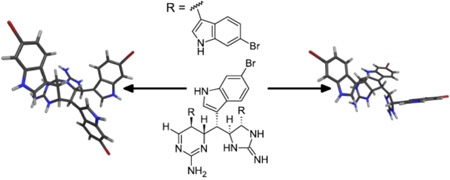
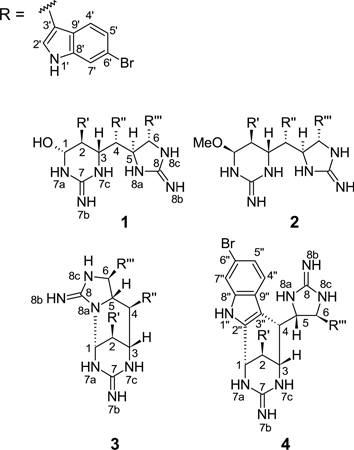
Abstract
Four new tris-bromoindole cyclic guanidine alkaloids, araiosamines A–D, were isolated from the methanol extract of a marine sponge, Clathria (Thalysias) araiosa collected from Vanuatu. Their carbon skeletons delineate a new class of indole alkaloids apparently derived from a linear polymerization process involving a carbon-carbon bond formation. Comparison of the structures including the relative configurations suggests a common intermediate containing a dihydroaminopyrimidine moiety capable of undergoing various modalities of conjugate addition to yield unprecedented ring systems.
Introduction
Two oft-occurring motifs in non-peptidic alkaloids, both marine and terrestrial, are the presence of indole rings or of cyclic guanidines. The high prevalence of indole rings can be understood from their versatility in both binding interactions (through hydrogen bonding, aromatic stacking, hydrophobic, or dipolar interactions) and as a scaffold for chemical modification due to the incorporation of a heteroatom in the aromatic system. Although marine alkaloids containing two indole rings are fairly common,1 tris-indole alkaloids are far less frequently encountered though there are examples from sponges,2,3 ascidians,4 and marine bacteria.5 Properties of cyclic guanidines possibly contributing to their prevalence in natural products include a delocalized charge system and their tendency to form compact and rigid rings. In particular, a [4.3.0]-bicyclic core incorporating five-membered and six-membered guanidine rings features prominently in the paralytic toxin saxitoxin.6 The structural diversity of guanidines (cyclic or otherwise) and the variety of their biological activities have been recently reviewed.7
Marine sponges from the genus Clathria have been reported to produce many different classes of natural products including carotenoids,8 5-thio-D-mannose,9 brominated lipid amides,10 terpenoids such as the antiviral sulfated sterol clathsterol,11 ceramides,12 the cytotoxic microcionamides,13 and various alkaloids. Among the alkaloids are the pseudoanchynazines,14 unusual pteridine-containing indole alkaloids, the cyclic guanidine-containing mirabilins,15,16 and various batzelladine congeners.17 A specimen of Clathria (Thalysias) araiosa Hooper and Levi, 1993 was collected from Vanuatu whose extracts displayed mass spectral peaks indicative of multiple halogenations and the occurrence of a large number of nitrogen atoms. Subsequent isolation and identification of the relevant compounds led to the discovery of a new family of alkaloids incorporating three bromoindole moieties and both five- and six-membered cyclic guanidines. These compounds, whose structures suggest an unusual linear multimerization of tryptamine-like subunits via carbon-carbon bond formation, have been named araiosamines A–D (1–4).
Results and Discussion
Araiosamine A (1) was obtained as a white amorphous solid. A characteristic cluster of mass spectral isotope peaks with an intensity ratio of 1:2:2:1 at m/z 792 is consistent with the presence of three bromine atoms, according well with a formula of C32H29N9OBr3 derived from HRMS ([M + H]+ m/z 794.0034; calcd for C32H29N9O79Br281Br, 794.0025). This formula was supported by the combined 1H and 13C NMR data which indicated two quaternary heteroatom-bound sp2 carbons, 12 quaternary aromatic carbons, 12 aromatic methines, four heteroatom-bound aliphatic methines, and two aliphatic methines adjacent to aromatic systems. 2D [15N, 1H] HSQC was used to identify three types of protonated nitrogens (Table 1): three downfield (δN 134–137 ppm, δH 11.08–11.39 ppm), four at medium range (δN 84.6–104 ppm, δH 8.33–9.42 ppm), and two relatively upfield (δN 70–72 ppm, δH 6.95–7.95 ppm). The 1H and 15N shifts of the furthest downfield group were consistent with the presence of indole rings.18 Closer examination of the NMR data revealed three identical disubstituted indole rings with substituents at ring positions 2 or 3 and at 5 or 6. The latter was suggested by the occurrence of two doublet and one singlet 1H NMR signals for the benzenoid hydrogens. The presence of substituents at positions 3 and 6 in all rings was confirmed by strong COSY correlations between the exchangeable indole hydrogen and the pyrrole-bound hydrogen and by NOESY correlations from the exchangeable indole hydrogen to both the pyrrole-bound hydrogen and the benzenoid singlet hydrogen (Table 1). This substitution pattern was confirmed for all indole rings by HMBC (Table 1). Attachment of the three bromine atoms to position 6 of each of the indole rings was supported by the relatively high-field shift of the corresponding carbons (δC 114.1, 114.1, and 114.3 ppm, for C-6’, C-6”, and C-6”’, respectively, vs. 130 ppm expected with aliphatic substitution;19 Table 1). The presence of indole moieties in 1 is also consistent with a strong UV absorption at 288 nm (ε = 7800).
Table 1.
NMR data for compound 1 recorded at 600 MHz in DMSO-d6.
| Posn. | δH mult (J in Hz) |
δC | δN | COSY | NOESY | [13C, 1H] HMBC |
[13C, 1H] HMBCa |
|---|---|---|---|---|---|---|---|
| 1 | 4.86 dd (7.9, 7.5) | 77.2 CH | 2, 7a, 1-OH | 3, 7a, 1-OH, 4’(w)c | 3(w)c, 7, 3’ | ||
| 2 | 2.61 dd (10.2, 7.5) | 40.7 CH | 1, 3 | 1-OH, 2’(w)c | 3, 3’ | 1, 3, 4(w)c, 2’, 3’ | |
| 3 | 4.62 d (10.2) | 51.3 CH | 2, 4, 7c | 1, 4, 5, 6, 7c, 8a, 2’, 4’ | 1, 2, 3” | ||
| 4 | 3.24 d (9.5) | 42.0 CH | 3, 5 | 3, 6(s)c, 8a(w)c, 2”(vw)c, 4” | |||
| 5 | 4.43 dd (9.5, 3.2) | 63.1 CH | 4, 6, 8a | 3, 7c, 8a, 2” | 6, 8, 3”, 3”’ | ||
| 6 | 4.38 d (3.2) | 56.0 CH | 5,8c | 3, 4(s)c, 8c, 2”(vw)c, 2”’ | 8, 2”’ | 4, 5, 8, 2”’, 3”’ | |
| 7a | 8.36 s | 103.7 | 1(vw) | 1, 7b, 1-OH | 2 | ||
| 7b | 6.95 s | 72.0 | 7a, 7c | ||||
| 7 | –b | ||||||
| 7c | 8.33 s | 84.6 | 3 | 3, 5, 7b, 8a, 2” | 2 | ||
| 8a | 9.42 s | 89.3 | 5,8c | 3, 4(w)c, 5, 7c, 8b | 5, 6, 8 | ||
| 8b | 7.95 s | 70.6 | 8a, 8c | ||||
| 8 | 157.6 C | ||||||
| 8c | 8.46 s | 93.0 | 6,8a | 6, 8b | 5, 6, 8 | ||
| 1-OH | 6.28 d (7.9) | 1 | 1, 2(w)c, 7a | 1, 2 | |||
| 1’ | 11.36 br s | 134.9 | 2’ | 2’, 7’(w)c | 3’, 8’, 9’ | ||
| 2’ | 6.99 br s | 125.5 CH | 1’ | 2(w)c, 3, 1’ | 3’, 8’, 9’ | ||
| 3’ | 109.9 C | ||||||
| 4’ | 7.28 d (7.8) | 120.6 CH | 5’ | 1(w)c, 3, 5’ | 9’ | ||
| 5’ | 7.07 d (7.8) | 121.3 CH | 4’,7’ | 4’ | 7’ | 7’, 9’ | |
| 6’ | 114.1 C | ||||||
| 7’ | 7.63 d (2.2) | 114.2 CH | 5’ | 1’ | 5’, 6’, 9’ | 5’, 6’, 9’ | |
| 8’ | 137.4 C | ||||||
| 9’ | 125.2 C | ||||||
| 1” | 11.39 s | 137.0 | 2” | 2”, 7”(w)c | 3”, 9” | ||
| 2” | 7.13 s | 125.3 CH | 1” | 4(vw)c, 5, 6(vw)c, 7c, 1” | 3”, 8” | 3”, 8”, 9” | |
| 3” | 107.4 C | ||||||
| 4” | 6.74 d (8.6) | 119.9 CH | 5” | 4, 5” | 6”, 8” | ||
| 5” | 6.93 d (8.6) | 121.3 CH | 4” | 4” | 7” | 7”, 9” | |
| 6” | 114.1 C | ||||||
| 7” | 7.61 s | 113.9 C | 1”(w)c | 5”, 6”, 9” | 5”, 6”, 9” | ||
| 8” | 136.3 C | ||||||
| 9” | 127.3 C | ||||||
| 1”’ | 11.08 br s | 134.0 | 2”’ | 2”’, 7”’(w)c | 3”’, 8”’, 9”’ | ||
| 2”’ | 6.54 d (1.9) | 123.2 CH | 1”’ | 6, 1”’ | 3”’, 8”’, 9”’ | 3”’, 8”’, 9”’ | |
| 3”’ | 114.4 C | ||||||
| 4”’ | 6.56 d (8.8) | 119.3 CH | 5”’ | 5”’ | 3”’, 8”’ | 6”’, 8”’, 9”’ | |
| 5”’ | 6.97 d (8.8) | 121.3 CH | 4”’,7”’ | 4”’ | 7”’, 9”’ | 7”’, 9”’ | |
| 6”’ | 114.3 C | ||||||
| 7”’ | 7.47 s | 114.1 CH | 5”’ | 1”’(w)c | 5”’, 6”’, 9”’ | 5”’, 6”’, 9”’ | |
| 8”’ | 137.2 C | ||||||
| 9”’ | 123.2 C |
Correlations observed in CD3OD at 600 MHz.
Not observed in 2D [13C, 1H] HMBC in DMSO-d6.
s- strong; w- weak; vw- very weak.
The six aliphatic methines were sequentially connected to one another using COSY correlations suggesting a linear spin system spanning positions 1–6 (Table 1). Attachment of C-3’, C-3”, and C-3”’ of the three indole rings to positions 2, 4 and 6, respectively, was confirmed by HMBC correlations in methanol-d4 from H-1 to C-3’, from H-2 to C-2’ and C-3’, from H-3 to C-3”, from H-5 to C-3” and C-3”’, and from H-6 to C-2”’ and C-3”’ (Table 1). Carbons C-1, C-3, C-5, and C-6 exhibited low field signals (δC 77.2, 51.3, 63.1, and 56.0 ppm, respectively; Table 1) consistent with attachment to heteroatoms. This was further confirmed by observation of COSY correlations for each of H-1, H-3, H-5, and H-6 to nitrogen-bound protons at δH 8.36, 8.33, 9.42, and 8.46 ppm, respectively (H-7a, H-7c, H-8a, and H-8c in Table 1). In addition H-1 exhibited a COSY correlation with an exchangeable proton at δH 6.28 ppm (1-OH; Table 1) which exhibited no correlations in the 2D [15N, 1H] HSQC spectrum, suggesting that the corresponding hydrogen is attached to the sole oxygen atom in the molecule. The presence of a hemiaminal at position 1 is further supported by a low-field shift for C-1 (δC 77.2 ppm; Table 1) and a large 1JC–H for C-1/H-1 of 162 Hz (Table S2). The molecular fragments described thus far account for all heavy atoms except two carbons and two nitrogens and for 18 of the 22 units of unsaturation. This suggested that the nitrogen atoms attached to C-1, C-3, C-5, and C-6 must be bridged to one other by cyclic guanidine groups, consuming the last atoms and units of unsaturation. The presence of a guanidine group connecting N-8a and N-8c was supported by both NOESY and HMBC. Each of H-8a and H-8c (δH 9.42 and 8.46 ppm, respectively; Table 1) exhibited NOESY correlations with H-8b (δH 7.95 ppm) which in turn exhibited a 2D [15N, 1H] HSQC correlation with δN 70.6 ppm (Table 1). Furthermore, HMBC correlations in DMSO-d6 to C-8 (δC 157.6 ppm; Table 1) from H-8a, H-8c, and H-6 suggested that C-8 is bound to both N-8a and N-8c. This in turn suggested that the last remaining guanidine group included N-7a and N-7c, forming a six-membered ring and completing structure 1. This was further supported by mutual NOESY correlations from both H-7a and H-7c to H-7b at δH 6.95 ppm (Table 1). The occurrence of N-7a in a guanidine group was further confirmed by an HMBC correlation from H-1 to C-7 (δC 155.4 ppm; Table 1) in methanol-d4.
Due to the planar nature of the guanidine group, the six-membered cyclic guanidine appears to adopt a half-chair conformation. Furthermore H-1, H-2, and H-3 all have axial locations based on the large values of 3JH-1–H-2 and 3JH-2–H-3 (7.5 and 10.2 Hz, respectively; Table 1) and the presence of a strong NOESY correlation between H-1 and H-3 (Table 1). A moderately small value of 3JH-5–H-6 at 3.2 Hz (Table 1) suggested a trans substitution pattern for the five-membered cyclic guanidine, which likely has a planar conformation in which the bond C-5/C-6 adopts an eclipsed rotamer (Figure 1F). This substitution pattern was further supported by the presence of a strong NOESY correlation between H-4 and H-6 (Table 1), whereas H-4 and H-5 are anti to one another based on a 3JH-4–H-5 of 9.5 Hz (Table 1). Thus, although the relative configurations could be established independently for the groups 1/2/3 and 5/6, a more detailed analysis was necessary to relate the configurations of each of these groups to one another and to that of position 4. There are four relative configurations that can be obtained by combining the three groups 1/2/3, 4, and 5/6 that were each judged by their consistency with the scalar coupling and NOESY data.
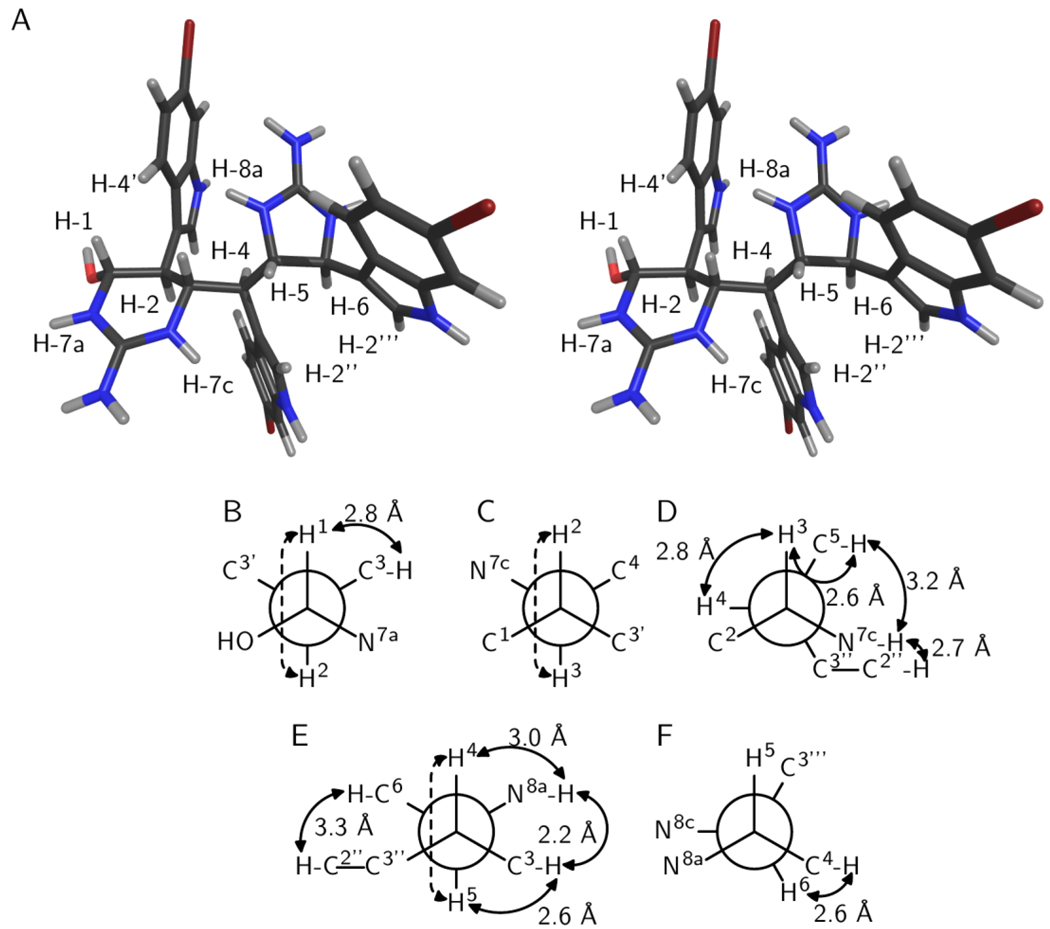
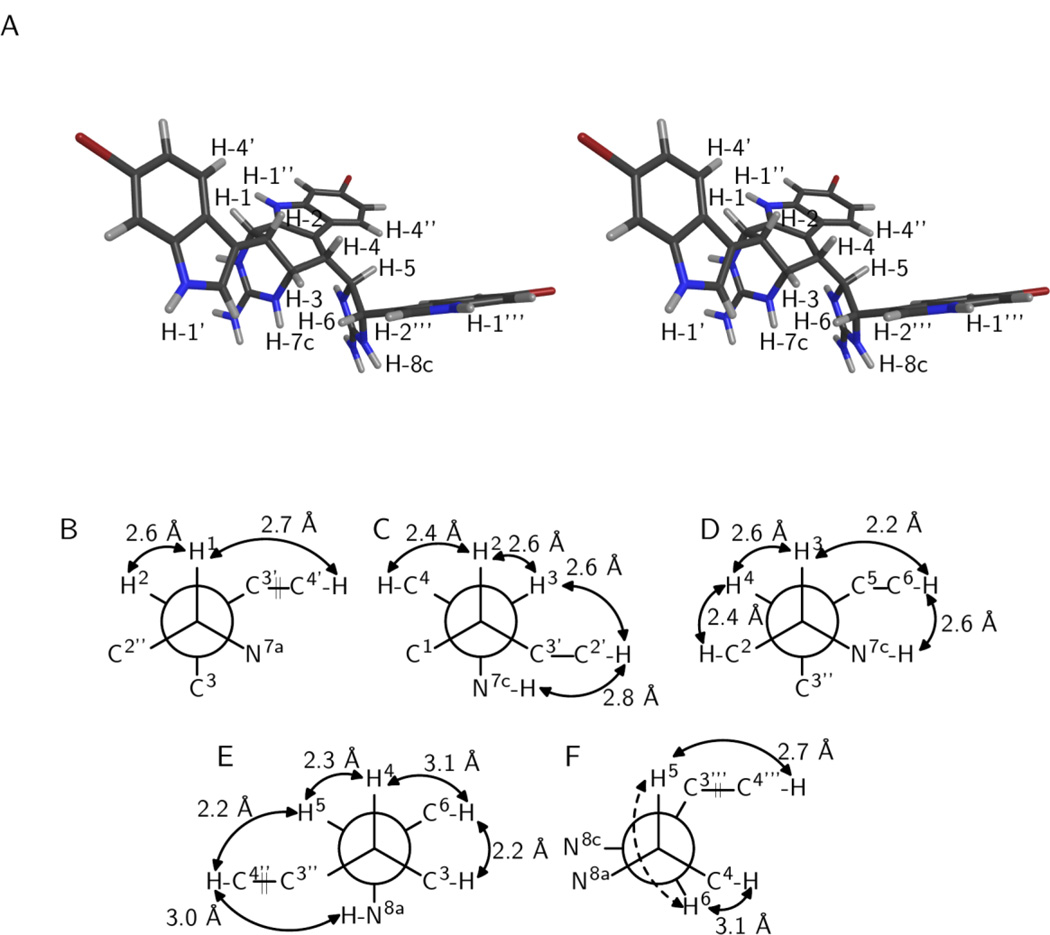
Figure 1.
Relative configuration of araiosamine A (1). (A) Cross-eyed stereogram of a low energy conformer that satisfies the experimental restraints. (B–F) Newman projections along the backbone bonds in the low energy conformer depicted in (A). Solid double-ended arrows indicate experimental NOE correlations were observed for the indicated pair. Distances in the model are shown by each arrow. Dashed arrows indicate 3JH–H > 8 Hz for the indicated pair. (B) Projection for bond C-1/C-2. (C) Projection for bond C-2/C-3. (D) Projection for bond C-3/C-4. (E) Projection for bond C-4/C-5. (F) Projection for bond C-5/C-6.
The bond C-3/C-4 adopts a conformation that places H-3 gauche to H-4 as evidenced by 3JH-3–H-4 < 3 Hz (Table 1) and a strong NOESY correlation for H-3/H-4 (Table 1; Figure 1D). The presence of reasonably strong NOESY correlations for H-5/H-7c and for H-5/H-3 (Table 1; Figure 1D) suggests that C-5 is gauche to both N-7c and H-3, respectively. The presence of a NOESY correlation for H-7c/H-2” (Table 1; Figure 1D) further suggests that C-3” is also gauche to N-7c, leaving H-4 as being anti to N-7c, and fixing the configuration of C-4 relative to C-3. Next considering bond C-4/C-5, a 3JH-4–H-5 of 9.8 Hz (Table 1) indicates that H-5 is anti to H-4 (Figure 1E). The aforementioned NOESY correlations H-3/H-5 and H-5/H-7c indicate that H-5 is gauche to C-3 (Figures 1D, 1E). The presence of a reasonably strong NOESY correlation H-3/H-8a further suggests that N-8a is gauche to C-3, leaving C-6 gauche to H-4 and anti to C-3, fixing the configuration of C-5 relative to C-4 (Figure 1E). Figure 1 illustrates the best conformation and configuration along with key experimental restraints that it satisfies. Thus, the relative configuration of 1 is 1-R*,2-S*,3-S*,4-S*,5-S*,6-S*.
Due to the presence of two freely rotatable bonds at C-3/C-4 and C-4/C-5, there are many possible combinations of relative configuration pairings among the groups C-1/C-2/C-3, C-4, and C-5/C-6 and bond rotamers for C-3/C-4 and C-4/C-5. To further probe whether the experimental observations noted above uniquely defined a configuration and conformation for 1, a series of molecular modeling calculations were carried out for all of the possible diastereomeric pairings of the groups noted above, namely bc (base configuration; 1-R*,2-S*,3-S*,4-R*,5-R*,6-R*), bc-4-ent (1-R*,2-S*,3-S*,4-S*,5-R*,6-R*), bc-5,6-ent (1-R*,2-S*,3-S*,4-R*,5-S*,6-S*), and bc-4,5,6-ent (1-R*,2-S*,3-S*,4-S*,5-S*,6-S*). Note that the experimentally determined configuration described above corresponds to bc-4,5,6-ent. Each base structure was subjected to two different types of conformational search. The first search was based on the use of simulated annealing to generate 200 base structures that were then each submitted to a Monte Carlo conformation search followed by energy minimization. The second search was based on the use of molecular dynamics simulation with implicit solvent at high temperature to sample accessible conformations, followed by selection of unique conformational families based on clustering analysis of the simulation trajectories,20 and energy minimization of the members of each cluster. Low energy conformers within 5 kcal/mol of the minimum energy conformer for each family were pooled together and compared based on backbone conformation (i.e. for carbons C-1 through C-6 and the corresponding cyclic guanidines). For each diastereomer, the lowest energy unique conformers were then subjected to ab initio structure optimization and single point energy calculation to allow for comparison of conformational energetics. The optimized structures for each diastereomer were then compared against the experimental NOE and 3JH–H measurements to identify all diastereomers that have low energy conformers that are consistent with experimental observations (Table S12). Of the four diastereomers, bc-4,5,6-ent was the only one that had conformers satisfying all experimental restraints. The bc-4,5,6-ent conformers satisfying all restraints were either the lowest energy conformer or within 2.2 kcal/mol of the lowest energy conformer depending on whether the PCM implicit solvent correction was excluded or included, respectively (Table S12). Of the other diastereomers, bc contained a conformer which satisfied all but one experimental observation, the NOE between H-7c and H-5, and bc-4-ent had a conformer that satisfied all experimental restraints except the NOE between H-3 and H-8a. The structure depicted in Figure 1A illustrates a low energy conformer of the bc-4,5,6-ent configuration that satisfied the experimental observations.
Araiosamine B (2) had a molecular formula of C33H30N9OBr3 (Δ = +1.4 ppm) as determined by HRMS which differs from the molecular formula of 1 by addition of CH2. Analysis of the NMR spectra for 2 revealed the presence of a methoxy group (1a; δH 3.24, δC 56.1 ppm; Table 2) that exhibited an HMBC correlation to C-1 (δC 83.6 ppm; Table S3), suggesting that 2 is the methyl ether of 1. The remaining features found in 1, namely three 6-bromoindole substituents, the presence of a linear six-carbon chain, and the presence of six-membered and five-membered cyclic guanidines, were all confirmed in 2 by analysis of the NMR spectra and comparsion to those of 1. In 2, however, H-1 is located equatorially although H-2 and H-3 are both located axially, as determined by 3JH-1–H-2 < 3 Hz and 3JH-2–H-3 > 11 Hz. Further evidence supporting an equatorial location for H-1 is provided by a NOESY correlation between H-1 and H-2 (Table S3). Based on similarity in scalar coupling constants and in NOESY correlations, the remaining stereocenters appear to have the same configurations as in 1.
Table 2.
NMR data for compounds 1, 2, 3 and 4 recorded at 600 MHz in DMSO-d6.
| 1 | 2 | 3 | 4 | |||||
|---|---|---|---|---|---|---|---|---|
| Posn. | δH mult (J in Hz) |
δC | δH mult (J in Hz) |
δC | δH mult (J in Hz) |
δC | δH mult (J in Hz) |
δC |
| 1 | 5.09 d (8.3) | 78.7 CH | 4.34 d (3.0) | 83.6 CH | 5.97 d (1.1) | 58.8 CH | 4.92 dd (2.3, 1.3) | 46.6 CH |
| 2 | 2.81 dd (9.6, 8.3) | 42.5 CH | 2.97 d (11.8) | 39.6 CH | 4.29 d (1.1) | 30.0 CH | 3.82 dd (2.7, 1.3) | 34.3 CH |
| 3 | 4.67 dd (9.6, 2.0) | 53.6 CH | 4.83 d (11.8) | 51.3 CH | 4.10 d (10.5) | 54.8 CH | 4.71 d (3.8) | 52.3 CH |
| 4 | 3.52 dd (9.8, 2.0) | 43.9 CH | 3.29 m | 44.6 CH | 3.83 d (10.5) | 48.9 CH | 3.87 dd (3.8, 2.4) | 46.3 CH |
| 5 | 4.60 dd (9.8, 5.5) | 64.9 CH | 4.57 dd (10.3, 5.0) | 65.3 CH | 4.76 dd (10.5, 9.6) | 60.3 CH | 5.23 dd (8.2, 2.4) | 65.1 CH |
| 6 | 4.64 d (5.5) | 58.4 CH | 4.50 d (5.0) | 59.2 CH | 5.11 d (9.6) | 60.6 CH | 5.29 d (8.2) | 56.3 CH |
| 1-OMe | N/A | N/A | 3.24 s | 56.1 CH3 | N/A | N/A | N/A | N/A |
| 7 | 155.4 C | 155.8 C | 153.0 C | 154.7 C | ||||
| 8 | 159.6 C | 159.9 C | –a | –a | ||||
| 2’ | 6.96 s | 126.2 CH | 7.03 s | 127.0 CH | 7.23 s | 124.3 CH | 7.29 s | 123.7 CH |
| 3’ | 110.9 C | 110.7 C | 108.4 C | 112.9 C | ||||
| 4’ | 7.15 d (8.5) | 121.0 CH | 7.20 d (8.5) | 122.1 CH | 7.47 d (8.4) | 120.3 CH | 7.53 d (8.5) | 120.2 CH |
| 5’ | 7.06 d (8.5) | 123.3 CH | 7.05 d (8.5) | 123.4 CH | 7.11 d (8.4) | 123.3 CH | 7.16 d (8.5) | 123.0 CH |
| 6’ | 116.0 C | 116.2 C | 116.3 C | 116.3 C | ||||
| 7’ | 7.60 s | 115.4 CH | 7.61 s | 115.3 CH | 7.52 s | 115.3 CH | 7.54 s | 115.3 CH |
| 8’ | 138.9 C | 139.0 C | 138.3 C | 138.7 C | ||||
| 9’ | 126.2 C | 126.9 C | 125.6 C | 126.3 C | ||||
| 2” | 7.00 s | 125.4 CH | 6.86 s | 126.1 CH | 6.86 s | 123.7 C | 139.3 C | |
| 3” | 108.5 C | 108.5 C | 113.2 C | 106.3 C | ||||
| 4” | 6.75 d (8.5) | 120.6 CH | 6.47 d (8.3) | 120.8 CH | 7.62 d (8.5) | 120.1 CH | 7.38 d (8.6) | 121.6 CH |
| 5” | 6.99 d (8.5) | 123.1 CH | 6.85 d (8.3) | 123.3 CH | 7.15 d (8.5) | 123.2 CH | 7.18 dd (8.4, 1.8) | 123.9 CH |
| 6” | 116.2 C | 116.3 C | 116.1 C | 116.7 C | ||||
| 7” | 7.56 s | 115.3 CH | 7.52 s | 115.2 CH | 7.42 s | 115.2 CH | 7.63 d (1.8) | 115.8 CH |
| 8” | 137.9 C | 138.1 C | 138.3 C | 139.3 C | ||||
| 9” | 128.6 C | 128.7 C | 125.9 C | 125.8 C | ||||
| 2”’ | 6.35 s | 124.4 CH | 6.28 s | 124.4 CH | 6.85 s | 125.6 CH | 7.41 s | 126.2 CH |
| 3”’ | 114.9 C | 115.4 C | 112.2 C | 113.6 C | ||||
| 4”’ | 6.71 d (8.4) | 120.1 CH | 6.59 d (8.6) | 120.4 CH | 7.37 d (8.5) | 120.9 CH | 7.46 d (8.5) | 120.2 CH |
| 5”’ | 6.97 d (8.4) | 123.3 CH | 6.92 d (8.6) | 123.3 CH | 7.02 d (8.5) | 123.1 CH | 7.23 dd (8.5, 1.6) | 123.7 CH |
| 6”’ | 116.0 C | 116.2 C | 115.9 C | 116.6 C | ||||
| 7”’ | 7.38 s | 115.3 CH | 7.38 s | 115.3 CH | 7.40 s | 115.1 CH | 7.61 d (1.6) | 115.9 CH |
| 8”’ | 139.0 C | 139.0 C | 138.8 C | 139.6 C | ||||
| 9”’ | 124.4 C | 124.7 C | 124.8 C | 125.3 C |
Not observed in 2D [13C, 1H] HMBC.
The mass spectrum of araiosamine C (3) exhibited the same isotopic pattern as observed for 1, indicating the presence of three bromine atoms in the molecule. The molecular formula obtained by HRMS, C32H26N9Br3 ([M + H]+ m/z 773.9937; calcd for C32H27N979Br3 773.9939), differed from that of 1 by loss of H2O. Analysis of the NMR data revealed 3 to have the same overall architecture of 1 with a linear chain of six carbons, 6-bromoindole rings attached at C-2, C-4, and C-6, and with 5-membered and 6-membered cyclic guanidine groups. This, combined with the molecular formula, suggested that 3 is similar to 1 but with an additional ring. Since all carbon-bound protons in 1 could be accounted for in 3, this in turn suggested that the additional cyclization occurs via nitrogen. Through combined use of COSY, NOESY, and 2D [15N, 1H] HSQC, it was possible to confirm the adjacency of H-1 and H-7a, of H-3 and H-7c, and of H-6 and H-8c (Table S5). Moreover, NOESY correlations from H-7b to both H-7a and H-7c confirmed their inclusion in a single guanidine group (Table S5). Likewise a NOESY correlation from H-8b to H-8c confirmed that a second guanidine group was present (Table S5). Taken together, the data suggested that N-8a had formed a new bond to carbon. Further evidence supporting this hypothesis came from the 2D [15N, 1H] HMBC acquired in CD3OH/TFA where a correlation was observed from H-8c to an unprotonated nitrogen at δN 107.2 ppm (N-8a; Table S7). Based on the loss of a hydroxyl substituent at C-1 for 3 compared to 1 and the apparent absence of changes in connectivity for all other chain carbons, C-1 was judged the most likely site of attachment for N-8a. Although the 13C NMR shift for C-1 in 3 has shifted upfield relative to the corresponding carbon in 1 (δC 58.8 vs. 78.7 ppm; Table 2), a large measured 1JH-1−C-1 (166 Hz; Table S5) suggests that C-1 in 3 is nevertheless still bound to two heteroatoms. Moreover the presence of a NOESY correlation from H-1 to H-5 supports the proposed cyclization (Table S5).
Of the observable scalar proton-proton coupling interactions in 3, only 3JH-3–H-4 and 3JH-4–H-5 remain essentially unchanged from the corresponding couplings in 1. The presence of a weak COSY correlation between H-1 and H-3 (in methanol-d4; Table S6) suggests a W-coupling interaction best explained if H-1 and H-3 are both located equatorially with respect to the six-membered cyclic guanidine ring. Under such an arrangement, the newly formed ring would be axially fused to the six-membered cyclic guanidine at C-1 and C-3 which is sterically favorable. An equatorial location for H-2 with respect to the cyclic guanidine ring is strongly suggested by a reasonably strong NOESY correlation from H-2 to H-5 (Table S5). Further evidence supporting this arrangement includes a weak NOESY correlation apparent at short mixing time from H-7c to H-2’ on the axially located indole ring (Table S5). Thus, although the ring conformation is inverted in 3 relative to 1, C-1, C-2, and C-3 all have the same configuration relative to one another as observed in 1.
The newly formed ring appears to adopt a boat conformation based on the aforementioned flagpole NOESY correlation from H-2 to H-5. This correlation also makes it possible to fix the configuration of C-5 relative to C-1, C-2, and C-3. The presence of a large 3JH-4–H-5 (10.3 Hz; Table S5) further suggests an anti relationship between H-4 and H-5, placing H-4 axially with respect to the newly formed ring. Additional support for this arrangement comes from NOESY interactions between H-2” on the equatorially located indole ring and both H-2 and H-5 (Table S5; Figure 2A). Finally, a larger 3JH-5–H-6 in 3 compared to 1 (9.3 Hz and 3.2 Hz, respectively; Table S5) admits the possibility of a change in relative configuration between C-5 and C-6 for 3 compared to 1. This does not appear to be the case, however, based on the presence of a reasonably strong NOESY correlation between H-4 and H-6 (Table S5). Given that H-4 is anti to H-5, if H-5 and H-6 were cis to one another then H-6 would be expected to also be located distal to H-4. Instead, the increase in 3JH-5–H-6 might be caused by a change in conformation in the five-membered cyclic guanidine that deviates from planarity and places H-5 and H-6 anti to one another. Such a conformation and configuration is further supported by a NOESY correlation from H-5 to H-4”’ (Table S5; Figure 2F). It should be noted that the configurations for all stereocenters in 3 are identical to those in 1. Figure 2 shows a summary of key experimental correlations supporting the stereochemical assignments for 3.
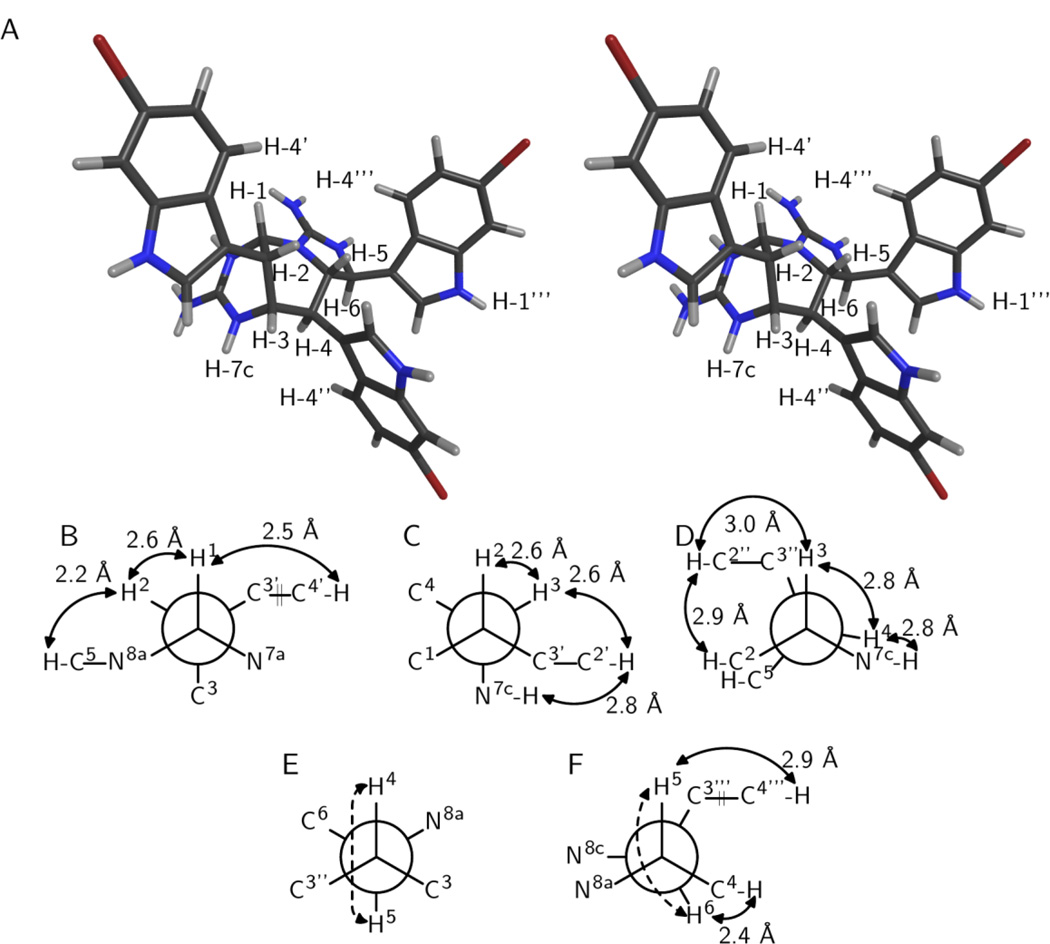
Figure 2.
Relative configuration of araiosamine C (3). (A) Cross-eyed stereogram of a model that satisfies the experimental restraints. (B–F) Newman projections along the backbone bonds in the model depicted in (A). Solid double-ended arrows indicate experimental NOE correlations were observed for the indicated pair. Distances in the model are shown by each arrow. Dashed arrows indicate 3JH–H > 8 Hz for the indicated pair. (B) Projection for bond C-1/C-2. (C) Projection for bond C-2/C-3. (D) Projection for bond C-3/C-4. (E) Projection for bond C-4/C-5. (F) Projection for bond C-5/C-6.
The molecular formula of araiosamine D (4) was suggested to be C32H26N9Br3 by HRMS ([M + H]+ m/z 773.9944, Δ +0.65 ppm), making 4 an isomer of 3. Compound 4 appeared to share the same overall architecture with the other araiosamines. Again, six aliphatic carbons form a linear spin system. The presence of the five-membered and six-membered cyclic guanidines could be confirmed by NOESY correlations from H-7b to both H-7a and H-7c and from H-8b to both H-8a and H-8c (Table S8) combined with COSY correlations H-1/H-7a, H-3/H-7c, H-5/H-8a, and H-6/H-8c (Table S8). Moreover, HMBC in CD3OH/TFA clearly showed correlations from all of H-6, H-8a, and H-8c into C-8 (Table S10). The three 6-bromoindole moieties could also be assigned in the NMR data, although one of the rings had one fewer methine than the others. In addition, the 13C NMR signal for C-1 was shifted significantly upfield compared to 1–3 (Table 2) and 1JH-1–C-1 was much lower than in the other compounds (148 Hz; Table S9), suggesting that C-1 no longer binds to two heteroatoms in 4. The nitrogen-bound hydrogen of one of the three indole rings exhibited no COSY correlations and had a NOESY correlation with H-1 (Table S8), suggesting that C-1 was connected to position 2 of one of the three indoles. Attachment between C-1 and C-2’ could be ruled out based on an HMBC correlation from H-2 to the protonated carbon C-2’ (Table S8). Moreover membership of C-1 in a strained four-membered ring is contraindicated by the small 1JH-1–C-1 mentioned previously. Attachment between C-1 and C-2”’ could be ruled out based on HMBC correlations in methanol-d4 from H-6 to C-2”’, C-3”’, and C-9”’; and from H-2”’ to C-6, C-3”’, and C-9”’ (Table S9). Thus, C-1 was determined to bond to C-2”, forming a six-membered ring in a [3.3.1] bicyclic system.
The pattern of scalar couplings for positions 1–6 was different for 4 compared to 1 with all scalar couplings being below 4 Hz except 3JH-5–H-6 which was 7.7 Hz (Table S8). Again, the COSY acquired in methanol-d4 exhibited a correlation between H-1 and H-3 (Table S9), suggesting that both hydrogens are located equatorially. Evidence for an axial location for the indole ring attached to C-2 was provided by a NOESY correlation from H-7c to H-2’ (Table S8). Despite the similarity in chemical shift of H-2 and H-4 (δH 3.77 and 3.71 ppm, respectively), a ROESY correlation was just resolved from the diagonal in DMSO-d6 (Figure S56), suggesting that H-2 and H-4 have a 1,3-diaxial relationship in the newly formed ring (Figures 3A, 3D). Further confirmation of this arrangement was provided by the presence of a reasonably strong NOESY correlation between H-7c and H-6 (Table S8; Figure 3D) which suggests that Newman projection along bond C-3/C-4 should place C-5 gauche to N-7c rather than anti, thus determining the configuration of C-4 (Figure 3D). H-5 displayed reasonably strong NOESY and ROESY correlations with H-4 and H-4” and a considerably weaker correlation to H-3 (Table S8), suggesting that H-5 is gauche to both H-4 and C-3” when considering the C-4/C-5 bond (Figure 3E). H-6 displays a strong NOESY correlation with H-3 in addition to the aforementioned correlation with H-7c (Table S8), suggesting that C-6 is gauche to C-3 in the projection along the C-4/C-5 bond (Figure 3E). The presence of a NOESY correlation between H-6 and H-4 (Table S8) further suggests that C-6 is gauche to H-4 (Figure 3E). In addition, a NOESY correlation at short mixing time (80 ms) between H-8a and H-4” suggests that N-8a is gauche to C-3” (Figure 3E), thus fixing the configuration of C-5 relative to the rest of the molecule. Finally, 3JH-5–H-6 is somewhat larger in 4 than in 1 (7.7 Hz and 3.2 Hz, respectively; Tables S8 and 1) again raising the question as to whether the change in scalar coupling is due to a change in relative configuration or to a change in ring conformation. Building a molecular model of the two possible configurations of C-6 with all other centers fixed and bond rotations optimized to be consistent with the experimental observations noted above strongly favors a trans substitution pattern for the five-membered cyclic guanidine. With trans substitution, the approximate internuclear distances from H-6 to H-3 and H-7c are 2.1 Å and 2.6 Å, respectively whereas with cis substitution the corresponding distances are 3.7 Å and 4.4 Å, respectively. Thus, the relative configuration for 4 is 1-S*,2-S*,3-R*,4-S*,5-S*,6-S*. Taking into account changes in priority due to the newly formed bond between C-1 and C-2”, the configuration for all centers in 4 are identical to the corresponding centers in 1. Key correlations are highlighted in Figure 3.
Figure 3.
Relative configuration of araiosamine D (4). (A) Cross-eyed stereogram of a model that satisfies the experimental restraints. (B–F) Newman projections along the backbone bonds in the model depicted in (A). Solid double-ended arrows indicate experimental NOE correlations were observed for the indicated pair. Distances in the model are shown by each arrow. Dashed arrows indicate 3JH–H > 8 Hz for the indicated pair. (B) Projection for bond C-1/C-2. (C) Projection for bond C-2/C-3. (D) Projection for bond C-3/C-4. (E) Projection for bond C-4/C-5. (F) Projection for bond C-5/C-6.
The araiosamines represent a new class among indole-containing alkaloids. The aminoimadazoline-containing portion of the araiosamines most closely resemble the discodermindoles21,22 and trachycladindoles,23 both of which were isolated from marine sponges. In the araiosamines, discodermindoles, and trachycladindoles, the 2-aminoimidazoline moieities incorporate carbons equivalent to C-2 and C-3 of tryptophan. The araiosamines differ from these simpler compounds, however, in that position 2 for each indole is not substituted whereas in discodermindole and trachycladindole the corresponding position is substituted with bromine and carboxylate, respectively. In addition, discodermindole and trachycladindole only incorporate one tryptophan equivalent whereas the araiosamines each incorporate three. Furthermore, the polycyclic cores of 3 and 4 are without precedent among synthetic or natural compounds.
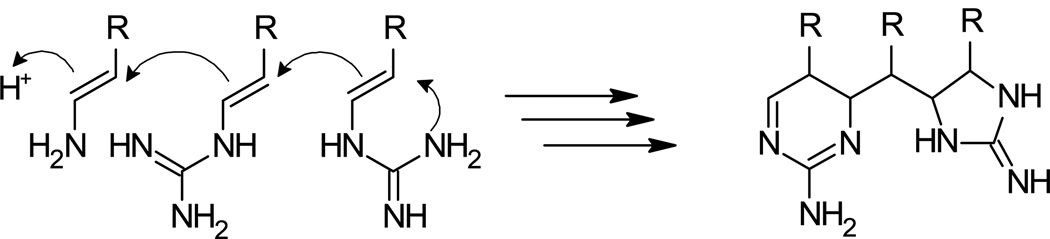
At the core of the araiosamines is a linear precursor formally obtained by bond formation between C-2 of one tryptophan unit and C-3 of a second unit. Although such a formal tryptophan dimer can be found in substructures of other natural products such as yuehchukene (derived from prenyl indole),24 violacein,25,26 the eusynstyelamides,27,28 and various aplysinopsin dimers,29,30 in the other natural products the interunit bonds co-occur with cyclization and are hence limited to dimerization whereas in the araiosamines this linkage appears to be a means of assembling a linear chain. One can envision a few modes of linking the units together such as Claisen condensation of 2-(1H-indol-3-yl)-ethanoic acid thioesters, a chain polymerization of (1H-indol-3-yl)-ethene units potentially initiated by nucleophilic attack of a guanidine group at the equivalent of C-6 (Figure 4), or a stepwise condensation of (1H-indol-3-yl)-ethene derivatives via free radical formation. The second of these possibilities is illustrated in Figure 4 which is essentially a biogenetic route originally proposed for the tachycladindoles23 extended to incorporate multiple (1H-indol-3-yl)-ethene units. The ability to form what are essentially tryptamine oligomers via carbon-carbon bond formation appears to be a novel means of achieving biogenetic diversity.
Figure 4.
Proposed biogenesis for the putative intermediate of the araiosamines.
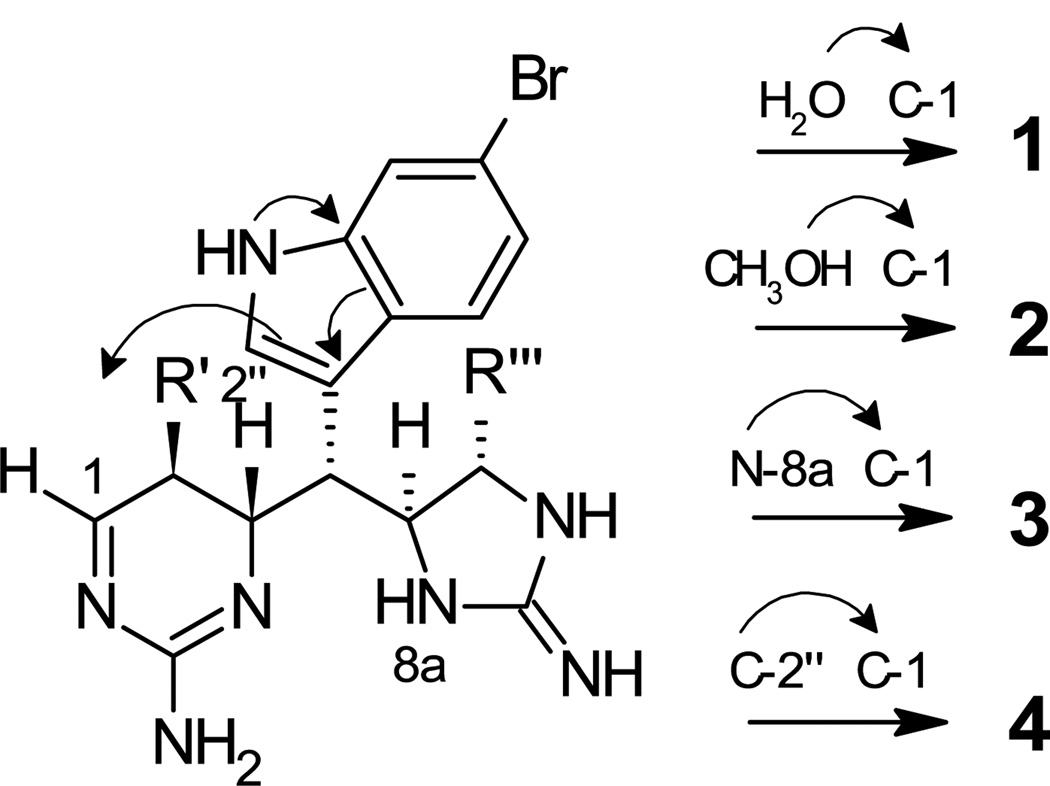
Each of the araiosamines apparently derives from a common precursor in which five of the six chiral centers are defined (Scheme 1). This putative precursor incorporates a dihydroaminopyrimidine ring structure that provides a platform for structural diversification. 1,4-Conjugate addition to C-1 of the precursor by either solvent or an internal nucleophile can account for the structures of 1–4. Such a mechanism provides an elegant means for constructing highly complex cyclic structures from a relatively simple precursor. It should be noted that dihydroaminopyrimidine moieties occur in other compounds isolated from Clathria including some mirabilins16 and some members of the batzelladine family,17 although in neither case is there an imine in direct conjugation with the guanidine group as in the hypothetical precursor in Scheme 1. It is also interesting to note that pseudoanchynazine A isolated from Clathria sp.14 might plausibly derive from a nucleophilic attack by position 2 of an indole ring on a biopterin analogue precursor, not unlike the cyclization proposed to lead to 4 (Scheme 1).
Scheme 1.
Proposed common intermediate for 1–4 and mechanisms for generating each compound. The arrows illustrate how the lone pair electrons on N-1” contribute to the nucleophilicity of C-2”.
The araiosamines were tested for phenotypic effects against zebrafish embryos, for antibacterial activity against Staphylococcus aureus, and in an HIV infection assay but did not exhibit any significant effects.
In conclusion, the araiosamines are exemplary of many of the best features of marine, particularly sponge, natural products. Their structures incorporate simple but distinctive repeated structural motifs such as brominated indole rings and cyclic guanidines, the means of combining biogenetic precursors is innovative and unique, and the apparent assembled precursor to the araiosamines is stereochemically and regiochemically compatible with intramolecular cyclization reactions yielding complex polycyclic scaffolds. Both the sponge species (Clathria (Thalysias) araiosa) and the location it was collected from (Vanuatu) have not been subjects of intensive chemical study, suggesting that focusing on underrepresented organisms and locales remains, even now, a viable route to identifying novel chemical entities.
Experimental Section
General Experimental Procedures
NMR spectra were acquired at 600, 150, and 60 MHz for 1H, 13C, and 15N, respectively, except for the 13C NMR spectra for 3 and 4 which were acquired at 200 MHz. Cryogenically cooled probes were used for all NMR experiments with a sample temperature of 26 °C. 1H and 13C NMR signals were referenced to residual solvent signals at 3.30 and 49.0 ppm, respectively, for methanol-d4 and CD3OH/TFA or at 2.50 and 39.5 ppm, respectively, for DMSO-d6. All HPLC purifications were carried out using HPLC grade solvents and UV detection. Semi-preparative scale purification was performed using a Phenomenex Luna C18 column (5 µm, 10 × 250 mm). Analytical scale purification was carried out using an Agilent Eclipse Plus C18 column (3.6 µm, 4.6 × 150 mm).
Animal Material
The specimen of Clathria (Thalysias) araiosa Hooper & Levi, 1993 was collected from Aore Island, Vanuatu (Sanma Province) in 2008. The collection site was located at S 15° 34.736’ and E 167° 08.221’. A voucher specimen (Van08-13-093) is maintained at the University of Utah, Department of Medicinal Chemistry, Salt Lake City, Utah, USA.
Extraction and Isolation
Frozen sponge material (448 g) was exhaustively extracted with MeOH (3 × 500 mL) to afford 8.84 g crude extract. A portion of this crude extract (5.06 g) was subjected to a solvent partition scheme to yield CHCl3, EtOAc, n-BuOH and aqueous extracts. The CHCl3 extract (214.5 mg) was further partitioned between MeOH and hexane. The MeOH soluble material was subjected to semi-preparative reversed phase HPLC chromatography using a gradient of 35–50% CH3CN in 0.5% TFA over 45 min to yield eight fractions (F1 through F8). Fraction F5 was subjected to semi-preparative reversed-phase HPLC using isocratic elution with 42% CH3CN in 0.5% TFA followed by two successive rounds of analytical reversed-phase HPLC (40% CH3CN in 0.5% TFA for both separations) to afford 1.0 mg araiosamine A (1). Araiosamine B (2) (1.1 mg) was obtained by semi-preparative reversed-phase HPLC of fraction F7 (42% CH3CN in 0.5% TFA) followed by analytical reversed-phase HPLC (40% CH3CN in 0.5% TFA). Fraction F8 was subjected to analytical reversed-phase HPLC (45% CH3CN in 0.5% TFA) to yield 1.1 mg araiosamine C (3). Fraction F6 was subjected to analytical reversed-phase HPLC (40–50% CH3CN in 0.5% TFA over 20 min) to provide araiosamine D (4) (0.8 mg).
Araiosamine A (1): white amorphous powder; [α]24D +20.6 (c 0.18, MeOH); UV (MeOH) λmax (ε) 228 (47000), 288 nm (7800); IR (film) νmax 3272, 1700, 1683, 1600, 1204, 1139, 802, 725 cm−1; 1H and 13C NMR data in Table 1; (+) HRMS with ESI source and TOF mass analyzer: [M + H]+ m/z 794.0034 (calcd for C32H29N9O79Br281Br, 794.0025, Δ +1.1 ppm).
Araiosamine B (2): white amorphous powder; [α]24D +7.3 (c 0.22, MeOH); UV (MeOH) λmax (ε) 228 (17000), 286 nm (3300); IR (film) νmax 3239, 2924, 1682, 1444, 1201, 1140, 1026, 841, 802, 720 cm−1; 1H and 13C NMR data in Table 2; (+) HRMS with ESI source and TOF mass analyzer: [M + H]+ m/z 808.0192 (calcd for C33H31N9O79Br281Br, 808.0181, Δ +1.4 ppm).
Araiosamine C (3): white amorphous powder; [α]24D +25 (c 0.1, MeOH); UV (MeOH) λmax (ε) 226 (48000), 286 nm (8800); IR (film) νmax 3116, 2922, 1672, 1616, 1558, 1540, 1200, 1134, 801, 724 cm−1; 1H and 13C NMR data in Table 2; (+) HRMS with ESI source and FT mass analyzer: [M + H]+ m/z 773.9937 (calcd for C32H27N979Br3 773.9939, Δ −0.3 ppm).
Araiosamine D (4): white amorphous powder; [α]24D +5.7 (c 0.2, MeOH); UV (MeOH) λmax (ε) 228 (32000), 288 nm (6400); IR (film) νmax 3174, 2916, 1676, 1445, 1197, 1137, 801, 721 cm−1; 1H and 13C NMR data in Table 2; (+) HRMS with ESI source and FT mass analyzer: [M + H]+ m/z 773.9944 (calcd for C32H27N979Br3, 773.9939, Δ +0.6 ppm).
Molecular Modeling
Four different diastereomers of 1 were generated that represented all combinations of the three groups for which relative configuration could be unambiguously determined from the scalar coupling and NOE data (C-1/C-2/C-3, C-4, and C-5/C-6, respectively): bc (base configuration; 1-R*,2-S*,3-S*,4-R*,5-R*,6-R*), bc-4-ent (1-R*,2-S*,3-S*,4-S*,5-R*,6-R*), bc-5,6-ent (1-R*,2-S*,3-S*,4-R*,5-S*,6-S*), and bc-4,5,6-ent (1-R*,2-S*,3-S*,4-S*,5-S*,6-S*). The initial structures for each diastereomer were generated using the Avogadro molecular editor (version 1.01)31 with minimization using the MMFF94 force field.32 Two parallel strategies were pursued for generating conformers for each of the four diastereomers noted above. The first strategy made use of the Tinker suite of molecular mechanics programs version 5.033 that was adapted to implement the MMFF94 potential function.34 For each diastereomer, 200 conformers were generated by simulated annealing followed by Monte Carlo torsion angle search. Simulated annealing was carried out using the ANNEAL program with an initial temperature of 2000 K and a final temperature of 0 K with 2×105 steps used both for equilibration and linear cooling. The step size was 1 fs and a unity multiplier was used for atomic masses. Monte Carlo conformational search on the annealed structures was carried out in torsional space using the MONTE program. For each annealed structure, 300 iterations were used. Steepest descent energy minimization was applied at a simulated temperature of 298 K to an RMS gradient of 0.01 kcal/mol. For each structure generated by simulated annealing, only the lowest energy conformer found by Monte Carlo search was retained for analysis. For each diastereomer, the 200 structures generated were then clustered into families by RMSD comparison of the heavy atoms making up the backbone (C-1 through C-6 and the cyclic guanidine groups). The lowest energy member of each family was analyzed for congruence with the experimental data. The second strategy to generate conformers used the AMBER molecular dynamics package.35 The initial structures, with both guanidines protonated, were parameterized with the general AMBER force field36 using the Antechamber tool. Charges were determined using the AM1-BCC method. Improper torsion parameters were added to the guanidinium groups to improve rigidity. Molecular dynamics simulations of the parameterized diastereomers were carried out with PMEMD using the generalized Born solvation model (igb=1) and an infinite nonbonded cutoff. Temperature was regulated using randomly seeded Langevin dynamics with a collision frequency of 1 ps−1. Bonds involving hydrogen atoms were constrained using the SHAKE algorithm. A time step of 1 fs was used and trajectory coordinates were recorded every picosecond. Initially the structures were minimized for 1000 steps using the steepest descent method, followed by 1000 steps of conjugate gradient minimization. The structures then were smoothly heated from 10 K to 600 K over the course of 1 ns. Final production runs at 600 K lasted 50 ns. The conformers generated during the production simulations were sorted using the cluster analysis features in Ptraj.20 Using a subset of every tenth frame, the trajectories of each diastereomer were split into 10 clusters using the average linkage algorithm with the RMS distance metric. Each frame of each cluster was then minimized (900 steps steepest descent, 100 steps conjugate gradient) and an average energy was obtained for each cluster.
For each of the four diastereomers, a representative set of conformers was chosen that covered all of the different backbone conformations observed. These were then subjected to ab initio structure optimization and single point energy calculation to allow comparison of the relative energy level of each conformer for the various diastereomers. These calculations were performed using the Gaussian09 program.37 Geometry optimization was performed first at the HF/3-21G level, followed by DFT optimization and frequency calculation at the B3LYP/6-311G(d,p) level. Optimization to an energy minimum was confirmed by the absence of negative frequencies. Single point energies of the optimized structures were computed at the MP2/6-311G(d,p) level, both with and without the Polarizable Continuum Model (using methanol as the solvent). Each optimized conformer was then analyzed for how well it adhered to experimental observations.
Supplementary Material
Acknowledgment
This work was supported by NIH CA36622 (C.M.I) and NIH GM54501590 (T.E.C.). Funding for the Varian NMR spectrometers was provided by NSF grant DBI-0002806 and NIH grants RR06262 and RR14768. We thank the University of Colorado Denver School of Medicine NMR Core Facility for data acquisition. The UC Denver NMR facilities are funded by the University of Colorado Comprehensive Cancer Center and the UC Denver Program in Structural Biology and Biophysics. The 800 MHz NMR spectrometer was purchased with funds provided by the W.M. Keck Foundation. We would also like to acknowledge computer time from the NSF TeraGrid TG-MCA01S027 and the Center for High Performance Computing at the University of Utah.
Footnotes
Supporting Information Available: NMR data and complete assignments for 1–4 recorded in various solvents; CD spectra in methanol; models used for structure analysis. This material is available free of charge via the Internet at http://pubs.acs.org.
References
- 1.Gul W, Hamann MT. Life Sci. 2005;78:442–453. doi: 10.1016/j.lfs.2005.09.007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Bifulco G, Bruno I, Minale L, Riccio R, Calignano A, Debitus C. J. Nat. Prod. 1994;57:1294–1299. doi: 10.1021/np50111a020. [DOI] [PubMed] [Google Scholar]
- 3.Bifulco G, Bruno I, Riccio R, Lavayre J, Bourdy G. J. Nat. Prod. 1995;58:1254–1260. doi: 10.1021/np50122a017. [DOI] [PubMed] [Google Scholar]
- 4.Foderaro TA, Barrows LR, Lassota P, Ireland CM. J. Org. Chem. 1997;62:6064–6065. [Google Scholar]
- 5.Veluri R, Oka I, Wagner-Döbler I, Laatsch H. J. Nat. Prod. 2003;66:1520–1523. doi: 10.1021/np030288g. [DOI] [PubMed] [Google Scholar]
- 6.Schantz EJ, Ghazarossian VE, Schnoes HK, Strong FM, Springer JP, Pezzanite JO, Clardy J. J. Am. Chem. Soc. 1975;97:1238–1239. doi: 10.1021/ja00838a045. [DOI] [PubMed] [Google Scholar]
- 7.Berlinck RGS, Burtoloso ACB, Trindade-Silva AE, Romminger S, Morais RP, Bandeira K, Mizuno CM. Nat. Prod. Rep. 2010;27:1871–1907. doi: 10.1039/c0np00016g. [DOI] [PubMed] [Google Scholar]
- 8.Tanaka Y, Katayama T. Nippon Suisan Gakkaishi. 1976;42:801–805. [Google Scholar]
- 9.Capon RJ, MacLeod JK. J. Chem. Soc., Chem. Commun. 1987:1200–1201. [Google Scholar]
- 10.Ohta S, Okada H, Kobayashi H, Oclarit JM, Ikegami S. Tetrahedron Lett. 1993;34:5935–5938. [Google Scholar]
- 11.Rudi A, Yosief T, Loya S, Hizi A, Schleyer M, Kashman Y. J. Nat. Prod. 2001;64:1451–1453. doi: 10.1021/np010121s. [DOI] [PubMed] [Google Scholar]
- 12.Xiao D-J, Deng S-Z, Zeng L-M. Zhongshan Daxue Xuebao, Ziran Kexueban. 2002;41:111–114. [Google Scholar]
- 13.Davis RA, Mangalindan GC, Bojo ZP, Antemano RR, Rodriguez NO, Concepcion GP, Samson SC, De G, Dennis, Cruz LJ, Tasdemir D, Harper MK, Feng X, Carter GT, Ireland CM. J. Org. Chem. 2004;69:4170–4176. doi: 10.1021/jo040129h. [DOI] [PubMed] [Google Scholar]
- 14.Zuleta IA, Vitelli ML, Baggio R, Garland MT, Seldes AM, Palermo JA. Tetrahedron. 2002;58:4481–4486. [Google Scholar]
- 15.Capon RJ, Miller M, Rooney F. J. Nat. Prod. 2001;64:643–644. doi: 10.1021/np000564g. [DOI] [PubMed] [Google Scholar]
- 16.El-Naggar M, Conte M, Capon RJ. Org. Biomol. Chem. 2010;8:407–412. doi: 10.1039/b915624k. [DOI] [PubMed] [Google Scholar]
- 17.Laville R, Thomas OP, Berrue F, Marquez D, Vacelet J, Amade P. J. Nat. Prod. 2009;72:1589–1594. doi: 10.1021/np900244g. [DOI] [PubMed] [Google Scholar]
- 18.Martin GE, Hadden CE. J. Nat. Prod. 2000;63:543–585. doi: 10.1021/np9903191. [DOI] [PubMed] [Google Scholar]
- 19.Pretsch E, Clerc T, Seibl J, Simon W. Tables of Spectral Data for Structure Determination of Organic Compounds. 2nd ed. Berlin: Springer-Verlag; 1989. [Google Scholar]
- 20.Shao J, Tanner SW, Thompson N, Cheatham TE., III J. Chem. Theory Comp. 2007;3:2312–2334. doi: 10.1021/ct700119m. [DOI] [PubMed] [Google Scholar]
- 21.Sun HH, Sakemi S. J. Org. Chem. 1991;56:4307–4308. [Google Scholar]
- 22.Cohen J, Paul G, Gunasekera S, Longley R, Pomponi S. Pharm. Biol. (Lisse, Netherlands) 2004;42:59–61. [Google Scholar]
- 23.Capon RJ, Peng C, Dooms C. Org. Biomol. Chem. 2008;6:2765–2771. doi: 10.1039/b803455a. [DOI] [PubMed] [Google Scholar]
- 24.Kong YC, Cheng KF, Cambie RC, Waterman PG. J. Chem. Soc., Chem. Commun. 1985:47–48. [Google Scholar]
- 25.Ballantine JA, Beer RJS, Crutchley OJ, Dodd GM, Palmer DR. Proc. Chem. Soc., London. 1958:232–233. [Google Scholar]
- 26.Laatsch H, Thomson RH, Cox PJ. J. Chem. Soc., Perkin Trans. 2. 1984:1331–1339. [Google Scholar]
- 27.Swersey JC, Ireland CM, Cornell LM, Peterson RW. J. Nat. Prod. 1994;57:842–845. doi: 10.1021/np50108a027. [DOI] [PubMed] [Google Scholar]
- 28.Tapiolas DM, Bowden BF, Abou-Mansour E, Willis RH, Doyle JR, Muirhead AN, Liptrot C, Llewellyn LE, Wolff CWW, Wright AD, Motti CA. J. Nat. Prod. 2009;72:1115–1120. doi: 10.1021/np900099j. [DOI] [PubMed] [Google Scholar]
- 29.Bialonska D, Zjawiony JK. Mar. Drugs. 2009;7:166–183. doi: 10.3390/md7020166. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Dai J, Jiménez JI, Kelly M, Williams PG. J. Org. Chem. 2010;75:2399–2402. doi: 10.1021/jo902566n. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31. http://avogadro.openmolecules.net.
- 32.Halgren TA. J. Comput. Chem. 1996;17:616–641. [Google Scholar]
- 33.Ponder JW, Richards FM. J. Comput. Chem. 1987;8:1016–1024. [Google Scholar]
- 34. http://dasher.wustl.edu/tinker/
- 35.Case DA, Cheatham TE, III, Darden T, Gohlke H, Luo R, Merz KM, Onufriev A, Simmerling C, Wang B, Woods RJ. J. Comput. Chem. 2005;26:1668–1688. doi: 10.1002/jcc.20290. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Wang J, Wolf RM, Caldwell JW, Kollman PA, Case DA. J. Comput. Chem. 2004;25:1157–1174. doi: 10.1002/jcc.20035. [DOI] [PubMed] [Google Scholar]
- 37.Frisch MJ, Trucks GW, Schlegel HB, Scuseria GE, Robb MA, Cheeseman JR, Scalmani G, Barone V, Mennucci B, Petersson GA, Nakatsuji H, Caricato M, Li X, Hratchian HP, Izmaylov AF, Bloino J, Zheng G, Sonnenberg JL, Hada M, Ehara M, Toyota K, Fukuda R, Hasegawa J, Ishida M, Nakajima T, Honda Y, Kitao O, Nakai H, Vreven T, Montgomery J, J A, Peralta JE, Ogliaro F, Bearpark M, Heyd JJ, Brothers E, Kudin KN, Staroverov VN, Kobayashi R, Normand J, Raghavachari K, Rendell A, Burant JC, Iyengar SS, Tomasi J, Cossi M, Rega N, Millam JM, Klene M, Knox JE, Cross JB, Bakken V, Adamo C, Jaramillo J, Gomperts R, Stratmann RE, Yazyev O, Austin AJ, Cammi R, Pomelli C, Ochterski JW, Martin RL, Morokuma K, Zakrzewski VG, Voth GA, Salvador P, Dannenberg JJ, Dapprich S, Daniels AD, Farkas Ö, Foresman JB, Ortiz JV, Cioslowski J, Fox DJ. Gaussian 09, Revision A.1. Wallingford, CT: Gaussian, Inc.; 2009. [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.