Abstract
Caenorhabditis elegans sur-8 encodes a positive regulator of Ras signaling. We investigated the mechanism by which the human Sur-8 homolog can positively regulate Ras–MAP kinase signaling in mammalian cells. Sur-8 expression enhances Ras- or EGF-induced Raf and ERK activation but has no effect on ERK activation induced by active Raf or MEK. Furthermore, Sur-8 expression does not increase AKT or JNK activation. Sur-8 interacts with Ras and Raf and is able to form a ternary complex with the two proteins. Thus, Sur-8 may function as a scaffold that enhances Ras–MAP kinase signal transduction by facilitating the interaction between Ras and Raf.
Keywords: Sur-8, RTK, Ras, Raf, ERK
Ras relays signals from receptor tyrosine kinases (RTKs) to activation of the mitogen-activated protein kinase (MAPK) pathway (van der Geer et al. 1994; Robinson and Cobb 1997). Ras directly interacts with and activates Raf, which phosphorylates and activates MEK. MEK is a dual-specific kinase that selectively phosphorylates and activates the extracellular signal-regulated kinase, ERK, a MAPK family member. Ras also interacts with numerous other downstream molecules, although the functional significance of many of the Ras-interacting proteins is not clear (Katz and McCormick 1997; Vojtek and Der 1998). However, it has been shown that PI3K and RalGDS are physiological targets of Ras (Rodriguez-Viciana et al. 1994; Urano et al. 1996).
Genetic studies in Caenorhabditis elegans and Drosophila demonstrate that the Ras–MAPK pathway is essential for signal transduction of RTK (Chang et al. 1994; Sternberg and Han 1998). Recent genetic studies have identified additional molecules that may regulate Ras–MAPK signaling. Sieburth et al. (1998) isolated sur-8 by screening for mutations that can suppress the Multivulva phenotype of an activated let-60 Ras mutation. Selfors et al. (1998) identified soc-2, which is identical to sur-8, by isolating suppressors of clr-1, which encodes a receptor tyrosine phosphatase. soc-2 mutations also suppress the activity of an hyperactive mutation in egl-15 FGF receptor gene, suggesting that soc-2 functions in RTK signaling. Although single mutation of sur-8 in an otherwise wild-type background causes no obvious phenotype, mutation of sur-8 dramatically enhances the weak loss-of-function mutation in mpk-1 (Sieburth et al. 1998), indicating that sur-8 is required for optimal Ras–MAPK signaling. These properties of sur-8 are similar to other worm genes such as ksr (Kornfeld et al. 1995; Sundaram and Han 1995) and sur-6 PP2A-B (Sieburth et al. 1999). Because double mutations in sur-8 and ksr essentially eliminate vulval induction in C. elegans, the functions of these two genes together are essential for Ras signaling.
The deduced amino acid sequence of sur-8 contains multiple leucine-rich repeats (LRR), which are present in numerous proteins and may mediate protein–protein interaction (Kobe and Deisenhofer 1993). LRR is found in the yeast adenylyl cyclase (Kataoka et al. 1985), which is a direct downstream target of Ras, and the LRR sequences in the yeast adenylyl cyclase mediate its interaction with Ras (Field et al. 1990; Suzuki et al. 1990). Genetic data suggest that sur-8 functions in RTK signaling, most likely downstream of Ras and upstream of Raf. Both C. elegans and mammalian Sur-8 proteins were shown to directly bind to Ras (Sieburth et al. 1998); however, the biochemical mechanism of Sur-8 in Ras signaling has not been determined.
The human Sur-8 cDNA can functionally complement the sur-8 mutation in C. elegans (Sieburth et al. 1998), suggesting that the Sur-8 function is conserved. We tested the biochemical functions of human Sur-8 on Ras-MAPK signaling. Data in this report suggest that Sur-8 may modulate the potency and specificity of Ras signaling.
Results
Sur-8 enhances the activation of ERK by EGF or Ras
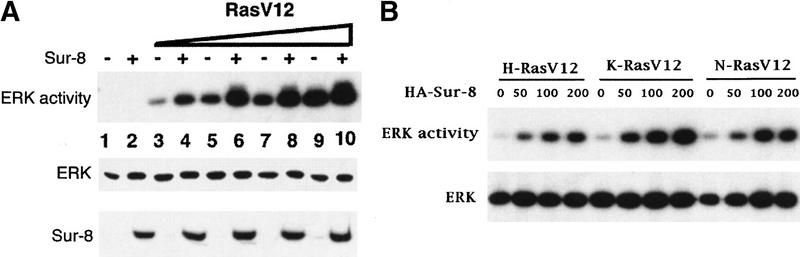
We tested the effect of expressing human Sur-8 on the activity of ERK in mammalian cells. Myc-tagged ERK1 was cotransfected into 293 cells with Ras and Sur-8 followed by immunoprecipitation and an in vitro kinase assay. Sur-8 alone had no significant effect on ERK activity (Fig. 1A). However, in the presence of Ras, Sur-8 significantly enhanced the ability of Ras to activate ERK. The enhancement of ERK activity by Sur-8 was most significant at low concentrations of Ras signal (Fig. 1A). Sur-8 had similar effects on ERK activation in response to H-Ras, K-Ras, and N-Ras (Fig. 1B). Sur-8 enhanced ERK activation in a dose-dependent manner (Fig. 1B).
Figure 1.
Sur-8 enhances ERK activation. (A) Myc-tagged ERK1 (200 ng) was cotransfected with increasing amounts of RasV12 in the presence or absence of Sur-8 (500 ng) as indicated. The amounts of RasV12 (all Ras plasmids used are K-Ras except as described) used for transfection are 0 (lanes 1,2), 5 ng (lanes 3,4), 10 ng (lanes 5,6), 20 ng (lanes 7,8), and 50 ng (lanes 9,10). Myc–ERK was immunoprecipitated and kinase activity was determined using GST–Elk-1 as a substrate (top). Myc–ERK in the immunoprecipitate was determined by anti-ERK Western blot (middle). Expression of HA–Sur-8 in cell lysates was detected by anti-HA Western blot (bottom). (B) Dose-dependent effect of Sur-8 on ERK activity. Cells were cotransfected with 1 ng of H-RasV12, K-RasV12, or N-RasV12, and 200 ng of Myc–ERK. The amount of HA–Sur-8 (ng DNA) cotransfected is indicated. (C) Sur-8 enhances EGF-stimulated ERK activity. The 293 cells were cotransfected with 200 ng of Myc–ERK and 500 ng of HA–Sur-8. After cells were starved in serum-free medium overnight, the cells were stimulated for 5 min by indicated concentrations of EGF. Myc–ERK was immunoprecipitated and activity was determined. (D) Mutations at Cys-260 and Glu457 of Sur-8 compromised its functions in ERK activation. Sur-8 or mutants (50 and 500 ng) were cotransfected with Myc–ERK and RasV12. Immunoprecipitated ERK activity and expression levels are shown at top and middle, respectively. HA–Sur-8 expression was detected as above (bottom).
The effect of Sur-8 on EGF-stimulated ERK activation was also examined. At low EGF concentrations, cotransfection of Sur-8 significantly increased EGF-induced ERK activity (Fig. 1C). However, at a high concentration of EGF (100 ng/ml), maximum ERK activation was observed and Sur-8 expression had no significant effect (data not shown). Together, these results indicate that Sur-8 has a positive function in RTK and Ras-induced ERK activation.
A genetic screen isolated two loss-of-function mutants of C. elegans sur-8 as Ras suppressors (Sieburth et al. 1998). The corresponding mutations, C260Y and E457K, were created in human Sur-8, and their abilities to enhance ERK activity were examined. Both C260Y and E457K mutants significantly reduced Sur-8's ability to enhance Ras-stimulated ERK activation (Fig. 1D). Thus, the ability of Sur-8 to enhance the ERK activation may be relevant to its in vivo function as a positive modulator of the Ras–MAPK signaling pathway.
Sur-8 acts to up-regulate Raf activity
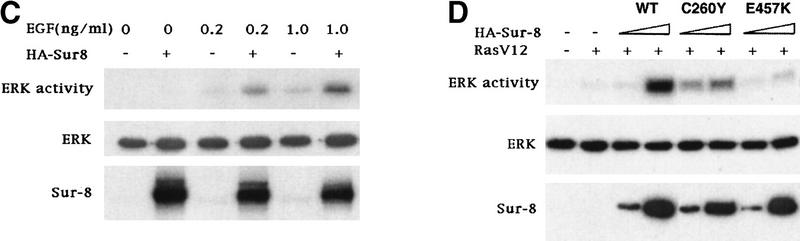
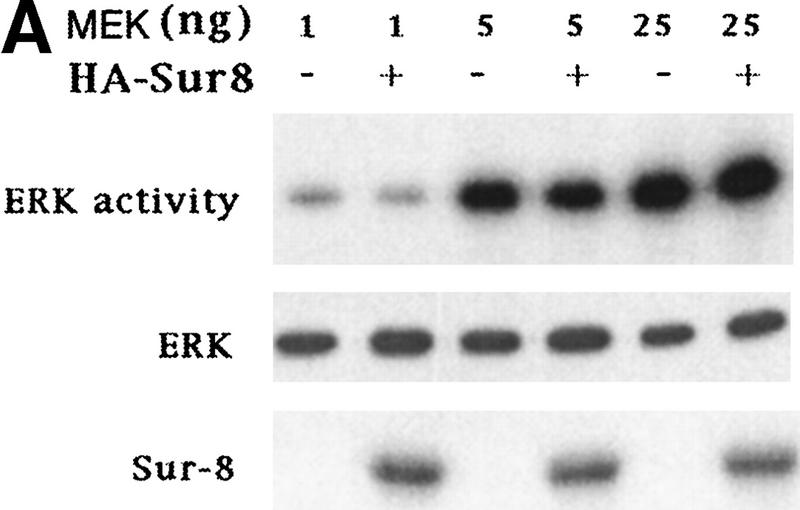
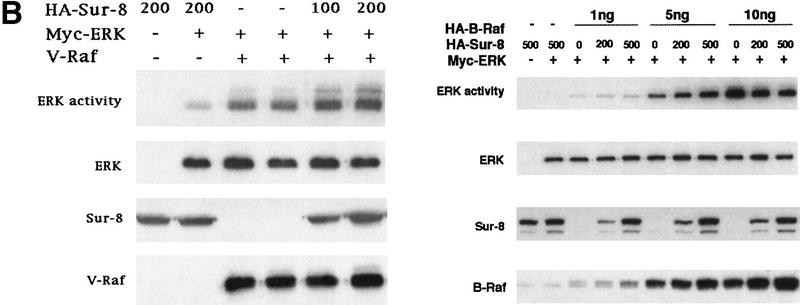
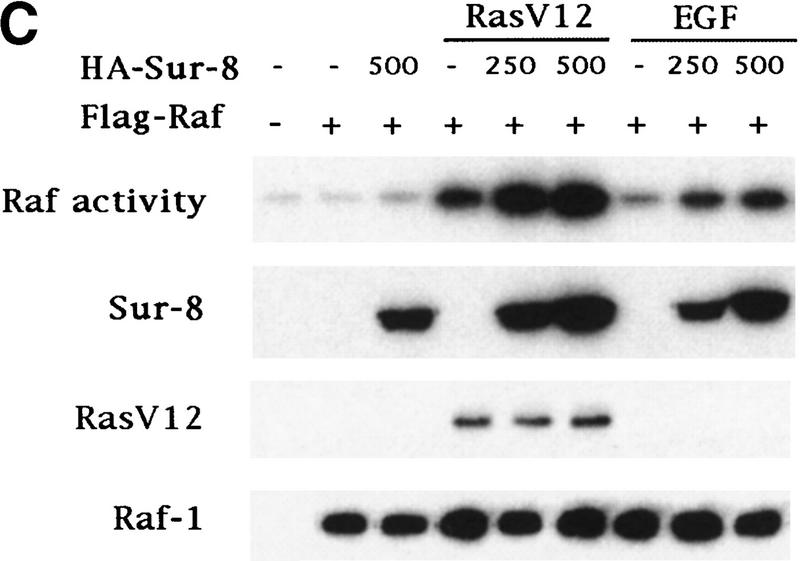
To determine at which step Sur-8 affects ERK activation in mammalian cells, we examined the effect of Sur-8 on ERK activation induced by active Raf or active MEK. Sur-8 had no significant effect on ERK activation stimulated by a constitutively active MEK (Fig. 2A; Mansour et al. 1994) , suggesting that Sur-8 functions upstream of MEK. Sur-8 showed no significant effect on either V-Raf (Howe et al. 1992; Kyriakis et al. 1992) or active B-Raf-induced (A. Vojtek, unpubl.) ERK activation (Fig. 2B), supporting the notion that Sur-8 acts upstream of Raf. To test this hypothesis further, we examined whether Sur-8 enhances Raf activation. Sur-8 enhanced RasV12-stimulated, as well as EGF-induced, Raf activity (Fig. 2C). Therefore, these data are consistent with a model that Sur-8 functions upstream of Raf.
Figure 2.
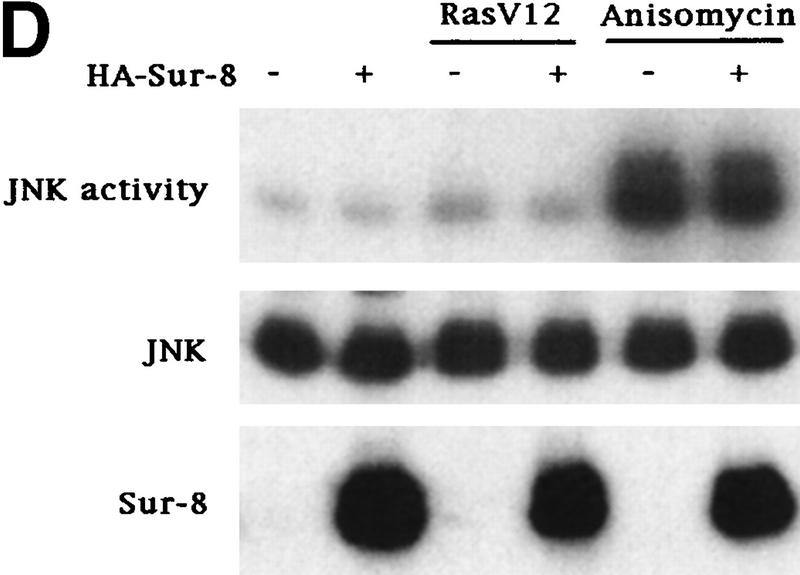
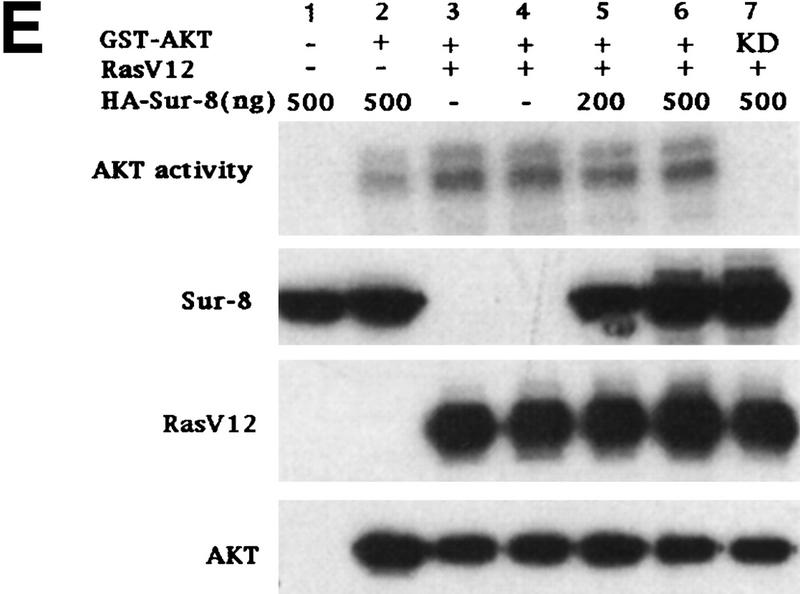
Sur-8 specifically enhances the ERK pathway and functions upstream of Raf. (A) Sur-8 has no effect on MEK-induced ERK activation. The 293 cells were cotransfected with 500 ng of HA–Sur-8, 200 ng of Myc–ERK and varied amounts of a constitutively active MEK mutant as indicated. ERK activity was determined as in Fig. 1A. (B) Sur-8 has no effect on Raf-induced ERK activation. Myc–ERK (200 ng) was cotransfected with V-Raf (10 ng, left) or varied amounts of active B-Raf mutant (right). Cotransfection of Sur-8 did not further increase ERK activity. (C) Sur-8 enhances Raf activity in response to RasV12 or EGF stimulation. 293 cells were cotransfected with 400 ng of Flag–Raf, 20 ng of HA–RasV12, and varied amounts of HA–Sur-8 as indicated. Cells were stimulated with EGF for 3 min where indicated. Flag–Raf was immunoprecipitated with anti-Flag and activity was determined by the coupled kinase assay. Expression of Sur-8 and Ras in lysates and the levels of Raf in the immunoprecipitates were confirmed by Western blot. (D) Sur-8 has no effect on JNK activation. HA–JNK (300 ng) was transfected into cells. Cotransfection with RasV12 or treatment with anisomycin was used to stimulate JNK activity. HA–JNK was immunoprecipitated and assayed for kinase activity using GST–Jun as a substrate. (E) RasV12-induced AKT activation is not enhanced by Sur-8. Wild-type GST–AKT (100 ng) was cotransfected into cells with HA–RasV12 and varied amounts of Sur-8 as indicated. GST–AKT were purified and kinase activity was determined using histone 2B as a substrate. (Lane 7) Negative control of the kinase dead mutant (KD) of GST–AKT.
Sur-8 does not enhance activation of AKT or other MAPK by RasV12
Mammalian cells contain several MAPK pathways. To test the effect of Sur-8 on other MAPK, we determined JNK activity when Sur-8 was coexpressed. JNK is strongly activated by environmental stress such as anisomycin but is weakly activated by Ras (Fig. 2D; Derijard et al. 1994; Kyriakis et al. 1994). JNK activation induced by RasV12 was not enhanced by Sur-8 cotransfection (Fig. 2D). Similarly, the activation of JNK by anisomycin was not augmented by Sur-8 cotransfection. Moreover, Sur-8 had no effect on p38 MAPK activation (data not shown).
In addition to the Raf–ERK pathway, Ras activates multiple downstream targets (Vojtek and Der 1998). For example, Ras activates PI3K, thereby leading to activation of the protein kinase AKT (Marte and Downward 1997). Coexpression of Sur-8 had no significant effect on Ras-induced AKT activity (Fig. 2E). These results further demonstrate that Sur-8 specifically enhances the Ras–ERK pathway.
Sur-8 interacts with Ras and Raf
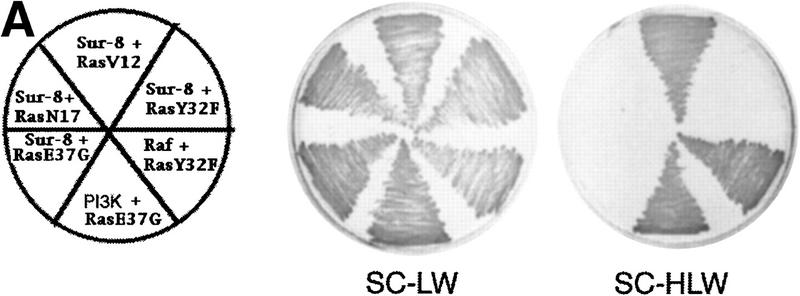
Previous data have indicated that Sur-8 has strong interaction with K-Ras and N-Ras and weak interaction with H-Ras (Sieburth et al. 1998). We further tested the interaction between Sur-8 and Ras or other GTPases (Table 1). Sur-8 interacts with all three Ras isoforms by the yeast two-hybrid assays (Table 1) and coimmunoprecipitations (data not shown). In addition, Sur-8 interacts with the activated RasV12 mutants but not the dominant-negative RasN17 mutants. Ras interacts with downstream targets via its effector domain. We tested the interaction between Sur-8 and several Ras effector domain mutants. Mutations of Y32F, P34G, T35S, V45E, and A59E in RasV12 eliminated its interaction with Sur-8 (Table 1, an example is shown in Fig. 3A) without affecting Raf binding. Mutation of E37G in Ras abolished binding to both Sur-8 and Raf. Many Ras targets, including Raf and RalGDS, are also able to interact with Rap (Macara et al. 1996; Vojtek and Der 1998), a closely related member of the Ras family GTPases. Sur-8 showed no interaction with Rap,TC21, which interacted with Raf in the same assay, or Rac1. Our results demonstrate that Sur-8 specifically interacts with Ras but not with other related small GTPases and mutations of the Ras effector domain abolish its interaction with Sur-8. Furthermore, only the active mutant RasV12 but not the dominant-negative mutant RasN17 shows a positive interaction with Sur-8.
Table 1.
Interaction between Sur-8 and Ras mutants in the yeast two-hybrid system
| Target bait
|
Sur-8
|
Raf
|
PI3K
|
RalGDS
|
|---|---|---|---|---|
| K-RasV12 | ++++ | ++++ | ++++ | ++++ |
| K-RasN17 | − | − | − | − |
| K-RasV12Y32F | − | ++++ | − | − |
| K-RasV12P34G | − | ++++ | − | − |
| K-RasV12T35S | − | ++++ | − | − |
| K-RasV12E37G | − | − | ++++ | |
| K-RasV12V45E | − | ++++ | ++++ | |
| K-RasV12A59E | − | ++++ | − | |
| H-RasV12 | +++ | ++++ | ++++ | ++++ |
| H-RasN17 | − | − | − | − |
| N-RasV12 | ++ | ++++ | ++++ | ++++ |
| N-RasN17 | − | − | − | − |
| RapV12 | − | +++ | +++ | |
| RacV12 | − | − | ||
| TC21 | − | +++ | ||
| Sur-8 | − | − | ||
| Lamin | − | − |
(Blankspace) Experiments not performed.
Figure 3.
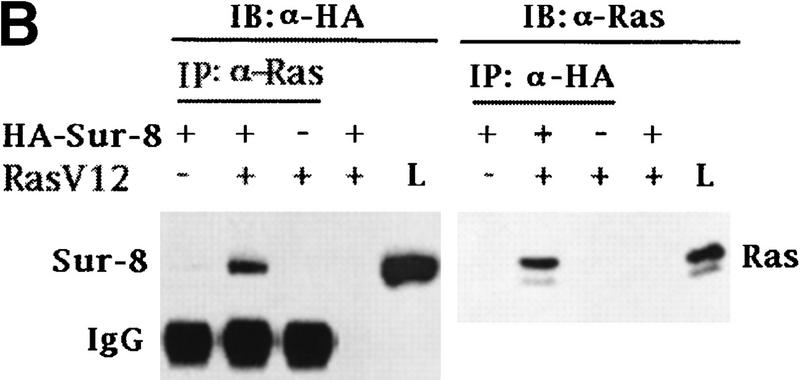
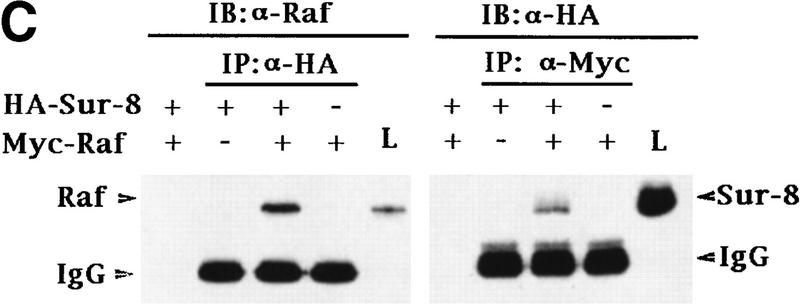
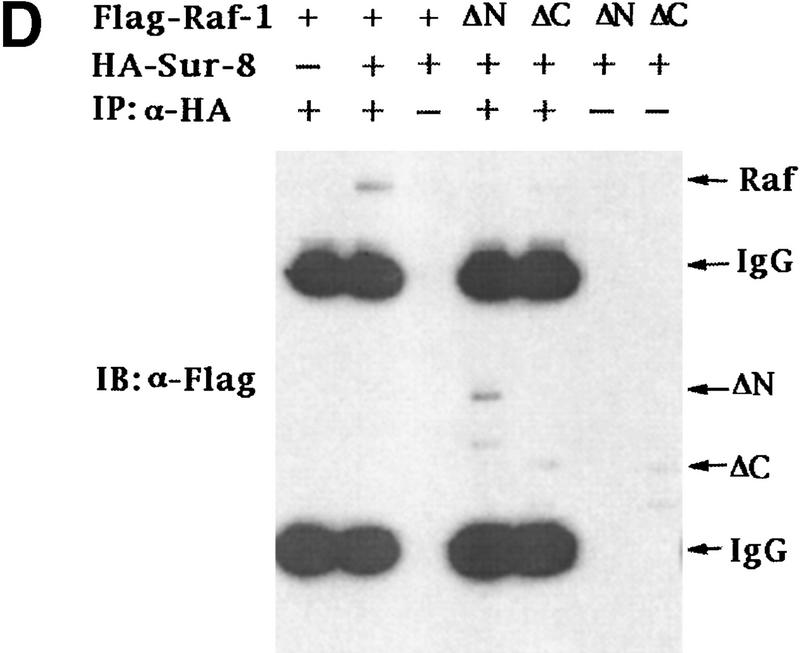
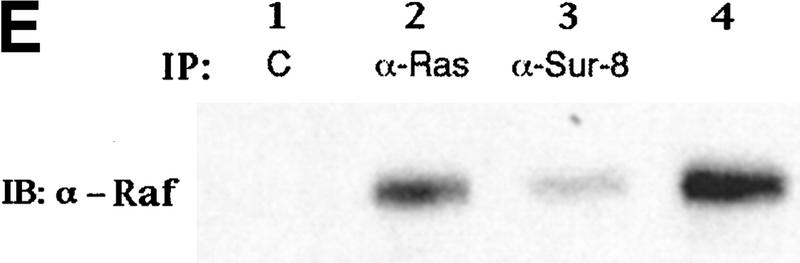
Sur-8 interacts with Ras and Raf. (A) Interactions between Sur-8 and Ras mutants in the yeast two-hybrid assays. Growth on SC-HLW medium indicates a positive interaction. RasY32F and RasE37G are in the RasV12 mutant background. (B) Coimmunoprecipitation between Sur-8 and RasV12. A total of 500 ng of HA–Sur-8 was cotransfected into 293 cells with 200 ng of RasV12 as indicated. Thirty-six hours after transfection, cells lysates were immunoprecipitated with anti-Ras antibody (left) or anti-HA (right) as indicated. The immunoprecipitates were blotted with anti-HA (left) or anti-Ras (right). (Lane L) Cell lysate controls. (IP, IB) Immunoprecipitation and immunoblot, respectively. (IgG) The immunoglobulin band. (C) Interaction between Sur-8 and Raf were determined by coimmunoprecipitation of transfected HA–Sur-8 and Myc–Raf. (Lane L) The cell lysate control. (D) The carboxy-terminal domain of Raf interacts with Sur-8. Flag-tagged Raf full length, ΔN (residues 325–648), ΔC (residues 1–256) were cotransfected with HA–Sur-8. HA–Sur-8 was immunoprecipitated by anti-HA followed by Western blot with anti-Flag antibody. Negative controls without anti-HA antibody were included. Bands corresponding to full length, ΔC, and ΔN of Raf are indicated. (E) Coimmunoprecipitation between endogenous Sur-8 and Raf. The 293 cell lysates were immunoprecipitated with control serum (lane 1), anti-Ras (lane 2), or anti-Sur-8. The immunoprecipitates were blotted with anti-Raf. (Lane 4) Control of cell lysates.
The in vivo interaction between Sur-8 and Ras was tested by coimmunoprecipitation experiments. When cotransfected into 293 cells, HA–Sur-8 and Ras were reciprocally coimmunoprecipitated (Fig. 3B).
Our biochemical data indicate that Sur-8 might function between Ras and Raf, although no positive interaction between Raf and Sur-8 was observed by the yeast two-hybrid assay (data not shown). We tested the possibility that Sur-8 may interact with Raf and other molecules acting downstream of Ras, and found that HA–Sur-8 was coimmunoprecipitated with Raf (Fig. 3C). Reciprocal immunoprecipitation confirmed the interaction between Sur-8 and Raf. Moreover, endogenous Raf and Sur-8 proteins interact in vivo. Anti-Sur-8 antibody coimmunoprecipitated endogenous Raf (Fig. 3E). As a control, anti-Ras antibody also coimmunoprecipitated Raf, whereas the control antibody did not. Raf consists of an amino-terminal domain, which is responsible for interaction with Ras (Vojtek et al. 1993), and a carboxy-terminal kinase domain. The carboxy-terminal domain (Δ) but not the amino-terminal domain (ΔC) of Raf retained a positive interaction with Sur-8 (Fig. 3D). These results show that different Raf domains are responsible for interacting with Ras and Sur-8. However, our data cannot exclude the possibility that other proteins mediate the interaction between Sur-8 and Raf.
Sur-8 can form a complex with Ras and Raf
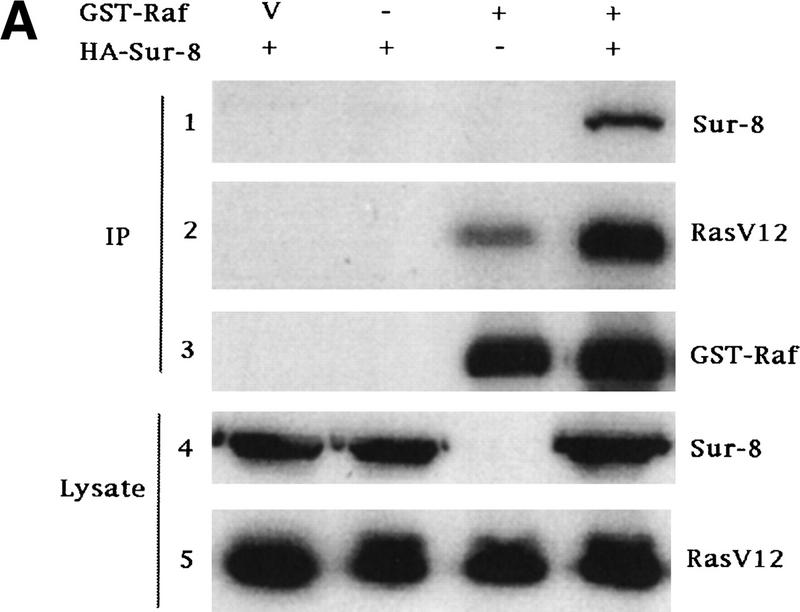
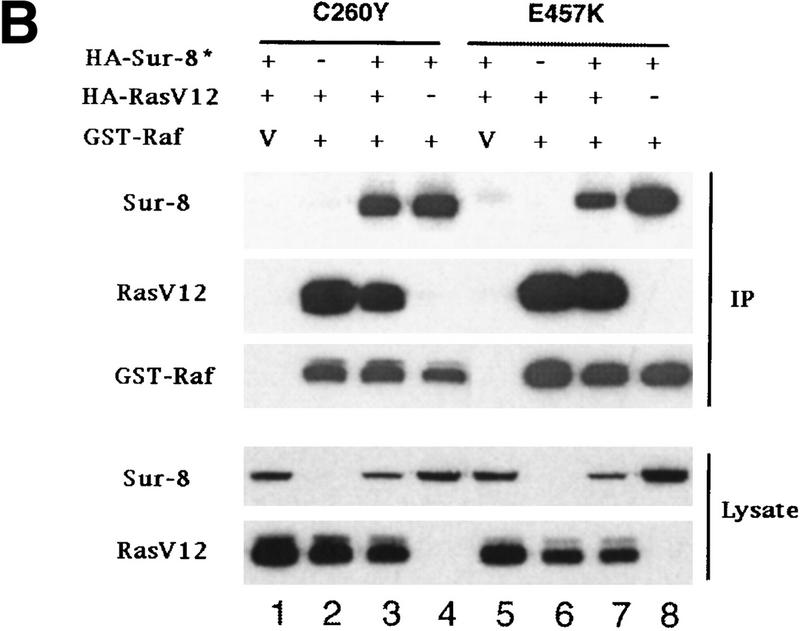
The above results demonstrate that Ras, Raf, and Sur-8 can interact with each other. A possible mechanism by which Sur-8 stimulates Raf activation is by promoting Ras–Raf complex formation. To test this hypothesis, Ras-Raf interaction was examined in the presence or absence of Sur-8 coexpression. Sur-8 significantly increased the amount of RasV12 copurified with GST–Raf (Fig. 4A). However, neither Sur-8C260Y nor E457K mutant can enhance the interaction between Ras and GST–Raf (Fig. 4B). These results suggest that the ability of Sur-8 to enhance Ras–Raf interaction correlates with its activity to enhance Ras signaling.
Figure 4.
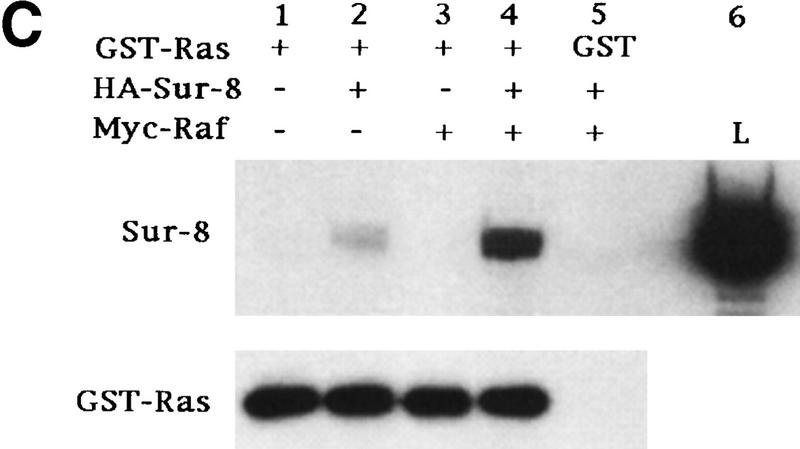
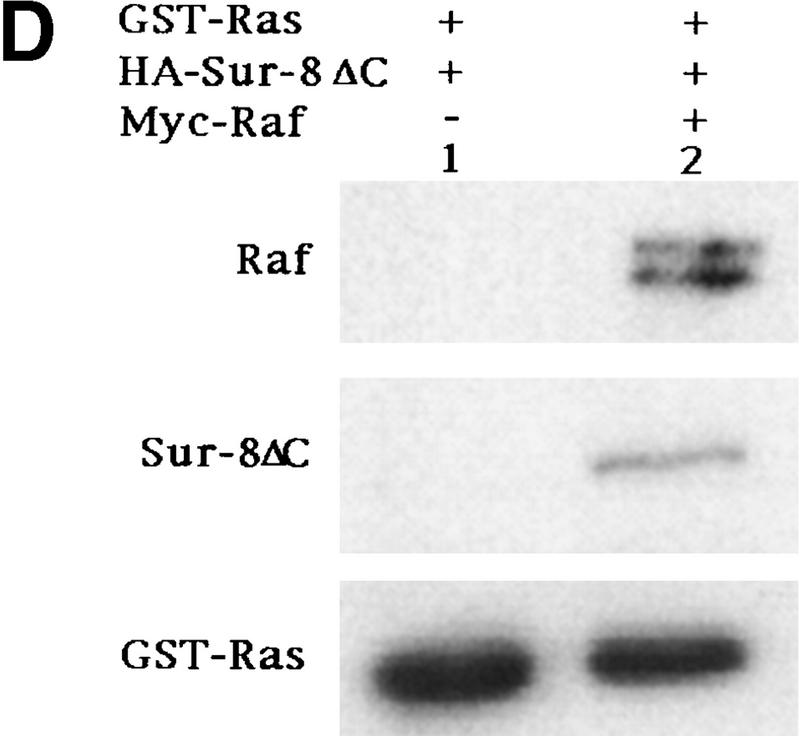
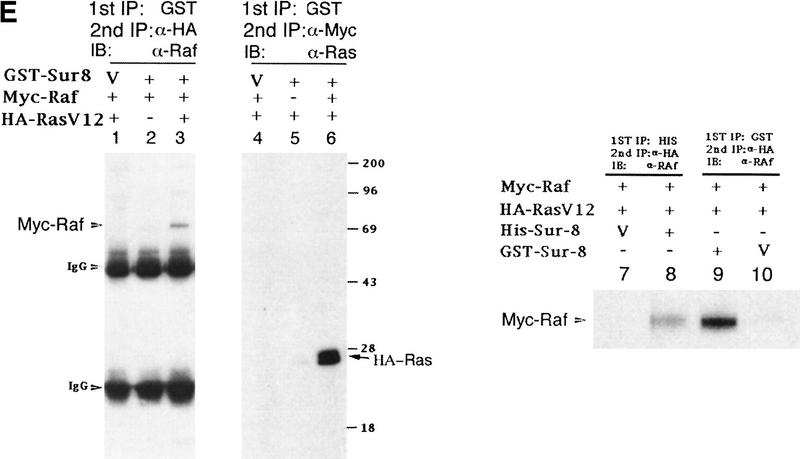
Sur-8 forms a complex with Ras and Raf. (A) Sur-8 enhances the interaction between Ras and Raf. HA–RasV12 was cotransfected with GST–Raf in the presence or absence of HA–Sur-8. GST–Raf was purified by glutathione–agarose resin. The copurified Ras and HA–Sur-8 were detected by anti-HA for HA–Ras (second panel from top) and HA–Sur-8 (top) Western blots. The GST–Raf in the precipitation was detected by an anti-Raf Western blot (middle). Expressions of HA–Sur-8 and HA–RasV12 in cell lysates were shown in bottom two panels. (B) Sur-8C260Y and E457K mutants fail to enhance the interaction between Ras and Raf. Experiments are similar to A. (HA–Sur-8*) Sur-8C260Y in lanes 1–4 and Sur-8E457K in lanes 5–8, respectively. (C) Raf enhances Sur–8-Ras interaction in vitro. Cell lysates expressing HA–Sur-8 or Myc–Raf were mixed with purified GST–RasV12 protein (5 μg). GST–RasV12 and associated proteins were purified by glutathione–agarose and Western blotted with anti-HA to detect HA–Sur-8 (top) and anti-GST to detect GST–RasV12 (bottom). (Lane 6) A control of cell lysate containing HA–Sur-8. (D) Raf bridges the interaction between Ras and Sur-8ΔC. GST–Ras (5 μg) was incubated with cell lysates containing HA–Sur-8ΔC (containing residues 1–366) in the absence (lane 1) or presence (lane 2) of Raf containing cell lysates. HA–Sur-8ΔC (middle) and Raf (top) copurified by GST–Ras were detected by Western blot. GST–Ras coprecipitated HA–Sur-8ΔC only in the presence of Raf. (E) Sur-8, Ras, and Raf form a ternary complex. The 293 cells were transfected with various plasmids as indicated. Cell lysates were subjected to purification by glutathione–agarose beads. The purified proteins were eluted by glutathione and subjected to immunoprecipitation with anti-HA for HA–Ras followed by Western blot with anti-Raf (left). Similarly, proteins eluted from glutathione–agarose beads were immunoprecipitated with anti-Myc for Myc–Raf followed by Western blot with anti-Ras (middle). (Right) Experimental results using His–Sur-8 (lane 8) along with GST–Sur-8 (lane 9). His–Sur-8 complex was first purified by nickel agarose and eluted with imidazole. The eluted samples were immunoprecipitated with anti-Myc and followed by Western blot with anti-Raf.
To further confirm the formation of Ras–Raf–Sur-8 complex, we performed in vitro binding experiments using purified GST–Ras and cell lysates containing Raf and Sur-8. Addition of Raf enhanced the association between GST–Ras and Sur-8 (Fig. 4C). The carboxy-terminal domain of Sur-8 is required for interaction with Ras but not required for Sur-8–Raf interaction (Sieburth et al. 1998) (data not shown). A Sur-8ΔC mutant showed no interaction with Ras (Fig. 4D, lane 1). Addition of Raf enhanced the interaction between Sur-8ΔC and Ras, suggesting that Raf is capable of bridging this interaction (Fig. 4D).
We further characterized the formation of a Ras, Raf, and Sur-8 complex in vivo by sequential coimmunoprecipitation experiments. GST–Sur-8, HA–RasV12, and Myc–Raf were cotransfected into 293 cells. GST–Sur-8 and its associated proteins were purified, eluted with glutathione, and subjected to a second immunoprecipitation with anti-HA antibodies. The anti-HA immunoprecipitates were then subjected to Western blot with anti-Raf to detect Myc–Raf. Myc–Raf was present after sequential immunoprecipitation (Fig. 4E, lane 3). Experiments were also performed first by purification of GST–Sur-8, followed by immunoprecipitation with anti-Myc to precipitate Myc–Raf, then blotted with anti-Ras to detect HA–Ras. Ras is present in the sample after sequential purification with GST–Sur-8 and Myc-Raf (Fig. 4E, lane 6). GST–Sur-8 may form a dimer and could complicate the interpretation of the above results. Similar experiments using His–Sur-8 demonstrated that Raf protein was present after the sequential precipitations with nickel agarose (for His–Sur-8) and anti-HA (for HA–Ras) (Fig. 4E), consistent with experiments using GST–Sur-8. These data provide strong evidence that Ras, Raf, and Sur-8 together can form a ternary complex in vivo.
Discussion
Here we demonstrate that human Sur-8 can form a complex with Ras and Raf, and specifically enhances ERK activation. Our observations support the model that Sur-8 acts between Ras and Raf. Sur-8 alone does not significantly stimulate the ERK activity in the absence of EGF or RasV12. The effect of Sur-8 is most significant at low concentrations of EGF or Ras. Therefore, Sur-8 enhances the signal strength of RTK and Ras by facilitating the Ras–Raf complex formation. Genetic properties of sur-8 in C. elegans are consistent with this notion. sur-8 mutations alone do not cause obvious mutant phenotypes, but they are effective suppressors of activated Ras mutations and drastically enhance weak mutations in MAPK or Raf, indicating that Sur-8 is required for optimal Ras–MAP kinase signaling (Sieburth et al. 1998). Our data demonstrate that Sur-8 may modulate the signal strength of Ras and RTK. Sur-8 may also modulate the specificity of physiological responses. For example, Sur-8 selectively increases the signal from RTK or Ras to Raf but not to other downstream effectors such as PI3K.
The Ras–MAPK pathway represents one of the best-characterized signal transduction pathways. However, the molecular mechanism of Raf activation is not completely understood (Morrison and Cutler 1997). Ras–Raf interaction is important for Raf activation, but such interaction alone is not sufficient to activate Raf in vitro. Raf activation requires additional events, such as stimulatory/inhibitory phosphorylation and association of other molecules (Morrison and Cutler 1997). Phosphorylation plays a key role in Raf activation. However, the kinase responsible for Raf activation in response to Ras or growth factors is largely unknown. Both PKC and Pak3 have been implicated to phosphorylate and activate Raf (Kolch et al. 1993; King et al. 1998). It is possible that Sur-8 may recruit a Raf-activating kinase to the complex, thereby facilitating Raf-activation. Recent genetic and biochemical data indicate that phosphorylation could also lead to Raf inactivation (Morrison and Cutler 1997; Sieburth et al. 1999; Zimmermann and Moelling 1999). Thus, it is equally possible that Sur-8 may recruit a protein phosphatase, such as PP2A (Sieburth et al. 1999), to the complex and facilitate the specific dephosphorylation of the inhibitory sites in Raf. Future biochemical studies of Sur-8 functions will provide new insights of the Ras signal transduction.
Materials and methods
Cell cultures and transfections
Human 293 cells were cultured in DMEM supplemented with 10% (vol/vol) FBS. Transfections of 293 cells were done by LipofectAmine (GIBCO BRL) methods in six-well plates.
Plasmid constructions
H-Ras, K-Ras, and N-Ras cDNA were subcloned into the vector pcDNA3HA or pCDNA3. GST-tagged constructs, GST–Ras, GST–Raf, and GST–Sur-8 were obtained by subcloning into bacterial-expressing vector pGEX-KG (Guan and Dixon 1991) or mammalian-expressing vector pEBG-3X. JNK/pcDNA3HA, Supmek/pcDNA3, Raf/pRK5 Myc, pcDNA3 Flag, and Myc-tagged ERK1, Gal4–Elk-1, Gal4–LUC, and CMV–lacZ were laboratory stocks. Constitutively active HA–AKT was obtained from Dr. Anne Vojtek (University of Michigan, Ann Arbor). Constitutively active HA–B-Raf/pcDNA3HA was created by site-directed mutagenesis in which Ser-364, Thr-428, and Ser-439 were replaced by alanine residues (A. Vojtek, unpubl.). Human Sur-8 cDNA was reported previously (Sieburth et al. 1998). Point mutations and deletions of Ras, Sur-8, and Raf were created by site-directed mutagenesis or PCR, and were confirmed by DNA sequencing. The plasmids used for yeast two-hybrid interaction were in pVP16 or plexA-Ade vector as targets or baits (Vojtek and Hollenberg 1995), respectively.
Kinase assays
Myc–ERK1 was transfected into 293 cells. Twenty-four hours after transfections, cells were starved in serum-free medium for 12 hr. Cells were lysed in lysis buffer (10 mm Tris -HCl at pH 7.5, 2 mm EDTA, 150 mm NaCl, 1% NP-40, 50 mm NaF, 1 mm sodium vanadate, 1 mm PMSF, 2 μg/ml aprotinin, 2 μg/ml leupeptin). A total of 1 μg of 9E10 antibody (anti-Myc) was added in each immunoprecipitation reaction. Kinase activities of ERK were measured using GST–Elk-1 as a substrate following the previously described method (Zheng and Guan 1994).
Raf-1 kinase activity was assessed by the coupled MEK/ERK activation assays according to previously described methods (Wu et al. 1996). Briefly, Flag-Raf was immunoprecipitated and used to activate 0.1 μg of GST–MEK1. The activated GST–MEK1 was removed from the protein G-bound Raf and used to activate 0.1 μg of GST–ERK1. Phosphorylation of GST–Elk-1 by activated GST–ERK1 was determined.
HA–JNK kinase protein was immunoprecipitated by α-HA antibody. The procedure used to assay JNK activity was basically the same as ERK activity assay, except that GST–Jun was used as a substrate.
HA–AKT was transfected into 293 cells together with RasV12 and Sur-8 plasmids. About 24 hr after transfection, cells were starved in serum-free DMEM overnight. Cells were then lysed in lysis buffer (20 mm Tris at pH 7.5, 150 mm NaCl, 10% glycerol, 1% Nonidet P-40, 10 mm NaF, 1 mm sodium Vanadate, 1 mm Na4O7P2, 1 mm PMSF, 2 μg/ml aprotinin, 2 μg/ml leupeptin). Immunoprecipitation was performed by addition of monoclonal anti-HA antibody (1–2 μg). Immune complexes were collected with protein A agarose and washed three times with lysis buffer and once with kinase buffer without ATP (20 mm HEPES at pH 7.4, 10 mm MgCl2, 1 mm DTT). Kinase reaction was performed in 20 μl of kinase reaction 50 μm ATP (with 1.5 μm Ci[32P]ATP) and 2 μg of histone 2B as a substrate. Reaction was incubated at 30°C for 20 min and followed by analysis on SDS-PAGE and autoradiography.
Coimmunoprecipitation and Western blot
Transfected 293 cells were lysed in IP buffer containing protease inhibitors (10 mm Tris-HCl at pH 7.5, 5 mm EGTA, 150 mm NaCl, 1% NP-40, 20 mm NaF, 1 mm Sodium Vanadate, 1 mm PMSF, 10 μg/ml aprotinin, 10 μg/ml leupeptin). Cell extracts were incubated with antibody for 2 hr at 4°C. Protein A or G beads were added in IP reactions and incubated for an additional hour. Immunoprecipitates were collected by centrifugation, washed four times with IP buffer, and analyzed by Western blot. For GST fusion proteins, cell extracts were incubated with glutathione–agarose beads directly for 1–2 hr at 4°C. The beads were then washed four times with lysis buffer, and eluted with 5 mm glutathione. The eluate was analyzed by immunoblot using indicated antibodies. For coimmunoprecipitation of endogenous proteins, exponentially growing 293 cells were used. The anti-Sur-8 antiserum was a generous gift of Dr. Mike Stern (Yale University, New Haven, CT) (Selfors et al. 1998).
For analysis of complex formation, HA–RasV12, Myc–Raf, and GST–Sur-8 were cotransfected into 293 cells. pEBG vector was cotransfected with HA–RasV12 and Myc–Raf as a control. Cell extracts were incubated with glutathione–agarose beads. GST fusion and associated proteins were eluted by 5 mm glutathione and subjected to immunoprecipitation with either anti-HA or anti-Myc antibodies. For experiments using His–Sur-8, nickel agarose was used in the first precipitation. The bound proteins were eluted with 50 mm imidazole, followed by immunoprecipitation with anti-HA antibody (for HA–Ras). Finally, the immunoprecipitated samples were analyzed by Western blot against either anti-Ras or anti-Raf antibody.
In vitro binding
In vitro binding was performed in binding buffer [50 mm Tris-HCl at pH 7.5, 150 mm NaCl, 1.0 mm DTT, 1% Triton X-100, 5 mm MgCl2, 25 μm ZnCl2, and 0.2% (wt/vol) BSA]. Purified GST-fusion proteins or cell lysates containing GST-fusion proteins were incubated with cell lysates expressing proteins of interest. The mixture was incubated for ∼6 hr and then incubated with glutathione–agarose for 1–2 hr. The beads were washed four times with binding buffer, once with binding buffer without BSA, and then eluted with 5 mm glutathione and analyzed by SDS-PAGE and Western blot.
Acknowledgments
We thank Drs. Derek Sieburth and Qun Sun for the Sur-8 plasmids, Kevin Pumiglia for Raf-1 plasmids, and M. Stern for anti-soc-2 antibodies. This work was supported by grants from National Institutes of Health (K.L.G. and M.H.) and the MacArthur Fellowship (K.L.G.).
The publication costs of this article were defrayed in part by payment of page charges. This article must therefore be hereby marked “advertisement” in accordance with 18 USC section 1734 solely to indicate this fact.
Footnotes
E-MAIL kunliang@umich.edu; FAX (734) 763-4581.
References
- Chang HC, Karim FD, O'Neill EM, Rebay I, Solomon NM, Therrien M, Wassarman DA, Wolff T, Rubin GM. Ras signal transduction pathway in Drosophila eye development. Cold Spring Harb Symp Quant Biol. 1994;59:147–153. doi: 10.1101/sqb.1994.059.01.018. [DOI] [PubMed] [Google Scholar]
- Derijard B, Hibi M, Wu IH, Barrett T, Su B, Deng T, Karin M, Davis RJ. JNK1: A protein kinase stimulated by UV light and Ha-Ras that binds and phosphorylates the c-Jun activation domain. Cell. 1994;76:1025–1037. doi: 10.1016/0092-8674(94)90380-8. [DOI] [PubMed] [Google Scholar]
- Field J, Xu HP, Michaeli T, Ballester R, Sass P, Wigler M, Colicelli J. Mutations of the adenylyl cyclase gene that block RAS function in Saccharomyces cerevisiae. Science. 1990;247:464–467. doi: 10.1126/science.2405488. [DOI] [PubMed] [Google Scholar]
- Guan KL, Dixon JE. Eukaryotic proteins expressed in Escherichia coli: An improved thrombin cleavage and purification procedure of fusion proteins with glutathione S-transferase. Anal Biochem. 1991;192:262–267. doi: 10.1016/0003-2697(91)90534-z. [DOI] [PubMed] [Google Scholar]
- Howe LR, Leevers SJ, Gomez N, Nakielny S, Cohen P, Marshall CJ. Activation of the MAP kinase pathway by the protein kinase raf. Cell. 1992;71:335–342. doi: 10.1016/0092-8674(92)90361-f. [DOI] [PubMed] [Google Scholar]
- Kataoka T, Broek D, Wigler M. DNA sequence and characterization of the S. cerevisiae gene encoding adenylate cyclase. Cell. 1985;43:493–505. doi: 10.1016/0092-8674(85)90179-5. [DOI] [PubMed] [Google Scholar]
- Katz ME, McCormick F. Signal transduction from multiple Ras effectors. Curr Opin Genet Dev. 1997;7:75–79. doi: 10.1016/s0959-437x(97)80112-8. [DOI] [PubMed] [Google Scholar]
- King AJ, Sun H, Diaz B, Barnard D, Miao W, Bagrodia S, Marshall MS. The protein kinase Pak3 positively regulates Raf-1 activity through phosphorylation of serine 338. Nature. 1998;396:180–183. doi: 10.1038/24184. [DOI] [PubMed] [Google Scholar]
- Kobe B, Deisenhofer J. Crystal structure of porcine ribonuclease inhibitor, a protein with leucine-rich repeats. Nature. 1993;366:751–756. doi: 10.1038/366751a0. [DOI] [PubMed] [Google Scholar]
- Kolch W, Heidecker G, Kochs G, Hummel R, Vahidi H, Mischak H, Finkenzeller G, Marme D, Rapp UR. Protein kinase C alpha activates RAF-1 by direct phosphorylation. Nature. 1993;364:249–252. doi: 10.1038/364249a0. [DOI] [PubMed] [Google Scholar]
- Kornfeld K, Hom DB, Horvitz HR. The ksr-1 gene encodes a novel protein kinase involved in Ras-mediated signaling in C. elegans. Cell. 1995;83:903–913. doi: 10.1016/0092-8674(95)90206-6. [DOI] [PubMed] [Google Scholar]
- Kyriakis JM, App H, Zhang XF, Banerjee P, Brautigan DL, Rapp UR, Avruch J. Raf-1 activates MAP kinase-kinase. Nature. 1992;358:417–421. doi: 10.1038/358417a0. [DOI] [PubMed] [Google Scholar]
- Kyriakis JM, Banerjee P, Nikolakaki E, Dai T, Rubie EA, Ahmad MF, Avruch J, Woodgett JR. The stress-activated protein kinase subfamily of c-Jun kinases. Nature. 1994;369:156–160. doi: 10.1038/369156a0. [DOI] [PubMed] [Google Scholar]
- Macara IG, Lounsbury KM, Richards SA, McKiernan C, Bar-Sagi D. The Ras superfamily of GTPases. FASEB J. 1996;10:625–630. doi: 10.1096/fasebj.10.5.8621061. [DOI] [PubMed] [Google Scholar]
- Mansour SJ, Matten WT, Hermann AS, Candia JM, Rong S, Fukasawa K, Vande Woude GF, Ahn NG. Transformation of mammalian cells by constitutively active MAP kinase kinase. Science. 1994;265:966–970. doi: 10.1126/science.8052857. [DOI] [PubMed] [Google Scholar]
- Marte BM, Downward J. PKB/Akt: Connecting phosphoinositide 3-kinase to cell survival and beyond. Trends Biochem Sci. 1997;22:355–358. doi: 10.1016/s0968-0004(97)01097-9. [DOI] [PubMed] [Google Scholar]
- Morrison DK, Cutler RE. The complexity of Raf-1 regulation. Curr Opin Cell Biol. 1997;9:174–179. doi: 10.1016/s0955-0674(97)80060-9. [DOI] [PubMed] [Google Scholar]
- Robinson MJ, Cobb MH. Mitogen-activated protein kinase pathways. Curr Opin Cell Biol. 1997;9:180–186. doi: 10.1016/s0955-0674(97)80061-0. [DOI] [PubMed] [Google Scholar]
- Rodriguez-Viciana P, Warne PH, Dhand R, Vanhaesebroeck B, Gout I, Fry MJ, Waterfield MD, Downward J. Phosphatidylinositol-3-OH kinase as a direct target of Ras. Nature. 1994;370:527–532. doi: 10.1038/370527a0. [DOI] [PubMed] [Google Scholar]
- Selfors LM, Schutzman JL, Borland CZ, Stern MJ. soc-2 encodes a leucine-rich repeat protein implicated in fibroblast growth factor receptor signaling. Proc Natl Acad Sci. 1998;95:6903–6908. doi: 10.1073/pnas.95.12.6903. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sieburth DS, Sun Q, Han M. SUR-8, a conserved Ras-binding protein with leucine-rich repeats, positively regulates Ras-mediated signaling in C. elegans. Cell. 1998;94:119–130. doi: 10.1016/s0092-8674(00)81227-1. [DOI] [PubMed] [Google Scholar]
- Sieburth DS, Sundaram M, Howard RM, Han M. A PP2A regulatory subunit positively regulates Ras-mediated signaling during Caenorhabditis elegans vulval induction. Genes & Dev. 1999;13:2562–2569. doi: 10.1101/gad.13.19.2562. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sternberg PW, Han M. Genetics of RAS signaling in C. elegans. Trends Genet. 1998;14:466–472. doi: 10.1016/s0168-9525(98)01592-3. [DOI] [PubMed] [Google Scholar]
- Sundaram M, Han M. The C. elegans ksr-1 gene encodes a novel Raf-related kinase involved in Ras-mediated signal transduction. Cell. 1995;83:889–901. doi: 10.1016/0092-8674(95)90205-8. [DOI] [PubMed] [Google Scholar]
- Suzuki N, Choe HR, Nishida Y, Yamawaki-Kataoka Y, Ohnishi S, Tamaoki T, Kataoka T. Leucine-rich repeats and carboxyl terminus are required for interaction of yeast adenylate cyclase with RAS proteins. Proc Natl Acad Sci. 1990;87:8711–8715. doi: 10.1073/pnas.87.22.8711. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Urano T, Emkey R, Feig LA. Ral-GTPases mediate a distinct downstream signaling pathway from Ras that facilitates cellular transformation. EMBO J. 1996;15:810–816. [PMC free article] [PubMed] [Google Scholar]
- van der Geer P, Hunter T, Lindberg RA. Receptor protein-tyrosine kinases and their signal transduction pathways. Annu Rev Cell Biol. 1994;10:251–337. doi: 10.1146/annurev.cb.10.110194.001343. [DOI] [PubMed] [Google Scholar]
- Vojtek AB, Der CJ. Increasing complexity of the Ras signaling pathway. J Biol Chem. 1998;273:19925–19928. doi: 10.1074/jbc.273.32.19925. [DOI] [PubMed] [Google Scholar]
- Vojtek AB, Hollenberg SM. Ras-Raf interaction: Two-hybrid analysis. Methods Enzymol. 1995;255:331–342. doi: 10.1016/s0076-6879(95)55036-4. [DOI] [PubMed] [Google Scholar]
- Vojtek AB, Hollenberg SM, Cooper JA. Mammalian Ras interacts directly with the serine/threonine kinase Raf. Cell. 1993;74:205–214. doi: 10.1016/0092-8674(93)90307-c. [DOI] [PubMed] [Google Scholar]
- Wu X, Noh SJ, Zhou G, Dixon JE, Guan KL. Selective activation of MEK1 but not MEK2 by A-Raf from epidermal growth factor-stimulated Hela cells. J Biol Chem. 1996;271:3265–3271. doi: 10.1074/jbc.271.6.3265. [DOI] [PubMed] [Google Scholar]
- Zheng CF, Guan KL. Activation of MEK family kinases requires phosphorylation of two conserved Ser/Thr residues. EMBO J. 1994;13:1123–1131. doi: 10.1002/j.1460-2075.1994.tb06361.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zimmermann S, Moelling K. Phosphorylation and regulation of Raf by Akt (protein kinase B) Science. 1999;286:1741–1744. doi: 10.1126/science.286.5445.1741. [DOI] [PubMed] [Google Scholar]