Abstract
Cullin RING ligases (CRLs), the most prolific class of ubiquitin ligase enzymes, are multimeric complexes that regulate a wide range of cellular processes. CRL activity is regulated by CAND1 (Cullin-associated Nedd8-dissociated protein 1), an inhibitor that promotes the dissociation of substrate receptor components from the CRL. We demonstrate here that COMMD1 (copper metabolism MURR1 domain-containing 1), a factor previously found to promote ubiquitination of various substrates, regulates CRL activation by antagonizing CAND1 binding. We show that COMMD1 interacts with multiple Cullins, that the COMMD1-Cul2 complex cannot bind CAND1, and that, conversely, COMMD1 can actively displace CAND1 from CRLs. These findings highlight a novel mechanism of CRL activation and suggest that CRL regulation may underlie the pleiotropic activities of COMMD1.
Keywords: E3 Ubiquitin Ligase, Protein Degradation, Ubiquitin, Ubiquitin Ligase, Ubiquitination, CAND1, COMMD1, Cul1, Cul2, Cullin
Introduction
Ubiquitin, a highly conserved 76-amino acid polypeptide, can be conjugated to substrate proteins through an enzymatic machinery present in all eukaryotic cells (1). Ubiquitination can lead to a variety of outcomes, but a particularly notable event is the degradation of the target protein by the proteasome. Ubiquitination of the target protein involves a multistep enzymatic process that requires a ubiquitin ligase, also referred to as an E3 enzyme (2). It is estimated that there are ∼500–1,000 ubiquitin ligases (3–5), and among them, a particularly prolific group are the Cullin RING ligases (CRLs)3 (6). The core CRL complex contains a Cullin protein (such as Cul1, 2, 3, 4A, 4B, or 5 in mammals), and a RING box protein (Rbx1 or Rbx2). In addition, a variety of substrate binding subunits specific to each Cullin contributes to a large repertoire of complexes, estimated at ∼300 distinct ligases (7–9). These complexes are designated by the Cullin and the specific substrate subunit that they contain. For example, the well known ligase complex that targets IκB-α, which contains Cul1 and the substrate receptor protein βTrCP, is designated as CRL1-βTrCP.
Structurally, the carboxyl-terminal globular domain of the Cullin protein binds to Rbx1 or Rbx2, whereas the amino-terminal region is a rigid stalk formed by three unique five–helix bundle structures called Cullin repeats 1, 2, and 3 (10). The amino-terminal Cullin repeat 1 is responsible for recruiting the substrate receptor complex (SRC). Displacement of this complex is a major mechanism of inhibition of these ligases and is mediated by CAND1 (Cullin-associated Nedd8-dissociated), a large protein that interacts with all three Cullin repeats and the carboxyl-terminal domain (11–13). Formation of the CRL-CAND1 complex is in turn regulated by the ubiquitin-like protein Nedd8, which upon conjugation to the carboxyl-terminal domain prevents CAND1 binding to the Cullin protein (11, 12). On the other hand, the assembled Cul1-CAND1 complex cannot be effectively neddylated because of steric occlusion of the lysine acceptor site by CAND1. Therefore, the mechanism of dissociation of the Cul1-CAND1 complex remains poorly understood, and the existence of cellular factors that control CRL-CAND1 interactions has been proposed (13).
COMMD (copper metabolism MURR1 domain-containing) proteins are pleiotropic factors present in a wide range of eukaryotic organisms (14) and are defined by the presence of the carboxyl-terminal COMM domain (15, 16). The best studied of these factors is COMMD1, which participates in copper metabolism (17), NF-κB-mediated transcription (18), adaptation to hypoxia (19), and electrolyte transport (20, 21). A large deletion in the canine COMMD1 gene, which abolishes protein expression, leads to pathologic copper accumulation, cirrhosis, and liver failure in Bedlington terriers (17). Although humans with pathologic copper accumulation caused by COMMD1 mutations have not been identified (22, 23), a role for this gene in modulating the phenotype of Wilson's disease has been proposed (24). Moreover, COMMD1 has been found to have copper binding activity in vitro (25) and can modulate the maturation of the copper-containing enzyme SOD1 (26). In addition, COMMD1 has been found to play a role in tumor invasion acting as a regulator of both hypoxia-inducible factor (HIF) and NF-κB (27). Decreased expression of COMMD1 in cancer was found to promote tumor invasion in a variety of settings and was associated with a negative impact on patient survival.
Although the mechanisms underlying these pleiotropic activities of COMMD1 remain unclear, this protein has been shown to be an activating co-factor for a Cul2-containing ligase that ubiquitinates NF-κB/RelA (15, 16). However, a broader paradigm that explains the pleiotropic activities of COMMD proteins has not been defined. Here, we report that these factors bind to multiple CRLs and that the prototype member, COMMD1, can activate ligase function through displacement of the CRL inhibitor CAND1.
EXPERIMENTAL PROCEDURES
Plasmids
Expression vectors for Cullin proteins, CAND1, and Skp1 (all of human origin) were generated by PCR amplification of the corresponding coding sequence, which was then introduced into the pEBB-FLAG and pEBG vectors. For Cul1, Pallino-FLAG-Cul1 was used as template (kindly provided by Dr. Michele Pagano). The following IMAGE cDNA clones were obtained from Open Biosystems and used as templates for Cul3, Cul4A, Cul4B, Cul5, Cul7, CAND1, and Skp1: 5784147, 3537176, 5269392, 30331132, 5580027, 5265409, and 6672613, respectively. COMMD-EYFP constructs were generated by subcloning EYFP into pEBB-FLAG expression vectors previously described (15). The plasmid pEBB-COMMD1 (M110A/H134A)-GST was generated by site-directed mutagenesis. Deletion constructs for Cul2 were generated by PCR using pEBB-FLAG-Cul2 as template (16), with the amino acid boundaries of the encoded mutant proteins being: Cul2 1–415, 109–415, 109–745, 415–745, 1–150 (R1), 151–270 (R2), 271–386 (R3), and 151–386 (R2-R3). His6-tagged versions of COMMD1 and Skp1 were subcloned from the corresponding pEBB vectors into pET30a using the BamHI and NotI sites. All other plasmids have been previously described (15, 16, 28, 29).
Cell Culture and Transfection
HEK 293 cells and HeLa cells were obtained from ATCC. U2OS cells with stable shRNA-mediated repression of COMMD1 have been previously reported (16). HEK 293 cells stably expressing GST or GST-Cul2 were generated by lentiviral infection and selection as previously described (16). All of the cell lines were cultured in DMEM supplemented with 10% FBS and l-glutamine (2 mm). Biotin (Sigma; 4 μm) was added to the medium when fusion proteins with the biotinylation target peptide were expressed. A standard calcium phosphate transfection protocol was used to transfect HEK 293 cells (29). HeLa cells were transfected using FuGENE (Roche Applied Science) according to the manufacturer's instructions.
Confocal and Fluorescence Microscopy
HeLa cells were plated in chambered coverglass plates and transfected with the indicated COMMD-EYFP plasmids (0.5 μg/well). The cells were stained with Hoechst 33342 (8 μm) for 30 min, and images were obtained with a Zeiss LSM 510 META confocal microscope equipped with a Chameleon XR NIR laser.
Immunoblotting and Protein Precipitation
Cell lysate preparation, immunoprecipitations, GSH precipitations, and immunoblotting were performed as previously described (15, 28). For the COMMD1-Cul1 interaction, the cells were sonicated in detergent-free buffer E (25 mm HEPES, 100 mm NaCl, 1 mm EDTA, 5% glycerol). Interactions with Cul2 and Cul3 were studied using buffer F (PBS, 0.8% Nonidet P-40). Other lysis buffer conditions are indicated in the figure legends where appropriate. The precipitation of RelA after denatured lysis of nuclei, which was used to detect ubiquitinated RelA (see Fig. 6), has been previously described (30). The following antibodies were utilized in our studies: CAND1 (Novus, H00055832-MO1; Santa Cruz, sc-10672), COMMD1 (15), Cul1 (Santa Cruz, sc-12761), Cul2 (Zymed Laboratories Inc., 51-1800), Cul3 (kindly provided by Dr. Matthias Peter) (31), Elongin C (BioLegend, 613101), FLAG (Sigma, A8592 and F1804), GST (Santa Cruz, sc-459), HA (Sigma, H6533; Covance, MMS101R), RelA (Santa Cruz, sc-372), and Rbx1 (LabVision, 127-075-160).
FIGURE 6.
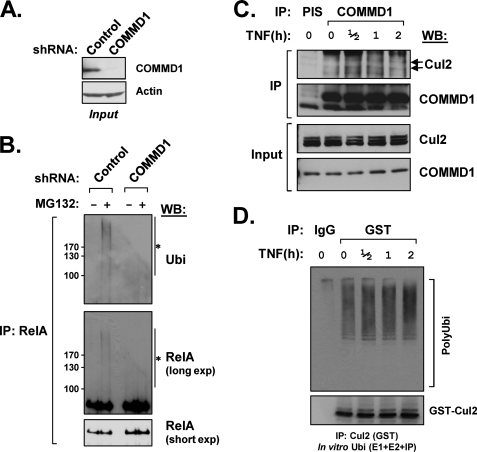
COMMD1 is required for the ubiquitination of specific targets in vivo. A, Western blot demonstrating stable shRNA-mediated repression of COMMD1 in U2OS cells. B, RelA ubiquitination is greatly impaired in COMMD1-deficient cells. U2OS cells were treated with the proteasome inhibitor MG-132 as indicated, and cell nuclei were isolated and then lysed in a denaturing buffer. RelA was immunoprecipitated from nuclear extracts and immunoblotted for ubiquitin (top panel) or for RelA itself (middle and bottom panels). The asterisk indicates the high molecular weight smear consistent with ubiquitinated RelA. C, TNF stimulates COMMD1-Cul2 binding. HEK 293 cells were treated with TNF (1,000 units/ml) for the indicated time intervals, and endogenous COMMD1 was subsequently precipitated from cell lysates as in Fig. 1A. The recovery of endogenous Cul2 was determined by immunoblotting. D, cells stably expressing GST-Cul2 (see supplemental Fig. S3 for details) were treated with TNF (1,000 units/ml) for the indicated time points. Thereafter, GST-Cul2 was immunoprecipitated with a GST antibody. The recovered material was utilized as a source for E3 activity as previously. WB, Western blot; IP, immunoprecipitation; PIS, preimmune serum; Ubi, ubiquitination.
Recombinant Protein Preparation
HA-tagged Skp1 and COMMD1 proteins were expressed in HEK 293 cells and purified from cell lysates. After incubation with an anti-HA affinity matrix (Roche Applied Science), the purified material was extensively washed four times with RIPA lysis buffer and then eluted in dissociation buffer (40 mm HEPES, pH 7.9, 60 mm potassium acetate, 10% glycerol) containing HA peptide as a competitor (1 mg/ml). Elution was performed three times at 37 °C with agitation. The combined eluate was used for further experiments. His6-tagged COMMD1 or Skp1 were expressed and purified from Escherichia coli (BL21 strain, Stratagene) using the nickel-nitrilotriacetic acid purification system (Invitrogen). The final imidazole eluate was loaded onto an Amicon centrifuge filter (Millipore) to eliminate imidazole, and the purified protein was redissolved in PBS buffer with 10% glycerol through sequential filtration and buffer exchange.
Bimolecular Affinity Purification of the Cul2-COMMD1 Complex
HEK 293 cells were transfected with GST or GST-tagged Cul2 and COMMD1 fused to TB (a biotinylation target peptide), grown in biotin-supplemented media, and lysed in Triton buffer 2 days later. The Cul2-COMMD1 complex was purified through sequential affinity purification using glutathione and streptavidin columns as previously described (29).
In Vitro Ubiquitination Assays
Each reaction mixture consisted of recombinant HA-human ubiquitin (2.5 μg), human His6-E1 (Uba1, 50 ng), various human E2 enzymes as indicated (UbcH5a, 5b, 5c, 7, and 10, 100 ng; UbcH3, 150 ng), and an ATP regenerating buffer (all obtained from Boston Biochem). These were mixed in reaction buffer (40 mm HEPES, pH 7.9, 60 mm potassium acetate, 2 mm DTT, 5 mm MgCl2, 10% glycerol) and agitated at 30 °C for 60 min (16). Polyubiquitin chain formation was detected by Western blotting with an anti-HA antibody as described above.
In Vitro Displacement of CAND1
The CAND1 dissociation reaction was based on the prior report by Zheng et al. (12). For displacement of endogenous CAND1 (see Fig. 4B), HEK 293 cells were transfected with GST-Cul1 or GST-Cul2 and lysed in a hypotonic buffer (20 mm HEPES, pH 7.2, 5 mm KCl, 1.5 mm MgCl2, 0.5 mm DTT) using a Dounce apparatus. The lysate was cleared by centrifugation at 27,000 × g (4 °C for 30 min). In the case of displacement of overexpressed FLAG-CAND1 (see Fig. 4, C and D), the cells were similarly transfected but lysed in a Triton X-100-containing buffer (25 mm HEPES, pH 7.9, 100 mm NaCl, 10% glycerol, 1% Triton X-100). All of the buffers were supplemented with protease inhibitors (Roche Applied Science) and DTT (10 mm). Cul1 or Cul2 complexes were purified by GSH-Sepharose (GE Healthcare). The GSH beads were aliquoted and incubated at 30 °C for 30 min with ATP in dissociation buffer (40 mm HEPES, pH 7.9, 60 mm potassium acetate, 10% glycerol), with the indicated recombinant proteins (Skp1 or COMMD1). Thereafter, the beads were washed twice with Triton X-100 lysis buffer (25 mm HEPES, 100 mm NaCl, 1 mm EDTA, 10% glycerol, 1% Triton X-100). The leftover material was utilized for Western blot analysis.
FIGURE 4.
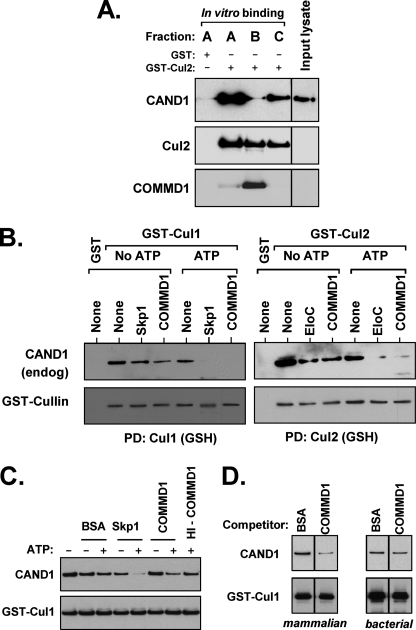
COMMD1 promotes the dissociation of CAND1 from CRL complexes. A, COMMD1 prevents CAND1 binding to Cul2. Fractions A, B, and C were prepared as depicted in Fig. 3A, mixed for 2 h with a fresh HEK 293 cell lysate expressing FLAG-CAND1, washed, and immunoblotted for CAND1, Cul2, or COMMD1. B, COMMD1 can displace CAND1 from Cul1 or Cul2 complexes. GST-Cul1 or GST-Cul2 were expressed in HEK 293 cells and purified through a GSH affinity matrix. This material was then mixed with HA-Skp1 or HA-COMMD1 purified from mammalian cells. After incubation at 30 °C with ATP (15 mm), the GSH beads were washed, and the presence of endogenous CAND1 still bound to Cul1 or Cul2 was determined by immunoblotting. PD, pulldown. C, the activity of COMMD1 is heat-labile. GST-Cul1 complexes were purified as before from cells also co-expressing FLAG-CAND1. After a displacement reaction (performed as previously but using 7.5 mm ATP), the presence of remaining FLAG-CAND1 bound to Cul1 was determined by immunoblotting. HI-COMMD1, heat inactivated COMMD1 (95 °C for 10 min). D, recombinant COMMD1 made in E. coli is devoid of activity. Shown is the same reaction as in C, but utilizing mammalian HA-COMMD1 or bacterially made His6-COMMD1. Only the ATP containing reactions are shown; each subpanel was run in the same gel.
RESULTS
COMMD1 Associates with CRL Complexes
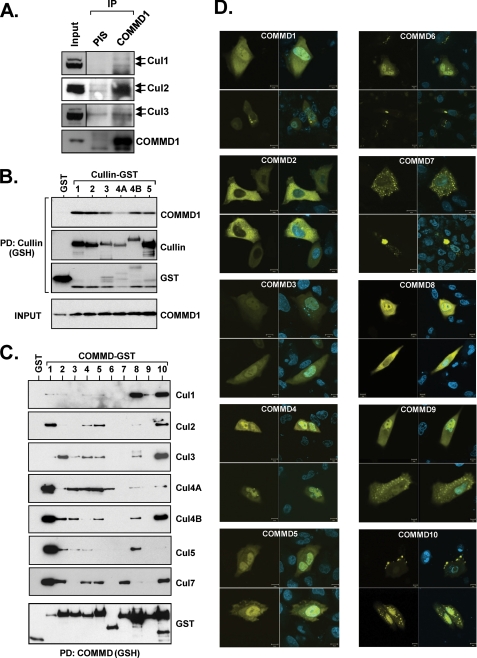
Given the diverse activities of COMMD1 (32), we initially explored the possibility that this factor might play a broader role in the regulation of ubiquitination of various targets by CRLs. To this end, we examined in more detail the interaction between COMMD1 and other Cullin proteins. We found that endogenous COMMD1 could be co-precipitated with endogenous Cul1, Cul2, or Cul3 (Fig. 1A). The Cul1-COMMD1 interaction was best visualized in a detergent-free buffer, whereas the Cul2 and Cul3 interactions were best seen with a glycerol-free buffer containing low amounts of Nonidet P-40 (see “Experimental Procedures”). These interactions were also evident using a buffer containing 1% Triton X-100 (supplemental Fig. S1). Utilizing expression vectors for all five canonical Cullins (Cul1, 2, 3, 4A, 4B, and 5), we could easily recapitulate the interaction between COMMD1 and all of the Cullin family members tested (Fig. 1B).
FIGURE 1.
COMMD1 and other COMMD proteins associate with CRL complexes. A, endogenous co-immunoprecipitation (IP) of COMMD1 and CRLs. COMMD1 was immunoprecipitated from HEK 293 cell lysates. The precipitated material was probed for the indicated Cullins. PIS, preimmune serum. B, COMMD1 binds to other Cullin family members. COMMD1 was expressed in HEK 293 cells together with Cullin proteins fused to GST, which were precipitated as above and immunoblotted for COMMD1. PD, pulldown. C, COMMD proteins interact with Cullins. In each experiment, one FLAG-tagged Cullin was expressed together with COMMD proteins fused to GST, which were subsequently precipitated. The presence of the Cullin in the precipitates was determined by immunoblotting with the FLAG antibody. D, cellular distribution of COMMD proteins. The indicated COMMD proteins fused to EYFP were expressed in HeLa cells. The cells were counterstained with Hoechst and imaged in a confocal microscope. The scale bar corresponds to 10 μm.
Other COMMD Proteins Also Interact with CRLs
Other COMMD proteins have largely undefined functions at this point. Using a similar approach, it was found that other COMMDs may also interact with CRLs (Fig. 1C). Interestingly, their patterns of Cullin preference were not identical.
Several COMMDs are known to display differential tissue expression (15), yet several of them are ubiquitous and form heterodimers. To try to understand the potential redundancy in this system, we examined the cellular distribution of COMMD proteins expressed with a fused EYFP tag (Fig. 1D). Although some similarities were found for several family members, certain COMMD proteins displayed specific and unique cellular distribution. For example, in the case of COMMD2, the protein was consistently excluded from the nucleus, whereas COMMD7 demonstrated a unique punctuate pattern.
COMMD1 Associates with Active CRLs in a Copper-independent Manner
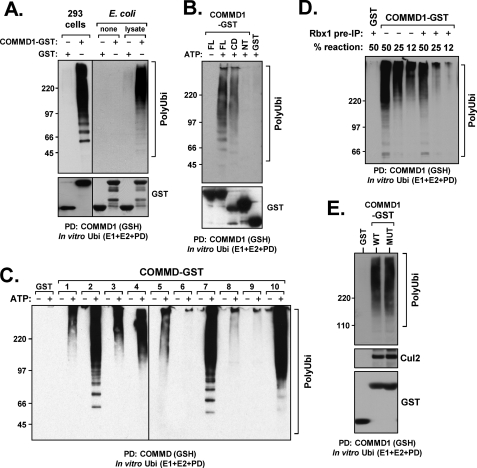
Consistent with the interaction between COMMD1 and CRLs, we found that COMMD1 could precipitate ubiquitin ligase activity when extracted from mammalian cells. COMMD1 was expressed and affinity-purified from HEK 293 cells or E. coli, and the recovered proteins were offered to an in vitro reaction containing the E1 and E2 (UbcH5a) enzymes, recombinant ubiquitin, and an ATP regenerating buffer. Polyubiquitin chain formation in the presence of ATP was readily detectable when COMMD1 precipitated from mammalian cells was provided as a source of ubiquitin ligase activity (Fig. 2A, left panels). However, if COMMD1 was precipitated and washed in RIPA buffer, this activity was lost, consistent with the fact that COMMD1-CRL interactions are abolished in this buffer (data not shown). On the other hand, recombinant COMMD1 prepared in E. coli was devoid of ligase activity, but when the protein was incubated with a mammalian lysate and then washed, ligase activity was reconstituted (Fig. 2A, right panels). These results suggested that COMMD1 interacts with cellular factors that provide ligase activity.
FIGURE 2.
COMMD1 associates with active CRL complexes in a copper-independent manner. A, COMMD1 precipitates cellular factors with ubiquitin ligase activity. In vitro ubiquitination reactions were supplemented with precipitated proteins as indicated. Polyubiquitin chain formation was determined by Western blotting as an indication of ubiquitin ligase activity. COMMD1-GST or GST were expressed in HEK 293 cells (left panels) or prepared recombinantly in E. coli (right panels). Recombinant proteins either were offered directly (none) or were first mixed with a mammalian lysate and washed prior to the reaction (lysate). PD, pulldown. B, COMMD1 precipitates ubiquitin ligase activity through the COMM domain. COMMD1 full-length (FL), its amino terminus (NT), or the COMM domain (CD) fused to GST were precipitated from transfected HEK 293 cells and added to an in vitro ubiquitination reaction as in A. C, COMMD proteins precipitate ubiquitin ligase activity. COMMD proteins fused to GST were precipitated and added to an in vitro ubiquitination reaction as in A. D, depletion of Rbx1 reduces COMMD1-associated ligase activity. COMMD1-GST was utilized as a source of ubiquitin ligase activity as before. The protein was expressed in HEK 293 cells and either precipitated directly (−) or following immunodepletion of FLAG-Rbx1 (+). IP, immunoprecipitation. E, copper binding by COMMD1 is not involved in E3 and Cul2 binding. WT COMMD1 or an M110A/H134A mutant (MUT) unable to bind copper were expressed in HEK 293 cells. After precipitation, the protein complexes were examined for associated E3 activity as before (upper panel) and for the presence of co-precipitated endogenous Cul2 (middle panel). Ubi, ubiquitination.
Next, we observed that the COMM domain of COMMD1, which is responsible for binding to Cul2 (16), was also necessary and sufficient for recovering ligase activity (Fig. 2B, COMM Domain or CD lane). Given the conservation of the COMM domain across all COMMD proteins and our finding of COMMD-CRL interactions, we speculated that other family members may be similarly linked to active ubiquitin ligases. Indeed, when precipitated from mammalian cells, several COMMD proteins provided a source of ubiquitin ligase activity in vitro (Fig. 2C). The ligase activity varied as a function of the E2 offered to each COMMD protein (supplemental Fig. S2), consistent with their differential binding to Cullin family members.
Next, we examined whether CRL interactions were responsible for COMMD1-associated ligase activity. To test this possibility, COMMD1 and Rbx1 (the main RING finger subunit in CRLs) were co-expressed in HEK 293 cells, and Rbx1 was immunoprecipitated prior to assessing the E3 activity associated to COMMD1. Depletion of Rbx1 led to a significant drop in COMMD1-associated ubiquitin ligase activity (Fig. 2D), consistent with the notion that ligase activity associated with COMMD1 is due to its interaction with CRLs.
Finally, a copper binding activity for COMMD1 has been found in vitro, and residues that coordinate copper have been similarly mapped (25). We examined whether copper could modulate COMMD1-CRL interactions. A double mutation in COMMD1 (M110A/H134A) targeting copper-binding residues did not affect Cul2 binding or COMMD1-associated E3 activity (Fig. 2E). Similarly, the addition of copper to the growth medium did not affect these interactions (data not shown), indicating that CRL binding is copper-independent.
The Cul2-COMMD1 Complex Excludes the CRL Inhibitor CAND1
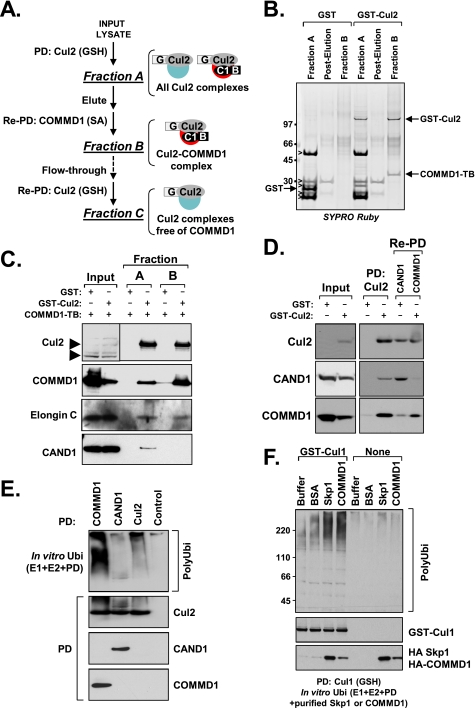
Altogether, the data indicated that COMMD1 associates with CRLs, a property that seems to be shared by other COMMD proteins. Nevertheless, the specific function for such interactions was not immediately apparent. To address the latter question, we utilized a bi-molecular tandem affinity purification scheme (29) to isolate a purified Cul2-COMMD1 complex from mammalian cells (Fig. 3A). Cul2 was purified using a glutathione-Sepharose column (fraction A); after elution, this material was offered to a streptavidin-agarose column for purification of the COMMD1 subfraction (fraction B). As expected, COMMD1 and Cul2 were readily demonstrated in fraction B (Fig. 3B), and based on the amounts recovered, this complex was estimated to represent 2–5% of all cellular Cul2. In addition, fractions A and B provided ubiquitin ligase activity in vitro (not shown here), and known components of the active ligase, such as Elongin C, were also present in both fractions (Fig. 3C). Interestingly, the CRL inhibitor CAND1 was readily identified in fraction A but was absent in Cul2-COMMD1 complexes (Fig. 3C).
FIGURE 3.
The Cul2-COMMD1 complex excludes the CRL inhibitor CAND1. A, purification strategy used to isolate the Cul2-COMMD1 complex. Cul2 fused to GST (G) and COMMD1 fused to a biotinylation tag (B) were expressed in HEK 293 cells. Cul2 was purified through a GSH-Sepharose column (fraction A) and subsequently eluted. COMMD1 was precipitated from the eluted material using streptavidin (SA)-agarose beads (fraction B). Cul2 remaining in the flow-through after the streptavidin-agarose column was reprecipitated with a GSH column (Re-PD: Cul2 (GSH)) resulting in fraction C. PD, pulldown. B, Cul2-COMMD1 complex purification. Proteins from fractions A and B were separated by SDS-PAGE and stained with SYPRO Ruby; the identity of major bands was determined by paired Western blot analysis (nonspecific bands indicated by open arrowheads). C, the Cul2-COMMD1 complex excludes CAND1. Fractions A and B were immunoblotted for Cul2, COMMD1, Elongin C, and CAND1. Endogenous Cul2 and GST-Cul2 are noted by arrowheads to the left of the top panel. D, COMMD1 and CAND1 exist in separate Cul2 cellular pools. GST-Cul2, COMMD1 fused to a biotinylation tag, and FLAG-CAND1 were co-expressed in HEK 293 cells. Cul2 was purified through a GSH affinity column (PD: Cul2) and subsequently eluted. COMMD1 or CAND1 were then precipitated from the eluate using SA agarose beads or FLAG antibody, respectively. The resulting fractions were immunoblotted for Cul2, CAND1, or COMMD1. E, Cul2 complexes containing COMMD1 are active. GST-Cul2 was precipitated from transfected HEK 293 cells utilizing GSH beads (Cul2) or using protein G beads as a control (Control). FLAG-COMMD1 or FLAG-CAND1 co-expressed with GST-Cul2 were precipitated utilizing the FLAG antibody. After adjusting for equal Cul2 recovery, the resulting material was added to an in vitro ubiquitination reaction as in Fig. 2A and immunoblotted for ubiquitin. F, purified COMMD1 potentiates Cul1 E3 activity in vitro. GST-tagged Cul1 was expressed in HEK 293 cells and purified using GSH affinity matrix. Purified HA-Skp1 or HA-COMMD1 prepared from mammalian lysates was then added to the recovered Cul1 complex, and this preparation was used as a source for E3 activity as before.
Next, we sought to ascertain whether the lack of CAND1 in the Cul2-COMMD1 complex was truly due to the purification of a distinct cellular fraction of Cul2 rather than an artifact of the purification scheme itself. To address this question, Cul2 was first purified, eluted, and subsequently offered to a second column for precipitation of either COMMD1 or CAND1. Although COMMD1 and CAND1 were readily co-precipitated with Cul2 in the initial fraction, the second purification step of either COMMD1 or CAND1 resulted in the depletion of the other protein (Fig. 3D), consistent with the existence of mutually exclusive pools of COMMD1 and CAND1 when bound with Cul2 in cells.
COMMD1 Promotes CRL E3 Activity
Given the exclusion of CAND1 from Cul2-COMMD1 complexes, we reasoned that COMMD1 might bind to or promote a more active CRL state. To test this notion, Cul2 was co-expressed with COMMD1 and subsequently precipitated from cell lysates directly or through COMMD1. The CRL inhibitor CAND1 was included as a negative control. At the end of the precipitation and after adjusting for equivalent Cul2 recovery, we utilized the isolated complexes as a source for E3 ligase activity in an in vitro ligase reaction. These experiments demonstrated that Cul2 associated with COMMD1 was the most active isolate, whereas the CAND1 fraction was relatively devoid of enzymatic activity (Fig. 3E).
The preferential binding of COMMD1 to a CAND1-free pool of Cul2 could be responsible for these findings; alternatively, COMMD1 could promote an active conformation for the complex. Therefore, we explored further whether COMMD1 could potentiate CRL E3 activity in vitro. GST-Cul1 that was immunoprecipitated from mammalian cells and its E3 activity was examined in vitro as before. In these assays, its substrate receptor adaptor Skp1 or COMMD1 were added to the in vitro reaction. As can be seen in Fig. 3F, the addition of COMMD1 or Skp1 led to enhanced Cul1 E3 activity. This was not due to additive E3 activity precipitated by COMMD1 or Skp1, because there was no associated E3 activity bound to COMMD1 or Skp1 when these preparations were made with RIPA buffer (Fig. 3F, right half of the gel). Altogether, this indicated that COMMD1 binds to active CRL complexes and can lead to their activation in vitro. However, cellular deficiency of COMMD1 did not substantially affect endogenous CRL2 activity (supplemental Fig. S3), potentially because of redundant effects by other COMMDs expressed in these cells.
COMMD1 Promotes the Dissociation of CAND1 from CRL Complexes
Akin to the distinct fractionation between CAND1 and substrate receptor complexes (11–13), our data indicated that CAND1 and COMMD1 do not coexist in the same CRL complexes. To test this notion further, we examined whether the purified Cul2-COMMD1 complex was capable of binding CAND1 in vitro. To that end, after purification of fraction B, unbound complexes were offered again to a glutathione-Sepharose column to isolate Cul2 devoid of COMMD1 (fraction C; Fig. 3C). All of these preparations were mixed with a fresh lysate containing expressed CAND1, and binding in vitro was assessed by co-precipitation. This experiment indicated that total Cul2 complexes or Cul2 complexes devoid of COMMD1 (fractions A and C) bound CAND1 well, but the Cul2-COMMD1 complex (fraction B) could not bind CAND1 efficiently (Fig. 4A). This was the case despite similar loading of Cul2 among these fractions and the additional purification steps required to isolate fraction C, strongly suggesting that these results were independent of the purification scheme itself.
Next we examined whether COMMD1 could facilitate the dissociation of CAND1 from CRLs. To test this notion, GST-tagged Cul1 or Cul2 was precipitated from transfected HEK 293 cells resulting in the co-precipitation of endogenous CAND1. Next, the precipitated Cul1 or Cul2 complexes were mixed with purified COMMD1 or their respective substrate receptor adaptors, Skp1 or Elongin C, respectively. After incubation at 30 °C and thorough washing, the presence of CAND1 still associated to either Cul1 or Cul2 was assessed by immunoblotting. As has been reported before, substrate receptor adaptors were able to promote the dissociation of CAND1 (12, 33), particularly when the reaction was supplemented with ATP (Fig. 4B). Importantly, the same was true of COMMD1, which promoted CAND1 dissociation from either Cul1 or Cul2 (Fig. 4B). Interestingly, this displacement reaction was abrogated when COMMD1 had been heat-inactivated (Fig. 4C). In addition, recombinant COMMD1 made in E. coli was devoid of activity compared with a mammalian preparation (Fig. 4D). This is in contrast with Skp1, which, as previously described (12), was active irrespective of it being prepared from E. coli lysates (data not shown).
COMMD1 Binds to CRLs in a Manner Distinct from Substrate Receptors
Structural information indicates that Skp1 and CAND1 bind to an overlapping region of Cullin repeat 1 (R1) in Cul1 (13). CAND1 also binds to the carboxyl-terminal domain of Cul1 over its neddylation site and makes extensive contacts with R2 and R3 (13, 34) (Fig. 5F). This information explains why CAND1 does not bind to neddylated Cullins and why CAND1 and SRC binding are mutually exclusive.
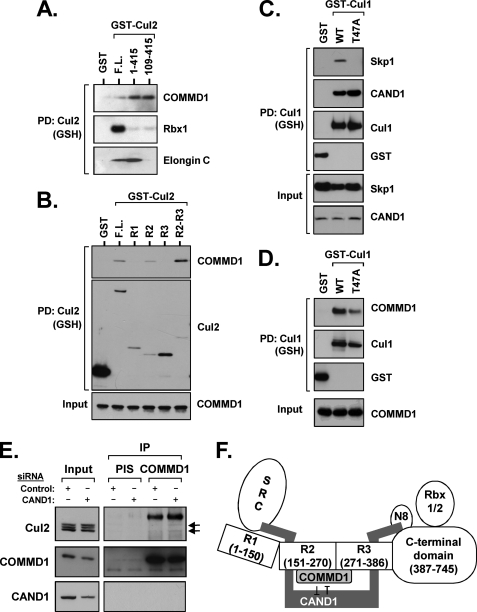
FIGURE 5.
COMMD1 binds to CRLs in a manner distinct from substrate receptors. A, full-length and specific deletion mutants of Cul2 fused to GST were expressed in HEK 293 cells and precipitated from cell lysates with glutathione-Sepharose (GSH) beads. The recovered material was immunoblotted for endogenous COMMD1, Rbx1, and Elongin C. PD, pulldown. B, COMMD1 binds to Cullin repeat 2 in Cul2. COMMD1 was expressed in HEK 293 cells together with Cul2 deletion mutants fused to GST (R1, R2, or R3, Cullin repeats 1, 2, or 3, respectively), which were precipitated as in A and immunoblotted for COMMD1. C and D, Cul1 can bind to COMMD1 independently of Skp1. Cul1 WT or a T47A point mutant fused to GST were expressed in HEK 293 cells. Their interactions with HA-Skp1 or FLAG-CAND1 were tested by co-precipitation (C). The same Cul1 proteins were expressed together with HA-COMMD1 and Cul1-COMMD1 interactions were examined by co-precipitation (D). E, CAND1 deficiency promotes COMMD1-Cul2 binding. HEK 293 cells were transfected with siRNA duplexes targeting CAND1. Endogenous COMMD1 was immunoprecipitated (IP), and its interaction with endogenous Cul2 was evaluated as in Fig. 1A. F, model depicting the various domains in canonical Cullin proteins, their known interacting partners, and the competitive binding by COMMD1 and CAND1 for R2. The amino acid boundaries for R1, R2, and R3 in Cul2 are also depicted. N8, NEDD8.
Next, we tried to ascertain whether competitive binding for a shared domain on Cul2 may be involved in the ability of COMMD1 to displace CAND1. First, it was noted that neither the amino terminus nor the carboxyl terminus of Cul2 were required for COMMD1 binding (Fig. 5A). Instead, amino acids 109–415 were found to be sufficient to mediate the COMMD1 interaction. Utilizing an alignment between Cul2 and Cul1 and the published crystal structure of Cul1 as a framework (10), we predicted the boundaries of Cul2 R1, R2, and R3 (Fig. 5F). COMMD1 was found to bind preferentially to R2 and not to R1 (Fig. 5B).
Based on this finding, we predicted that COMMD1-CRL binding occurs independently of the SRC. To test this notion, we introduced a previously reported point mutation in R1 of Cul1 (10), which disrupted Skp1 binding (Fig. 5D). Interestingly, the T47A mutant lost the ability to interact with Skp1 but bound to CAND1 (Fig. 5C), consistent with the incomplete overlap between CAND1 and Skp1 binding surfaces (10, 13). Importantly, this Cul1 mutant retained its ability to interact with COMMD1 (Fig. 5D), indicating that this interaction is independent of the SRC.
Consistent with the competitive nature of the COMMD1-CAND1 interactions with CRLs, we observed that siRNA-mediated reduction of CAND1 expression increased the Cul2-COMMD1 endogenous binding (Fig. 5E). On the other hand, depletion of COMMD1 by siRNA had little effect on the CAND1-Cul1 interaction (data not shown), which may be explained by the potential redundancy of the nine other COMMD family members.
COMMD1 Is Required for the Ubiquitination of Specific Targets in Vivo
We next set out to address the role of COMMD1 on the ubiquitination of endogenous proteins in vivo. We examined this question in the context of the NF-κB signaling pathway. As reported previously, we found that COMMD1 is required for NF-κB/RelA ubiquitination (16, 35, 36). Cells with decreased levels of COMMD1 (Fig. 6A) demonstrated a profound decrease in the ubiquitination of endogenous RelA (Fig. 6B). In addition, the pro-inflammatory cytokine TNF stimulated the binding of COMMD1 to Cul2 (Fig. 6C), consistent with its role in promoting NF-κB/RelA ubiquitination and degradation as a mechanism to terminate signaling (16, 35, 36). Interestingly, TNF stimulation had only modest effects on total CRL2 activity (Fig. 6D), suggesting that the TNF induction of COMMD1/Cul2 binding more likely activates specific CRL2 complexes, such as those targeting RelA and possibly other targets.
DISCUSSION
Altogether, our studies indicate that COMMD1 can modulate CRL activity through its ability to displace CAND1. This finding provides a mechanism for the ability of COMMD1 to promote protein ubiquitination and degradation (16, 36, 37). The data also suggest that COMMD1 activates specific CRL complexes involved in the turnover of specific products, such as RelA. This activation seems to be regulated by the inducible interaction between COMMD1 and certain CRL complexes under specific conditions, as in the case of induced COMMD1-Cul2 interactions after TNF treatment.
In the present model of CRL regulation, CAND1 exerts its inhibitory function on all Cullins, potentially regulating ∼300 different ligase complexes by preventing the full assembly of the active ligase (9). This broad activity would have a myriad of effects on cellular physiology, and therefore factors that control CAND1-CRL regulation in a more restricted fashion would make physiologic sense. Indeed, others have proposed the existence of such regulation based on the unexplained variable affinity of CAND1 for different Cullins in different tissues (8). Based on the data presented, our view is that the COMMD protein family, with its variable pattern of tissue and cellular expression and the unique Cullin binding preferences of each COMMD family member, could provide fine tuning of CRL activation by countering CAND1 binding.
Given the myriad of activities regulated by the ∼300 CRL complexes existent in mammals, it is anticipated that COMMD proteins would likely affect many physiological processes. Indeed, consistent with this notion, COMMD1 has been implicated in a variety of seemingly unrelated processes such as NF-κB regulation, sodium and electrolyte transport, hypoxia responses, and copper excretion. In most instances, the mechanism has been linked to protein ubiquitination, in agreement with the notion that COMMD1 and other COMMD proteins regulate CRL ubiquitin ligase complexes. Our view is that the biochemical events dissected in this study provide a mechanistic explanation for the various biological activities identified for COMMD1.
Similar to what has been reported for substrate receptor complexes (11, 12, 33), COMMD1 binding to CRLs displaces CAND1, leading to an enzymatically active complex. Based on the structure of Cul1, we speculate that this is mediated by competitive binding for the highly conserved R2 region (Fig. 5F), which interacts extensively with CAND1 (13). The competitive nature of COMMD1/CAND1 binding to CRLs is extensively demonstrated using various in vitro systems, and the relevance of these findings is validated by the increased binding of COMMD1 to Cul2 in CAND1-deficient cells.
It is important to note that the biochemical events that lead to CRL-CAND1 dissociation remain quite enigmatic. In the case of both COMMD1 and the SRC components Skp1 or Elongin C, the reaction requires ATP supplementation, yet the basis for this ATP requirement is not evident, because none of these factors have ATPase or kinase activity. In addition, a CRL1-CAND1 complex formed by E. coli-expressed recombinant proteins cannot be dissociated by Skp1 or neddylation co-factors (13), and this reaction only takes place when the CRL-CAND1 complex is isolated from mammalian cells (12, 33). For all of these reasons, we speculate that additional factors present in the mammalian preparations are required for CAND1 dissociation, and these factors may explain the requirement for ATP in the reaction. Similarly, we found that COMMD1 was active in this reaction only when extracted from mammalian cells, suggesting either that post-translational modifications of COMMD1 are necessary or that co-factor(s) associated with COMMD1 in mammalian cells are required for this process.
Recent data indicate that the recycling of CRLs between the CAND1-bound and SRC-bound states facilitates the loading of rare SRCs that would otherwise be outcompeted by more abundant co-factors (38). Although the SRC can dissociate the CRL1-CAND1 complex in vitro, various in vivo experiments indicate that other factors regulate this interaction. Disrupting Cul1-SRC complex formation in vivo has limited effects on CRL1-CAND1 interactions (39). Similarly, inhibiting CRL neddylation in cells did not promote CRL1-CAND1 interactions, nor did it destabilize CRL1-SRC complexes (40). These observations support the notion that other factors regulate CRL-CAND1 interactions in vivo, which is in keeping with the findings of this study.
At the present time, it remains unclear whether COMMD1 acts to synergize the SRC recycling or whether it constitutes a parallel activation pathway. Until the precise mechanisms of CAND1 displacement are further elucidated, addressing these questions will remain problematic with our current systems. Nevertheless, the fact that COMMD1 binds to Cullins in a manner that is distinct from that used by SRCs suggests that these two mechanisms of CAND1 displacement may be related but are still independent from each other.
Supplementary Material
Acknowledgments
We are grateful to Michele Pagano and Peter Kaiser for providing plasmids that were useful for these studies. We also express our thanks to Matthias Peter for kindly providing the Cul3 antibody utilized in our studies.
This work was supported, in whole or in part, by National Institutes of Health Grant R01 DK073639. This work was also supported by Crohn's and Colitis Foundation of America Grant SRA 2737 and a University of Texas Southwestern Medical Center DOCS Award (to E. B.).

The on-line version of this article (available at http://www.jbc.org) contains supplemental Figs. S1–S3.
- CRL
- Cullin RING ligase
- SRC
- substrate receptor complex
- RIPA
- radioimmune precipitation assay.
REFERENCES
- 1. Weissman A. M. (2001) Nat. Rev. Mol. Cell Biol. 2, 169–178 [DOI] [PubMed] [Google Scholar]
- 2. Wilkinson C. R. (2002) Trends Cell Biol. 12, 545–546 [DOI] [PubMed] [Google Scholar]
- 3. Joazeiro C. A., Weissman A. M. (2000) Cell 102, 549–552 [DOI] [PubMed] [Google Scholar]
- 4. Patterson C. (2002) Sci. STKE 2002, pe4. [DOI] [PubMed] [Google Scholar]
- 5. Coscoy L., Ganem D. (2003) Trends Cell Biol. 13, 7–12 [DOI] [PubMed] [Google Scholar]
- 6. Petroski M. D., Deshaies R. J. (2005) Nat. Rev. Mol. Cell. Biol. 6, 9–20 [DOI] [PubMed] [Google Scholar]
- 7. Willems A. R., Schwab M., Tyers M. (2004) Biochim. Biophys. Acta 1695, 133–170 [DOI] [PubMed] [Google Scholar]
- 8. Bosu D. R., Kipreos E. T. (2008) Cell Div. 3, 7 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9. Duda D. M., Scott D. C., Calabrese M. F., Zimmerman E. S., Zheng N., Schulman B. A. (2011) Curr. Opin. Struct Biol. 21, 257–264 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10. Zheng N., Schulman B. A., Song L., Miller J. J., Jeffrey P. D., Wang P., Chu C., Koepp D. M., Elledge S. J., Pagano M., Conaway R. C., Conaway J. W., Harper J. W., Pavletich N. P. (2002) Nature 416, 703–709 [DOI] [PubMed] [Google Scholar]
- 11. Liu J., Furukawa M., Matsumoto T., Xiong Y. (2002) Mol. Cell 10, 1511–1518 [DOI] [PubMed] [Google Scholar]
- 12. Zheng J., Yang X., Harrell J. M., Ryzhikov S., Shim E. H., Lykke-Andersen K., Wei N., Sun H., Kobayashi R., Zhang H. (2002) Mol. Cell 10, 1519–1526 [DOI] [PubMed] [Google Scholar]
- 13. Goldenberg S. J., Cascio T. C., Shumway S. D., Garbutt K. C., Liu J., Xiong Y., Zheng N. (2004) Cell 119, 517–528 [DOI] [PubMed] [Google Scholar]
- 14. Maine G. N., Burstein E. (2007) Cell Cycle 6, 672–676 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15. Burstein E., Hoberg J. E., Wilkinson A. S., Rumble J. M., Csomos R. A., Komarck C. M., Maine G. N., Wilkinson J. C., Mayo M. W., Duckett C. S. (2005) J. Biol. Chem. 280, 22222–22232 [DOI] [PubMed] [Google Scholar]
- 16. Maine G. N., Mao X., Komarck C. M., Burstein E. (2007) EMBO J. 26, 436–447 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17. Klomp A. E., van de Sluis B., Klomp L. W., Wijmenga C. (2003) J. Hepatol. 39, 703–709 [DOI] [PubMed] [Google Scholar]
- 18. Ganesh L., Burstein E., Guha-Niyogi A., Louder M. K., Mascola J. R., Klomp L. W., Wijmenga C., Duckett C. S., Nabel G. J. (2003) Nature 426, 853–857 [DOI] [PubMed] [Google Scholar]
- 19. van de Sluis B., Muller P., Duran K., Chen A., Groot A. J., Klomp L. W., Liu P. P., Wijmenga C. (2007) Mol. Cell. Biol. 27, 4142–4156 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20. Biasio W., Chang T., McIntosh C. J., McDonald F. J. (2004) J. Biol. Chem. 279, 5429–5434 [DOI] [PubMed] [Google Scholar]
- 21. Drévillon L., Tanguy G., Hinzpeter A., Arous N., de Becdelièvre A., Aissat A., Tarze A., Goossens M., Fanen P. (2011) PLoS One 6, e18334 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22. Müller T., van de Sluis B., Zhernakova A., van Binsbergen E., Janecke A. R., Bavdekar A., Pandit A., Weirich-Schwaiger H., Witt H., Ellemunter H., Deutsch J., Denk H., Müller W., Sternlieb I., Tanner M. S., Wijmenga C. (2003) J. Hepatol 38, 164–168 [DOI] [PubMed] [Google Scholar]
- 23. Coronado V. A., Bonneville J. A., Nazer H., Roberts E. A., Cox D. W. (2005) Clin. Genet. 68, 548–551 [DOI] [PubMed] [Google Scholar]
- 24. Stuehler B., Reichert J., Stremmel W., Schaefer M. (2004) J. Mol. Med. 82, 629–634 [DOI] [PubMed] [Google Scholar]
- 25. Narindrasorasak S., Kulkarni P., Deschamps P., She Y. M., Sarkar B. (2007) Biochemistry 46, 3116–3128 [DOI] [PubMed] [Google Scholar]
- 26. Vonk W. I., Wijmenga C., Berger R., van de Sluis B., Klomp L. W. (2010) J. Biol. Chem. 285, 28991–29000 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27. van de Sluis B., Mao X., Zhai Y., Groot A. J., Vermeulen J. F., van der Wall E., van Diest P. J., Hofker M. H., Wijmenga C., Klomp L. W., Cho K. R., Fearon E. R., Vooijs M., Burstein E. (2010) J. Clin. Invest. 120, 2119–2130 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28. Burstein E., Ganesh L., Dick R. D., van De Sluis B., Wilkinson J. C., Klomp L. W., Wijmenga C., Brewer G. J., Nabel G. J., Duckett C. S. (2004) EMBO J. 23, 244–254 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29. Maine G. N., Gluck N., Zaidi I. W., Burstein E. (2009) Cold Spring Harb. Protoc. 10.1101/pdb.prot5318 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30. Mao X., Gluck N., Li D., Maine G. N., Li H., Zaidi I. W., Repaka A., Mayo M. W., Burstein E. (2009) Genes Dev. 23, 849–861 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31. Sumara I., Quadroni M., Frei C., Olma M. H., Sumara G., Ricci R., Peter M. (2007) Dev. Cell 12, 887–900 [DOI] [PubMed] [Google Scholar]
- 32. van de Sluis B., Groot A. J., Wijmenga C., Vooijs M., Klomp L. W. (2007) Cell Cycle 6, 2091–2098 [DOI] [PubMed] [Google Scholar]
- 33. Bornstein G., Ganoth D., Hershko A. (2006) Proc. Natl. Acad. Sci. U.S.A. 103, 11515–11520 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34. Duda D. M., Borg L. A., Scott D. C., Hunt H. W., Hammel M., Schulman B. A. (2008) Cell 134, 995–1006 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35. Li H., Wittwer T., Weber A., Schneider H., Moreno R., Maine G. N., Kracht M., Schmitz M. L., Burstein E. (2011) Oncogene, in press [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36. Geng H., Wittwer T., Dittrich-Breiholz O., Kracht M., Schmitz M. L. (2009) EMBO Rep. 10, 381–386 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37. van de Sluis B., Groot A. J., Vermeulen J., van der Wall E., van Diest P. J., Wijmenga C., Klomp L. W., Vooijs M. (2009) PLoS One 4, e7332 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38. Schmidt M. W., McQuary P. R., Wee S., Hofmann K., Wolf D. A. (2009) Mol. Cell 35, 586–597 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39. Chua Y. S., Boh B. K., Ponyeam W., Hagen T. (2011) PLoS One 6, e16071 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40. Lee J. E., Sweredoski M. J., Graham R. L., Kolawa N. J., Smith G. T., Hess S., Deshaies R. J. (2011) Mol. Cell Proteomics 10, DOI 10.1074/mcp.M110.006460 [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.