination in the mouse. EMBO J. 2009;28:2616–2624. doi: 10.1038/emboj.2009.207. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Baudat F, et al. PRDM9 is a major determinant of meiotic recombination hotspots in humans and mice. Science. 2010;327:836–840. doi: 10.1126/science.1183439. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Parvanov ED, Petkov PM, Paigen K. Prdm9 controls activation of mammalian recombination hotspots. Science. 2010;327:835. doi: 10.1126/science.1181495. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Borde V, et al. Histone H3 lysine 4 trimethylation marks meiotic recombination initiation sites. EMBO J. 2009;28:99–111. doi: 10.1038/emboj.2008.257. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Shi Y, et al. Histone demethylation mediated by the nuclear amine oxidase homolog LSD1. Cell. 2004;119:941–953. doi: 10.1016/j.cell.2004.12.012. [DOI] [PubMed] [Google Scholar]
- 9.Lan F, et al. S. pombe LSD1 homologs regulate heterochromatin propagation and euchromatic gene transcription. Mol Cell. 2007;26:89–101. doi: 10.1016/j.molcel.2007.02.023. [DOI] [PubMed] [Google Scholar]
- 10.Di Stefano L, Ji JY, Moon NS, Herr A, Dyson N. Mutation of Drosophila Lsd1 disrupts H3-K4 methylation, resulting in tissue-specific defects during development. Curr Biol. 2007;17:808–812. doi: 10.1016/j.cub.2007.03.068. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Rudolph T, et al. Heterochromatin formation in Drosophila is initiated through active removal of H3K4 methylation by the LSD1 homolog SU(VAR)3-3. Mol Cell. 2007;26:103–115. doi: 10.1016/j.molcel.2007.02.025. [DOI] [PubMed] [Google Scholar]
- 12.Katz DJ, Edwards TM, Reinke V, Kelly WG. A C. elegans LSD1 demethylase contributes to germline immortality by reprogramming epigenetic memory. Cell. 2009;137:308–320. doi: 10.1016/j.cell.2009.02.015. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Nottke A, Colaiácovo MP, Shi Y. Developmental roles of the histone lysine demethylases. Development. 2009;136:879–889. doi: 10.1242/dev.020966. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Karagiannis TC, El-Osta A. Chromatin modifications and DNA double-strand breaks: The current state of play. Leukemia. 2007;21:195–200. doi: 10.1038/sj.leu.2404478. [DOI] [PubMed] [Google Scholar]
- 15.Gartner A, Milstein S, Ahmed S, Hodgkin J, Hengartner MO. A conserved checkpoint pathway mediates DNA damage-induced apoptosis and cell cycle arrest in C. elegans. Mol Cell. 2000;5:435–443. doi: 10.1016/s1097-2765(00)80438-4. [DOI] [PubMed] [Google Scholar]
- 16.Gartner A, Boag PR, Blackwell TK. Germline survival and apoptosis. WormBook. 2008 doi: 10.1895/wormbook.1.145.1. Sep 4:1–20. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Dernburg AF, et al. Meiotic recombination in C. elegans initiates by a conserved mechanism and is dispensable for homologous chromosome synapsis. Cell. 1998;94:387–398. doi: 10.1016/s0092-8674(00)81481-6. [DOI] [PubMed] [Google Scholar]
- 18.Colaiácovo MP, et al. Synaptonemal complex assembly in C. elegans is dispensable for loading strand-exchange proteins but critical for proper completion of recombination. Dev Cell. 2003;5:463–474. doi: 10.1016/s1534-5807(03)00232-6. [DOI] [PubMed] [Google Scholar]
- 19.Mets DG, Meyer BJ. Condensins regulate meiotic DNA break distribution, thus crossover frequency, by controlling chromosome structure. Cell. 2009;139:73–86. doi: 10.1016/j.cell.2009.07.035. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Smolikov S, et al. Synapsis-defective mutants reveal a correlation between chromosome conformation and the mode of double-strand break repair during Caenorhabditis elegans meiosis. Genetics. 2007;176:2027–2033. doi: 10.1534/genetics.107.076968. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Cheung VG, Sherman SL, Feingold E. Genetics. Genetic control of hotspots. Science. 2010;327:791–792. doi: 10.1126/science.1187155. [DOI] [PubMed] [Google Scholar]
- 22.Schumacher B, Hofmann K, Boulton S, Gartner A. The C. elegans homolog of the p53 tumor suppressor is required for DNA damage-induced apoptosis. Curr Biol. 2001;11:1722–1727. doi: 10.1016/s0960-9822(01)00534-6. [DOI] [PubMed] [Google Scholar]
- 23.Takanami T, Mori A, Takahashi H, Higashitani A. Hyper-resistance of meiotic cells to radiation due to a strong expression of a single recA-like gene in Caenorhabditis elegans. Nucleic Acids Res. 2000;28:4232–4236. doi: 10.1093/nar/28.21.4232. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Rinaldo C, Bazzicalupo P, Ederle S, Hilliard M, La Volpe A. Roles for Caenorhabditis elegans rad-51 in meiosis and in resistance to ionizing radiation during development. Genetics. 2002;160:471–479. doi: 10.1093/genetics/160.2.471. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Stergiou L, Eberhard R, Doukoumetzidis K, Hengartner MO. NER and HR pathways act sequentially to promote UV-C-induced germ cell apoptosis in Caenorhabditis elegans. Cell Death Differ. 2011;18:897–906. doi: 10.1038/cdd.2010.158. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Brenner S. The genetics of Caenorhabditis elegans. Genetics. 1974;77:71–94. doi: 10.1093/genetics/77.1.71. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Eimer S, Lakowski B, Donhauser R, Baumeister R. Loss of spr-5 bypasses the requirement for the C.elegans presenilin sel-12 by derepressing hop-1. EMBO J. 2002;21:5787–5796. doi: 10.1093/emboj/cdf561. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Derry WB, Putzke AP, Rothman JH. Caenorhabditis elegans p53: Role in apoptosis, meiosis, and stress resistance. Science. 2001;294:591–595. doi: 10.1126/science.1065486. [DOI] [PubMed] [Google Scholar]
- 29.Schumacher B, et al. Translational repression of C. elegans p53 by GLD-1 regulates DNA damage-induced apoptosis. Cell. 2005;120:357–368. doi: 10.1016/j.cell.2004.12.009. [DOI] [PubMed] [Google Scholar]
- 30.MacQueen AJ, Colaiácovo MP, McDonald K, Villeneuve AM. Synapsis-dependent and -independent mechanisms stabilize homolog pairing during meiotic prophase in C. elegans. Genes Dev. 2002;16:2428–2442. doi: 10.1101/gad.1011602. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Whetstine JR, et al. Reversal of histone lysine trimethylation by the JMJD2 family of histone demethylases. Cell. 2006;125:467–481. doi: 10.1016/j.cell.2006.03.028. [DOI] [PubMed] [Google Scholar]
- 32.Reinke V, Gil IS, Ward S, Kazmer K. Genome-wide germline-enriched and sex-biased expression profiles in Caenorhabditis elegans. Development. 2004;131:311–323. doi: 10.1242/dev.00914. [DOI] [PubMed] [Google Scholar]
- 33.Kunitomo H, Uesugi H, Kohara Y, Iino Y. Identification of ciliated sensory neuron-expressed genes in Caenorhabditis elegans using targeted pull-down of poly(A) tails. Genome Biol. 2005;6:R17. doi: 10.1186/gb-2005-6-2-r17. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Pauli F, Liu Y, Kim YA, Chen PJ, Kim SK. Chromosomal clustering and GATA transcriptional regulation of intestine-expressed genes in C. elegans. Development. 2006;133:287–295. doi: 10.1242/dev.02185. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Gaudet J, Mango SE. Regulation of organogenesis by the Caenorhabditis elegans FoxA protein PHA-4. Science. 2002;295:821–825. doi: 10.1126/science.1065175. [DOI] [PubMed] [Google Scholar]
- 36.Wang J, Kim SK. Global analysis of dauer gene expression in Caenorhabditis elegans. Development. 2003;130:1621–1634. doi: 10.1242/dev.00363. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.