Abstract
Chronological gene expression patterns of biofilm-forming cells are important to understand bioactivity and pathogenicity of biofilms. For Porphyromonas gingivalis ATCC 33277 biofilm formation, the number of genes differentially regulated by more than 1.5-fold was highest during the growth stage (312/2,090 genes), and some pathogen-associated genes were time-dependently controlled.
TEXT
Oral biofilms contain multiple bacterial species and cause opportunistic infections like dental caries and periodontal disease (3, 5). Porphyromonas gingivalis, a Gram-negative oral anaerobe, is distributed throughout oral biofilms and is one of the major pathogens in severe forms of marginal and refractory periapical periodontitis (14, 16). Global gene analysis using DNA microarray was performed on the planktonic cells of P. gingivalis ATCC 33277, and the total number of genes was reported to be 2,090 (11). Lo et al. (10) compared the global gene expression in a P. gingivalis strain W50 biofilm after 40 days of incubation with that of its planktonic counterparts grown in the same continuous culture relative to the gene expression data from strain W83. They found that in biofilm cells, genes involved in growth and metabolic activity were downregulated. However, time course gene expression changes during P. gingivalis biofilm formation have never been studied.
Biofilm formation is a dynamic and sequential process involving attachment, maturation, and detachment (17). It is important, therefore, to understand the changes in time course gene expression during biofilm growth. Recent microarray analyses of biofilms revealed that hundreds of genes, including many uncharacterized genes, are differentially expressed in biofilms; if fully characterized, they might provide insights into the genetic basis for biofilm formation (13). P. gingivalis biofilm 40 to 80 μm thick was formed on disks. A pump was used for 4 to 40 days to perfuse cell culture medium into the flow cell model (1, 4, 10, 15). Because it is possible to sequentially collect biofilm samples from initial adhesion to maturation, this model was chosen for the present study. The novel flow cell model is also valuable because of its high reproducibility and its ability to recover more biofilm-forming cells than the modified Robbins device (MRD) model (15).
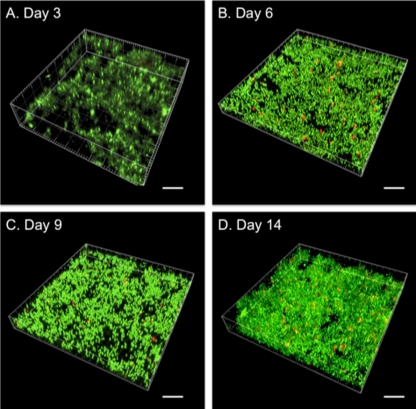
The conditions used for the biofilm cell culture of P. gingivalis ATCC 33277 are described in the supplemental material. The biofilms were allowed to form for 14 days on hydroxyapatite (HA) powders using the flow cell model (see Fig. S1 in the supplemental material). The measurement of optical density at 550 nm (OD550) and the confocal laser scanning microscopic (CLSM) observations were performed at 3, 6, 9 and 14 days. The OD550 value was lowest at day 3 and then increased sequentially to day 14 (Fig. 1). Individual P. gingivalis cells sparsely adhered to the disks at day 3 (Fig. 2 A); between days 6 and 14, three-dimensional biofilm growth and a few red-stained dead cells were seen (Fig. 2B, C, and D). The volume of the biofilm was largest at day 14. Combining the images taken during biofilm growth (Fig. 2) with the OD values (Fig. 1) showed that the density of the biofilm-forming cells increased from days 3 to 6 and again from days 9 to 14 (Fig. 2).
Fig. 1.
P. gingivalis biofilm growth rate. Growth of P. gingivalis biofilm was analyzed quantitatively using the optical density method. The growth rate was not constant.
Fig. 2.
CLSM images of P. gingivalis biofilm. P. gingivalis biofilm images at 3 (A), 6 (B), 9 (C), and 14 (D) days. Scale bar, 10 μm. Live and dead cells are stained with green and red, respectively. The proportion of live to dead cells was 98.67% ± 0.10% to 1.33% ± 0.10% (A), 97.78% ± 0.40% to 2.22% ± 0.40% (B), 98.22% ± 0.17% to 1.78% ± 0.17% (C), and 98.75% ± 0.19% to 1.25% ± 0.19% (D).
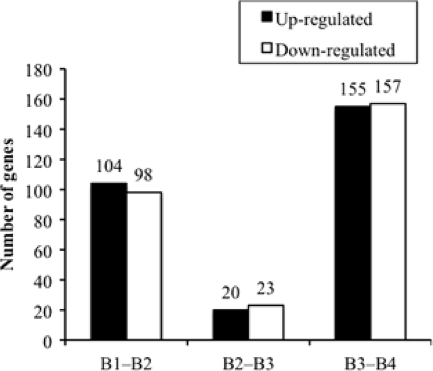
The HA powders were rinsed and ultrasonicated for 30 min at 4°C in 20 ml of distilled water to release biofilm cells from the powders into the liquid. After isolation from the suspension of biofilm cells, the RNA was analyzed using the microarray (see supplemental material for details) and the gene expression data were normalized, generated, and assessed using a previously described method (see supplemental materials). The microarray data were deposited in the Center for Information Biology Gene Expression Database (CIBEX). The RNA samples harvested from the biofilm-forming cells after 3, 6, 9, and 14 days were named B1, B2, B3, and B4, respectively. The numbers of differentially expressed genes (DEGs) that were differentially regulated by more than 1.5-fold between each time point (P < 0.01, Welch t test) are shown in Fig. 3. The number of DEGs was highest between 9 days (B3) and 14 days (B4), when 155 genes were upregulated and 157 genes were downregulated. It is likely that the processes of biofilm maturation and of extracellular matrix formation are the processes that need changes in gene expression most.
Fig. 3.
Time-dependent changes for the numbers of differentially expressed genes of P. gingivalis biofilm-forming cells. The numbers of differentially expressed genes significantly upregulated (black bar) and downregulated (white bar) by more than 1.5-fold (P < 0.001, Welch t test) are shown.
The 52 genes that are known to be associated with the five virulence factors of P. gingivalis were selected, and the fold change data for each of them are shown in Table 1 (see also the supplemental material). Because the deletion of the gingipain-related genes kgp and/or rgpA rgpB promotes biofilm formation (6, 8), they might be predicted to be downregulated during biofilm growth. However, in this study, the expression of gingipain-related genes was for the most part, constant; an exception was a subtle upregulation of PGN_0023 and rgpB, but this result was not significant (P < 0.01) (Table 1). An rgpA rgpB double mutant of P. gingivalis ATCC 33277 possessed very few fimbriae on the cell surface (7, 12). Although the deletion of both rgpA and rgpB had an impact on biofilm formation caused by defects in the fimbriae, the growth stage of the biofilm had no effect on the gene expression of the gingipain-related genes. During biofilm growth, fimA was upregulated between B1 and B2 and significantly downregulated in the following 3 days. A previous report found that the FimA fimbriae promoted initial biofilm formation but exerted a restraining regulation on biofilm maturation (8). These results suggested that in the early stage of biofilm formation the upregulation of fimA may encourage FimA fimbria production, thereby accelerating the attachment of the cells to the surface; suppression of fimA, on the other hand, may function as a switch for biofilm maturation. Deletion of clpXP, one of the members of the heat shock protein family, caused overexpression of mfa1 and accelerated biofilm formation (2), and while the wild-type strain formed microcolonies, an mfa1 mutant did not (9). Our results are consistent with those of the earlier studies and support the speculation that mfa1 acts from the B1 stage to the B2 stage. Lipopolysaccharide (LPS)-related PGN_2019 was significantly upregulated between B1 and B2, and seven genes were significantly downregulated between B3 and B4. It is likely that, at the later stages of biofilm formation, the downregulation of the fimbria- and LPS-related genes might be associated with the detachment of cells.
Table 1.
Fold change values for the differentially expressed genes associated with the virulence factors of P. gingivalis
| Virulence factor | Locus tag | Fold changesa |
Gene name | Product nameb | ||
|---|---|---|---|---|---|---|
| B1–B2c | B2–B3c | B3–B4c | ||||
| Gingipain | PGN_0023 | 1.54 | −1.68 | 1.56 | Hypothetical protein | |
| PGN_0295 | −1.32 | 1.41 | 1.05 | C-terminal domain of Arg- and Lys-gingipainproteinase | ||
| PGN_0778 | −1.20 | −1.24 | 1.45 | porT | Membrane-associated protein PorT | |
| PGN_1466 | 1.52 | −1.26 | 1.30* | rgpB | Arginine-specific cysteine proteinase RgpB | |
| PGN_1728 | −1.12 | 1.09 | 1.03 | kgp | Lysine-specific cysteine proteinase Kgp | |
| PGN_1768 | 1.25 | −1.11 | 1.09 | Putative DNA-binding response regulator/sensorhistidine kinase | ||
| PGN_1970 | 1.08 | −1.05 | 1.15 | rgpA | Arginine-specific cysteine proteinase RgpA | |
| PGN_2065 | 1.14 | −1.20 | 1.14 | Putative Lys- and Rgp-gingipain domain protein | ||
| Hemagglutinin | PGN_0435 | −1.24 | 1.03 | 1.27 | Probable partial hemagglutinin-related protein | |
| PGN_0436 | 1.03 | 1.16 | −1.20 | Probable partial hemagglutinin-related protein | ||
| PGN_0561 | 1.21 | 1.01 | −1.03 | prtT | Trypsinlike proteinase PrtT | |
| PGN_0900 | 1.00 | 1.20 | 1.02 | Thiol protease | ||
| PGN_1115 | 1.13 | −1.08 | 1.02 | Putative hemagglutinin | ||
| PGN_1519 | 1.03 | −1.18 | 1.17 | Hemagglutinin-related protein | ||
| PGN_1556 | −1.24 | 1.06 | −1.11 | Putative hemagglutinin | ||
| PGN_1733 | −1.23* | −1.06 | 1.13 | hagA | Hemagglutinin protein HagA | |
| PGN_1904 | 1.28 | 1.06 | −1.21 | hagB | Hemagglutinin protein HagB | |
| PGN_1906 | 1.28* | 1.04 | −1.17 | hagC | Hemagglutinin protein HagC | |
| PGN_2024 | −1.11 | −1.05 | 1.09 | Putative hemagglutinin | ||
| Fimbriae | PGN_0180 | 1.89* | −1.80* | 1.39 | fimA | FimA type I fimbrilin |
| PGN_0183 | 1.60* | −1.54* | 1.03 | fimC | Minor component FimC | |
| PGN_0184 | −1.20 | 1.01 | 1.40 | fimD | Minor component FimD | |
| PGN_0185 | −1.38* | 1.11 | 1.36 | fimE | Minor component FimE | |
| PGN_0287 | 1.62* | −1.21 | 1.13 | mfa1 | Mfa1 fimbrilin | |
| PGN_0288 | 1.68* | −1.13 | −1.22* | Hypothetical protein | ||
| LPS | PGN_0206 | 1.26 | −1.33 | 1.47* | Putative lipid A disaccharide synthase | |
| PGN_0376 | 1.00 | −1.06 | 1.33* | 2-Dehydro-3-deoxyphosphooctonate aldolase | ||
| PGN_0524 | −1.21 | 1.21 | −1.09 | Hypothetical protein | ||
| PGN_0544 | 1.07 | −1.14 | 1.43* | 3-Deoxy-d-manno-octulosonic acid transferase | ||
| PGN_0679 | 1.42 | −1.32 | −1.08 | Putative tetraacyldisaccharide 4′kinase | ||
| PGN_0696 | 1.10 | −1.03 | 1.28* | Probable hydrolase | ||
| PGN_0777 | 1.27 | −1.57* | 1.32 | Probable glycosyl transferase | ||
| PGN_1054 | −1.20 | −1.08 | 1.82* | vimF | Virulence-modulating gene F | |
| PGN_1134 | −1.08 | −1.10 | 1.38 | Hypothetical protein | ||
| PGN_1235 | −2.19* | 1.83* | −1.37 | porS | Membrane protein PorS | |
| PGN_1239 | −1.51 | 1.10 | −1.10 | Probable lipopolysaccharide biosynthesisglycosyltransferase | ||
| PGN_1240 | −1.12 | 1.37 | −1.24 | Hypothetical protein | ||
| PGN_1251 | −1.48 | 1.28 | 1.03 | Probable glycosyltransferase | ||
| PGN_1255 | 1.04 | 1.28 | −2.64* | Putative heptosyltransferase | ||
| PGN_1302 | −1.25 | 1.29 | 1.16 | Hypothetical protein | ||
| PGN_1310 | −2.84 | 1.64 | −1.73* | Glycogen synthase | ||
| PGN_1481 | −1.17 | 1.08 | 1.33 | Putative polysaccharide biosynthesis protein | ||
| PGN_1614 | −1.53* | 1.31 | −1.72* | UDP-glucose 4-epimerase | ||
| PGN_1713 | −1.09 | −1.09 | 1.43 | Hypothetical protein | ||
| PGN_1718 | 1.03 | 1.16 | −2.09* | Probable UDP-2,3-diacylglucosamine hydrolase | ||
| PGN_1736 | −1.28 | 1.12 | −1.03 | Putative glycogen synthase | ||
| PGN_1750 | 1.59 | −1.25 | −1.36* | Putative 3-deoxy-d-manno-octulosonatecytidylyltransferase | ||
| PGN_2018 | 3.45 | −1.37 | −2.74* | Putative UDP-N-acetylglucosamineacyltransferase | ||
| PGN_2019 | 2.64* | −1.19 | −2.32* | UDP-3-O-[3-hydroxymyristoyl] N-acetylglucosaminedeacetylase | ||
| PGN_2020 | 1.37 | −1.11 | −1.60* | UDP-3-O-[3-hydroxymyristoyl] glucosamine N-acyltransferase | ||
| PGN_2086 | −1.07 | 1.00 | 1.23 | Probable acetyltransferase | ||
| Capsule | PGN_1100 | −1.15 | 1.22* | −1.29* | Putative capsule biosynthesis protein CapA | |
Bold text indicates a fold change of more than 1.5 or less than −1.5. *, P < 0.001 for the Welch t test. Shaded text indicates that the changes satisfy two criteria: a fold change of more than 1.5 or less than −1.5 and a statistically significant difference (P < 0.01, Welch t test).
The product name is from the P. gingivalis ATCC 33277 genome database (http://www.ncbi.nlm.nih.gov/sites/entrez?Db=genome&Cmd=Retrieve&dopt=Protein+Table&list_uids=22372).
B1, B2, B3, and B4 refer to the RNA samples harvested from the biofilm-forming cells after 3, 6, 9, and 14 days, respectively.
In conclusion, many genes were found to be time-dependently regulated during P. gingivalis ATCC 33277 biofilm growth, suggesting that they may be related to the pathogenicity of P. gingivalis.
Database sequence accession number.
The entire set of microarray data has been deposited in the Center for Information Biology Gene Expression Database (CIBEX) (http://cibex.nig.ac.jp/index.jsp) under accession number CBX149.
Supplementary Material
Acknowledgments
This study was supported by Grants-in-Aid for Scientific Research (20249076 and 21390508) by the Japan Society for the Promotion of Science.
Footnotes
Supplemental material for this article may be found at http://aem.asm.org/.
Published ahead of print on 29 July 2011.
REFERENCES
- 1. Asahi Y., et al. 2010. Effects of N-acyl homoserine lactone analogues on Porphyromonas gingivalis biofilm formation. J. Periodontal. Res. 45:255–261 [DOI] [PubMed] [Google Scholar]
- 2. Capestany C. A., Tribble G. D., Maeda K., Demuth D. R., Lamont R. J. 2008. Role of the Clp system in stress tolerance, biofilm formation, and intracellular invasion in Porphyromonas gingivalis. J. Bacteriol. 190:1436–1446 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3. Costerton J. W., Stewart P. S., Greenberg E. P. 1999. Bacterial biofilms: a common cause of persistent infections. Science 284:1318–1322 [DOI] [PubMed] [Google Scholar]
- 4. Davey M. E. 2006. Techniques for the growth of Porphyromonas gingivalis biofilms. Periodontol. 2000 42:27–35 [DOI] [PubMed] [Google Scholar]
- 5. Donlan R. M., Costerton J. W. 2002. Biofilms: survival mechanisms of clinically relevant microorganisms. Clin. Microbiol. Rev. 15:167–193 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6. Grenier D., et al. 2003. Effect of inactivation of the Arg- and/or Lys-gingipain gene on selected virulence and physiological properties of Porphyromonas gingivalis. Infect. Immun. 71:4742–4748 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7. Kadowaki T., et al. 1998. Arg-gingipain acts as a major processing enzyme for various cell surface proteins in Porphyromonas gingivalis. J. Biol. Chem. 273:29072–29076 [DOI] [PubMed] [Google Scholar]
- 8. Kuboniwa M., et al. 2009. Distinct roles of long/short fimbriae and gingipains in homotypic biofilm development by Porphyromonas gingivalis. BMC Microbiol. 9:105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9. Lin X., Wu J., Xie H. 2006. Porphyromonas gingivalis minor fimbriae are required for cell-cell interactions. Infect. Immun. 74:6011–6015 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10. Lo A. W., et al. 2009. Comparative transcriptomic analysis of Porphyromonas gingivalis biofilm and planktonic cells. BMC Microbiol. 9:18. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11. Naito M., et al. 2008. Determination of the genome sequence of Porphyromonas gingivalis strain ATCC 33277 and genomic comparison with strain W83 revealed extensive genome rearrangements in P. gingivalis. DNA Res. 15:215–225 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12. Nakayama K., Yoshimura F., Kadowaki T., Yamamoto K. 1996. Involvement of arginine-specific cysteine proteinase (Arg-gingipain) in fimbriation of Porphyromonas gingivalis. J. Bacteriol. 178:2818–2824 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13. Niba E. T., Naka Y., Nagase M., Mori H., Kitakawa M. 2007. A genome-wide approach to identify the genes involved in biofilm formation in E. coli. DNA Res. 14:237–246 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14. Noguchi N., Noiri Y., Narimatsu M., Ebisu S. 2005. Identification and localization of extraradicular biofilm-forming bacteria associated with refractory endodontic pathogens. Appl. Environ. Microbiol. 71:8738–8743 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15. Noiri Y., et al. 2003. Effects of chlorhexidine, minocycline, and metronidazole on Porphyromonas gingivalis strain 381 in biofilms. J. Periodontol. 74:1647–1651 [DOI] [PubMed] [Google Scholar]
- 16. Noiri Y., Li L., Yoshimura F., Ebisu S. 2004. Localization of Porphyromonas gingivalis-carrying fimbriae in situ in human periodontal pockets. J. Dent. Res. 83:941–945 [DOI] [PubMed] [Google Scholar]
- 17. Sauer K., Camper A. K., Ehrlich G. D., Costerton J. W., Davies D. G. 2002. Pseudomonas aeruginosa displays multiple phenotypes during development as a biofilm. J. Bacteriol. 184:1140–1154 [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.