Abstract
Mutations in the interdomain linker of the gene for the AraC regulatory protein of Escherichia coli that severely interfere with the protein's ability either to repress or to activate transcription have been found. These mutations have relatively small effects on the dimerization domain's ability to bind arabinose or to dimerize the protein or on the DNA-binding domain's affinity for a single DNA half-site. The linker mutations, however, dramatically change the affinity of AraC for binding to two direct-repeat DNA half-sites. Less dramatically, the induction-deficient linker variants also display altered DNA sequence selectivity. These results show that changing the sequence of the interdomain linker can profoundly affect the dimerization domain–DNA-binding domain interactions in AraC. The smaller effects on the functions of the individual domains could be the direct result of the linker alterations but more likely are the indirect result of the altered dimerization domain–DNA-binding domain interactions. In summary, the linker does not simply function as a passive and flexible connector between the domains of AraC but, instead, is more directly involved in the protein's dimerization domain–DNA-binding domain interactions.
INTRODUCTION
The classical genetic studies of Englesberg and collaborators (5, 6, 32) showed that the product of the Escherichia coli araC gene positively and negatively regulates expression of the araBAD genes in response to the presence of arabinose. Subsequent biochemical and biophysical studies have provided in vitro assays for the protein's activities and have provided much information about the structure, properties, and function of the protein (reviewed in references 28 and 29). Each monomer of the dimeric AraC protein (35) contains two structurally and functionally separate and distinct domains (Fig. 1): a dimerization domain (DD) that both dimerizes the protein and binds arabinose and a DNA-binding domain (DBD) (1, 9, 27) that both binds to DNA and provides the interactions with RNA polymerase that stimulate transcription initiation at the ara pBAD promoter (40). In vivo in the absence of arabinose, AraC binds in trans to two well-separated DNA half-sites, araO2 (O2) and araI1 (I1) the binding represses pBAD activity by forming a DNA loop in which pBAD is inactive (4, 16, 18, 19, 21). When arabinose appears, AraC protein shifts from predominantly looping the DNA to largely binding in cis to the adjacent direct-repeat DNA half-sites I1 and I2, from which the AraC protein stimulates both the binding of RNA polymerase and its transition to an open complex.
Fig. 1.
The repressing and inducing states of the araCBAD regulatory region and the domain organization of a single subunit of the AraC homodimer showing the locations of the important structural features. CAP, catabolite gene activator protein; RNAP, RNA polymerase.
In vitro, purified AraC protein has been found to bind with high affinity to direct-repeat DNA half-sites, inverted-repeat half-sites, and direct-repeat half-sites whose separation has been increased from 4 bases to 14 bases (2). Furthermore, the affinity of AraC for all these sites increases in the presence of arabinose (2). Finally, the DNA-binding and dimerization/arabinose-binding domains can function independently of one another (1, 8, 13, 27).
The in vivo activities of the protein, the ability of the two domains to function independently, the arabinose-dependent increase in DNA-binding affinity for the various orientations of two direct-repeat half-sites, the activity of two DBDs connected by a 20-residue linker (13), the activity of two DBDs dimerized by a coiled-coil domain (1), and the DNA looping and unlooping properties of AraC (4, 19) are well explained by a mechanism in which, only in the absence of arabinose, the DBDs are restrained by being held to the DD in positions and orientations that favor DNA looping (8, 27). A recent physical study directly demonstrated a central component of the proposed mechanism for AraC function, which is that in the absence of arabinose there is indeed a DD-DBD interaction, that it has about the appropriate strength, and also that the presence of arabinose weakens or eliminates the interaction (10). The increased freedom of the DBDs that is promoted by the presence of arabinose presumably allows them to orient such that they can easily bind to either of the adjacent DNA half-sites in a direct-repeat or an inverted orientation.
Preliminary mutational studies of the linker of AraC were performed (7) once the approximate domain boundaries in AraC had been established with genetic studies (1, 9). These mutational studies suggested that changes of a single amino acid in the linker produced minor effects but that changes of multiple residues might be capable of producing a substantial effect. Unfortunately, the majority of the mutations that produced significant effects included changes to amino acid residues that are now known from the X-ray structure of the dimerization domain of AraC and the nuclear magnetic resonance structure of the DNA-binding domain of AraC to lie within regions of stable secondary structure (24, 34, 37). Therefore, with more precise knowledge of the location of the interdomain linker in AraC and the more recent development of assays for activities of the individual domains (26, 36), we have reexamined the consequences of mutating the interdomain linker of AraC.
The experiments reported here show that mutations in the interdomain linker of AraC can dramatically interfere with the protein's ability to repress or to activate the pBAD promoter. While the mutations strongly alter the activities of the protein that require the simultaneous binding to two DNA half-sites, the mutations have much smaller effects on the intrinsic activities of either of the individual domains. That is, the DD/arabinose-binding domains of the mutant proteins largely retained their ability to bind arabinose, and the DBDs largely retained their ability to bind to a single I1 half-site. Thus, by elimination, since the DBDs themselves have shown no evidence of DBD-DBD interactions even up to near millimolar concentrations (24), the linker mutations must primarily affect the protein's DD-DBD interactions. Neither the induction-defective linker variants nor the repression-defective linker variants possess dramatic and obvious properties such as an excess or deficit of small, large, or charged amino acids. Therefore, our understanding of the basis of the new linker mutations ultimately will require additional and more detailed structural analysis.
MATERIALS AND METHODS
DNA oligonucleotides.
Synthetic DNAs used in this study are named for the half-sites that they contain, and the bases that comprise the AraC binding sites are underlined in this section. The fluorescently (Cy5) labeled direct repeat I1-I1 contains two I1 half-sites with the sequence of 5′-Cy5-GCCATAGCATTTTTATCCATAAGATTAGCATTTTTATCCATACCTC annealed to its complement, and unlabeled direct-repeat I1-I1 competitor DNA contained the same sequence. Cy3-labeled direct-repeat I1-I2 was 5′-Cy3-GCCATAGCATTTTTATCCATAAGATTAGCGGATCCTACCTGACCTC annealed to its complement. 6-Carboxyfluorescein (FAM)-labeled direct-repeat I1-O2 was 5′-FAM-ACCATAGCATTTTTATCCATAAGATGAAACCAATTGTCCATACCTC annealed to its complement. These synthetic direct-repeat half-sites maintain the same half-site spacing that occurs in the operon. The Cy5-labeled I1 half-site was 5′-Cy5-GCTAGCATTTTTATCCATACCTCCCTC annealed to its complement.
Flexibly linked half-sites for the study of binding, similar to that which occurs in DNA looping, were composed of three DNA strands (25). The oligonucleotides form a complex consisting of the two double-stranded AraC DNA half-sites separated by 24 bases of single-stranded DNA. Thus, one strand of the flexibly linked half-sites consists of the 24 bases flanked on each end by the sequences of the two half-sites. To this was hybridized the complementary sequences of the two half-sites. A previous study (25) found that a 24-base single-stranded linker (L24) was sufficient to eliminate the arabinose dependence of binding characteristic of DNA looping. For clarity, the DNA sequences that remain single stranded after annealing are presented in lowercase. All flexibly linked DNAs contained 5′-Cy5-GTCTTATGGATAAAAATGCTAGGGT. This DNA strand was annealed to 5′-ACCCTAGCATTTTTATCCATAAGACctactggtaccgtctcatgcatagACCCGAAACCAATTGTCCATAAGACc and 5′-GTCTTATGGACAATTGGTTTCGGGT for I1-L24-O2 or was annealed to 5′-ACCCTAGCATTTTTATCCATAAGACctactggtaccgtctcatgcatagACCCTAGCGGATCCTACCTGAAGACc and 5′-GTCTTCAGGTAGGATCCGCTAGGGT for I1-L24-I2. The single-stranded DNA linker is identical in all constructs, and the sequence of the single-stranded DNA and the presence of a single 3′ base overhang are not believed to affect the ability of AraC to bind to these flexibly linked constructs. The length and sequence of the single-stranded DNA linker were identical to those of the one used in a previous study (25).
Isolation of linker mutations in AraC.
The coding region of AraC was cloned between the NcoI and SacI sites of the pET24d vector (Novagen) and transfected into AraC− AraBAD+ strain SH321 (12). QuikChange mutagenesis reactions were used to randomize residues 169 to 171, 169 to 173, 170 to 172, 171 to 173, 172 to 174, 172 to 175, 172 to 176, 173 to 175, 174 to 175, 175, and 176 and to make the N168P variant of AraC. These mutations were made in the Y31V variant of AraC to minimize aggregation in the in vitro experiments (37). The Y31V mutation does not alter the in vivo phenotype of AraC and, with the exception of its decreased tendency to aggregate, does not appear to alter the behavior of AraC.
Determination of in vivo phenotype of AraC linker mutants.
For the initial screening of the randomized linker mutants, the resulting DNAs were used to transform AraC− host SH321, and the transformants were spot tested for the resulting phenotypes on minimal arabinose, minimal arabinose-fucose, or tetrazolium indicator plates (30). Mutations with a noninducible phenotype (Ind−) were red on tetrazolium-arabinose and unable to grow on minimal arabinose medium, and mutations that were unable to repress transcription from pBAD (Rep−), which are also called constitutive, were able to grow on minimal arabinose-fucose plates. The level of expression of the arabinose isomerase in the fucose-resistant mutants was assayed in cells grown on minimal glycerol-arabinose-Casamino Acids medium (30).
Purification of WT AraC and its linker variants.
The proteins with mutations in the linkers and also containing the Y31V AraC background mutation and wild-type (WT) AraC, which lacked the Y31V mutation, were overexpressed from the pET24d vector in the E. coli strain BL21(DE3). The Y31V mutation was not included in the WT AraC construct because WT AraC does not significantly aggregate under the conditions and concentrations used in this study, and a previous study indicated that Y31V had no significant effects on the behavior of AraC beyond the improved aggregation profile (37).
Cells containing AraC or one of its mutants were grown at 37°C in YT medium (30) with 40 μg/ml kanamycin to an optical density at 600 nm of ∼0.6 to 1.5. The cells were transferred to room temperature (∼20°C) shakers, and 13 mM arabinose was added to the cultures of WT AraC and Ind− mutant cells but not the Rep− mutant cell cultures. After 15 min, the cells were induced by the addition of isopropyl-β-d-thiogalactopyranoside (IPTG) to a concentration of 0.4 mM IPTG and were grown for between 3 h and overnight. Cells were pelleted by centrifugation and stored either at 4°C or at −80°C.
The cell pellets were resuspended in ∼40 ml of lysis buffer, 50 mM Na-phosphate, pH 6.5, 300 mM NaCl, 1 mM EDTA, pH 7.0, 13 mM arabinose, 5% glycerol, and 1 mM dithiothreitol. During the purification of WT AraC and the Ind− mutant strains but not the Rep− mutant strains, MgCl2 was added to a concentration of 0.2 mM. Immediately prior to lysis, phenylmethylsulfonyl fluoride in 100% ethanol was added to a final concentration of 0.1 mM, and a sprinkle of DNase was added. The resuspended cells were lysed in an Avestin EmulsiFlex-C3 cell homogenizer, diluted to ∼90 to 100 ml with 20 mM Na-phosphate, pH 6.5, buffer containing 10 mM arabinose, and then centrifuged at 10,000 to 12,000 rpm for 15 to 20 min at 4°C to pellet cell debris. The supernatant from the first centrifugation step was centrifuged for an additional 15 to 20 min at 10,000 to 12,000 rpm. The supernatant from the second centrifugation step was loaded at 0.5 to 3 ml/min onto a 5-ml HiTrap heparin HP column (Amersham Biosciences) and eluted at concentrations of ∼400 to 600 mM NaCl using a 50-ml gradient of 150 to 1,000 mM NaCl in 20 mM Na-phosphate, pH 6.5, buffer containing 10 mM arabinose. The Rep− AraC-containing fractions from the heparin column were pooled and diluted ∼2- to 3-fold with buffer containing 20 mM Tris-HCl, pH 7.5, and 10 mM arabinose, loaded at 1 to 3 ml/min onto a 5-ml HiTrap Q HP column (Amersham Biosciences), and retrieved from the column flowthrough. This flowthrough was then loaded onto a 5-ml HiTrap heparin HP column. The Rep− mutants of AraC-containing strains were eluted from the column in a 24-ml gradient of 150 to 750 mM NaCl in 20 mM Na-phosphate, pH 6.5, buffer containing 10 mM arabinose. After the final purification step, glycerol was added to 30% in the AraC-containing fractions, and the proteins were stored at −80°C.
Determination of active fraction of AraC.
The concentration of purified AraC was calculated using an extinction coefficient of 48,150 M−1 cm−1 per monomer calculated from the amino acid composition (11). The activity of AraC was determined by titration against known concentrations of direct-repeat I1-I1 using electrophoretic mobility shift assays in which the DNA concentrations were well above the Kd (dissociation constant) of the interaction (15). These activity experiments were performed in 50 to 150 mM KCl binding buffer containing 10 mM Tris-acetate, pH 7.4, 1 mM EDTA, pH 7.0, 5% glycerol, 0.05% NP-40, and 0.01 mg/ml bovine serum albumin in either the presence or absence of 50 mM arabinose. Freshly thawed aliquots of AraC and its linker mutant proteins contained at least 30% and typically more than 60% active dimers. Aliquots were discarded after a few months of storage at 4°C or after their activity fell below ∼15% active dimers. To estimate the concentration of active monomer required for half-site affinity assays, we assumed that active and inactive subunits randomly and independently assort and that the activities of subunits in a dimer are independent of one another, in which case the fraction of active monomers is the square root of the fraction of active dimers. The concentration of AraC used throughout the study is the concentration of active dimers or, in the case of the half-site affinity experiments, active monomers rather than the concentration of total protein.
Arabinose-binding assay.
The affinity of AraC for arabinose was determined by monitoring the arabinose-induced changes in average emission wavelength of the intrinsic tryptophan fluorescence of AraC by excitation at 295 nm and collection of the emission spectra between 320 nm and 370 nm (26, 37). Typically, these experiments were performed in 133 mM KCl in 89% strength binding buffer that lacked NP-40, in which 500 nM AraC was titrated with increasing concentrations of arabinose. The change in the average emission wavelength of AraC was fitted to a Langmuir binding isotherm that accounted [ztotal] ≠ [zfree] (equation 1), where z is the concentration of titrant. In equation 1, F is the concentration of the fluorescent species, xo is the initial value of the fluorescent signal, Δx is the change in the fluorescent signal, and X¯ is the fraction bound. Equation 1 was used to fit all equilibrium binding constants presented in this paper.
| (1) |
I1 half-site DNA-binding assay.
The affinity of AraC for I1 half-site DNA was determined by monitoring the change in the fluorescence anisotropy of Cy5-labled DNA that occurs upon protein binding. The fluorescence anisotropy was measured by exciting the Cy5 fluorophore at 645 nm and integrating its emission at 660 nm for 30 s. In either the presence or absence of 50 mM arabinose, 5 to 10 nM Cy5-labeled I1 half-site DNA in 150 mM KCl binding buffer was titrated with increasing concentrations of AraC. The affinities were determined from the fit to equation 1.
I1-I1 direct-repeat DNA-binding assay.
The affinity of AraC for direct-repeat I1-I1 DNA was determined using electrophoretic mobility shift assays (14, 25), in which the binding was performed at sufficiently high salt concentrations (200 to 700 mM KCl) that the DNA-binding affinity of AraC was on the order of or weaker than the molar concentration of DNA. For each titration, in either the presence or absence of 50 mM arabinose, 0.02 to 0.4 nM Cy5-labeled I1-I1 DNA in binding buffer containing between 200 and 700 mM KCl, as indicated, was allowed to equilibrate with 8 to 15 different concentrations of AraC for at least 10 min. The range of concentrations of active AraC dimers used for each Kd measurement typically was from 10−10 M to 10−6 M. Immediately before the sample was loaded onto the gel, at least a 2-fold excess of unlabeled competitor I1-I1 DNA was added and the samples were loaded within 30 s into the wells of the 6% gels. After electrophoresis, the fluorescently labeled DNA bands were visualized on a Typhoon 9410 variable-mode imager and quantitated using ImageQuant TL software (Amersham Biosciences). Typically, the gel images were subjected to the 5-by-5 median filtering algorithm in ImageQuant TL, and the background noise was subtracted using the rolling-ball method. The affinities were determined from the fit to equation 1.
Relative DNA-binding affinities for direct-repeat versus flexibly linked half-sites.
The relative affinities of AraC for flexibly linked half-sites, where the flexible linker was L24, a 24-base single-stranded DNA linker (Cy5-I1-L24-O2 or Cy5-I1-L24-I2), and for direct-repeat half-sites (Cy3-I1-I2 or FAM-I1-O2) were determined by including both DNAs at 4 to 5 nM in the same reaction mixtures with 3 to 5 nM AraC and determining the bound/free ratio of each DNA species (31) after electrophoresis, as in the standard electrophoretic mobility shift assay. Arabinose, if present, was at 25 mM, and the binding buffer contained 75 mM KCl and 2 mM Mg acetate. The protein and DNAs were allowed to equilibrate for 25 min before the gels were loaded. In this type of binding experiment, the relative affinities of AraC for I1-I2 or I1-L24-I2 DNA is proportional to the ratio of the bound to free DNA bands for each of two DNA species (31) (equation 2).
| (2) |
Interpretation of this experiment required the assumption that the salt dependence of DNA binding is similar for the two DNA species.
Half-site DNA sequence selectivity.
The pairwise relative affinities of AraC for two DNAs containing different types of half-sites (the Cy5-I1-I1 or Cy3-I1-I2 half-site or the FAM-I1-O2 half-site) were determined in the same manner as the relative affinities of AraC for direct-repeat versus flexibly linked half-sites.
RESULTS
Isolation of linker mutations.
AraC was cloned into the pET24d overexpression plasmid vector to facilitate both physiological and biochemical analyses of the effects of the mutations. The former was facilitated by our finding that sufficient AraC protein is synthesized by the plasmid during normal cell growth in our K-12 tester strain, SH321, to regulate the chromosomal pBAD promoter normally without explicitly inducing the T7 overexpression promoter. The latter was possible because upon induction of the T7 promoter, high levels of AraC are synthesized, thus facilitating purification of the protein and subsequent biochemical assays.
Oligonucleotide-directed mutagenesis was used to randomize different regions of the interdomain linker. Plasmid DNA from randomly chosen colonies was sequenced, and genetic screening was performed on those containing alterations only in the linker region. Figure 2 shows the sequence of the region, the domain boundaries, the sequences of the mutants isolated, and their phenotypes. Seven of the interdomain linker mutations with regulatory effects were chosen for further study: the three Rep− mutations ESL → MSP, SLH → FDA, and LHP → NYL and the four Ind− mutations HPPM → LKDR, M175P, D176F, and D176G.
Fig. 2.
Sequence of the interdomain linker region of AraC, the mutations isolated, and their resulting phenotypes. Gray shading, wild-type induction and repression behavior; light gray shading, Rep− or Ind−.
The DD is not significantly affected by linker mutations.
Because the region indicated to be a linker in Fig. 2 displayed no regular secondary structure and, in fact, varied in structure from one monomer to the next in the X-ray structure studies of the dimerization domain and because the DNA-binding domain lacking the linker is functional (24, 34, 37), we conclude that the linker region is not an integral part of either the dimerization or DBD. It therefore seems unlikely that the primary effect of linker mutations is on the intrinsic activities of either domain. Nevertheless, we examined the individual activities of the two domains of AraC: arabinose binding of the DD and DNA binding to a single DNA half-site of the DNA-binding domain.
The functions of the DDs of AraC are to bind to the inducer, arabinose, and to dimerize the protein (1). If the linker mutations do not significantly alter these functions of the DD, then compared to WT protein, the mutant protein should have a similar affinity for arabinose and should still dimerize the protein. This is what we found experimentally.
To measure the binding of arabinose to the protein, we monitored the slight arabinose-induced shift in the average fluorescence emission wavelength from the tryptophan residues naturally present in AraC (26, 37). Figure 3 shows typical data obtained with wild-type AraC and three Rep− linker mutant proteins. Table 1 presents the apparent dissociation constants of the mutant proteins for arabinose. The affinities of the linker variants for arabinose differed from the affinity of WT AraC by less than 2-fold, which demonstrates that the linker mutations do not significantly affect the arabinose-binding properties of AraC.
Fig. 3.
Binding of arabinose by wild-type AraC and Rep− mutant proteins. The average emission wavelength of the intrinsic tryptophan fluorescence of AraC or a Rep− variant as a function of molar arabinose concentration.
Table 1.
Apparent affinities of AraC and its variants for arabinosea
| WT | Rep− |
Ind− |
|||||
|---|---|---|---|---|---|---|---|
| MSP | FDA | NYL | LKDR | M175P | D176G | D176F | |
| 340 ± 30 | 400 ± 100 | 360 ± 10 | 320 ± 20 | 440 ± 30b | 500 ± 30 | 550 ± 30b | 490 ± 50 |
Affinities are in μM, and the errors are estimated from fitting the binding curve as described in Materials and Methods.
The Kapp was measured in buffer containing 133 mM KCl for all samples except D176G and LKDR, which were measured in the presence of 100 mM KCl.
The mutant proteins possessed elution properties similar to those of wild-type AraC during purification, suggesting that they were dimeric at the concentrations encountered during purification. More definitively, however, when binding to DNA containing two half-sites, e.g., I1-I1, no nonlinearity in binding versus protein concentration, which would be indicative of a monomer-dimer equilibrium, was ever observed.
The DBD is not significantly affected by linker mutations.
As described in the previous section, to confirm that the linker mutations did not substantially change the biochemical activities of the individual domains of AraC, we also tested whether the linker mutations altered the affinity of a DBD for a single DNA half-site. We measured binding to Cy5-labeled I1 half-site DNA using fluorescence anisotropy. Typical data are shown in Fig. 4. Table 2 presents the affinities of the repression-negative (constitutive) mutant proteins. As their phenotype differs from that of wild-type AraC in the absence of arabinose, their DNA-binding affinities were measured in the absence of arabinose. For analogous reasons, the induction-negative mutant proteins were measured in the presence of arabinose (Table 2). WT AraC was measured in both the absence and presence of arabinose for comparison purposes. Only moderate changes in DNA-binding affinity are observed for most of the mutant proteins. The D176 mutant proteins show larger changes in half-site affinity, but anomalous behavior of the binding curves for these mutant proteins at high concentrations and a fine white precipitate that was visible at the end of the titration suggest that these proteins were aggregating during the course of the titrations. The other experiments in this study involving D176 did not require such high protein concentrations and did not display any evidence of aggregation.
Fig. 4.
Binding of an I1 half-site by wild-type AraC and the Rep− variants. The fluorescence anisotropy of a Cy5-labeled I1 half-site as a function of the total molar concentration of WT AraC or a Rep− variant.
Table 2.
Affinities of WT and linker variants for an I1 half-site DNAa
| WTb | Rep− |
WTc | Ind− |
|||||
|---|---|---|---|---|---|---|---|---|
| MSPb | FDAb | NYLb | LKDRc | M175Pc | D176Gc | D176Fc | ||
| 46 ± 3 | 9.2 ± 0.6 | 28 ± 3 | 16 ± 1 | 8 ± 1 | 14 ± 2 | 9.3 ± 0.8 | 700 ± 100d | 160 ± 20d |
Affinities are in nM, and the errors are estimated from fitting the single binding curve as described in Materials and Methods.
Affinity was measured in the absence of arabinose.
Affinity was measured in the presence of arabinose.
Showed evidence of aggregation.
Mutant proteins' affinities for direct-repeat DNA are significantly affected.
As described above, we found that the intrinsic activities, that is, activity that does not depend upon another domain or subunit, of the domains of the linker variants were largely like those of the wild-type domains. Therefore, we tested whether the linker mutant proteins possessed markedly altered DNA-binding activities when both domains were required to act together, as is necessary for high-affinity binding to direct-repeat half-sites. Because preliminary experiments suggested that some of the mutant proteins bind to direct-repeat DNA significantly more weakly than wild-type AraC, it was important to ensure that the affinity being measured was that for binding in which both of the subunits of AraC contact both of the half-sites rather than binding by a single DBD to a single half-site. For this reason, we measured binding to the high-affinity direct-repeat I1-I1 site rather than the weaker I1-I2 site that is found at the pBAD promoter.
In buffers containing physiological concentrations of salt, the affinity of wild-type AraC for direct-repeat I1-I1 DNA is too high to be conveniently measured with the gel-based electrophoretic mobility shift assay using fluorescently labeled DNA. Since at salt concentrations above 200 mM the DNA-binding affinity of AraC varies approximately with the 13th power of the concentration (22), it was possible to bring the affinities of WT AraC and its linker mutants for I1-I1 DNA to values convenient for assay. Typically, we used salt concentrations of about 300 mM. To maximize the accuracy of the measurements, it was necessary to measure the DNA-binding affinities of each mutant protein in somewhat different salt concentrations and, in several cases, to extrapolate a mutant protein's affinity to the 300 mM salt concentration chosen for comparison of the affinities.
Figure 5 shows a typical gel from which a dissociation constant at one salt concentration could be determined. Normally, we determined the Kd using at least six different salt concentrations. From these the affinity and dependence on salt concentration were determined. Figure 6 shows the measured affinities as a function of salt concentration for wild-type AraC and the linker variants, where it can be seen that the Rep− mutant proteins bind direct-repeat half-sites more tightly than WT AraC does in the absence of arabinose and the Ind− mutant proteins bind direct-repeat half-sites less tightly than WT AraC does in the presence of arabinose. The salt dependence of binding to direct-repeat half-sites is similar for WT AraC and all of the mutant proteins, and the small differences in the slopes in Fig. 6 are likely the result of experimental error associated with measuring the Kd of the interaction. The affinities of the variants interpolated or extrapolated to 300 mM are presented in Table 3.
Fig. 5.
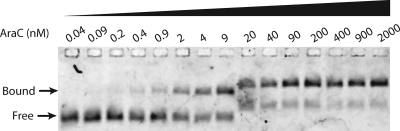
A typical electrophoretic mobility shift assay experiment to measure the affinity of AraC for direct-repeat I1-I1 half-sites. Cy5-labeled I1-I1 DNA was incubated with increasing amounts of WT AraC. The different migration rates are due to time differences between when the first 8 lanes and the last 7 lanes were loaded, which resulted in different lengths of time for the electrophoresis. The fraction bound was fit to equation 1, and for the experiment shown, the Kd of WT AraC binding to I1-I1 DNA in 500 mM KCl binding buffer was 2.7 nM.
Fig. 6.
Binding of I1-I1 direct-repeat DNA by wild-type AraC and its linker variants. Binding of WT AraC (A and B), Rep− AraC (A), and Ind− AraC (B) to Cy5-labled I1-I1 direct-repeat DNA in the absence of arabinose (A) and in its presence (B) as a function of the molar concentration of KCl. Each data point represents the Kd (M) from fitting the fraction bound and the error associated with the fit of a single titration curve to equation 1. The lines represent the best fits of the data points to straight lines, as described previously (22).
Table 3.
Affinities of AraC and its variants for I1-I1 direct-repeat DNAa
| −arabinose |
+arabinose |
|||||||
|---|---|---|---|---|---|---|---|---|
| WT | Rep− |
WT | Ind− |
|||||
| MSP | FDA | NYL | LKDR | M175P | D176G | D176F | ||
| 0.7 ± 0.5 | 0.1 ± 0.2b | 0.3 ± 0.1 | 0.1 ± 0.1b | 0.02 ± 0.01b | 3 ± 4 | 0.6 ± 0.8 | 3 ± 1 | 30 ± 30 |
Affinities are in nM and were either interpolated or extrapolated to 300 mM KCl.
A Kd that was extrapolated to 300 mM KCl.
The Ind− mutant protein's altered direct-repeat DNA-binding affinities are not just the result of the decreased affinity of the DBDs for a half-site.
Conceivably, the relatively small weakening in the affinities of the Ind− mutant proteins for a half-site measured earlier (Table 2) could be sufficient to explain the relatively large effects on binding to two half-sites measured and described in the previous section (Table 3). This follows, because the binding of AraC to two half-sites involves two DBDs, and therefore, the effects, like those measured earlier for binding to a single half-site, would be squared for binding to two half-sites; see the work of Timmes et al., for example (36). We therefore compared (Table 4) the measured affinity of the Ind− variants for binding to I1-I1 direct-repeat half-sites to an affinity calculated by assuming that the only effect of a linker mutation is on the intrinsic affinity of the DBDs. That is, the effect on the affinity of a variant binding to DNA containing the I1-I1 direct repeat compared to the affinity of the wild type will be the square of the mutation's effect on binding to a single I1 half-site. If the actual affinity change of a variant greatly exceeds this predicted change, then we can conclude that the linker mutation does more than alter the intrinsic binding activities of the individual DBDs. Such mutations would have to alter the ability of both DBDs in an AraC dimer to simultaneously contact two adjacent DNA half-sites. Such a change could be a result of the mutations altering the intrinsic structure of the interdomain linker or altering the flexibility of the linker or a result of altered domain-linker interactions. Ultimately, however, whatever the detailed, atomic-level mechanism, the linker mutations affect the ability of the arabinose-binding status of the DD to be communicated to the DBD. Thus, the effects of the mutations can be summarized to be affecting the DD-DBD interactions. As shown in Table 4, the affinities of the induction-negative mutant proteins predicted on the basis of the above-mentioned assumptions differ substantially from their experimentally measured affinities. We thus conclude that the linker mutations have altered the DD-DBD interactions, presumably by affecting linker flexibility or structure. We did not include the data from the D176 mutant protein in reaching this conclusion because we consider these data to be suspect due to aggregation, as mentioned above.
Table 4.
Comparison of the predicted and interpolated affinities of AraC and the Ind− variants for I1-I1 direct-repeat DNA
| −arabinose |
+arabinose |
|||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| WT | LKDR | M175P | D176G | D176F | WT | LKDR | M175P | D176G | D176F | |
| Measureda | 0.7 ± 0.5 | 50 ± 900 | 30 ± 50 | 200 ± 1,000 | 200 ± 500 | 0.02 ± 0.01 | 3 ± 4 | 0.6 ± 0.8 | 3 ± 1 | 30 ± 30 |
| Predictedb | 0.63 ± 0.08 | 0.027 ± 0.003 | 0.33 ± 0.06 | 5.9 ± 0.8 | 29 ± 3 | 0.019 ± 0.005 | 0.026 ± 0.004 | 0.026 ± 0.004 | 150c ± 80 | 8c ± 2 |
| Ratio of measured/predicted | 0.9 ± 0.6 | 1,700 ± 200 | 80 ± 10 | 35 ± 5 | 8 ± 1 | 1.0 ± 0.2 | 50 ± 20 | 20 ± 4 | 0.02c ± 0.01 | 3.6c ± 0.9 |
Affinities are in nM and were either interpolated or extrapolated to 300 mM KCl.
Affinities are nM, using a factor of 3 × 105 to correct for salt concetration and local concentration effects, assuming that the salt dependence of direct-repeat binding and the local effective concentration are similar for WT AraC and all of its linker variants. The errors associated with fold differences were calculated using error propagation.
Evidence of aggregation in the half-site affinity experiment.
One might expect that the Ind− linker mutants would not show an arabinose response. Table 4, however, shows that they do. Apparently, their inability to stimulate transcription in the presence of arabinose is a result of their very low affinity, even in the presence of arabinose, for direct-repeat half-sites. As shown in Table 4, their affinity for direct-repeat half-sites in the presence of arabinose is lower than the affinity of wild-type AraC in the absence of arabinose. Since wild-type AraC does not induce in the absence of arabinose, it is not surprising that the Ind− mutants do not induce in the presence of arabinose.
Linker variants alter the looping versus induction preference of AraC.
As described in the introduction (Fig. 1), the presence of arabinose causes AraC to switch between the repressing, looped state, where, apparently as a result of DBD-DD interactions, its two DBDs are bound to two half-sites that are separated by 210 bp, and the inducing state, where the domains are bound to two adjacent direct-repeat half-sites. In this section, we demonstrate in another way that the linker mutations alter the ability of the two DBDs of AraC to function together. In this experiment, two DNAs were simultaneously present in the DNA-binding assays. The first DNA consisted of an I1 half-site separated from an O2 half-site by 24 bases of flexible, single-stranded linker DNA and was named I1-L24-O2. Binding to this DNA mimics DNA looping by AraC (25, 38, 39). The long and flexible single-stranded DNA connector between the two DNA half-sites allows the two DBDs of AraC to bind to this DNA even while they are held in orientations incompatible with binding to adjacent direct-repeat half-sites. As a result, binding to this DNA is rather tight, even if AraC is trapped in a conformation incompatible with binding to adjacent direct-repeat DNA half-sites. The second DNA studied was I1-I2. Binding to this mimics that which is required for activation of induction from ara pBAD. When both DNAs are incubated in the same reaction, the ratio of the AraC bound to free DNA for each species equals the ratio of the affinities for the two DNAs (see Materials and Methods). The resulting data (first row of Table 5) show that in comparison to wild-type AraC, the Rep− variants prefer, by a factor of about 4, to bind to direct-repeat I1-I2 DNA than to bind to I1-L24-O2 DNA. Conversely, in these competition experiments, the Ind− variants slightly prefer, by an average factor of 2, to bind to I1-L24-O2 DNA than to bind to direct-repeat I1-I2 DNA. These results are consistent with the mutants' in vivo behaviors. We note that the absolute values of these ratios cannot be used to directly predict the in vivo phenotypes of the various forms of AraC because the substrate DNAs are imperfect mimics of the actual DNA in its in vivo environment, consisting of many other proteins, different salt concentrations, different lengths and flexibilities, and the presence of DNA supercoiling. Thus, only the direction of the changes of the ratios should be considered, and even then, the ratios do not lead to unequivocal conclusions. The results suggest that in comparison to wild-type AraC, the Rep− proteins activate transcription because they bind more tightly to the direct-repeat I1-I2 half-sites and that the Ind− proteins do not activate transcription because they bind more weakly to the direct-repeat I1-I2 half-sites.
Table 5.
Relative preferencea of WT AraC and its linker variants for direct-repeat versus flexibly linked half-sites
| −arabinose |
+arabinose |
||||||||
|---|---|---|---|---|---|---|---|---|---|
| WT | Rep− |
WT | Ind− |
||||||
| MSP | FDA | NYL | LKDR | M175P | D176G | D176F | |||
| I1-I2 vs I1-L24-O2 | 0.12 ± 0.02 | 0.41 ± 0.07 | 0.51 ± 0.03 | 0.49 ± 0.03 | 1.0 ± 0.2 | 0.21 ± 0.04 | 0.70 ± 0.04 | 0.5 ± 0.1 | 0.5 ± 0.1 |
| I1-I2 vs I1-L24-I2 | 1.5 ± 0.2 | 3.1 ± 0.8 | 4.2 ± 0.3 | 2.7 ± 0.5 | 3 ± 1 | 0.8 ± 0.1 | 1.8 ± 0.5 | 1.3 ± 0.3 | 1.2 ± 0.2 |
| I1-O2 vs I1-L24-O2 | 0.6 ± 0.1 | 3.4 ± 0.8 | 2.7 ± 0.2 | 4.8 ± 0.7 | 1.5 ± 0.2 | 1.7 ± 0.2 | 5.6 ± 2.3 | 7 ± 1 | 7 ± 1 |
Errors are the 95% confidence interval of between 3 and 9 repetitions of the measurement, using the same stock solutions but independent binding reactions.
The experiments described in the paragraph above involved two differences between the two DNAs used. The first was that the DNA used for looping contained I1 and O2 half-sites, while the direct-repeat DNA contained I1 and I2 half-sites, and the second was looping versus binding to direct-repeat half-sites. To further separate the factors responsible for the Ind− and Rep− phenotypes of the linker mutant proteins, we examined looping using DNAs with the same half-site composition and also examined the sequence selectivity of the variants when binding to direct-repeat half-sites.
To determine directly whether the linker variants altered the ability of AraC to loop DNA in the absence of any DNA sequence selectivity effects, we compared binding between I1-I2 and I1-L24-I2 and between I1-O2 and I1-L24-O2. In these experiments in the absence of arabinose, the Rep− variants' preference for looping (repression) was reduced compared to that of wild-type AraC (Table 5, rows 2 and 3), thus demonstrating unequivocally that the repression-negative variants have reduced abilities to bind to DNA in the looping state compared to binding to adjacent half-sites.
In the half-site selectivity experiments, binding to I1-O2 and I1-I2 was compared directly in single binding reactions as described in the previous section, as were binding to I1-O2 and I1-I1 and binding to I1-I1 and I1-I2. The results of the pairwise comparisons are presented in Table 6. Somewhat unexpectedly in the context of direct-repeat half-sites, Ind− linker mutations displayed appreciably altered DNA sequence selectivity (see Discussion), with an apparent decrease in affinity for I2. The sequence selectivity of the Rep− mutant proteins, however, was found to be similar to that of wild-type AraC, with only 0.8- to 3.6-fold differences between the Rep− variants and WT AraC. Since this experiment shows that the Rep− mutations did not significantly alter the DNA sequence selectivity of AraC and the results of the previous experiment (Table 5) showed that the Rep− mutations altered the looping preference, the Rep− mutations must have reduced the stability of the loop-forming structure of AraC. Apparently, the altered linker weakens the loop-favoring DD-DBD interaction, and this allows the protein to bind to direct-repeat half-sites without incurring the energetic cost(s) of breaking the interaction between the domains. As a result, the protein binds to direct-repeat half-sites with an increased affinity.
Table 6.
Half-site selectivity in relative preferencea of WT AraC and linker variants when binding to direct-repeat half-sites
| arab | WT | Rep− |
Ind− |
||||||
|---|---|---|---|---|---|---|---|---|---|
| MSP | FDA | NYL | LKDR | M175P | D176G | D176F | |||
| I1-O2vs I1-I2 | − | 7 ± 1 | 6.7 ± 0.5 | 9 ± 4 | 9 ± 4 | 10 ± 5 | 10 ± 3 | 16 ± 5 | 16 ± 2 |
| + | 1.95 ± 0.05 | 1.6 ± 0.2 | 1.7 ± 0.9 | 1.6 ± 0.3 | 7 ± 3 | 11 ± 4 | 11 ± 7 | 6 ± 3 | |
| I1-I1 vs I1-O2 | − | 4 ± 1 | NDc | ND | 4 ± 3 | 10 ± ND | 9 ± ND | 12 ± ND | 10 ± ND |
| + | 2.5 ± 0.3 | ND | ND | 2.5 ± 0.3 | 3.2 ± 0.4 | 3.2 ± 0.4 | 1.9 ± 0.6 | 2.1 ± 0.4 | |
| I1-I1 vs I1-I2 | − | 18 ± 3 | 9 ± 3 | 5 ± 3 | 6 ± 2 | 60 ± 40 | 60 ± 30 | 100 ± 40 | 30 ± 10 |
| + | 2.3 ± 0.3 | 1.8 ± 0.5 | 2.4 ± 0.3 | 2.3 ± 0.3 | 23 ± 6 | 14 ± 2 | 1 ± 3 | 21 ± 9 | |
Errors are the 95% confidence interval of between 2 and 9 repetitions of the measurement, using the same stock solutions but independent binding reactions. When only one measurement was made, no confidence interval was provided.
ara, arabinose.
ND, not determined.
The behavior of the Ind− mutants in this looping study is more difficult to interpret. When binding to I1 and I2 half-sites, the Ind− mutant proteins appear to have simply altered the looping preference of AraC, but when binding to I1 and O2 half-sites, the Ind− mutant proteins do not appear to have altered the looping preference of AraC. These results and the DNA sequence selectivity results suggest that Ind− mutations have altered more than just the looping preference of AraC.
A potential mechanism for Rep− mutants.
A significant fraction of the repression-negative, that is, the constitutive, mutants replaced residues of the linker with the helix-breaking residues of glycine or proline (Fig. 2). These mutations occurred in the first part of the interdomain linker ahead of the pair of proline residues that lie in the latter half of the linker. It is conceivable, therefore, that in the inducing state, the upstream helix ends at about residue 168, as seen in the crystal structures of the DD (33, 37), but that in the repressing state, the helix extends to the first proline in the interdomain linker, residue 173. Hence, introducing a proline or glycine residue between residues 168 and 173 could end the helix early and produce a repression-negative protein. We tested this idea by replacing the asparagine at position 168 of the linker with proline. The resulting mutant is indeed unable to repress. In vivo, the N168P AraC protein activated transcription to levels of 60% and 110% of fully induced WT AraC in the absence and presence of arabinose, respectively. The behavior of the N168P AraC thus lends support to the hypothesis that the N terminus of the interdomain linker may need to be helical in order for AraC to properly repress transcription.
DISCUSSION
Prior to the work described in this paper, it was unclear whether the interdomain linker plays an active role in modulating AraC's ability to activate transcription or if, instead, the linker functions as a simple unstructured tether connecting the DBD to the DD. Here we have demonstrated that the interdomain linker of AraC does play an active role in regulating transcription from pBAD. It does so primarily by altering the looping preference of AraC but, in the Ind− mutants, also by altering DNA sequence selectivity.
Our new results are somewhat surprising, in light of the earlier work on AraC (7, 9). Previously, it was found that the interdomain linker was relatively tolerant to amino acid substitutions. Roughly the same was found in studies of the AraC homologs RhaR and RhaS (17) and the AraC-like proteins Rns (20) and VirF (23). Studies on RhaR and RhaS found that some alanine substitutions interfered with normal regulation and that swapping the predicted interdomain linkers from RhaR to RhaS produced measurable but modest effects, and it was concluded from those studies that, overall, the linkers did not play a direct role in activating transcription (17). With Rns it was found that replacing the Rns linker with the RhaS linker results in a protein with WT-like behavior (20). A mutagenesis study on the AraC-like protein VirF found that the A154T interdomain linker mutation produced only a small reduction in the ability of the protein to activate transcription, and therefore, it was concluded that the interdomain linker was not critical for the function of VirF (23).
Linker mutations alter the looping preference of AraC.
Mutations in the interdomain linker of AraC could alter the function of the individual domains of AraC or could alter the ability of the two DNA-binding domains to function together either in binding to the I1 and O2 half-sites and forming the DNA loop, which leads to repression, or in binding to the adjacent I1 and I2 half-sites, which leads to induction. The experiments presented in this paper show that linker variants displaying profound regulatory defects were not greatly altered in the ligand binding abilities of either their DD (Table 1) or DBD (Table 2) or in dimerization of AraC. This leaves alterations in the abilities of the two DNA-binding domains to function together as the primary cause of both the induction- and repression-negative phenotypes observed. Because DBD-DBD interactions have not been observed, even at high concentrations (24), the altered behavior of the linker mutations is likely to be a result of altered linker-DBD or altered linker-DD or altered DD-DBD interactions rather than the result of altered DBD-DBD interactions.
As recently reviewed (28), in the current view of the mechanism of AraC unlooping in the presence of arabinose, at least one of the DBDs of AraC must reposition with respect to the DD to allow binding to direct-repeat DNA half-sites, as occurs upon induction at pBAD. This repositioning breaks a DD-DBD interaction present in the loop-favoring conformation of AraC. Consistent with this model, we found that the linker variants behaved differently than WT AraC when binding to direct-repeat half-sites (Tables 3, 5, and 6).
The results of our multiple DNA-binding measurements suggest that the linker variants have lost either their repression or their induction capabilities as a result of an impaired or enhanced ability to maintain the DD-DBD interaction necessary for the loop-favoring conformation of AraC. The repression-negative linker variants would be impaired in their ability to immobilize the DBDs in the loop-favoring positions or orientations. Consequently, less free energy would be required to break the bonds holding them in this state and to shift them from the loop-favoring state to the inducing state. This would be reflected in a tighter binding to direct-repeat half-sites than wild-type protein. Conversely, in the induction-negative linker mutants, the DBDs of AraC would be held more tightly in the repression state and extra energy would be required to break these interactions. The higher-than-normal energy required to break the loop-favoring DD-DBD interactions in the induction-negative mutants would be reflected in weaker than wild-type affinity for binding to direct-repeat half-sites. Table 3 shows that, indeed, these expectations are met.
By altering the strength of the DD-DBD interaction, the linker mutations altered the looping preference of AraC. A more direct comparison of the looping preference of the linker mutant proteins was achieved by determining the relative binding preference for DNAs in direct competition experiments. These experiments also indicated that the loop-favoring DD-DBD interaction has been altered by mutations in the interdomain linker. For example, Table 5 shows that linker mutations have altered the relative amount of direct-repeat I1-I2 DNA bound compared to that of either I1-L24-O2 or I1-L24-I2. This finding supports the interpretation that mutations in the interdomain linker have altered the relative strength of the loop-favoring DD-DBD interaction. The Rep− or Ind− mutations would weaken and the Ind− mutations would strengthen this loop-favoring DD-DBD interaction. Thus, each class of linker mutations would alter the proteins' preference for binding to distally located DNA half-sites and forming a DNA loop that represses transcription relative to binding to direct-repeat DNA half-sites and activating transcription.
In addition to demonstrating their altered looping versus binding to direct-repeat half-sites, our experiments also revealed that the induction-negative mutant proteins are also somewhat altered in their DNA sequence selectivity (Table 6). Because the linker is not intimately included in the structure of the DBD, it is hard to imagine how its alteration could directly affect the structure of the DNA-contacting region of the domain. It therefore seems more likely that the selectivity effects are indirect and are the result of altered DD-DBD interactions that slightly distort the DBD or partially occlude the DNA-contacting surface of the DBD.
A potential mechanism for the Rep− AraC mutants.
The newly substituted amino acids in the Rep− AraC mutants tended to be helix breakers, whereas this class of amino acids was rare in variants that behaved like WT AraC (Fig. 2). This property suggested that the last helix of the DD may extend into the interdomain linker and that this helical extension is important for maintaining the loop-favoring repression state. If this idea were correct, then introducing a single proline substitution (N168P) at the beginning of the interdomain linker should produce Rep− (constitutive) behavior. Indeed, we found that N168P AraC was Rep−. The failure to find N168P in previous extensive searches for constitutive mutants (3) likely results from the fact that mutating an Asn to Pro or to Gly requires two nucleotide changes.
Ending the helix earlier, as suggested above, could increase the effective length of the linker and therefore decrease the effective concentration of one domain in the presence of the other. Alternatively, altering the amino acid composition in the region could alter the flexibility of the linker. In either case, the result would be a weakening of the DD-DBD interaction between the DBD and the DD, which would make it more difficult for the Rep− mutant protein to maintain the loop-favoring state.
Conclusions.
The data presented in this paper demonstrate that mutations in the linker did not significantly affect the function of either the DD or the DBD to bind to its respective ligand. Instead, the effect of the linker mutations appeared when both of the DNA-binding domains of AraC were required to act together. Although it has not been rigorously demonstrated by the experiments presented here, it appears most likely that the linker mutations that were studied primarily lead to an alteration of the interactions between the DD and DBD and, for the Ind− mutants, secondarily alter the DNA sequence selectivity. The simplest explanation for all the observations is that mutations in the linker alter the flexibility, length, or structure of the interdomain linker, and this, in turn, modulates the strength of the DD-DBD interaction. The altered strength of the DD-DBD interaction in the linker mutants would primarily affect the ability of AraC to reposition its DBDs, and this, in turn, alters the ability of the linker mutants to maintain the repressing conformation or adopt the inducing conformation in vivo. The interdomain linker of AraC can no longer be considered a simple, passive tether. Instead, the interdomain linker plays a more active role in regulating the repressing and inducing activities of AraC.
ACKNOWLEDGMENTS
We thank Stephanie Cole for developing the half-site affinity method and for assistance and comments on the manuscript. We thank Michael Rodgers and Katie Frato for developing the purification process for AraC protein.
We also thank NIH (grant GM018277) for the support that allowed the initial isolation of the mutants.
Footnotes
Published ahead of print on 12 August 2011.
REFERENCES
- 1. Bustos S. A., Schleif R. F. 1993. Functional domains of the AraC protein. Proc. Natl. Acad. Sci. U. S. A. 90:5638–5642 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2. Carra J. H., Schleif R. F. 1993. Variation of half-site organization and DNA looping by AraC protein. EMBO J. 12:35–44 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3. Dirla S., Chien J. Y., Schleif R. 2009. Constitutive mutations in the Escherichia coli AraC protein. J. Bacteriol. 191:2668–2674 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4. Dunn T. M., Hahn S., Ogden S., Schleif R. F. 1984. An operator at −280 base pairs that is required for repression of araBAD operon promoter: addition of DNA helical turns between the operator and promoter cyclically hinders repression. Proc. Natl. Acad. Sci. U. S. A. 81:5017–5020 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5. Englesberg E., Squires C., Meronk F., Jr 1969. The l-arabinose operon in Escherichia coli B-r: a genetic demonstration of two functional states of the product of a regulator gene. Proc. Natl. Acad. Sci. U. S. A. 62:1100–1107 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6. Englesberg E., Irr J., Power J., Lee N. 1965. Positive control of enzyme synthesis by gene C in the l-arabinose system. J. Bacteriol. 90:946–957 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7. Eustance R. J., Schleif R. F. 1996. The linker region of AraC protein. J. Bacteriol. 178:7025–7030 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8. Eustance R. J., Schleif R. F. 1996. In vivo association of protein fragments giving active AraC. Proteins 25:501–505 [DOI] [PubMed] [Google Scholar]
- 9. Eustance R. J., Bustos S. A., Schleif R. F. 1994. Reaching out. Locating and lengthening the interdomain linker in AraC protein. J. Mol. Biol. 242:330–338 [DOI] [PubMed] [Google Scholar]
- 10. Frato K. E., Schleif R. F. 2009. A DNA-assisted binding assay for weak protein-protein interactions. J. Mol. Biol. 394:805–814 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11. Gill S. C., von Hippel P. H. 1989. Calculation of protein extinction coefficients from amino acid sequence data. Anal. Biochem. 182:319–326 [DOI] [PubMed] [Google Scholar]
- 12. Hahn S., Dunn T., Schleif R. 1984. Upstream repression and CRP stimulation of the Escherichia coli l-arabinose operon. J. Mol. Biol. 180:61–72 [DOI] [PubMed] [Google Scholar]
- 13. Harmer T., Wu M., Schleif R. 2001. The role of rigidity in DNA looping-unlooping by AraC. Proc. Natl. Acad. Sci. U. S. A. 98:427–431 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14. Hendrickson W., Schleif R. 1985. A dimer of AraC protein contacts three adjacent major groove regions of the araI DNA site. Proc. Natl. Acad. Sci. U. S. A. 82:3129–3133 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15. Hendrickson W., Schleif R. F. 1984. Regulation of the Escherichia coli l-arabinose operon studied by gel electrophoresis DNA binding assay. J. Mol. Biol. 178:611–628 [DOI] [PubMed] [Google Scholar]
- 16. Huo L., Martin K. J., Schleif R. 1988. Alternative DNA loops regulate the arabinose operon in Escherichia coli. Proc. Natl. Acad. Sci. U. S. A. 85:5444–5448 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17. Kolin A., Jevtic V., Swint-Kruse L., Egan S. M. 2007. Linker regions of the RhaS and RhaR proteins. J. Bacteriol. 189:269–271 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18. Lee D. H., Schleif R. F. 1989. In vivo DNA loops in araCBAD: size limits and helical repeat. Proc. Natl. Acad. Sci. U. S. A. 86:476–480 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19. Lobell R. B., Schleif R. F. 1990. DNA looping and unlooping by AraC protein. Science 250:528–532 [DOI] [PubMed] [Google Scholar]
- 20. Mahon V., Smyth C. J., Smith S. G. 2010. Mutagenesis of the Rns regulator of enterotoxigenic Escherichia coli reveals roles for a linker sequence and two helix-turn-helix motifs. Microbiology 156:2796–2806 [DOI] [PubMed] [Google Scholar]
- 21. Martin K., Huo L., Schleif R. F. 1986. The DNA loop model for ara repression: AraC protein occupies the proposed loop sites in vivo and repression-negative mutations lie in these same sites. Proc. Natl. Acad. Sci. U. S. A. 83:3654–3658 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22. Martin K. J., Schleif R. F. 1987. Equilibrium DNA-binding of AraC protein. Compensation for displaced ions. J. Mol. Biol. 195:741–744 [DOI] [PubMed] [Google Scholar]
- 23. Porter M. E., Dorman C. J. 2002. In vivo DNA-binding and oligomerization properties of the Shigella flexneri AraC-like transcriptional regulator VirF as identified by random and site-specific mutagenesis. J. Bacteriol. 184:531–539 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24. Rodgers M. E., Schleif R. 2009. Solution structure of the DNA binding domain of AraC protein. Proteins 77:202–208 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25. Rodgers M. E., Schleif R. 2008. DNA tape measurements of AraC. Nucleic Acids Res. 36:404–410 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26. Ross J. J., Gryczynski U., Schleif R. 2003. Mutational analysis of residue roles in AraC function. J. Mol. Biol. 328:85–93 [DOI] [PubMed] [Google Scholar]
- 27. Saviola B., Seabold R., Schleif R. F. 1998. Arm-domain interactions in AraC. J. Mol. Biol. 278:539–548 [DOI] [PubMed] [Google Scholar]
- 28. Schleif R. 2010. AraC protein, regulation of the l-arabinose operon in Escherichia coli, and the light switch mechanism of AraC action. FEMS Microbiol. Rev. 34:779–796 [DOI] [PubMed] [Google Scholar]
- 29. Schleif R. 2003. AraC protein: a love-hate relationship. Bioessays 25:274–282 [DOI] [PubMed] [Google Scholar]
- 30. Schleif R. F., Wensink P. C. 1981. Practical methods in molecular biology. Springer-Verlag, New York, NY [Google Scholar]
- 31. Seabold R. R., Schleif R. F. 1998. Apo-AraC actively seeks to loop. J. Mol. Biol. 278:529–538 [DOI] [PubMed] [Google Scholar]
- 32. Sheppard D. E., Englesberg E. 1967. Further evidence for positive control of the l-arabinose system by gene araC. J. Mol. Biol. 25:443–454 [DOI] [PubMed] [Google Scholar]
- 33. Soisson S. M., MacDougall-Shackleton B., Schleif R., Wolberger C. 1997. Structural basis for ligand-regulated oligomerization of AraC. Science 276:421–425 [DOI] [PubMed] [Google Scholar]
- 34. Soisson S. M., MacDougall-Shackleton B., Schleif R., Wolberger C. 1997. The 1.6 Å crystal structure of the AraC sugar-binding and dimerization domain complexed with d-fucose. J. Mol. Biol. 273:226–237 [DOI] [PubMed] [Google Scholar]
- 35. Steffen D., Schleif R. 1977. Overproducing AraC protein with lambda-arabinose transducing phage. Mol. Gen. Genet. 157:333–339 [DOI] [PubMed] [Google Scholar]
- 36. Timmes A., Rodgers M., Schleif R. 2004. Biochemical and physiological properties of the DNA binding domain of AraC protein. J. Mol. Biol. 340:731–738 [DOI] [PubMed] [Google Scholar]
- 37. Weldon J. E., Rodgers M. E., Larkin C., Schleif R. F. 2007. Structure and properties of a truely apo form of AraC dimerization domain. Proteins 66:646–654 [DOI] [PubMed] [Google Scholar]
- 38. Wu M., Schleif R. 2001. Mapping arm-DNA-binding domain interactions in AraC. J. Mol. Biol. 307:1001–1009 [DOI] [PubMed] [Google Scholar]
- 39. Wu M., Schleif R. 2001. Strengthened arm-dimerization domain interactions in AraC. J. Biol. Chem. 276:2562–2564 [DOI] [PubMed] [Google Scholar]
- 40. Zhang X., Schleif R. 1998. Catabolite gene activator protein mutations affecting activity of the araBAD promoter. J. Bacteriol. 180:195–200 [DOI] [PMC free article] [PubMed] [Google Scholar]