Abstract
As one of major epigenetic changes to inactivate tumor suppressor genes in human carcinogenesis, promoter hypermethylation was proposed as a marker to define novel tumor suppressor genes and predict the prognosis of cancer patients. In the present study, we found KL (klotho) as a novel tumor suppressor gene silenced through promoter hypermethylation in gastric cancer, the second leading cause of cancer death worldwide. KL expression was downregulated in primary gastric carcinoma tissues (n=22, p<0.05) and all of gastric cancer cells lines examined. Ectopic expression of KL inhibited the growth of gastric cancer cells partially through the induction of apoptosis, demonstrating a tumor suppressive role of KL in gastric cancer. Demethylation with 5-aza-2'-deoxycytidine (Aza) increased KL expression and KL promoter was hypermethylated in gastric cancer cell lines as well as some of primary gastric carcinoma tissues (47/99) but none of normal gastric tissues. Importantly, promoter methylation of KL was significantly associated with the poor outcome of gastric cancer patients (p=0.025, Log-rank test), highlighting the relevance of epigenetic inactivation of KL in gastric carcinogenesis. As a summary, we found that KL is a novel tumor suppressor gene epigenetically inactivated in gastric cancer and promoter methylation of KL could be used to predict the prognosis of gastric cancer patients.
Keywords: KL, gastric cancer, promoter hypermethylation
Introduction
In addition to genetic and epigenetic changes within tumor-initiating cells, the microenviron-ment plays important roles in cancer development. Tumors frequently developed from the chronic inflammation. The interaction between tumor-initiating cells and immune cells could eventually render the inflammatory microenvi-ronment permissive to tumor development [1-2]. Such tumor permissive microenvironments are enriched with various cytokines and growth factors such as insulin and insulin-like growth factor-1 (IGF-1). These bioactive soluble factors play important roles in the progression of tumors by activating multiple intracellular signaling pathways important to cellular proliferation and differentiation.
As a hormone similar in molecular structure to insulin, IGF-1 functions in a paracrine/autocrine fashion. Binding of IGF-1 to IGF-1R, a receptor tyrosine kinase, could initiate multiple intracellular signaling critical for cell growth and survival, such as PI3K/Akt signaling pathways [3]. By doing so, IGF-1 can promote cancer progression. The risk of cancer development is higher among people with raised concentrations of IGF-1. IGF-1 was thus proposed as a target for cancer treatment.
IGF-1 initiated signaling pathways are also subjected to intracellular or extracellular regulators. For example, IGF-1 can interact with IGF binding proteins (IGFBPs). IGFBPs are usually inhibitory to the function of IGF-1 since they bind IGF-1 at a higher affinity than it binds to its receptor. Therefore, increases of IGFBPs are often associated with the decrease of IGF-1 activity and reduced risk for cancer development [3-4]. In addition, IGFBPs were found to be inactivated through promoter hypermethylation in many human cancers including gastric cancer [5]. Recently, KL was found as an inhibitor of IGF-1 pathways [6-8], indicating that KL might be relevant to the cancer development by remodeling the interaction of tumor-initiating cells with the microenvironment. Indeed, KL was found to act as a tumor suppressor in human breast cancer and cervical carcinoma [9-10]. In the current study, we would like to understand the relevance of KL to human carcinogenesis.
Material and methods
Cell lines and tissue samples
All cell lines and primary tissues were obtained as previously described [11-12]. All cell lines were cultured in RPMI 1640 medium (Invitrogen, Carlsbad, CA, USA) supplemented with 10% fetal bovine serum and incubated at 5% CO2, 37°C and 95% humidity. For plasmid transfection, FuGENE 6 (Roche Applied Science, Mannheim, Germany) were used following the protocol provided.
Pharmacological demethylation
Cells were treated for 72 h with 5 μM 5-aza-2'-deoxycytidine (Aza) (Sigma, St Louis, MO, USA), a widely used methyltransferase inhibitor. Aza was replenished every 24 h. An equivalent concentration of the vehicle (DMSO) was used as the control. The cells were then harvested for total RNA and genomic DNA extraction.
Total RNA and genomic DNA extraction
Total RNA and genomic DNA were extracted using Trizol reagent (Invitrogen) following manufacturer's instruction. The concentrations of RNA and DNA were quantified by NanoDrop 1000 (Nanodrop, Wilmington, USA).
RT-PCR and quantitative real-time RT-PCR
Reverse transcription reaction was performed using 1 μg of total RNA with High Capacity cDNA Reverse Transcription kit (Applied Biosystems, Foster City, CA, USA). KL expression levels were determined by conventional RT-PCR with GoTaq polymerase (Promega, Madison, WI, USA) and quantitative real-time PCR using SYBR Green Master Mix Kit (Applied Biosystems). Human glyceraldehyde-3-phosphate dehydrogenase (hGAPDH) was used as an internal control of RNA integrity [11]. Primers used for KL were KL-F: 5'- ACCTGGTGGCGCACAACC and KL-R: 5'-TTGGCAAACCAACCTAGTACA.
Bisulfite treatment of DNA and methylation analysis
Methylation status of KL was determined by MSP (methylation specific PCR) and COBRA (combined bisulfite restriction analysis) using bisulfite modified genomic DNA as the template [12]. Genomic DNA was bisulphite-treated with Zymo DNA Modification Kit (Zymo Research, Orange, CA, USA) according to the protocol provided by the manufacturer. MSP was carried out for 40 cycles with annealing temperature at 62°C, as previously described [13]. For COBRA, Bstu I was used to digest the PCR products. Primers used for MSP were KL-MF: 5'-GTCGTCGTTGTAGTTCGTTATC and KL-MR: 5'-CAACAAACGCCGATAATAACG. Primers used for USP (unmethylation specific PCR) were KL-UF: 5'-TTGTTGTTGTTGTAGTTTGTTATT and KL-UR: 5'-CCAACAAACACCAATAATAAC. Primers used for COBRA were KL-BF: 5'-GGGTTTTTTTTAGGGTAT TTTTTT and KL-BR: 5'- GACAAATCCCAATA ATA-CAAAATA.
Western blot analysis
Proteins were resolved on 10-12.5% SDS-PAGE minigels and transferred onto Hybond C nitrocellulose membranes (Amersham Life Science, Buckinghamshire, UK). Membranes were blocked by 5% dry-milk containing TBS-T (Tris-Buffered Saline Tween-20) and probed with primary antibodies in blocking buffer overnight at 4°C. Finally, membranes were incubated with secondary antibodies conjugated with HRP (horse-radish peroxidase) and signals were visualized with enhanced chemiluminescence (Amersham Life Science). Membranes were reprobed with antibodies against b-actin (Cell Signaling Technology) as the loading control.
Cell growth assay
AGS and MKN28 cells transfected with empty vector or pcDNA3.1-KL kindly provided by Prof. Carmeda R. Abraham) were used for monolayer colony formation assay and growth analysis. Cells were cultured 24 h in a 12-well plate (1.0 × 105/well) and transfected with pcDNA3.1-KL or empty pcDNA3.1 using FuGENE 6 (Roche). After 48 h, the transfectants were re-plated in triplicate and cultured for 14-20 days in complete RPMI1640 medium containing G418 (500 μg/ ml). Surviving colonies were stained with Gentian Violet after methanol fixation and visible colonies (≥ 50 cells) were counted. The experiments were repeated three times. For growth analysis, 3-(4,5-dimethylthiazol-2-yl)-5-(3-carboxymethonyphenol)-2-(4-sulphophenyl)-2H-tetrazolium (MTS) assay was used. Cells transiently transfected with pcDNA3.1-KL or empty pcDNA3.1 were measured by a non-radioactive proliferation assay based on the ability of metabolically active cells to convert MTS into formazan using the CellTiter 96® AQueous Assay kit (Promega, Madison, WI, USA). The quantity of formazan was measured at 490 nm absorbance after one hour of incubation with CellTiter 96® AQueous One Solution Reagent following the instructions provided.
Flowcytometry analysis
Cells transfected with empty vector or KL-expressing vectors were stained with PI (propidium iodide) and FITC labeled Annexin-V and analyzed by Flow Cytometry (BD Biosciences, Franklin Lakes, NJ, USA). Cells negative for PI but positive for FITC are early apoptotic cells while cells positive for both PI and FITC are cells in late apoptosis or necrosis.
Statistical analysis
The difference of KL mRNA expression between tumor and adjacent non-tumor tissues was analyzed by the Wilcoxon matched pairs test. The χ2 tests were used for comparison of patient characteristics and distributions of methylation by vital status. The probability of overall survival was calculated with the Kaplan-Meier method and differences between curves were evaluated with the Log-rank test. Multivariate Cox proportional hazards models were also constructed to estimate the RRs (Relative risks) for various parameters including KL promoter methylation, with adjustments of other parameters such as gender, Lauren type, differentiation and TNM stage. All analyses were performed using SPSS for Windows, version 14.0, software. P-value < 0.05 was taken to define the statistical significance.
Results
KL is downregulated in gastric cancer
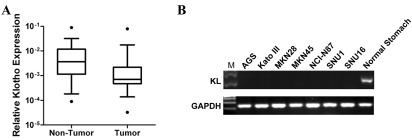
In order to know whether KL is relevant to gastric carcinogenesis, expression of KL in primary gastric carcinoma and adjacent non-tumor tissues were determined by real-time RT-PCR. KL was significantly downregulated in primary gastric carcinoma tissues (n=22, p<0.05, Wilcoxon matched pairs test) (Figure 1A). Consistently, KL expression in a panel of gastric carcinoma cell lines were also reduced (Figure 1B), indicating that KL might play as a tumor suppressor in gastric cancer.
Figure 1.
KL is downregulated in gastric cancer. (A) KL expression in primary gastric carcinoma tissues and adjacent non-tumor tissues were determined by real-time RT-PCR (n=22, p<0.05, Wilcoxon matched pairs test). GAPDH was used for the normalization. (B) KL expression in gastric cancer cell lines were determined by RT-PCR. GAPDH was used the loading control.
KL functions as a tumor suppressor in gastric cancer
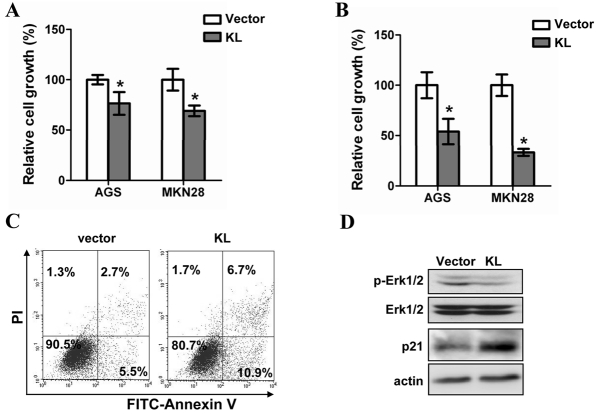
To confirm such assumption, we evaluated the growth of gastric cancer cell lines before and after restoration of KL expression. Both transient and stable KL transfection led to a significant growth inhibition of two independent gastric cancer cell lines (Figure 2A and B). Such a growth inhibition was accompanied by the suppression of Erk phosphorylation (Figure 2C) as well as the increase of apoptosis and p21 expression (Figure 2D)
Figure 2.
KL functions as a tumor suppressor in gastric cancer. (A) The growth of gastric cancer cell lines transiently transfected with pcDNA3.1-KL or empty vector were determined by MTS assay. The asterisks indicate statistical difference (p<0.05, Student's t test). (B) Colony formation assay was applied to analyze the long-term effect of KL expression on the growth of gastric cancer cells. The asterisks indicate statistical difference (p<0.05, Student's t test). (C) Cells stained with PI and FITC-Annexin-V were subjected to Flowcytometry analysis. (D) The amount of phosphory-lated Erk-1/2 and p21 were determined by western blotting. Pan-Erk-1/2 and GAPDH were used as the loading control respectively.
KL is epigenetic silenced in gastric cancer
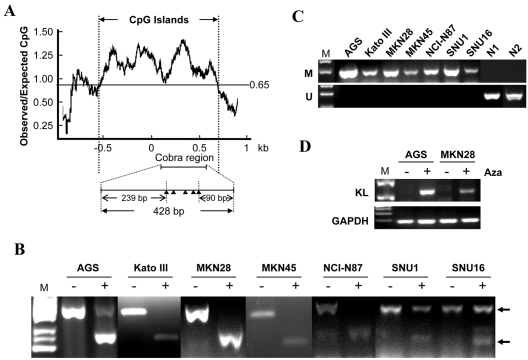
After confirming the tumor suppressor function of KL in gastric caner, we would like to know the mechanism responsible for the downregulation of KL in gastric cancer. A typical CpG island (CGI) was found in the promoter region of KL (Figure 3A), indicating that the expression of KL might be regulated by promoter hypermethylation. Indeed, both COBRA (combined bisulfite restriction analysis) and MSP (methylation specific PCR) revealed that KL promoter was hypermethylated in all gastric cancer cell lines (Figure 3B and C). To further confirm the relevance of promoter hypermethylation to the downregulation of KL in gastric cancer cells, KL expression in gastric cancer cells before and after pharmaceutical demethylation were determined by RT-PCR. KL expression was significantly upregulated after Aza treatment (Figure 3D), highlighting the contribution of promoter hypermethylation to KL downregulation.
Figure 3.
KL downregulation in gastric cancer cells was mediated by promoter hypermethylation. The schematic structure of the KL CGI was shown in (A) CGI was plotted by GeneTool program. Black triangles indicate BstU I sites. Promoter methylation of KL were analyzed by Cobra (B) and MSP (C), respectively. M: methylation specific PCR; U: un-methylation specific PCR. N1 and N2 are normal stomach tissues. (D) The expression of KL before and after Aza treatment were analyzed by RT-PCR. GAPDH was used as the loading control.
Promoter methylation of KL is associated with poor outcome of gastric cancer patients
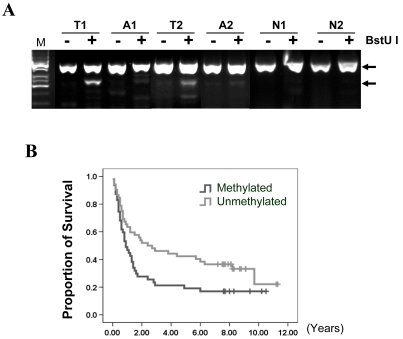
We further evaluated the methylation of KL promoter in primary gastric carcinoma tissues. Notably, methylation of KL promoter occurred in 46% (47/99) gastric cancer patients but not normal gastric tissues (0/10) (Figure 4A). The association of KL promoter hypermethylation with clinicopathologic characters of gastric cancer patients was summarized in Table 1. There was no significant correlation between the methylation of KL promoter and clinicopathologic features such as gender, Lauren type, differentiation, or pathologic stage. Interestingly, KL promoter hypermthylation seems to be associated with the survival status of gastric cancer patients. The percentage of KL promoter hypermethylation were 32% (8/25) in survived patients and 53% (39/74) in another group of patients (p=0.073, Chi-square test). We therefore analyzed the effect of KL promoter hypermthylation on survival of gastric cancer patients using Kaplan-Meier survival and Cox regression analyses. Indeed, KL promoter hypermethylation was significantly associated with the poor outcome of gastric cancer patients (p=0.025, Log-rank test) (Figure 4B). The multivariate Cox regression analysis indicated that KL promoter hypermethylation is an independent prognosis factor of gastric cancer patients (RR: 3.87, 95% CI: 1.27-11.77, p=0.017) (Table 2).
Figure 4.
Methylation of KL promoter was associated with poor outcome of gastric cancer patients. (A) Methylation of KL promoter were analyzed by Cobra as Figure 3B. T1 and T2 are tumor tissues; A1 and A2 are adjacent non-tumor tissues; N1 and N2 are normal stomach tissues. (B) Survival curves were plotted based on Kaplan-Meier survival analysis. Methylation status of KL promoter was used as the variate to separate two lines (n=99, p=0.025, Log-rank test).
Table 1.
Clinicopathologic features of KL methylation in gastric cancer
| Characteristics | Methylated |
Unmethylated |
p-value | ||
|---|---|---|---|---|---|
| n | % | n | % | ||
| Gender | |||||
| M | 24 | 43.6 | 31 | 56.4 | |
| F | 23 | 52.3 | 21 | 47.7 | 0.393 |
| H. pylori infection | |||||
| Positive | 16 | 50 | 16 | 50 | |
| Negative | 16 | 51.6 | 15 | 48.4 | 0.898 |
| Lauren | |||||
| Diffuse | 13 | 43.3 | 17 | 56.7 | |
| Intestinal | 19 | 47.5 | 21 | 52.5 | 0.579 |
| Differentiation | |||||
| Poor (or no differentiation) | 20 | 51.3 | 19 | 48.7 | |
| Well or moderate | 11 | 37.9 | 18 | 62.1 | 0.274 |
| TNM stage | |||||
| I | 7 | 46.7 | 8 | 53.3 | |
| II | 10 | 62.5 | 6 | 37.5 | 0.283 |
| III | 11 | 36.7 | 19 | 63.3 | |
| IV | 15 | 57.7 | 11 | 42.3 | |
Table 2.
Multivariate Cox regression analysis of potential prognostic factors for gastric cancer patients
| Variable | RR (95% CI) | p-value |
|---|---|---|
| Gender | ||
| Male | 1.00 | 0.762 |
| Female | 1.19(0.38-3.70) | |
| Lauren type | ||
| Diffuse | 2.32(0.58-9.31) | 0.234 |
| Intestinal | 1.00 | |
| Differentiation | ||
| Poor | 1.00 | 0.548 |
| Well or moderate | 0.66(0.17-2.57) | |
| TNM stage | ||
| I | 0.05(0.01-0.38) | 0.004 |
| II | 0.04(0.10-0.57) | <0.001 |
| III | 0.48(0.26-0.89) | 0.020 |
| IV | 1.00 | |
| KL | ||
| Methylated | 3.87 (1.27 - 11.77) | 0.017 |
| Unmethylated | 1.00 | |
Discussion
Gastric cancer remains the second leading cause of cancer death worldwide. The development of gastric cancer, similar to other cancers, is a multiple-stage process in which the accumulation of genetic and epigenetic changes finally leads to the clonal selection of variant progenies with the most aggressive growth properties [2, 14-16].
The epigenetic changes such as promoter DNA methylation can induce the inactivation of tumor suppressor genes that are important to prevent carcinogenesis. Promoter methylation was therefore proposed as a marker to identify new tumor suppressor genes [17-20]. We and others have previously found many cancer-related genes frequently silenced through promoter hypermethylation in gastric cancer [11, 21-26]. In addition, promoter methylation can facilitate the progression of cancers by conferring certain growth advantages to the affected cells. Thus, promoter methylation of some tumor suppressor genes can also be used as the prognosis factor to predict the outcome of cancer patients [11, 27].
In the current study, we found KL as a new tumor suppressor genes inactivated in gastric cancer through promoter hypermethylation. By inhibiting the activity of insulin/IGF-1 pathways, KL functions as a tumor suppressor in human breast cancer and cervical carcinoma [9-10]. In addition to its well-characterized function to inhibiting insulin/IGF-1 signaling, KL was found to act as the secreted Wnt antagonist [10]. Therefore, its function loss may contribute to aberrant activation of the canonical Wnt pathway in addition to insulin/IGF-1 signaling. Both pathways are important to the development and progression of many cancers including gastric cancer. In deed, estoration of KL expression in gastric cancer cells indeed suppressed cell growth and Erk phosphorylation, induced apoptosis and increased p21 expression (Figure 2).
In consistent with our finding, KL was found to be downregulated in cancers of bladder [28], breast [29-30],lung [31], kindey [32], and melanoma [33], mesothelioma [34]. Its downregulation was also associated with poor outcome of patients with breast cancer [35] and glioblastoma [35-36].
Similarly, other inhibitors of IGF-1 signaling such as IGFBPs were found to be inactivated through promoter hypermethylation in many cancers including gastric cancer [5]. Methylation of IGFBP3 and other IGFBPs correlated with cellular immortalization as well as drug resistance [37-38]. In addition, promoter methylation of IGFBPs were proposed to be a useful prognostic marker for disease progression and death in multiple cancers such as ovarian cancer, breast cancer, and lung cancer [39-41]. Interestingly, we found that methylation of KL promoter was also useful in the outcome prediction of gastric cancer patients. It would be important to know whether such a biomarker can be used to monitor the progression of precancerous lesion.
In summary, we identified KL as a new tumor suppressor gene epigenetically inactivated in gastric cancer. Promoter hypermthylation is an independent prognosis factor to predict the outcome of gastric cancer patients.
Acknowledgments
The project was supported by National Natural Science Foundation of China (81071963) to XW and Fundamental Research Funds for the Central Universities to HJ.
References
- 1.Tysnes BB, Bjerkvig R. Cancer initiation and progression: involvement of stem cells and the microenvironment. Biochim Biophys Acta. 2007;1775:283–297. doi: 10.1016/j.bbcan.2007.01.001. [DOI] [PubMed] [Google Scholar]
- 2.Wang X, Jin H. The epigenetic basis of Warburg effect. Epigenetics. 2010;5 doi: 10.4161/epi.5.7.12662. [DOI] [PubMed] [Google Scholar]
- 3.Samani AA, Yakar S, Le Roith D, Brodt P. The role of the IGF system in cancer growth and metastasis: overview and recent insights. Endocr Rev. 2007;28:20–47. doi: 10.1210/er.2006-0001. [DOI] [PubMed] [Google Scholar]
- 4.LeRoith D, Helman L. The new kid on the block(ade) of the IGF-1 receptor. Cancer Cell. 2004;5:201–202. doi: 10.1016/s1535-6108(04)00054-6. [DOI] [PubMed] [Google Scholar]
- 5.Tomii K, Tsukuda K, Toyooka S, Dote H, Hanafusa T, Asano H, Naitou M, Doihara H, Kisimoto T, Katayama H, Pass HI, Date H, Shimizu N. Aberrant promoter methylation of insulin-like growth factor binding protein-3 gene in human cancers. Int J Cancer. 2007;120:566–573. doi: 10.1002/ijc.22341. [DOI] [PubMed] [Google Scholar]
- 6.Kurosu H, Yamamoto M, Clark JD, Pastor JV, Nandi A, Gurnani P, McGuinness OP, Chikuda H, Yamaguchi M, Kawaguchi H, Shimomura I, Takayama Y, Herz J, Kahn CR, Rosenblatt KP, Kuro-o M. Suppression of aging in mice by the hormone Klotho. Science. 2005;309:1829–1833. doi: 10.1126/science.1112766. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Kuro-o M, Matsumura Y, Aizawa H, Kawaguchi H, Suga T, Utsugi T, Ohyama Y, Kurabayashi M, Kaname T, Kume E, Iwasaki H, Iida A, Shiraki-Iida T, Nishikawa S, Nagai R, Nabeshima YI. Mutation of the mouse klotho gene leads to a syndrome resembling ageing. Nature. 1997;390:45–51. doi: 10.1038/36285. [DOI] [PubMed] [Google Scholar]
- 8.Chen CD, Podvin S, Gillespie E, Leeman SE, Abraham CR. Insulin stimulates the cleavage and release of the extracellular domain of Klotho by ADAM10 and ADAM17. Proc Natl Acad Sci U S A. 2007;104:19796–19801. doi: 10.1073/pnas.0709805104. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Wolf I, Levanon-Cohen S, Bose S, Ligumsky H, Sredni B, Kanety H, Kuro-o M, Karlan B, Kaufman B, Koeffler HP, Rubinek T. Klotho: a tumor suppressor and a modulator of the IGF-1 and FGF pathways in human breast cancer. Oncogene. 2008;27:7094–7105. doi: 10.1038/onc.2008.292. [DOI] [PubMed] [Google Scholar]
- 10.Lee J, Jeong DJ, Kim J, Lee S, Park JH, Chang B, Jung SI, Yi L, Han Y, Yang Y, Kim KI, Lim JS, Yang I, Jeon S, Bae DH, Kim CJ, Lee MS. The anti-aging gene KLOTHO is a novel target for epigenetic silencing in human cervical carcinoma. Mol Cancer. 2010;9:109. doi: 10.1186/1476-4598-9-109. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Liu X, Wang X, Zhang J, Lam EK, Shin VY, Cheng AS, Yu J, Chan FK, Sung JJ, Jin HC. Warburg effect revisited: an epigenetic link between glycolysis and gastric carcinogenesis. Oncogene. 2010;29:442–450. doi: 10.1038/onc.2009.332. [DOI] [PubMed] [Google Scholar]
- 12.Liu X, Lam EK, Wang X, Zhang J, Cheng YY, Lam YW, Ng EK, Yu J, Chan FK, Jin H, Sung JJ. Promoter hypermethylation mediates down-regulation of thiamine receptor SLC19A3 in gastric cancer. Tumour Biol. 2009;30:242–248. doi: 10.1159/000243767. [DOI] [PubMed] [Google Scholar]
- 13.Wang X, Lau KK, So LK, Lam YW. CHD5 is down-regulated through promoter hypermethylation in gastric cancer. J Biomed Sci. 2009;16:95. doi: 10.1186/1423-0127-16-95. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Hanahan D, Weinberg RA. The hallmarks of cancer. Cell. 2000;100:57–70. doi: 10.1016/s0092-8674(00)81683-9. [DOI] [PubMed] [Google Scholar]
- 15.Ushijima T, Sasako M. Focus on gastric cancer. Cancer Cell. 2004;5:121–125. doi: 10.1016/s1535-6108(04)00033-9. [DOI] [PubMed] [Google Scholar]
- 16.Ponder BA. Cancer genetics. Nature. 2001;411:336–341. doi: 10.1038/35077207. [DOI] [PubMed] [Google Scholar]
- 17.Jones PA, Baylin SB. The fundamental role of epigenetic events in cancer. Nat Rev Genet. 2002;3:415–428. doi: 10.1038/nrg816. [DOI] [PubMed] [Google Scholar]
- 18.Esteller M. Epigenetics in cancer. N Engl J Med. 2008;358:1148–1159. doi: 10.1056/NEJMra072067. [DOI] [PubMed] [Google Scholar]
- 19.Chan SL, Cui Y, van Hasselt A, Li H, Srivastava G, Jin H, Ng KM, Wang Y, Lee KY, Tsao GS, Zhong S, Robertson KD, Rha SY, Chan AT, Tao Q. The tumor suppressor Wnt inhibitory factor 1 is frequently methylated in nasopharyngeal and esophageal carcinomas. Lab Invest. 2007;87:644–650. doi: 10.1038/labinvest.3700547. [DOI] [PubMed] [Google Scholar]
- 20.Cheng YY, Jin H, Liu X, Siu JM, Wong YP, Ng EK, Yu J, Leung WK, Sung JJ, Chan FK. Fibulin 1 is downregulated through promoter hypermethylation in gastric cancer. Br J Cancer. 2008;99:2083–2087. doi: 10.1038/sj.bjc.6604760. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Vogiatzi P, Vindigni C, Roviello F, Renieri A, Giordano A. Deciphering the underlying genetic and epigenetic events leading to gastric carcinogenesis. J Cell Physiol. 2007;211:287–295. doi: 10.1002/jcp.20982. [DOI] [PubMed] [Google Scholar]
- 22.Ushijima T. Epigenetic field for cancerization. J Biochem Mol Biol. 2007;40:142–150. doi: 10.5483/bmbrep.2007.40.2.142. [DOI] [PubMed] [Google Scholar]
- 23.Cheng YY, Jin H, Liu X, Siu JM, Wong YP, Ng EK, Yu J, Leung WK, Sung JJ, Chan FK. Fibulin 1 is downregulated through promoter hypermethylation in gastric cancer. Br J Cancer. 2008 doi: 10.1038/sj.bjc.6604760. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Jin H, Wang X, Ying J, Wong AH, Li H, Lee KY, Srivastava G, Chan AT, Yeo W, Ma BB, Putti TC, Lung ML, Shen ZY, Xu LY, Langford C, Tao Q. Epigenetic identification of ADAMTS18 as a novel 16q23.1 tumor suppressor frequently silenced in esophageal, nasopharyngeal and multiple other carcinomas. Oncogene. 2007;26:7490–7498. doi: 10.1038/sj.onc.1210559. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Wang LJ, Jin HC, Wang X, Lam EK, Zhang JB, Liu X, Chan FK, Si JM, Sung JJ. ZIC1 is downregulated through promoter hypermethylation in gastric cancer. Biochem Biophys Res Commun. 2009;379:959–963. doi: 10.1016/j.bbrc.2008.12.180. [DOI] [PubMed] [Google Scholar]
- 26.Yu J, Tao Q, Cheung KF, Jin H, Poon FF, Wang X, Li H, Cheng YY, Rocken C, Ebert MP, Chan AT, Sung JJ. Epigenetic identification of ubiq-uitin carboxyl-terminal hydrolase L1 as a functional tumor suppressor and biomarker for hepatocellular carcinoma and other digestive tumors. Hepatology. 2008;48:508–518. doi: 10.1002/hep.22343. [DOI] [PubMed] [Google Scholar]
- 27.Yu J, Cheng YY, Tao Q, Cheung KF, Lam CN, Geng H, Tian LW, Wong YP, Tong JH, Ying JM, Jin H, To KF, Chan FK, Sung JJ. Methylation of protocadherin 10, a novel tumor suppressor, is associated with poor prognosis in patients with gastric cancer. Gastroenterology. 2009;136:640–651. doi: 10.1053/j.gastro.2008.10.050. e641. [DOI] [PubMed] [Google Scholar]
- 28.Sanchez-Carbayo M, Socci ND, Lozano J, Saint F, Cordon-Cardo C. Defining molecular profiles of poor outcome in patients with invasive bladder cancer using oligonucleotide microarrays. J Clin Oncol. 2006;24:778–789. doi: 10.1200/JCO.2005.03.2375. [DOI] [PubMed] [Google Scholar]
- 29.Radvanyi L, Singh-Sandhu D, Gallichan S, Lovitt C, Pedyczak A, Mallo G, Gish K, Kwok K, Hanna W, Zubovits J, Armes J, Venter D, Hakimi J, Shortreed J, Donovan M, Parrington M, Dunn P, Oomen R, Tartaglia J, Berinstein NL. The gene associated with trichorhinophalangeal syndrome in humans is overexpressed in breast cancer. Proc Natl Acad Sci U S A. 2005;102:11005–11010. doi: 10.1073/pnas.0500904102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Karnoub AE, Dash AB, Vo AP, Sullivan A, Brooks MW, Bell GW, Richardson AL, Polyak K, Tubo R, Weinberg RA. Mesenchymal stem cells within tumour stroma promote breast cancer metastasis. Nature. 2007;449:557–563. doi: 10.1038/nature06188. [DOI] [PubMed] [Google Scholar]
- 31.Su LJ, Chang CW, Wu YC, Chen KC, Lin CJ, Liang SC, Lin CH, Whang-Peng J, Hsu SL, Chen CH, Huang CY. Selection of DDX5 as a novel internal control for Q-RT-PCR from microarray data using a block bootstrap re-sampling scheme. BMC Genomics. 2007;8:140. doi: 10.1186/1471-2164-8-140. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Gumz ML, Zou H, Kreinest PA, Childs AC, Belmonte LS, LeGrand SN, Wu KJ, Luxon BA, Sinha M, Parker AS, Sun LZ, Ahlquist DA, Wood CG, Copland JA. Secreted frizzled-related protein 1 loss contributes to tumor phenotype of clear cell renal cell carcinoma. Clin Cancer Res. 2007;13:4740–4749. doi: 10.1158/1078-0432.CCR-07-0143. [DOI] [PubMed] [Google Scholar]
- 33.Talantov D, Mazumder A, Yu JX, Briggs T, Jiang Y, Backus J, Atkins D, Wang Y. Novel genes associated with malignant melanoma but not benign melanocytic lesions. Clin Cancer Res. 2005;11:7234–7242. doi: 10.1158/1078-0432.CCR-05-0683. [DOI] [PubMed] [Google Scholar]
- 34.Gordon GJ, Rockwell GN, Jensen RV, Rheinwald JG, Glickman JN, Aronson JP, Pottorf BJ, Nitz MD, Richards WG, Sugarbaker DJ, Bueno R. Identification of novel candidate oncogenes and tumor suppressors in malignant pleural mesothelioma using large-scale transcriptional profiling. Am J Pathol. 2005;166:1827–1840. doi: 10.1016/S0002-9440(10)62492-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Pawitan Y, Bjohle J, Amler L, Borg AL, Egyhazi S, Hall P, Han X, Holmberg L, Huang F, Klaar S, Liu ET, Miller L, Nordgren H, Ploner A, Sandelin K, Shaw PM, Smeds J, Skoog L, Wedren S, Bergh J. Gene expression profiling spares early breast cancer patients from adjuvant therapy: derived and validated in two population-based cohorts. Breast Cancer Res. 2005;7:R953–964. doi: 10.1186/bcr1325. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Nutt CL, Mani DR, Betensky RA, Tamayo P, Cairncross JG, Ladd C, Pohl U, Hartmann C, McLaughlin ME, Batchelor TT, Black PM, von Deimling A, Pomeroy SL, Golub TR, Louis DN. Gene expression-based classification of malignant gliomas correlates better with survival than histological classification. Cancer Res. 2003;63:1602–1607. [PubMed] [Google Scholar]
- 37.Fridman AL, Rosati R, Li Q, Tainsky MA. Epigenetic and functional analysis of IGFBP3 and IGFBPrP1 in cellular immortalization. Biochem Biophys Res Commun. 2007;357:785–791. doi: 10.1016/j.bbrc.2007.04.019. [DOI] [PubMed] [Google Scholar]
- 38.Ibanez de Caceres I, Cortes-Sempere M, Moratilla C, Machado-Pinilla R, Rodriguez-Fanjul V, Manguan-Garcia C, Cejas P, Lopez-Rios F, Paz-Ares L, de CastroCarpeno J, Nistal M, Belda-Iniesta C, Perona R. IGFBP-3 hypermethylation-derived deficiency mediates cisplatin resistance in non-small-cell lung cancer. Oncogene. 2010;29:1681–1690. doi: 10.1038/onc.2009.454. [DOI] [PubMed] [Google Scholar]
- 39.Smith P, Nicholson LJ, Syed N, Payne A, Hiller L, Garrone O, Occelli M, Gasco M, Crook T. Epigenetic inactivation implies independent functions for insulin-like growth factor binding protein (IGFBP)-related protein 1 and the related IGFBPL1 in inhibiting breast cancer phenotypes. Clin Cancer Res. 2007;13:4061–4068. doi: 10.1158/1078-0432.CCR-06-3052. [DOI] [PubMed] [Google Scholar]
- 40.Wiley A, Katsaros D, Fracchioli S, Yu H. Methylation of the insulin-like growth factor binding protein-3 gene and prognosis of epithelial ovarian cancer. Int J Gynecol Cancer. 2006;16:210–218. doi: 10.1111/j.1525-1438.2006.00299.x. [DOI] [PubMed] [Google Scholar]
- 41.Chang YS, Wang L, Liu D, Mao L, Hong WK, Khuri FR, Lee HY. Correlation between insulin-like growth factor-binding protein-3 promoter methylation and prognosis of patients with stage I non-small cell lung cancer. Clin Cancer Res. 2002;8:3669–3675. [PubMed] [Google Scholar]