Abstract
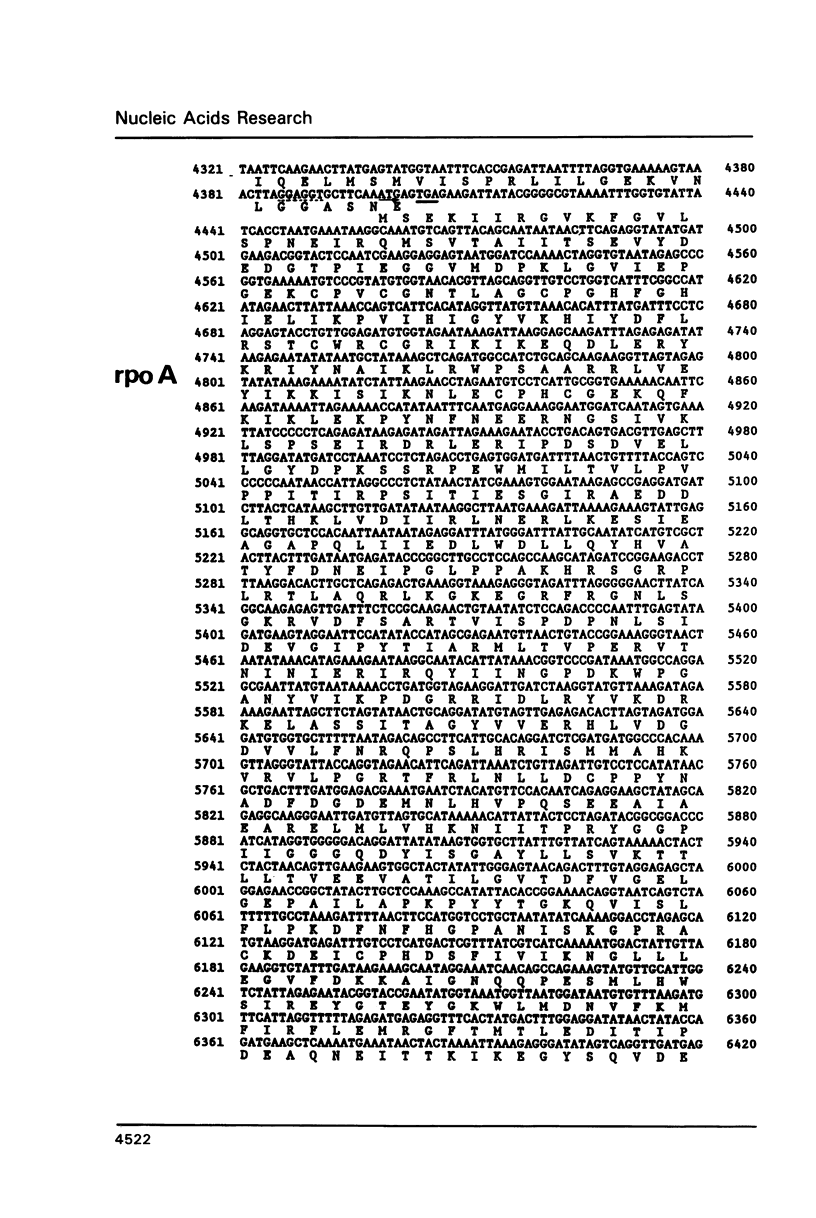
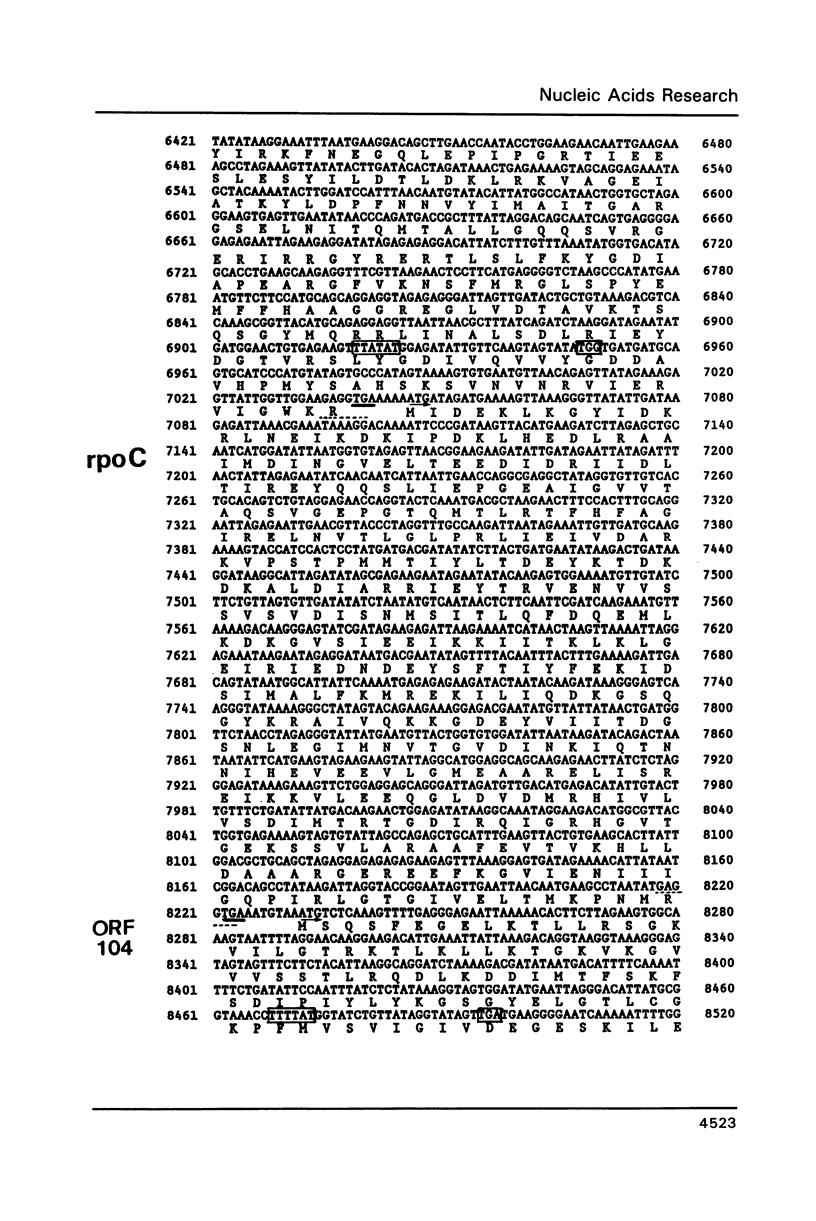
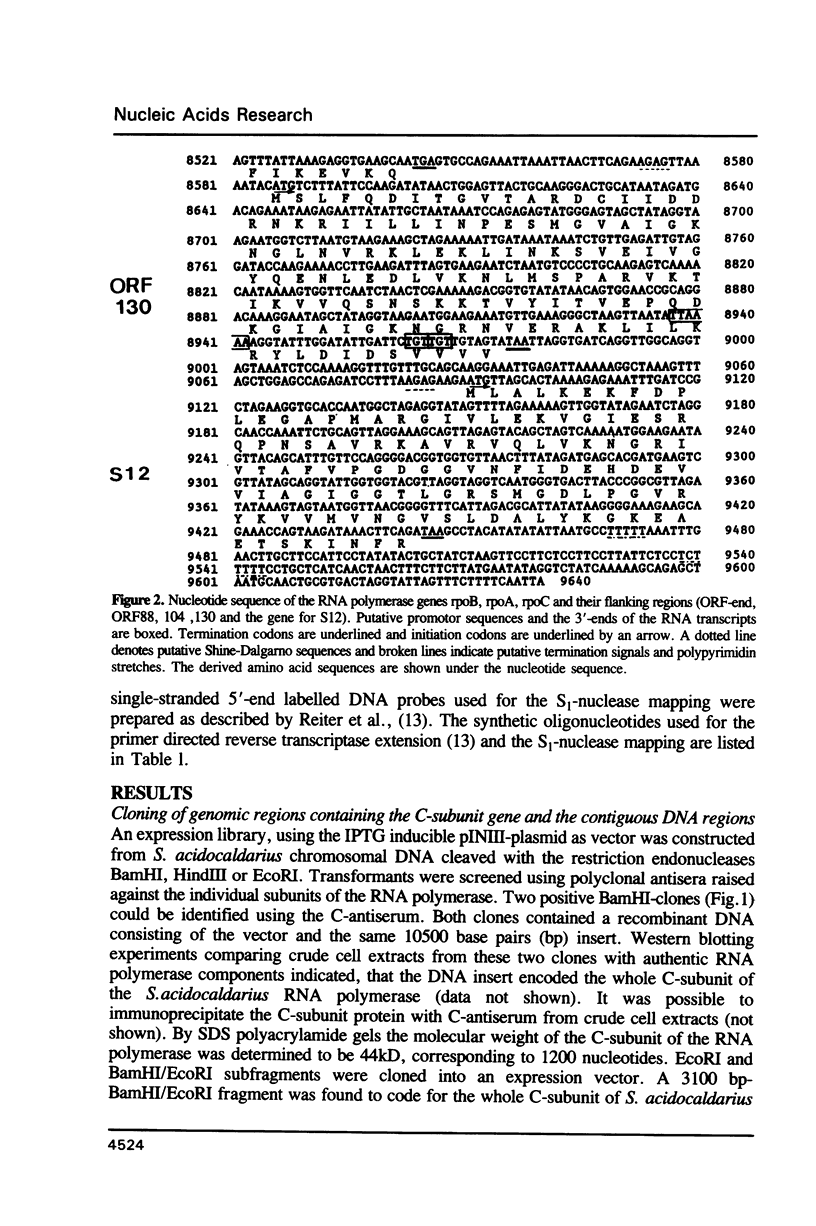
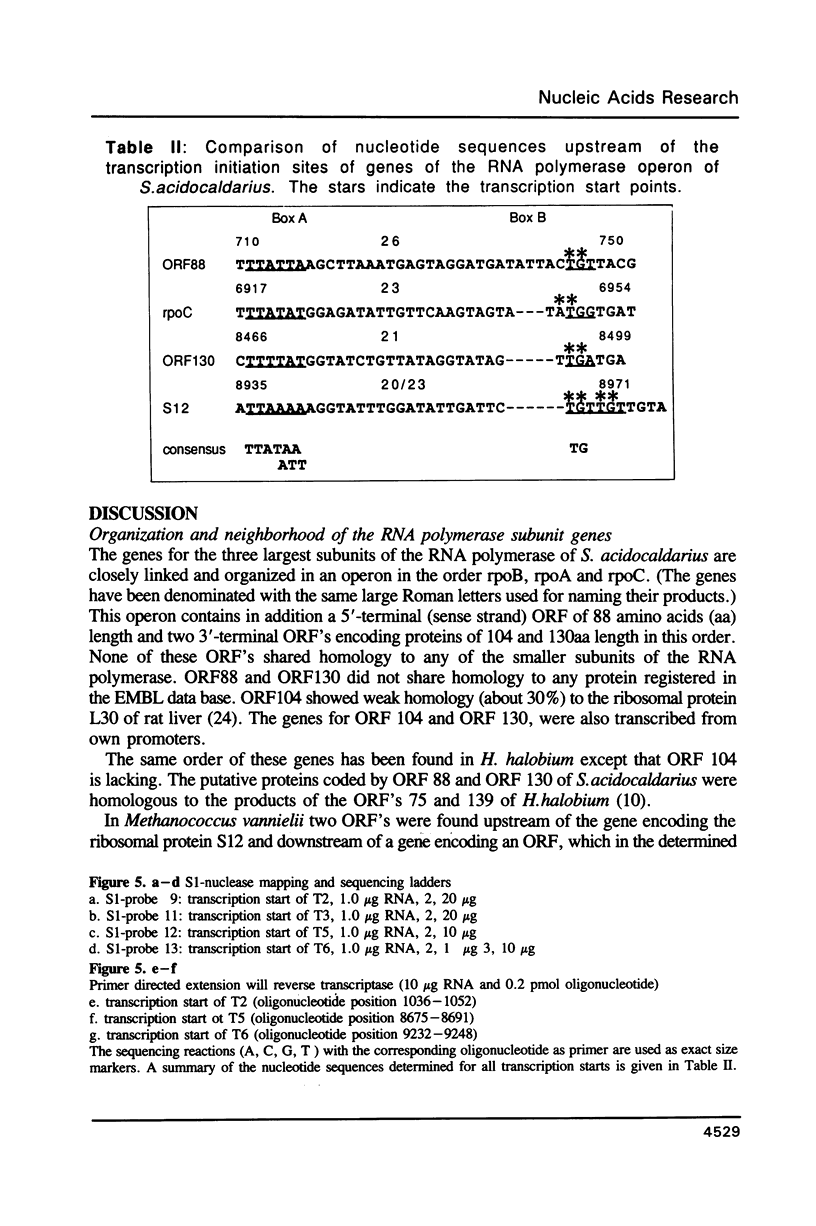
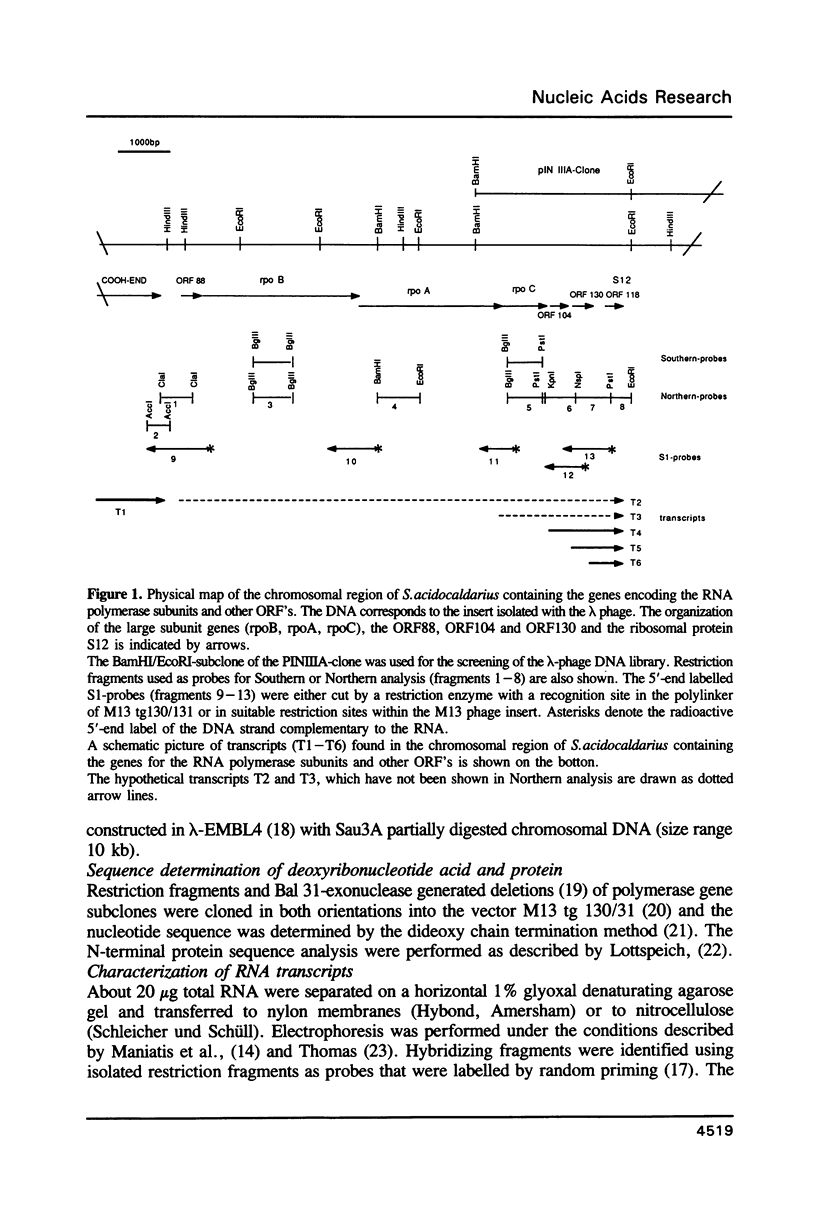
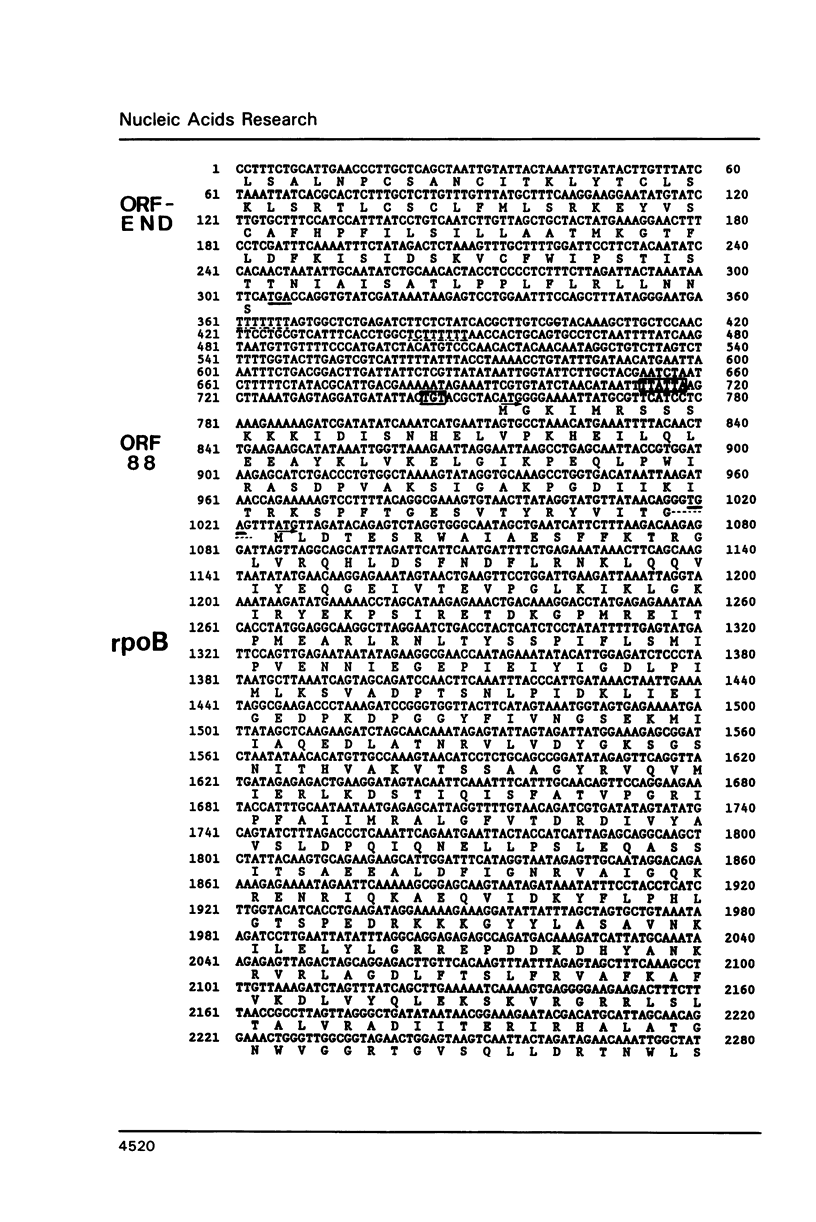
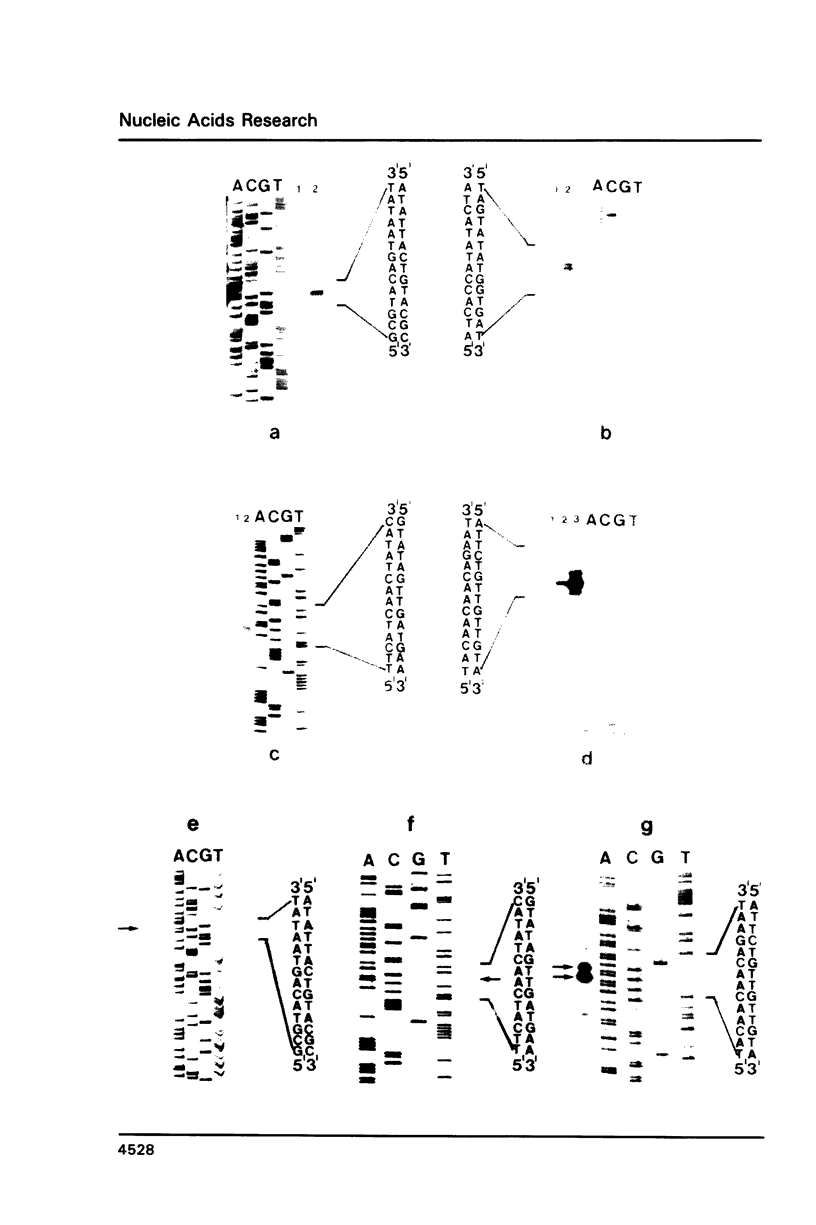
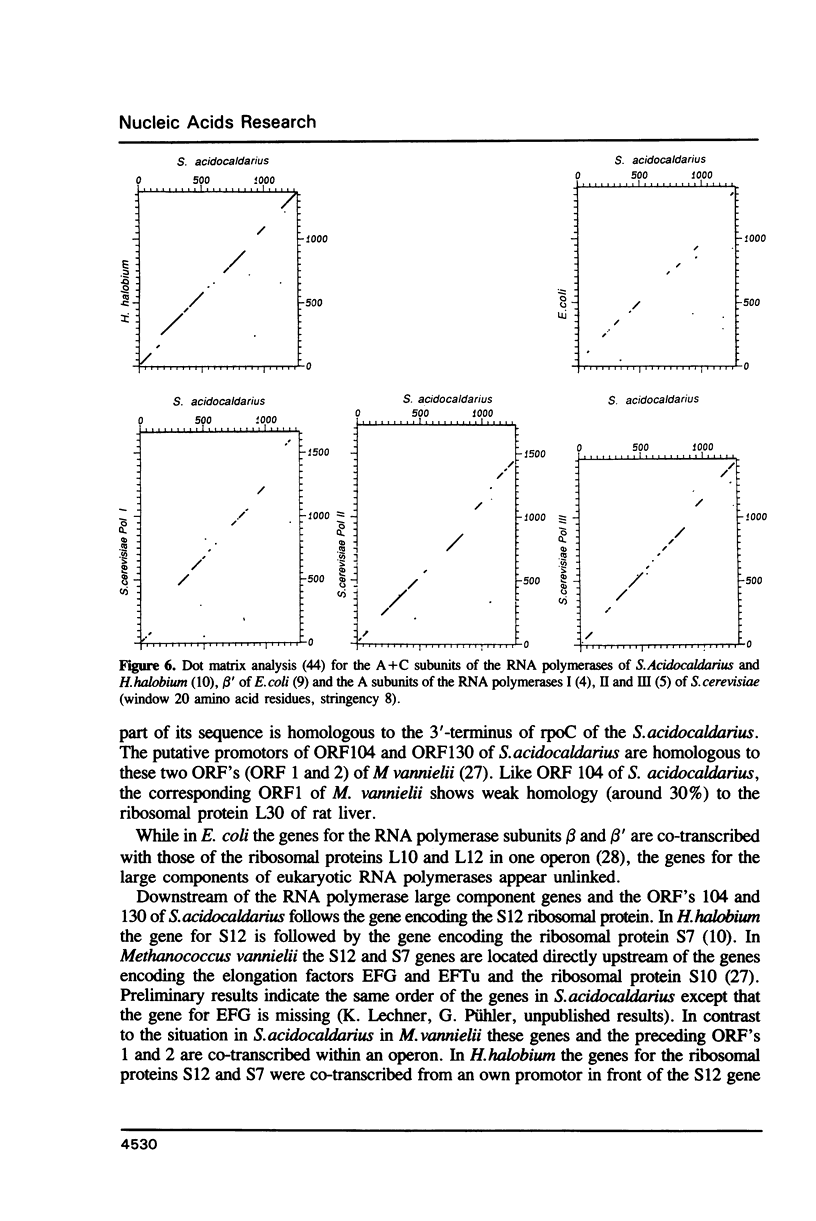
The genes for the three large subunits A, B and C, of the DNA-dependent RNA polymerase of the archaebacterium Sulfolobus acidocaldarius DSM 639, were identified and characterized. The three genes follow each other immediately in the order B-A-C, which corresponds to that found in the rpoBC operon of the Escherichia coli genome. The transcription products formed in vivo were studied by Northern analysis and the start-points were determined by S1-nuclease mapping and primer directed extension analysis. The three RNA polymerase subunit genes were co-transcribed together with an open reading frame (ORF) of 88 amino acid residues length situated immediately upstream of the B gene and two ORFs of 104 and 130 amino acid residues following the C gene (together 8500 nucleotides). The following ORF, encoding a protein of 118 amino acids homologous to the ribosomal protein S12 of E. coli, was weakly transcribed with the large co-transcript and strongly from an own promoter. The derived amino acid sequence of the B-subunit was found to be homologous to the B- (second largest) subunits of the eukaryotic nuclear polymerases I, II and III and to the eubacterial beta-subunit. The combined A + C-subunits correspond to the A- (largest) subunits of the eukaryotic RNA polymerases I, II and III and to the eubacterial beta'-subunit. The amino acid sequence similarity of the Sulfolobus subunits to the eukaryotic components is clearly higher than to the E. coli subunit.
Full text
PDF













Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Ahearn J. M., Jr, Bartolomei M. S., West M. L., Cisek L. J., Corden J. L. Cloning and sequence analysis of the mouse genomic locus encoding the largest subunit of RNA polymerase II. J Biol Chem. 1987 Aug 5;262(22):10695–10705. [PubMed] [Google Scholar]
- Allison L. A., Moyle M., Shales M., Ingles C. J. Extensive homology among the largest subunits of eukaryotic and prokaryotic RNA polymerases. Cell. 1985 Sep;42(2):599–610. doi: 10.1016/0092-8674(85)90117-5. [DOI] [PubMed] [Google Scholar]
- Berghöfer B., Kröckel L., Körtner C., Truss M., Schallenberg J., Klein A. Relatedness of archaebacterial RNA polymerase core subunits to their eubacterial and eukaryotic equivalents. Nucleic Acids Res. 1988 Aug 25;16(16):8113–8128. doi: 10.1093/nar/16.16.8113. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bucher P., Trifonov E. N. Compilation and analysis of eukaryotic POL II promoter sequences. Nucleic Acids Res. 1986 Dec 22;14(24):10009–10026. doi: 10.1093/nar/14.24.10009. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dassarma S., Rajbhandary U. L., Khorana H. G. Bacterio-opsin mRNA in wild-type and bacterio-opsin-deficient Halobacterium halobium strains. Proc Natl Acad Sci U S A. 1984 Jan;81(1):125–129. doi: 10.1073/pnas.81.1.125. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Devereux J., Haeberli P., Smithies O. A comprehensive set of sequence analysis programs for the VAX. Nucleic Acids Res. 1984 Jan 11;12(1 Pt 1):387–395. doi: 10.1093/nar/12.1part1.387. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Falkenburg D., Dworniczak B., Faust D. M., Bautz E. K. RNA polymerase II of Drosophila. Relation of its 140,000 Mr subunit to the beta subunit of Escherichia coli RNA polymerase. J Mol Biol. 1987 Jun 20;195(4):929–937. doi: 10.1016/0022-2836(87)90496-7. [DOI] [PubMed] [Google Scholar]
- Feinberg A. P., Vogelstein B. A technique for radiolabeling DNA restriction endonuclease fragments to high specific activity. Anal Biochem. 1983 Jul 1;132(1):6–13. doi: 10.1016/0003-2697(83)90418-9. [DOI] [PubMed] [Google Scholar]
- Frischauf A. M., Lehrach H., Poustka A., Murray N. Lambda replacement vectors carrying polylinker sequences. J Mol Biol. 1983 Nov 15;170(4):827–842. doi: 10.1016/s0022-2836(83)80190-9. [DOI] [PubMed] [Google Scholar]
- Glass R. E., Jones S. T., Nene V., Nomura T., Fujita N., Ishihama A. Genetic studies on the beta subunit of Escherichia coli RNA polymerase. VIII. Localisation of a region involved in promoter selectivity. Mol Gen Genet. 1986 Jun;203(3):487–491. doi: 10.1007/BF00422074. [DOI] [PubMed] [Google Scholar]
- Guo L. H., Yang R. C., Wu R. An improved strategy for rapid direct sequencing of both strands of long DNA molecules cloned in a plasmid. Nucleic Acids Res. 1983 Aug 25;11(16):5521–5540. doi: 10.1093/nar/11.16.5521. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hartmann G. R., Biebricher C., Glaser S. J., Grosse F., Katzameyer M. J., Lindner A. J., Mosig H., Nasheuer H. P., Rothman-Denes L. B., Schäffner A. R. Twelfth Fritz Lipmann lecture. Initiation of transcription--a general tool for affinity labeling of RNA polymerases by autocatalysis. Biol Chem Hoppe Seyler. 1988 Sep;369(9):775–788. [PubMed] [Google Scholar]
- Huet J., Schnabel R., Sentenac A., Zillig W. Archaebacteria and eukaryotes possess DNA-dependent RNA polymerases of a common type. EMBO J. 1983;2(8):1291–1294. doi: 10.1002/j.1460-2075.1983.tb01583.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kieny M. P., Lathe R., Lecocq J. P. New versatile cloning and sequencing vectors based on bacteriophage M13. Gene. 1983 Dec;26(1):91–99. doi: 10.1016/0378-1119(83)90039-2. [DOI] [PubMed] [Google Scholar]
- Kjems J., Garrett R. A. Novel expression of the ribosomal RNA genes in the extreme thermophile and archaebacterium Desulfurococcus mobilis. EMBO J. 1987 Nov;6(11):3521–3530. doi: 10.1002/j.1460-2075.1987.tb02678.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Leffers H., Gropp F., Lottspeich F., Zillig W., Garrett R. A. Sequence, organization, transcription and evolution of RNA polymerase subunit genes from the archaebacterial extreme halophiles Halobacterium halobium and Halococcus morrhuae. J Mol Biol. 1989 Mar 5;206(1):1–17. doi: 10.1016/0022-2836(89)90519-6. [DOI] [PubMed] [Google Scholar]
- Miller J., McLachlan A. D., Klug A. Repetitive zinc-binding domains in the protein transcription factor IIIA from Xenopus oocytes. EMBO J. 1985 Jun;4(6):1609–1614. doi: 10.1002/j.1460-2075.1985.tb03825.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mémet S., Gouy M., Marck C., Sentenac A., Buhler J. M. RPA190, the gene coding for the largest subunit of yeast RNA polymerase A. J Biol Chem. 1988 Feb 25;263(6):2830–2839. [PubMed] [Google Scholar]
- Nakanishi O., Oyanagi M., Kuwano Y., Tanaka T., Nakayama T., Mitsui H., Nabeshima Y., Ogata K. Molecular cloning and nucleotide sequences of cDNAs specific for rat liver ribosomal proteins S17 and L30. Gene. 1985;35(3):289–296. doi: 10.1016/0378-1119(85)90007-1. [DOI] [PubMed] [Google Scholar]
- Nomura M., Morgan E. A. Genetics of bacterial ribosomes. Annu Rev Genet. 1977;11:297–347. doi: 10.1146/annurev.ge.11.120177.001501. [DOI] [PubMed] [Google Scholar]
- Nomura T., Ishihama A., Kajitani M., Takahashi T., Nakada N., Yoshinaga K. Promoter selectivity of Escherichia coli RNA polymerase. II: Altered promoter selection by mutant holoenzymes. Mol Gen Genet. 1984;193(1):8–16. doi: 10.1007/BF00327407. [DOI] [PubMed] [Google Scholar]
- Ohama T., Yamao F., Muto A., Osawa S. Organization and codon usage of the streptomycin operon in Micrococcus luteus, a bacterium with a high genomic G + C content. J Bacteriol. 1987 Oct;169(10):4770–4777. doi: 10.1128/jb.169.10.4770-4777.1987. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Olsen G. J., Pace N. R., Nuell M., Kaine B. P., Gupta R., Woese C. R. Sequence of the 16S rRNA gene from the thermoacidophilic archaebacterium Sulfolobus solfataricus and its evolutionary implications. J Mol Evol. 1985;22(4):301–307. doi: 10.1007/BF02115685. [DOI] [PubMed] [Google Scholar]
- Ovchinnikov YuA, Monastyrskaya G. S., Gubanov V. V., Guryev S. O., Salomatina I. S., Shuvaeva T. M., Lipkin V. M., Sverdlov E. D. The primary structure of E. coli RNA polymerase, Nucleotide sequence of the rpoC gene and amino acid sequence of the beta'-subunit. Nucleic Acids Res. 1982 Jul 10;10(13):4035–4044. doi: 10.1093/nar/10.13.4035. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Panka D., Dennis D. RNA polymerase. Direct evidence for two active sites involved in transcription. J Biol Chem. 1985 Feb 10;260(3):1427–1431. [PubMed] [Google Scholar]
- Post L. E., Nomura M. DNA sequences from the str operon of Escherichia coli. J Biol Chem. 1980 May 25;255(10):4660–4666. [PubMed] [Google Scholar]
- Reiter W. D., Palm P., Zillig W. Analysis of transcription in the archaebacterium Sulfolobus indicates that archaebacterial promoters are homologous to eukaryotic pol II promoters. Nucleic Acids Res. 1988 Jan 11;16(1):1–19. doi: 10.1093/nar/16.1.1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Reiter W. D., Palm P., Zillig W. Transcription termination in the archaebacterium Sulfolobus: signal structures and linkage to transcription initiation. Nucleic Acids Res. 1988 Mar 25;16(6):2445–2459. doi: 10.1093/nar/16.6.2445. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sanger F., Nicklen S., Coulson A. R. DNA sequencing with chain-terminating inhibitors. Proc Natl Acad Sci U S A. 1977 Dec;74(12):5463–5467. doi: 10.1073/pnas.74.12.5463. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schnabel R., Thomm M., Gerardy-Schahn R., Zillig W., Stetter K. O., Huet J. Structural homology between different archaebacterial DNA-dependent RNA polymerases analyzed by immunological comparison of their components. EMBO J. 1983;2(5):751–755. doi: 10.1002/j.1460-2075.1983.tb01495.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shine J., Dalgarno L. The 3'-terminal sequence of Escherichia coli 16S ribosomal RNA: complementarity to nonsense triplets and ribosome binding sites. Proc Natl Acad Sci U S A. 1974 Apr;71(4):1342–1346. doi: 10.1073/pnas.71.4.1342. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sweetser D., Nonet M., Young R. A. Prokaryotic and eukaryotic RNA polymerases have homologous core subunits. Proc Natl Acad Sci U S A. 1987 Mar;84(5):1192–1196. doi: 10.1073/pnas.84.5.1192. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Thomas P. S. Hybridization of denatured RNA transferred or dotted nitrocellulose paper. Methods Enzymol. 1983;100:255–266. doi: 10.1016/0076-6879(83)00060-9. [DOI] [PubMed] [Google Scholar]
- Thomm M., Wich G. An archaebacterial promoter element for stable RNA genes with homology to the TATA box of higher eukaryotes. Nucleic Acids Res. 1988 Jan 11;16(1):151–163. doi: 10.1093/nar/16.1.151. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wich G., Hummel H., Jarsch M., Bär U., Böck A. Transcription signals for stable RNA genes in Methanococcus. Nucleic Acids Res. 1986 Mar 25;14(6):2459–2479. doi: 10.1093/nar/14.6.2459. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yeats S., McWilliam P., Zillig W. A plasmid in the archaebacterium Sulfolobus acidocaldarius. EMBO J. 1982;1(9):1035–1038. doi: 10.1002/j.1460-2075.1982.tb01292.x. [DOI] [PMC free article] [PubMed] [Google Scholar]