Abstract
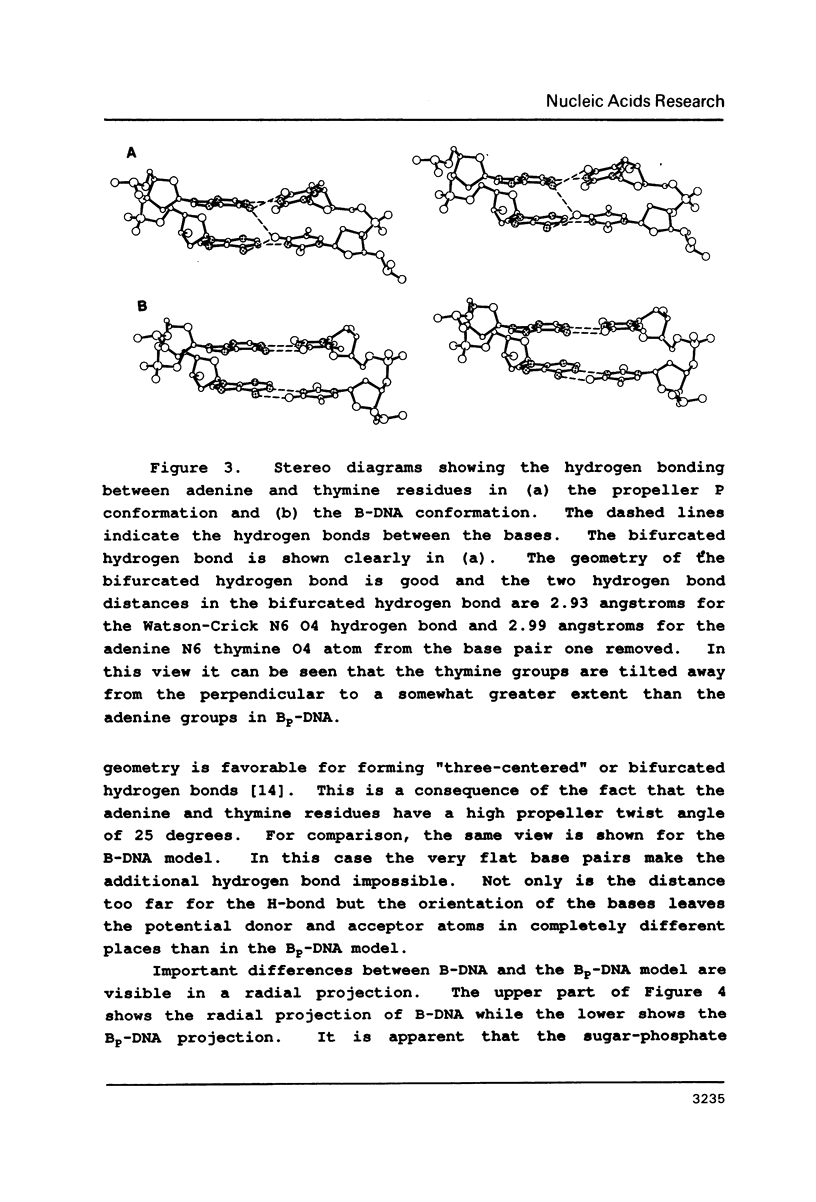
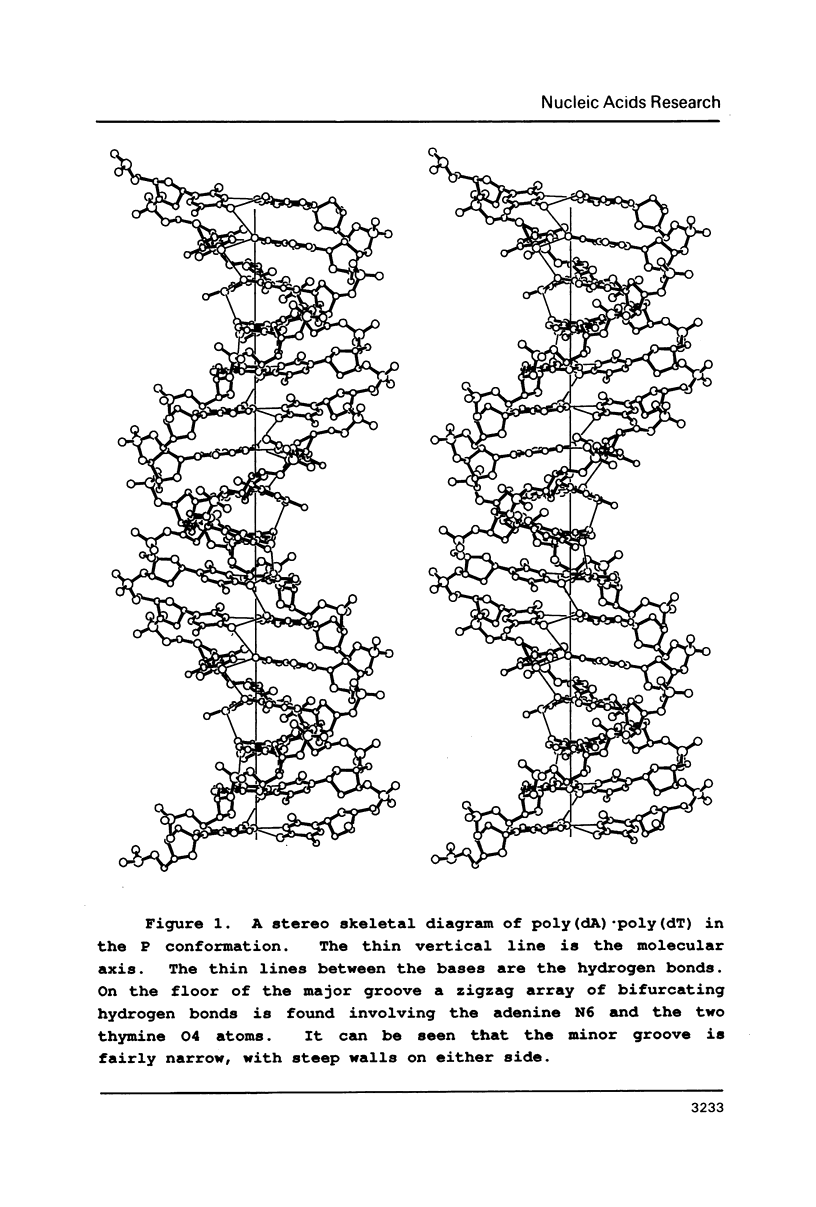
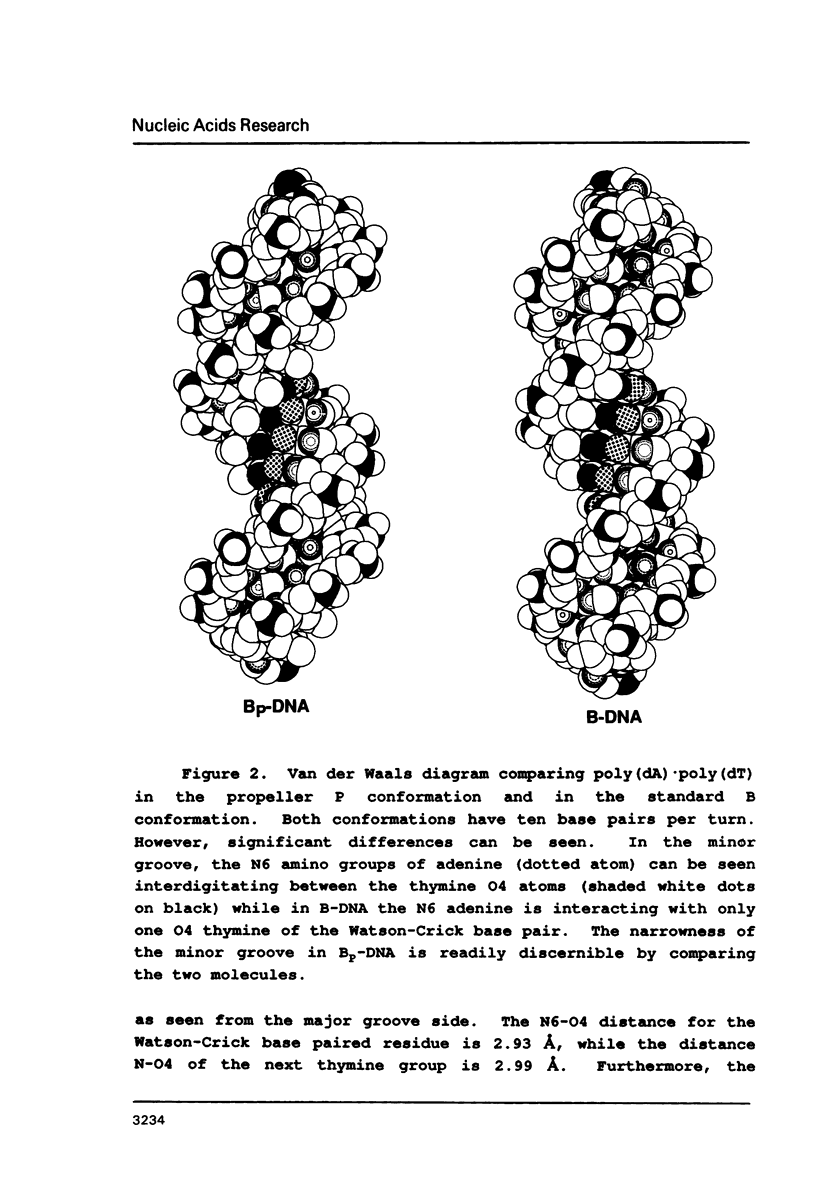
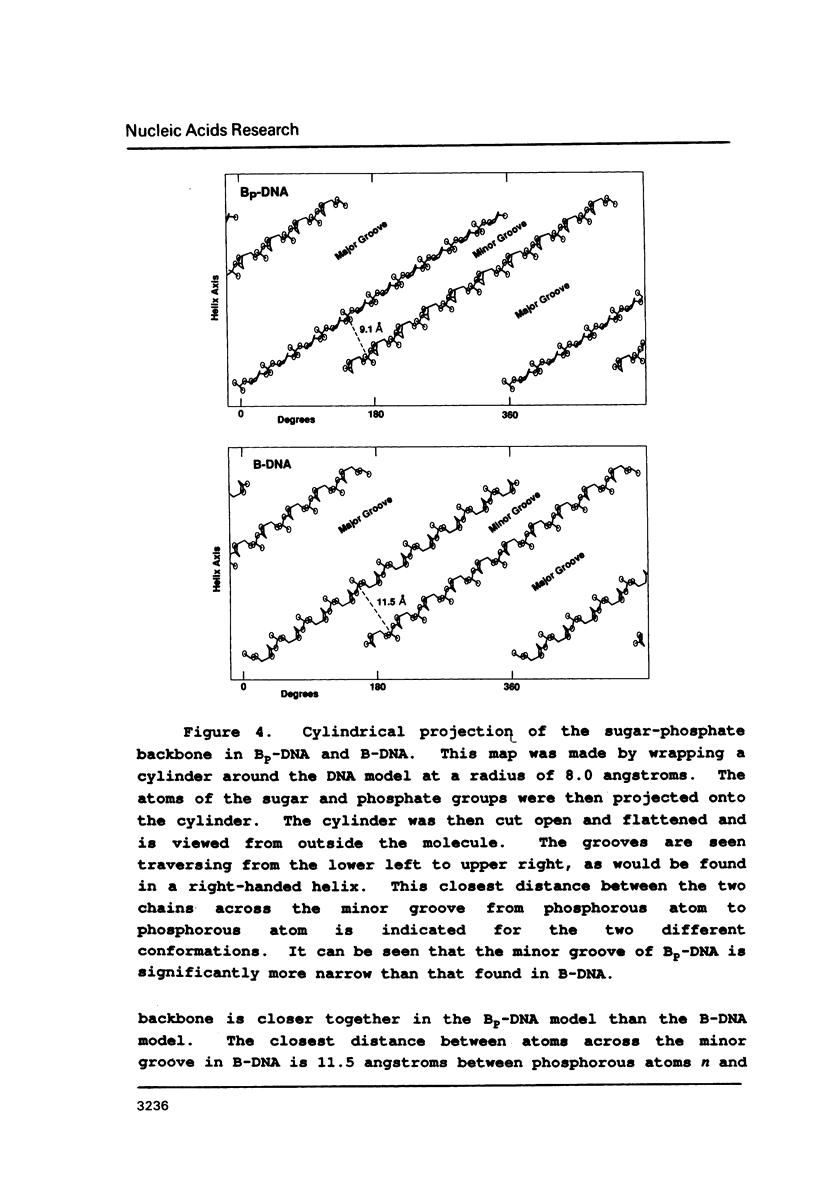
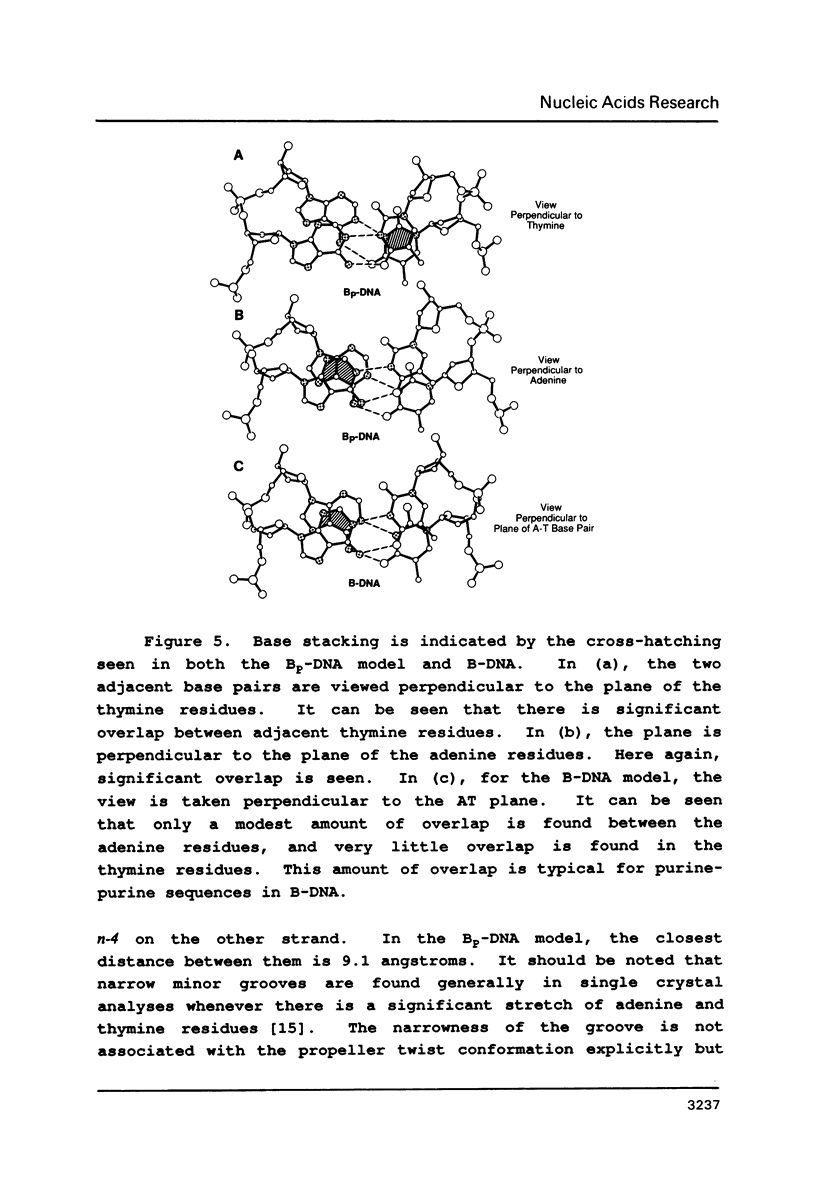
Physical properties of the DNA duplex, poly(dA).poly(dT) differ considerably from the alternating copolymer poly(dAT). A number of molecular models have been used to describe these structures obtained from fiber X-ray diffraction data. The recent solutions of single crystal DNA dodecamer structures with segments of oligo-A.oligo-T have revealed the presence of a high propeller twist in the AT regions which is stabilized by the formation of bifurcated (three-center) hydrogen bonds on the floor of the major groove, involving the N6 amino group of adenine hydrogen bonding to two O4 atoms of adjacent thymine residues on the opposite strand. Here we show that it is possible to incorporate the features of the single crystal analysis, specifically high propeller twist, bifurcated hydrogen bonds, and a narrow minor groove, as well as the close interstrand NMR signal between adenine HC2 and ribose HC1' of the opposite strand, into a model that is fully compatible with the diffraction data obtained from poly(dA).poly(dT).
Full text
PDF










Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Alexeev D. G., Lipanov A. A., Skuratovskii IYa Poly(dA).poly(dT) is a B-type double helix with a distinctively narrow minor groove. 1987 Feb 26-Mar 4Nature. 325(6107):821–823. doi: 10.1038/325821a0. [DOI] [PubMed] [Google Scholar]
- Arnott S., Chandrasekaran R., Hall I. H., Puigjaner L. C. Heteronomous DNA. Nucleic Acids Res. 1983 Jun 25;11(12):4141–4155. doi: 10.1093/nar/11.12.4141. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Arnott S., Selsing E. Structures for the polynucleotide complexes poly(dA) with poly (dT) and poly(dT) with poly(dA) with poly (dT). J Mol Biol. 1974 Sep 15;88(2):509–521. doi: 10.1016/0022-2836(74)90498-7. [DOI] [PubMed] [Google Scholar]
- Behling R. W., Kearns D. R. 1H two-dimensional nuclear Overhauser effect and relaxation studies of poly(dA).poly(dT) Biochemistry. 1986 Jun 3;25(11):3335–3346. doi: 10.1021/bi00359a037. [DOI] [PubMed] [Google Scholar]
- Coll M., Aymami J., van der Marel G. A., van Boom J. H., Rich A., Wang A. H. Molecular structure of the netropsin-d(CGCGATATCGCG) complex: DNA conformation in an alternating AT segment. Biochemistry. 1989 Jan 10;28(1):310–320. doi: 10.1021/bi00427a042. [DOI] [PubMed] [Google Scholar]
- Coll M., Frederick C. A., Wang A. H., Rich A. A bifurcated hydrogen-bonded conformation in the d(A.T) base pairs of the DNA dodecamer d(CGCAAATTTGCG) and its complex with distamycin. Proc Natl Acad Sci U S A. 1987 Dec;84(23):8385–8389. doi: 10.1073/pnas.84.23.8385. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Diekmann S., Wang J. C. On the sequence determinants and flexibility of the kinetoplast DNA fragment with abnormal gel electrophoretic mobilities. J Mol Biol. 1985 Nov 5;186(1):1–11. doi: 10.1016/0022-2836(85)90251-7. [DOI] [PubMed] [Google Scholar]
- Diekmann S., Zarling D. A. Unique poly(dA).poly(dT) B'-conformation in cellular and synthetic DNAs. Nucleic Acids Res. 1987 Aug 11;15(15):6063–6074. doi: 10.1093/nar/15.15.6063. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Drew H. R., Dickerson R. E. Structure of a B-DNA dodecamer. III. Geometry of hydration. J Mol Biol. 1981 Sep 25;151(3):535–556. doi: 10.1016/0022-2836(81)90009-7. [DOI] [PubMed] [Google Scholar]
- Fratini A. V., Kopka M. L., Drew H. R., Dickerson R. E. Reversible bending and helix geometry in a B-DNA dodecamer: CGCGAATTBrCGCG. J Biol Chem. 1982 Dec 25;257(24):14686–14707. [PubMed] [Google Scholar]
- Hagerman P. J. Sequence dependence of the curvature of DNA: a test of the phasing hypothesis. Biochemistry. 1985 Dec 3;24(25):7033–7037. doi: 10.1021/bi00346a001. [DOI] [PubMed] [Google Scholar]
- Kim S. H., Sussman J. L. pi turn is a conformational pattern in RNA loops and bends. Nature. 1976 Apr 15;260(5552):645–646. doi: 10.1038/260645a0. [DOI] [PubMed] [Google Scholar]
- Kintanar A., Klevit R. E., Reid B. R. Two-dimensional NMR investigation of a bent DNA fragment: assignment of the proton resonances and preliminary structure analysis. Nucleic Acids Res. 1987 Jul 24;15(14):5845–5862. doi: 10.1093/nar/15.14.5845. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kopka M. L., Yoon C., Goodsell D., Pjura P., Dickerson R. E. Binding of an antitumor drug to DNA, Netropsin and C-G-C-G-A-A-T-T-BrC-G-C-G. J Mol Biol. 1985 Jun 25;183(4):553–563. doi: 10.1016/0022-2836(85)90171-8. [DOI] [PubMed] [Google Scholar]
- Langridge R. Nucleic acids and polynucleotides. J Cell Physiol. 1969 Oct;74(2 Suppl):1–20. doi: 10.1002/jcp.1040740403. [DOI] [PubMed] [Google Scholar]
- Marini J. C., Levene S. D., Crothers D. M., Englund P. T. Bent helical structure in kinetoplast DNA. Proc Natl Acad Sci U S A. 1982 Dec;79(24):7664–7668. doi: 10.1073/pnas.79.24.7664. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nelson H. C., Finch J. T., Luisi B. F., Klug A. The structure of an oligo(dA).oligo(dT) tract and its biological implications. Nature. 1987 Nov 19;330(6145):221–226. doi: 10.1038/330221a0. [DOI] [PubMed] [Google Scholar]
- Peck L. J., Wang J. C. Sequence dependence of the helical repeat of DNA in solution. Nature. 1981 Jul 23;292(5821):375–378. doi: 10.1038/292375a0. [DOI] [PubMed] [Google Scholar]
- Rhodes D., Klug A. Sequence-dependent helical periodicity of DNA. Nature. 1981 Jul 23;292(5821):378–380. doi: 10.1038/292378a0. [DOI] [PubMed] [Google Scholar]
- Rhodes D. Nucleosome cores reconstituted from poly (dA-dT) and the octamer of histones. Nucleic Acids Res. 1979;6(5):1805–1816. doi: 10.1093/nar/6.5.1805. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sarma M. H., Gupta G., Sarma R. H. Untenability of the heteronomous DNA model for poly(dA).poly(dT) in solution. This DNA adopts a right-handed B-DNA duplex in which the two strands are conformationally equivalent. A 500 MHz NMR study using one dimensional NOE. J Biomol Struct Dyn. 1985 Jun;2(6):1057–1084. doi: 10.1080/07391102.1985.10507624. [DOI] [PubMed] [Google Scholar]
- Wing R., Drew H., Takano T., Broka C., Tanaka S., Itakura K., Dickerson R. E. Crystal structure analysis of a complete turn of B-DNA. Nature. 1980 Oct 23;287(5784):755–758. doi: 10.1038/287755a0. [DOI] [PubMed] [Google Scholar]
- Wu H. M., Crothers D. M. The locus of sequence-directed and protein-induced DNA bending. Nature. 1984 Apr 5;308(5959):509–513. doi: 10.1038/308509a0. [DOI] [PubMed] [Google Scholar]


