Abstract
We have characterized mRNA expression and transcription of the mouse α- and β-globin loci during development. S1 nuclease and primary transcript in situ hybridization analyses demonstrate that all seven murine globin genes (ζ, α1, α2, εy, βH1, βmaj, and βmin) are transcribed during primitive erythropoiesis, however transcription of the ζ, εy, and βH1 genes is restricted to the primitive erythroid lineage. Transcription of the βmaj and βmin genes in primitive cells is EKLF-dependent demonstrating EKLF activity in embryonic red cells. Novel kinetic analyses suggest that multigene expression in the β locus occurs via alternating single-gene transcription whereas coinitiation cannot be ruled out in the α locus. Transcriptional activation of the individual murine β genes in primitive cells correlates inversely with their distance from the locus control region, in contrast with the human β locus in which the adult genes are only activated in definitive erythroid cells. The results suggest that the multigene expression mechanism of alternating transcription is evolutionarily conserved between mouse and human β globin loci but that the timing of activation of the adult genes is altered, indicating important fundamental differences in globin gene switching.
Keywords: Globin genes, transcription, locus control region, gene competition, mechanism of transcriptional control, in vivo
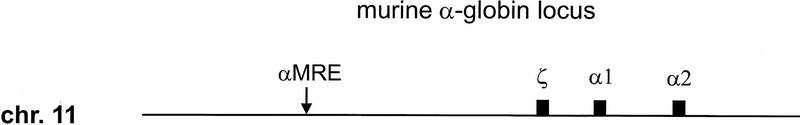
The murine α- and β-globin loci are multigene clusters located on chromosomes 11 and 7, respectively (Fig. 1). They are highly homologous to their human counterparts in organizational structure and function and represent paradigms for the study of developmental gene regulation. The α locus consists of three genes (ζ, α1, and α2) that are dependent for expression on the major regulatory element (αMRE) which appears as an erythroid-specific DNase I-hypersensitive site ∼26-kb upstream of the ζ gene (Gourdon et al. 1995). The murine β locus consists of four functional genes [εy, βH1, β-major (βmaj), and β-minor (βmin)] that are controlled by the locus control region (LCR), a series of five DNase I-hypersensitive sites in erythroid chromatin located 5–25 kb upstream of the εy gene (Moon and Ley 1990; Hug et al. 1992; Jimenez et al. 1992; Fiering et al. 1995). The genes are arranged in the order of their developmental expression, as are their human homologs. Embryonic yolk sac-derived erythroid cells coexpress high levels of both ζ- and α-globin mRNA (Leder et al. 1992) and primarily εy and βH1 β-like globin mRNA with small amounts of βmaj and βmin (Brotherton et al. 1979; Chui et al. 1979; Wawrzyniak and Popp 1987; Whitelaw et al. 1990). At 11.5 days of gestation the major site of erythropoiesis in the developing embryo switches from the yolk sac to the fetal liver. This switch in site is coincident with a change to definitive gene expression in both the α and β clusters leading to predominant expression of the α1 and α2 genes and the βmaj and βmin genes. Although the small amount of βmaj and βmin expression in embryonic cells appears to be genuine and not caused by maternal contamination (Wawrzyniak and Popp 1987) it is unclear whether the embryonic genes are expessed in early fetal liver cells (Wong et al. 1983; Whitelaw et al. 1990).
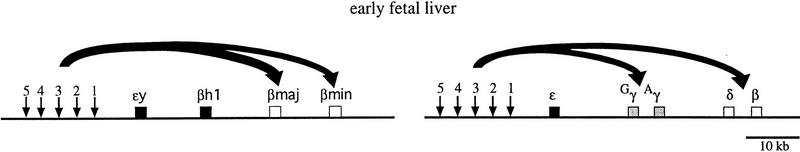
Figure 1.
Schematic diagram of the murine α-globin and β-globin loci. Genes are represented by solid boxes and vertical arrows represent DNase I hypersensitive sites of the αMRE and β-globin LCR.
The human globin loci have been studied more thoroughly, facilitated by the use of transgenic mice. The β LCR has been shown to be required for the initial activation of the locus and provides erythroid-specific, high-level, copy-number-dependent, position-independent expression to linked genes (Grosveld et al. 1987). Studies with the α locus have shown that HS-40 is required for expression of the α-like genes (Bernet et al. 1995) but transgenic results have suggested that additional sequences are required for developmental position-independent expression (Higgs et al. 1990; Sharpe et al. 1992; Gourdon et al. 1994). A detailed model of the mechanism of developmental regulation of the β-like genes has been proposed based on the observations of gene competition for LCR function (Enver et al. 1990; Hanscombe et al. 1991; Peterson and Stamatoyannopoulos 1993; Wijgerde et al. 1995, 1996; Dillon et al. 1997) and single gene activation by the LCR (Wijgerde et al. 1995; Gribnau et al. 1998). These data argue strongly in favor of a looping model in which the LCR forms direct, mutually exclusive chromatin interactions with gene-local regulatory elements to activate transcription of a single gene. Multiple genes are coexpressed in cis through alternating transcriptional periods of single genes (Wijgerde et al. 1995; Gribnau et al. 1998) suggesting that LCR–gene interactions are dynamic but also semistable, persisting on the order of several minutes (Wijgerde et al. 1995; Dillon et al. 1997).
It has therefore been suggested that two parameters determine the transcriptional output and hence expression level of a given gene during development. The first is the frequency with which the LCR productively contacts a particular gene and the second is the stability of that interaction. The frequency of LCR–gene contact has been proposed to be dependent on distance from the LCR (Dillon et al. 1997). The relative distance between two competing genes and the LCR has been shown to be important in controlling both the level and timing of expression (Enver et al. 1990; Hanscombe et al. 1991; Peterson and Stamatoyannopoulos 1993). Dillon et al. (1997) measured the effects of distance on the frequency of LCR–gene interactions by comparing genes of equal stability at varying positions in the locus in combination with primary transcript in situ hybridization. Insertion of a β gene into more LCR-proximal positions resulted in that gene being transcriptionally activated more often and at the expense of the equivalent downstream gene in relation to the difference in distance.
The stability of the LCR–gene interaction has been proposed to be determined by the transcription factor environment. Targeted disruption of the erythroid Kruppel-like factor (EKLF) has shown that it is required for transcription of only the adult-type β-globin genes (Nuez et al. 1995; Perkins et al. 1995). EKLF binds selectively with high affinity to the CCACCC element present in the promoters of the mouse and human adult-type β-globin genes (Donze et al. 1995). Studies with compound human globin locus transgenic/EKLF knockout mice have shown that reductions in EKLF expression in heterozygous and homozygous knockout mice lead to decreased expression of β-globin and reciprocally increased expression of γ-globin mRNA (Wijgerde et al. 1996; Perkins et al. 1996). We have shown that these changes are caused by reductions in the number of transcriptionally active β genes in the fetal liver population with reciprocally increased numbers of active γ genes (Wijgerde et al. 1996). These studies have been interpreted to suggest that reduced EKLF levels lead to a decrease in the stability of the LCR–β gene complex. Reduction in the amount of time that the LCR complexes with the β gene allows more frequent interaction with the γ genes.
Here we present detailed characterization of the developmental expression and transcriptional regulation of the murine α- and β-globin loci at the single-cell level. The results of in situ hybridization and novel kinetic analyses suggest that transcriptional regulation of the mouse β genes is mediated by a similar dynamic chromatin interaction mechanism as has been proposed for the human β locus. However, unlike the human locus transcription of the adult β-like genes occurs in embryonic cells and hence is only suppressed partially and not silenced through competition for the LCR. Similar analyses of the α-globin locus indicate that a coactivational mechanism of multiple genes in cis cannot be ruled out.
Results
Developmental expression
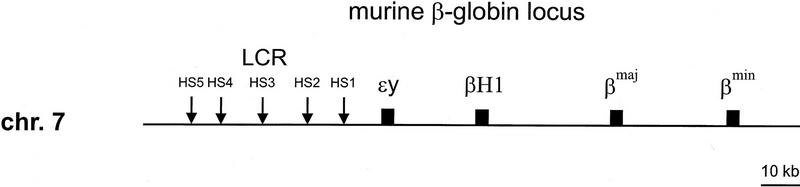
RNA samples were collected from embryonic and fetal erythroid tissues at different stages of development and subjected to S1 nuclease protection assay to determine the expression pattern for the murine α- and β-globin genes (Fig. 2). Previous studies have suggested that small amounts of the adult β-like globin genes are expressed in embryonic cells (Brotherton et al. 1979; Chui et al. 1979; Wawrzyniak and Popp 1987; Whitelaw et al. 1990). Quantitative PhosphorImager analysis of S1 assays (Fig. 2A) shows that in 10.5-day embryonic blood the level of εy is 60% and βH1 34% of total β-like globin (Table 1). βmaj–βmin (the β S1 probe does not distinguish between βmaj and βmin) is detected at levels of ∼6%. Expression of the α genes at 10.5 days is comprised of 46% ζ and 54% α mRNA (the α S1 probe does not distinguish between α1 and α2) (Fig. 2B; Table 1). This is in agreement with previously published results (Whitelaw et al. 1990). At day 11.5 of gestation the main site of erythropoiesis changes from the embryonic yolk sac to the fetal liver, which is the site of definitive erythropoiesis. Analysis of fetal liver RNA at 12.5 days shows that 91% of the β-like mRNA is βmaj–βmin, and εy and βH1 are still detectable at 7% and 2%, respectively. The appearance of small amounts of βH1 and especially εy mRNA in the early fetal liver as late as 14.5 days has been suggested to indicate that expression of these genes is not restricted to yolk sac-derived cells (Whitelaw et al. 1990). However, this residual mRNA may also be caused by contamination of fetal livers with circulating embryonic cells. A definitive conclusion is only possible through individual cell analyses with cell-type markers and morphological analysis (see below). The levels of α-globin mRNA in 12.5-day fetal liver cells also shows a switch from roughly equal ζ and α expression in the embryonic cells to 99% α and only 1% ζ mRNA. ζ mRNA is undetectable by 14.5 days, suggesting but not proving that ζ transcription is restricted to embryonic cells.
Figure 2.
S1 nuclease protection assays. Total RNA from the indicated developmental timepoints (10.5-day whole embryo and 12.5-, 14.5-, 16.5-day fetal liver and adult blood) was subjected to S1 nuclease protection assay as described in Materials and Methods with probes of equal specific activity. (A) β locus probes εy, βH1, and βm (βmaj and βmin). (B) α locus probes ζ and α (α1 and α2). The positions of protected fragments are indicated on the left.
Table 1.
Quantitative S1 nuclease protection analysis
|
|
10.5 blood
|
12.5 fetal liver
|
14.5 fetal liver
|
16.5 fetal liver
|
|---|---|---|---|---|
| εy | 60 | 7 | <1 | 0 |
| βH1 | 34 | 2 | <1 | 0 |
| βmaj–βmin | 6 | 91 | 99 | 100 |
| ζ | 46 | 1 | 0 | 0 |
| α1–α2 | 54 | 99 | 100 | 100 |
PhosphorImager quantitation of S1 nuclease protection assays shown in Fig. 2. The numbers represent the percentage of total β-globin or α-globin mRNA expression for the individual β and α genes.
Primary transcript in situ hybridization
We have shown previously that coexpression of the human γ- and β-globin genes in transgenic mouse fetal erythroid cells containing a single, complete human β-globin locus (Strouboulis et al. 1992) is achieved through alternating transcriptional periods of individual genes (Wijgerde et al. 1995). Primary transcript in situ hybridization with gene-specific intron probes showed that the vast majority of loci (88%) had only single gene signals. Approximately 12% of loci displayed two gene signals in cis, which we proposed represented a recent switch in gene activation. To ensure significance in this type of analysis the hybridization efficiency must be extremely high to guarantee the detection of nearly all transcriptionally active genes.
We determined the hybridization efficiency of the mouse ζ, α, εy, and βH1 globin intron probes by hybridizing them to 10.5-day primitive erythroid cells (Fig. 3A,B; and Table 2). The ζ probes detect primary transcript signals at 96% of the loci. The α-globin intron probes that recognize the primary transcripts of both highly homologous α1 and α2-globin genes demonstrate detectable signals at 89% of the loci (Table 2). Double-label experiments with both the ζ- and α-globin probes shows that a signal (α, ζ, or both) is detected at >95% of the loci. If all three α-globin genes are constitutively transcribed in primitive cells then our hybridization efficiencies are very high, at least >90%. It is possible that some of the genes are off at certain times; in this case we would conclude that our efficiency is even higher.
Figure 3.
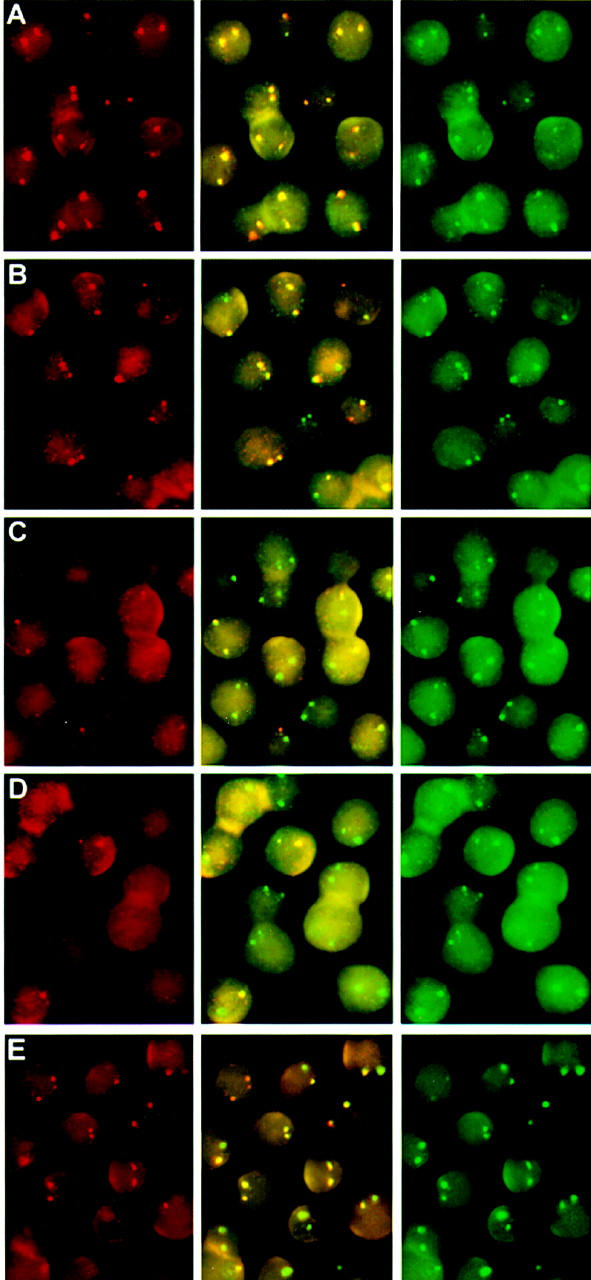
Primary transcript in situ hybridization on embryonic and fetal erythroid tissues. Double-label in situ hybridizations were performed with the indicated probe sets on 10.5-day blood (A–D) and 13.5-day fetal liver cells (E) as described in Material and Methods. Three separate images are shown for each hybridization; (left) red signal; (right) green signal; (middle) overlay of red and green. (A) ζ in red and α in green. (B) βH1 in red and εy in green. (C) βmaj in red and εy in green. (D) βmin in red and εy in green. (E) βmin in red and βmaj in green. The photos are composites of CCD images that represent a single focal plane and therefore may not show both loci from an individual cell in focus.
Table 2.
Quantitation of transcriptional cell types and percentages of loci with transcription signals for the individual globin genes
| Transcriptional cell types
|
Cells (%)
|
Loci with single, double, or not detected signals (%)
|
Transcriptional cell types
|
Cells (%)
|
Loci with single, double, or not detected signals (%)
|
Transcriptional cell types
|
Cells (%)
|
% loci with single,double, or not detected signals
|
||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| ○εy
|
βH1•
|
εy ○
|
βH1 •
|
εy/βH1
|
N.D.
|
•ζ
|
α○
|
ζ •
|
α ○
|
ζ/α
|
N.D.
|
○βmaj
|
βmin•
|
βmaj ○
|
βmin •
|
βmaj/βmin
|
N.D.
|
|||
| • | 1 | 0 | 0.5 | 0 | 0.5 | • | 0.4 | 0.2 | 0 | 0 | 0.2 | • | 1 | 0 | 0.5 | 0 | 0.5 | |||
| • | • | <1 | 0 | 1 | 0 | 0 | • | • | 3 | 3 | 0 | 0 | 0 | • | • | <1 | 0 | 1 | 0 | 0 |
| •* | 4 | 0 | 2 | 2 | 0 | •* | 13 | 6.5 | 0 | 6.5 | 0 | •* | 3 | 0 | 1.5 | 1.5 | 0 | |||
| 37 | 0 | 0 | 37 | 0 | 76 | 0 | 0 | 76 | 0 | 69 | 0 | 0 | 69 | 0 | ||||||
| ○* | 31 | 15.5 | 0 | 15.5 | 0 | ○* | 4 | 0 | 2 | 2 | 0 | ○* | 14 | 7 | 0 | 7 | 0 | |||
| • | ○* | 3 | 1.5 | 0 | 1.5 | 0 | • | ○* | 2 | 1 | 1 | 0 | 0 | • | ○* | <1 | 0.5 | 0.5 | 0 | 0 |
| 6 | 0 | 0 | 3 | 3 | 0.45 | 0 | 0 | 0.2 | 0.2 | 8 | 0 | 0 | 4 | 4 | ||||||
| ○ | ○ | 14 | 14 | 0 | 0 | 0 | ○ | ○ | 1 | 0 | 1 | 0 | 0 | ○ | ○ | 2 | 2 | 0 | 0 | 0 |
| ○ | 3 | 1.5 | 0 | 0 | 1.5 | ○ | 0.15 | 0 | 0.1 | 0 | 0.1 | ○ | 2 | 1 | 0 | 0 | 1 | |||
| Total | 31.5 | 3.5 | 59 | 5 | Total | 11 | 4 | 85 | 0.5 | Total | 10.5 | 3 | 81.5 | 5.5 | ||||||
| total % ɛy signals | 31.5 + | 0 + | 59 + | 0 = 91 | total % α signals | 0 + | 4 + | 85 + | 0 = 89 | total % βmaj signals | 10.5 + | 0 + | 81.5 + | 0 = 92 | ||||||
| total % βH1 signals | 0 + | 3.5 + | 59 + | 0 = 63 | total % ζ signals | 11 + | 0 + | 85 + | 0 = 96 | total % βmin signals | 0 + | 3 + | 81.5 + | 0 = 85 | ||||||
Double label in situ hybridizations as shown in Fig. 3 were counted and the percentages of the nine different transcriptional cell types are shown. (○, •) The different primary transcript signals for the individual globin genes as indicated at the top of each column. The signals present on each of the homologous chromosomes are shown. (Overlapping ○, and • ). Loci with two different primary transcripts signals in cis. The percentages of individual loci with single, double or not detectable (N.D.) in situ signals contained within these cells are shown in the four columns on the right. The totals of these columns (single, double, or not detected) are shown directly beneath them. The appropriate values were summed up to arrive at the total percent of the specific gene signals (bottom two lines of each table). Results were taken from 10.5-day embryonic blood cells for εy, βH1, ζ, and α. βmaj and βmin results are from 13.5-day fetal liver cells. Shown are the percentages of loci with transcription signals for the indicated genes.
Cells with different transcription signals or combinations of signals on each locus.
We performed similar quantitations for the εy and βH1 globin primary transcript probes (Fig. 4B, Table 2). Approximately 91% of the loci have an εy signal and 63% have a βH1 signal at 10.5 days. Calculation of the relative percent of εy versus βH1 primary transcript signals (59% and 40%, respectively) yields comparable percentages to the relative amount of mRNA as determined by S1 analysis (60% and 34% of total β-like globin, respectively; Table 1). This close correlation between the relative percentages of transcriptionally active genes and mRNA expression suggests that the number of fully active genes in the population determines the level of mRNA expression. This was also found to be the case in the human β locus in which the results indicated that a gene is either fully on or off (Wijgerde et al. 1995, 1996; Dillon et al. 1997).
Figure 4.
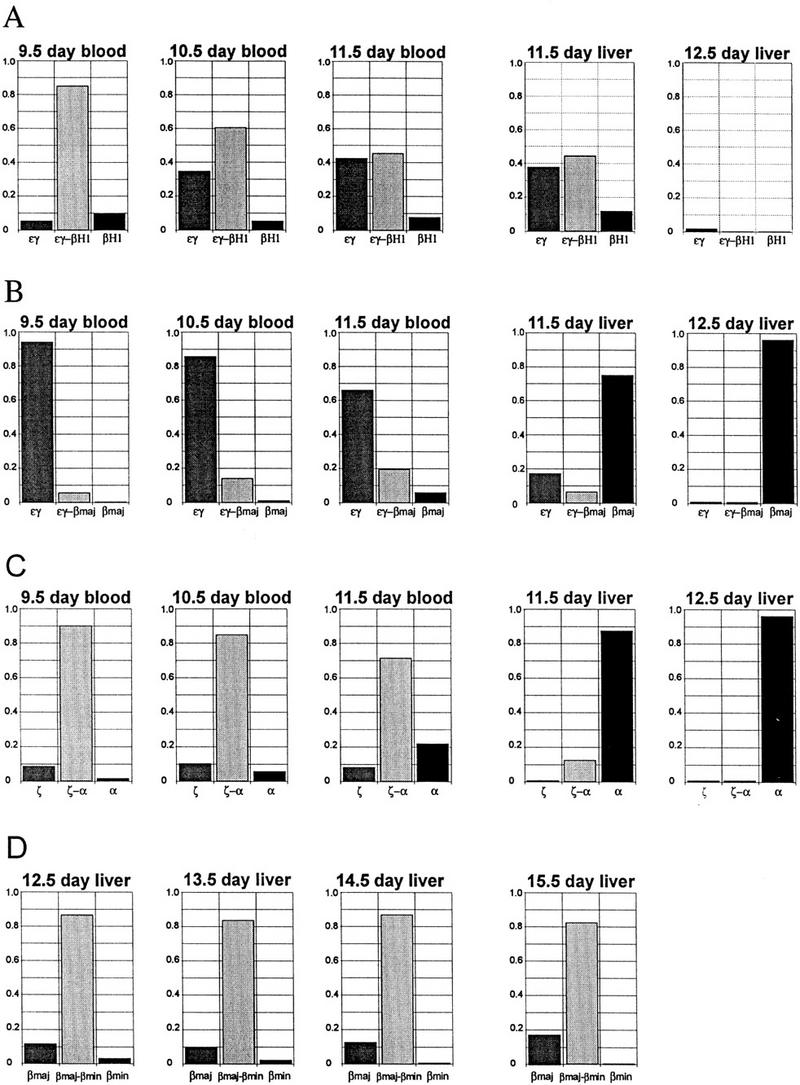
Quantitation of loci with single or double in situ primary transcript signals during embryonic and fetal development for the indicated genes. Double-label in situ hybridizations as shown in Fig. 3 were quantitated and plotted as the percentage of loci with single or double signals at the timepoints indicated. (A) εy vs. βH1. (B) εy vs. βmaj. (C) ζ vs. α. (D) βmaj vs. βmin. The values represent the percent of individual loci with detectable single or double signals and therefore do not include the small percentage of loci with no detectable signals (see Table 2).
Globin gene transcription in primitive erythroid cells
Gene competition in the mouse β-globin locus has been suggested by a naturally occurring deletion of the βmaj gene and its gene-local regulatory elements (Skow et al. 1983) which leads to increased expression of the βmin gene in homozygotes (Curcio et al. 1986). In contrast, targeted disruption of the βmaj gene via insertion of a selectable marker leads to perinatal lethality with no increase in βmin expression (Shehee et al. 1993;). In the α locus, marker gene insertion into the ζ gene leads to reductions in the level of α gene expression in the fetal and adult erythroid cells suggesting that the marker gene may be competing for activation with the α genes (Leder et al. 1997).
To investigate the mechanism of coexpression in the mouse α- and β-globin loci and characterize the pattern of transcriptional regulation we performed primary transcript in situ hybridization, on 9.5-, 10.5-, and 11.5-day peripheral blood. Oligonucleotide probes specific for the introns of εy, βH1, βmaj, βmin, ζ, and α were used in double-label experiments. Primary transcript in situ hybridization shows that embryonic blood cells from days 9.5–11.5 transcribe all four β-like globin genes to varying degrees (Fig. 4A–C; Table 2). Each of the loci in a single cell can have a different primary transcript signal or combination of signals (Table 2) demonstrating that, like the human β genes, each locus responds independently to the same trans-acting factor environment. Almost all loci (91%; Table 2) have signals for the εy gene, which remains fairly constant in embryonic blood cells throughout the period from 9.5 to 11.5 days (Fig. 4A). The εy gene is closest to the LCR and is the most highly expressed globin gene during embryonic erythropoiesis (Fig. 2A). The εy and βH1 foci appear frequently in cis (i.e., double signals on one locus) decreasing from 85% at 9.5 days to 45% at 11.5 days of gestation (Figs. 3B; 4A) as a result of the decrease in βH1 gene transcription from 95% at 9.5 days to 53% at 11.5 days. Thus, fewer βH1 genes are being transcribed as development proceeds.
Adult β gene transcription is observed in a small percentage of the βmaj genes in 9.5-day embryonic blood, increasing to 25% in 11.5-day blood (Figs. 3C; 4B). Three percent of the βmin genes had primary transcript signals at 10.5 days (Fig. 3D) and increased to ∼10% in 11.5-day blood. The βmaj and βmin foci nearly always appear in cis with an εy signal in the early primitive cells, but by 11.5 days (blood) ∼20% of the βmaj signals are single signals (Fig. 4B). These results of β gene transcription in the mouse globin locus are markedly different from those of the human transgene locus. The early transcription of the adult mouse β genes (βmaj and βmin) is in contrast to the human locus in which transcription of the adult β-globin gene is not detected in embryonic cells but is restricted to fetal-derived erythroid cells. These results show that the entire murine β-globin locus is activatable in embryonic erythroid cells.
In situ transcription analysis of the ζ and α genes in embryonic cells demonstrates that both genes are transcriptionally active as expected but also reveals a high percentage of double signals in cis starting at 90% in 9.5-day red cells and decreasing to ∼70% at 11.5-days (Fig. 4C). The percentage of α genes with a signal remains fairly constant ∼90% throughout the embryonic period but the percentage of ζ gene signals decreases from 99% of all ζ genes at 9.5 days to 80% at 11.5 days (blood), resulting in fewer double signals and more single α gene signals. As in the β locus the pattern of transcriptional activity of the genes in the α locus does not remain static from 9.5 to 11.5 days.
εy, βH1, and ζ gene transcription is restricted to primitive erythroid cells
The same probes were used to analyze gene transcription in fetal liver erythroid cells at 11.5 and 12.5 days of gestation (Fig. 4A–C). Livers were isolated from fetal mice at the time points indicated and distrupted gently and prepared for in situ hybridization as described in Materials and Methods. In situ analysis of 11.5- and 12.5-day fetal liver cells shows that the βmaj and βmin genes are now the most highly expressed genes, which is in agreement with the S1 analysis (Fig. 2A). At 11.5 days there is a small percentage of erythroid cells that continue to transcribe the εy, βH1, and ζ genes in the fetal liver preparations (Fig. 4A–C). It varies from one preparation to the next (average 15%) but by 12.5 days it is reduced to near zero. In situ analysis in 12.5-day peripheral blood in which >75% of the cells are nucleated primitive erythrocytes shows that transcription of the globin genes is switched off dramatically (not shown). The near complete disappearance of εy, βH1, and ζ gene signals in the 12.5-day fetal liver preparations (Fig. 4A–C) suggested that those signals in the 11.5-day fetal liver preparations arose from contaminating embryonic erythrocytes. In addition the cells that display foci for the εy, βH1, and ζ genes in the 11.5-day fetal liver slides are distinct microscopically from the bulk of fetal liver cells as they display a high degree of autofluoresence, a characteristic of embryonic cells.
We confirmed that these cells were indeed embryonic and not fetal derived by three separate experiments. Hybridization of 11.5-day fetal liver slides with probes for εy, ζ, and βmaj show that a small percentage of autofluorescent erythroid cells (∼15%) transcribe both the εy and βH1 genes indicating that their expression is restricted to a subpopulation of cells in the fetal liver preparations (Fig. 5A–C). Transcription signals for mouse ζ and εy in cells from 11.5-day fetal-liver preparations from a homozygous transgenic line that contains a single copy of the human β-globin locus, were completely separate from the cells that had transcription signals for the fetal-restricted human β gene (Wijgerde et al. 1995) (not shown). Finally, in situ analysis of 11.5-day blood and fetal liver cells from an EKLF−/− mouse with εy and βmaj probes showed that the lack of βmaj transcription because of the EKLF knockout (Nuez et al. 1995; Perkins et al. 1995) did not lead to the appearance of εy transcription foci in the fetal liver cells. This indicates that the εy gene is silenced and not in competition with the βmaj gene in fetal cells (data not shown). Interestingly, βmaj transcription foci were not present in the EKLF−/− embryonic blood cells indicating that transcription of the βmaj gene is dependent on EKLF activity in primitive cells. We conclude from these experiments that the cells with mouse ζ and/or εy, and by inference those with βH1 signals, are embryonically derived erythroid cells and that transcription of these genes is restricted to the primitive lineage and does not occur in true definitive cells.
Figure 5.
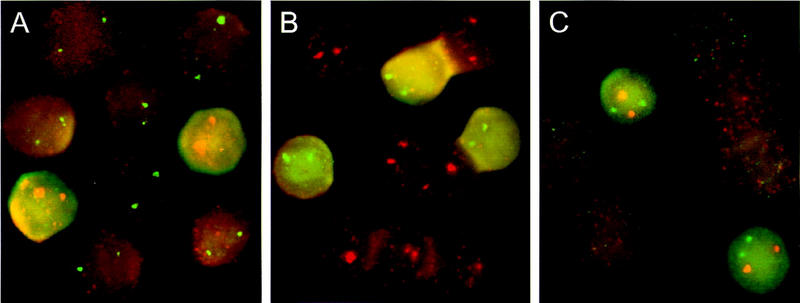
Primary transcript in situ analysis of 11.5-day fetal liver preparations. Cells were hybridized with (A) ζ in red and βmaj in green; (B) εy in green and βmaj in red; (C) ζ in red and εy in green. Note the relatively high autofluorescence in cells transcribing εy and/or ζ genes compared to the cells transcribing the βmaj gene.
Globin gene transcription in definitive erythroid cells
Presumably the mouse ζ, εy, and βH1 genes are autonomously silenced during erythroid development as are the human ε (Raich et al. 1990) and ζ genes (Liebhaber et al. 1996). Definitive cells derived from the fetal liver express only the α1 and α2 genes from the α locus and the βmaj and βmin genes from the β locus. As mentioned previously the α1 and α2 genes are highly homologous even in their intervening sequences precluding the use of gene-specific intron probes to distinguish them.
The βmaj and βmin intron sequences are divergent and gene-specific probes were used to detect primary transcripts from these genes separately in double-label in situ hybridizations (Fig. 3E). Quantitation of both βmaj and βmin signals together shows that signals are detected at 95% of the loci in 13.5-day fetal liver indicating that the probe efficiency is very high (Table 2). Separate quantitation shows that 92% of the βmaj alleles and 85% of the βmin alleles have primary transcript signals (Table 2). Approximately 81% of the loci have βmaj and βmin signals in cis (double signals), 10% have βmaj alone and 3% have βmin alone (Table 2). A cell-by-cell analysis shows that ∼17% of the cells transcribe a different combination of genes, whereas the trans-acting factor environment is the same (Table 2; indicated by *). The results from fetal liver cells from 12.5–15.5 days show that the percentage of βmin transcription foci is declining during development (Fig. 4D), which fits well with the previously reported changes in levels of β-maj and β-min proteins during fetal development (Whitney 1977; Alter and Goff 1980; Wawrzyniak and Popp 1987).
Coactivation or alternating single-gene activation
The relatively high percentage of double signals in the mouse α and β loci suggested that the mechanism of multigene expression in these loci may be different from that proposed for the human β locus. In the human β locus the LCR is thought to flip-flop between genes to alternately activate transcription. Primary transcript in situ hybridization analysis of γ and β transcription in the early fetal liver cells showed that 85% of the loci have single-gene signals (Wijgerde et al. 1995). The fact that nearly all cells had human γ and β mRNA in their cytoplasm indicated that alternation must occur and suggested that the small amount of double signals were caused by a recent switch from γ to β or vice versa. The results from the mouse α and β loci could be interpreted to indicate that transcription is coinitiated from multiple globin genes in cis in most cells. However, there are two indications from the data that contest this conclusion in the case of the β locus. First, a significant proportion of the cells have loci that are responding differently to the same trans-acting factor environment (38% at 10.5 days and 17% at 13.5 days, Table 2). Second, there is a significant proportion of loci with only single-gene signals (35% at 10.5 days and 14% at 13.5 days, Table 2). These results suggest the possibility that the individual genes in the mouse β locus may be alternating, albeit more often than in the human β locus (Wijgerde et al. 1995).
We therefore designed an experiment to provide further evidence of coinitiation or alternating transcription of multiple genes in the mouse globin loci making use of the inhibitor of transcriptional elongation DRB (5,6-dichloro-1-β-d-ribofuranosyl-benzimidazole). Previous studies have shown that DRB does not affect initiation of transcription (Fraser et al. 1978; Marshall and Price 1992) but prematurely aborts elongating transcripts 400–600 bp from the initiation site (Chodosh et al. 1989; Marshall et al. 1996) by inhibiting the activity of the P-TEFb kinase, which phosphorylates the carboxy-terminal domain (CTD) of Pol II (Marshall et al. 1996). The effect of DRB treatment on globin gene transcription in mouse erythroleukemia cells has been reported previously (Tweeten and Molloy 1981). The results demonstrate that DRB causes premature termination without affecting initiation of transcription. This was tested in transgenic mice by briefly exposing fetal liver cells to DRB and analyzing initiation using promoter-proximal probes and elongation using probes to distal regions of the human γ- and β-globin primary transcripts via in situ hybridization (Gribnau et al. 1998). In situ signals with probes that hybridize to intron 1, located in the first 300 bases of the primary transcript are still visible after 15 min of DRB treatment, whereas probes that hybridize to intron 2, 600–1200 bases 3′ of the initiation site are no longer visible after 7 min of DRB treatment.
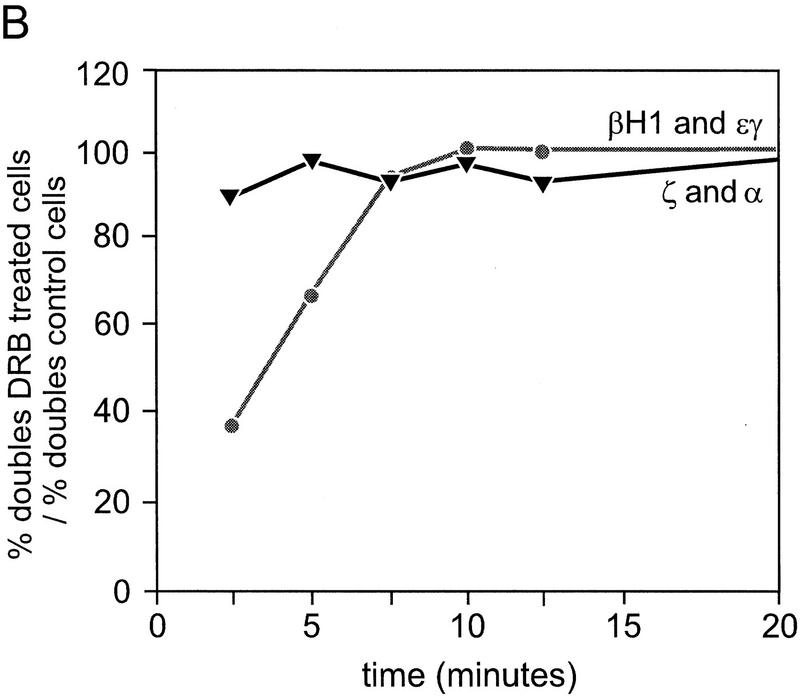
What makes DRB inhibition of elongation particularly useful is the fact that it is reversible, allowing elongation to proceed normally after removal of DRB. We tested this in day 10.5 embryonic blood cells by first treating cells with DRB for 15 min and then measured the reappearance of transcription elongation at intervals after washing out the DRB (Fig. 6). As a measure of recovery of the cells we recorded the percentage of cells in which both homologs had reappearing signals, using probes to intron 2 of the mouse εy and βH1 primary transcripts. The results show that elongation signals reappear on both homologs in 67% of the cells, 2.5 min after washing out the DRB (Fig 6I; trans curve). This result illustrates two very important points. First, that elongation is restored very rapidly after removal of DRB and second, that the vast majority of cells that will recover already have elongation signals on both homologs. This shows that the elongation signals are not reappearing randomly throughout the available loci in the population of cells, but instead are contained in a subset of recovering cells in the early time points. The late time points show that the percentage of cells with trans elongation signals (signals on both homologs) are equal to untreated control values (82%), indicating that the cells are fully recovered and not adversely affected by the brief drug treatment.
Figure 6.
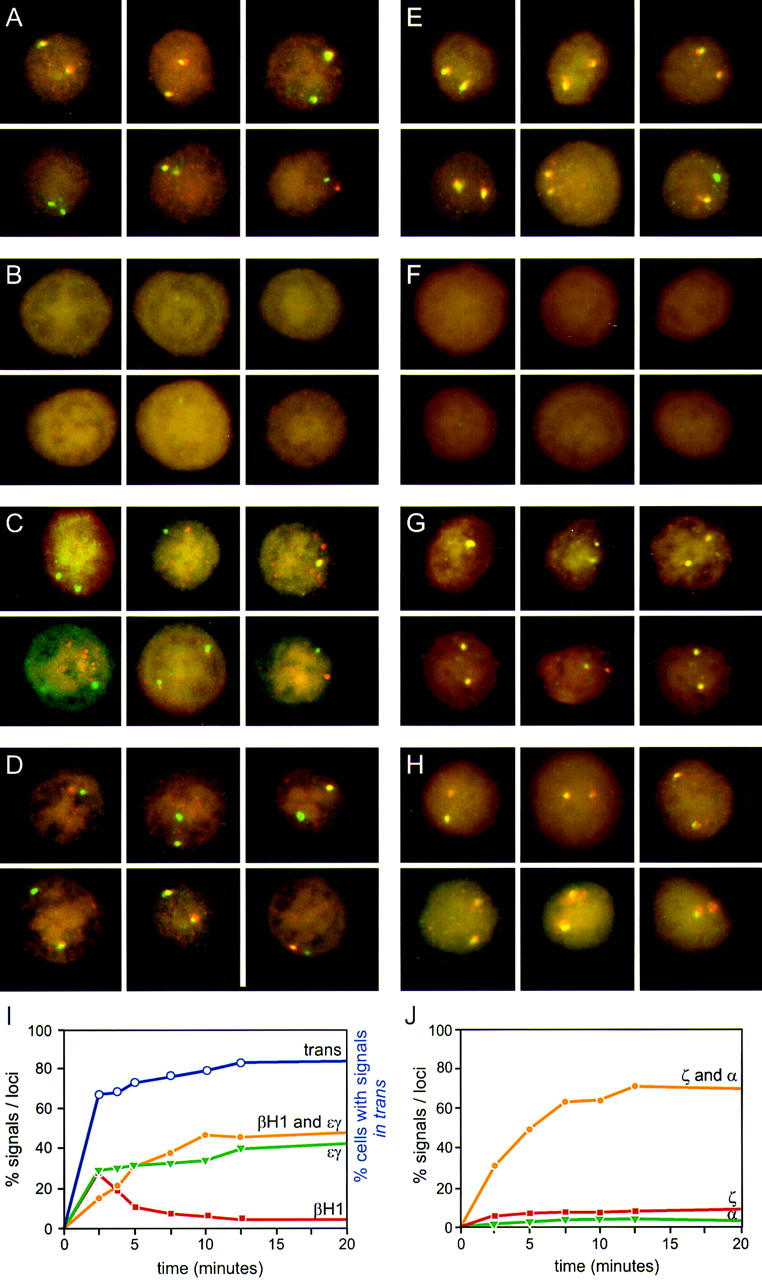
Kinetic analysis of reappearance of primary transcript signals after DRB inhibition of elongation. Blood cells from 10.5-day embryos were pulse treated with DRB as described in Materials and Methods and hybridized with mouse εy and βH1 primary transcript probes (A–D), and mouse ζ and α globin primary transcript probes (E–H). (A,E) Prior to DRB treatment. (B,F) Cells treated with DRB for 15 min. (C,G) 5 min after washing out DRB. (D,H) 20 min after washing out DRB. The percent of single and double signals are presented from the indicated timepoints in I (εy and βH1) and J (ζ and α). Also shown is the percentage of cells with signals on both loci (trans curve; I) which is a measure of the recovery of cells after DRB treatment. All values represent the percentages of total loci or cells, respectively. The data represent the results of a single experiment was repeated four times in the case of β and three times in the case of α, yielding essentially the same curve shapes. Some variability is observed in the position of the curves along the x-axis, because of the inherent difficulties in obtaining short time points.
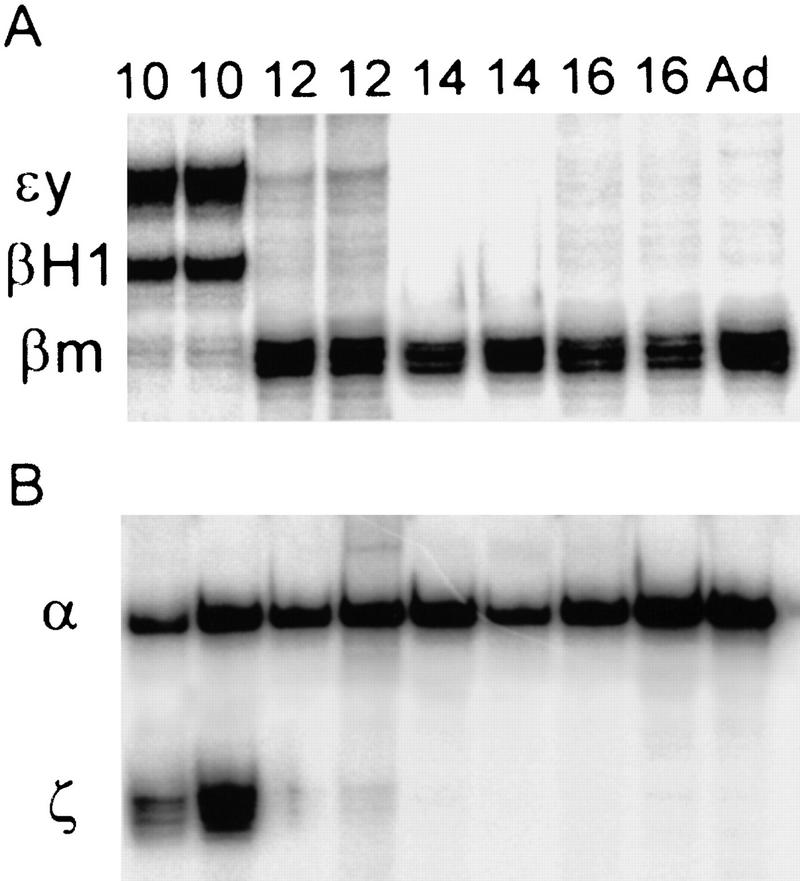
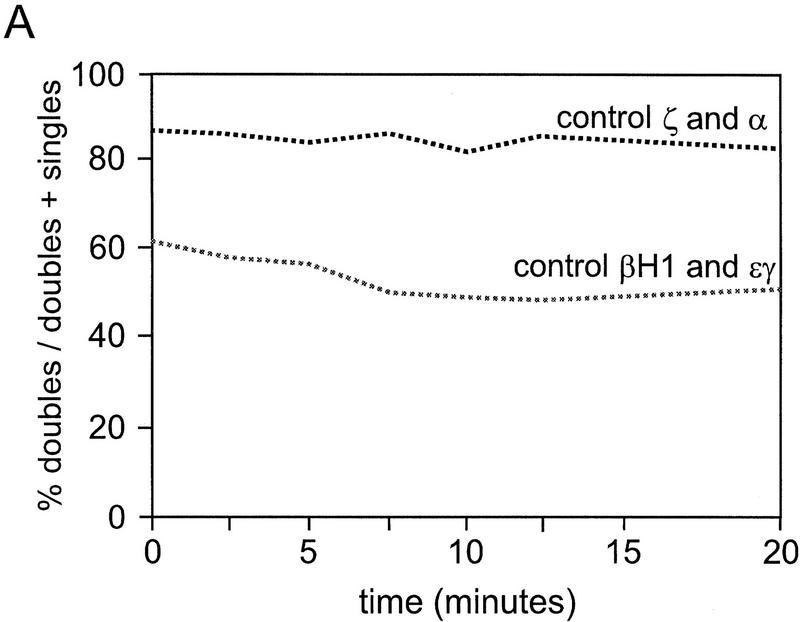
We reasoned that if coinitiation in cis was occurring in loci that have double signals, then restoration of elongation after DRB treatment should result in the simultaneous reappearance of the same relative percentages of single- and double-gene signals. If alternation of transcription is the cause of the double signals, then single signals should reappear first and be followed by the reappearance of double signals after a lag that reflects the extra time required to produce a second signal in cis. Reappearing elongation signals were quantitated for ζ- and α-globin and εy and βH1 globin genes at the indicated time points. The cells in the untreated panels (Fig. 6A,E) show the normal distribution of single versus double signals for the εy and βH1 genes and the ζ and α genes, respectively. After 15 min of DRB treatment no primary transcript signals are detectable (Fig. 6B,F). Figure 6, C and G, shows an early time point after removal of the DRB block (5 min after wash). Note that the majority of signals in the case of the β genes are single-gene signals per locus, whereas for the α genes, most signals are double signals. A graphical analysis of the quantitation of loci with single- or double-gene signals as a percentage of all loci for the β and α genes are shown in Figure 6, I and J, respectively. In the case of the β genes, a higher than normal proportion of single signals are present in the early time points, whereas the double signals are underrepresented. As time progresses, the single signals are converted to double signals (most apparent from the βH1 curve). The results for the α genes are strikingly different. The double and single signals reappear at approximately the same rate and with the same relative proportions as in control cells. The percentage of double signals does not significantly change in untreated control cells during the time course of the experiment (Fig. 7A). Comparison of the reappearance of double signals for the β and α genes by plotting the percentage of double signals/total signals, for DRB-treated cells normalized to control values shows that the double β signals are much slower to reappear when compared to the double α signals (Fig. 7B).
Figure 7.
Percentage of double signals in control cells and reappearance of double signals in DRB-treated cells. (A) Double signals in untreated control cells are plotted as a percent of total signals for the α genes (ζ and α) and the β genes (εy and βH1). (B) The reappearance of double signals in the DRB-treated cells are plotted as a percent of the control values in A.
The fact that the relative amount of double α versus single α or ζ signals is the same as in the untreated cells, regardless of the number of cells that have recovered, suggests that a coinitiation mechanism could explain multigene expression in the mouse α locus. However, a transcriptional mechanism that involves very frequent alternations (i.e., alternations occurring every minute) would give the same result. The results support a mechanism of alternating transcription in the mouse β-globin locus that is similar to that proposed for the human β-globin locus (Wijgerde et al. 1995).
Discussion
We have used a combination of S1 nuclease protection assays on total RNA and primary transcript in situ hybridization in erythroid cells to analyze the gene expression and transcription patterns of the murine α and β globin genes. The results show that mRNAs from the embryonic εy, βH1, and ζ-globin genes are expressed at high levels in primitive erythroid cells and are still detectable at low levels in the early fetal liver until 12.5–14.5 days of gestation. Primary transcript in situ hybridization indicates that the small amount of residual expression of these genes in early fetal liver preparations is caused by the presence of a subpopulation of cells that transcribe both the εy and ζ genes. These cells display a high degree of autofluoresence and do not transcribe the fetal-restricted human β gene indicating that they are circulating embryonic cells and not true definitive cells derived from the fetal liver. We conclude from this data that transcription of the εy, βH1, and ζ-globin genes is restricted to the primitive erythroid lineage.
In contrast, the α1, α2, and to a lesser extent the βmaj and βmin genes are expressed in the primitive lineage and are the only globin genes expressed in definitive cells. α1 and α2 mRNA makes up ∼50% of the embryonic α globin at 10.5 days, whereas βmaj and βmin expression is ∼6%. In the case of the β locus where all four genes are distinguishable, the results show that they are transcribed at frequencies that are inversely proportional to their distances from the LCR. These results are reminiscent of those of Dillon et al. (1997) in which a marked β gene was placed at different positions in the human globin locus and the effect of distance on transcription of the introduced gene and the other globin genes was measured. The results showed that altering the distance of a gene from the LCR had an effect on the frequency of transcription of that gene. An LCR-proximal gene was transcribed more often than a distal gene with reciprocal consequences for the remaining downstream genes in the locus. Those results suggested that the LCR activates gene transcription by forming direct chromatin interactions with gene-local regulatory elements and that the frequency of contact between two or more competing genes and the LCR is controlled in part by differences in distance.
The results highlight an important difference in the pattern of developmental regulation between the mouse and human β loci. In the mouse locus βmaj and βmin gene transcription is activated in the primitive lineage, whereas human β gene transcription is restricted to definitive erythroid cells in transgenics. The human β gene is thought to be silenced in primitive cells in part through competition for the LCR by the more LCR-proximal ε and γ genes. A competition mechanism could also be operating in the mouse β-globin locus, which is suggested by the DRB experiments, and could account for the observed inverse correlation between percentage of transcriptionally active genes and distance from the LCR in primitive cells. One might expect from these results that if the human β gene in its distal location in the locus is activatable in primitive erythroid cells then some transcription should be detected in situ. Clearly the human β gene is activatable in primitive cells when placed next to the LCR (Enver et al. 1990; Hanscombe et al. 1991) or in the position of the ε gene (Dillon et al. 1997) but not in its wild-type location (Wijgerde et al. 1995) or when placed just 5′ of the δ globin gene (Dillon et al. 1997). These results suggest the possibility that the distal part of the human locus that contains the δ and β genes is not accessible to LCR activation in primitive cells. This concept is supported by results from multicopy transgenic mice that contain head-to-tail tandemly integrated copies of a complete 70-kb human globin locus in which the β gene is juxtaposed on the 3′ side by an LCR (Milot et al. 1996). These mice do not express β mRNA in the primitive erythroid cells even though the β gene would be closer to the downstream LCR than the γ genes. It is important to note here that the LCR can activate the β gene in a 5-kb fragment when it is placed 5′ of HS5 (Zafarana et al. 1995) i.e., it works in both orientations. Taken together with the results presented in this paper that indicate that the βmaj and βmin genes are partially suppressed but not silenced by gene competition in primitive cells, it suggests that the human β gene is silenced in primitive cells through another mechanism that may involve epigenetic chromatin modification (Fig. 8). It has been proposed that the human β locus may be divided into distinct chromatin subdomains (see Collins and Weissman 1984 and references therein). This idea gains support from experiments with somatic cell hybrids between primitive transgenic erythroid cells and MEL cells that show that γ gene expression is retained through several hybrid cell divisions before eventually switching to β expression (Papayannopoulou et al. 1986; Stanworth et al. 1995; see also Fraser et al. 1998). The above discussion is not intended to suggest that competition plays no role in transcriptional regulation of the β-globin genes. Competition is clearly operating in the human locus when multiple genes (or domains) are activatable, such as in primitive cells between ε and γ and in early definitive cells between γ and β.
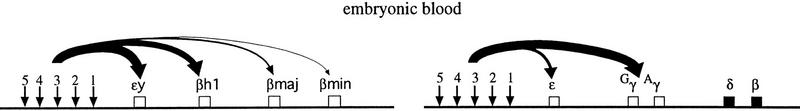
Figure 8.
Models of developmental switching in the mouse and human β globin loci. Schematic diagrams of the mouse (left) and human (right) β globin loci are shown during embryonic (top) and early fetal (bottom) erythropoiesis. Vertical arrows denote DNase I hypersensitive sites of the LCRs. The curved arrows signify an interaction between the LCR and an individual globin gene. The relative thickness, of the arrows represents the percent of transcriptionally active genes in the population of erythroid cells. Open boxes represent activatable genes. Closed boxes represent genes that are silenced or in an inaccessible chromatin conformation (or both). Gray boxes (γ genes, fetal liver) reflect the possibility that the γ genes in some early fetal liver cells may be silenced (i.e., cells committed to β transcription; Wijgerde et al. 1996).
The DRB experiments presented here suggest that alternating transcriptional activation of the mouse β genes is occurring, which may be indicative of gene competition for the LCR. Thus the results presented can be explained easily by the direct chromatin interaction mechanism between the LCR and an individual β gene that has been proposed for the human β locus. Alternative explanations would require the postulation of a number of other parameters. We propose a model in which the entire mouse β locus is accessible to LCR activation in primitive cells and that the individual genes are transcribed at frequencies that are inversely proportional to their distance from the LCR caused largely by gene competition (Fig. 8). In definitive cells the εy and βH1 genes are no longer activatable because of silencing that may involve chromatin modification allowing the adult genes to be transcribed more frequently. Thus we propose that switching in the mouse locus involves shutting off the embryonic genes. In contrast, the switch to definitive erythropoiesis in the human transgene β locus involves ‘opening up’ of the adult δ–β domain through chromatin modification, making it accessible to activation by the LCR in competition with the γ genes.
The DRB results with the mouse α locus are more difficult to interpret but are clearly different from those of the β locus. The results of targeted disruption of the ζ and α1 genes by insertion of a PGK–Neo cassette (Leder et al. 1997) are strongly indicative of gene competition for the αMRE. Insertion into the ζ gene had a more profound effect on α1 and α2 gene expression than did insertion into the α1 gene, clearly indicating that relative position of the PGK–Neo cassette with respect to the αMRE is important. It has been firmly established that insertion of an activatable promoter (or gene) between a regulatory element and its normal target promoter leads to decreased activation of the downstream gene (Kim et al. 1992; Fiering et al. 1995; Hug et al. 1996; Pham et al. 1996; Dillon et al. 1997) indicating that gene order and/or relative distance from the regulatory element are key parameters in determining the competitive ability of a gene. We cannot conclude that the mouse α genes are alternately transcribed or coinitiated from our results, however the accumulated data suggest that they may be alternating frequently in a competitive mechanism for interaction with the αMRE. Obviously experiments that are able to show a direct interaction between the LCR or αMRE and an individual gene will allow firm conclusions regarding this type of mechanism.
Materials and methods
Preparation of RNA and S1 nuclease protection assay
RNA was prepared from 10.5-day embryos, 12.5- and 14.5-day fetal liver, and adult blood and subjected to S1 nuclease assay as described previously (Weaver and Weissman 1979). The εy, βH1, and adult β probes used were those described by Lindenbaum and Grosveld (1990). The mouse α and ζ probes were a 300-bp BamHI fragment from plasmid GSE 1454 and a 300-bp ApaI–AvaII fragment subcloned from cosmid cML1 (Kielman et al. 1994), respectively. S1 nuclease-protected bands were quantitated with a PhosphorImager (Molecular Dynamics).
Primary transcript in situ hybridization
Primary transcript in situ hybridization to detect transcriptional activity of the mouse β-globin genes in 9.5-, 10.5-, and 11.5- blood; and 11.5-, 12.5-, and 13.5-day fetal liver cells was performed as described by Wijgerde et al. (1995). Embryonic blood cells were collected in PBS, fetal liver cells were disrupted in PBS (by pipetting up and down several times) and spotted on poly-l-lysine coated slides. Slides were placed in fixative (4% formaldehyde, 5% acetic acid in saline) for 20 min at room temperature then washed three times in PBS at room temperature and stored in 70% ethanol at −20°C. Slides were pretreated for hybridization by rinsing in Tris/saline (0.1m Tris at pH 7 and 0.85% saline) and incubated in 0.01% pepsin, and 10 mm HCl at 37°C for 5 min. The slides were rinsed briefly in H2O and fixed again for 5 min in 3.7% formaldehyde in PBS at room temperature. After a PBS wash slides were dehydrated in ethanol (70%, 90%, and 100%, respectively, 3 min each). Slides were hybridized in 25% formamide, 2×SSC, 1 mm EDTA, 5× Denhardt’s, 50 mm NaH2PO4/NaHPO4 (pH 7.0) and 200 ng of sheared salmon sperm DNA using 1–5 ng/μl of the appropriate oligonucleotide probes at 37°C overnight. After hybridization slides were washed in PBS for 30 min at 37°C and rinsed briefly in TST (Tris/saline with 0.05% TWEEN 20). The slides were then incubated in Tris/saline containing 1% blocking reagent (TSB) (Boehringer) for 30 min at room temperature. The slides were incubated subsequently with TSB containing the appropriate primary antibody (or Avidin D Texas red for biotinylated oligos) for 30 min at room temperature and washed two times for 5 min in TST. The previous steps were repeated with fluorescently labeled secondary (biotinylated goat anti-avidin D for biotinylated oligos) and tertiary antibodies. After the final washing step the slides were dehydrated (70%, 90%, and 100% ethanol), air-dried, and mounted witha 1:1 mixture of 1% DABCO, 0.4 μm DAPI (Sigma), 90% glycerol, 0.02% sodium azide, 10 mm Tris (pH 8.0) with Vectashield (Vector labs).
Quantitation of primary transcript signals was done by counting at least 400 cells for each timepoint using an epiflourescence microscope (Leitz). The figures presented were created with a CCD camera (Hamamatzu).
Oligonucleotide probes
Antisense oligonucleotide probes that recognize the intronic sequences of the respective globin gene primary transcripts were labeled with digoxygenin, dinitrophenol, or biotin haptens at both ends and in the middle as indicated. εy 1–4 (digoxygenin): CTCAGAATTCTTGATTTCCCTAGCTCTTTGTACCTAAAAAACAATTCTTCAGCCATTCACTGTCACCCTTACTGGGACCAATTAATTAACTTTGACAGCACTCTCTTCATATTACTCTCCATATAAATCCATGATAAATTTTATCACGGTTGTCTTGCAAGACTTTTCTTCAACATCAATAAATAGGACCGCGCAAAA. βH1 1–4 (biotin): CAAAACCCT-ATAGAAACCCTGGAAATTTCTGCCATGCATAAGGATAATTTTGGACCCATGGACTCTAACATCTGTCAAGGCATTGCCAATCACAGTCTCAAAATGCTGGGCGCTCACTCAAATCTGCACCCAAATCATTGTTGCCCACAAGCATAGATGTATTAATTTATAAAAACATACTCCTTTTTAAAAAAGATCCA. βmaj 2–4 (dinitrophenol): GAACTCTTGTCAACACTCCACACACAGTCATGGAGACTGCTCCCTAGAATATGGGAAGTAAATAACCAGAGCTTAATTAATTTAGTAAAATGCAACTGGAGACAAATTATTATAAGAATCCTATGTCAAACAGAATTTATATGTAAAATA. βmin 1–4 (biotin): TATGAAGTAGAGCAACAATACAAGATGCTGAAGGCCGATTTCAAATGGAAACTGTGGAAAGGATCAGAGAATCATTTATCTTTTTGTCCTCAGAGTAAGAAAAAAAGAATTATTCTATGACACACAAAATTTAGCCACAAAATATACTCTGGTAAAATGGCAGCTGGGTTCTACTGGTCAATTTTGATAAGAATTATTCT. ζ1–5 (biotin): CCTTCTCAGTGGCTTCTCCTCACTAACTGCTCTTTGTCACTTCTGTTCATGGAAGACTCTGGTGAGCTCTGTGAATGCCAGCCCACCTCCTTTAGTAACAACCCCAAGAGTGATGTTACTATTTGCTGTTGCACAAGGGTCTACAAAGGGGATTTGATGCCTCCAGCCTCCAATGGCACCCATGCCTGCGCTCG. α1–3(di-goxygenin): TCACAGAAAAGCATAGTTAGAAGCTGCCCACTGAGCGAGTGCCAGGTCCATTAGCCCTTCCTAGGGGT-CCCAGATGCCGCCTGCCAGGTCCCTGCTCCCCTTCCTGGGACCACTATGTTCCCTGCCTTGGGCACGAGGACCCT.
Reversible inhibition of elongation with DRB
Blood was collected from 10.5 day embryos in PBS and diluted to ∼106 cells/ml. Half of the cells were treated with 100 μm DRB (5,6 dichloro-1-β-d-ribofuranosyl-benzimidazole, Sigma) for 15 min at 37°C, the other half were used as untreated controls. The cells were then diluted with several volumes of ice-cold PBS, spun at 200g, and washed two more times with ice-cold PBS and resuspended in PBS at ∼106 cells/ml at 25°C. Aliquots were removed at 0, 2.5, 5, 7.5, 10, and 20 min after washing, spotted onto poly-l-lysine slides (Sigma) and prepared for in situ hybridization as described.
Acknowledgments
We thank S. Philipsen and A. van den Wijngaard for critical reading of this manuscript. We are grateful to M. Wijgerde, T McMorrow, E. Milot, M. Bender, and D. Higgs for helpful discussions and L. Braam for animal care. We are indebted to M. Kielman for the generous gift of the mouse α globin cosmid. T.T. is supported by the Nederlandse Organisatie voor Wetenschappelijk Onderzoek (N.W.O.). J.G. is supported by the Jeantet Foundation. This work is supported by the N.W.O.
The publication costs of this article were defrayed in part by payment of page charges. This article must therefore be hereby marked ‘advertisement’ in accordance with 18 USC section 1734 solely to indicate this fact.
Footnotes
E-MAIL fraser@ch1.fgg.eur.n1; FAX 31-10-436-0225.
References
- Alter BP, Goff SC. A murine model for the switch from fetal to adult hemoglobin during ontogeny. Blood. 1980;56:1100–1105. [PubMed] [Google Scholar]
- Bernet A, Sabatier S, Picketts D, Ouazana R, Morle F, Higgs D, Godet J. Targeted inactivation of the major positive regulatory element (HS-40) of the human α-globin gene locus. Blood. 1995;86:1202–1211. [PubMed] [Google Scholar]
- Brotherton T, Chui DH, Gauldie J, Patterson M. Hemoglobin ontogeny during normal mouse fetal development. Proc Natl Acad Sci. 1979;76:2853–2857. doi: 10.1073/pnas.76.6.2853. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chodosh LA, Fire A, Samuels M, Sharp PA. DRB inhibits transcription elongation by RNA polymerase II in vitro. J Biol Chem. 1989;264:2250–2257. [PubMed] [Google Scholar]
- Chui DH, Brotherton T, Gauldie B. Hemoglobin ontogeny in fetal mice. Adult hemoglobin in yolk sac derived erythrocytes. In: Stamatoyannopoulos G, Nienhuis A, editors. Cellular and molecular regulation of hemoglobin switching. New York, NY: Alan R. Liss; 1979. pp. 213–225. [Google Scholar]
- Collins FS, Weissman SM. The molecular genetics of human hemoglobin. Prog Nucleic Acids Res Mol Biol. 1984;31:315–461. doi: 10.1016/s0079-6603(08)60382-7. [DOI] [PubMed] [Google Scholar]
- Curcio MJ, Kantoff P, Schafer MP, Anderson WF, Safer B. Compensatory increase in levels of βminor globin in murine β thalassemia is under translational control. J Biol Chem. 1986;261:16126–16132. [PubMed] [Google Scholar]
- Dillon N, Trimborn T, Strouboulis J, Fraser P, Grosveld F. The effect of distance on long-range chromatin interactions. Mol Cell. 1997;1:131–139. doi: 10.1016/s1097-2765(00)80014-3. [DOI] [PubMed] [Google Scholar]
- Donze D, Townes TM, Bieker JJ. Role of erythroid Kruppel like factor in human γ to β-globin switching. J Biol Chem. 1995;270:1955–1959. doi: 10.1074/jbc.270.4.1955. [DOI] [PubMed] [Google Scholar]
- Enver T, Raich N, Ebens AJ, Papayannopoulou T, Constantini F, Stamatoyannopoulos G. Developmental regulation of human fetal-to-adult globin gene switching in transgenic mice. Nature. 1990;344:309–313. doi: 10.1038/344309a0. [DOI] [PubMed] [Google Scholar]
- Fiering S, Epner E, Robinson K, Zhuang Y, Telling A, Hu M, Martin D, Enver T, Ley T, Groudine M. Targeted deletion of 5′HS2 of the murine β-globin LCR reveals that it is not essential for proper regulation of the β-globin locus. Genes & Dev. 1995;9:2203–2213. doi: 10.1101/gad.9.18.2203. [DOI] [PubMed] [Google Scholar]
- Fraser NW, Sehgal PB, Darnell J. DRB-induced premature termination of late adenovirus transcription. Nature. 1978;272:590–593. doi: 10.1038/272590a0. [DOI] [PubMed] [Google Scholar]
- Fraser P, Gribnau J, Trimborn T. Mechanisms of developmental regulation in globin loci. Curr Opin Hematol. 1998;5:139–144. doi: 10.1097/00062752-199803000-00009. [DOI] [PubMed] [Google Scholar]
- Gourdon G, Sharpe J, Wells D, Wood W, Higgs D. Analysis of a 70 kb segment of DNA containing the human ζ and α-globin genes linked to their regulatory element (HS-40) in transgenic mice. Nucleic Acids Res. 1994;22:4139–4147. doi: 10.1093/nar/22.20.4139. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gourdon G, Sharpe J, Higgs D, Wood W. The mouse α-globin locus regulatory element. Blood. 1995;86:766–775. [PubMed] [Google Scholar]
- Gribnau J, deBoer E, Trimborn T, Wijgerde M, Milot E, Grosveld F, Fraser P. Chromatin interaction mechanism of transcriptional control in vivo. EMBO J. 1998;17:6020–6027. doi: 10.1093/emboj/17.20.6020. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Grosveld F, Blom van Assendelft G, Greaves D, Kolias G. Position independent high-level expression the human β-globin gene. Cell. 1987;51:975–985. doi: 10.1016/0092-8674(87)90584-8. [DOI] [PubMed] [Google Scholar]
- Hanscombe O, Whyatt D, Fraser P, Yannoutsos N, Greaves D, Dillon N, Grosveld F. Importance of globin gene order for correct developmental expression. Genes & Dev. 1991;5:1387–1394. doi: 10.1101/gad.5.8.1387. [DOI] [PubMed] [Google Scholar]
- Higgs D, Wood W, Jarman A, Sharpe J, Lida J, Pretorius I, Ayyub H. A major positive regulatory region located far upstream of the human α globin gene locus. Genes & Dev. 1990;4:4679–4689. doi: 10.1101/gad.4.9.1588. [DOI] [PubMed] [Google Scholar]
- Hug B, Moon A, Ley T. Structure and function of the murine β-globin locus control region 5′ HS3. Nucleic Acids Res. 1992;20:5771–5778. doi: 10.1093/nar/20.21.5771. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hug B, Wesselschmidt R, Fiering S, Bender M, Epner E, Groudine M, Ley T. Analysis of mice containing a targeted deletion of β-globin locus control region 5′ hypersensitive site 3. Mol Cell Biol. 1996;16:2906–2912. doi: 10.1128/mcb.16.6.2906. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jimenez G, Gale K, Enver T. The mouse β-globin locus control region: Hypersensitive sites 3 and 4. Nucleic Acids Res. 1992;20:5797–5803. doi: 10.1093/nar/20.21.5797. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kielman MF, Smits R, Bernini LF. Localization and characterization of the mouse α globin locus control region. Genomics. 1994;21:431–433. doi: 10.1006/geno.1994.1289. [DOI] [PubMed] [Google Scholar]
- Kim CG, Epner EM, Forrester WC, Groudine M. Inactivation of the human β globin gene by targeted insertion into the β globin locus control region. Genes & Dev. 1992;6:928–937. doi: 10.1101/gad.6.6.928. [DOI] [PubMed] [Google Scholar]
- Leder A, Kuo A, Shen M, Leder P. In situ hybridization reveals coexpression of embryonic and adult β-globin genes in the earliest murine erythrocyte progenitors. Development. 1992;116:1041–1049. doi: 10.1242/dev.116.4.1041. [DOI] [PubMed] [Google Scholar]
- Leder A, Daugherty C, Whitney B, Leder P. Mouse zeta- and alpha-globin genes: Embryonic survival, alpha-thalassemia, and genetic background effects. Blood. 1997;90:1275–1282. [PubMed] [Google Scholar]
- Liebhaber S, Wang Z, Cash F, Monks B, Russel E. Developmental silencing of the embryonic ζ-globin gene: Concerted action of the promoter and 3′-flanking region combined with stage-specific silencing by the transcribed segment. Mol Cell Biol. 1996;16:2637–2646. doi: 10.1128/mcb.16.6.2637. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lindenbaum MH, Grosveld F. An in vitro globin gene switching model based on differentiated embryonic stem cells. Genes & Dev. 1990;4:2075–2085. doi: 10.1101/gad.4.12a.2075. [DOI] [PubMed] [Google Scholar]
- Marshall NF, Price DH. Control of formation of two distinct classes of RNA polymerase II elongation complexes. Mol Cell Biol. 1992;12:2078–2090. doi: 10.1128/mcb.12.5.2078. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Marshall NF, Peng J, Xie Z, Price DH. Control of RNA polymerase II elongation potential by a novel carboxyl-terminal domain kinase. J Biol Chem. 1996;271:27176–27183. doi: 10.1074/jbc.271.43.27176. [DOI] [PubMed] [Google Scholar]
- Milot E, Strouboulis J, Trimborn T, Wijgerde M, de Boer E, Langeveld A, Tan-Un K, Vergeer W, Yannoutsos N, Grosveld F, Fraser P. Heterochromatin effects on the frequency and duration of LCR-mediated gene transcription. Cell. 1996;87:105–114. doi: 10.1016/s0092-8674(00)81327-6. [DOI] [PubMed] [Google Scholar]
- Moon A, Ley T. Conservation of the primary structure, organization and function of the human and mouse β-globin locus-activating regions. Proc Natl Acad Sci. 1990;87:7693–7697. doi: 10.1073/pnas.87.19.7693. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nuez B, Michalovich D, Bygrave A, Ploemacher R, Grosveld F. Defective haematopoiesis in fetal liver resulting from inactivation of the EKLF gene. Nature. 1995;375:316–318. doi: 10.1038/375316a0. [DOI] [PubMed] [Google Scholar]
- Papayannopoulou T, Brice M, Stamatoyannopoulos G. Analysis of human hemoglobin switching in MEL X human fetal erythroid cell hybrids. Cell. 1986;46:469–476. doi: 10.1016/0092-8674(86)90667-7. [DOI] [PubMed] [Google Scholar]
- Perkins AC, Sharpe AH, Orkin S. Lethal β-thalassemia in mice lacking the erythroid CACCC-transcription factor EKLF. Nature. 1995;375:318–322. doi: 10.1038/375318a0. [DOI] [PubMed] [Google Scholar]
- Perkins AC, Gaensler KML, Orkin SH. Silencing of human fetal globin expression is impaired in the absence of the adult β-globin gene activator protein EKLF. Proc Natl Acad Sci. 1996;93:12267–12271. doi: 10.1073/pnas.93.22.12267. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Peterson KR, Stamatoyannopoulos G. Role of gene order in developmental control of human gamma- and beta-globin gene expression. Mol Cell Biol. 1993;13:4836–4843. doi: 10.1128/mcb.13.8.4836. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pham CT, MacIvor DM, Hug BA, Heusel JW, Ley TJ. Long range disruption of gene expression by a selectable marker cassette. Proc Natl Acad Sci. 1996;93:13090–13095. doi: 10.1073/pnas.93.23.13090. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Raich N, Enver T, Nakamoto B, Josephson B, Papayannopoulou T, Stamatoyannopoulos G. Autonomous developmental control of human embryonic globin gene switching in transgenic mice. Science. 1990;250:1147–1149. doi: 10.1126/science.2251502. [DOI] [PubMed] [Google Scholar]
- Sharp J, Can-Thomas P, Lida J, Ayyub H, Wood W, Higgs D. Analysis of the human α-globin upstream regulatory element (HS-40) in transgenic mice. EMBO J. 1992;11:4564–4570. doi: 10.1002/j.1460-2075.1992.tb05558.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shehee R, Oliver P, Smithies O. Lethal thalassemia after insertional disruption of the mouse β major gene. Proc Natl Acad Sci. 1993;90:3177–3181. doi: 10.1073/pnas.90.8.3177. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Skow L, Burkhart B, Johnson F, Popp R, Goldberg S, Anderson W, Barnett L, Lewis S. A mouse model for β-thalassemia. Cell. 1983;34:1043–1052. doi: 10.1016/0092-8674(83)90562-7. [DOI] [PubMed] [Google Scholar]
- Stanworth SJ, Roberts NA, Sharpe JA, Sloane-Stanley JA, Wood W. Established epigenetic modifications determine the expression of developmentally regulated globin genes in somatic cell hybrids. Mol Cell Biol. 1995;15:3969–3978. doi: 10.1128/mcb.15.8.3969. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Strouboulis J, Dillon N, Grosveld F. Developmental regulation of a complete 70-kb human β-globin locus in transgenic mice. Genes & Dev. 1992;6:1857–1864. doi: 10.1101/gad.6.10.1857. [DOI] [PubMed] [Google Scholar]
- Tweeten KA, Molloy GR. Induction of premature termination of transcription of the mouse β globin gene by DRB. Nucleic Acids Res. 1981;9:3307–3319. doi: 10.1093/nar/9.14.3307. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wawrzyniak C, Popp R. Expression of the two adult β-globin genes in mouse yolk sac and fetal liver erythrocytes. Dev Biol. 1987;119:299–301. doi: 10.1016/0012-1606(87)90231-4. [DOI] [PubMed] [Google Scholar]
- Weaver RF, Weissman C. Mapping of RNA by a modification of th Berk-Sharp procedure: The 5′ termini of 15s β globin mRNA precursor and mature 10s β globin mRNA have identical map co-ordinates. Nucleic Acids Res. 1979;6:1175–1193. doi: 10.1093/nar/7.5.1175. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Whitelaw E, Tsai S, Hogben P, Orkin S. Regulated expression of globin chains and erythroid transcription factor GATA-1 during erythropoiesis in the developing mouse. Mol Cell Biol. 1990;10:6596–6606. doi: 10.1128/mcb.10.12.6596. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Whitney JB. Differential control of the synthesis of two hemoglobin β chains in normal mice. Cell. 1977;12:863–871. doi: 10.1016/0092-8674(77)90150-7. [DOI] [PubMed] [Google Scholar]
- Wijgerde M, Grosveld F, Fraser P. Transcription complex stability and chromatin dynamics in vivo. Nature. 1995;377:209–213. doi: 10.1038/377209a0. [DOI] [PubMed] [Google Scholar]
- Wijgerde M, Gribnau J, Trimborn T, Nuez B, Philipsen S, Grosveld F, Fraser P. The role of EKLF in human β-globin gene competition. Genes & Dev. 1996;10:2894–2902. doi: 10.1101/gad.10.22.2894. [DOI] [PubMed] [Google Scholar]
- Wong PMC, Chung S, White JS, Reicheld SM, Patterson M, Clarke BJ, Chui DHK. Adult hemoglobins are synthesized in murine fetal hepatic erythropoietic cells. Blood. 1983;62:1280–1288. [PubMed] [Google Scholar]
- Zafarana G, Raguz S, Pruzina S, Grosveld F, Meijer D. The regulation of human β-globin gene expression: The analysis of hypersensitive site 5 (HS5) in the LCR. In: Stamatoyannopoulos G, editor. Molecular biology of hemoglobin switching. Proceedings of the Ninth Conference on Hemoglobin Switching. Andover, Hants, UK: Intercept Ltd.; 1995. pp. 39–44. [Google Scholar]