Abstract
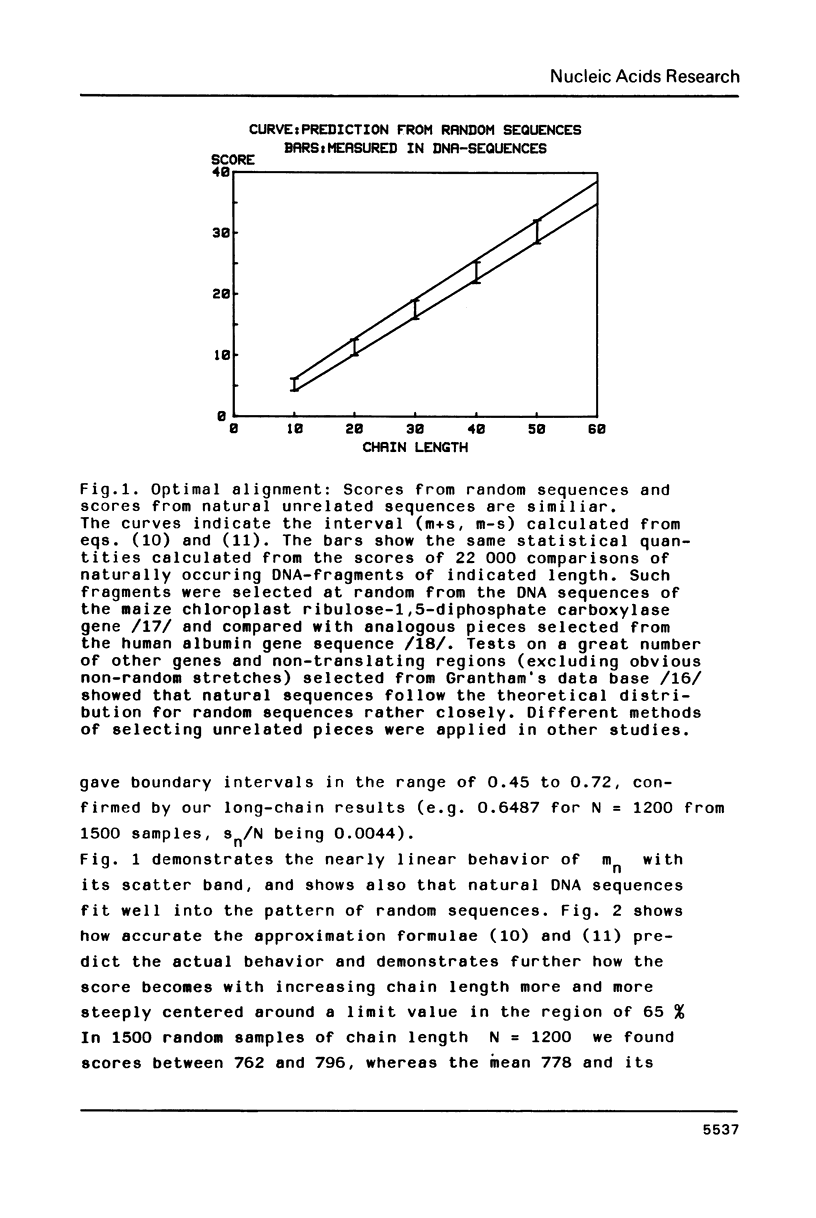
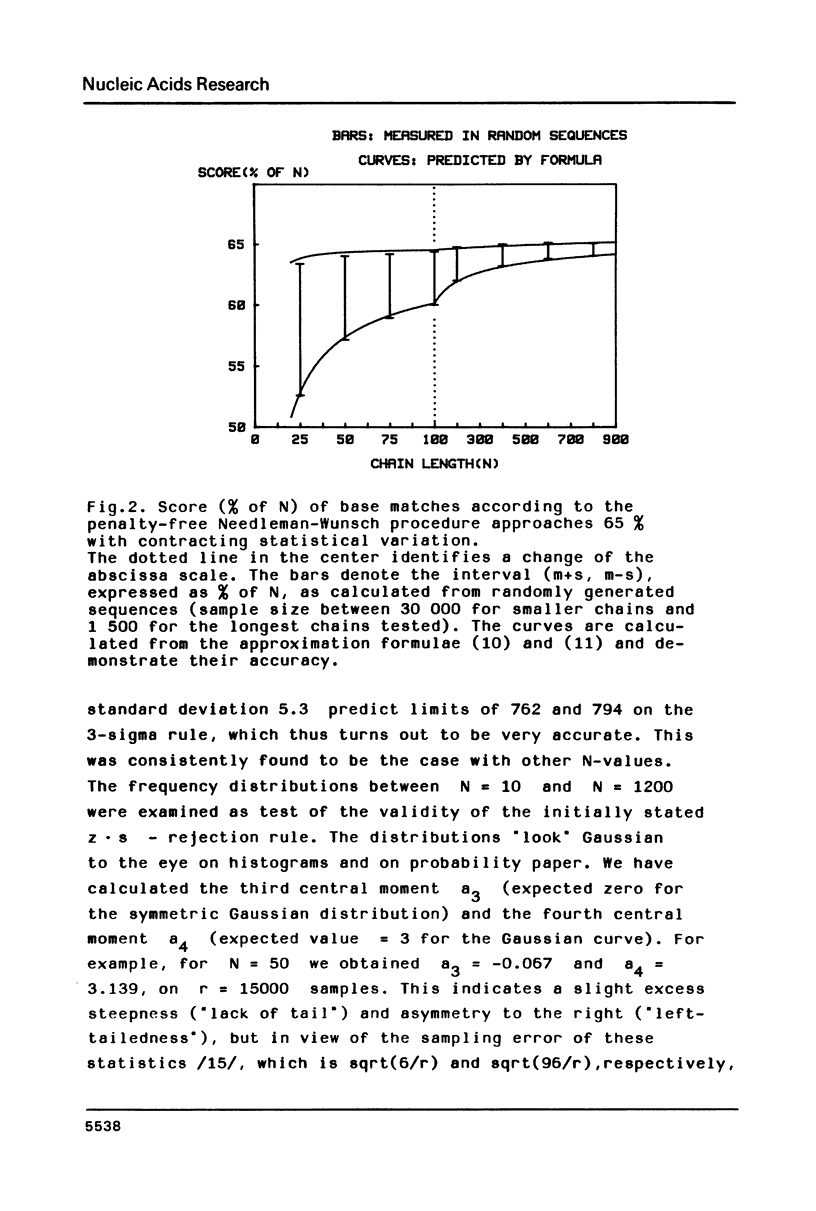
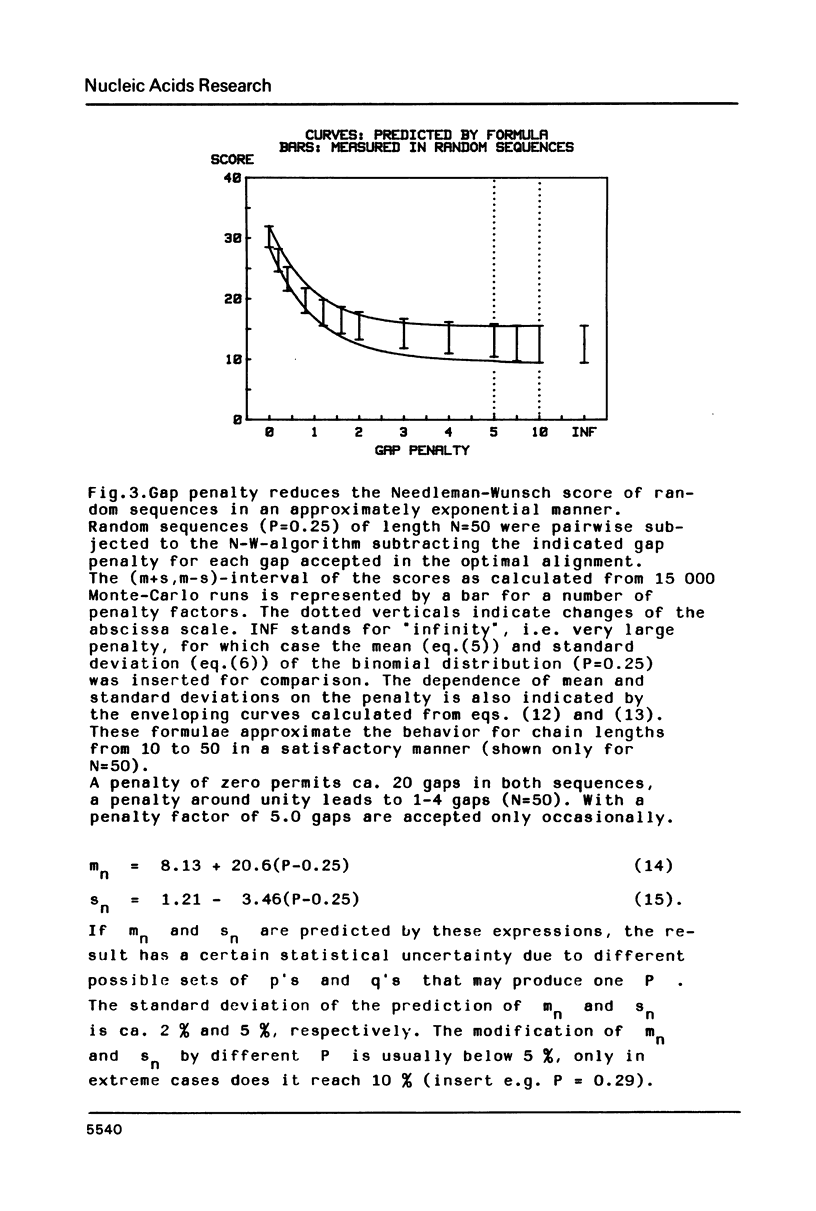
The statistical behavior of the similarity score for unrelated DNA sequences calculated as letter-by-letter comparison or from various forms of optimal alignment was studied. It was found that natural DNA-sequences from a data base and true random sequences show the same statistical behavior in terms of such scores. This makes it possible to adopt a simple criterion for the rejection of fortuitous similarity. It is based on the mean and standard deviation of chance scores whose expected values, depending on chain length, gap penalty and probability of letter coincidence, may be calculated from formulae given in the paper.
Full text
PDF











Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Fitch W. M., Smith T. F. Optimal sequence alignments. Proc Natl Acad Sci U S A. 1983 Mar;80(5):1382–1386. doi: 10.1073/pnas.80.5.1382. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Goad W. B., Kanehisa M. I. Pattern recognition in nucleic acid sequences. I. A general method for finding local homologies and symmetries. Nucleic Acids Res. 1982 Jan 11;10(1):247–263. doi: 10.1093/nar/10.1.247. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hunt L. T., Dayhoff M. O. A surprising new protein superfamily containing ovalbumin, antithrombin-III, and alpha 1-proteinase inhibitor. Biochem Biophys Res Commun. 1980 Jul 31;95(2):864–871. doi: 10.1016/0006-291x(80)90867-0. [DOI] [PubMed] [Google Scholar]
- Lawn R. M., Adelman J., Bock S. C., Franke A. E., Houck C. M., Najarian R. C., Seeburg P. H., Wion K. L. The sequence of human serum albumin cDNA and its expression in E. coli. Nucleic Acids Res. 1981 Nov 25;9(22):6103–6114. doi: 10.1093/nar/9.22.6103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Needleman S. B., Wunsch C. D. A general method applicable to the search for similarities in the amino acid sequence of two proteins. J Mol Biol. 1970 Mar;48(3):443–453. doi: 10.1016/0022-2836(70)90057-4. [DOI] [PubMed] [Google Scholar]
- Sankoff D., Cedergren R. J. A test for nucleotide sequence homology. J Mol Biol. 1973 Jun 15;77(1):169–164. doi: 10.1016/0022-2836(73)90369-0. [DOI] [PubMed] [Google Scholar]
- Sankoff D. Matching sequences under deletion-insertion constraints. Proc Natl Acad Sci U S A. 1972 Jan;69(1):4–6. doi: 10.1073/pnas.69.1.4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Staden R. An interactive graphics program for comparing and aligning nucleic acid and amino acid sequences. Nucleic Acids Res. 1982 May 11;10(9):2951–2961. doi: 10.1093/nar/10.9.2951. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wilbur W. J., Lipman D. J. Rapid similarity searches of nucleic acid and protein data banks. Proc Natl Acad Sci U S A. 1983 Feb;80(3):726–730. doi: 10.1073/pnas.80.3.726. [DOI] [PMC free article] [PubMed] [Google Scholar]



