Abstract
Cell surfaces are fertile ground for chemists and material scientists to manipulate or augment cell functions and phenotypes. This not only helps to answer basic biology questions but also has diagnostic and therapeutic applications. In this review, we summarize the most recent advances in the engineering of the cell surface. In particular, we focus on the potential applications of surface engineered cells for 1) targeting cells to desirable sites in cell therapy, 2) programming assembly of cells for tissue engineering, 3) bioimaging and sensing, and ultimately 4) manipulating cell biology.
The interface of chemistry, material science and biology presents many opportunities for developing innovative tools to answer fundamental biology questions and diagnose/treat various diseases. The past decade has witnessed an explosion in the development of novel materials and methods, including some on a nano-scale, which can be applied to fields including tissue engineering, diagnostics, drug delivery, and medical devices. A particularly exciting subject in the field is the direct engineering and manipulation of living cells, the basic structural and functional unit of living organisms. Excellent examples include controlling cell fate by biomaterial scaffolds1,2, labeling cells with molecular and nanoparticle probes for imaging and visualization of cellular processes and molecular pathways3–5, delivery of diverse species into cells6,7, and patterning cells for drug discovery8–10. This review focuses on the engineering of the cell surface, a fertile ground for chemists and material scientists to manipulate cell phenotypes and biological fates. This will open up new avenues for biologists to interrogate basic cellular functions, or advance cell-based therapies and diagnostics. The cell membrane is a highly heterogeneous and dynamic environment comprising mainly proteins, lipids, and carbohydrates, which mediate cell-cell and cell-niche communication and intracellular signaling, ultimately governing cell fate11,12. To date, cell surface engineering has primarily been a subject of molecular biology. However, tools developed by chemists and material scientists provide simple alternatives to the genetic and biosynthetic approaches. Specifically, targeting molecules, molecular and nanoparticle probes, polymer patches, and nanostructures can be introduced onto the cell surface by conjugation (covalently and non-covalently), or by utilizing biomolecular recognition such as antibody/antigen and biotin/streptavidin interactions. These tools greatly enhance our ability to use and manipulate cells, and answer biological questions. This review highlights the most recent developments in this field with particular focuses on the use of cell surface engineering strategies;
targeting cells to desirable sites in cell therapy,
programming assembly of cells with substrates or cells in tissue engineering,
bioimaging and sensing, and
manipulating cell biology.
Cell encapsulation, cell-matrix interactions, and the detailed mechanisms of chemical reactions involved in cell surface modification, which are subjects of several recent excellent reviews11,13–19, will not be discussed here.
Cell targeting
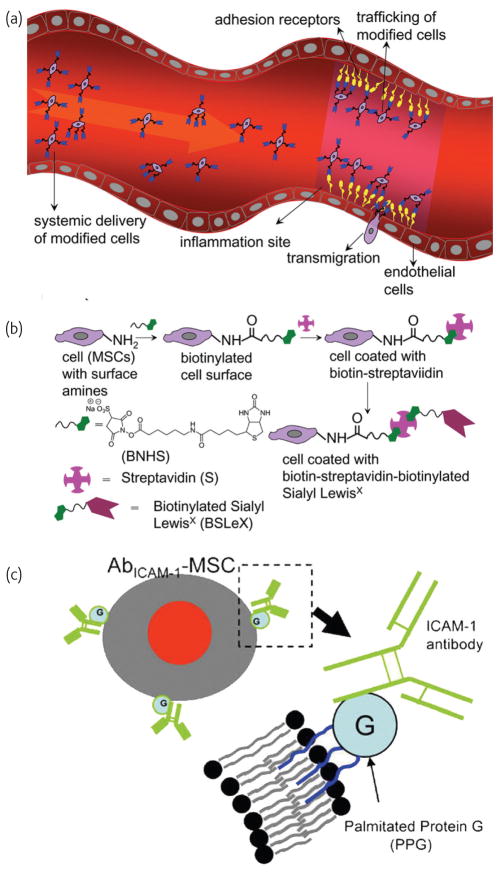
Delivery of systemically infused cells, particularly stem/progenitor cells, into damaged or diseased tissue holds enormous promise for the treatment of a variety of diseases worldwide20. For instance, hematopoietic stem cell (HSC) transplantation (i.e. bone marrow transplantation), which has been used in clinic for several decades to treat blood diseases and cancer, requires systemically infused HSCs to home to the bone marrow from the blood. Mesenchymal stem cells (MSCs) are similarly believed to home to various sites of inflammation and injury in the body after being systemically infused. MSCs are capable of differentiating into connective tissue cells types that can produce bone, cartilage and fat, and produce immunomodulatory cytokines, are currently being tested in clinical trials for treatment of numerous diseases including graft versus host disease, myocardial infarction, multiple sclerosis, and skeletal tissue repair, among others20. Unfortunately, systemically administered therapeutic cells home to the target sites at low efficiencies (typically < 1%), which is due to, at least in part, the lack of expression (or loss during culture expansion) of key homing receptors21. Therefore, efforts have been made to introduce cell homing ligands onto cell membrane. Cell homing ligands (on the homing cell) and receptors (on the endothelium) allow homing cells to tether, roll, adhere and then transmigrate on endothelium as part of the cell homing cascade. Methods include genetic22–24 and enzymatic engineering25, treatment with cytokines26 and, as will be discussed here in detail, chemical approaches27,28 (Fig. 1a). For instance, we have recently developed a simple platform technology to chemically attach cell adhesion molecules to MSC surface to improve homing efficiency to inflamed tissues27. Specifically, this chemical approach (Fig. 1b) involves a stepwise process including;
Fig. 1.
a) Schematic showing targeting of cells from blood to inflamed endothelium expressing receptors that bind to ligands engineered on the cell surface. Two methods of engineering the homing ligand onto the cell surfaces, using covalent and noncovalent chemistry, are shown in b) and c), respectively. b) Stepwise process of attaching SLeX to MSCs using biotin-streptavidin linkers, and c) incorporation of palmitated protein G into the cell membrane and subsequent conjugation with an ICAM-1 antibody.(a) and b) reprinted with permission from27. ©ACS 2008. c) adapted with permission from28. ©Elsevier 2009).
treatment of cell amine groups with sulfonated biotinyl-Nhydroxy- succinimide (NHS) to introduce biotin groups on the cell surface;
addition of streptavidin that binds to the biotin on the cell surface and presents unoccupied binding sites, and
attachment of biotinylated homing ligands. In our model system, a biotinylated sialyl Lewis X (SLeX), a ligand that binds to P and E selectins expressed on the inflamed endothelium and allows cells to roll on the endothelial layer, was conjugated on the MSC surface.
The SLeX modified MSCs demonstrated a robust rolling response on a P-selectin coated substrate under shear stress conditions in vitro27 and on inflamed endothelium in vivo (Sarkar et al. unpublished). In addition, homing ligands can be introduced onto the cell membrane using non-covalent chemical modification28,29. In a recent work from Dennis and co-workers28, MSCs were treated with palmitated protein G where the palmitate chain was incorporated into lipid bilayer via hydrophobic interactions and protein G provides generic binding sites for antibodies (Fig. 1c). In their proof-of-concept work, intercellular cell adhesion molecule-1 (ICAM-1) antibodies were conjugated onto MSCs which enabled the cells to bind to ICAM-1, a critical adhesion molecule expressed on activated endothelium. Note that these chemical approaches to modify the cell surface and immobilize required ligands are not limited to MSCs or the above-mentioned ligands. Rather they should have broad implications on cell therapies that utilize systemic administration and require targeting of cells to specific tissues. Presumably, if we know which receptors are present on endothelium in a targeted anatomical site, we could deliver cells specifically to that site by engineering the complementary ligands onto the cell surface.
Programmed cell-substrate and cell-cell assembly
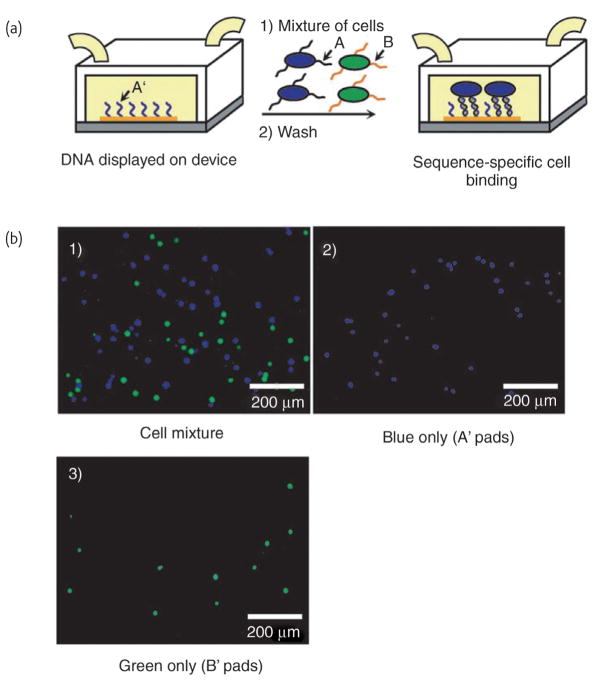
The precise 3D arrangement and orientation of cells and their substrate (e.g. extracellular matrix), is a key criterion in cell-based assays and microfluidic devices as well as tissue engineering where one goal is to mimic the 3D complexity of native tissues30. Towards this end, chemists and material scientists have begun to engineer cells with biorecognition moieties that are not possessed by native cells to control cell-cell and cell-substrate interactions31,32. Francis and colleagues have demonstrated the use of single-stranded DNA (ssDNA) to mediate cell adhesion on a substrate33. In their work, ssDNA was attached onto cells by a Staudinger ligation reaction between azides (delivered to cell surface glycans via a metabolism process)34 and phosphine groups modified on ssDNA or by a simple NHS ester (on DNA) and NH2 (on cell surface) chemistry35. The ssDNAs attached on cells directed the assembly of cells on complementary DNA-attached substrate via DNA hybridization (Fig. 2). Such DNA-modified cells can also be assembled, on a single cell level, on microfabricated patterned sensor devices that may be used for single cell analyses, such as measuring cellular electrical and metabolic activation. In a more recent work from Mathies and co-workers36, an array of lithographically patterned iridium oxide pH microelectrodes was enclosed within a microfluidic channel on which ssDNA was attached using silane chemistry. Such devices could then capture complementary DNAmodified cells and measure the single cell metabolism using pH as an indicator. They demonstrated that such devices are able to discriminate between healthy primary T cells and cancerous Jurkat T cells that have a higher metabolism.
Fig. 2.
DNA sequence-specific capture cells on substrate. a) General scheme of experiment where blue and green dye-labeled Jurkat cell populations were functionalized with DNA sequence A or B, respectively. An equal number of cells were combined and the mixture was introduced into a microfluidic device with DNA sequence bound to one surface. Only cells with complementary DNA sequences should bind postwash. b) Fluorescence images of 1) the cell mixture prewash; 2) an A′-coated pad postwash; 3) a B′-coated pad postwash. (Reprinted with permission from33. ©Wiley-VCH Verlag GmbH & Co. KGaA 2006).
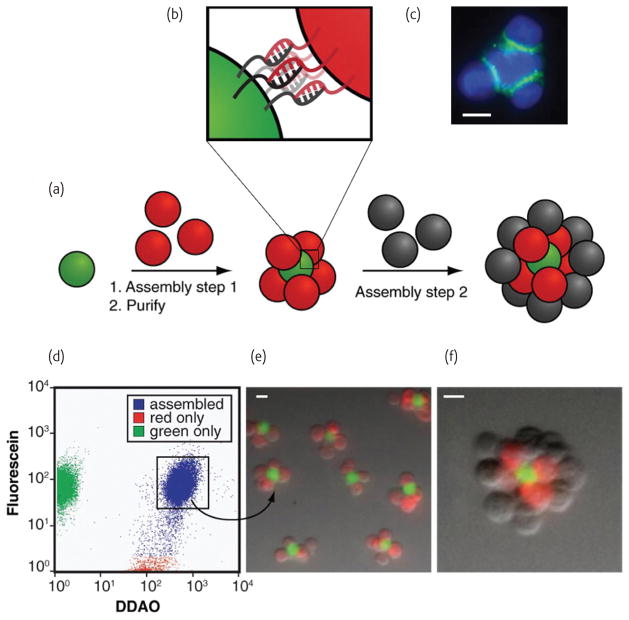
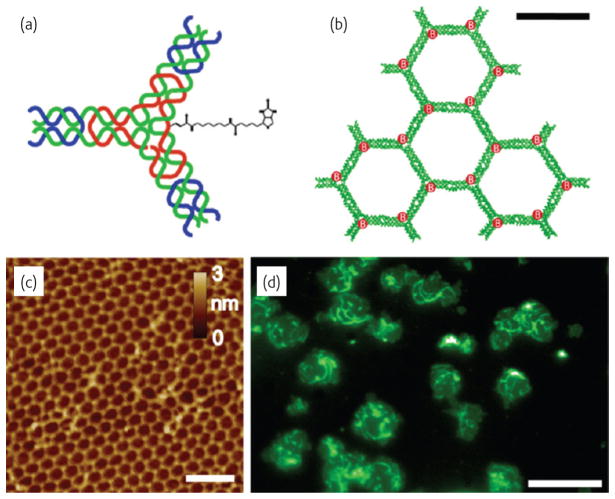
The DNA-based approach can also be used for the programmable assembly of 3-D cell structures, which may find applications in synthesizing artificial tissues37,38. Recently, Gartner and Bertozzi elegantly demonstrated that DNA-modified cells can be assembled into microtissues in a well-defined manner (i.e. structures were controlled by varying the DNA site density on the cell surface, cell concentration, and DNA sequence)38. Briefly, by adjusting the stoichiometry (i.e. 1:50) of two Jurkat cell population modified with complementary DNA strands, single cell clusters were produced where the limiting cell type was surrounded by cells added in excess (Fig. 3a,b). Interestingly, using fluorescent labeled DNA, they showed that DNA strands are clustered at the cell-cell interface (Fig. 3c), confirming their specific role in cell-cell assembly. Significantly, such assembled cell clusters could be isolated and purified using flow cytometry, and subsequently used as the core for further iterative cell assembly, i.e. adding another layer of DNA-modified cells (Fig. 3d–f). More importantly, they demonstrated that cells within the assembled microtissues communicated via a paracrine mechanism, which is a prerequisite for the development of therapeutically functional tissues. Specifically, in their proof-of-concept model, a CHO cell line engineered to express growth factor interleukin-3 (IL-3) was used as a first building block and the second was an untransformed hematopoietic progenitor cell line (FL5.12) whose survival and replication depends on the presence of IL-3. Cell composites were assembled using the above-mentioned DNA approach and then embedded within a 3-dimensional agarose matrix. After 16 h, the cell structures underwent accelerated growth due to the presence of IL-3. By contrast, when CHO cells lacking the gene coding in IL-3 were used in the microtissue, FL5.12 cells showed no growth and instead underwent a morphological change corresponding to the absence of IL-3 induced apoptosis. Finally, other advantages of using DNA assembly in such systems include their reversibility and versatility with respect to engineering multiple orthogonal cell-cell interactions. In addition to ssDNA, self-assembled cargo-carrying DNA arrays (Fig. 4) can also be attached onto cell surface (Fig. 5a). Reich and colleagues attached hexagonal DNA arrays on cells using streptavidin and antibody as bridges39. In their study, it was observed that one or two DNA array patches often attached to one cell, and that single DNA arrays often bridged cells (Fig. 5c–e). Occasionally, micron-sized DNA arrays seemed capable of binding to and enveloping multiple cells into larger cellular structures (Fig. 5b). These larger structures may have the potential to form the foundation for the construction of tissues or organs for transplantation in tissue engineering.
Fig. 3.
a) General scheme of multilayered cell assembly using DNA hybridization. (b)Schematic of DNA hybridization between 2 cells (shown in red and green) c) Fluorescence image of a cluster of assembled cells (blue) with intercellular DNA linkages (green). Clustering of DNA crosslinkers cellular interface can be observed. d) Gating of assembled cells in fluorescence activated flow cytometry. e) Fluorescence image showing assembled cells isolated by flow cytometry. f) Fluorescence image of an assembled cell cluster from (e) which underwent a second assembly step to acquire another layer of cells. All scale bars are 10μm. (Reprinted with permission from38. ©PNAS 2009)
Fig. 4.
Assembly of fluorescent DNA hexagonal arrays. (a) Diagram of the biotinylated three-point-star monomer used to form the DNA arrays. (b) Diagram of an 18 unit biotinylated hexagonal DNA array (30 nm scale bar). (c) AFM image of a biotinylated hexagonal DNA array (100 nm scale bar). (d) Biotinylated hexagonal DNA arrays labeled with YOYO 1 DNA staining dye on a mica surface (10 μm scale bar). (Reprinted with permission from39. ©ACS 2009).
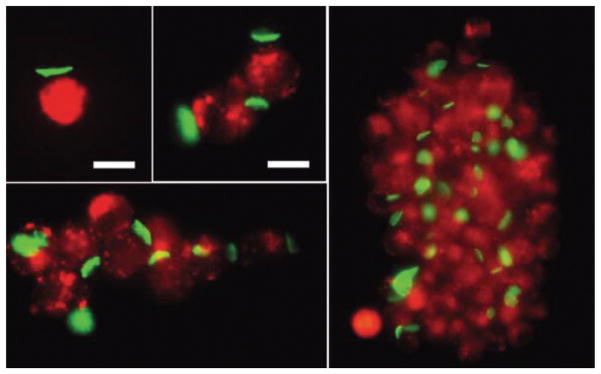
Fig. 5.
(a) Image of fluorescent DNA hexagonal arrays on HeLa cells, acquired by confocal microscopy. DNA arrays are green and HeLa cell surfaces red; HeLa cytoplasm is blue (50 μm scale bar). (b) Image of f a large DNA array binding and enveloping multiple HeLa cells, acquired by confocal microscopy. DNA arrays are green and HeLa cell surfaces red; HeLa cytoplasm is blue (50 μm scale bar). (c–e) Fluorescent DNA arrays bound to HeLa cells visualized in DIC mode (10 μm scale bar). (Reprinted with permission from39. ©ACS 2009).
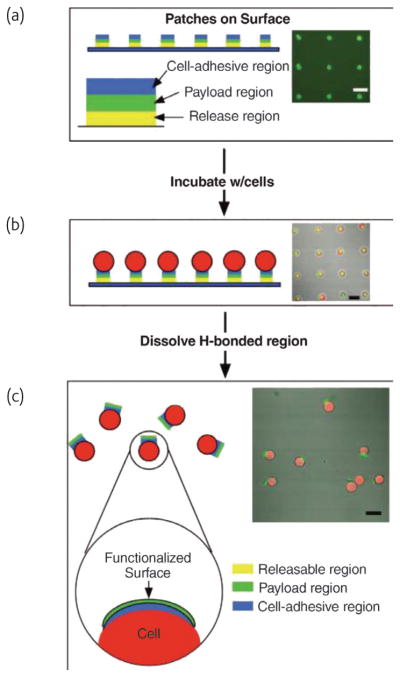
Engineered cell-cell interactions in a bottom-up tissue engineering approach can also be achieved by attaching cell-adhesive polyelectrolyte patches on the cell surface. Swiston et al. recently used a photolithographic-pattering technique to engineer multilayer polymer patches containing a payload component (i.e. super-paramagnetic nanoparticles) onto T cell surfaces (Fig. 6)40. The magnetic nanoparticles allowed cells to be spatially manipulated using an external magnetic field. Importantly, patch-functionalized cells could be programmed to assemble into sheet, rings, or cluster structures. Patches that are cell-adhesive may allow the polymerization of cellular species, with individual cells being the monomers and the patches serving as bifunctional linkages (Fig. 7), for construction of macroscopic, functional tissues.
Fig. 6.
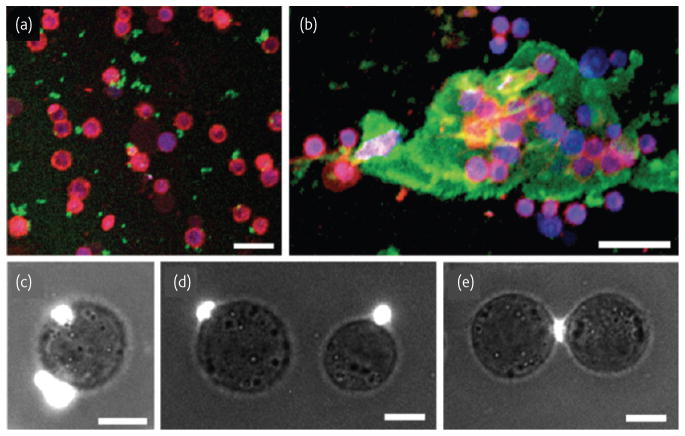
Overview of a cell functionalization scheme, using cell-adhesive polyelectrolyte patches. In this particular example, cell adhesive region, payload region and release region are chitosan-hyaluronic acid, FITCpoly( allylamine hydrochloride), poly(methylacrylic acid)-poly(Nisopropylacrylamide), respectively. (a) A regular array of surface-bound patches spaced 50 μm apart. Payload regions are labeled green. After cell incubation and attachment (b), a majority (85 ± 3%) of the surface bound patches are occupied. Cell cytoplasm is labeled red. (c) After the temperature is reduced to 4°C for 30 min, the patches are released from the surface while remaining attached to the cell membrane. All scale bars are 25 μm. (Reprinted with permission from40. ©ACS 2008).
Fig. 7.
An overlay of fluorescent microscopy images demonstrating different degrees of cell aggregation mediated by polymer patches. Cells and polymer patches are labeled in red and green, respectively. Scale bar is 10 μm. (Reprinted with permission from40. ©ACS 2008).
Bioimaging and sensing
One particularly important application of cell surface engineering is to introduce sensing and imaging probes to visualize cellular and molecular events both in cells and living organisms. The Bertozzi group at University of California, Berkerley has pioneered the use of chemical reporters for such purposes with a focus on those that can be incorporated through metabolic pathways. Such systems have been summarized recently by the same group in several excellent reviews which we refer to the interested readers11,16–19,41.
An example of their technology is the engineering of chromatic polydiacetylene (PDA) polymer patches onto cell surfaces for imaging structural perturbations of the membrane bilayer42. Specifically, vesicles comprising of diacetylene and lipid molecules including 1,2-dimyristoyl-sn-glycero-3-phosphoethanolamine and 1,2-dimyristoyl-sn-glycero-3-[phospho-rac-(1-glycerol)] were first formed in solution. These lipid vesicles were then fused with cell membrane and PDA was formed in situ upon UV-mediated polymerization. The conjugated PDA nanopatches on cells exhibited an intense blue color (Fig. 8a, left tube) and underwent a blue-to-red color change (Fig. 8a, right tube) upon structural perturbations of the cell membrane due to conformational transitions in the conjugated (ene–yne) polymer backbone. During this blue-to-red color transition, the initial non-fluorescent PDA also emitted fluorescence at 560 nm and 640 nm (Fig. 8b,c). The PDA nanopatches can therefore be used to visualize the cell bilayer perturbations induced by, for example, lidocaine, polymyxin-B, cholesterol, and oleic acid and to screen toxic pesticides and other environmentally sensitive small molecules43.
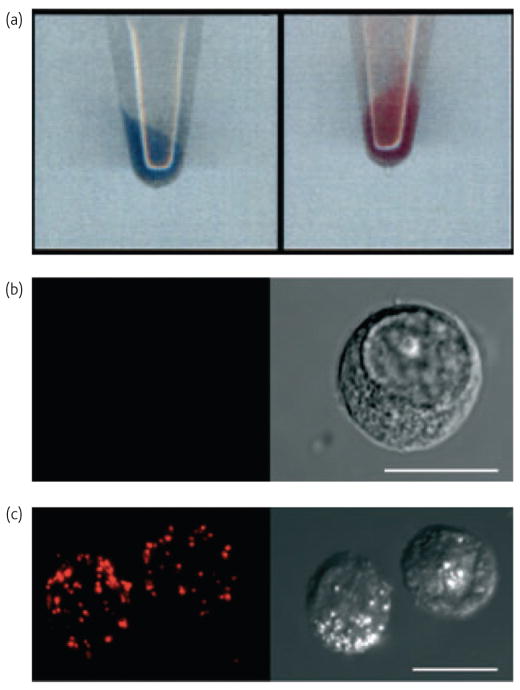
Fig. 8.
a) Photographs of tubes containing PDA-labeled cells after sedimentation. Left tube: blue cells before treatment; right tube: cells after incubation with oleic acid (180 μM), b) and c) are confocal microscopy fluorescence images (left) and transmission differential interference contrast (DIC) microscopy images (right) of PDA-labeled U937 cells. b) and c) show cells before and after polymyxin-B (50 μM) treatment, respectively. Scale bars in all images correspond to 10 μm. (Reprinted with permission from42. ©Wiley-VCH Verlag GmbH & Co. KGaA 2005).
Manipulating cell biological fate
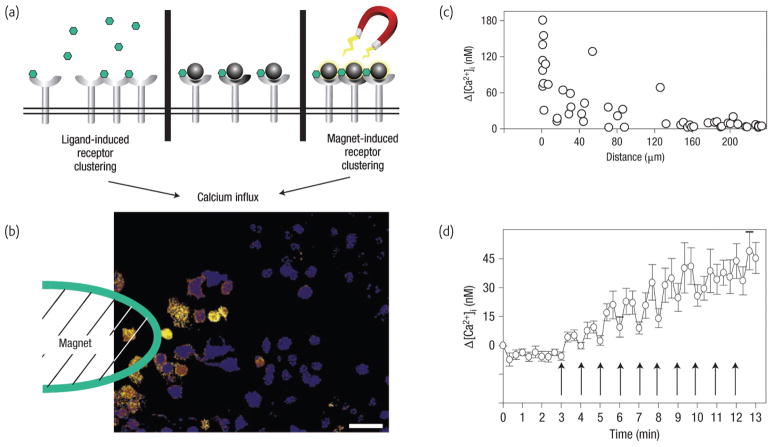
One ultimate goal of engineering the cell surface is to use externally applied chemical moieties and materials to control cell biology. As an initial effort, Ingber and co-workers have demonstrated the use of magnetic nanoparticles on cell membrane to regulate cellular signaling pathways44. Specifically, mast cells express membrane FcεRI receptors that can bind to the Fc region of IgE molecules (Fig. 9a). Multivalent antigens (dinitrophenyl, DNP) that bind to IgE can then induce FcεRI receptor oligomerization on cell membrane whereas monovalent antigens do not. The FcεRI receptor clustering quickly triggers an intracellular signaling response characterized by a rapid rise of cytosolic Ca2+, which in turn leads to the local inflammatory response of mast cells. Instead of using antigen ligand to induce such signaling pathway, Ingber and colleagues immobilized DNP-attached magnetic nanoparticles45 on IgE-tethered mast cells44 (Fig. 9a). They ensured that only one DNP molecule was attached on each particle so that receptor clustering does not occur. When an external magnetic field is applied, magnetic nanoparticles clustered by themselves on the cell membrane, which caused the oligomerization of FcεRI receptors, a process which could be visualized by the local Ca2+ concentration change (Fig. 9b,c). Significantly, such magnetic nanoparticle mediated cellular signaling is reversible: switching on and off the magnetic field directly corresponds to a local Ca2+ concentration oscillation (Fig. 9d).
Fig. 9.
Nanomagnetic control of receptor signal transduction. a) The biochemical mechanism that stimulates downstream signalling (left) involves the binding of multi-valent ligands (represented by green hexagons) that induce oligomerization of individual IgE/FcεRI receptor complexes. In the magnetic switch, monovalent ligand-coated magnetic nanobeads (dark grey circles), similar in size to individual FcεRI receptors, bind individual IgE/FcεRI receptor complexes without inducing receptor clustering (centre). However, applying a magnetic field that magnetizes the beads and pulls them into tight clusters (right) rapidly switches on receptor oligomerization and calcium signalling. b) The pseudocoloured microfluorimetric image shows the local induction of calcium signaling (yellow) in cells near the tip of the electromagnet within 20 s of the field being applied. Scale bar is 50 μm. c) Quantification of peak changes in intracellular calcium relative to time 0 measured within individual cells during a 1-min pulse of applied magnetic force as a function of the distance of the tip from the cell surface. d) Effect of a rapid cyclical magnetic stimulation regimen (40 s on, 20 s off) on intracellular calcium signaling. (Reprinted with permission from44. ©NPG 2008).
Conclusion and perspectives
While promising, cell surface engineering research is still in its infancy and many issues remain to be addressed. One technical challenge is adapting chemical modification methods to suit the highly dynamic and heterogeneous environment of the cell. For instance, when chemical and material moieties are introduced on the cell surface, one has to consider the following factors: site density, distribution, mobility, and stability (shedding or internalization) that may be critical for down-stream applications. Furthermore, the newly introduced moieties on the cell surface should be accessible for the relevant application (e.g. binding to receptors on other cells), and not hindered by steric effects27 brought about by conformational change or the presence of other cell surface proteins.
It is important to consider and assess how cell surface modifications and introduction of external materials impact the cell’s phenotype. Most of the above-mentioned examples have characterized the impact of cell surface modification on cell viability but more detailed characterization is required. We studied the impact of SLeX modification on MSCs’ native functions including adhesion, proliferation, viability, differentiation, and in vivo trans-endothelium migration27. Rubner et al. studied the influence of cell membrane-attached polymer patches on T cell viability and migration40. Overall, it appears possible to modify cell membranes with specific ligands, micron sized patches, and nanoparticles46, without significantly altering cell phenotype. Nevertheless, more detailed analysis is required including examination of how specific modifications may impact gene expression profiles.
During the past decade, chemists and materials scientists have joined biologists to physically and chemically manipulate and engineer cell membranes for a variety of applications. Desirable chemical functionalities and materials can be introduced onto the cell surface by covalent and noncovalent chemistry as well through specific biological recognition events including antibody/antigen or ligand/receptor interactions. Cell surface engineering represents a powerful new approach that will likely find broad utilities in tissue engineering, cell therapy, cell-based assays and devices, drug delivery and discovery, biosensing/bioimaging, cell separation, and manipulation of cell biological fate.
Acknowledgments
This work was funded by grants from the National Institute of Health (R03DE019191) and the American Heart Association (#0970178N) to JMK. WZ is funded by the Human Frontier Science Program (HFSP).
References
- 1.Huebsch N, Mooney DJ. Nature. 2009;462(7272):426. doi: 10.1038/nature08601. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Zhao W, Karp JM. Chem Bio Chem. 2009;10(14):2308. doi: 10.1002/cbic.200900445. [DOI] [PubMed] [Google Scholar]
- 3.Anker JN, et al. Nat Mater. 2008;7(6):442. doi: 10.1038/nmat2162. [DOI] [PubMed] [Google Scholar]
- 4.Zhao W, Karp JM. Nat Mater. 2009;8(6):453. doi: 10.1038/nmat2463. [DOI] [PubMed] [Google Scholar]
- 5.Giepmans BNG, et al. Science. 2006;312(5771):217. doi: 10.1126/science.1124618. [DOI] [PubMed] [Google Scholar]
- 6.Schmidt U, et al. FASEB J. 2001;15(9):1646. doi: 10.1096/fj.00-0645fje. [DOI] [PubMed] [Google Scholar]
- 7.Akin D, et al. Nature Nanotech. 2007;2(7):441. doi: 10.1038/nnano.2007.149. [DOI] [PubMed] [Google Scholar]
- 8.Chen S, Smith LM. Langmuir. 2009;25(20):12275. doi: 10.1021/la9017135. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Kang L, et al. J Biomed Mater Res. 2009;Part A doi: 10.1002/jbm.a.32557. (Article online in advance of print) [DOI] [Google Scholar]
- 10.Falconnet D, et al. Biomaterials. 2006;27(16):3044. doi: 10.1016/j.biomaterials.2005.12.024. [DOI] [PubMed] [Google Scholar]
- 11.Mahal LK, Bertozzi CR. Chemistry & Biology. 1997;4(6):415. doi: 10.1016/s1074-5521(97)90193-9. [DOI] [PubMed] [Google Scholar]
- 12.Kellam B, et al. Chem Soc Rev. 2003;32:327. doi: 10.1039/b211643j. [DOI] [PubMed] [Google Scholar]
- 13.Orive G, et al. Biomaterials. 2006;27(20):3691. doi: 10.1016/j.biomaterials.2006.02.048. [DOI] [PubMed] [Google Scholar]
- 14.Beck J, et al. Tissue Eng. 2007;13(3):589. doi: 10.1089/ten.2006.0183. [DOI] [PubMed] [Google Scholar]
- 15.Stevens MM, George JH. Science. 2005;310(5751):1135. doi: 10.1126/science.1106587. [DOI] [PubMed] [Google Scholar]
- 16.Cook BN, Bertozzi CR. Bioorg Med Chem. 2002;10(4):829. doi: 10.1016/s0968-0896(01)00376-5. [DOI] [PubMed] [Google Scholar]
- 17.Hang HC, Bertozzi CR. Acc Chem Res. 2001;34(9):727. doi: 10.1021/ar9901570. [DOI] [PubMed] [Google Scholar]
- 18.Laughlin ST, Bertozzi CR. Proc Nat Acad Sci USA. 2009;106(1):12. doi: 10.1073/pnas.0811481106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Agard NJ, Bertozzi CR. Acc Chem Res. 2009;42(6):788. doi: 10.1021/ar800267j. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Karp JM, Teo GSL. Cell Stem Cell. 2009;4(3):206. doi: 10.1016/j.stem.2009.02.001. [DOI] [PubMed] [Google Scholar]
- 21.Rombouts WJC, Ploemacher RE. Leukemia. 2003;17(1):160. doi: 10.1038/sj.leu.2402763. [DOI] [PubMed] [Google Scholar]
- 22.Kumar S, et al. Gene Ther. 2010;17:105. doi: 10.1038/gt.2009.116. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Brenner S, et al. Stem Cells. 2004;22(7):1128. doi: 10.1634/stemcells.2003-0196. [DOI] [PubMed] [Google Scholar]
- 24.Zhang D, et al. J Mol Cell Cardiol. 2008;44(2):281. doi: 10.1016/j.yjmcc.2007.11.010. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Sackstein R, et al. Nat Med. 2008;14(2):181. doi: 10.1038/nm1703. [DOI] [PubMed] [Google Scholar]
- 26.Chavakis E, et al. J Mol Cell Cardiol. 2008;45(4):514. doi: 10.1016/j.yjmcc.2008.01.004. [DOI] [PubMed] [Google Scholar]
- 27.Sarkar D, et al. Bioconjugate Chem. 2008;19(11):2105. doi: 10.1021/bc800345q. [DOI] [PubMed] [Google Scholar]
- 28.Ko IK, et al. Biomaterials. 2009;30(22):3702. doi: 10.1016/j.biomaterials.2009.03.038. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Rabuka D, et al. J Am Chem Soc. 2008;130(18):5947. doi: 10.1021/ja710644g. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Khademhosseini A, et al. Scientific American. 2009;300:64. doi: 10.1038/scientificamerican0509-64. [DOI] [PubMed] [Google Scholar]
- 31.Borisenko GG, et al. Nucleic Acids Res. 2009;37(4):e28. doi: 10.1093/nar/gkn1034. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Kamitani R, et al. Chem Bio Chem. 2009;10(2):230. doi: 10.1002/cbic.200800621. [DOI] [PubMed] [Google Scholar]
- 33.Chandra RA, et al. Angew Chem Int Ed. 2006;45(6):896. doi: 10.1002/anie.200502421. [DOI] [PubMed] [Google Scholar]
- 34.Prescher JA, et al. Nature. 2004;430(7002):873. doi: 10.1038/nature02791. [DOI] [PubMed] [Google Scholar]
- 35.Hsiao SC, et al. Langmuir. 2009;25(12):6985. doi: 10.1021/la900150n. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Douglas ES, et al. Lab Chip. 2009;9(14):2010. doi: 10.1039/b821690h. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Paul ADB, et al. Biotechnol Bioeng. 2007;97(6):1617. doi: 10.1002/bit.21343. [DOI] [PubMed] [Google Scholar]
- 38.Gartner ZJ, Bertozzi CR. Proc Nat Acad Sci USA. 2009;106(12):4606. doi: 10.1073/pnas.0900717106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Koyfman AY, et al. J Am Chem Soc. 2009;131(40):14237. doi: 10.1021/ja9015638. [DOI] [PubMed] [Google Scholar]
- 40.Swiston AJ, et al. Nano Lett. 2008;8(12):4446. doi: 10.1021/nl802404h. [DOI] [PubMed] [Google Scholar]
- 41.Prescher JA, Bertozzi CR. Nat Chem Biol. 2005;1(1):13. doi: 10.1038/nchembio0605-13. [DOI] [PubMed] [Google Scholar]
- 42.Orynbayeva Z, et al. Angew Chem Int Ed. 2005;44(7):1092. doi: 10.1002/anie.200462393. [DOI] [PubMed] [Google Scholar]
- 43.Mech A, et al. Chem Res Toxicol. 2009;22(1):90. doi: 10.1021/tx800206t. [DOI] [PubMed] [Google Scholar]
- 44.Mannix RJ, et al. Nature Nanotech. 2008;3(1):36. doi: 10.1038/nnano.2007.418. [DOI] [PubMed] [Google Scholar]
- 45.Yung CW, et al. Lab Chip. 2009;9(9):1171. doi: 10.1039/b816986a. [DOI] [PubMed] [Google Scholar]
- 46.Cheng H, et al. ACS Nano. doi: 10.1021/nn901319y. (in press) [DOI] [Google Scholar]