Abstract
Meiotic chromosomes are organized about a proteinaceous core that forms between replicated sister chromatids. We have isolated a Caenorhabditis elegans gene, him-3, which encodes a meiosis-specific component of chromosome cores with some similarity to the yeast lateral element protein Hop1p. Antibodies raised against HIM-3 localize the protein to condensing chromosomes in early prophase I and to the cores of both synapsed and desynapsed chromosomes. In RNA interference experiments, chromosomes appear to condense normally in the absence of detectable protein but fail to synapse and form chiasmata, indicating that HIM-3 is essential for these processes. Hypomorphs of him-3, although being synapsis proficient, show severe reductions in the frequency of crossing-over, demonstrating that HIM-3 has a role in establishing normal levels of interhomolog exchange. Him-3 mutants also show defects in meiotic chromosome segregation and the persistence of the protein at the chromosome core until the metaphase I–anaphase I transition suggests that HIM-3 may play a role in sister chromatid cohesion. The analysis of him-3 provides the first functional description of a chromosome core component in a multicellular organism and suggests that a mechanistic link exists between the early meiotic events of synapsis and recombination, and later events such as segregation.
Keywords: C. elegans, chromosome core, synapsis, crossing-over, segregation
Meiosis is two specialized cell divisions that result in daughter cells with half the chromosome number of the parental cell. This is accomplished by a single round of DNA replication followed by two divisions, anaphase I and II. Anaphase I is unique in that it segregates homologous chromosomes, each composed of a pair of sister chromatids joined by a single kinetochore, from each other. Most organisms use crossing-over, cytologically evident as chiasmata, to direct the segregation of chromosomes at this division. A decrease in the frequency of crossing-over generally leads to an increase in the nondisjunction of chromosome pairs (for review, see Hawley 1988). For exchange to occur in most organisms, homologous chromosomes must find one another in the prophase nucleus and align themselves throughout their lengths, a process that usually culminates in the formation of a tripartite proteinaceous structure, the synaptonemal complex (SC) (Moses 1968; von Wettstein et al. 1984; Heyting 1996). A prerequisite to the assembly of the mature SC is the polymerization of single proteinaceous axes between sister chromatids, which have been referred to as axial elements or chromosome cores (Moses 1968). The axial elements of homologous chromosomes become aligned and equidistantly separated by the proteins that constitute the central region of the SC and are then referred to as lateral elements. The conserved, highly ordered nature of the SC has raised questions concerning its function and protein composition. Although the localization of axis-associated proteins during meiosis has been described at the ultrastructural level, their roles in meiotic processes such as pairing, recombination, and segregation have been the topic of much debate, because functional information is lacking for the most promising candidates, and few components have been identified in genetically tractable organisms in which functional analyses are feasible.
The largest advances in understanding the functions of chromosome axis-associated proteins during synapsis have been made in Saccharomyces cerevisiae. The RED1 protein associates with the cores of unsynapsed meiotic chromosomes and the SC, and then dissociates upon dissolution of the SC (Smith and Roeder 1997). Red1p is required for the assembly of the axial element and for the recruitment of another meiosis-specific chromosome component, Hop1p (Hollingsworth et al. 1990). Hop1p localizes to pachytene chromosomes in a Red1p-dependent manner and dissociates before SC disassembly (Hollingsworth et al. 1990; Smith and Roeder 1997). HOP1 mutants are defective in many aspects of chromosome metabolism, including chromosome condensation (Loidl et al. 1994), recombination, and synapsis (Hollingsworth and Byers 1989). However, neither Red1p nor Hop1p show a localization pattern on meiotic chromosomes that is consistent with a role as a fundamental building block of the axial element: Red1p distribution is relatively discontinuous along the axial/lateral element (Smith and Roeder 1997), and the localization of Hop1 to pachytene chromosomes is sparse or undetectable (Hollingsworth et al. 1990; Smith and Roeder 1997). The best characterized structural candidate is the hamster Cor1 protein and its rat homolog, SCP3. Cor1/SCP3 appears in the unsynapsed axial cores, in the lateral element of the SC, and in the cores of desynapsed chromosomes; at metaphase I, Cor1 localizes to sister centromeres, and during the transition to anaphase II, it dissociates completely (Dobson et al. 1994). This pattern of localization suggests that the protein functions in both sister chromatid arm and centromere cohesion (for review, see Bickel and Orr-Weaver 1996), a role supported by its ultrastructural localization in the lateral element (Moens 1987; Schalk et al. 1998). The persistence of Cor1/SCP3 at the axial element from early in prophase I to the anaphase II transition raises intriguing questions about the functions of this class of protein during meiosis.
Meiosis in Caenorhabditis elegans is marked by the same highly conserved features found in other sexually reproducing organisms (Albertson et al. 1997). The earliest stages of meiosis involve the recognition and alignment of homologous chromosome pairs (n = 6), and correct chromosome segregation is dependent on crossing- over and the subsequent formation of chiasmata (Hodgkin et al. 1979; Zetka and Rose 1992,1995a; Villeneuve 1994; Dernburg et al. 1998). Recently, it has been shown that crossing-over in C. elegans, like in Drosophila melanogaster, requires the activity of a yeast SPO11 homolog (Dernburg et al. 1998; McKim et al. 1998; McKim and Hayashi-Hagihara 1998), a member of a conserved family of proteins that may catalyze meiosis-specific double-stranded DNA breaks (Keeney et al. 1997). In the absence of this protein in flies and worms, reciprocal exchange (and gene conversion in at least Drosophila) is eliminated, but the assembly of the SC proceeds normally, indicating that, unlike yeast, these processes are not essential for SC formation (for review, see Roeder 1997; Dernburg et al. 1998; McKim et al. 1998).
The identification of meiosis-specific yeast chromosome components that function in highly conserved meiotic processes, like SC formation, raised the hopes that their counterparts could be identified in other organisms on the basis of homology. The identification of such homologs has not been forthcoming and mutationally defined meiotic chromosome core proteins in higher eukaryotes have remained elusive. In this study we present the molecular and genetic analyses of a meiosis-specific chromosome component in C. elegans; HIM-3 shares some similarity to yeast Hop1p and is required for synapsis and chiasma formation, indicating a possible conservation of function between the two proteins at the level of SC assembly. However, the temporal and spatial distribution of HIM-3 during meiosis suggests that it is an integral component of the chromosome core and has been adapted to function in different processes in the nematode than in yeast. him-3 hypomorphs are defective in crossing-over but proficient in synapsis, disclosing an influence on the recombination pathway independent of SC formation. him-3(e1256) mutants are also defective in chromosome segregation, and this, in conjunction with the continued association of HIM-3 with the chromosome core until the metaphaseI–anaphase I transition, suggests that the protein plays a role in sister chromatid arm cohesion. These mutant phenotypes suggest that HIM-3 may be part of a mechanistic link that exists between the early prophase events of synapsis and recombination, and later events such as segregation.
Results
him-3 mutants are defective in meiotic crossing-over
A high incidence of XO males (known as a Him phenotype) among the self-fertilization progeny of XX hermaphrodites indicates a defect in X-chromosome segregation. A subset of the group of recessive mutations that produce this phenotype also produces a large number of arrested aneuploid embryos, characteristic of mutations that decrease meiotic recombination and increase chromosome nondisjunction (for review, see Zetka and Rose 1995b). The existing two EMS-induced alleles of him-3, e1256 and e1147, differ in the severity of a similar spectrum of phenotypes and affect meiosis in both sexes (Hodgkin et al. 1979). Hermaphrodites homozygous for either allele produce normal numbers of fertilized eggs but differ in the frequency of their survival. In the case of e1256, 80% of the fertilized eggs laid die as embryos, and of the surviving progeny, 11.5% are male (data not shown). The phenotype of e1147 homozygotes is less severe; they produce normal numbers of viable progeny, and only 3.5% of the progeny are male (data not shown). To examine in detail the effects of the two existing alleles on crossover frequencies, several intervals on chromosome I and on the X chromosome were examined (Table 1). The severity of the reduction in the frequency of crossing-over in e1147 homozygotes was interval dependent and ranged from no significant change to a decrease to 63% of the wild-type frequency. Crossing-over in e1256 homozygotes was severely reduced to ∼50% in all intervals tested. This reduction was reflected by an increase in the number of achiasmate chromosomes (univalents) present in oocytes. The average number of DAPI-stained bodies present in the oocytes of e1256 hermaphrodites was 9.6 ± 1.3 (61 nuclei were scored), compatible with the presence, on average, of three bivalents and six univalents (2n = 12).
Table 1.
Crossing-over is reduced in him-3 mutants
| Genotype
|
Wild type
|
Dpy
|
Unc
|
cM (95% CI)
|
|---|---|---|---|---|
| Chromosome I | ||||
| dpy-5 unc-101/+ + | 1436 | 95 | 123 | 11.2 (9.7–12.7) |
| dpy-5 unc-101/+ +; him-3(e1256)/him-3(e1256) | 471 | 15 | 13 | 4.4 (3.0–6.2) |
| dpy-5 unc-101/+ +; him-3(e1147)/him-3(e1147) | 1934 | 164 | 164 | 12.5 (12.0–12.9) |
| unc-101 unc-54/+ + | 1959 | 222 | Unc-101 | 16.6 (14.4–18.8) |
| unc-101 unc-54/+ +; him-3(e1256)/him-3(e1256) | 512 | 25 | Unc-101 | 7.2 (4.8–10.4) |
| unc-101 unc-54/+ +; him-3(e1147)/him-3(e1147) | 1347 | 99 | Unc-101 | 10.5 (8.7–13.1) |
| Chromosome X | ||||
| unc-1 dpy-7/+ + | 1421 | 200 Dpy | 224 Unc | 21.8 (20.9–22.7) |
| unc-1 dpy-7/+ +; him-3(e1256)/him-3(e1256) | 778 | 54 Dpy | 74 Unc | 12.1 (10.4–14.7) |
| unc-1 dpy-7/+ +; him-3(e1147)/him-3(e1147) | 1587 | 201 Dpy | 222 Unc | 19.5 (18.9–21.8) |
| dpy-7 unc-3/+ + | 1793 | 250 Dpy | 277 Unc | 21.5 (20.5–22.3) |
| dpy-7 unc-3/+ +; him-3(e1256)/him-3(e1256) | 1924 | 152 Dpy | 177 Unc | 12.6 (12.1–13.0) |
| dpy-7 unc-3/+ +; him-3(e1147)/him-3(e1147) | 2060 | 166 Dpy | 191 Unc | 13.1 (12.7–13.6) |
him-3 encodes a HOP1-like protein
The him-3 locus was genetically mapped to the unc-44 unc-24 interval of chromosome IV by three-factor and deficiency mapping experiments (data not shown). The lesion associated with him-3(e1147) was fortuitously identified during the cloning of the adjacent gene, pgl-1, on the cosmid ZK381 (Kawasaki et al. 1998). Further sequencing revealed that both alleles of him-3 contain C to T transitions in the first exon, yielding a Pro to Ser substitution at amino acid 16 in e1147 and a Thr to Met substitution at amino acid 24 in e1256. The presence of these mutations in the coding region, in conjunction with the results of RNA interference experiments (see below), has led us to conclude that the him-3 locus corresponds to the open reading frame ZK381.1 identified by the Genome Sequencing Consortium.
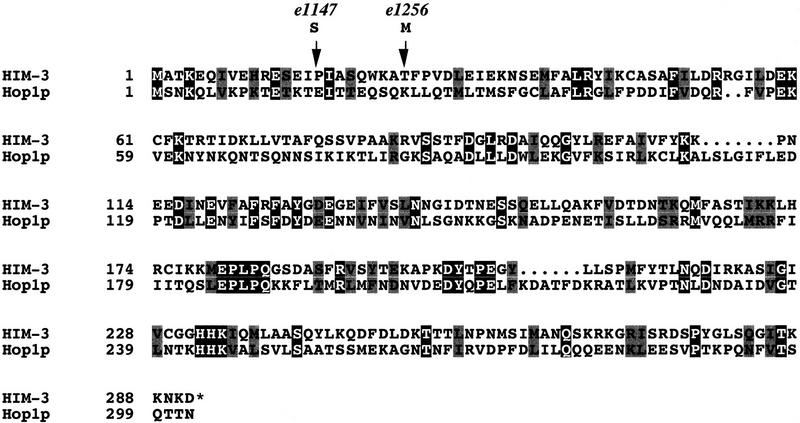
The him-3 cDNA contains a single long open reading frame of 873 nucleotides encoding a 291-amino-acid polypeptide (Fig. 1) of a predicted molecular mass of 33.1 kD. The him-3 cDNA sequence possesses an SL1-transpliced leader sequence (Krause and Hirsh 1987) and confirmed the ATG start codon, exon/intron splice sites, and the TAG translation termination signal predicted by GeneFinder. The distance from the SL1 sequence of the him-3 cDNA to the putative polyadenylation signal (AATAAA; Proudfoot and Brownlee 1976), located 204 nucleotides 3′ to the stop codon, corresponds well to the size of the 1.2-kb major transcript observed on Northern blots (Fig. 2). BLAST database searches indicate that the full-length protein sequence of HIM-3 shares 16% identity and 31% similarity to the amino-terminal 302 amino acids of Hop1p of S. cerevisiae (Fig. 1), a component of meiotic chromosomes that is essential for the formation of the SC (Hollingsworth and Byers 1989; Hollingsworth et al. 1990). A comparison of the HIM-3 and Hop1p sequences finds that they differ in two major respects: Hop1p is larger (605 vs. 291 amino acids) and has a Cys2/Cys2 finger motif with DNA-binding activity that is required for its function (Hollingsworth et al. 1990; Kironmai et al. 1998).
Figure 1.
The predicted amino acid sequence of the him-3 cDNA (GenBank accession no. AF172351) in an alignment with the first 302 amino acids of Hop1p (Hollingsworth et al. 1990), showing identical amino acids in black boxes and conserved amino acids in gray boxes. The positions of the missense mutations associated with the e1147 (amino acid 16) and e1256 (amino acid 24) are shown by arrows (see text for details).
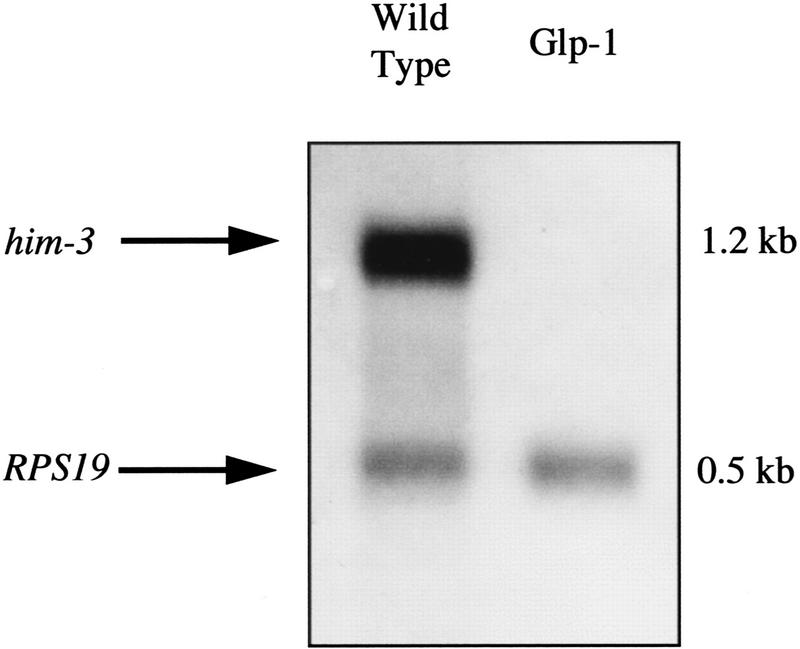
Figure 2.
him-3 expression is germ line specific. Poly(A)+ RNA was isolated from wild-type and Glp-1 hermaphrodites, a Northern blot was prepared, and it was probed with digoxigenin-labeled him-3 RNA. glp-1(q224ts) homozygotes shifted to the restrictive temperature are defective in the signal required for germ-line proliferation and produce only six to eight germ cells (Austin and Kimble 1987). The him-3 mRNA is detectable only in wild type, which possess about 1500 germ cells as adults. The transcript of the ribosomal protein gene RPS19 was used as a control for the amount of mRNA loaded in each lane.
him-3 RNA interference disrupts synapsis and chiasma formation
Because both existing him-3 alleles are missense mutations that do not change the accumulation and localization of HIM-3 (see below), RNA interference (RNAi) was used as a tool to eliminate the function of the endogenous him-3 gene and to predict the null phenotype (Fire et al. 1998).
Wild-type L4 hermaphrodites were injected with double-stranded full-length him-3 RNA, and the total number of fertilized eggs that they laid were counted (Table 2). Injected mothers from this experiment segregated 27% arrested embryos and 2.2% male progeny among the survivors. Fifteen F1 progeny from one individual exhibited a stronger meiotic nondisjunction phenotype in the germ line; the vast majority of the F2 individuals died as embryos (95.2%), and of the surviving progeny, 20% were male. The difference in the severity of the phenotypes observed in the two generations may be attributed to the fact that the injected hermaphrodites were at the fourth larval stage; spermatogenesis concludes in L4, and gametogenesis switches to continual oogenesis. Thus, the chromosome complement of the sperm in injected animals was normal, and only oogenesis was disrupted by the RNAi. In the case of the F1 individuals, both oogenesis and spermatogenesis were affected, and the resulting phenotype is likely to be a closer approximation of the null phenotype of him-3. The effectiveness of the RNA was corroborated by staining with anti-HIM-3; HIM-3 was localized to the chromosomes of wild-type control nuclei but could not be detected in the nuclei exhibiting the severe RNAi-induced phenotype (data not shown).
Table 2.
RNA interference phenotypes of him-3
| Individuals scored
|
Eggs laid
|
Dead eggs
|
Male progeny
|
Frequency of dead eggs
|
Frequency of males
|
|---|---|---|---|---|---|
| 24 P0 | 3760 | 1035 | 61 | 0.275 | 0.02 |
| 15 F1 | 1379 | 1314 | 13 | 0.952 | 0.20 |
The large frequency of arrested embryos and XO individuals segregated by the injected hermaphrodites and their progeny is suggestive of a disruption in crossing-over or chromosome segregation. To investigate the defect, the DAPI-stained gonads of injected hermaphrodites were cytologically analyzed by fluorescence microscopy (Fig. 3). The transparent gonad of C. elegans presents a precise temporal succession of nuclei passing through premeiotic events, terminating at diakinesis in hermaphrodite oocytes and in sperm maturation in males. Nuclei at each stage of meiosis have a distinct morphology that can be observed easily by staining with DAPI (Francis et al. 1995). In adults of both sexes, the most distal region of the gonad arm contains nuclei undergoing mitosis, followed by a region of transition from the mitotic cell cycle through the early stages of meiotic prophase. This is followed by a large tract of nuclei at the pachytene stage, in which the synapsed homologous chromosomes are visible as equidistantly spaced parallel ribbons. The next population of nuclei are going through diplotene; the bivalents desynapse and chromatin condenses until diakinesis (Schedl 1997).
Figure 3.
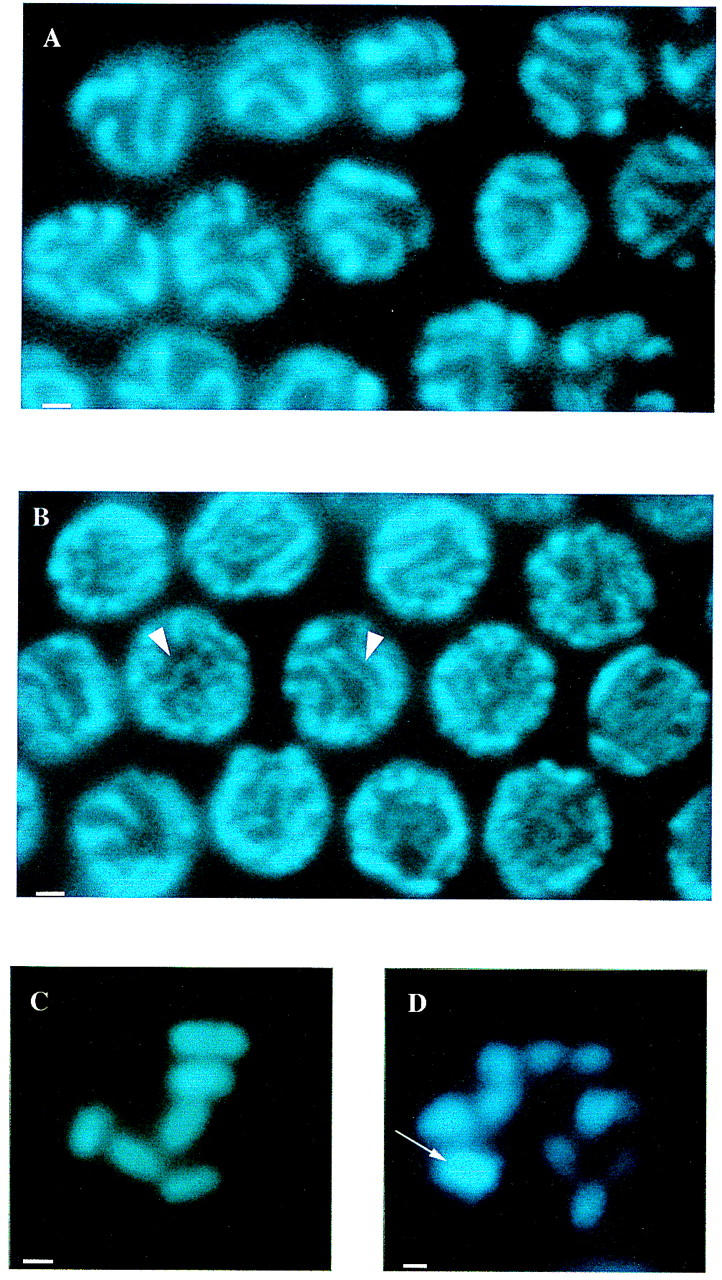
DAPI staining of RNAi nuclei shows that him-3 is essential for synapsis and chiasma formation. (A) Oocyte nuclei at the pachytene stage from wild-type hermaphrodites showing synapsed homologous chromosomes. (B) Nuclei from the pachytene zone of the germ line of hermaphrodites injected with him-3 RNA. Arrowheads indicate examples of regions of unsynapsed chromosomes. (C) The six bivalents observed in wild-type oocyte diakinesis representing 12 chromosomes joined by chiasmata. (D) RNA oocytes at diakinesis showing 12 univalents, indicating an absence of chiasmata. Arrow shows a signal from two overlapping univalents, located on different focal planes. Scale bars, 1 μm.
The chromosomes of RNAi nuclei progressively condensed from early prophase I to diakinesis (data not shown) and the univalents that appeared at diakinesis (see below) were approximately half the size of the bivalents present in control nuclei (Fig. 3C,D), suggesting that chromatin condensation was not significantly affected by the RNAi. In the pachytene region of the gonad, however, the nuclei appeared disorganized and the parallel, ribbon-like alignment of synapsed chromosomes was absent, suggesting that the RNAi had disrupted chromosome synapsis. To provide empirical evidence that the chromosomes were unsynapsed, we measured the width of the DAPI-stained chromosomes in pachytene nuclei of injected hermaphrodites and age-matched controls. The mean width of the DAPI-stained pachytene bands of injected animals (0.29 μm ± 0.05) was half the mean observed in age-matched controls (0.56 μm ± 0.06), indicating that the chromosomes were unsynapsed. The consequence of the defect at pachytene was the appearance of univalents at diakinesis; in oocyte nuclei where individual chromosomes could be resolved unambiguously, 12 univalents were observed in the nuclei of injected animals and 6 bivalents were observed in wild-type nuclei, indicating that in the absence of detectable HIM-3, chiasmata failed to form or failed to be maintained. The resulting achiasmate phenotype is identical to that reported for the null allele of the C. elegans spo-11 homolog, which encodes an enzyme required for the initiation of recombination and for which it has been shown that the lack of chiasmata can be attributed to a failure to form crossovers (Dernburg et al. 1998). Because him-3 hypomorphic mutants reduce the frequency of crossing-over, we postulate that the absence of chiasmata in the oocytes of injected animals stems from a general failure in homologous crossing-over. From these results, we conclude that in the absence of HIM-3, chromosomes condense but fail to synapse and cross over.
HIM-3 is a meiosis-specific component of chromosomes
To determine whether him-3 is expressed in adult somatic tissues, the level of him-3 mRNA present in wild-type hermaphrodites and in Glp-1 hermaphrodites lacking a germ line was compared (Fig. 2). The him-3 probe identifies a single 1.2-kb transcript in wild-type worms, but no transcript could be detected in Glp-1 adult hermaphrodites, indicating that him-3 is not expressed at a detectable level in somatic tissues but is abundantly expressed in the germ line. Moreover, the few surviving euploid F2 progeny from RNAi experiments developed normally and appeared wild type in phenotype.
To examine the localization of the protein in the germ line, L3 larvae and microdissected gonads from wild-type hermaphrodites and males were stained with an antibody raised against the carboxy-terminal portion of the HIM-3 protein. All three gamete types of C. elegans (oocyte, male spermatocyte, and hermaphrodite spermatocyte) show meiosis-specific expression of HIM-3 (Fig. 4). No protein could be detected in somatic nuclei at any stage of development or in the distal tip nuclei that are undergoing mitotic divisions. A strong signal first appears in the transition zone where HIM-3 is associated with the chromatin of early meiotic prophase nuclei. Localization of HIM-3 becomes more discrete in pachytene and diplotene as the chromosomes synapse and desynapse. The protein remains associated with the chromosomes of oocytes at diakinesis; however, it could no longer be detected in postmeiotic sperm. In L3 hermaphrodite larvae, the majority of the germ line nuclei are in the mitotic cell cycle, and the few entering meiosis are destined to become sperm. At this stage, the antibody identifies the first nuclei that are entering the pachytene stage of meiosis but does not stain the mitotic nuclei, indicating that the expression of HIM-3 during hermaphrodite spermatogenesis is identical to its expression during oogenesis and male spermatogenesis. These results have led us to conclude that HIM-3 is a meiosis-specific component of chromosomes.
Figure 4.
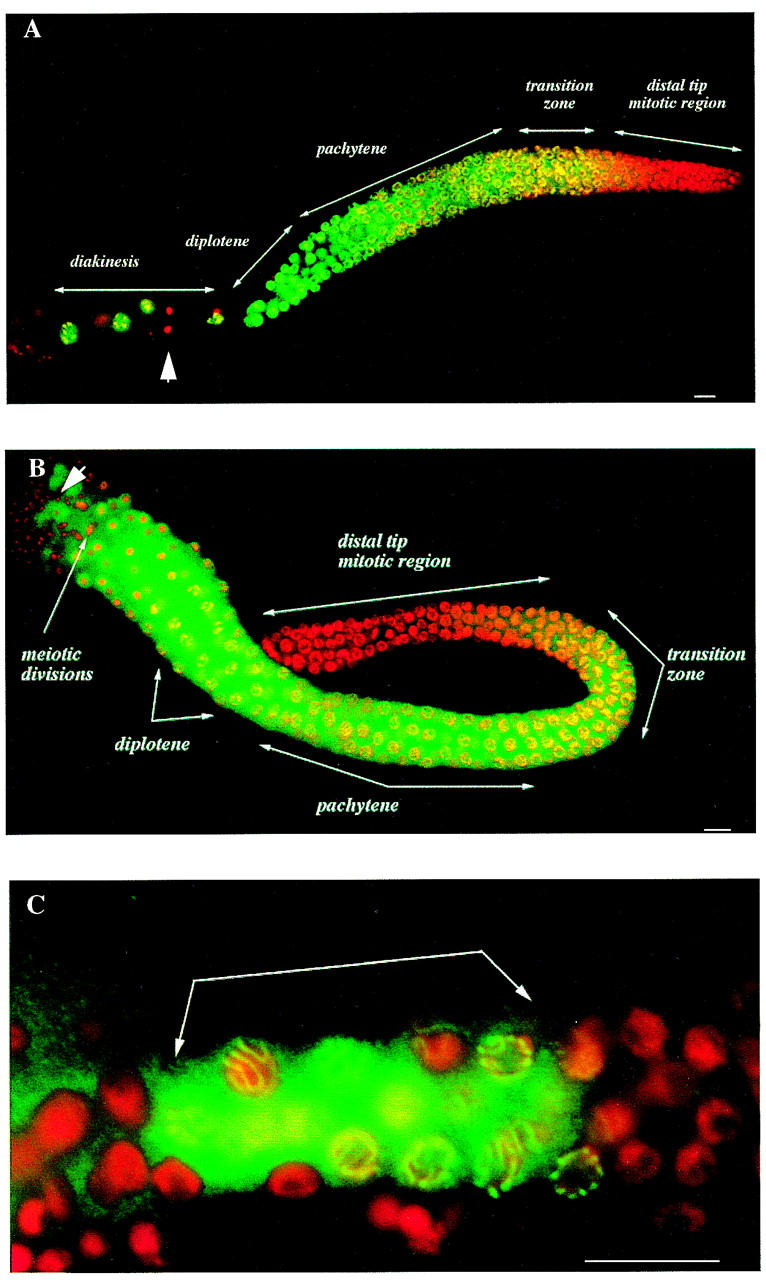
Expression of HIM-3 within the germ lines is meiosis specific. Merged images of microdissected gonad arms and an L3 larva stained with anti-HIM-3 (green) and DAPI (red). (A) Localization of HIM-3 during oogenesis. The adult hermaphrodite germ line has five distinct zones of differentiation (see text for details). Oocytes pause at diakinesis of prophase I in the proximal region of the gonad arm before ovulation. (B) Localization of HIM-3 during male spermatogenesis. The first four zones of differentiation are similar to those observed in adult hermaphrodites; however, the nuclei progress through the meiotic divisions. (C) Localization of HIM-3 during hermaphrodite spermatogenesis. An L3 larva showing HIM-3 expression only in pachytene nuclei (arrows). HIM-3 staining is not detected in the germ line nuclei undergoing mitotic divisions or in the somatic nuclei of the gonad (large arrowhead in A) or in mature sperm (large arrowhead in B). Scale bar, 10 μm.
The him-3 protein localizes to the chromosome core and associates with both unsynapsed and synapsed chromosomes
The observation that HIM-3 immunocolocalizes with meiotic chromosomes and is required for synapsis suggests that it has a structural role at the chromosome core. In whole-mount preparations of gonads, HIM-3 staining is first detected in the nuclei of the transition zone where it overlaps the pattern of DAPI staining, indicating that it is loaded onto the chromosomes early in prophase of meiosis I. To circumvent the problem of overlapping signals from different nuclei in the region, nuclei in the early stages of meiotic prophase were released from dissected gonads and then spread on slides before staining (Fig. 5A–C). HIM-3 localization assumes three general patterns that appear to be temporally distinct: nuclei with punctate staining, nuclei with longer stretches of contiguous staining that merge in places, and nuclei in which six bands (representing the six synapsed bivalents) are evident. The discrete localization of HIM-3 to the chromosomes indicates that it is not a major constituent of bulk chromatin. These staining patterns (and in particular the association of HIM-3 to the cores of unsynapsed chromosomes) are consistent with the interpretation that HIM-3 associates with the axial element (Moses 1968). The punctate pattern of HIM-3 staining is likely to represent an early stage in axial element assembly, which evolves into a contiguous stretch of staining in nuclei in which assembly is more advanced. As synapsis proceeds, the number of single elements is reduced until HIM-3 is localized to cores of the six synapsed chromosomes at pachytene.
Figure 5.
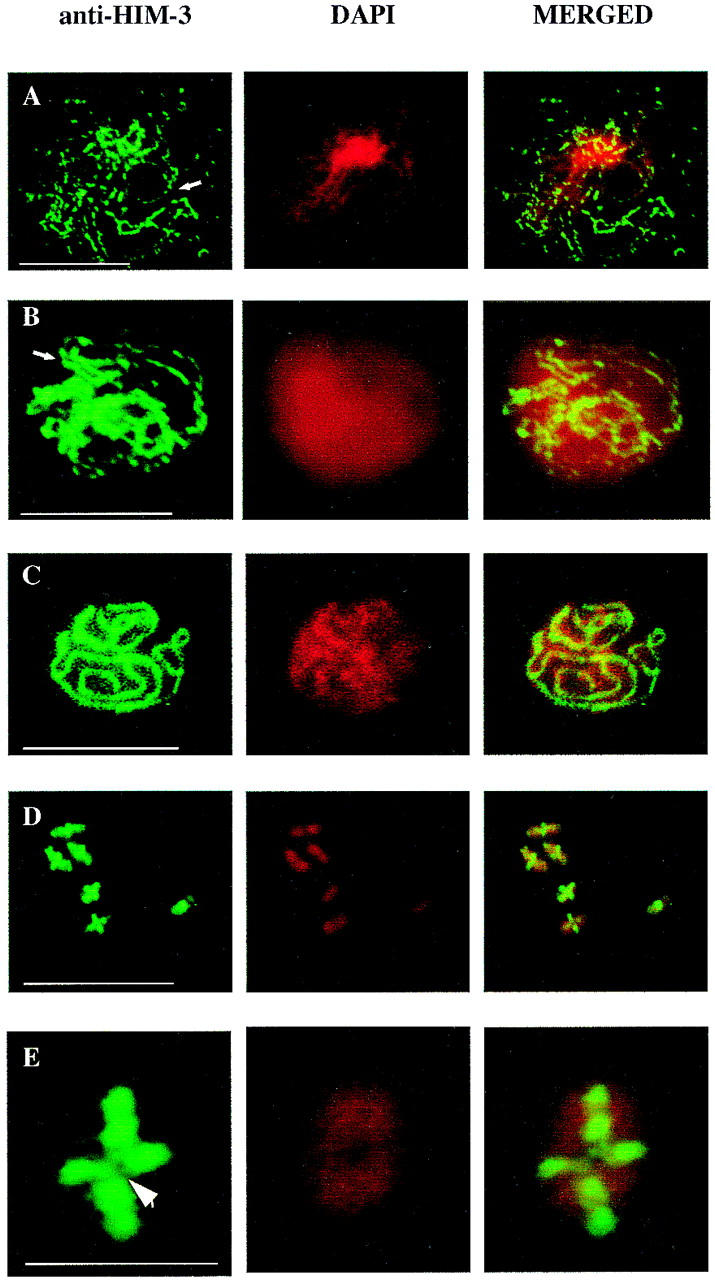
HIM-3 associates with the chromosome core. (A–C) Immunofluorescence micrographs of squashed meiotic prophase chromosomes assembling lateral elements and SC stained with anti-HIM-3. (A) Arrow shows the formation of a stretch of lateral element early in prophase that becomes contiguous in B as the chromosomes continue to condense. The diffuse appearance of the DAPI staining is probably the result of a variation in the fixation time. (C) HIM-3 colocalizes with the synaptonemal complex of the pachytene chromosomes. Scale bars, 10 μm. (D–E) Chromosomes at diakinesis stained with anti-HIM-3 showing that HIM-3 remains at the chromosome core after desynapsis. Scale bar, 10 μm. (E) A magnification of a single bivalent where HIM-3 is associated with the chromosome core and is not present at the junction of the bivalent occupied by the chiasma (arrowhead). Scale bar, 1 μm.
Because HIM-3 localizes to the axial core of wild-type meiotic chromosomes, one might expect that the phenotypes associated with loss-of-function alleles of him-3, particularly the severity of the decrease in meiotic crossing-over associated with e1256, would be the result of a disruption in synapsis. Immunofluorescence studies of e1256 and e1147 homozygotes revealed, however, that the chromosomes were synapsed at pachytene and the localization of the protein was indistinguishable from the wild-type pattern (data not shown). The mutant proteins are loaded normally onto condensing chromosomes early in prophase and localized to the SC of the fully synapsed chromosomes at pachytene and to the cores of both bivalents and univalents at diplotene, diakinesis, and metaphase I. Because RNA interference experiments suggest that the null phenotype of him-3 is asynapsis, we conclude that the amino acid substitutions in the first exon associated with e1147 and e1256 influence the frequency of interhomolog exchange rather than synapsis per se.
HIM-3 is released from the chromosome core at the metaphase—anaphase I transition
Although Hop1p and HIM-3 both localize to synapsed chromosomes, the localization of HIM-3 differs in that it remains at the chromosome core joining the sister chromatids at diakinesis and permits the localization of the chiasma that anchors the bivalent after desynapsis (Fig. 5D). No univalents and no bivalents containing two chiasmata were observed in the hundreds of wild-type oocyte nuclei examined during the course of this study. This observation correlates well with the single crossover predicted to occur between homologs based on genetic distances (ACeDB). Comparison of the genetic and physical maps has found that nearly 90% of the autosomal crossovers occur in the terminal thirds of the chromosomes (Barnes et al. 1995; for review, see Albertson et al. 1997). By measuring the position of the chiasma on 10 randomly chosen chromosomes (chiasma-distal tip length/total length), we observed that 9 occurred in the terminal 40% of the chromosome (0.27, 0.31, 0.33, 0.35, 0.37, 0.38, 0.38, 0.39, 0.40) and one occurred in the terminal 45%. Bivalents with truly terminal chiasmata were not observed. These results are compatible with the genetic data and indicate that the positions of chiasmata are reliable indicators of the physical positions of genetic exchanges and that the chiasmata do not terminalize.
After ovulation and fertilization, the oocyte pronucleus begins a complex series of events, including spindle formation, in preparation for the completion of the meiotic divisions. HIM-3 remains tightly associated with the oocyte chromosomes at metaphase I but is not detectable in the postmeiotic sperm pronucleus (Fig. 6A). Because the oocyte divisions are difficult to capture in embryo preparations, we studied the behavior of HIM-3 in the male gonad, which presents a large number of nuclei at different stages of the meiotic divisions. HIM-3 was associated with the chromosome core of prometaphase–metaphase I bivalents of sperm nuclei; however, no protein could be detected at the chromosome core once the transition to anaphase I was made (Fig. 6B). The dissociation of HIM-3 from the core of the chromosomes at the metaphase I–anaphase I transition is consistent with a role for the protein in sister chromatid arm cohesion, which is released at this time (for review, see Bickel and Orr-Weaver 1996).
Figure 6.
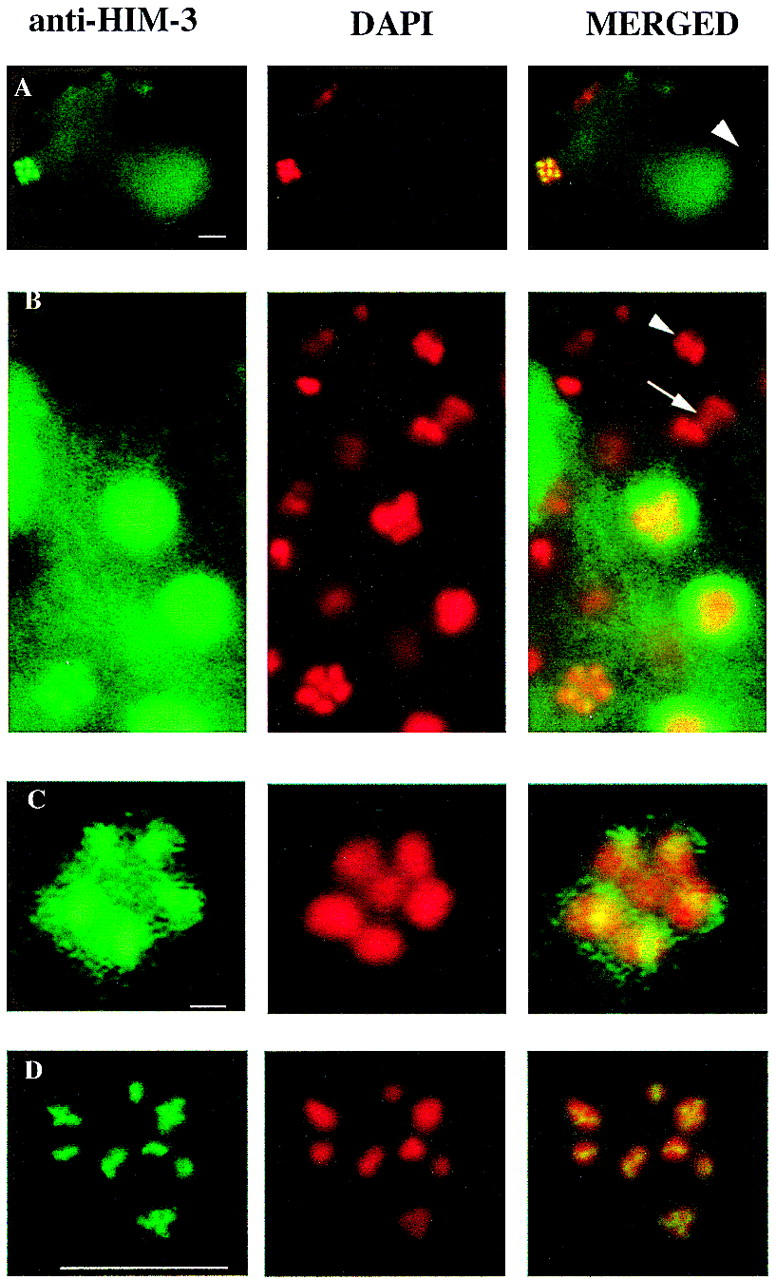
HIM-3 remains associated with the chromosome core until the metaphase I–anaphase I transition. Immunofluorescence micrographs of oocyte nuclei at metaphase and spermatocyte nuclei undergoing the meiotic divisions stained with anti-HIM-3. (A) Fertilized embryo showing that HIM-3 stains the oocyte metaphase I nucleus (left) but not the sperm nucleus that has completed meiosis (right, arrowhead). Scale bar, 10 μm. (B) Spermatocyte nuclei of males undergoing meiotic divisions. The X chromosome is centrally located in a ring formed by the autosomal bivalents at metaphase I (Albertson and Thomson 1993) in the lower left corner. The single arrow shows the lagging X chromosome, which moves to one pole at anaphase I (Albertson and Thomson 1993). HIM-3 is associated with the cores of metaphase I chromosomes but abruptly dissociates by anaphase I and is not detectable on any of the chromosomes at this time or at the second meiotic division (arrowhead). Other nuclei at earlier stages stain with anti-HIM-3, indicating that the gonad is permeable to the antibody. Scale bar, 10 μm. (C) A magnification of the metaphase I nucleus in B showing that HIM-3 is associated with the ring of five autosomal bivalents, but is not detectable above background levels on the single X chromosome. Scale bar, 1 μm. (D) Oocyte of him-3(e1256) showing that HIM-3 associates with the cores of univalents. Scale bar, 10 μm.
A him-3 hypomorph is defective in meiotic chromosome segregation
Mutations that disrupt meiotic sister chromatid cohesion result in the nondisjunction of chromosomes (for review, see Miyazaki and Orr-Weaver 1994). Based on the persistence of HIM-3 into metaphase I, we reasoned that if HIM-3 promotes sister chromatid cohesion, chromosome nondisjunction may occur in the presence of a defective protein. To examine this possibility, unc-42/unc-42 (V); lon-2 + / + dpy-7 (X); him-3(e1256)/him-3(e1256)(IV) hermaphrodites were mated to wild-type males and the non-Unc-42 cross progeny scored. Exceptional Lon-2 and Dpy-7 hermaphrodite progeny result from the fertilization of an ovum bearing two marked X chromosomes by a nullo-X sperm and are diagnostic of X-chromosome nondisjunction. It is not possible to determine at which division the nondisjunction events took place (see Materials and methods for details) because either end of the chromosome can assume kinetic activity at meiosis I in C. elegans (Albertson and Thomson 1993). e1256 homozygotes segregated 10 exceptional Lon-2 hermaphrodites and 4 Dpy-7 hermaphrodites (among 1365 wild-type hermaphrodites and 32 wild-type males), yielding an estimated frequency of 0.007. No exceptional progeny were recovered from wild-type controls (among 978 wild-type hermaphrodites and 1 wild-type male), indicating that a significant frequency of X-chromosome nondisjunction occurs in e1256 oocytes. Three of the 10 Lon-2 hermaphrodites recovered segregated Lon-2 Dpy-7 progeny, indicating that the nondisjunction event involved exchange chromosomes. The estimated frequency is likely to be an underestimate; although X-chromosome nondisjunction should lead to equal numbers of diplo-X and nullo-X gametes, the former are significantly rarer than the latter in all HIM strains tested, suggesting that diplo-X ova have reduced viability (Hodgkin et al. 1979) or that X chromosomes are lost when they fail to be included in reformed daughter nuclei. Because HIM-3 could no longer be detected on wild-type chromosomes after the transition to anaphase I and all of the nondisjunction events recovered can be explained by chromosome missegregation at the first division, we feel the most consistent interpretation of the nondisjunction phenotype associated with e1256 is a defect in sister chromatid arm cohesion.
HIM-3 fails to localize to the obligately achiasmate X chromosome in the male
During the course of this study we observed that HIM-3 associated with the cores of univalents in the oocytes of recombination-defective mutants including Him-3 (Fig. 6D), Him-14, and Spo-11 (data not shown). In C. elegans males, half of the meiotic nuclei contain a single X chromosome and half are nullo-X. We could detect no HIM-3 antigen at the core of the X chromosome at metaphase (Fig. 6B,C), suggesting that either its organization differs from that of the autosomal bivalent axial elements or that chromosomes without pairing partners generally fail to recruit HIM-3. Because the univalents observed in recombination-defective mutant backgrounds differ from the single X chromosomes in spermatocytes in that they possess intact pairing partners and undergo synapsis (Dernburg et al. 1998; this paper), we examined the ability of two free duplications, sDp2 (I;f) and sDp3 (III;f), to recruit HIM-3. In 20 oocyte nuclei of each genotype, we observed HIM-3 staining at the cores of both duplications, suggesting that the duplications were organized about a single core, like the other chromosomes. The association of HIM-3 with univalents and with sDp2, which is incapable of pairing for recombination with chromosome I because it lacks the HRR (homolog recognition region; Rose et al. 1984), indicates that HIM-3 localizes to both recombination competent and incompetent chromosomes. Its recruitment to chromosomes cores is not ubiquitous, however, because it is absent at the core of the lone X chromosome in male spermatocytes. This suggests that X-chromosome organization in the male is different from the autosomes and that HIM-3 is not required for its condensation during meiosis.
Discussion
Synapsis in C. elegans requires HIM-3
RNA interference experiments indicate that HIM-3 is essential for synapsis because in the absence of the protein, chromosomes fail to synapse. Because HOP1 mutants also fail to form an SC (Hollingsworth and Byers 1989), both proteins perform an essential and possibly conserved function necessary for synapsis. A recent alignment of proteins functioning in mitotic checkpoints, chromosome synapsis, and DNA repair, including HIM-3 and Hop1p, has suggested the existence of a common conserved structural domain, the HORMA domain (for Hop1p, Rev7p, and MAD2; Aravind and Koonin 1998). However, the function of this region, which includes almost the entire protein sequence of HIM-3 (236 of 239 amino acids) and excludes the DNA-binding domain of Hop1p, is unknown. The role of HIM-3 at the chromosome core appears to be structural. No previously described catalytic or DNA-binding domain is evident in the protein sequence, and the localization of HIM-3 during meiosis mirrors the pattern one would expect of an integral building block; the protein associates with condensing chromatin in the earliest stages of prophase I and localizes discretely to the cores of both synapsed and desynapsed chromosomes.
HIM-3 is essential for crossing-over and chiasma formation
We have shown that the recombination-defective phenotype of him-3 mutants occurs in different contexts: the failure to synapse and form chiasmata in the absence of the protein and the failure of synapsed chromosomes to undergo appropriate levels of exchange in the hypomorphic mutants. RNA interference experiments have shown that in the absence of the him-3 protein, homologous chromosomes fail to synapse at pachytene, no chiasmata are formed, and the chromosomes appear as univalents at diakinesis. This observation is most easily reconciled with a model in which exchanges simply do not occur between unsynapsed chromosomes in C. elegans. In this context, the effect of HIM-3 on recombination is indirect; crossing-over is eliminated because HIM-3 is required for synapsis. In null mutants of HOP1, however, interhomolog exchange is severely reduced (Hollingsworth and Byers 1989) but not eliminated, experimental evidence that has contributed to the model that synapsis is not essential for recombination in budding yeast (for review, see Roeder 1997).
The observation that, in the absence of HIM-3, chromosomes fail to synapse or form chiasmata, however, does not preclude a direct influence of HIM-3 on the recombination pathway. In him-3(e1256) homozygotes, crossing-over was reduced in all intervals tested at least twofold, and 80% of the embryos died as a result of aneuploidy. Surprisingly, the pachytene chromosomes were synapsed along their lengths, and the mutant protein was localized to the chromosomes in a manner indiscernible from wild type. At diakinesis, however, univalents appeared in oocytes, indicating that the mutation results in a reduced number of chiasmata.
Several models could explain the effect of mutant HIM-3 on crossing-over despite the presence of a SC, but any explanation is complicated by the probability that components of the axial element are involved in numerous interdependent processes. First, one role of the axial element may be to provide a structure to fix the chromatin such that preferred sites of exchange are exposed; it is formally possible that, in the presence of mutant HIM-3, this access is impeded and the frequency of crossing-over is reduced. The lack of an obvious DNA-binding motif in the protein, however, and the normal size of the chromosomes at pachytene and diakinesis in RNA interference nuclei suggest that HIM-3 is unlikely to play a major role in chromatin condensation. Second, him-3 mutants might be defective in the process of crossover maturation, but evidence exists that is incompatible with such a role. The decrease in the frequency of crossing-over associated with him-3(e1256) mutants is proportionally reflected in an increase in the number of univalents at diakinesis, suggesting that the interhomolog recombination events that do occur are correctly processed to yield chiasmata. Furthermore, rad-4, a radiation and MMS (methyl methanesulfonate)-sensitive mutant, nonspecifically suppresses the nondisjunction phenotype of several recombination-defective him mutations, including him-3 (Hartman and Herman 1982). This suggests that the rad-4 effect is due to an increase in DNA double-strand breaks and that these events are effectively processed to yield chiasmata in the presence of mutant HIM-3, thereby decreasing the frequency of nondisjunction. A third possibility, which we favor, is that HIM-3 has a role in establishing the preference for nonsister chromatids as partners for exchange events, a function that has also been ascribed to Hop1p (Schwacha and Kleckner 1994). In the presence of the mutant protein, one would predict that a normal number of events are initiated but that these would occur disproportionately between sister chromatids, a consequence of which would be the lack of a chiasma between homologs to direct segregation at metaphase I.
HIM-3 promotes chromosome segregation and may function in sister chromatid cohesion at the first meiotic division
The observation that HIM-3 remains bound to the chromosome core after desynapsis is suggestive of a function for HIM-3 that is distinct from synapsis or partner choice during recombination. The localization pattern of HIM-3 is similar to Cor1, a mammalian component of the axial element that is a promising candidate sister chromatid cohesion molecule, but for which no functional information exists. Two types of sister chromatid cohesion exist during the meiotic divisions (Maguire et al. 1993; Bickel and Orr-Weaver 1996; Nicklas 1997; for review, see Miyazaki and Orr-Weaver 1994). The first provides cohesion along the arms from prophase I to the onset of anaphase I, which stabilizes chiasmata and allows the proper orientation of homologous kinetochores to opposite poles. The second maintains the physical association between sister kinetochores until their segregation at the equational division. The loss of sister chromatid cohesion during meiosis occurs in two successive steps to reflect these dual functions: At the first division cohesion is lost between sister chromatid arms, and at the second division it is lost between sister chromatid centromeres. Like HIM-3, Cor1 appears in the unsynapsed axial cores, in the lateral element of the SC, and in the cores of desynapsed chromosomes (Dobson et al. 1994). At metaphase I, Cor1 localizes to sister centromeres and then finally dissociates during the transition to anaphase II, consistent with functions in both sister chromatid arm cohesion and centromere cohesion. During metaphase I, HIM-3 remains at the axial core, but once the transition to anaphase I has been completed, the protein can no longer be detected on the chromosomes, a localization pattern suggestive of a role for HIM-3 in promoting sister chromatid arm cohesion. Consistent with this interpretation, we recovered a significant frequency of nondisjunction events from e1256 oocytes that could originate from the missegregation of exchange chromosomes at anaphase I. The meioisis-specific expression of HIM-3 and the failure to detect prematurely separated chromatids in e1256 or RNAi oocyte nuclei (12 chromosomes rather than 24 chromatids are visible) indicate that the protein is not necessary for sister chromatid cohesion in mitosis and prophase I of meiosis. Instead, HIM-3 may act to promote arm cohesion within bivalents to stabilize chiasmata and establish tension at the metaphase I plate (Maguire 1974; Nicklas 1997; for review, see Carpenter 1994). In future studies of HIM-3, it will be of particular interest to determine whether the release of the protein during the metaphase I – anaphase I transition is functionally related to the release of sister chromatid arm cohesion, thereby demonstrating a role for HIM-3 in chiasma binding. An interesting observation in this respect is the failure of the obligately achiasmate X chromosome in the male to recruit the him-3 protein.
In this study we have found that three distinct phenotypes are associated with defects in him-3: asynapsis, reductions in interhomolog exchange, and meiotic nondisjunction. How can a single protein, which appears to be structural, disrupt such temporally distinct processes? A simple model that reconciles these observations is that during meiosis I, HIM-3 has two functions at the chromosome core: to promote sister chromatid cohesion and to act at the interface between the axial element and components of the SC to facilitate its assembly. In the absence of him-3, the chromosomes would be predicted to condense normally but fail to synapse, similar to the results of RNAi experiments. In him-3 hypomorphs, which assemble SC but show reduced levels of crossing-over and increased meiotic nondisjunction, the phenotypes can be explained by a disruption of sister chromatid cohesion. Because the protein is produced and loaded normally onto the chromosomes, the SC is assembled and the chromosomes synapse. However, the defect in the protein disrupts sister chromatid cohesion, resulting in a reduction in nonsister chromatid recombination during prophase I and the missegregation of exchange chromosomes during anaphase I. This scenario derives some support from the observation that mutations in RED1, whose protein product is required for sister chromatid cohesion during yeast meiosis (Bailis and Roeder 1998), result in an increase in recombination between sister chromatids (Schwacha and Kleckner 1997), in asynapsis, and in the failure of crossovers to ensure proper segregation (Rockmill and Roeder 1990), indicating that defects in sister chromatid cohesion can disrupt several meiotic processes.
Materials and methods
General methods and strains
C. elegans strains were cultured following the methods described by Brenner (1974). Experiments were performed at 20°C unless otherwise noted. The mutations used in this study were obtained from either the Caenorhabditis Genetic Center (University of Minnesota, St. Paul) or Ann Rose (University of British Columbia, Vancouver). The glp-1(q224) strain was the gift of Judith Kimble (University of Wisconsin, Madison). The following mutations were used in this study: LG I, dpy-5(e61), unc-101(m1), unc-54(e190); LG III, glp-1(q224ts); LG IV, him-3(e1147), him-3 (e1256); LG X, unc-1(e719), dpy-3(e27), lon-2(e678), dpy-7(e88), unc-3(e151).
Molecular analysis of him-3
him-3(+) cDNAs were isolated by screening a Lambda Zap cDNA library provided by R. Barstead (Barstead and Waterston 1989). Two independent isolates yielded clones of the same size (1.2 kb). cDNA H3.1 was sequenced in its entirety on one strand and compared with the genomic sequence generated by the Genome Sequencing Consortium using the GeneFinder program. Standard techniques were used to obtain the genomic and cDNA sequence of him-3 (Sambrook et al. 1989). To identify the lesions associated with him-3(e1147 ) and him-3(e1256), genomic DNA prepared from mutant homozygotes and wild-type controls was used to amplify him-3 sequences by PCR using standard conditions (Sambrook et al. 1989). Two PCR products that spanned the entire region were directly sequenced using gene-specific primers. The mutation observed in e1147 was confirmed by independent sequencing in two laboratories. Sequence comparisons and alignments were obtained by the use of GCG (version 8; Devereux et al. 1984) and BLAST (Altschul et al. 1990) programs. BLAST was accessed either through the National Center for Biotechnology Information (NCBI) web site or through the Genome Sequencing Center.
Northern blot analysis
Mixed stage wild-type RNA was isolated as described by Sulston and Hodgkin (1988). RNA from glp-1(q224) homozygotes lacking a germ line was isolated as follows: Glp-1 hermaphrodites were grown in a 1-liter liquid culture at the permissive temperature (15°C) until saturation. The culture was allowed to settle overnight at 4°C, and the pellet of worms was treated with hypochlorite to isolate embryos. These embryos were placed back in liquid culture and shifted to the restrictive temperature (25°C) and allowed to develop until adulthood. Total RNA was isolated using Trizol (Life Technologies) according to the manufacturer’s instructions. Poly(A)+ RNA was isolated using PolyATract System IV (Promega). Electrophoresis of RNA in denaturing formaldehyde gels and transfer to Tropilon Plus positively charged nylon membranes (Tropix) followed the procedures described by Sambrook et al. (1989). Digoxigenin-labeled full-length RNA probes were prepared from the him-3 cDNA using a DIG RNA-labeling kit and hybridized using the conditions recommended by the manufacturer for the CDP–Star chemiluminescent substrate (Boehringer Mannheim).
Production of antibodies
PCR fragments encoding the amino-terminal HIM-3 amino acids 1–209 and the carboxy-terminal 107–291 were cloned in-frame with the six histidine-encoding tag of the pQE32 vector (Qiagen). The inserts of the resulting plasmids (pMZ14 and pMZ13, respectively) were sequenced to ensure no errors were introduced during the PCR. The recombinant protein was expressed and purified by nickel-nitrilo-tri-acetic acid (Ni–NTA) metal-affinity chromatography under denaturing conditions as described by Qiagen and dialyzed against phosphate-buffered saline (PBS). Rabbits were immunized according to standard protocols (Harlow and Lane 1988) with either the amino-terminal or carboxy-terminal protein fragment. The resulting sera identified both the fusion protein and a band at the length predicted for HIM-3 on a Western blot containing mixed-stage C. elegans extracts (data not shown). The specificity of HIM-3 antibodies was demonstrated by the following criteria: Identical staining patterns were observed from three different polyclonal antisera (one generated against the amino-terminal portion and two against the carboxy-terminal portion), no staining was observed using preimmune sera (or with the FITC conjugate alone), and no staining was detected in RNA interference experiments. To reduce the background obtained during staining of whole worms or microdissected gonads, sera raised against the carboxy-terminal portion of the protein were affinity-purified against nitrocellulose-bound antigen and eluted with 0.2 m glycine (pH 2.8)/1 mm EGTA (Sambrook et al. 1989). All antibody stainings presented in this study use this purified preparation.
Antibody stainings
Antibody staining of whole larvae was performed using the method described by Finney and Ruvkun (1990). For in situ antibody staining of intact gonads, specimens were prepared according to Francis et al. (1995). To examine early prophase nuclei, several cuts were made to the distal tip of the gonad with a syringe to release the nuclei. Nuclei were spread by placing a 20 × 20-mm coverslip over the specimens, incubating on dry ice for 10 min, popping off the coverslip and fixing in cold methanol and acetone, air-drying, and washing twice in PBS. To examine the post-fertilization meiotic divisions, embryos were prepared as described by Miller and Shakes (1995). In all cases, specimens were incubated with anti-HIM-3 diluted 1:100 in AbA (1% BSA, 0.5% Triton-X-100, 0.05% sodium azide, 1 mm EDTA in PBS) overnight at 4°C, washed three times for 10 min in AbB (AbA with 0.1% BSA), incubated for 30 min at room temperature with FITC-conjugated AffiniPure goat anti-rabbit IgG (1:200; Jackson) diluted in AbA, washed three times for 10 min in PBS/0.1% Tween 20, and mounted in Vectashield mounting medium (Vector) containing 2 μg/ml DAPI.
Specimens were observed using a Zeiss Axioplan epifluorescence microscope, and images were taken with a Hamamatsu chilled CCD camera (C5985) and processed with the Argus-20 Image Processor (Hamamatsu). DAPI and immunofluorescence images were processed, merged, and differentiated with false color using Adobe Photoshop (4.0).
RNA interference experiments
To investigate the null phenotype of him-3, RNA experiments were performed as described previously (Fire et al. 1998). Interference with endogenous him-3 was assayed in a wild-type genetic background. Two other him-3-like genes have been identified in C. elegans but were not expected to be affected by interference with him-3 RNA because of their limited sequence similarity. The specificity of the RNAi is supported by the failure of other injected dsRNAs to produce the him-3 RNAi phenotype (Fire et al. 1998; F. Palladino, pers. comm.). Early L4 hermaphrodites were injected with 250 μg/ml dsRNA in the pachytene zone of each gonad arm and transferred to fresh culture plates every 12–16 hr over the course of 3 days. The eggs laid in every brood were counted, and the number and sex of surviving progeny were scored 3 days later. No interference phenotype was detectable during early broods, as unaffected nuclei are cleared from the gonad at this time. However, a strong interference phenotype appeared from the third or fourth egg lay onwards.
To examine the interference phenotype cytologically in pachytene nuclei, injected worms were cultured until the first arrested embryos could be detected on the plates. The gonads of the injected hermaphrodites and age-matched wild-type controls were then microdissected, fixed, and stained as described above. Once mounted, pictures were taken from the same region of the pachytene zone of wild-type and injected animals. Images were processed to normalize contrast and brightness between the two data sets, and the width of three nonoverlapping DAPI-stained bands were measured in 20 random nuclei from each set. When possible, the widest and narrowest point of each band was measured; however, band width was very uniform in each set.
Measuring the frequency of crossing-over
Recombination frequencies in the hermaphrodite were measured by scoring the number of recombinant progeny of a cis-heterozygote, using the formula P = 1 − (1 − 2R)1/2, where P was the frequency of recombination, and R was the number of visible recombinant individuals divided by the total progeny number (Brenner 1974). Because the double homozygote class was not scored because of its reduced viability, the total progeny number was estimated by 4/3 X (number of wild types + one recombinant class). In some intervals, recovery of only one recombinant class was possible, and in these instances, R = 2 X (one recombinant class) divided by the total progeny number. Male progeny were not included in the calculations because they could not be progeny tested. The 95% confidence intervals (CIs) were estimated using the statistics of Crow and Gardner (1959).
Measuring meiotic nondisjunction
Meiotic nondisjunction is most easily measured in an assay for the matriclinous inheritance of X-linked markers from the hermaphrodite oocyte. lon-2 and dpy-7 are centrally located markers 4 map units apart (M. Zetka, unpubl.). To determine whether him-3(e1256) mutants are defective in segregation, him-3/him-3(IV); unc-42/ +(IV); lon-2/0(X) males were mated to him-3/him-3; unc-42/unc-42; dpy-7/dpy-7 hermaphrodites, and the resulting Unc-42 progeny (of the genotype him-3/him-3; unc-42/unc-42; lon-2 + /+ dpy-7) were picked as immature L4s and mated to wild-type males. These hermaphrodites were individually plated after 24 hr, and all their non-Unc-42 progeny were counted (3 X hermaphrodites were not scored). One hermaphrodite exhibited a segregation pattern characteristic of a 3 X individual and was removed from the data set. Exceptional Lon-2 segregants were then progeny tested to determine whether they were recombinant (lon-2 dpy-7/lon-2 + ) or nonrecombinant (lon-2/lon-2). Because either end of the chromosome can have kinetic activity at the first division (Albertson and Thomson 1993), it is impossible to determine at which division the nondisjunction event occurred. For example, there are four possible ways of generating an ovum with two lon-2 marked chromosomes: (1) an exchange between the markers on a chromosome where the right end assumes kinetic activity (resulting in lon-2 +/+ + and lon-2 dpy-7/ + dpy-7 chromosomes), followed by nondisjunction at the first division and correct segregation at the second, (2) an exchange between the markers on a chromosome in which the left end assumes kinetic activity (resulting in lon-2 + /lon-2 dpy-7 and + + / + dpy-7 chromosomes), followed by correct segregation at the first division and nondisjunction at the second, (3) an exchange event between lon-2 and the left centromere (yielding two lon-2 + / + dpy-7 chrmosomes), followed by nondisjunction at the first division and correct segregation at the second, and (4) nondisjunction at the second division. To provide an indication of the frequency of nondisjunction, the total progeny number was estimated by 2 X [wild types (XX + XO) + Lon-2 (XX) + Dpy-7(XX)]. This calculation was based on the recovery of Lon-2 hermaphrodites and the assumptions that nondisjunction occurred at the first meiotic division and that the left end adopted centromeric activity.
Acknowledgments
We would like to thank Adelaide Carpenter, Kim McKim, Anne Villeneuve, Ann Rose, Abby Dernburg, and members of our laboratory for discussion and critical reading of the manuscript. We are especially grateful to Peter Moens for technical advice and encouragement. Some strains were supplied by the Caenorhabditis Genetics Center, which is funded by the National Institutes of Health (NIH) National Center for Research Resources. This research was supported by NIH grant GM34059 to S.S. and by Swiss National Foundation grant 31-40776.94 to F.M.
The publication costs of this article were defrayed in part by payment of page charges. This article must therefore be hereby marked ‘advertisement’ in accordance with 18 USC section 1734 solely to indicate this fact.
Footnotes
E-MAIL mczetka@ucdavis.edu; FAX (530) 752-1185.
References
- Albertson DG, Thomson JN. Segregation of holocentric chromosomes at meiosis in the nematode, Caenorhabditis elegans. Chromosome Res. 1993;1:15–26. doi: 10.1007/BF00710603. [DOI] [PubMed] [Google Scholar]
- Albertson DG, Rose AM, Villeneuve AM. Chromosome organization, mitosis, and meiosis. In: Riddle DL, Blumenthal T, Meyer BJ, Priess JR, editors. C. elegans II. Cold Spring Harbor, NY: Cold Spring Harbor Laboratory Press; 1997. pp. 47–78. [PubMed] [Google Scholar]
- Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ. Basic local alignment search tool. J Mol Biol. 1990;215:403–410. doi: 10.1016/S0022-2836(05)80360-2. [DOI] [PubMed] [Google Scholar]
- Aravind L, Koonin EV. The HORMA domain: A common structural denominator in mitotic checkpoints, chromosome synapsis and DNA repair. Trends Biochem. 1998;32:284–286. doi: 10.1016/s0968-0004(98)01257-2. [DOI] [PubMed] [Google Scholar]
- Austin J, Kimble, J J. glp-1 is required in the germ line for regulation of the decision between mitosis and meiosis in Caenorhabditis elegans. Cell. 1987;51:589–599. doi: 10.1016/0092-8674(87)90128-0. [DOI] [PubMed] [Google Scholar]
- Bailis JM, Roeder GS. Synaptonemal complex morphogenesis and sister-chromatid cohesion require Mek1-dependent phosphorylation of a meiotic chromosomal protein. Genes & Dev. 1998;12:3551–3563. doi: 10.1101/gad.12.22.3551. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Barnes TM, Kohara Y, Coulson A, Hekimi S. Meiotic recombination, non-coding DNA and genomic organization in Caenorhabditis elegans. Genetics. 1995;141:159–179. doi: 10.1093/genetics/141.1.159. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Barstead RJ, Waterston RH. The basal component of the nematode dense body is viniculin. J Biol Chem. 1989;264:10177–10185. [PubMed] [Google Scholar]
- Bickel SE, Orr-Weaver TL. Holding chromatids together to ensure they go their separate ways. Bioessays. 1996;18:293–300. doi: 10.1002/bies.950180407. [DOI] [PubMed] [Google Scholar]
- Brenner S. The genetics of Caenorhabditis elegans. Genetics. 1974;77:71–94. doi: 10.1093/genetics/77.1.71. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Carpenter AT. Chiasma function. Cell. 1994;77:959–962. doi: 10.1016/0092-8674(94)90434-0. [DOI] [PubMed] [Google Scholar]
- Crow EL, Gardner RS. Confidence intervals for the expectation of a Poisson variable. Biometrika. 1959;46:441–453. [Google Scholar]
- Dernburg AF, McDonald K, Moulder G, Barstead R, Dresser M, Villeneuve AM. Meiotic recombination in C. elegans initiates by a conserved mechanism and is dispensible for homologous chromosome synapsis. Cell. 1998;94:387–398. doi: 10.1016/s0092-8674(00)81481-6. [DOI] [PubMed] [Google Scholar]
- Devereux J, Haeberli P, Smithies O. A comprehensive set of sequence analysis programs for the VAX. Nucleic Acids Res. 1984;12:387–395. doi: 10.1093/nar/12.1part1.387. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dobson MJ, Pearlman RE, Karaiskakis A, Spyropoulos B, Moens PB. Synaptonemal complex proteins: Occurrence, epitope mapping and chromosome disjunction. J Cell Sci. 1994;107:2749–2760. doi: 10.1242/jcs.107.10.2749. [DOI] [PubMed] [Google Scholar]
- Finney M, Ruvkun GB. The unc-86 gene product couples cell lineage and cell identity in C. elegans. Cell. 1990;63:895–905. doi: 10.1016/0092-8674(90)90493-x. [DOI] [PubMed] [Google Scholar]
- Fire A, Xu S, Montgomery MK, Kostas SA, Driver SE, Mello CC. Potent and specific genetic interference by double-stranded RNA in Caenorhabditis elegans. Nature. 1998;391:806–811. doi: 10.1038/35888. [DOI] [PubMed] [Google Scholar]
- Francis R, Barton MK, Kimble J, Schedl T. gld-1, a tumor suppresor gene required for oocyte development in Caenorhabditis elegans. Genetics. 1995;139:579–606. doi: 10.1093/genetics/139.2.579. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Harlow E, Lane D. Antibodies: A laboratory manual. Cold Spring Harbor, NY: Cold Spring Harbor Laboratory Press; 1988. pp. 100–114. [Google Scholar]
- Hartman PS, Herman RK. Radiation-sensitive mutants of Caenorhabditis elegans. Genetics. 1982;102:159–178. doi: 10.1093/genetics/102.2.159. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hawley RS. Exchange and chromosomal segregation in eukaryotes. In: Kucherlapati R, Smith GR, editors. Genetic recombination. Washington, DC.: American Society for Microbiology; 1988. pp. 497–527. [Google Scholar]
- Heyting C. Synaptonemal complexes: Structure and function. Curr Opin Cell Biol. 1996;8:389–396. doi: 10.1016/s0955-0674(96)80015-9. [DOI] [PubMed] [Google Scholar]
- Hodgkin J, Horvitz HR, Brenner S. Nondisjunction mutants of the nematode Caenorhabditis elegans. Genetics. 1979;91:67–94. doi: 10.1093/genetics/91.1.67. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hollingsworth NM, Byers B. HOP1: A yeast meiotic pairing gene. Genetics. 1989;121:445–462. doi: 10.1093/genetics/121.3.445. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hollingsworth NM, Goetsch L, Byers B. The HOP1 gene encodes a meiosis-specific component of yeast chromosomes. Cell. 1990;61:73–84. doi: 10.1016/0092-8674(90)90216-2. [DOI] [PubMed] [Google Scholar]
- Kawasaki I, Shim YH, Kirchner J, Kaminker J, Wood WB, Strome S. PGL-1, a predicted RNA-binding component of germ granules, is essential for fertility in C. elegans. Cell. 1998;94:635–645. doi: 10.1016/s0092-8674(00)81605-0. [DOI] [PubMed] [Google Scholar]
- Keeney S, Giroux CN, Kleckner N. Meiosis-specific DNA double-strand breaks are catalyzed by Spo11, a member of a widely conserved protein family. Cell. 1997;88:375–384. doi: 10.1016/s0092-8674(00)81876-0. [DOI] [PubMed] [Google Scholar]
- Kironmai KM, Muniyappa K, Friedman DB, Hollingsworth NM, Byers B. DNA-binding activities of Hop1 protein, a synaptonemal complex component from Saccharomyces cerevisiae. Mol Cell Biol. 1998;18:1424–1435. doi: 10.1128/mcb.18.3.1424. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Krause M, Hirsh D. A trans-spliced leader sequence on actin mRNA in Caenorhabditis elegans. Cell. 1987;49:753–761. doi: 10.1016/0092-8674(87)90613-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Loidl J, Klein F, Scherthan H. Homologous pairing is reduced but not abolished in asynaptic mutants of yeast. J Cell Biol. 1994;125:1191–1200. doi: 10.1083/jcb.125.6.1191. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Maguire MP. Letter: The need for a chiasma binder. J Theor Biol. 1974;48:485–487. doi: 10.1016/s0022-5193(74)80017-2. [DOI] [PubMed] [Google Scholar]
- Maguire MP, Riess RW, Paredes AM. Evidence from a maize desynaptic mutant points to a probable role of synaptonemal complex central region components in provision for subsequent chiasma maintenance. Genome. 1993;36:797–807. doi: 10.1139/g93-105. [DOI] [PubMed] [Google Scholar]
- McKim KS, Hayashi-Hagihara A. mei-W68 in Drosophila melanogaster encodes a Spo11 homolog: Evidence that the mechanism for initiating meiotic recombination is conserved. Genes & Dev. 1998;12:2932–2942. doi: 10.1101/gad.12.18.2932. [DOI] [PMC free article] [PubMed] [Google Scholar]
- McKim KS, Green MB, Sekelsky JJ, Chin G, Steinberg C, Khodosh R, Hawley RS. Meiotic synapsis in the absence of recombination. Science. 1998;279:876–878. doi: 10.1126/science.279.5352.876. [DOI] [PubMed] [Google Scholar]
- Miller DM, Shakes DC. Immunofluorescence microscopy. In: Epstein HF, Shakes DC, editors. Caenorhabditis elegans: Modern biological analysis of an organism. San Diego, CA: Academic Press; 1995. pp. 365–394. [Google Scholar]
- Miyazaki WY, Orr-Weaver TL. Sister-chromatid cohesion in mitosis and meiosis. Annu Rev Genet. 1994;28:167–187. doi: 10.1146/annurev.ge.28.120194.001123. [DOI] [PubMed] [Google Scholar]
- Moens PB, Heyting C, Dietrich AJJ, van Raamsdonk W, Chen Q. Synaptonemal complex antigen location and conservation. J Cell Biol. 1987;105:93–103. doi: 10.1083/jcb.105.1.93. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Moses MJ. Synaptinemal complex. Annu Rev Genet. 1968;2:363–412. [Google Scholar]
- Nicklas RB. How cells get the right chromosomes. Science. 1997;275:632–637. doi: 10.1126/science.275.5300.632. [DOI] [PubMed] [Google Scholar]
- Proudfoot NJ, Brownlee GG. 3′ non-coding region sequences in eucaryotic messenger RNA. Nature. 1976;263:211–214. doi: 10.1038/263211a0. [DOI] [PubMed] [Google Scholar]
- Rockmill B, Roeder GS. Meiosis in asynaptic yeast. Genetics. 1990;126:563–574. doi: 10.1093/genetics/126.3.563. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Roeder GS. Meiotic chromosomes: It takes two to tango. Genes & Dev. 1997;11:2600–2621. doi: 10.1101/gad.11.20.2600. [DOI] [PubMed] [Google Scholar]
- Rose AM, Baillie DL, Curran J. Meiotic pairing behavior of two free duplications of linkage group I in Caenorhabditis elegans. Mol & Gen Genet. 1984;195:52–56. doi: 10.1007/BF00332723. [DOI] [PubMed] [Google Scholar]
- Sambrook J, Fritsch EF, Maniatis T. In: Molecular cloning: A laboratory manual. Nolan C, editor. 1989. pp. 7.43–45. , 7.49–50, and 18.17–18. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, NY. [Google Scholar]
- Schalk JAC, Dietrich AJJ, Vink ACG, Offenberg HH, van Aalderen M, Heyting C. Localization of SCP2 and SCP3 protein molecules within the synaptonemal complexes of the rat. Chromosoma. 1998;107:540–548. doi: 10.1007/s004120050340. [DOI] [PubMed] [Google Scholar]
- Schedl T. Developmental genetics of the germ line. In: Riddle DL, Blumenthal T, Meyer BJ, Priess JR, editors. C. elegans II. Cold Spring Harbor, NY: Cold Spring Harbor Laboratory Press; 1997. pp. 241–269. [PubMed] [Google Scholar]
- Schwacha A, Kleckner N. Identification of joint molecules that form frequently between homologs but rarely between sister chromatids during yeast meiosis. Cell. 1994;76:51–63. doi: 10.1016/0092-8674(94)90172-4. [DOI] [PubMed] [Google Scholar]
- ————— Interhomolog bias during meiotic recombination: Meiotic functions promote a highly differentiated interhomolog-only pathway. Cell. 1997;90:1123–1135. doi: 10.1016/s0092-8674(00)80378-5. [DOI] [PubMed] [Google Scholar]
- Smith AV, Roeder GS. The yeast Red1 protein localizes to the cores of meiotic chromosomes. J Cell Biol. 1997;136:957–967. doi: 10.1083/jcb.136.5.957. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sulston J, Hodgkin J. Methods. In: Wood WB, editor. The nematode Caenorhabditis elegans. Cold Spring Harbor, NY: Cold Spring Harbor Laboratory; 1988. pp. 587–606. [Google Scholar]
- Villeneuve AM. A cis-acting locus that promotes crossing over between X chromosomes in Caenorhabditis elegans. Genetics. 1994;136:887–902. doi: 10.1093/genetics/136.3.887. [DOI] [PMC free article] [PubMed] [Google Scholar]
- von Wettstein D, Rasmussen SW, Holm PB. The synaptonemal complex in genetic segregation. Annu Rev Genet. 1984;18:331–431. doi: 10.1146/annurev.ge.18.120184.001555. [DOI] [PubMed] [Google Scholar]
- Zetka MC, Rose AM. The meiotic behavior of an inversion in Caenorhabditis elegans. Genetics. 1992;131:321–332. doi: 10.1093/genetics/131.2.321. [DOI] [PMC free article] [PubMed] [Google Scholar]
- ————— Mutant rec-1 eliminates the meiotic pattern of crossing over in Caenorhabditis elegans. Genetics. 1995a;141:1339–1349. doi: 10.1093/genetics/141.4.1339. [DOI] [PMC free article] [PubMed] [Google Scholar]
- ————— The genetics of meiosis in Caenorhabditis elegans. Trends Genet. 1995b;11:27–31. doi: 10.1016/s0168-9525(00)88983-0. [DOI] [PubMed] [Google Scholar]