Abstract
Background
Stevens-Johnson syndrome (SJS) and Toxic Epidermal Necrolysis (TEN) are rare but extremely severe cutaneous adverse drug reactions in which drug-specific associations with HLA-B alleles were described.
Objectives
To investigate genetic association at a genome-wide level on a large sample of SJS/TEN patients.
Methods
We performed a genome wide association study on a sample of 424 European cases and 1,881 controls selected from a Reference Control Panel.
Results
Six SNPs located in the HLA region showed significant evidence for association (OR range: 1.53-1.74). The haplotype formed by their risk allele was more associated with the disease than any of the single SNPs and was even much stronger in patients exposed to allopurinol (ORallopurinol = 7.77, 95%CI = [4.66; 12.98]). The associated haplotype is in linkage disequilibrium with the HLA-B*5801 allele known to be associated with allopurinol induced SJS/TEN in Asian populations.
Conclusion
The involvement of genetic variants located in the HLA region in SJS/TEN is confirmed in European samples, but no other locus reaches genome-wide statistical significance in this sample that is also the largest one collected so far. If some loci outside HLA play a role in SJS/TEN, their effect is thus likely to be very small.
Background
Adverse drug reactions are a major public health issue as they represent an important cause of morbidity and mortality [1]. Skin lesions are frequent expressions of adverse drug reactions with, for some drugs, up to 10% of cutaneous reactions observed [2,3]. SJS and TEN are severe cutaneous adverse reactions characterized by the development of acute exanthema which progresses towards limited (in SJS) or widespread (in TEN) blistering and erosion of the skin and mucous membranes [4]. SJS and TEN are thus considered to be two different forms of the same disease with TEN representing the most severe form [5]. The incidence of SJS/TEN is estimated to be of approximately 1-2 patients per million individuals per year [6]. It is thus a very rare disease but with a high morbidity and mortality (reaching up to 45% in TEN) that requires intensive treatment.
SJS/TEN is not associated with a single drug or a single group of drugs but several different drugs have been involved. A limited number of drugs however are more often associated with the disease: antibacterial sulfonamides (especially sulfamethoxazole), allopurinol, which is the most frequent drug involved [7], carbamazepine, lamotrigine, phenobarbital, phenytoine, non-steroidal anti-inflammatory drugs (NSAIDs) of the oxicam type and neviparine [4,8]. Only a small number of individuals exposed to these "highly suspected" drugs develop the disease and a genetic susceptibility has been suggested [9-12].
An association with HLA was reported more than 20 years ago [12,13]. More recently, studies in the Han Chinese population have involved the HLA-B locus with very strong drug-specific associations: the HLA-B*1502 allele was found in all carbamazepine-induced SJS/TEN patients [14] and the HLA-B*5801 in all allopurinol-induced SJS/TEN patients [15]. An investigation of HLA-B associations in European samples did not detect the association between the HLA-B*1502 allele and carbamazepine-induced SJS/TEN but did report a strong but not complete association between HLA-B*5801 and the allopurinol-induced disease [16,17]. Furthermore, it has also been suggested that HLA genetic predisposition may not be the same for SJS and TEN and that it might thus be important to take into account the disease severity in association tests [18].
Apart from HLA, several other candidate genes have been tested for association with SJS/TEN, mostly genes involved in the immune response, in the inflammation process or in drug metabolism [18-21]. However, no association with these genes has been consistently found and it has been suggested that rather than focusing on candidate genes, a genome-wide association study might provide more insights into the genetic susceptibility of adverse drug reactions [21].
In the context of the RegiSCAR project (European Registry of Severe Cutaneous Adverse Reactions to Drugs and Collection of Biological Samples) DNA of 563 cases of SJS/TEN was collected. This is the largest available sample of SJS/TEN patients in the world with accurate medical information regarding the severity of the disease and the history of drug intake. These patients were genotyped at the Centre National de Genotypage (CNG) using Illumina 317 K chips and a Genome Wide Association Study (GWAS) was conducted against controls selected from the CNG European Reference Control Panel [22].
Methods
Patients
A total of 563 cases (226 males and 337 females, sex-ratio = 0.67) were included in this study. They were collected as part of the "European Registry of Severe Cutaneous Adverse Reactions" (RegiSCAR) (see http://regiscar.uni-freiburg.de/) in six countries (Austria (1 case), France (184 cases), Germany (331 cases), Israel (14), Italy (26) and The Netherlands (7)). All of them had a diagnosis of SJS (268 cases, 48%), SJS/TEN overlap (181 cases, 32%) or TEN (114 cases, 20%) validated as probable or definitive by the expert committee of RegiSCAR blindly from information on drug exposure.
For each patient, written informed consent was obtained and a blood sample was taken for genomic DNA extraction that was carried out at the CEPH-Fondation Jean Dausset (France) (http://www.cephb.fr) until 2005 and at the biobank of the CIC-Henri Mondor / Créteil after 2005.
The 563 patients were genotyped on Illumina 317 K chips in France at the CNG (http://www.cng.fr) for a total of 318,127 SNPs (among which 309,091 were located on autosomes). Stringent quality-control (QC) was performed using Plink v1.06 [23] that led to the exclusion of 68 out of the 563 cases for low genotyping (MIND > 0.05) and 15,088 SNPs (14,343 autosomal SNPs) for missing data (GENO > 0.05). After QC, a total of 495 affected individuals were considered in the analysis for whom genotypes on 303,039 SNPs (294,748 located on autosomes) were available. The total genotyping rate in these 495 remaining individuals was 0.95.
Controls
Controls were selected from the CNG European Reference Panel collected at the CNG [22]. This panel includes 5,847 unrelated individuals from 13 European countries who are all genotyped on Illumina 317 K chips. Since patients collected as part of the RegiSCAR project were mostly from France and Germany, we decided to consider only the 1,881 individuals from the Reference Control panel originating from these two countries (653 from Germany and 1,228 from France).
Quality control and principal component analysis of genotype data
A second quality control of the data was performed where markers that were missing in more than 5% of the controls or had significant different missing rates between cases and controls were removed. A total of 25,834 autosomal markers were excluded.
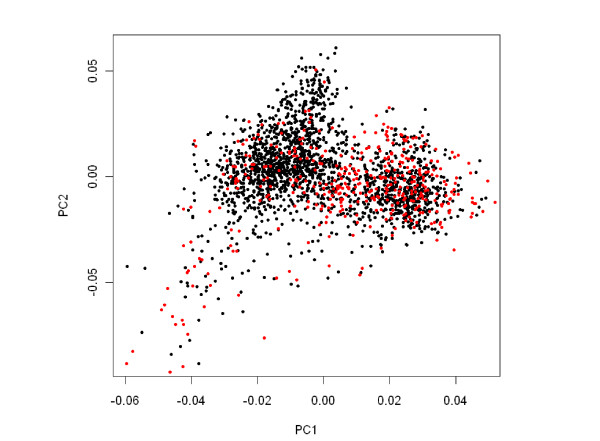
To assess the level of population stratification in the sample, a principal component analysis (PCA) was performed using the genotypes of the 495 cases and 1,881 controls for a sub-panel of 35,232 markers obtained after pruning based on linkage disequilibrium using Plink (option indep-pairwise 50 5 0.1). The program smartPCA from the Eigenstrat package [24,25] was used with the default options. A total of 71 patients were excluded from the study as they were found to be outliers. Those were mostly individuals with a suspected African or Asian ancestry. Plots of the first two principal components (PCs) are provided in Figure 1.
Figure 1.
Plots of top two principal components for cases (in red) and controls (in black).
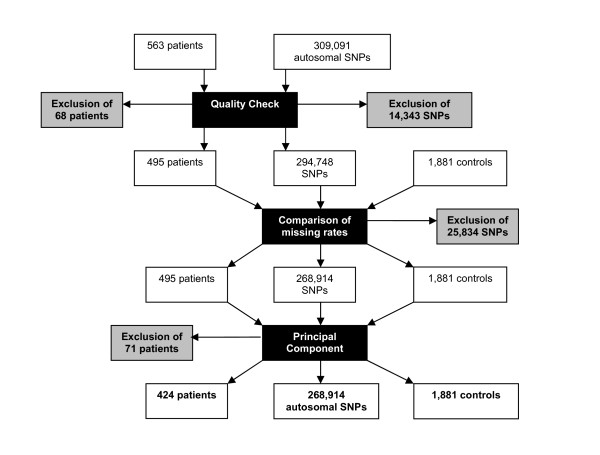
As shown in the flowchart in Figure 2, a total of 268,914 autosomal SNPs were kept for the association testing in a sample of 424 SJS/TEN patients and 1,881 controls.
Figure 2.
Flowchart summarizing the different steps of the quality control of the data.
Drug exposure
Information on drug exposure was collected by interviewing patients about their drug intakes and by consulting medical records. Reactions were considered to be potentially caused by allopurinol, carbamazepine or phenytoin if the drug has been taken between 4 and 10 days before the onset of the disease and if the drug has not been started more than 42 days before onset. This resulted in a group of 57 patients (13.4%) with potentially allopurinol-induced reactions, a group of 25 patients (5.9%) with potentially carbamazepine-induced reactions and a group of 19 patients (4.5%) with potentially phenytoin-induced reactions. Because of the low number in the latter two groups we decided to only compare the allopurinol group and the group with other drugs to the controls in this study. Note that because of the stringent time criterion used to determine allopurinol as culprit drug there might be some patients with an allopurinol-induced reaction left in the group of patients with a culprit drug other than allopurinol. For this reason, this analysis was further enforced by an additional analysis where any allopurinol exposed patient (irrelevant of time of usage) was assigned to the allopurinol group resulting in a higher sensitivity with respect to allopurinol-induced reaction while the specificity is decreased.
Association testing
Association was tested using the PC-corrected Armitage-trend test implemented in Eigenstrat [24]. A sensitivity analysis was performed to determine the number of PCs that should be accounted for and we decided to adjust only on the first two PCs (the genomic control coefficient lambda [26] was 1.023 when adjusting on the top 2 PCs, 1.019 when adjusting on the top 10 PCs and 1.016 when adjusting on the top 20).
Odds-ratios (OR) and confidence intervals of the different genotypes were computed using the logistic regression model implemented in Plink v1.06 [23].
To study the influence of the drug involved in the disease, a second study was performed where the multinomial logistic model implemented in Stata v10 [27] was used as in [28]. The disease status D was coded in 3 classes: 0 unaffected (controls), 1 affected with a suspected allopurinol-induced disease, 2 affected with another suspected drug induced disease. At each SNP, an additive model was considered and two ORs were computed to model the effect of the SNP among allopurinol-induced affected individuals (ORallopurinol) and among other affected individuals (ORotherdrugs). The null hypothesis tested was one of no effect of the SNP in any of the strata, i.e. ORallopurinol = ORotherdrugs = 1. It was tested using a 2 degrees-of-freedom chi-square test. The significance of the difference between ORallopurinol and ORotherdrugs was also tested in this model using the corresponding one degree-of-freedom chi-square test.
In the same way, a multinomial logistic model was used to determine whether genetic differences exist depending on the disease severity. A four class disease status was used: 0 unaffected (controls), 1 SJS, 2 SJS/TEN overlap and 3 TEN.
Furthermore, haplotype association was tested using Plink v1.06 [23].
For all the logistic regression modelling and haplotype association testing, an adjustment on the first two PCs obtained from the PCA of the genotype data was used.
Results
The clinical details of the 424 patients remaining after quality-control are summarized in Table 1: 61.3% are female, 47.2% have a SJS phenotype, 34.4% a SJS/TEN overlap phenotype and 18.4% a TEN phenotype. A majority of the patients was sampled from France (110 patients) and Germany (277 patients). History of drug intake was carefully monitored and a subgroup of 57 patients (13.4%) with potentially allopurinol-induced reactions was identified. There is no difference in phenotype distribution between males and females but there are some differences depending on the drug and the country where samples were collected. There is a lack of TEN among allopurinol-induced patients (test of homogeneity of the three disease forms in allopurinol induced versus non-allopurinol induced partients: chi-square = 7.89, p-value = 0.019) and a lack of TEN among patients from Germany as compared to patients from France (chi-square = 23.68, p-value = 7.2 10-6). The two variables however are not independent since allopurinol is more used in Germany than in some other countries and in particular France.
Table 1.
Description of the sub-sample of 424 patients selected for the analysis
| Number of patients | SJS | SJS/TEN | TEN | Total |
|---|---|---|---|---|
| Total | 200 (47.2%) | 146 (34.4%) | 78 (18.4%) | 424 |
| Females | 124 | 89 | 47 | 260 (61.3%) |
| Allopurinol-induced | 33 | 21 | 3 | 57 (13.4%) |
| From France | 41 | 32 | 37 | 110 (25.9%) |
| From Germany | 145 | 97 | 35 | 277 (65.3%) |
| From Other countries | 14 | 17 | 6 | 37 (8.7%) |
The number of patients (proportion of the total of 424 patients in percents) with the different phenotypes is presented in the total sample, among the female patients and as a function of the country where patients were identified.
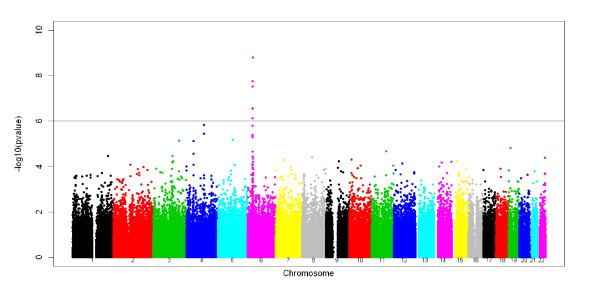
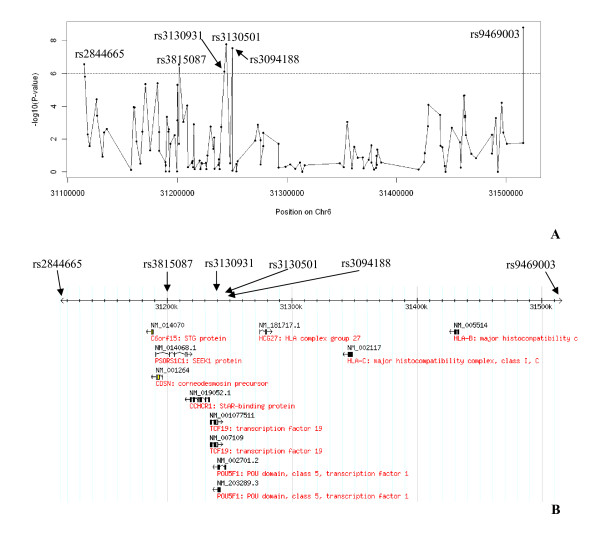
For association testing, the 424 patients were compared to the 1,881 controls from the reference panel originating from France (1,228 individuals) and Germany (653 individuals) who were also the closest controls to the cases on the PCA plot (Figure 1). We then tested each SNP for association controlling on the first two PCs of the PCA using Eigenstrat [24]. Results are presented in Figure 3. One genome-wide significant signal was detected in the MHC region on chromosome 6 where 6 SNPs showed False Discovery Rate adjusted q-values below 5% (see Figure 4 and Table 2). The most significant SNP, rs9469003, is located at position 31,515,807, ~85 kb upstream of the HLA-B locus and has an OR of 1.73 (95%CI = [1.44;2.08]). The five other SNPs are all located in a more telomeric region, 250 kb apart from rs9469003, in between positions 31,114,834 and 31,250,224 (see Figure 5). There is very limited Linkage Disequilibrium (LD) between these five SNPs and the top one and the six SNPs defined 13 haplotypes with a frequency of 1% or above. The haplotype CACGAC formed by the risk allele at each locus showed the strongest association with the disease with an OR of 2.84 (95%CI = [2.03; 3.98]) that is significantly higher than the one observed for rs9469003 only. To dissect further this haplotypic association, the haplotype conditioning test implemented in Plink v1.06 was used and we found that none of the 6 SNPs by itself has an independent effect nor explains the haplotype association.
Figure 3.
Results of the genome-wide association screen. The -log(p-value) of the PC-corrected Armitage-trend test implemented in Eigenstrat are plotted for the 268,818 autosomal SNPs that passed the QC with the different colors representing the different chromosome. The horizontal line represents the 10-6 p-value threshold that also corresponds to a False Discovery Rate q-value of 5%.
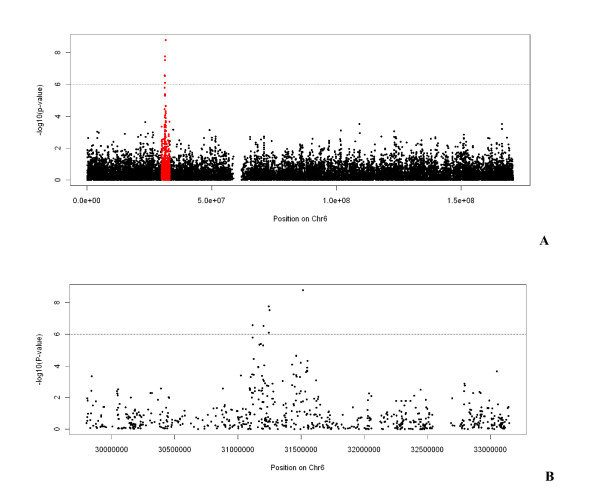
Figure 4.
Results of the association test on chromosome 6. A The -log(p-value) of the PC-corrected Armitage-trend test implemented in Eigenstrat are plotted for the 18,1278 SNPs located on chromosome 6 as a function of their position on the chromosome in base pairs (Build 36). SNPs located in the HLA region are highlighted in red. B Detail of the association signals for the 706 SNPs located in the HLA region. The horizontal line represents the 10-6 p-value threshold.
Table 2.
List of most significant SNPs associated with SJS/TEN
| SNP | Chr | Position Build 36 | RiskAllele (Other Allele) | Risk Allele Frequency in Cases | Risk Allele Frequency in Controls | PC corrected Trend testb | p-valueb | q-valuec | OddsRatiod | 95% CId | Pvalue of HW test | Annotationa |
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| rs2844665 | 6 | 31114834 | C (T) | 0.72 | 0.62 | 26.46 | 2.69 10-7 | 1.48 10-2 | 1.54 | 1.30-1.82 | 0.77 | C6orf205 flanking_3UTR |
| rs3815087 | 6 | 31201566 | A (G) | 0.31 | 0.21 | 26.32 | 2.89 10-7 | 1.54 10-2 | 1.53 | 1.29-1.80 | 0.08 | PSORS1C1 UTR |
| rs3130931 | 6 | 31242867 | C (T) | 0.77 | 0.69 | 24.41 | 7.78 10-7 | 3.34 10-2 | 1.54 | 1.29-1.84 | 0.66 | POU5F1 Intron |
| rs3130501 | 6 | 31244432 | G (A) | 0.83 | 0.74 | 31.78 | 1.73 10-8 | 0.22 10-2 | 1.74 | 1.43-2.13 | 0.59 | POU5F1 Intron |
| rs3094188 | 6 | 31250224 | A (C) | 0.72 | 0.63 | 30.75 | 2.93 10-8 | 0.26 10-2 | 1.59 | 1.34-1.88 | 0.06 | POU5F1 Flanking_5UTR |
| rs9469003 | 6 | 31515807 | C (T) | 0.24 | 0.15 | 36.41 | 1.60 10-9 | 0.04 10-2 | 1.73 | 1.44-2.08 | 0.003 | HCP5 flanking_5UTR |
a Annotations were obtained from the Illumina annotation file.
b Results obtained using the test implemented in Eigenstrat
c q-values (FDR adjusted p-values) were obtained by using the R package fdrtool [50].
d Results obtained using plink logistic regression test accounting for the top 2 PC.
Figure 5.
Association results in the 401 kb region encompassing the top six signals. A The -log(p-value) of the PC-corrected Armitage-trend test are plotted against the chromosome 6 positions for the 126 SNPs located in the region. B Listing of genes located in this region based on the HapMap web-browser.
Using a multinomial logistic regression approach, we found that the association with haplotype CACGAC was much stronger in the subgroup of patients with an allopurinol induced disease (ORallopurinol = 7.77, 95%CI = [4.66; 12.98]) than in other patients (ORotherdrugs = 1.92, 95%CI = [1.40;2.64]). The equality of odds-ratio (ORallopurinol = ORotherdrugs) was strongly rejected (p = 6.56 × 10-7). Results were the same in France and Germany a shown in Additional File 1 Tables S1 and S2.
To determine how much of the observed signal could be explained by the known association between allopurinol-induced SJS/TEN and HLA-B*5801, we took advantage of the availability of HLA-B two-digit resolution genotypes for 74 of the 424 patients [16]. Among the 74 HLA-B genotyped patients, 11 were carriers of an HLA-B58 allele and had an allopurinol-induced disease. The frequency of the CACGAC haplotype is increased in HLA-B58 carriers as compared to non HLA-B58 carriers (32.44% versus 10.44%, p-value of the test adjusted on the top 2 PC = 0.0052) but the linkage disequilibrium is not complete. Note however that given the small sample size of HLA-B typed individuals, the power was quite low to detect this difference (power of 60% at a type-one error rate level of 5%).
Discussion
Discovering genes involved in severe cutaneous adverse reactions and especially SJS/TEN is a major challenge for pharmacogenetics as these reactions, when not fatal, are a sword of Damocles for those who already had the disease and fear to take any drug. Despite important efforts, only genes located in the HLA region have been identified so far. One possible explanation for this lack of success is the limited sample sizes of patients available. Indeed SJS/TEN is fortunately a very rare disease and in most studies, sample sizes rarely exceed one or two hundreds of patients, making it difficult to investigate more than a few candidate genes. Through a collaborative effort, the RegiSCAR group was able to collect detailed medical information and DNA of more than half a thousand of patients from Europe who were genotyped on Illumina 317 K chips. By comparing their genotypes to the ones of 1,881 controls genetically matched for the country of origin, we were able to study the SJS/TEN association at a genome-wide level on European samples. Apart from six SNPs located in the HLA region, no other locus was found to be associated with the disease at a high enough significance level to ensure it is not false positive due to the multiple tests performed. Sample sizes of half a thousand are not enough to ensure a good power to detect common variants with small effects (OR in the range between 1.1 and 1.3) similar to those identified in several common diseases such as diabetes or different cancers [29] where larger samples are easier to collect. However, given the 424 patients remaining after Quality Control, the study was powerful enough to detect common variants with modest effects (the power exceeded 80% to detect variants with an allele frequency above 15% conferring ORs above 1.7 under a multiplicative model). The fact that only the HLA region is detected, suggests that there might not be any other common variant that confer a substantial increase in disease risk and could thus be of interest as a predictive factor. These power computations might however be too optimistic as they are based on the assumption that the same genetic variants might be involved in cases of SJS/TEN associated to different drugs. If we consider that there might exist some levels of genetic heterogeneity depending on the drugs involved as it is the case for HLA associations, the power can be dramatically reduced. For example, considering only the 57 allopurinol-exposed cases, the power to detect a variant of frequency 15% conferring an OR of 2 is then only 5% at a nominal type-one error rate value of 10-6. On the other hand, the power to detect a similar effect than the one observed at rs9469003 (i.e., risk allele of frequency of 15% with an associated OR of ~4 among allopurinol-induced SJS/TEN) is 99%, showing that the study was not underpowered to detect strong effects even if they were restricted to a small subset of patients.
It is true however that we might have missed some variants with important effects, especially those with minor allele frequencies below 5% that are not well covered by SNP-chips and it could thus be of interest to investigate the association with rare variants. If the "common disease-common variant hypothesis" was believed to explain the susceptibility to multifactorial diseases [30,31], results from GWAS have shown that common variants only explain a minor part of most common disease heritabilities. It has been suggested that rare variants could explain at least part of this "missing heritability" and indeed, rare variants have been found in several diseases [32-36]. Interestingly these rare variants are often functional variants with a direct impact on the protein functionality and they usually confer a much stronger increase in disease risk than common variants. They are also more likely to be affected by some moderate levels of negative selection [37,38]. For diseases such as adverse drug reactions, it is not unlikely that the genes and variants involved could be under selection. Indeed if most of the drugs have been introduced too recently to be directly responsible for selective pressures, they are often derived from nutrients that have been consumed by humans for a long time. Thus, genes involved in drug metabolism can potentially be involved in natural selection and this was confirmed by previous studies [39-41]. However, the implication of drug metabolism genes in SJS/TEN remains to be established.
Concerning the association with HLA detected in this sample, we confirmed that it is drug-specific with the strongest association found for the group of patients where allopurinol is suspected to be the cause of the disease. However the disease association was still detectable after exclusion of all patients exposed to allopurinol (whether considered as causing the reaction or not) (OR = 2.13, 95% CI = [1.41; 3.23]). This suggests that different HLA alleles are probably involved depending on the drug, and the 6 SNPs identified could be those that were in linkage disequilibrium (LD) with several of these HLA alleles. To further explore this hypothesis, we tried to impute HLA-B genotypes from the SNP data [42] but we were not successful because of the presence of multiple rare alleles at this locus such as HLA-B58 itself that is only present in one of the European families genotyped by de Bakker et al. [43] precluding the possibility to find appropriate tags for this allele using this sample. To overcome the problem, we tried to determine tag SNPs using our own HLA-B genotyped sample that was enriched in HLA-B58 but we did not find any SNP or group of SNPs that was correlated enough with the HLA-B58 allele (the maximum r2 value was below 20%). This poor HLA-B tagging ability of the SNPs could certainly explain why the association found here in relation with allopurinol is much weaker than the one reported in studies on European samples using resolved HLA-B alleles [16,44]. However, even in these two latter studies, the association with HLA-B*5801 is much weaker than the one found in Han Chinese [15] or Thai populations [45] where all patients with allopurinol-induced SJS/TEN are carriers of this allele. It is thus possible that the HLA genetic determinants involved in SJS/TEN are not the HLA-B alleles themselves but some loci in LD with them. Further studies in this genomic region would certainly be necessary to better understand the mechanisms involved.
On a methodological point of view, this study illustrates how Reference Control Panels could be used to test for association when only cases are genotyped. Only patients were genotyped here and controls were selected from the CNG European Control Panel [22] based on their country of origin that was either France and Germany, since those were the two main countries where the cases come from. We also tried to use the whole panel of controls and correct for population stratification using Principal Component Analysis (PCA) and an adjustment on up to 20 PCs instead of the 2 used here. We found that including controls that were more genetically distant to the cases does not really improve the signal in the HLA region and instead could lead to false positive results in other genomic regions. This is somewhat in contradiction with the results recently reported by Zhuang et al. [46] and will probably need to be further investigated. We also used some alternative approaches where controls were genetically matched to cases using either a distance based on the identity by state as described by Guan et al. [47] or a distance derived from the top PCs of the PCA performed to identify axes of variations similar to the one proposed by Luca et al. [48]. Tests accounting for the matching units were then performed and we found that only the HLA region pointed out with significance levels in the same order of magnitude as the ones obtained here. These matching strategies might however be of interest as they could allow the analysis of more heterogeneous samples by including for example those non-European RegiSCAR cases that were excluded here and matching them to individuals from the CEPH-Human Genome Diversity Panel [49].
Conclusion
Our study confirms the involvement of genetic variants located in the HLA region in the susceptibility to SJS/TEN in European samples, especially in association with allopurinol. No difference is seen depending on the disease severity and no other locus reaches genome-wide association in this sample that is also the largest one collected so far.
Competing interests
The authors declare that they have no competing interests.
Authors' contributions
MM, MS, JCR, AH, ML designed the study, DZ, ML supervised the genotyping, LN, YL, JCR, MM, PS were responsible for clinical data collection, RK, EG performed the statistical analysis, EG wrote the manuscript and all authors contributed to and approved the final draft.
Supplementary Material
Supporting material for manuscript "Genome-Wide Association study of Stevens-Johnson Syndrome and Toxic Epidermal Necrolysis in Europe" Results of association tests for the top SNP (Table S1) and for the most associated haplotype (Table S2) after stratification on the country of origin.
Contributor Information
Emmanuelle Génin, Email: emmanuelle.genin@inserm.fr.
Martin Schumacher, Email: ms@imbi.uni-freiburg.de.
Jean-Claude Roujeau, Email: jean-claude.roujeau@wanadoo.fr.
Luigi Naldi, Email: luigi.naldi@gised.it.
Yvonne Liss, Email: dzh@uniklinik-freiburg.de.
Rémi Kazma, Email: remi.kazma@gmail.com.
Peggy Sekula, Email: ps@imbi.uni-freiburg.de.
Alain Hovnanian, Email: alain.hovnanian@inserm.fr.
Maja Mockenhaupt, Email: dzh@uniklinik-freiburg.de.
Acknowledgements
We are indebted to all patients whose participation made this study possible as well as to their treating physicians for the great cooperation. We thank the Centre National de Génotypage (CNG) and especially Mark Lathrop and Diana Zelenika for conducting the genotyping study and providing the control data. We also thank Alexis Sidoroff (Austria; data acquisition), Alexander Hellmer (Germany; data management), Konrad Bork, Uwe-Frithjof Haustein, Dieter Vieluf (Germany; clinical case review/expert committee), Davide Zenoni (Italy; data acquisition) and Jan Nico Bouwes Bavinck (The Netherlands; data acquisition) for their support in the study.
The RegiSCAR-study was funded by grants from the European Commission (QLRT-2002-01738), GIS-Institut des Maladies Rares and INSERM (4CH09G) in France, and by a consortium of pharmaceutical companies (Bayer Vital, Boehringer-Ingelheim, Cephalon, GlaxoSmithKline, MSD Sharp and Dohme, Merck, Novartis, Pfizer, Roche, Sanofi-Aventis, Servier). Maja Mockenhaupt received the Else Kröner Memorial Stipendium for support of clinical research through Else Kröner-Fresenius-Foundation. The Centre National de Génotypage (CNG), Paris, France, provided the funding for the genotyping.
References
- Lazarou J, Pomeranz BH, Corey PN. Incidence of adverse drug reactions in hospitalized patients: a meta-analysis of prospective studies. Jama. 1998;279(15):1200–1205. doi: 10.1001/jama.279.15.1200. [DOI] [PubMed] [Google Scholar]
- Svensson CK, Cowen EW, Gaspari AA. Cutaneous drug reactions. Pharmacol Rev. 2001;53(3):357–379. [PubMed] [Google Scholar]
- Wolverton SE. Update on cutaneous drug reactions. Adv Dermatol. 1997;13:65–84. [PubMed] [Google Scholar]
- Roujeau JC, Stern RS. Severe adverse cutaneous reactions to drugs. N Engl J Med. 1994;331(19):1272–1285. doi: 10.1056/NEJM199411103311906. [DOI] [PubMed] [Google Scholar]
- Auquier-Dunant A, Mockenhaupt M, Naldi L, Correia O, Schroder W, Roujeau JC. Correlations between clinical patterns and causes of erythema multiforme majus, Stevens-Johnson syndrome, and toxic epidermal necrolysis: results of an international prospective study. Arch Dermatol. 2002;138(8):1019–1024. doi: 10.1001/archderm.138.8.1019. [DOI] [PubMed] [Google Scholar]
- Rzany B, Mockenhaupt M, Baur S, Schroder W, Stocker U, Mueller J, Hollander N, Bruppacher R, Schopf E. Epidemiology of erythema exsudativum multiforme majus, Stevens-Johnson syndrome, and toxic epidermal necrolysis in Germany (1990-1992): structure and results of a population-based registry. J Clin Epidemiol. 1996;49(7):769–773. doi: 10.1016/0895-4356(96)00035-2. [DOI] [PubMed] [Google Scholar]
- Halevy S, Ghislain PD, Mockenhaupt M, Fagot JP, Bouwes JN Bavinck, Sidoroff A, Naldi L, Dunant A, Viboud C, Roujeau JC. Allopurinol is the most common cause of Stevens-Johnson syndrome and toxic epidermal necrolysis in Europe and Israel. J Am Acad Dermatol. 2008;58(1):25–32. doi: 10.1016/j.jaad.2007.08.036. [DOI] [PubMed] [Google Scholar]
- Mockenhaupt M, Viboud C, Dunant A, Naldi L, Halevy S, Bouwes JN Bavinck, Sidoroff A, Schneck J, Roujeau JC, Flahault A. Stevens-Johnson syndrome and toxic epidermal necrolysis: assessment of medication risks with emphasis on recently marketed drugs. The EuroSCAR-study. J Invest Dermatol. 2008;128(1):35–44. doi: 10.1038/sj.jid.5701033. [DOI] [PubMed] [Google Scholar]
- Pirmohamed M, Park BK. Genetic susceptibility to adverse drug reactions. Trends Pharmacol Sci. 2001;22(6):298–305. doi: 10.1016/S0165-6147(00)01717-X. [DOI] [PubMed] [Google Scholar]
- Melsom RD. Familial hypersensitivity to allopurinol with subsequent desensitization. Rheumatology (Oxford) 1999;38(12):1301. doi: 10.1093/rheumatology/38.12.1301. [DOI] [PubMed] [Google Scholar]
- Pellicano R, Silvestris A, Iannantuono M, Ciavarella G, Lomuto M. Familial occurrence of fixed drug eruptions. Acta Derm Venereol. 1992;72(4):292–293. [PubMed] [Google Scholar]
- Roujeau JC, Huynh TN, Bracq C, Guillaume JC, Revuz J, Touraine R. Genetic susceptibility to toxic epidermal necrolysis. Arch Dermatol. 1987;123(9):1171–1173. doi: 10.1001/archderm.123.9.1171. [DOI] [PubMed] [Google Scholar]
- Roujeau JC, Bracq C, Huyn NT, Chaussalet E, Raffin C, Duedari N. HLA phenotypes and bullous cutaneous reactions to drugs. Tissue Antigens. 1986;28(4):251–254. doi: 10.1111/j.1399-0039.1986.tb00491.x. [DOI] [PubMed] [Google Scholar]
- Chung WH, Hung SI, Hong HS, Hsih MS, Yang LC, Ho HC, Wu JY, Chen YT. Medical genetics: a marker for Stevens-Johnson syndrome. Nature. 2004;428(6982):486. doi: 10.1038/428486a. [DOI] [PubMed] [Google Scholar]
- Hung SI, Chung WH, Liou LB, Chu CC, Lin M, Huang HP, Lin YL, Lan JL, Yang LC, Hong HS, Chen MJ, Lai PC, Wu MS, Chu CY, Wang KH, Chen CH, Fann CS, Wu JY, Chen YT. HLA-B*5801 allele as a genetic marker for severe cutaneous adverse reactions caused by allopurinol. Proc Natl Acad Sci USA. 2005;102(11):4134–4139. doi: 10.1073/pnas.0409500102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lonjou C, Borot N, Sekula P, Ledger N, Thomas L, Halevy S, Naldi L, Bouwes-Bavinck JN, Sidoroff A, de Toma C, Schumacher M, Roujeau JC, Hovnanian A, Mockenhaupt M. RegiSCAR study group. A European study of HLA-B in Stevens-Johnson syndrome and toxic epidermal necrolysis related to five high-risk drugs. Pharmacogenetics and genomics. 2008;18(2):99–107. doi: 10.1097/FPC.0b013e3282f3ef9c. [DOI] [PubMed] [Google Scholar]
- Lonjou C, Thomas L, Borot N, Ledger N, de Toma C, LeLouet H, Graf E, Schumacher M, Hovnanian A, Mockenhaupt M, Roujeau JC. RegiSCAR Group. A marker for Stevens-Johnson syndrome ...: ethnicity matters. Pharmacogenomics J. 2006;6(4):265–268. doi: 10.1038/sj.tpj.6500356. [DOI] [PubMed] [Google Scholar]
- Pirmohamed M, Arbuckle JB, Bowman CE, Brunner M, Burns DK, Delrieu O, Dix LP, Twomey JA, Stern RS. Investigation into the multidimensional genetic basis of drug-induced Stevens-Johnson syndrome and toxic epidermal necrolysis. Pharmacogenomics. 2007;8(12):1661–1691. doi: 10.2217/14622416.8.12.1661. [DOI] [PubMed] [Google Scholar]
- Miller JW. Of race, ethnicity, and rash: the genetics of antiepileptic drug-induced skin reactions. Epilepsy Curr. 2008;8(5):120–121. doi: 10.1111/j.1535-7511.2008.00263.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Abe R. Toxic epidermal necrolysis and Stevens-Johnson syndrome: soluble Fas ligand involvement in the pathomechanisms of these diseases. J Dermatol Sci. 2008;52(3):151–159. doi: 10.1016/j.jdermsci.2008.06.003. [DOI] [PubMed] [Google Scholar]
- Pirmohamed M. Genetic factors in the predisposition to drug-induced hypersensitivity reactions. Aaps J. 2006;8(1):E20–26. doi: 10.1208/aapsj080103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Heath SC, Gut IG, Brennan P, McKay JD, Bencko V, Fabianova E, Foretova L, Georges M, Janout V, Kabesch M. et al. Investigation of the fine structure of European populations with applications to disease association studies. Eur J Hum Genet. 2008;16(12):1413–1429. doi: 10.1038/ejhg.2008.210. [DOI] [PubMed] [Google Scholar]
- Purcell S, Neale B, Todd-Brown K, Thomas L, Ferreira MA, Bender D, Maller J, Sklar P, de Bakker PI, Daly MJ, Sham PC. PLINK: a tool set for whole-genome association and population-based linkage analyses. American journal of human genetics. 2007;81(3):559–575. doi: 10.1086/519795. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Price AL, Patterson NJ, Plenge RM, Weinblatt ME, Shadick NA, Reich D. Principal components analysis corrects for stratification in genome-wide association studies. Nature genetics. 2006;38(8):904–909. doi: 10.1038/ng1847. [DOI] [PubMed] [Google Scholar]
- Patterson N, Price AL, Reich D. Population structure and eigenanalysis. PLoS genetics. 2006;2(12):e190. doi: 10.1371/journal.pgen.0020190. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Devlin B, Roeder K. Genomic control for association studies. Biometrics. 1999;55(4):997–1004. doi: 10.1111/j.0006-341X.1999.00997.x. [DOI] [PubMed] [Google Scholar]
- StataCorp. College Station, TX. StataCorp LP; 2007. Stata Statistical Software: Release 10. [Google Scholar]
- Kazma R, Babron MC, Genin E. Genetic association and gene-environment interaction: a new method for overcoming the lack of exposure information in controls. Am J Epidemiol. pp. 225–235. [DOI] [PubMed]
- Manolio TA, Brooks LD, Collins FS. A HapMap harvest of insights into the genetics of common disease. J Clin Invest. 2008;118(5):1590–1605. doi: 10.1172/JCI34772. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Reich DE, Lander ES. On the allelic spectrum of human disease. Trends Genet. 2001;17(9):502–510. doi: 10.1016/S0168-9525(01)02410-6. [DOI] [PubMed] [Google Scholar]
- Lohmueller KE, Pearce CL, Pike M, Lander ES, Hirschhorn JN. Meta-analysis of genetic association studies supports a contribution of common variants to susceptibility to common disease. Nature genetics. 2003;33(2):177–182. doi: 10.1038/ng1071. [DOI] [PubMed] [Google Scholar]
- Bodmer W, Bonilla C. Common and rare variants in multifactorial susceptibility to common diseases. Nature genetics. 2008;40(6):695–701. doi: 10.1038/ng.f.136. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Manolio TA, Collins FS, Cox NJ, Goldstein DB, Hindorff LA, Hunter DJ, McCarthy MI, Ramos EM, Cardon LR, Chakravarti A, Cho JH, Guttmacher AE, Kong A, Kruglyak L, Mardis E, Rotimi CN, Slatkin M, Valle D, Whittemore AS, Boehnke M, Clark AG, Eichler EE, Gibson G, Haines JL, Mackay TF, McCarroll SA, Visscher PM. Finding the missing heritability of complex diseases. Nature. 2009;461(7265):747–753. doi: 10.1038/nature08494. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gorlov IP, Gorlova OY, Sunyaev SR, Spitz MR, Amos CI. Shifting paradigm of association studies: value of rare single-nucleotide polymorphisms. American journal of human genetics. 2008;82(1):100–112. doi: 10.1016/j.ajhg.2007.09.006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- McCarthy MI. Exploring the unknown: assumptions about allelic architecture and strategies for susceptibility variant discovery. Genome medicine. 2009;1(7):66. doi: 10.1186/gm66. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fearnhead NS, Winney B, Bodmer WF. Rare variant hypothesis for multifactorial inheritance: susceptibility to colorectal adenomas as a model. Cell cycle (Georgetown, Tex. 2005;4(4):521–525. doi: 10.4161/cc.4.4.1591. [DOI] [PubMed] [Google Scholar]
- Boyko AR, Williamson SH, Indap AR, Degenhardt JD, Hernandez RD, Lohmueller KE, Adams MD, Schmidt S, Sninsky JJ, Sunyaev SR, White TJ, Nielsen R, Clark AG, Bustamante CD. Assessing the evolutionary impact of amino acid mutations in the human genome. PLoS genetics. 2008;4(5):e1000083. doi: 10.1371/journal.pgen.1000083. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kryukov GV, Pennacchio LA, Sunyaev SR. Most rare missense alleles are deleterious in humans: implications for complex disease and association studies. American journal of human genetics. 2007;80(4):727–739. doi: 10.1086/513473. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Luca F, Bubba G, Basile M, Brdicka R, Michalodimitrakis E, Rickards O, Vershubsky G, Quintana-Murci L, Kozlov AI, Novelletto A. Multiple advantageous amino acid variants in the NAT2 gene in human populations. PloS one. 2008;3(9):e3136. doi: 10.1371/journal.pone.0003136. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Patin E, Barreiro LB, Sabeti PC, Austerlitz F, Luca F, Sajantila A, Behar DM, Semino O, Sakuntabhai A, Guiso N, Gicquel B, McElreavey K, Harding RM, Heyer E, Quintana-Murci L. Deciphering the ancient and complex evolutionary history of human arylamine N-acetyltransferase genes. American journal of human genetics. 2006;78(3):423–436. doi: 10.1086/500614. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wilson JF, Weale ME, Smith AC, Gratrix F, Fletcher B, Thomas MG, Bradman N, Goldstein DB. Population genetic structure of variable drug response. Nature genetics. 2001;29(3):265–269. doi: 10.1038/ng761. [DOI] [PubMed] [Google Scholar]
- Leslie S, Donnelly P, McVean G. A statistical method for predicting classical HLA alleles from SNP data. American journal of human genetics. 2008;82(1):48–56. doi: 10.1016/j.ajhg.2007.09.001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- de Bakker PI, McVean G, Sabeti PC, Miretti MM, Green T, Marchini J, Ke X, Monsuur AJ, Whittaker P, Delgado M, Morrison J, Richardson A, Walsh EC, Gao X, Galver L, Hart J, Hafler DA, Pericak-Vance M, Todd JA, Daly MJ, Trowsdale J, Wijmenga C, Vyse TJ, Beck S, Murray SS, Carrington M, Gregory S, Deloukas P, Rioux JD. A high-resolution HLA and SNP haplotype map for disease association studies in the extended human MHC. Nature genetics. 2006;38(10):1166–1172. doi: 10.1038/ng1885. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kazeem GR, Cox C, Aponte J, Messenheimer J, Brazell C, Nelsen AC, Nelson MR, Foot E. High-resolution HLA genotyping and severe cutaneous adverse reactions in lamotrigine-treated patients. Pharmacogenetics and genomics. 2009;19(9):661–665. doi: 10.1097/FPC.0b013e32832c347d. [DOI] [PubMed] [Google Scholar]
- Tassaneeyakul W, Jantararoungtong T, Chen P, Lin PY, Tiamkao S, Khunarkornsiri U, Chucherd P, Konyoung P, Vannaprasaht S, Choonhakarn C, Pisuttimarn P, Sangviroon A, Tassaneeyakul W. Strong association between HLA-B*5801 and allopurinol-induced Stevens-Johnson syndrome and toxic epidermal necrolysis in a Thai population. Pharmacogenetics and genomics. 2009;19(9):704–709. doi: 10.1097/FPC.0b013e328330a3b8. [DOI] [PubMed] [Google Scholar]
- Zhuang JJ, Zondervan K, Nyberg F, Harbron C, Jawaid A, Cardon LR, Barratt BJ, Morris AP. Optimizing the power of genome-wide association studies by using publicly available reference samples to expand the control group. Genetic epidemiology. 2010. Advance Online. [DOI] [PMC free article] [PubMed]
- Guan W, Liang L, Boehnke M, Abecasis GR. Genotype-based matching to correct for population stratification in large-scale case-control genetic association studies. Genetic epidemiology. 2009;33(6):508–517. doi: 10.1002/gepi.20403. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Luca D, Ringquist S, Klei L, Lee AB, Gieger C, Wichmann HE, Schreiber S, Krawczak M, Lu Y, Styche A, Devlin B, Roeder K, Trucco M. On the use of general control samples for genome-wide association studies: genetic matching highlights causal variants. American journal of human genetics. 2008;82(2):453–463. doi: 10.1016/j.ajhg.2007.11.003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cann HM, de Toma C, Cazes L, Legrand MF, Morel V, Piouffre L, Bodmer J, Bodmer WF, Bonne-Tamir B, Cambon-Thomsen A, Chen Z, Chu J, Carcassi C, Contu L, Du R, Excoffier L, Ferrara GB, Friedlaender JS, Groot H, Gurwitz D, Jenkins T, Herrera RJ, Huang X, Kidd J, Kidd KK, Langaney A, Lin AA, Mehdi SQ, Parham P, Piazza A, Pistillo MP, Qian Y, Shu Q, Xu J, Zhu S, Weber JL, Greely HT, Feldman MW, Thomas G, Dausset J, Cavalli-Sforza LL. A human genome diversity cell line panel. Science (New York, NY. 2002;296(5566):261–262. doi: 10.1126/science.296.5566.261b. [DOI] [PubMed] [Google Scholar]
- Strimmer K. fdrtool: a versatile R package for estimating local and tail area-based false discovery rates. Bioinformatics (Oxford, England) 2008;24(12):1461–1462. doi: 10.1093/bioinformatics/btn209. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Supporting material for manuscript "Genome-Wide Association study of Stevens-Johnson Syndrome and Toxic Epidermal Necrolysis in Europe" Results of association tests for the top SNP (Table S1) and for the most associated haplotype (Table S2) after stratification on the country of origin.