Abstract
A simple and highly efficient route to the FKBP-binding domain (FKBD) from the natural product rapamycin has been developed, which entails a sequence of ozonolysis/Baeyer-Villiger/Wittig reactions. The newly synthesized FKBD may serve as a core to assemble hybrid macrocyclic libraries for the discovery of novel probes of protein function and to synthesize new ligands for the FKBP family of proteins.
Keywords: FKBP, Rapamycin, FKBP-binding domain
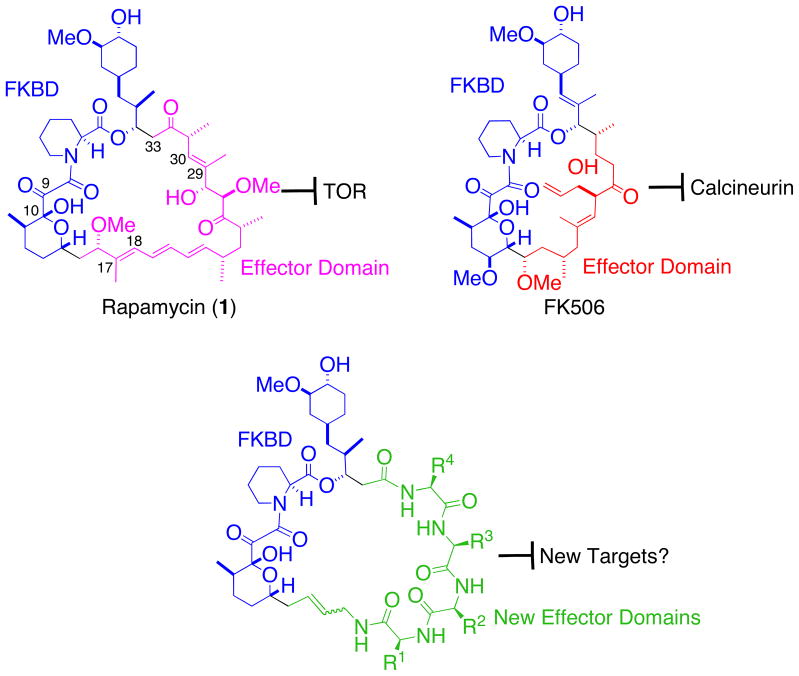
The natural products FK506 and rapamycin possess potent immunosuppressive and antitumor (rapamycin) activities and they share a unique and extraordinary mode of action—through inducing the formation of ternary complexes between an abundant cellular enzyme known as FKBP and their respective protein targets.1–5 Upon entering cells, both FK506 and rapamycin form binary complexes with FKBP. While the FKBP-FK506 complex binds to and inhibits the enzymatic activity of the protein phosphatase calcineurin, blocking calcium-dependent intracellular signaling emanating from the T cell receptor,1 the FKBP-rapamycin complex binds to a distinct target—the protein kinase mTOR that serves as a nodal point in signaling leading to the regulation of a multitude of cellular processes including cell proliferation.2–5 Extensive biochemical and structural studies have revealed that both FK506 and rapamycin can be divided into two structural and functional domains, an FKBP-binding domain (FKBD) and a so-called effector domain. A comparison of the structures of FK506 and rapamycin revealed that they share a nearly identical FKBD but each possesses a distinct effector domain (Fig. 1). In FK506 and rapamycin, Nature has taught us that by swapping the effector domain of FK506 with that of rapamycin, it is possible to change the target from calcineurin to TOR, which bear no sequence, functional or structural similarities to each other.6–9 We were thus inspired to ask the question: Is it possible to target other proteins by grafting new structures onto FKBD of rapamycin and FK506?
Figure 1.
Rapamycin, FK506, their FKBP-binding domains (blue), effector domains (pink and red) and generic structures of the proposed new hybrid combinatorial cyclic libraries.
To answer this question, we will need to synthesize sufficiently large combinatorial hybrid libraries that display random peptides or other building blocks on FKBD. As a first step of this exploration, we needed access to large quantities of FKBD. Although synthetic versions of FKBD can be employed,10 we felt it necessary to compare the relative affinities of the “natural” FKBD with the synthetic versions before embarking on the synthesis of libraries with an optimal version of FKBD. In one of our library designs, we envisioned the use of ring-closing metathesis to achieve the macrocyclization, calling for the use of an FKBD that is equipped with a terminal olefin.
Previous syntheses of FKBD from rapamycin had involved the initial cleavage of the inherently labile ester bond at C-34 of rapamycin, which necessitated the reassembly of the two resulting pieces of FKBD.11 Should the same route be followed, it would not only increase the number of steps, be less elegant and efficient, it would also significantly increase the demand for the not-so-cheap rapamycin as the starting material. After attempting a number of potential routes, we developed a simple and efficient method to prepare the desired FKBD from rapamycin while keeping the entire FKBD intact throughout the process.
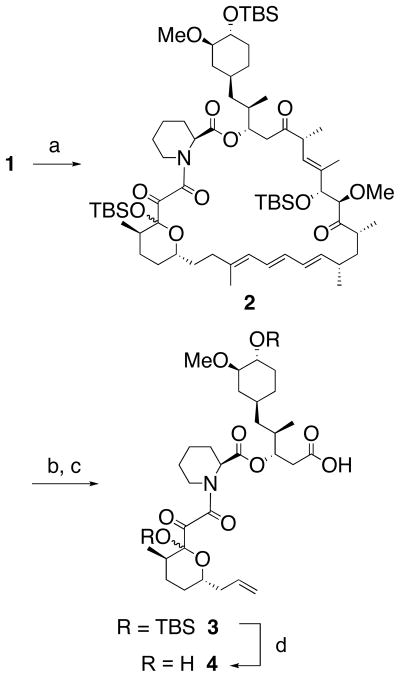
Our preparation of FKBD commenced with the treatment of rapamycin (1) with TBSOTf and Et3N to afford the fully protected epimers (2) in quantitative yield (Scheme 1). According to an earlier report by Goulet et al.,12 protection of the hydroxyl at the hemiketal center (C-10) of rapamycin is not necessary for the following ozonolysis step. However, the corresponding C-10 unprotected FKBD fragment was cleaved between C-9 and C-10 when subjected to Baeyer-Villiger oxidation.12 Thus, we deemed the protection of the C-10 hydroxyl to be an absolute requirement to succeed in our strategy. The rapamycin derivative 2 was then subjected to ozonolysis in CH2Cl2 at −68 °C and subsequently treated with 35% H2O2 to give an intermediate containing a carboxylic acid at one end and a peroxyhemiacetal or aldehyde at the other end of the FKBD fragment.13,14 After a brief flash-column chromatography over silica gel, the crude product was directly subjected to Wittig olefination to yield product 3 in 54% combined yield.15,16 Deprotection of the TBS protecting groups with trifluoroacetic acid and H2O at 0 °C gave the final product FKBD (4) in 96% yield.17 It is noteworthy that neither TABF nor HF·pyridine were able to accomplish a clean deprotection of the two silyl groups. Thus, FKBD was prepared from rapamycin in just four steps with 52% overall yield.
Scheme 1.
Preparation of FKBD from rapamycin. (a) TBSOTf, Et3N, CH2Cl2, r.t., 100%; (b) O3, CH2Cl2, −68 °C; then 35% H2O2, r.t.; (c) CH3PPh3Br, t-BuOK, THF, 0 °C (54% b–c); (d) CF3COOH, H2O, 0 °C (96%).
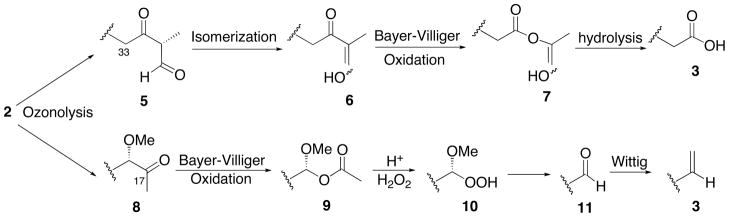
We were unable to isolate and completely characterize the intermediates after ozonolysis and the Baeyer-Villiger reaction due in part to the complexity of the reaction products. The complexity was further compounded by the propensity of the FKBD to exhibit rotamerism and in this case we were also dealing with a pair of epimers. The most likely intermediates in the course of these two tandem steps (ozonolysis/Baeyer-Villiger) can be predicted (Scheme 2). Ozonolysis of the C29-C30 double bond gave rise to an aldehyde intermediate (5) that could undergo facile enolization to give 6. This rendered 6 with an exclusive migratory aptitude in the Baeyer-Villiger rearrangement to give ester 7. Hydrolysis of 7 led to the formation of the carboxylic group in 3. Concurrently, ozonolysis of C17-C18 double bond yielded the ketone intermediate 8. Baeyer-Villiger oxidation of 8 led to the formation, possibly via ester 9, of the peroxyhemiacetal (10) that, upon hydrolysis, gave the corresponding aldehyde 11, which is ready to undergo Wittig reaction to form the terminal olefin in 3. An interesting feature of this procedure is that the same set of reactions occurred concomitantly at both ends of FKBD, but led to the formation of distinct functional groups--a carboxylic acid and an olefinic group. This simultaneous chemical transformation of two groups significantly reduced the number of steps required to prepare FKBD, hence the high overall yield.
Scheme 2.
Proposed reaction intermediates.
In addition to serving our purpose for the synthesis of FKBD-containing hybrid combinatorial libraries, high-affinity ligands of FKBP isoforms may find use as probes for a number of biological processes, as FKBP themselves have been implicated in the regulation of neuronal cell differentiation, ion channels, and Ras post-translational modification among others.18–22 The easy access to FKBD could thus allow for the synthesis of potent and possibly isoform-selective FKBP ligands that are devoid of immunosuppressive and antiproliferative activity of FK506 and rapamycin.
In conclusion, a simple and highly efficient process for the preparation of FKBD from rapamycin has been developed using a one-pot ozonolysis and Baeyer-Villiger reaction as the key step.
Acknowledgments
This work was supported by the NIH Director’s Pioneer Award (DP1OD006795 to JOL). We are grateful to Drs. Zufeng Guo, Manisha Das and members of the Liu Lab for helpful discussions. We thank Drs. Carol Greider and Paul Talalay for generous provision of temporary lab space for this work.
Footnotes
Publisher's Disclaimer: This is a PDF file of an unedited manuscript that has been accepted for publication. As a service to our customers we are providing this early version of the manuscript. The manuscript will undergo copyediting, typesetting, and review of the resulting proof before it is published in its final citable form. Please note that during the production process errors may be discovered which could affect the content, and all legal disclaimers that apply to the journal pertain.
References and Notes
- 1.Liu J, Farmer JD, Jr, Lane WS, Friedman J, Weissman I, Schreiber SL. Cell. 1991;66:807–815. doi: 10.1016/0092-8674(91)90124-h. [DOI] [PubMed] [Google Scholar]
- 2.Heitman J, Movva NR, Hall MN. Science. 1991;253:905–909. doi: 10.1126/science.1715094. [DOI] [PubMed] [Google Scholar]
- 3.Brown EJ, Albers MW, Shin TB, Ichikawa K, Keith CT, Lane WS, Schreiber SL. Nature. 1994;369:756–758. doi: 10.1038/369756a0. [DOI] [PubMed] [Google Scholar]
- 4.Sabatini DM, Erdjument-Bromage H, Lui M, Tempst P, Snyder SH. Cell. 1994;78:35–43. doi: 10.1016/0092-8674(94)90570-3. [DOI] [PubMed] [Google Scholar]
- 5.Chen Y, Chen H, Rhoad AE, Warner L, Caggiano TJ, Failli A, Zhang H, Hsiao CL, Nakanishi K, Molnar-Kimber KL. Biochem Biophys Res Commun. 1994;203:1–7. doi: 10.1006/bbrc.1994.2140. [DOI] [PubMed] [Google Scholar]
- 6.Bierer BE, Somers PK, Wandless TJ, Burakoff SJ, Schreiber SL. Science. 1990;250:556–9. doi: 10.1126/science.1700475. [DOI] [PubMed] [Google Scholar]
- 7.Griffith JP, Kim JL, Kim EE, Sintchak MD, Thomson JA, Fitzgibbon MJ, Fleming MA, Caron PR, Hsiao K, Navia MA. Cell. 1995;82:507–22. doi: 10.1016/0092-8674(95)90439-5. [DOI] [PubMed] [Google Scholar]
- 8.Kissinger CR, Parge HE, Knighton DR, Lewis CT, Pelletier LA, Tempczyk A, Kalish VJ, Tucker KD, Showalter RE, Moomaw EW, et al. Nature. 1995;378:641–4. doi: 10.1038/378641a0. [DOI] [PubMed] [Google Scholar]
- 9.Choi J, Chen J, Schreiber SL, Clardy J. Science. 1996;273:239–42. doi: 10.1126/science.273.5272.239. [DOI] [PubMed] [Google Scholar]
- 10.Holt DA, Juengo JI, Yamashita DS, Oh H, Konialian AL, Yen H, Rozamus LW, Brandt M, Bossard MJ, Levy MA, Eggleston DS, Liang J, Schultz LW, Stout TJ, Clardy J. J Am Chem Soc. 1993;115:9925–9938. [Google Scholar]
- 11.Chakraborty TK, Weber HP, Nicolaou KC. Chem Biol. 1995;2:157–61. doi: 10.1016/1074-5521(95)90070-5. [DOI] [PubMed] [Google Scholar]
- 12.Goulet MT, Boger J. Tetrahedron Lett. 1990;31:4845–4848. [Google Scholar]
- 13.Moulines J, Bats JP, Lamidey AM, Silva ND. Helv Chim Acta. 2004;87:2695–2705. [Google Scholar]
- 14.Note: Shortly after the addition of H2O2 the reaction mixture attains a pH of 2. Oxidative cleavage of the ozonides at C-17, C-19 and C-21 ought to produce two equivalents of oxalic acid. With the excess of hydrogen peroxide around peroxalic acid can be generated in situ and it can mediate the Baeyer-Villiger rearrangemnets. We have observed this to be a convenient and efficient process rather than isolating the ozonolysis product and subjecting to Baeyer-Villiger reaction using peracetic acid or mCPBA.
- 15.Getty SJ, Berson JA. J Am Chem Soc. 1990;112:1652–1653. [Google Scholar]
- 16.Wong JCY, Lacombe P, Sturino CF. Tetrahedron Lett. 1999;40:8751–8754. [Google Scholar]
- 17.Synthetic Procedures:(2) To a solution of rapamycin (999 mg, 1.09 mmol) and anhydrous Et3N (0.6 mL, 4.45 mmol) in anhydrous CH2Cl2 (3 mL) was added TBSOTf (0.85 mL, 3.79 mmol). The solution was stirred for 1 h at r.t., and then quenched with H2O. The resulting mixture was diluted with EtOAc (120 mL), washed with saturated NaHCO3 and brine, dried and concentrated in vacuo. The crude residue was subjected to silica gel column chromatography (hexane-EtOAc, 8:1) to afford 2 (1.37 g, 100%) as a yellow solid: 1H NMR (400 MHz, CDCl3): δ 6.39–5.82 (m, 4 H), 5.52–4.91 (m, 4 H), 4.45–3.98 (m, 2 H), 3.83 (d, J = 4.0 Hz, 1 H), 3.40 (s, 3 H), 3.27 (s, 3 H), 3.10 (s, 3 H), 2.89–2.83 (m, 1 H), 2.66–2.40 (m, 3 H), 2.23–2.19 (m, 3 H), 1.70 (s, 3 H), 0.17–0 (m, 18 H); 13C NMR (100 MHz, CDCl3): δ 210.3, 206.6, 197.9, 169.5, 167.0, 139.3, 139.0, 136.6, 132.0, 130.8, 127.4, 126.7, 126.4, 101.7, 86.6, 84.3, 83.2, 78.0, 75.7, 74.3, 67.7, 58.1, 57.8, 56.1, 51.4, 45.8, 44.1, 41.4, 40.8, 40.6, 40.2, 38.2, 36.8, 35.7, 34.0, 33.9, 33.0, 32.0, 30.1, 27.1, 26.6, 26.3, 26.0, 25.9, 25.8, 25.7, 24.9, 22.0, 21.0, 19.3, 18.2, 18.1, 15.7, 15.5, 14.0, 13.9, 11.0, −2.5, −3.2, −4.5, −4.6, −4.7, −4.9; HR-ESIMS calcd for C69H121O13NSi3Na [M + Na]+ 1278.8043, found 1278.8049.(3) To a solution of silyl derivative 2 (1.20 g, 0.95 mmol) in CH2Cl2 (15 mL) at −68 °C was bubbled O3 until the blue color persisted. 35% H2O2 (15 mL) was then added and the stirring was continued for another 14 h at r.t.. The solution was diluted with EtOAc (120 mL), washed with brine, dried and concentrated to afford an oil, which was purified using silica gel chromatography (hexane-EtOAc, 4:1, then CH2Cl2-MeOH, 20:1) to give an epimeric mixture (710 mg) as a white solid. The epimeric mixture from the ozonolysis reaction stated above (210 mg, 0.26 mmol) was dissolved in anhydrous THF (2 mL), and then added to a freshly prepared Wittig reagent at 0 °C, which in turn was prepared from CH3PPh3+Br− (560 mg, 1.57 mmol) and t-BuOK (140 mg, 1.25 mmol) in anhydrous THF (5 mL). After stirring for 10 min, the reaction mixture was quenched with 5% HCl. The resulting mixture was diluted with EtOAc (60 mL), washed with brine, dried and concentrated. The crude residue was purified by silica gel chromatography (toluene-EtOAc, 4:1, then toluene-EtOAc-AcOH, 4:1:0.025) to provide 3 (124 mg, 54%) as a white solid: 1H NMR (400 MHz, CDCl3): 5.84–5.68 (m, 1 H), 5.29–4.89 (m, 4 H), 4.42–4.29 (m, 1 H), 4.06–3.85 (m, 1 H), 3.40 (s, 3 H), 2.93–2.48 (m, 4 H), 2.34–2.13 (m, 4 H), 0.87 (s, 9 H), 0.19–0.02 (m, 12 H); 13C NMR (100 MHz, CDCl3): δ 197.7, 197.4, 176.0, 175.4, 169.7, 169.2, 167.2, 166.4, 135.3, 134.3, 117.2, 116.6, 102.0, 101.6, 84.5, 84.4, 75.7, 75.2, 74.6, 70.5, 70.3, 58.1, 58.0, 56.6, 56.5, 51.8, 44.4, 40.3, 38.9, 38.6, 36.5, 36.4, 36.2, 36.0, 35.1, 34.7, 33.9, 33.34, 33.25, 33.1, 31.4, 30.9, 30.6, 29.7, 29.4, 27.6, 27.3, 26.84, 26.76, 26.6, 26.3, 25.9, 24.8, 24.4, 22.8, 21.3, 20.7, 19.4, 18.2, 15.82, 15.78, 15.3, 14.9, −2.5, −2.7, −2.9, −3.0, −4.5, −4.7; HR-ESIMS calcd for C42H75O10NSi2Na [M + Na]+ 832.4827, found 832.4833.(4) To a mixture of CF3CO2H (0.8 mL) and H2O (0.2 mL) at 0 °C was added olefin 3 (32 mg, 0.040 mmol). After stirring for 2 h, the mixture was concentrated in vacuo and purified by silica gel chromatography (CH2Cl2-MeOH-AcOH, 20:1:0.2) to offer 4 (22 mg, 96%) as a white solid: 1H NMR (400 MHz, CDCl3): 5.82–5.68 (m, 1 H), 5.27–4.90 (m, 4 H), 4.45–4.32 (m, 1 H), 4.02–3.85 (m, 1 H), 3.40 (s, 3 H), 2.99–2.94 (m, 1 H), 2.65–2.51 (m, 2 H); 13C NMR (100 MHz, CDCl3): δ 197.8, 194.8, 175.1, 175.0, 171.0, 170.0, 169.8, 169.34, 169.26, 167.6, 166.8, 165.7, 135.1, 134.8, 134.3, 117.4, 117.1, 116.7, 99.3, 98.5, 97.7, 84.4, 75.7, 75.2, 75.0, 73.9, 70.2, 70.1, 56.5, 56.4, 51.5, 44.5, 43.1, 41.5, 41.4, 40.2, 39.1, 38.9, 35.1, 35.0, 34.4, 34.3, 34.1, 33.4, 33.2, 31.3, 30.8, 30.6, 29.7, 27.5, 27.1, 26.6, 26.4, 26.3, 25.0, 24.6, 24.4, 21.1, 20.9, 20.8, 16.7, 16.1, 15.9, 15.7, 15.3, 15.1, 15.0; HR-ESIMS calcd for C30H47O10NNa [M + Na]+ 604.3098, found 604.3093.
- 18.Lyons WE, George EB, Dawson TM, Steiner JP, Snyder SH. Proc Natl Acad Sci USA. 1994;91:3191–5. doi: 10.1073/pnas.91.8.3191. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Brillantes AB, Ondrias K, Scott A, Kobrinsky E, Ondriasova E, Moschella MC, Jayaraman T, Landers M, Ehrlich BE, Marks AR. Cell. 1994;77:513–23. doi: 10.1016/0092-8674(94)90214-3. [DOI] [PubMed] [Google Scholar]
- 20.Shou W, Aghdasi B, Armstrong DL, Guo Q, Bao S, Charng MJ, Mathews LM, Schneider MD, Hamilton SL, Matzuk MM. Nature. 1998;391:489–92. doi: 10.1038/35146. [DOI] [PubMed] [Google Scholar]
- 21.Xin HB, Senbonmatsu T, Cheng DS, Wang YX, Copello JA, Ji GJ, Collier ML, Deng KY, Jeyakumar LH, Magnuson MA, Inagami T, Kotlikoff MI, Fleischer S. Nature. 2002;416:334–8. doi: 10.1038/416334a. [DOI] [PubMed] [Google Scholar]
- 22.Ahearn IM, Tsai FD, Court H, Zhou M, Jennings BC, Ahmed M, Fehrenbacher N, Linder ME, Philips MR. Mol Cell. 2011;41:173–85. doi: 10.1016/j.molcel.2011.01.001. [DOI] [PMC free article] [PubMed] [Google Scholar]