Abstract
The crustacean sinus gland (SG) is a well-defined neuroendocrine site that produces numerous hemolymph-borne agents including the most complex class of endocrine signaling molecules—neuropeptides. Via a multifaceted mass spectrometry (MS) approach, 70 neuropeptides were identified including orcokinins, orcomyotropin, crustacean hyperglycemic hormone (CHH) precursor-related peptides (CPRPs), red pigment concentrating hormone (RPCH), pigment dispersing hormone (PDH), proctolin, RFamides, RYamides, and HL/IGSL/IYRamide. Among them, 15 novel orcokinins, 9 novel CPRPs, one novel orcomyotropin, one novel Ork/Orcomyotropin-related and one novel PDH were de novo sequenced via collision induced dissociation (CID) from the SG of a model organism Callinectes sapidus. Electron transfer dissociation (ETD) was used for sequencing of intact CPRPs due to their large size and charge state. Capillary isoelectric focusing (CIEF) was employed for separation of members of the orcokinin family which is one of the most abundant neuropeptide families observed in the SG. Collectively, our study represents the most complete characterization of neuropeptides of the SG and provides a foundation for future investigation of the physiological function of neuropeptides in the SG of C. sapidus.
Keywords: De novo sequencing, Callinectes sapidus, sinus glands, mass spectrometry, neuropeptides, CPRP, Orcokinins, capillary isoelectric focusing, CIEF, ETD, CID
Introduction
Neuropeptides are the largest and most diverse group of endocrine signaling molecules in the nervous system. These signaling peptides are critical for initiation and regulation of numerous physiological processes such as feeding, reproduction and development.1, 2 Mass spectrometry (MS), with unique advantages such as high sensitivity, high throughput, chemical specificity and the capability of de novo sequencing with limited genomic information, is well suited for the detection and sequencing of neuropeptides in endocrine glands, and simultaneously provides the potential for information on post-translational modifications.3–6
The sinus glands (SG) are located between the medulla interna and medulla externa of the eyestalk in C. sapidus. It is a well-known neuroendocrine site that synthesizes and secretes peptide hormones regulating various physiological activities such as molting, hemolymph glucose levels, integument color changes, contraction on visceral muscles, eye pigment movements and hydro-mineral balance.7 Our previous neuropeptidomic studies of SG in several crustacean species including Cancer borealis,8–11 Carcinus maenas,12 and Homarus americanus13, 14 used multiple MS strategies combining MALDI-based high resolution accurate mass profiling, biochemical derivatization, and nanoscale separation coupled to tandem MS, for de novo sequencing. In addition, several previous studies reported a limited number of neuropeptides in C. sapidus.15–17 In the current study, we explore the neuropeptidome of the SG in the blue crab C. sapidus, a vital species of economic importance on the seafood market worldwide. In total, 70 neuropeptides are identified including 27 novel neuropeptides, and among them, the crustacean hyperglycemic hormone precursor-related peptides (CPRPs) and orcokinins represent major neuropeptide families known in the SG. Therefore, we focus our effort on the comprehensive characterization of these two highly important classes of signaling peptides in the present study.
Orcokinins were originally purified from the crayfish Orconectes limosus.18 They are comprised of a family of neuropeptides with the conserved N-terminus sequence NFDEIDR and C-terminal variations that can occur at residues eight, nine, or thirteen.19, 20 Orcokinins have been identified in a large number of crustaceans18, 19, 21–27 and several insect species.28–30 These studies indicate that this peptide family has been conserved throughout invertebrates and suggest their potential roles in complex physiological processes. Further physiological studies show the effect of orcokinins in the stomatogastric ganglion of the lobster Homarus americanus23 and their myotropic effects in the hindgut of crayfish O. limosus.18, 31 As one of the most abundant neuropeptides in crustacean SG,8, 21, 27 comprehensive profiling and characterization of the orcokinin family represents an important first step toward a better understanding of the functional roles of this peptide family. However, given their conserved amino acid sequences and similar molecular weights, the large number and chemical complexity of orcokinins in the SG requires the use of highly efficient separation techniques prior to MS detection for comprehensive characterization.
Among the separation techniques which have been coupled with MS, capillary isoelectric focusing (CIEF) is an emerging tool for the focusing and separation of proteins and peptides.32–36 Here, we present a modified CIEF system for neuropeptide analysis by adding two membrane-coated joints near both ends of the capillary to prevent carrier ampholyte band shifts and interferences from gas bubbles to retain high separation efficiency with high sensitivity. The combination of this modified membrane assisted CIEF (MA-CIEF) system with MALDI-FTMS provides a superior analytical platform to enrich and separate orcokinins for efficient detection.
The crustacean hyperglycemic hormone (CHH) and its precursor-related peptide (CPRP) are produced concurrently during the cleavage of CHH from the CHH preprohormone protein.37 The CPRP peptide is localized between the signal peptide and the CHH sequence and is separated from the CHH by a dibasic cleavage site. The functions of CPRPs are currently unknown.38 However, the complete characterization of CPRPs provides a foundation for future experiments elucidating their functional roles in crustaceans. The cDNA of CHH preprohormone in SG of C. sapidus has been characterized,39 but the direct detection of CPRP has not been reported due to its relatively large size and possible post-translational modifications. While small fragments of CPRPs can be identified using nanoLC-ESI-Q-TOF tandem mass spectrometry, the intact peptide is difficult to sequence due to the large molecular weight of CPRPs.
Peptidomics traditionally uses collision induced dissociation (CID) for tandem MS experiments for de novo sequencing. Recently, an alternative peptide fragmentation method has been developed using the transfer of an electron termed electron transfer dissociation (ETD).40, 41 ETD involves a radical anion with low electron affinity to be transferred to peptide cation, which results in more uniform cleavage along the peptide backbone and reduced sequence discrimination and thus provides improved sequencing for larger peptides compared to CID.42 Specifically, for an ion trap ETD, the fluoranthene radical anion is allowed to react with peptide cations in the three dimensional trap. The resulting dissociation cleaves at the N-Cα bond to produce c- and z•-type product ions. The peptidome of model organisms like C. sapidus can be further explored and expanded utilizing ETD as a complementary fragmentation technique to CID. An improved coverage for peptidomic analysis has been reported by using ETD as an additional fragmentation method.43 It was observed that enzymatically produced peptides with a higher m/z produced improved sequence coverage using ETD. This observation, and subsequent development termed decision tree analysis, determined that a charge state of ≥6 all peptides, endogenous or enzymatic, should be fragmented via ETD.44 In the present study, the highly charged peptide CPRP with an intact molecular weight of 3837 Da readily produces plus six charge states, which is highly amenable to fragmentation by ETD and produces remarkably improved fragmentation and thus improved sequence coverage when compared to CID. In this report, we developed a MS-based platform utilizing multiple tandem mass spectrometry techniques and enhanced separation methods to demonstrate the discovery and characterization of CPRP and orcokinin neuropeptides in the sinus glands of the blue crab C. sapidus.
Experimental Section
Chemical and materials
Acetic acid, sodium hydroxide, acetonitrile (ACN) and methanol were purchased from Fisher Scientific (Pittsburgh, PA). Formic acid and hydroxypropyl cellulose (HPC) were from Sigma-Aldrich (St. Louis, MO). 2,5-dihydroxybenzoic acid (DHB) was obtained from Alfa Aesar. (Ward Hill, MA). Parafilm “M” was obtained from Pechiney Plastic Packaging (Menasha, WI). Fused-silica capillary with 75 μm i.d. and 360 μm o.d. was purchased from Polymicro Technologies (Phoenix, AZ). Millipore C18 Ziptip column was used for sample cleaning, and all water used in this study was deionized water (18.2 MΩ·cm) prepared from a Milli-Q Millipore system (Billerica, MA). The physiological saline consisted of 440 mM NaCl, 11mM KCl, 26 mM MgCl2, 13 mM CaCl2, 11 mM Trizma base, and 5 mM maleic acid in pH 7.45.
Animals and dissection
C. sapidus (blue crab) were obtained from commercial food market and maintained without food in artificial sea water at 10–12 ºC. Animals were cold-anesthetized by packing on ice for 15–30min before dissection. The optic ganglia, with the SG attached, was dissected in chilled (~4ºC) physiological saline and individual SGs were isolated from the optic ganglia by micro-dissection and immediately placed in acidified methanol (90% methanol, 9% glacial acetic acid, and 1% water) and stored at −80ºC until tissue extraction.
Direct tissue preparation and tissue homogenization
SGs were dissected and prepared for MALDI MS analysis, followed by rinsing in 10 mg/mL of 2, 5-dihydroxybenzoic acid (DHB), to remove the extracellular salts associated with the tissue sample. Tungsten needles and fine forceps were used to dissect and transfer small pieces of tissue onto a MALDI sample plate followed by the application of 150mg/ml DHB matrix solution (in 50:50 acetone: water, v/v) for MALDI FTMS and 5 mg/ml α-cyano-4-hydroxycinnamic acid (CHCA) for MALDI-TOF/TOF MS. Acidified methanol was used during the homogenization of SGs. The homogenized SGs were immediately centrifuged at 16,100 ×g using an Eppendorf 5415D tabletop centrifuge (Eppendorf AG). The pellet was washed using acidified methanol and combined with the supernatant and further dried in a Savant SC 110 SpeedVac concentrator (Thermo Electron Corporation). The resulting dried sample was re-suspended with a minimum amount (20 μL) of 0.1% formic acid
Fractionation of homogenates using reversed phase (RP)-HPLC
The re-suspended extracts were then vortexed and briefly centrifuged. The resulting supernatants were subsequently fractionated via high performance liquid chromatography (HPLC). HPLC separations were performed using a Rainin Dynamax HPLC system equipped with a Dynamax UV-D II absorbance detector (Rainin Instrument Inc., Woburn, MA). The mobile phases included: Solution A (deionized water containing 0.1% formic acid) and Solution B (acetonitrile containing 0.1% formic acid). About 20 μL of extract was injected onto a Macrosphere C18 column (2.1 mm i.d. × 250 mm length, 5 μm particle size; Alltech Assoc. Inc., Deerfield, IL). The separation consisted of a 120 minute gradient of 5–95% Solution B. Fractions were automatically collected every two minutes using a Rainin Dynamax FC-4 fraction collector (Rainin Instrument Inc., Woburn, MA). The fractions were then concentrated using in Savant SC 110 SpeedVac concentrator (Thermo Electron Corporation) and re-suspended with minimum amount of 0.1% formic acid.
MALDI-FTMS and direct tissue analyses
MALDI-FTMS experiments were performed on a Varian/IonSpec ProMALDI Fourier transform mass spectrometer (Lake Forest, CA) equipped with a 7.0 Tesla actively-shielded superconducting magnet. The FTMS instrument contains an external MALDI source where the ions from multiple laser shots can be accumulated in the external hexapole storage trap before the ions are transferred to the ICR cell via a quadrupole ion guide. A 355 nm Nd: YAG laser (Laser Science, Inc., Franklin, MA) was used to create ions in an external source.
Off-line analysis of HPLC fractions was performed by spotting 0.3 μl of each individual HPLC fraction of interest on the MALDI sample plate and adding 0.3 μl of the saturated DHB. The resulting mixture was allowed to crystallize at room temperature. The MALDI-FTMS analysis was then performed as described above.
MALDI-TOF/TOF
A model 4800 MALDI-TOF/TOF analyzer (Applied Biosystems, Framingham, MA) equipped with a 200 Hz, 355 nm Nd:YAG laser was used for direct peptide profiling in SG sample and HPLC fraction screening. Acquisitions were performed in positive ion reflectron mode. Instrument parameters were set using the 4000 Series Explorer software (Applied Biosystems). Mass spectra were obtained by averaging 1000 laser shots covering mass range m/z 500–4000. MS/MS was achieved by 1 kV collision induced dissociation (CID) using air. For sample analysis, 0.4 μl of sample was spotted on MALDI plate first and allowed to dry followed by the addition of 0.4 μl matrix which consists of a saturated solution of CHCA in 70% acetonitrile.
Nano-LC-ESI-Q-TOF MS/MS
Nanoscale LC-ESI-Q-TOF MS/MS was performed using a Waters nanoAcquity UPLC system coupled to a Q-TOF Micro mass spectrometer (Waters Corp., Milford, MA). Chromatographic separations were performed on a homemade C18 reversed phase capillary column (75 μm internal diameter × 100 mm length, 3 μm particle size, 100 Å). The mobile phases used were: 0.1% formic acid in deionized water (A); 0.1% formic acid in ACN (B). An aliquot of 5.0 μl of a tissue extract or HPLC fraction was injected and loaded onto the trap column (Zorbax 300SB-C18 Nano trapping column, Agilent Technologies, Santa Clara, CA) using 95% mobile phase A and 5% mobile phase B at a flow rate of 10 μl/min for 10 minutes. For neuropeptides, the linear gradient was from 5% to 45% buffer B over 90 min. A data dependent acquisition was employed for the MS survey scan and the selection of three precursor ions and subsequent MS/MS of the selected parent ions. The MS scan range was from m/z 400–1800 and the MS/MS scan was from m/z 50–1800.
NanoLC coupled to MS/MS by CID and ETD
Setup for RP nanoLC separation
The setup for nanoLC consisted of an Eksigent nanoLC Ultra (Dublin, CA) with a 10 μL injection loop. Mobile phase A consisted of H2O in 0.1% formic acid and mobile phase B consisted of ACN (Fisher Scientific, Optima LC/MS grade, Fair Lawn, NJ). Injections consisted of 5 μL of prepared tissue extract onto an Agilent Technologies Zorbax 300 SB-C18 5 μm, 5 × 0.3 mm trap cartridge (Santa Clara, CA) at a flow rate of 5 μL/min for 5 minutes at 95% A/5% B. Separation was performed on a Waters HPLC column with 3 μm Atlantis dC18 75 μm × 150 mm (Milford, MA) on a gradient from 5 to 45% mobile phase B at 250 nL/min, over 90 minutes at room temperature. Emitter tips were pulled from 75 μm inner diameter 360 μm outer diameter capillary tubing (Polymicro Technologies, Phoenix, AZ) using a commercial laser puller model P-2000 (Sutter Instrument Co., Novato, CA).
Mass spectrometry data acquisitions
An ion-trap mass spectrometer (amaZon ETD, Bruker Daltonics, Bremen, Germany) equipped with an on-line nanospray source was used for mass spectrometry data acquisition. Hystar (Version 3.2, Bruker Daltonics, Bremen, Germany) was used to couple and control Eksigent nanoLC software (Dublin, CA; Version 3.0 Beta) for MS acquisition for all experiments. Smart parameter setting (SPS) was set to m/z 700 for CID and m/z 500 for ETD, compound stability and trap drive level were set at 100%. Optimization of the nanospray source resulted in dry gas temperature, 125°C, dry gas, 4.0 L/min, capillary voltage, −1300 V, end plate offset, −500 V, MS/MS fragmentation amplitude, 1.0V, and SmartFragmentation was set at 30–300%. Data were generated in data dependent mode with strict active exclusion set after two spectra and released after one minute. CID MS/MS fragmentation was set to fragment the six most abundant MS peaks with a preference for doubly charged ions and excluded singly charged ions. ETD MS/MS fragmentation was set to fragment the four most abundant MS peaks with no preference for specifically charged ions except for excluding singly charged ions. For MS generation the ion charge control (ICC) target was set to 200,000, maximum accumulation time, 50.00 ms, four spectral average, acquisition range of m/z 300–1500, and scan speed (enhanced resolution) of 8,100 m/z s−1. For MS/MS generation the ICC target was set to 200,000, maximum accumulation time, 50.00 ms, three spectral averages, acquisition range of m/z 100–2000, and scan speed (Ultrascan) of 32,000 m/z s−1. The ETD reagent parameters were set to 400,000 ICC target for the fluoranthene radical anion. For ETD MS/MS acquisition the fluoranthene radical anion has an m/z of 210 and therefore the m/z 210 was removed from the resulting ETD MS/MS spectra, and m/z 160 was set as the MS/MS low m/z cut-off.
Peptide prediction, de novo sequencing and database searches
De novo sequencing using ESI-Q-TOF was performed via a combination of MassLynx™ 4.1 PepSeq software (Waters) and manual sequencing. Tandem mass spectra acquired from the Q-TOF were first deconvoluted using MaxEnt 3 software (Waters) to convert multiply charged ions into their singly charged forms. The resulting spectra were pasted into the PepSeq window for sequencing analysis. The candidate sequences generated by the PepSeq software were compared and evaluated for homology with previously known peptides. The online program blastp (National Center for Biotechnology Information, Bethesda, MD; http://www.ncbi.nlm.nih.gov/BLAST/) was used to search the existing NCBI crustacean protein database, using the candidate peptide sequences as queries. For all searches, the blastp database was set to non-redundant protein sequences (i.e. nr) and restricted to crustacean sequences (i.e. taxid: 6657). For each of the proteins putatively identified via blastp, the BLAST score and BLAST-generated E-value for significant alignment are provided in the appropriate subsection of the results. Peptides with partial sequence homology were selected for further examination by comparing theoretical MS/MS fragmentation spectra generated by PepSeq with the raw MS/MS spectra. If the fragmentation patterns did not match well, manual sequencing was performed. For CID and ETD data via amaZon ion-trap mass spectrometer, the MS data were processed with DataAnalysis (Version 4.0, Bruker Daltonics Bremen, Germany). Deviations in parameters from the default Protein Analysis in DataAnalysis were as follows: intensity threshold, 1000, maximum number of compounds, 1E7, and retention time window 0.5 minutes. These parameter changes assisted in providing better quality spectra for sequencing. Identification of peptides was performed using Mascot (Version 2.3, Matrix Science, London, U.K.). Searches were performed against a custom crustacean database; no enzyme selected; amidated C-terminal as variable modifications; peptide tolerance mass error 1.2 Da; MS/MS mass error tolerance was 0.6 Da.
Fabrication of CIEF system
75 μm i.d. × 60 cm fused-silica capillary was flushed by 1 M NaOH for 10 min and dried with air before treatment. Hydroxypropyl cellulose (HPC) was dissolved in water to the final concentration of 5% (w/w) and was used to coat the capillary inner wall in an oven by heating the capillary from 60 °C to 140 °C over 16 minutes followed by 20 minute stabilization. Immediately following this procedure, the capillary was removed from the oven and washed with water and dried before further processing. Cellulose acetate membrane-coated porous joints were made 3 cm from both ends of the HPC-coated capillary as we have previously described.45 Briefly, a small fracture section was made on the capillary fixed on a 1 cm × 0.3 cm plastic slide and was further covered by cellulose acetate solution (12% in acetone, w/v). Under a gentle stream of air a uniform cellulose acetate membrane could form over the fracture section. This porous joint was then placed into a plastic vial (Fisher Scientific, Pittsburgh, PA) with about 2 cm of capillary stretched out of the vial from a small hole on the bottom. The 1% acetic acid solution was filled in the vial and the electrode wire was inserted into the vial to form an electrical connection.
CIEF procedure and sample collection
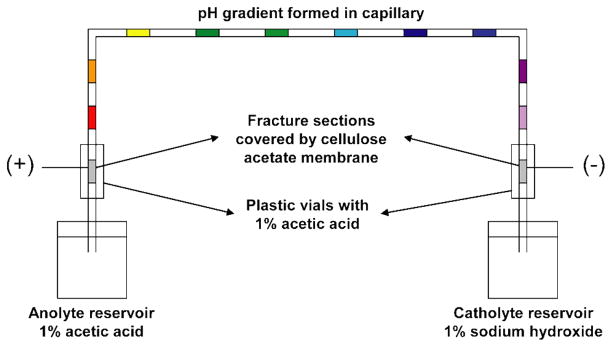
CIEF was performed by using TriSep-2100 HV power supplier from Unimicro Technologies (Pleasanton, CA). The 1% acetic acid solution and 1% sodium hydroxide solution were chosen as anolyte and catholyte, respectively. The 1% acetic acid solution was filled into the plastic vials containing membrane-coated joints, and electrodes were placed into these vials so that a circuit was formed via the porous joints, as shown in Figure 1. Prior to CIEF, the capillary was flushed with water and dried under air flow in sequence. A constant voltage of 20 kV was then applied to the capillary for 10–12 min depending to the samples. After focusing, the fractions were mobilized by air pressure. A MALDI plate pre-coated with Parafilm M was used for sample collection as described previously.46 The matrix was pre-spotted on the MALDI plate with 120 mg/ml of DHB. For each of the spots, 250 nL of matrix solution was mixed with approximately 150 nL of an individual collected fraction, and co-crystallized before MALDI-FTMS analysis.
Figure 1. Schematic illustration of the membrane-assisted CIEF setup enhanced with cellulose acetate membrane-coated porous joints.
The sample was mixed with 0.5% of Pharmalyte and filled into the capillary, which was then inserted into the anolyte and catholyte reservoirs with acid and base, respectively. A high voltage power supply was connected via the two membrane-coated joints near capillary ends to form a circuit, so that the shadowed sections were free from electrical field during focusing and formed “plugs” to prevent sample loss. After focusing, the reservoirs were removed, and the fractions were mobilized by air pressure and collected on a MALDI plate with pre-spotted matrix.
Results and Discussion
Comprehensive Neuropeptidomic Study in the Sinus Gland of C. sapidus Using Complementary Mass Spectral Techniques
In this study, the high resolution and mass accuracy of MALDI–FTMS, high sensitivity of MALDI-TOF/TOF and capability of de novo sequencing of nanoLC-ESI-Q-TOF were used to identify 70 neuropeptides consisting of 12 neuropeptide families from the SG. Although 43 neuropeptides were previously identified in C. borealis and C. maenas, here we report the successful identification of 27 novel peptides de novo sequenced for the first time from the SG of C. sapidus, as shown in Table 1. Multiple complementary mass spectrometric instruments have been employed, each offering overlapped yet distinct coverage due to different ionization preference and sequencing capabilities. As a comparison, 47 peptides were detected and sequenced using nanoESI-Q-TOF, 38 peptides were detected by MALDI-TOF/TOF and 53 were observed via MALDI-FTMS, while only 18 were commonly detected by all three instruments in the SG of C. sapidus. In addition, multiple sample preparation methods including direct tissue analysis, tissue extraction, off-line HPLC separation were used in this study to obtain a more complete neuropeptidome coverage. Direct tissue analysis provides a quick snapshot of the peptide profile as well as accurate mass measurements to identify numerous previously known neuropeptides; acidified methanol extraction offers a more complete profile of the peptides present in the whole tissue. Off-line reversed phase HPLC prior to MS analysis reduces the chemical complexity and thus expands the dynamic range of detectable neuropeptides using MS. Improved sample preparation results in enhanced neuropeptide detection from the HPLC fractions compared to crude extraction due to its significant signal suppression from lipids and chemical complexity. As an example, 8 peptides were detected from a single direct tissue, 21 were detected in crude extract and 53 were detected from HPLC fractions using MALDI-FTMS.
Table 1.
Neuropeptides identified in the sinus gland (SG) of Callinectes sapidus SG by multiple MS approaches
| Neuropeptide family | m/z | Peptide sequence | Q-TOF | FT | TOF/TOF |
|---|---|---|---|---|---|
| CabTRP | 934.4927 | APSGFLGMRa | + | + | − |
| 950.4921 | APSGFLGM(O)Ra | + | − | − | |
| 980.5027 | TPSGFLGM(O)Ramide | − | + | − | |
| CCAP | 956.3752 | PFCNAFTGCa | + | + | − |
| CPRP[1–9] | 975.5153 | RSAEGLGRMa | + | + | + |
| CPRP[1–10] | 1033.5207 | RSAEGLGRMG | + | + | + |
| CPRP[15–25] | 1216.6896 | SLKSDTVTPLR | + | − | + |
| CPRP[18–28] | 1221.6110 | SDTVTPLRGFE | + | + | − |
| CPRP[14–25] | 1287.7267 | ASLKSDTVTPLR | + | + | + |
| CPRP[1–12] | 1302.7059 | RSAEGLGRMGRL | + | + | + |
| CPRP[13–25] | 1400.8108 | LASLKSDTVTPLR | + | + | − |
| CPRP[1–13] | 1415.7900 | RSAEGLGRMGRLL | + | + | + |
| CPRP[22–36] | 1639.8075 | TPLRGFEGETGHPLE | + | + | − |
| Orcomyotropin | 1168.5422 | FPAFTTGFGSH | + | + | − |
| 1186.5164 | FDAFTTGFGHS | + | + | + | |
| Ork | 1027.4843 | EIDRSGFGF | + | + | − |
| 1066.4800 | NFDEIDRSA | + | + | − | |
| 1098.5214 | EIDRSGFGFA | + | + | − | |
| 1142.5113 | FDEIDRSGFG | + | − | − | |
| 1156.5269 | FDEIDRSGFA | + | + | − | |
| 1186.5375 | FDEIDRSSFA | + | + | − | |
| 1198.5487 | NFDEIDRSGFa | − | + | − | |
| 1199.5327 | NFDEIDRSGF | − | + | + | |
| 1213.5484 | DEIDRSGFGFA | − | + | + | |
| 1228.5593 | NFDEIDRSSFa | + | + | − | |
| 1256.5542 | NFDEIDRSGFG | + | + | + | |
| 1257.5382 | DFDEIDRSGFG | + | + | − | |
| 1270.5699 | NFDEIDRSGFA | + | + | + | |
| 1271.5539 | DFDEIDRSGFA | + | + | − | |
| 1286.5648 | NFDEIDRSSFG | + | + | + | |
| 1287.5488 | DFDEIDRSSFG | + | − | + | |
| 1300.5804 | NFDEIDRSSFA | + | + | − | |
| 1301.5644 | DFDEIDRSSFA | + | + | + | |
| 1360.6168 | FDEIDRSGFGFA | − | + | − | |
| 1403.6226 | NFDEIDRSGFGF | + | + | + | |
| 1433.6332 | NFDEIDRSSFGF | + | + | + | |
| 1474.6597 | NFDEIDRSGFGFA | + | + | + | |
| 1475.6437 | DFDEIDRSGFGFA | + | + | + | |
| 1502.6910 | NFDEIDRSGFGFV | + | + | + | |
| 1503.6750 | DFDEIDRSGFGFV | + | + | + | |
| 1504.6703 | NFDEIDRSSFGFA | + | + | + | |
| 1505.6543 | DFDEIDRSSFGFA | + | + | − | |
| 1532.7016 | NFDEIDRSSFGFV | + | + | + | |
| 1533.6856 | DFDEIDRSSFGFV | + | − | − | |
| 1547.6761 | NFDEIDRSSFGFN | + | + | + | |
| 1548.6601 | DFDEIDRSSFGFN | + | − | + | |
| 1554.6972 | NFDEIDRTGFGFH | − | + | + | |
| Ork/Orcomyotropin-related | 1062.5578 | TPRDIANLY | + | − | − |
| Others | 844.4788 | HLGSLYRa | − | + | + |
| 902.5458 | KIFEPLR | − | + | − | |
| 1259.7106 | KIFEPLRDKN | − | − | + | |
| PDH | 1190.5762 | pQELHVPEREA | + | − | − |
| 1927.1800 | NSELINSILGLPKVMNDAa | + | − | − | |
| proctolin | 649.3668 | RYLPT | − | + | + |
| RFa | 925.4890 | ETNFLRFa | − | − | + |
| 1005.5741 | GRPNFLRFa | − | + | − | |
| 1029.5629 | DHVPFLRFa | − | + | − | |
| 1045.5802 | GHRNFLRFa | − | − | + | |
| 1059.5959 | AHRNFLRFa | − | + | − | |
| 1104.6061 | GAHKNYLRFa | + | + | + | |
| 1116.5473 | YEQDFLRFa | − | + | − | |
| 1146.6055 | GYSKNYLRFa | − | + | + | |
| 1172.6323 | AYNRSFLRFa | − | + | + | |
| 1174.6368 | FTSKNYLRFa | − | − | + | |
| 1209.6123 | DENRNFLRFa | − | + | − | |
| 1273.6436 | YGSDRNFLRFa | − | − | + | |
| 1314.7753 | DARTAPLRLRFa | − | + | − | |
| RPCH | 930.3443 | pELNFSPGWa | + | − | − |
| RYa | 1030.4703 | pEGFYSQRYa | + | − | + |
| SIFa | 1381.7375 | GYRKPPFNGSIFa | − | + | − |
Peptides identified by de novo sequencing using ESI-Q-TOF, MALDI-FTMS and MALDI-TOF/TOF. Neuropeptides listed in red are novel peptides sequenced for the first time in this study. Lower case “a” at the C-terminal peptide indicates C-terminal amidation. CabTRP: Cancer borealis tachykinin-related peptide; CCAP: Crustacean cardioactive peptide; CPRP: CHH precursor-related peptide; PDH: pigment dispersing hormone; RPCH: red pigment concentrating hormone; Ork: Orcokinin
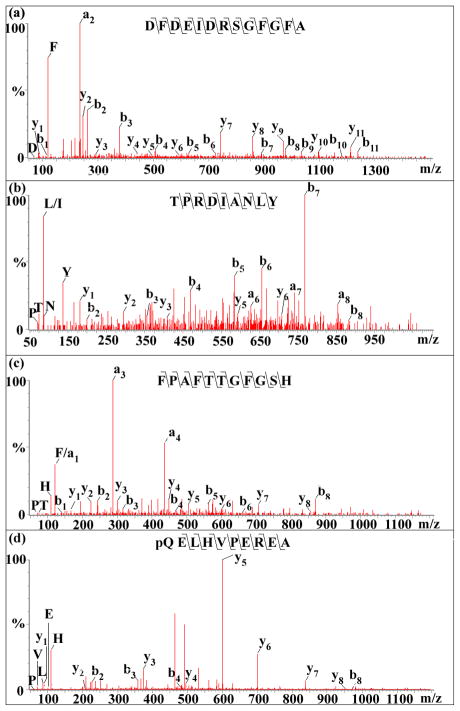
Among these novel peptides, there are 9 CPRPs and 15 orcokinins. Small pieces of detected CPRP fragments taken together cover the amino acid sequence of the intact CPRP RSAEGLGRMGRLLASKSDTVTPLRGFEGETGHPLE. Thirty-two orcokinins were sequenced from SG of C. sapidus, suggesting that the SG is rich in orcokinins. Fifteen novel orcokinins including 9 full length orcokinins such as DFDEIDRSGFG, DFDEIDRSSFG, DFDEIDRSSFA, NFDEIDRSSFGF, DFDEIDRSGFGFA, DFDEIDRSGFGFV, DFDEIDRSSFGFA, DFDEIDRSSFGFV and DFDEIDRSSFGFN, and 6 truncated orcokinins such as EIDRSGFGF, NFDEIDRSA, FDEIDRSGFG, FDEIDRSGFA, FDEIDRSSFA and DEIDRSGFGFA were sequenced by ESI-Q-TOF MS/MS. A spectrum of a novel orcokinin, DFDEIDRSGFGFA (m/z 1475.56), is shown in Figure 2(a). In addition, several novel peptides from other peptide families were sequenced including one orcokinin-related peptide TPRDIANLY (m/z 1062.56), one pigment dispersing hormone (PDH)-related peptide pQELHVPEREA (m/z 1190.58) and one orcomyotropin FPAFTTGFGSH (m/z 1168.54). The MS/MS fragmentation spectra of these three peptides sequenced by ESI-Q-TOF MS/MS are shown in Figure 2(b-d). As shown in Figure 3, one truncated orcokinin FDEIDRSSFA (m/z 1186.54) was detected in neuropeptide profile of the direct tissue analysis, suggesting that these truncated peptides are endogenously processed orcokinins in the SG due to minimal sample preparation involved in direct tissue analysis method as opposed to conventional tissue extract analysis. Collectively, our data not only significantly expand the catalog of peptide hormones known to be present in C. sapidus, but also provide a foundation for future studies of peptide functions in this important commercial species.
Figure 2.
MS/MS fragmentation spectra of several novel neuropeptides in C. sapidus SG by ESI-Q-TOF: (a) orcokinin (DFDEIDRSGFGFA m/z 1475.64), all y-ions and b-ions are detected; (b) Ork/Orcomyotropin-related peptide (TPRDIANLY m/z 1062.56), almost all b-ions from b2 to b8 are detected and are very abundant compared to the incomplete y-ion series; (c) orcomyotropin (FPAFTTGFGSH m/z 1168.54), y1-y8, b1-b8 except b7 are detected with the most abundant fragment ions being a3 and a4; (d) PDH-related peptide (pQELHVPEREA m/z 1190.57), y1-y8,b2-b4 and b8 are detected.
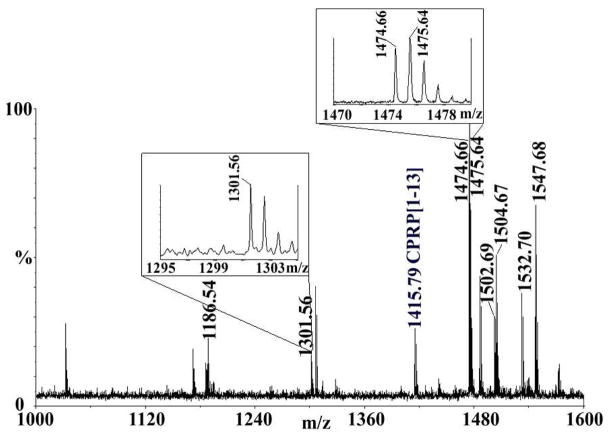
Figure 3.
Neuropeptide profiling of the sinus gland by direct tissue analysis using MALDI-TOF/TOF. The mass spectral insets with higher magnification highlight the existence of two N-terminal Asp-containing orcokinin neuropeptides: DFDEIDRSSFA (m/z 1301.56) and DFDEIDRSGFGFA (m/z 1475.56). A truncated CPRP fragment CPRP [1–13] RSAEGLGRMGRLL (m/z 1415.79) was labeled in blue.
Identification and Characterization of an Intact CPRP Using ETD
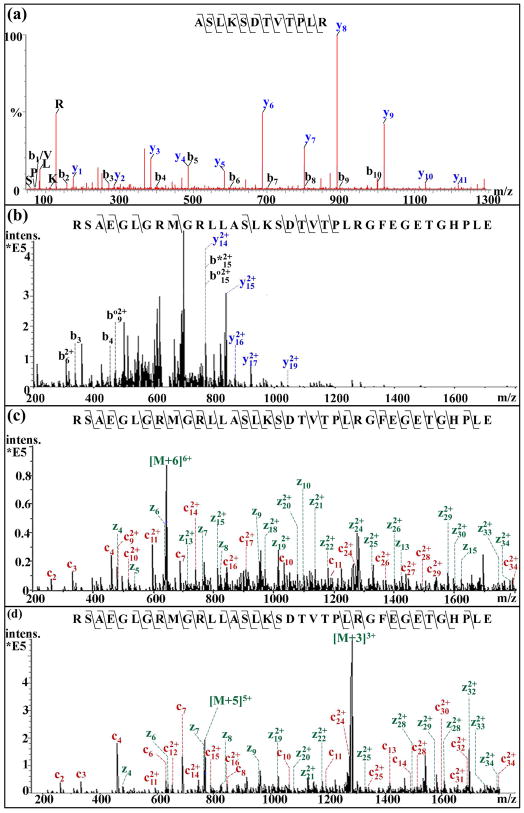
Using ESI-Q-TOF, 9 truncated CPRPs fragments were sequenced to cover the amino acid sequence of the intact CPRP peptide RSAEGLGRMGRLLASLKSDTVTPLRGFEGETGHPLE. Several truncated CPRP fragments such as CPRP [1–9] 975.52 (RSAEGLGRMa, data not shown) and CPRP [1–13]1415.79 (RSAEGLGRMGRLL) were detected in direct tissue analysis as shown in Figure 3 by MALDI TOF/TOF, suggesting the potential cleavage on prohormone involved in posttranslational proteolytic processing. More truncated CPRP fragments were expected to exist in the SG but were not detected in single tissue analysis, presumably due to their relatively lower abundances or poor ionization efficiency. Furthermore, the truncated CPRP fragments such as CPRP [14–25] ASLKSDTVTPLR resulted from a typical cleavage site on Arg as well. A representative CID MS/MS spectrum of CPRP [14–25] ASLKSDTVTPLR, is shown in Figure 4(a). CID using ESI-Q-TOF mainly produces fragmentation for doubly and triply charged molecules, which is adequate to sequence peptides with relatively small molecular weight (less than 2000 Da). However, CID fragmentation is insufficient to sequence the larger intact CPRP in a complex sample whose molecular mass is 3837 Da and therefore it is extremely difficult to directly sequence by traditional CID via ESI-Q-TOF. ETD is an attractive fragmentation alternative to sequence the highly charged CPRP peptide. In ETD, a protonated peptide reacts with an anion (fluoranthene radical) where the anion transfers an electron to the peptide cation and the resulting fragmentation occurs along the N-Cα bond by free-radical-driven cleavage. The large size of CPRP and numerous basic amino acid residues offer multiple sites to sequester charge make it well-suited for ETD fragmentation technique. Unlike CID which suffers from sequence-dependent fragmentation, ETD has no preferred cleavage sites except proline, which leads to a more uniform distribution of the fragment ions, thus providing superior sequence coverage for larger peptides.47 As shown in Figure 4(c, d), the extensive fragmentation patterns of ETD is observed compared to the limited and biased CID fragmentation in Figure 4(b). As evident from the comparison, ETD offers more uniform and extensive fragmentation than CID, thus providing enhanced coverage for large intact peptides such as CPRPs. Specifically, we observe 12.5% of the b- and y- cleavages for CID as compared to an average of 53.5% of c- and z•- type fragments. More than a four-fold increase in fragments using ETD suggests the utility of this relatively new tandem MS fragmentation method as a complementary technique for de novo sequencing of large intact endogenous peptides and novel neuropeptide discovery endeavors.
Figure 4.
A comparison of tandem MS spectra of truncated CPRP [14–25] (ASLKSDTVTPL, m/z 1287.66) by ESI-Q-TOF (a) and intact CPRP (MW 3837) by the amaZon ETD ion trap mass spectrometer for both CID and ETD fragmentation (b) MS2 of precursor ion with m/z 640 with charge state +6 by CID, * represent loss of NH3, º represent loss of H2O, (c) MS2 of precursor ion with m/z 640 with charge state +6 by ETD at (d) MS2 of precursor ion with m/z 768 with charge state +5 by ETD. For all labels in Figure 4, charge state +1 is not specified.
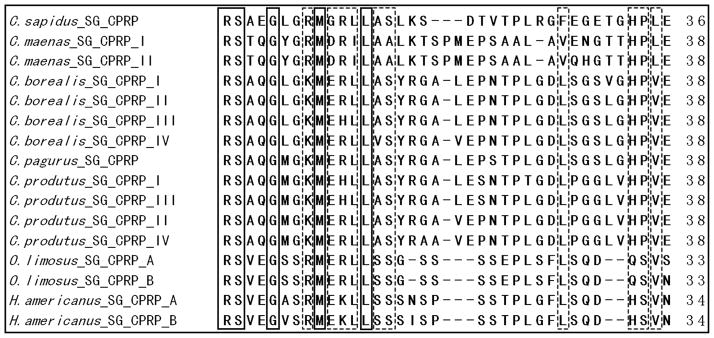
Figure 5 compares the sequences of CPRP in C. sapidus to those of other crustacean species. Significant sequence homology is observed at both the N- and C-termini of the CPRP sequences of these related species, with several amino acid residues being completely conserved among all of the crustacean species examined (highlighted with boxes). The extensive sequence homology among the CPRPs supports the hypothesis that they serve an important biological role and that this role is conserved. Although previous studies showed that the CPRP can act as a circulating hormone in the hemolymph and has a relatively long lifetime,38, 48 the precise physiological role(s) it plays in crustaceans remains unknown. Further investigations are needed to understand the functions of various CPRPs in multiple crustaceans.
Figure 5.
Amino acid sequence alignment of 16 CPRPs from seven crustacean species. Amino acid residues exhibiting 100% conservation are boxed with solid lines. Amino acid residues with significant, but less than 100%, conservation are boxed with dashed lines, indicating significant sequence homology among CPRPs in various crustacean species.
Discovery of N-terminal Asp-Orcokinins
The first orcokinin NFDEIDRSGFGFN was isolated from abdominal nerve cord extracts of the crayfish Orconectes limosus as a myoactive peptide acting on gut tissues.18 Subsequently, numerous orcokinins have been discovered from crustaceans, insects and other invertebrates 22, 28, 49–51 with the conserved N-terminal motif NFDEIDR. Unlike previously reported orcokinins, most of the novel full sequence orcokinins identified from this study, share the conserved N-terminal sequence of DFDEIDR except NFDEIDRSSFGF. One plausible explanation for this observation could be caused by deamidation of Asn during complex sample preparation procedures including tissue extraction, peptide purification, and separation. Alternatively, conserved amino-terminal Asp-containing orcokinins could be caused by a change of a single codon in orcokinins for this species; however, this is a hypothesis due to the lack of genetic information of pre-proorcokinin for this specific species. To investigate the origins of these amino-terminal Asp-containing orcokinins, direct tissue analysis was used to examine the resulting peptide profiles with minimal sample preparation involving only matrix application. As shown in Figure 3, via direct tissue analysis of SG by MALDI-TOF/TOF, 8 orcokinins were observed, including both more commonly found N-terminal Asn-containing orcokinins such as NFDEIDRSGFGFA (m/z 1474.66) and NFDEIDRSGFGFV (m/z 1502.52), and N-terminal Asp-containing orcokinins such as DFDEIDRSSFA (m/z 1301.56) and DFDEIDRSGFGFA (m/z 1475.56). A reduced number of N-terminal Asp-containing orcokinins were detected in the direct tissue analysis due to their low abundances and sample complexity in a single SG tissue. Nonetheless, the detection of DFDEIDRSSFA and DFDEIDRSGFGFA provided validation of N-terminal Asp-containing sequences in orcokinins detected in the SG of C. sapidus.
Profiling of Orcokinin Family Neuropeptides from the Sinus Gland of Callinectes sapidus Using MA-CIEF Separation Coupled to MALDI FTMS Detection
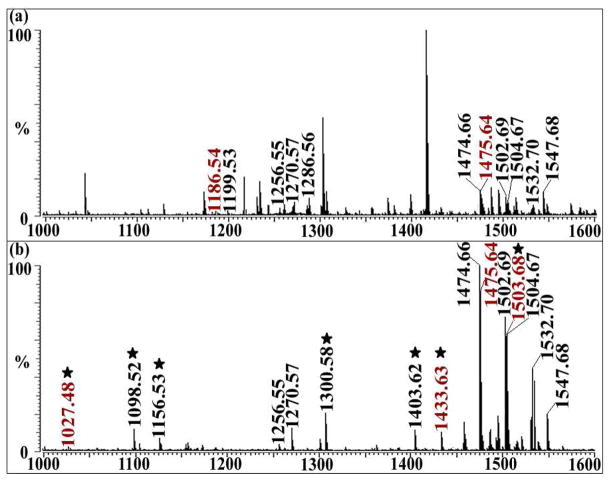
Orcokinin (NFDEIDRSGFGFN) and orcokinin homologues were first discovered as crustacean peptides with numerous physiological effects such as potent hindgut myostimulating factors,18 inhibiting the pyloric rhythm,23 and neurohormones in hemolymph,52 to name a few. Therefore, it is important to comprehensively identify and discover new orcokinins in complex animal nervous systems. However, due to the existence of multiple acidic amino acids such as D and E, orcokinins are less easy to ionize in the positive ion mode, which makes them difficult to detect. As shown in Figure 6(a), most orcokinins have relatively low peak intensity around 20% or lower by direct MALDI FTMS profiling. As shown in Table 1, a total of 32 orcokinin isoforms are identified from C. sapidus SG by combining multiple sample preparation methods, including direct tissue analysis, crude extraction, capillary electrophoresis and offline HPLC separation. With the large number of orcokinins in C. sapidus and their relatively poor ionization efficiencies, a reliable separation method with high efficiency is required. Membrane-assisted capillary isoelectric focusing (MA-CIEF) provides the separation needed for the large number of orcokinins in C. sapidus to be detected by MALDI-FTMS. MA-CIEF is a modified CIEF system developed in our lab, which provides great potential in MS-based proteomics53 and neuropeptide analysis.54 Compared with regular CIEF, the MA-CIEF method offers much faster separation and higher peak intensities. Orcokinin family neuropeptides feature unique pI values that are different from other neuropeptide families. It was found that with the MA-CIEF/MALDI-FTMS based platform, MS signals were significantly improved, and more orcokinins were detected compared with those by direct MALDI-FTMS profiling without any separation.54. As shown in Figure 6(a), only 11 orcokinins including FDEIDRSSFA (m/z 1186.59), NFDEIDRSGF (m/z 1199.65), NFDEIDRSGFG (m/z 1256.63), NFDEIDRSGFA (m/z 1270.65), NFDEIDRSSFG (m/z 1286.63), NFDEIDRSGFGFA (m/z 1474.75), DFDEIDRSGFGFA (m/z 1475.85), NFDEIDRSGFGFV (m/z 1502.79), NFDEIDRSSFGFA (m/z 1504.75), NFDEIDRSSFGFV (m/z 1532.81) and NFDEIDRSSFGFN (m/z 1547.74) were identified from 0.3 μL tissue extraction of SG using MALDI-FTMS with relatively low intensity. After separation by MA-CIEF, 2.5 μL of tissue extraction resulted in 24 fractions with each one containing around 100 nL of tissue extraction and 15 orcokinins were identified from a single fraction at a relatively high sensitivity. In Figure 6(b), the 7 peaks marked with stars are orcokinins not identified by direct profiling of tissue extract, including EIDRSGFGF (m/z 1027.48), EIDRSGFGFA (m/z 1098.53), FDEIDRSGFA (m/z 1156.58), NFDEIDRSSFA (m/z 1300.60), NFDEIDRSGFGF (m/z 1403.65), NFDEIDRSSFGF (m/z 1433.64), and DFDEIDRSGFGFV (m/z 1503.69). The profiling of all 24 fractions enabled identifications of 23 orcokinins, including all of the isoforms previously identified by direct profiling and 12 additional orcokinins which were not detected via direct profiling. Furthermore, only 2 novel peptides were detected by direct profiling of tissue extraction, in contrast 5 novel peptides were detected in a single fraction and a total of 9 novel peptides were detected by combining results from all the fractions by CIEF separation, highlighting the unique advantages of CIEF in discovery of novel peptides. After CIEF separation, isoforms of the orcokinin family with similar pI values were separated from other neuropeptides and focused to a narrow band corresponding to their similar pI values. Most neuropeptides in the crustacean nervous systems are basic, for example, RFamides have pI values higher than 12, and tackykinins have pI values higher than 14 (calculated from http://innovagen.net). Orcokinins, unlike other neuropeptides, are the most acidic neuropeptide family with pI value around 3–4, therefore they can be easily concentrated to a narrow section of the CIEF capillary and separated from other neuropeptides, which make CIEF extremely suitable for concentrating and separating orcokinins.
Figure 6.
Analysis of orcokinin family neuropeptides in the sinus gland (SG) of C. sapidus is enhanced by the use of membrane-assisted (MA)-CIEF/MALDI-FTMS. Orcokinins peptides are marked with molecular weight on the mass spectra. (a) Control sample without CIEF separation. (b) Fraction 16 which is rich in orcokinin family peptides after CIEF separation. Peptides with stars were only found with CIEF separation, and peptides labeled in red are novel peptides.
Conclusions
In this study, the neuropeptides in the SG of C. sapidus were identified and sequenced using multifaceted MS-based platforms. Collectively, 70 neuropeptides have been sequenced including 15 novel ones. ETD was performed to improve the sequence coverage of large endogenous neuropeptides with higher charge and larger size. The intact CPRP from C. sapidus was identified and characterized with 68% sequence coverage by MS for the first time. The discovery of N-terminal Asp-containing orcokinins was reported and discussed. A large number of orcokinins were discovered and a novel MA-CIEF configuration was employed to facilitate the separation of acidic amino acid-containing orcokinins. Our study not only significantly expands the catalog of peptide hormones known to be present in C. sapidus, but also provides a foundation for future studies of peptide function in this important aquaculture species.
Acknowledgments
The authors thank the Analytical Instrument Center at UW School of Pharmacy for the access of MALDI-FTMS and amaZon ETD ion trap instruments. The authors wish to thank the University of Wisconsin-Biotechnology Mass Spectrometry Facility for access to the MALDI-TOF/TOF instrument. This work was supported in part by the National Science Foundation grant (CHE-0957784), and National Institutes of Health through grant 1R01DK071801. L. Li acknowledges a Vilas Associate Award and an H. I. Romnes Faculty Research Fellowship.
References
- 1.Schwartz MW, Woods SC, Porte D, Jr, Seeley RJ, Baskin DG. Central nervous system control of food intake. Nature. 2000;404(6778):661–71. doi: 10.1038/35007534. [DOI] [PubMed] [Google Scholar]
- 2.Sweedler JV, Li L, Rubakhin SS, Alexeeva V, Dembrow NC, Dowling O, Jing J, Weiss KR, Vilim FS. Identification and characterization of the feeding circuit-activating peptides, a novel neuropeptide family of aplysia. J Neurosci. 2002;22(17):7797–808. doi: 10.1523/JNEUROSCI.22-17-07797.2002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Baggerman G, Cerstiaens A, De Loof A, Schoofs L. Peptidomics of the larval Drosophila melanogaster central nervous system. J Biol Chem. 2002;277(43):40368–40374. doi: 10.1074/jbc.M206257200. [DOI] [PubMed] [Google Scholar]
- 4.Desiderio DM. Mass spectrometric analysis of neuropeptidergic systems in the human pituitary and cerebrospinal fluid. J Chromatogr B. 1999;731(1):3–22. doi: 10.1016/s0378-4347(99)00172-3. [DOI] [PubMed] [Google Scholar]
- 5.Li LJ, Kelley WP, Billimoria CP, Christie AE, Pulver SR, Sweedler JV, Marder E. Mass spectrometric investigation of the neuropeptide complement and release in the pericardial organs of the crab, Cancer borealis. J Neurochem. 2003;87(3):642–656. doi: 10.1046/j.1471-4159.2003.02031.x. [DOI] [PubMed] [Google Scholar]
- 6.Li LJ, Moroz TP, Garden RW, Floyd PD, Weiss KR, Sweedler JV. Mass spectrometric survey of interganglionically transported peptides in Aplysia. Peptides. 1998;19(8):1425–1433. doi: 10.1016/s0196-9781(98)00094-1. [DOI] [PubMed] [Google Scholar]
- 7.Strand FL. Neuropeptides: regulators of physiological processes. 1. MIT Press; Cambridge, Mass: 1999. p. 658. [Google Scholar]
- 8.Fu Q, Goy MF, Li LJ. Identification of neuropeptides from the decapod crustacean sinus glands using nanoscale liquid chromatography tandem mass spectrometry. Biochem Biophys Res Comm. 2005;337(3):765–778. doi: 10.1016/j.bbrc.2005.09.111. [DOI] [PubMed] [Google Scholar]
- 9.Fu Q, Christie AE, Li LJ. Mass spectrometric characterization of crustacean hyperglycemic hormone precursor-related peptides (CPRPs) from the sinus gland of the crab, Cancer productus. Peptides. 2005;26(11):2137–2150. doi: 10.1016/j.peptides.2005.03.040. [DOI] [PubMed] [Google Scholar]
- 10.Ma MM, Chen RB, Ge Y, He H, Marshall AG, Li LJ. Combining bottom-up and top-down mass spectrometric strategies for de vovo sequencing of the crustacean hyperglycemic hormone from Cancer borealis. Anal Chem. 2009;81(1):240–247. doi: 10.1021/ac801910g. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Ma MM, Sturm RM, Kutz-Naber KK, Fu Q, Li LJ. Immunoaffinity-based mass spectrometric characterization of the FMRFamide-related peptide family in the pericardial organ of Cancer borealis. Biochem Biophys Res Commun. 2009;390(2):325–330. doi: 10.1016/j.bbrc.2009.09.122. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Ma MM, Bors EK, Dickinson ES, Kwiatkowski MA, Sousa GL, Henry RP, Smith CM, Towle DW, Christie AE, Li LJ. Characterization of the Carcinus maenas neuropeptidome by mass spectrometry and functional genomics. Gen Comp Endocrinol. 2009;161(3):320–334. doi: 10.1016/j.ygcen.2009.01.015. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Chen RB, Jiang XY, Conaway MCP, Mohtashemi I, Hui LM, Viner R, Li LJ. Mass spectral analysis of neuropeptide expression and distribution in the nervous system of the lobster Homarus americanus. J Proteome Res. 2010;9(2):818–832. doi: 10.1021/pr900736t. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Ma MM, Chen RB, Sousa GL, Bors EK, Kwiatkowski MA, Goiney CC, Goy MF, Christie AE, Li LJ. Mass spectral characterization of peptide transmitters/hormones in the nervous system and neuroendocrine organs of the American lobster Homarus americanus. Gen Comp Endocrinol. 2008;156(2):395–409. doi: 10.1016/j.ygcen.2008.01.009. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Klein JM, Mohrherr CJ, Sleutels F, Riehm JP, Rao KR. Molecular-cloning of 2 pigment-dispersing hormone (PDH) precursors in the blue crab Callinectes sapidus reveals a novel member of the PDH neuropeptide family. Biochem Biophys Res Commun. 1994;205(1):410–416. doi: 10.1006/bbrc.1994.2680. [DOI] [PubMed] [Google Scholar]
- 16.Krajniak KG. The identification and structure-activity relations of a cardioactive FMRFamide-related peptide from the blue crab Callinectes sapidus. Peptides. 1991;12(6):1295–1302. doi: 10.1016/0196-9781(91)90210-g. [DOI] [PubMed] [Google Scholar]
- 17.Lee KJ, Elton TS, Bej AK, Watts SA, Watson RD. Molecular cloning of a cDNA-encoding putative molt-inhibiting hormone from the blue crab, Callinectes sapidus. Biochem Biophys Res Commun. 1995;209(3):1126–1131. doi: 10.1006/bbrc.1995.1614. [DOI] [PubMed] [Google Scholar]
- 18.Stangier J, Hilbich C, Burdzik S, Keller R. Orcokinin - a novel myotropic peptide from the nervous-system of the crayfish, Orconectes limosus. Peptides. 1992;13(5):859–864. doi: 10.1016/0196-9781(92)90041-z. [DOI] [PubMed] [Google Scholar]
- 19.Skiebe P, Dreger M, Borner J, Meseke M, Weckwerth W. Immunocytochemical and molecular data guide peptide identification by mass spectrometry: Orcokinin and orcomyotropin-related peptides in the stomatogastric nervous system of several crustacean species. Cell Mol Biol. 2003;49(5):851–871. [PubMed] [Google Scholar]
- 20.Stemmler EA, Provencher HL, Guiney ME, Gardner NP, Dickinson PS. Matrix-assisted laser desorption/ionization Fourier transform mass spectrometry for the identification of orcokinin neuropeptides in crustaceans using metastable decay and sustained off-resonance irradiation. Anal Chem. 2005;77(11):3594–3606. doi: 10.1021/ac0502347. [DOI] [PubMed] [Google Scholar]
- 21.Dickinson PS, Stemmler EA, Barton EE, Cashman CR, Gardner NP, Rus S, Brennan HR, McClintock TS, Christie AE. Molecular, mass spectral, and physiological analyses of orcokinins and orcokinin precursor-related peptides in the lobster Homarus americanus and the crayfish Procambarus clarkii. Peptides. 2009;30(2):297–317. doi: 10.1016/j.peptides.2008.10.009. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Ma MM, Kutz-Naber KK, Li LJ. Methyl esterification assisted MALDI FTMS characterization of the orcokinin neuropeptide family. Anal Chem. 2007;79(2):673–681. doi: 10.1021/ac061536r. [DOI] [PubMed] [Google Scholar]
- 23.Li LJ, Pulver SR, Kelley WP, Thirumalai V, Sweedler JV, Marder E. Orcokinin peptides in developing and adult crustacean stomatogastric nervous systems and pericardial organs. J Comp Neurol. 2002;444(3):227–244. doi: 10.1002/cne.10139. [DOI] [PubMed] [Google Scholar]
- 24.Yasuda-Kamatani Y, Yasuda A. Identification of orcokinin gene-related peptides in the brain of the crayfish Procambarus clarkii by the combination of MALDI-TOF and on-line capillary HPLC/Q-Tof mass spectrometries and molecular cloning. Gen Comp Endocrinol. 2000;118(1):161–172. doi: 10.1006/gcen.1999.7453. [DOI] [PubMed] [Google Scholar]
- 25.Bungart D, Kegel G, Burdzik S, Keller R. Structure-activity-relationships of the crustacean myotropic neuropeptide orcokinin. Peptides. 1995;16(2):199–204. doi: 10.1016/0196-9781(94)00187-1. [DOI] [PubMed] [Google Scholar]
- 26.Bungart D, Hilbich C, Dircksen H, Keller R. Occurrence of analogs of the myotropic neuropeptide orcokinin in the shore crab, Carcinus maenas - evidence for a novel neuropeptide family. Peptides. 1995;16(1):67–72. doi: 10.1016/0196-9781(94)00145-v. [DOI] [PubMed] [Google Scholar]
- 27.Fu Q, Li LJ. Investigation of several unique tandem mass spectrometric fragmentation patterns of NFDEIDR, an orcokinin analog, and its N-terminal dimethylated form. Rapid Comm Mass Spectrom. 2006;20(4):553–562. doi: 10.1002/rcm.2337. [DOI] [PubMed] [Google Scholar]
- 28.Pascual N, Castresana J, Valero ML, Andreu D, Belles X. Orcokinins in insects and other invertebrates. Insect Biochem Mol Biol. 2004;34(11):1141–1146. doi: 10.1016/j.ibmb.2004.07.005. [DOI] [PubMed] [Google Scholar]
- 29.Hofer S, Homberg U. Orcokinin immunoreactivity in the accessory medulla of the cockroach Leucophaea maderae. Cell Tissue Res. 2006;325(3):589–600. doi: 10.1007/s00441-006-0155-y. [DOI] [PubMed] [Google Scholar]
- 30.Hofer S, Dircksen H, Tollback P, Homberg U. Novel insect orcokinins: Characterization and neuronal distribution in the brains of selected dicondylian insects. J Comp Neurol. 2005;490(1):57–71. doi: 10.1002/cne.20650. [DOI] [PubMed] [Google Scholar]
- 31.Dircksen H, Burdzik S, Sauter A, Keller R. Two orcokinins and the novel octapeptide orcomyotropin in the hindgut of the crayfish Orconectes limosus: Identified myostimulatory neuropeptides originating together in neurons of the terminal abdominal ganglion. J Exp Biol. 2000;203(18):2807–2818. doi: 10.1242/jeb.203.18.2807. [DOI] [PubMed] [Google Scholar]
- 32.Jensen PK, Pasa-Tolic L, Anderson GA, Horner JA, Lipton MS, Bruce JE, Smith RD. Probing proteomes using capillary isoelectric focusing-electrospray ionization Fourier transform ion cyclotron resonance mass spectrometry. Anal Chem. 1999;71(11):2076–2084. doi: 10.1021/ac990196p. [DOI] [PubMed] [Google Scholar]
- 33.Shen Y, Berger SJ, Anderson GA, Smith RD. High-efficiency capillary isoelectric focusing of peptides. Anal Chem. 2000;72(9):2154–2159. doi: 10.1021/ac991367t. [DOI] [PubMed] [Google Scholar]
- 34.Silvertand LH, Torano JS, van Bennekom WP, de Jong GJ. Recent developments in capillary isoelectric focusing. J Chromatogr A. 2008;1204(2):157–70. doi: 10.1016/j.chroma.2008.05.057. [DOI] [PubMed] [Google Scholar]
- 35.Silvertand LH, Torano JS, de Jong GJ, van Bennekom WP. Improved repeatability and matrix-assisted desorption/ionization - time of flight mass spectrometry compatibility in capillary isoelectric focusing. Electrophoresis. 2008;29(10):1985–96. doi: 10.1002/elps.200700434. [DOI] [PubMed] [Google Scholar]
- 36.Poitevin M, Morin A, Busnel JM, Descroix S, Hennion MC, Peltre G. Comparison of different capillary isoelectric focusing methods--use of “narrow pH cuts” of carrier ampholytes as original tools to improve resolution. J Chromatogr A. 2007;1155(2):230–6. doi: 10.1016/j.chroma.2007.02.013. [DOI] [PubMed] [Google Scholar]
- 37.Ollivaux C, Gallois D, Amiche M, Boscameric M, Soyez D. Molecular and cellular specificity of post-translational aminoacyl isomerization in the crustacean hyperglycaemic hormone family. FEBS J. 2009;276(17):4790–802. doi: 10.1111/j.1742-4658.2009.07180.x. [DOI] [PubMed] [Google Scholar]
- 38.Wilcockson DC, Chung JS, Webster SG. Is crustacean hyperglycaemic hormone precursor-related peptide a circulating neurohormone in crabs? Cell Tissue Res. 2002;307(1):129–138. doi: 10.1007/s00441-001-0469-8. [DOI] [PubMed] [Google Scholar]
- 39.Choi CY, Zheng J, Watson RD. Molecular cloning of a cDNA encoding a crustacean hyperglycemic hormone from eyestalk ganglia of the blue crab, Callinectes sapidus. Gen Comp Endocrinol. 2006;148(3):383–387. doi: 10.1016/j.ygcen.2006.03.003. [DOI] [PubMed] [Google Scholar]
- 40.Syka JE, Coon JJ, Schroeder MJ, Shabanowitz J, Hunt DF. Peptide and protein sequence analysis by electron transfer dissociation mass spectrometry. Proc Natl Acad Sci U S A. 2004;101(26):9528–33. doi: 10.1073/pnas.0402700101. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Coon JJ, Syka JEP, Schwartz JC, Shabanowitz J, Hunt DF. Anion dependence in the partitioning between proton and electron transfer in ion/ion reactions. Int J Mass Spectrom. 2004;236(1–3):33–42. [Google Scholar]
- 42.Xia Y, Gunawardena HP, Erickson DE, McLuckey SA. Effects of cation charge-site identity and position on electron-transfer dissociation of polypeptide cations. J Am Chem Soc. 2007;129(40):12232–43. doi: 10.1021/ja0736764. [DOI] [PubMed] [Google Scholar]
- 43.Altelaar AF, Mohammed S, Brans MA, Adan RA, Heck AJ. Improved identification of endogenous peptides from murine nervous tissue by multiplexed peptide extraction methods and multiplexed mass spectrometric analysis. J Proteome Res. 2009;8(2):870–6. doi: 10.1021/pr800449n. [DOI] [PubMed] [Google Scholar]
- 44.Swaney DL, McAlister GC, Coon JJ. Decision tree-driven tandem mass spectrometry for shotgun proteomics. Nat Methods. 2008;5(11):959–64. doi: 10.1038/nmeth.1260. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Wang J, Ma M, Chen R, Li L. Enhanced neuropeptide profiling via capillary electrophoresis off-line coupled with MALDI FTMS. Anal Chem. 2008;80(16):6168–6177. doi: 10.1021/ac800382t. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Wang JH, Chen RB, Ma MM, Li LJ. MALDI MS sample preparation by using paraffin wax film: Systematic study and application for peptide analysis. Anal Chem. 2008;80(2):491–500. doi: 10.1021/ac701614f. [DOI] [PubMed] [Google Scholar]
- 47.Coon JJ. Collisions or Electrons? Protein Sequence Analysis in the 21st Century. Anal Chem. 2009;81(9):3208–3215. doi: 10.1021/ac802330b. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Hsu YWA, Messinger DI, Chung JS, Webster SG, de la Iglesia HO, Christie AE. Members of the crustacean hyperglycemic hormone (CHH) peptide family are differentially distributed both between and within the neuroendocrine organs of Cancer crabs: implications for differential release and pleiotropic function. J Exp Biol. 2006;209(16):3241–3256. doi: 10.1242/jeb.02372. [DOI] [PubMed] [Google Scholar]
- 49.Ma MM, Wang JH, Chen RB, Li LJ. Expanding the crustacean neuropeptidome using a multifaceted mass spectrometric approach. J Proteome Res. 2009;8(5):2426–2437. doi: 10.1021/pr801047v. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Chen RB, Ma MM, Hui LM, Zhang J, Li LJ. Measurement of neuropeptides in crustacean hemolymph via MALDI mass spectrometry. J Am Soc Mass Spectrom. 2009;20(4):708–718. doi: 10.1016/j.jasms.2008.12.007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Behrens HL, Chen RB, Li LJ. Combining microdialysis, nanoLC-MS, and MALDI-TOF/TOF to detect neuropeptides secreted in the crab, Cancer borealis. Anal Chem. 2008;80(18):6949–6958. doi: 10.1021/ac800798h. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Bungart D, Dircksen H, Keller R. Quantitative determination and distribution of the myotropic neuropeptide orcokinin in the nervous system of astacidean crustaceans. Peptides. 1994;15(3):393–400. doi: 10.1016/0196-9781(94)90194-5. [DOI] [PubMed] [Google Scholar]
- 53.Simpson DC, Smith RD. Combining capillary electrophoresis with mass spectrometry for applications in proteomics. Electrophoresis. 2005;26(7–8):1291–1305. doi: 10.1002/elps.200410132. [DOI] [PubMed] [Google Scholar]
- 54.Zhang ZZ, Wang JH, Hui LM, Li LJ. Membrane-assisted capillary isoelectric focusing coupling with MALDI-FTMS for neuropeptide analysis. J Chromatogr A. 2011 doi: 10.1016/j.chroma.2011.05.072. In press. [DOI] [PMC free article] [PubMed] [Google Scholar]