Abstract
Purpose
One copy of the GALR1 locus on 18q is often deleted and expression is absent in some head and neck squamous cell carcinoma (HNSCC) cell lines. To determine if LOH and hypermethylation might silence the GALR1 gene, promoter methylation status and gene expression were assessed in a large panel of HNSCC cell lines and tumors.
Experimental Design
Promoter methylation of GALR1 in 72 cell lines and 100 primary tumor samples was analyzed using methylation-specific PCR (MSP). GALR1 expression and methylation status were analyzed further by real-time PCR and bisulfite sequencing analysis.
Results
The GALR1 promoter was fully or partially methylated in 38 of 72 HNSCC cell lines (52.7%) but not in the majority 18/20 (90.0%) of non-malignant lines. GALR1 methylation was also found in 38/100 (38%) primary tumor specimens. Methylation correlated with decreased GALR1 expression. In tumors methylation was significantly correlated with increased tumor size (P=0.0036), lymph-node status (P=0.0414), tumor stage (P=0.0037), cyclin D1 expression (P=0.0420), and p16 methylation (P=0.0494) and survival (P=0.045). Bisulfite sequencing of 36 CpG sites upstream of the transcription start site revealed that CpG methylation within transcription factor binding sites correlated with complete suppression of GALR1 mRNA. Treatment with TSA and 5-azacytidine restored GALR1 expression. In UM-SCC-23 cells that have total silencing of GALR1, exogenous GALR1 expression and stimulation with galanin suppressed cell proliferation.
Conclusions
Frequent promoter hypermethylation, gene silencing, association with prognosis, and growth suppression after re-expression support the hypothesis that GALR1 is a tumor suppressor gene in HNSCC.
Keywords: Gene silencing, tumor suppressor gene, human chromosome 18, G-protein coupled receptor, promoter hypermethylation, transcription factor binding sites, methyltransferase inhibitor, histone deacetylase inhibitor
Loss of heterozygosity (LOH) on chromosome 18q in head and neck squamous cell carcinoma (HNSCC) is associated with significantly decreased survival (1). Galanin receptor 1 (GALR1), a G-protein coupled receptor (GPCR) that maps to the commonly lost 18q23 region (2) exhibits aberrant expression in HNSCC cells with 18q LOH. Loss of one copy and inactivation of the remaining GALR1 gene would be consistent with it acting as a tumor suppressor gene. Abnormalities affecting G-protein coupled receptors have been implicated in many human tumors (3-5). Similarly, there is a growing literature implicating a variety of GPCR signaling pathways in head and neck cancer (6-14). GALR1 and its ligand, galanin, are expressed in normal keratinocytes, suggesting that loss of GALR1 plays a role in the development or progression of HNSCC (15). Furthermore, we found two cell lines with GALR1 mutations that affect the sixth transmembrane domain, a region known to affect GPCR function (15).
There are three galanin receptors; GALR1, GALR2 (17q25.3) and GALR3 (22q13.1) (16). Galanin activates the receptors and initiates signal transduction (17). GALR1 is reported to couple to heterotrimeric G-proteins of the Gi type, which inhibit cAMP (16). Galanin regulates many physiological functions in mammals, (15, 18) and may have a role in Alzheimer’s disease (19). GALR2 was reported to activate G12/13 proteins (20) that activate mitogenic cascades. However, galanin receptor function may depend on the cell type. Berger et al. (21) reported that exogenous expression of GALR2 inhibited cell proliferation and induced apoptosis in neuroblastoma cells, whereas in the same cell type expression of GALR1 only inhibited cell proliferation. Thus the functions of GALR1, GALR2 and GALR3, are not well understood. Our recent findings support a growth regulatory function for GALR1 since antibody blockade of this receptor enhances proliferation of on HNSCC cells (8) and restoration of GALR1 expression inhibits cell growth and tumor formation in HNSCC cells (22). Thus, GALR1 appears to act as a tumor suppressor in HNSCC.
Tumor suppressor genes may be inactivated by point mutations, homozygous deletions, or loss of heterozygosity and aberrant methylation (23). Methylation of CpG sites within promoter regions is often associated with silenced gene expression (24, 25). The GALR1 promoter is a TATA-less promoter containing GC-rich sequences that may be susceptible to DNA methylation and gene silencing (26).
In this study, we show for the first time that loss of GALR1 expression is associated with hypermethylation of key CpG sites within transcription factor binding domains and that expression can be restored after treatment with the demethylating agent, 5-Azacytidine and the histone deacetylase inhibitor, Trichostatin A (TSA). Moreover, assessment of primary tumor specimens confirmed that hypermethylation is as common in patient tumors as in cell lines, and is directly associated with tumor size and metastasis. Finally, restoration of GALR1 expression in HNSCC cells resulted growth inhibition in response to galanin stimulation, supporting the hypothesis that GALR1 is a tumor suppressor gene.
MATERIALS AND METHODS
Cell lines
DNA from 72 HNSCC cell lines established from patients at either the University of Michigan (62 UM-SCC) or the University of Turku (10 UT-SCC) was used for MSP analysis. The letter A after the cell line number (e.g. UM-SCC-10A) designates the primary tumor cell lines. Subsequent tumor lines from the same patients have a B designation. Fibroblasts from the original tumor specimen (15 samples) or transformed B-lymphocytes from the tumor cell line donors (3 samples) were used as the source of normal somatic DNA. Nonmalignant cells from the donors of UM-SCC and UT-SCC cell lines; have the same number, e.g. UM-SCC-6 and UM-6F (fibroblasts). Other control cells included normal human keratinocytes (NHK) and HPV16 transformed oral keratinocytes (HOK-16B) cells (a gift from Dr. No Hee Park) (27). cDNA from a normal human brain cDNA library (Invitrogen, Carlsbad, CA) was an additional control.
Tumor Specimens
DNA was isolated from specimens obtained at surgery from 100 primary HNSCC tumors. All patients were treated at the Department of Otolaryngology, Hamamatsu University School of Medicine, Hamamatsu, Japan. Clinical information including age, sex, smoking status, tumor size, lymph node status and stage were obtained from the clinical records. The mean age was 63.9 years (range 39-90), and the male:female ratio was 78:22. Primary tumor sites were: oral cavity (n=34), hypopharynx (n=24), larynx (n=20), oropharynx (n=11), and paranasal cavity (n=11).
Bisulfite Modification
Genomic DNA was extracted using the Wizard Genomic DNA Purification Kit (Promega, Madison, USA). Bisulfite modification of genomic DNA converts unmethylated cytidine residues to uradine residues that are then converted to thymidine during subsequent PCR (28). Methylated cytidine residues are not altered by bisulfite treatment. In brief, 1ug of genomic DNA was denatured with NaOH (final concentration, 0.2M), then incubated with sodium bisulfite (3M; Sigma, St. Louis, MO) (pH 5.0) and hydroquinone (10mM; Sigma, St. Louis, MO) at 55°C for 16 hours. Bisulfite-modified DNA was purified using the Wizard DNA Clean-Up System (Promega, Madison, MO). For DNA desulfonation, NaOH (final concentration, 0.3M) was added, incubated at room temperature for 5 min, ethanol precipitated and resuspended in 100 ul of autoclaved distilled water.
Methylation analysis of the GALR1 proximal promoter
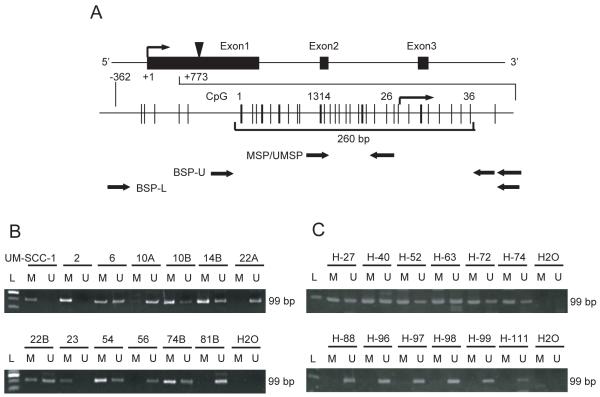
The GALR1 exon structure and the proximal promoter are shown in Figure 1A. An expanded view of the CpG rich proximal promoter and exon 1 extending from −362 to +773bp that includes both the transcription start site (TSS)(bent arrow) and the start codon (black arrow head), is shown below the exon map. The 260 bp region surrounding the TSS has 36 CpG sites. Potential transcription factor binding sites were detected within this region using TFSEARCH. Sites 13 and 14 are part of a consensus Ap-2 site and site 26 is within a consensus Sp1 site.
Figure 1. Diagrammatic representation of the GALR1 gene and its proximal promoter and GALR1 methylation analysis using the MSP assay.
(A) Fig 1A shows the GALR1 exon structure and CpG sites within expanded views of the promoter region relative to the transcription start site. Individual CpG sites are indicated as vertical lines. The bracket at the bottom encloses the 260 bp region that includes the transcription start site and the 36 individual CpG sites that were examined for frequency of methylation. The relative location of the primers used for methylation specific PCR and bisulfite sequencing are shown with black arrows. The transcription start site (TSS) is represented by a bent arrow. The translation start site (ATG) is represented by an arrowhead. (B) Representative examples of MSP of GALR1 in UM-SCC cell lines, showing samples that are fully methylated (UM-SCC-1, -2, -23), partially methylated (UM-SCC-6, -10B, 14B, -22B, -74B), or unmethylated (UM-SCC-10A, -22A, -56, -81B). (C) Representative examples of MSP of GALR1 in primary tumors from Hamamatsu University Hospital, showing samples that are methylated (H-27, H-40, H-52, H-63, H-72, H-74), or unmethylated (H-88, H-96, H-97, H-98, H-99, H-111).
Methylation Specific PCR analysis
Methylation in the region near the TSS was assessed using bisulfite treated DNA PCR amplified with methylation-specific PCR (MSP) primers; MSP-F (5′-GGT TCG CGG TAT TCG GTA GT-3′, upstream) and MSP-R (5′-TCG CCG CCC ACC TCC CGA CTA A-3′, downstream), and unmethylated DNA specific primers (UMSP); UMSP-F (5′-GGT TTG TGG TAT TTG GTA GT-3′, upstream) and UMSP-R (5′-TCA CCA CCC ACC TCC CAA CTA A-3′, downstream). Arrows show the binding locations of the MSP/UMSP primers below the expanded region (Figure 1A). The PCR conditions were 94°C for 5 min; 33cycles at 94°C for 30s, 60.0°C for 30s and 72°C for 40s; and a final extension at 72°C for 5 min. The 99bp PCR products were separated by electrophoresis through a 9% acrylamide gel and stained with ethidium bromide. To analyze the methylation status of the p16 gene and the RASSF1A gene, primers and conditions as described by Herman et al. and Kuroki et al. were used (28, 29).
Bisulfite Sequencing Analysis
The frequency of methylation at 36 individual CpG sites within the bracketed 260bp region was assessed on the upper or lower strands using bisulfite specific sequencing. Of the 36 CpG sites analyzed, numbers 1 to 27are upstream of the TSS and 28 to 36 are downstream (Figure 1A). Bisulfite sequencing PCR (BSP) primer pairs specific for modified upper and lower strand DNAs were designed to contain no CpG sites. Upper strand primers were BSP-U-F (5′-GAG GAG GAA AGG TAT TAA TGG ATG AGG AGG-3′), BSP-U-R (5′-ACT CTT TTA AAA AAC TTC CTA ATA ATA AAT-3′) and BSP-U-RN (5′-AAT ACC AAA AAC TTC TCT ACT ACA ACT AA-3′). In a second PCR reaction BSP-U-F and a nested primer, BSP-U-RN were used with 0.5ul of the PCR product as template. PCR conditions for both the first and second PCR were 94°C for 5 min; 35cycles at 94°C for 30s, 50.8°C for 30s and 72°C for 50s; with a final extension at 72C for 7 min. The lower strand primers were BSP-L-F (5′-AGG TTT TTT GGT GGT GGA TA-3′) and BSP-L-R (5′-ACC AAT TTT ACA AAA CTC CA-3′). The binding locations of the bisulfite sequencing primers is shown by arrows below the expanded region (Figure 1A). The lower strand PCR conditions were 94°C for 5 min; 40cycles at 94°C for 30s, 56.0°C for 30s and 72°C for 50s, and a final extension at 72°C for 5 min. The PCR products were separated by electrophoresis through a 1.5% agarose gel containing ethidium bromide, extracted from the gel with the QIAquick Gel Extraction Kit (QIAGEN, Hilden, Germany), cloned into pGEN T-Easy vector, and at least nine clones were sequenced by the ABI Model 3730 sequencer using T7 or SP6 primers.
RNA extraction and GALR1 RT-PCR
Total RNA from 13 HNSCC cell lines, including 9 with loss of one copy of chromosome 18q, one with no loss of 18q, and three for which 18q status is unknown, was isolated using the RNeasy Mini Kit (QIAGEN, Hilden, Germany). After DNase treatment, cDNA was generated using an oligo(dT)16 primer with Superscript II reverse transcriptase (Invitrogen, Carlsbad, CA). Primer sequences for amplifying the coding region of the human GALR1 cDNA (GenBank Accession AY541036) are as follows: RT-GALR1-sense 5′-CAC TTG CAT AAA AAG TTG AAG-3′ (location in coding sequence; 676-696), and RT-GALR1-antisense 5′-TTA TCA CAC ATG AGT ACA ATT GG-3′ (1053-1031) (30). The PCR product size was 378bp. The PCR conditions were 94°C for 8 min; with either 40 or 45cycles at 94°C for 30s, 52.0°C for 30s and 72°C for 45s; with a final extension at 72°C for 8 min. Primers for the GAPDH gene were described previously (30).
Quantitative reverse transcriptase PCR (Q-PCR) for GALR1
Q-PCR was performed with the ABI PRISM 7700 HT Sequence Detection System (Applied Biosystems, Foster City, CA). Inventoried Assays-on-Demand™ Gene Expression Products (Applied Biosystems, Foster City, CA), which passed quality control manufacturers specifications, were used as primers and probe (Hs00175668_m1). The cDNA was generated from DNase-treated total RNA using Random primers (Invitrogen, Carlsbad, CA) with Superscript II reverse transcriptase (Invitrogen, Carlsbad, CA). For each PCR evaluation, 10ul of diluted cDNA, 12.5ul of TaqMan Universal PCR Master Mix (Applied Biosystems, Foster City, CA), and 1.25ul of Assay Mix were added to a final volume of 25ul. The thermal cycler conditions were as follows: one cycle of preheating at 50°C for 2min, AmpliTaq Gold activation at 95°C for 10min, followed by 50cycles of denaturing at 95°C for 15sec, annealing/extension at 60°C for 60sec. Analysis was performed with ABI prism Sequence Detection System v1.7a software (Applied Biosystems, Foster City, CA) following the manufacturer’s instructions. For comparisons between samples, the mRNA expression of the target genes was normalized to GAPDH mRNA expression.
Immnohistochemistry
Five-um tumor sections were dewaxed with xylene, hydrated through graded alcohols, and rehydrated in water. Sections were microwaved in citrate buffer (pH 6.0) three times for five minutes, then endogenous peroxidase activity was blocked using 0.5% hydrogen peroxide in methanol for 30 minutes. After a 20% goat serum was applied to the sections for 10 minutes, they were incubated with monoclonal antibodies against cyclin D1 protein (1:100, 5D4: IBL. Fujioka, Japan), p53 protein (1:1000, Do7: DAKO. Copenhagen, Denmark) and PTEN (1:100, PN37: InVitrogen. Carlsbad, USA) at 4°C overnight (31).
Reactivation of GALR1 expression
Twelve hours after plating, cultures were incubated either for 48 hours with 5-Azacytidine (15ug/ml, 30ug/ml; Sigma, St Louis, MO), an inhibitor of DNA methyltransferase, or for 24 hours with 300nM Tricostatin A (TSA) (Sigma, St Louis, MO), an inhibitor of histone deacetylase, or for 48 hours with 5-Azacytidine followed by 24 hours with TSA. The medium was then removed and cultures were maintained in standard DMEM, which was replaced every other day for 2-5 days. Transcripts were optimal on day 4 or 5 (32, 33)
Transient transfection, Western blotting and cell proliferation assay
UM-SCC-23, a cell line established from a human laryngeal squamous cell carcinoma, was cultured in Dulbecco’s Modified Eagle Medium (DMEM) (Gibco, Grand Island, NY) supplemented with 10% heat-inactivated fetal bovine serum at 37°C in 5% CO2. The GALR1 sequence was obtained from a human cDNA library (Invitrogen, Carlsbad, CA), C-terminal HA-tagged, and subcloned into the pcDNA3 vector (Invitrogen) containing an internal ribosomal entry site (Ires) and GFP sequence. The pCMVIresGFP vector was used as a transfection control. The UM-SCC-23-GALR1 and UM-SCC-23-mock cells were established by transfecting with pCMVGALR1HAIresGFP or pCMVIresGFP respectively using lipofectamine (Invitrogen), followed by selection for GFP positive cells. Twenty-four hours after plating, stably transfected cells were fed with serum-free medium containing 0.1% BSA (SFM) for 24 hours to induce quiescence. Then, one uM galanin (ANASPEC, San Jose, CA) was added. Cell proliferation was measured by counting cells with a Coulter counter model Z1 (Beckman Coulter Inc., Hialeah, FL). Cells were lysed with 0.1% Nonidet-P 40 lysis buffer containing protease (Calbiochem, La Jolla CA) and phosphatase (Sigma) inhibitor cocktails. To avoid protein aggregation, cells lysates were treated with 1000 Units of N-Glycosidase F or a mock digestion without the enzyme (mock)(New England BioLabs, Beverly, MA), and subjected to electrophoresis without boiling. Equal amounts of protein were electrophoresed on 10% SDS-PAGE gels and transferred to Hybond-P (Amersham Biosciences, Buckinghamshire, United Kingdom). The mouse monoclonal anti-HA tag antibody (CONVANCE, Berkeley, CA) was used to detect exogenous GALR1. GAPDH was detected by mouse monoclonal anti-GAPDH (Chemicom international, Temecula, CA) as an internal control for protein loading. The membranes were incubated overnight with primary antibody at 4°C, followed by incubation with horseradish peroxidase conjugated rabbit anti-mouse secondary antibody (Amersham Biosciences).
Statistical analysis
The association between discrete variables and GALR1 methylation was tested by the Fisher’s exact probability test or the Mann-Whitney U test. In the colony formation assay, comparisons and tests for statistical significance were made by Student’s test. The 5-year overall survival rates were constructed using the method of Kaplan-Meier and analyzed by the log-rank test. Cox’s proportional hazards regression analysis, which involved age, tumor site, smoking status, stage grouping, cyclin D1 expression and GALR1 methylation, was used to identify the multivariate predictive value of the prognostic factors. A significant difference was identified when the probability was less than 0.05.
Results
Methylation Specific PCR
To determine if the GALR1 promoter is affected by methylation in HNSCC, methylation specific PCR analysis was carried out on bisulfite treated DNA from 72 UM-SCC and UT-SCC cell lines as well as from fibroblast, EBV-transformed B lymphocyte and normal keratinocyte samples. In addition, we analyzed bisulfite treated DNA from primary tumors of 100 head and neck squamous cell carcinoma patients in the Hamamatsu University Pathology archives. Forward primers for MSP and UMSP bind in the segment of the promoter that contains CpG dinucleotides 13-15, which correspond to a consensus Ap-2 site. Reverse primers for MSP and UMSP anneal to the segment that includes CpG dinucleotides 25-27 which correspond to a consensus Sp1 site (Figure 1A). These sites are immediately upstream of the TSS. Examples of the methylated and unmethylated PCR products from a representative subset of the cell lines are shown in Figure 1B and the results from all cell lines are summarized in Table 1. The subset included: nine cell lines (UM-SCC-2, -10A, -10B, -14B, -22A, -22B, -23, -54, -81B) previously shown to have LOH affecting the GALR1 locus, one with no 18q loss (UM-SCC-74B), and three (UM-SCC-1, -6, -56), that were not tested for 18q loss. In three cell lines (UM-SCC-1, -2, -23) only methylated alleles were detected (Figure 1B). Five other cell lines (UM-SCC-10B, -14B, -22B, -54 and -74B) had predominantly methylated alleles. In contrast, UM-SCC-81B, -10A, and 22A had only unmethylated alleles (Figure 1B). Of interest, GALR1 is mostly unmethylated in UM-SCC-10A and -22A whereas GALR1 is methylated in UM-SCC-10B and –22B that came from recurrent tumors in the same patients. Among 72 UM-SCC and UT-SCC cell lines tested, the GALR1 promoter was hypermethylated in 14 of 72 (19.4%) cases, partially methylated in 24 (33.3%), and unmethylated in 34 (47.2%). In contrast, 18 of 20 (90.0%) of the nonmalignant samples were unmethylated and only 2 (10.0%) were partially methylated (sTable 1). For comparison we also assessed the methylation status of p16 and RASSF1A, two tumor suppressor genes that are frequently reported to be silenced by methylation. As shown in sTable 1, like GALR1, p16 was fully or partially methylated in 38% of the tumors and RASSF1A in 30%. Among DNA samples from 100 previously untreated primary tumors tested with the same primers (Figure 1C), the GALR1 promoter was methylated in 38 of 100 (38.0%) cases and unmethylated in 62 (62.0%). Methylation of GALR1 significantly correlated with increased tumor size (P=0.0036), lymph node status (P=0.0414), tumor stage (P=0.0037), increased cyclin D1 expression (P=0.0420), p16 methylation (P=0.0494) and overall 5 yr survival (p=0.0448) (Table 1). There was no association with expression of p53, PTEN or RASSAF1. In multivariate analysis, taking into account, age, tumor site, smoking, tumor stage, and cyclin D1 expression, only GALR1 methylation and stage were significant predictors of poor survival (Table 2).
Table 1.
GALR1 Gene Methylation Status in HNSCC Primary Samples.
| Patient and tumor characteristics (n=100) |
Methylation | ||
|---|---|---|---|
| Present(n=38) | Absent (n=62) | P-value | |
| Age | |||
| 70 and older (29) | 9 | 20 | |
| Under 70 (71) | 29 | 42 | 0.2466 a |
| Gender | |||
| Male (78) | 27 | 51 | |
| Female (22) | 11 | 11 | 0.1438 a |
| Smoking status | |||
| Smoker (69) | 26 | 43 | |
| Non smoker (31) | 12 | 19 | 0.6280 a |
| Tumor site | |||
| Oral cavity & Oropharynx(45) | 14 | 31 | |
| Others(55) | 24 | 31 | 0.1407 a |
| Tumor size | |||
| T1 (10) | 2 | 8 | |
| T2 (39) | 9 | 30 | |
| T3 (19) | 10 | 9 | |
| T4 (32) | 17 | 15 | 0.0036 **b |
| Lympho-node status | |||
| N0 (44) | 11 | 33 | |
| N1 (22) | 11 | 11 | |
| N2 (27) | 13 | 14 | |
| N3 (7) | 3 | 4 | 0.0414 * b |
| Stage | |||
| I ( 6 ) | 2 | 4 | |
| II (22) | 3 | 19 | |
| III (17) | 5 | 12 | |
| IV (55) | 28 | 27 | 0.0037 ** b |
| p53 expression | |||
| Mutant type (51) | 20 | 31 | |
| Wild type (49) | 18 | 31 | 0.4804 a |
| PTEN expression | |||
| Low (31) | 13 | 18 | |
| High (69) | 25 | 44 | 0.3720 a |
| cyclin D1 expression | |||
| High (33) | 17 | 16 | |
| Low (67) | 21 | 46 | 0.0420*a |
| p16 methylation | |||
| Yes (54) | 25 | 29 | |
| No (46) | 13 | 33 | 0.0494* a |
| RASSF1A methylation | |||
| Yes (22) | 10 | 12 | |
| No (78) | 28 | 50 | 0.2831 a |
| Over all survival | |||
| (Kaplan-Meier, %) | 36.1 | 57.1 | 0.0448*c |
the Fisher’s exact probability test
the Mann-Whitney U test
the Log-rank test
Table 2.
Multivariate analysis of factors influencing survival, determined by Cox’s proportional hazards model.
| Variable | Relative risk | 95% CI | P-value |
|---|---|---|---|
| Age | |||
| 70 and older vs. <70 | 1.324 | 0.692-2.533 | 0.397 |
| Smoking status | |||
| Smoker vs. Non smoker | 1.127 | 0.560-2.270 | 0.737 |
| Tumor site | |||
| Oral cavity & Oropharynx vs. Others | 0.786 | 0.430-1.435 | 0.433 |
| Stage | |||
| I, II, III vs. IV | 1.995 | 1.044-3.811 | 0.037 * |
| cyclin D1 expression | |||
| High vs. Low | 0.533 | 0.223-1.270 | 0.155 |
| GALR1 methylation | |||
| Yes vs. No | 2.515 | 1.053-6.004 | 0.038 * |
CI : confidence interval
GALR1 expression in 13 UM-SCC cell lines tested for promoter methylation
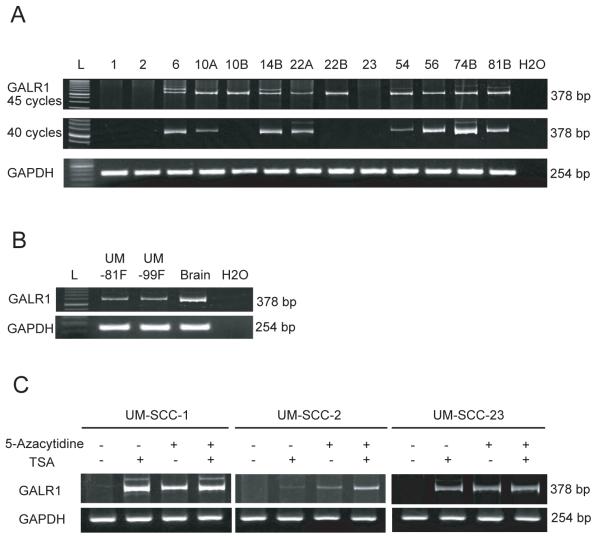
The thirteen UM-SCC cell lines shown in Figure 1 were tested for GALR1 expression using RT-PCR with either 40 or 45 cycles of PCR. Absence of GALR1 expression corresponded well with the methylation status shown in Figure 1. Three cell lines (UM-SCC-1, 2, and 23) had no detectable GALR1 mRNA even after 45 cycles as determined by absence of PCR product with this stringent test for message (Figure 2A, top). cDNA from normal fibroblasts and brain were GALR1 positive (Figure 2B), which is consistent with GALR1 expression in normal cells.
Figure 2. GALR1 expression by RT-PCR.
(A) Representative examples of RT-PCR for GALR1 expression in UM-SCC cell lines. PCR reactions were carried out at 40 and 45 cycles to identify cell lines with low or absent GALR1 expression. The housekeeping gene GAPDH was run as a control for RNA integrity. L, 50-base pair ladder in GALR1 and 100-base pair ladder in GAPDH. (B) Representative examples of RT-PCR for GALR1 expression in nonmalignant samples. (C) The effect of treatment with 5-Azacytidine or Trichostatin A (TSA) or both on GALR1 expression in 3 cell lines with densely methylated GALR1 is shown using RT-PCR. Controls were cells treated similarly but without 5-Azacytidine or TSA.
GALR1 expression after 5-Azacytidine and TSA Treatment
If DNA methylation and the associated deacetylation of histones that frequently accompanies methylation is responsible for silencing GALR1 gene expression, it should be possible to re-express the gene by reversing the epigenetic effects. We treated the three completely silenced cell lines (UM-SCC-1, -2 and -23) with methyltransferase (5-Azacytidine) and/or histone deacetylase (trichostatin A) inhibitors. In UM-SCC-1 and UM-SCC-23, either 5-Azacytidine or TSA was sufficient to induce GALR1 expression. For UM-SCC-2, only slight GALR1 expression was observed when TSA was used alone. Given alone 5-Azacytidine induced some expression, however when both drugs were used together GALR1 expression was more robust. Thus, all three-cell lines exhibited restored GALR1 expression after inhibition of methyltransferase and histone deacetylase (Figure 2C).
Bisulfite Sequencing Analysis
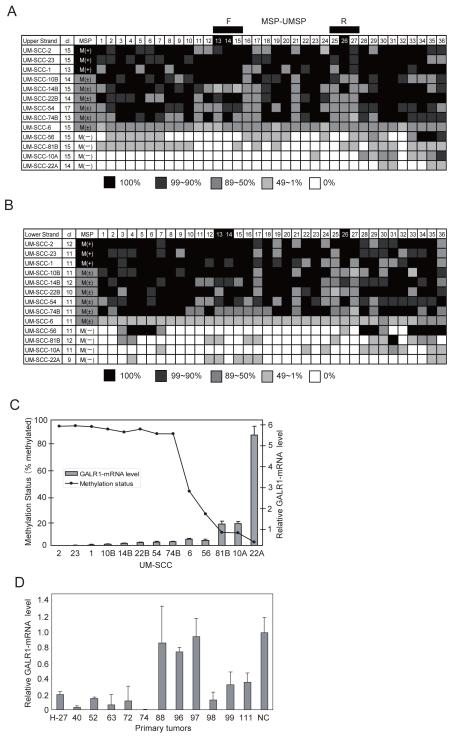
To examine the methylation of individual alleles and their association with GALR1 silencing, we focused on the 36 CpG sites surrounding the TSS (Figure 1A) using bisulfite sequencing PCR with the BSP primers for the upper (BSP-U-F/BSP-U-RN/BSP-U-R) and lower DNA strands (BSP-L-F/BSP-L-R)(as shown in Figure 1A). We sequenced nine or more clones from the upper and lower strands of the DNA from 13 cell lines representing hypermethylated, partially methylated or unmethylated examples. The percentage of methylated alleles for each of the 36 CpG sites, are shown in Figure 3A (top strand) and Figure 3B (bottom strand) for the 13 HNSCC cell lines.
Figure 3. Summary of bisulfite sequencing analysis and Quantitative real-time PCR.
(A) Bisulfite sequencing analysis of the upper strand. (B) Bisulfite sequencing analysis of the lower strand. The shading of each cell within the figure indicates the proportion of alleles that were found to be methylated by methylation specific sequencing. The key is shown below each panel. The numbers in the top row indicate the CpG dinucleotide positions (labeled 1-36 in Figure 1) in the region amplified by the bisulfite sequencing (BSP) primers. The location of the MSP primer binding sites (MSP/UMSP-F and MSP/UMSP-R) used for the data in Figure 3 are indicated by the black boxes above the figure. CpG sites 13, 14 and 26 are shaded in the top line to indicate that these sites are within the consensus transcription factor binding sites. The cell line numbers are shown the left column. The number of clones sequenced (cl) are given in the second column. The column headed MSP summarizes the methylation specific PCR results; M(+) indicates positive for the GALR1-densely methylated form by MSP; M(±), indicates some densely methylated and some unmethylated signal by MSP; M(−), negative for the GALR1-densely methylated form by MSP. (C) Comparison of methylation status and relative mRNA expression as assessed by quantitative RT-PCR of GALR1 in 13 UM-SCC cell lines. (D) Relative GALR1 mRNA expression as assessed by quantitative RT-PCR in 12 tumor specimens that were also evaluated for GALR1 promoter methylation. Specimens H-27, 40, 52, 63, 72, and 74 exhibited promoter methylation but H-88, 96, 97, 99, 111 did not (Figure 1C). NC-normal tissue control.
Methylation Status of Transcription Factor Binding Sites
CpG sites 13 and 14 (upper strand: 5′-GCCCGCGGC-3′, lower strand: 5′-GCCGCGGGC-3′) are within a near-perfect consensus AP-2 (Activating Protein 2) transcription factor binding site upstream of the transcription start site. DNA methylation within AP-2 binding sites is known to decrease the affinity of AP-2 binding (34). CpG site 26 is located within a consensus Sp1 (Selective Promoter Factor 1) binding site (upper strand: 5′- GGGCGG-3′, lower strand: 5′-CCGCCC-3′). Transcription factor binding to Sp1 sites is also known to be significantly decreased by methylation of the first two cytosines on the lower strand of the Sp1 recognition sequence (lower strand: 5′-CCGCCC-3′) (35). The frequency of methylated alleles corresponded to the MSP/UMSP results in Figure 1 and with the amplification of message shown in Figure 2.
Quantitative RT-PCR for GALR1
To quantify message expression in relation to promoter methylation generally and of the transcription factor binding sites real time RT-PCR was used. The three GALR1 negative cell lines, UM-SCC-1, -2, and -23, are methylated on 100% of CpG sites 13, 14, and 26, with only two exceptions; UM-SCC-1 at site 14 (lower strand) had 91% methylated alleles and UM-SCC-2 at site 13 (upper strand) had 93% methylated alleles. A high proportion of these alleles are also heavily methylated in (UM-SCC-10B, -14B, -22B, -54 and –74B), cell lines with barely detectable GALR1 message. In contrast, for the cell lines with more readily detectable message, the proportion of heavily methylated alleles falls below 50% and the level of methylation at sites 13, 14, or 26 (Figure 3A, 3B) also falls sharply with increasing message expression (Figure 3C). In fact, in cell lines with readily detectable GALR1 message expression, CpG sites 13-15 and 25-27 usually have only moderate methylation when compared to the non-expressing hypermethylated cell lines. No GALR1 message could be detected by Q-PCR in cell lines UM-SCC-1, -2, and -23 ( ΔΔCT value; 0.029+0.031, 0.001+0.001, 0.003+0.009)(Figure 3C). The average of the methylated alleles on the upper strand and lower strands in these cells lines was 94.8, 95.0% and 95.4%% respectively (Figure 3A, 3B). UM-SCC-10B, -14B, -22B, -54 and -74B had barely detectable expression of GALR1 by Q-PCR (ΔΔCT value; 0.051+0.023 to 0.182+0.023), and had a high proportion of methylated sites (averages-88.8% to 92.7%). UM-SCC-6, -10A, -22A, -56 and -81B, which express GALR1 expression by RT-PCR and Q-PCR (ΔΔCT value; 0.291+0.06 to 5.278+0.418), exhibited a range of methylation levels ranging from 2.8% to 43.2% of sequenced sites. Thus, promoter methylation showed considerable variability but was consistent with mRNA expression levels (Figure 3C). To determine if GALR1 methylation in tumor specimens also was associated with gene silencing we isolated mRNA from frozen tissue. GALR1 expression analyzed by quantitative RT-PCR (Figure 3D), showed that six tumors H-27, -40, -52, -63, -72, and -74 with strong methylation signals (Figure 1C), exhibited little or no mRNA expression. In contrast, 3/6 (H-88, -96, -97) tumors without GALR1 promoter methylation (Figure 1C) exhibited relatively robust mRNA expression (Figure 3D).
Suppression of cell growth after restoration of GALR1 expression
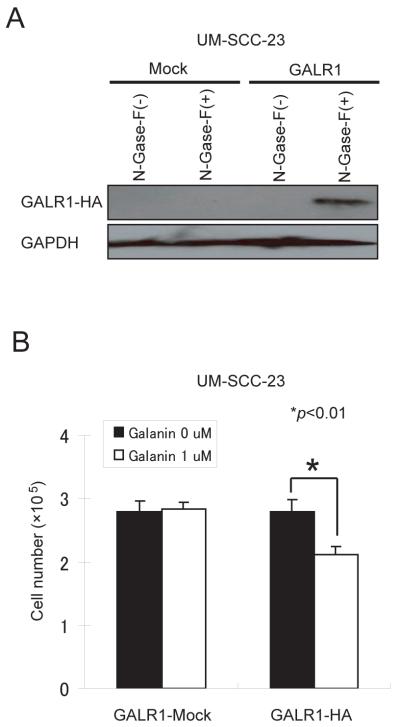
UM-SCC-23 cell line, that exhibits extensive hypermethylation and expresses no GALR1 message, was used for assessment of exogenous GALR1 re-expression and effects on cell proliferation in response to galanin stimulation. UM-SCC-23 cells transfected with wild-type GALR1 in the pCMVIresGFP or pCMVGALR1HAIresGFP constructs were assessed for GALR1 expression in western blot using antibody specific for the HA-tag (Figure 4A). Treatment with 1uM galanin suppressed cell proliferation in UM-SCC-23GALR1HA cells by 24% (p=0.0054) relative to the vector alone transfected cells (Figure 4B). Thus, GALR1 inhibits tumor cell proliferation in response to galanin stimulation.
Figure 4. Transient transfection, Western blotting and cell proliferation assay.
(A) Western blot analysis shows exogenous GALR1 expression in pCMVGALR1HAIresGFP and pCMVIresGFP transfected cells with or without treatment with N-Glycosidase F detected by antibody to the HA tag. (B) Relative colony formation ability for UM-SCC-23-GALR1 and UM-SCC-23-mock cells in response to treatment with galanin. In each case the number of Hygromycin B-resistant cells in vector-transfected controls was set at 100%. Cell proliferation was measured by counting cells with a Coulter counter model Z1 and results are presented as the means±SD (bars) of three separate experiments, each performed in triplicate.
Discussion
Genetic analysis of HNSCC has revealed loss of heterozygosity (LOH) at 3p, 8p, 9p, 11q, 13q and 18q alleles in a significant fraction of tumors (36). 18q LOH is linked to advanced stage and poor survival in HNSCC, suggesting that one or more genes on this chromosome are important in tumor behavior (1). The commonly lost region of 18q is 18q23, which was lost in 53% (D18S461) to 75% (D18S70) includes the GALR1 locus (37). Galanin and its receptors are variably expressed in HNSCC (15).
GALR1 is a G-protein coupled receptor. GPCRs with the typical seven transmembrane spanning domains and their ligands are involved in regulation of cellular physiology and complex behaviors mediated through numerous signal transduction and integration pathways (17, 38). We recently demonstrated that in HNSCC, galanin and GALR1 signal via ERK1/2 activation to inhibit expression of cyclin D1, upregulate expression of the cyclin dependent kinase inhibitors p27kip1 and p57kip2, and inhibit tumor cell proliferation and tumor growth (22). These findings support the concept that GALR1 regulates cell growth and functions as a tumor suppressor gene.
Some tumor suppressor genes exhibit homozygous loss, but in our studies of LOH we found no examples of homozygous loss of the GALR1 locus. Therefore we suspected that an epigenetic mechanism of gene silencing might be at work. In this study, we show that GALR1 expression is frequently absent in head and neck squamous cell carcinomas. Furthermore, expression-negative squamous cancers and cell lines exhibit hypermethylation of CpG islands in the GALR1 promoter region. In a survey of 62 UM-SCC and 10 UT-SCC cell lines using GALR1 promoter methylation specific PCR, we showed that 14 of 72 (19.4%) were completely methylated, and another 24/72 (33.3%) were partially methylated. Thus, more than half of the cell lines (52.7%) exhibit evidence of GALR1 epigenetic silencing. The frequency of promoter methylation of p16 22/72 (30.6%) and RASSF1A 10/72(13.9%), two other tumor suppressor genes, was similar to that of GALR1 in the UM-SCC and UT-SCC cell lines. Hypermethylation of the GALR1 promoter was also found in a similar proportion (38%) of primary head and neck tumor specimens, and suppression of GALR1 expression correlated with promoter methylation. Thus, GALR1 resembles other major TSGs in frequency of aberrant promoter methylation both in vitro and in vivo. Importantly, there is a significant correlation between GALR1 promoter methylation and tumor size, lymph-node status, tumor stage, cyclin D1 expression and p16 methylation status in primary samples. Moreover, in multivariate analysis, only GALR1 methylation and clinical stage were significantly associated with poor survival when age, smoking status, tumor site, and cyclin D1 expression were also considered. This is consistent with our earlier findings that LOH of the GALR1 locus on 18q is associated with poor prognosis (1). Many lines with LOH had little or no GALR1 expression, which is consistent with loss of one allele and silencing of the other. We did find strong GALR1 expression in UM-SCC-74A, a cell line with no LOH. Of interest, LOH was present in primary tumor cell lines from two patients, UM-SCC-10A and UM-SCC-22A, but methylation was low or absent and GALR1 expression was still detectable. However, in the recurrent or metastatic tumor cell lines from these same donors, UM-SCC-10B and UM-SCC-22B, the methylation signal was potent, suggesting that silencing of the remaining allele occurred with progression. The significant association of increased cyclin D1 expression with GALR1 methylation in these tumors is consistent with our previous in vitro study (22) that showed that GALR1 and galanin downregulate cyclin D1 expression. Thus, loss of GALR1 expression appears to be linked to loss of control of cyclin D1 expression which is consistent with tumor progression.
The MSP assay is a convenient and sensitive method to examine promoter methylation status and in this study it was fully concordant with mRNA expression. It became clear that although these cell lines have a strong promoter methylation signal, a critical feature in complete suppression of mRNA expression is whether or not the transcription factor binding sites are hypermethylated. Thus, the choice of the correct primer sets is an essential element for an effective MSP assay and should be designed based on bisulfite treated DNA sequencing and mRNA expression (39, 40) as we did in the current study. The methylation specific and unmethylated DNA specific primers each included 3 CpG sites, which spanned at least two transcription factor binding sites. AP-2 is a cell type-specific transcription factor expressed in neural crest and ectoderm-derived tissues, including craniofacial, skin and urogenital tissues. Although footprinting has shown some promiscuity in AP-2 sites, AP-2 protein homodimers bind consistently to a consensus palindromic sequence, 5′-GCCNNNGGC-3′. In the GALR1 promoter this sequence includes CpG sites 13 and 14 (41, 42). AP-2 has roles in both normal development and the development/progression of cancer (42). Methylation at CpGs within the AP-2-binding site distinguish the UM-SCC cell lines (such as UM-SCC-1, -2, and -23) that fail to express GALR1 mRNA from those such as UM-SCC-14B and -74B that do express GALR1. This is consistent with reports indicating that methylation of AP-2 CpG sites inhibits AP-2 binding and suppresses AP-2-regulated transcription of genes such as SOD2, E-cadherin and the proenkephalin-CAT gene (34, 43, 44). Sp1 sites are found in the promoter region of many housekeeping genes, indicating a role for the Sp1 as a basal transcription factor (45). Some reports indicate that methylation at Sp1 binding sites has no influence on Sp1 binding (35, 46). whereas others (28) used electrophoretic mobility shift assays to show that Sp1 binding to the Rb promoter can be abolished when the first two cytosines on the lower strand (5′-CCGCCC-3′) are methylated (34). Similarly, the methyl-CpG binding domain protein 1 (MBD1) and MBD1-containing chromatin-associated factor (MCAF) complex blocks transcription by affecting Sp1 in heavily methylated promoter regions (47). MCAF physically binds Sp1 and the general transcription apparatus like a positive regulator. MBD1-MCAF seems to interfere with Sp1-mediated activation of the transcription pre-initiation complexes in methylated DNA regions (47). The methyl groups on DNA could interfere with binding of specific transcription factors and been described as a mechanism for transcriptional inactivation of tumor suppressor genes in subsets of primary cancers. Within the CpG islands, there is typically a core promoter and a transcription start site and gene expression is completely repressed when this region becomes hypermethylated (48). Our results show that the methylation levels of CpG sites 13 and 14, on the upper strand in the AP-2 binding site, strongly influences the expression of GALR1 mRNA. In addition, it is likely that heavy methylation of the promoter region also causes inhibition of Sp1 binding and repression of GALR1 mRNA expression.
Furthermore, in 3/3 HNSCC cell lines that do not express any detectable GALR1 mRNA, the gene could be reactivated by treatment with 5-Azacytidine alone, TSA alone and/or the combination of 5-Azacytidine and TSA. Reactivation of heavily methylated cell lines with histone deacetylase inhibitors alone is somewhat unusual and indicates that chromatin silencing effects can be reversed even in the presence of DNA methylation. This also suggests that the combination of methylation and deacetylation act together to silence expression and that variability in DNA acetylation may account for some of the heterogeneity we observed in expression of GALR1 in lines with relatively strong methylation signals. Our data show that in HNSCC promoter hypermethylation of GALR1 is associated with loss of gene expression that can be reversed by treatment with methyltransferase and histone deacetylase inhibitors.
Our results are consistent with the hypothesis that GALR1 is a probable tumor suppressor gene in HNSCC. We demonstrate for the first time that expression of GALR1 mRNA is lost in HNSCC as a consequence of DNA methylation. Furthermore, silencing of the GALR1 gene by methylation may be a critical event in tumor progression of HNSCC, because 18q LOH is associated with tumor progression (36), and in this study GALR1 promoter methylation was observed to be greater in cell lines derived from secondary tumors in the same patient, e.g. UM-SCC-10A has a weaker MSP signal than UM-SCC-10B, and the same occurs in UM-SCC-22A and -22B. Furthermore, GALR1 promoter methylation was associated with larger primary tumor size, clinical stage, lymph node metastasis, cyclin D1 expression and p16 methylation, and poorer survival. In this paper we have shown that transcriptional repression is likely mediated by inhibition of AP-2 and Sp1 binding secondary to methylation of critical CpGs in the GALR1 promoter. Furthermore, re-expression of GALR1 in transfected cells results in galanin-induced inhibition of cell proliferation. Finally, GALR1 can be reactivated by altering chromatin modifications with methyltransferase and histone deacetylase inhibitors raising the promise of selective small molecule inhibitors of these enzymes as potential therapeutic agents in HNSCC.
Supplementary Material
Acknowledgements
We are grateful to Professor Mitsuyoshi Nakao (Department of Regeneration Medicine, Institute of Molecular Embryology and Genetics, Kumamoto University) and Professor Keith Kirkwood (Department of Periodontics and Oral Medicine at the University of Michigan) for helpful advice. This work was supported by NIH NIDCR (5 R01 DE12477)(TEC) and NIH NCI through the University of Michigan Head and Neck SPORE grant (1 P50 CA97248), the Cancer Center Support grant (5 P30 CA46592), and by NIH NIDCD Research Center Core grant (P30 DC05188).
References
- 1.Pearlstein RP, Benninger MS, Carey TE, et al. Loss of 18q predicts poor survival of patients with squamous cell carcinoma of the head and neck. Genes Chromosomes Cancer. 1998;21:333–9. doi: 10.1002/(sici)1098-2264(199804)21:4<333::aid-gcc7>3.0.co;2-#. [DOI] [PubMed] [Google Scholar]
- 2.Takebayashi S, Ogawa T, Jung KY, et al. Identification of new minimally lost regions on 18q in head and neck squamous cell carcinoma. Cancer research. 2000;60:3397–403. [PubMed] [Google Scholar]
- 3.Taborga M, Corcoran KE, Fernandes N, Ramkissoon SH, Rameshwar P. G-coupled protein receptors and breast cancer progression: potential drug targets. Mini reviews in medicinal chemistry. 2007;7:245–51. doi: 10.2174/138955707780059826. [DOI] [PubMed] [Google Scholar]
- 4.Touge H, Chikumi H, Igishi T, et al. Diverse activation states of RhoA in human lung cancer cells: contribution of G protein coupled receptors. International journal of oncology. 2007;30:709–15. [PubMed] [Google Scholar]
- 5.Spiegelberg BD, Hamm HE. Roles of G-protein-coupled receptor signaling in cancer biology and gene transcription. Current opinion in genetics & development. 2007;17:40–4. doi: 10.1016/j.gde.2006.12.002. [DOI] [PubMed] [Google Scholar]
- 6.Cheng Y, Lotan R. Molecular cloning and characterization of a novel retinoic acid-inducible gene that encodes a putative G protein-coupled receptor. J Biol Chem. 1998;273:35008–15. doi: 10.1074/jbc.273.52.35008. [DOI] [PubMed] [Google Scholar]
- 7.Gschwind A, Prenzel N, Ullrich A. Lysophosphatidic acid-induced squamous cell carcinoma cell proliferation and motility involves epidermal growth factor receptor signal transactivation. Cancer Res. 2002;62:6329–36. [PubMed] [Google Scholar]
- 8.Henson BS, Neubig RR, Jang I, et al. Galanin receptor 1 has anti-proliferative effects in oral squamous cell carcinoma. J Biol Chem. 2005;280:22564–71. doi: 10.1074/jbc.M414589200. [DOI] [PubMed] [Google Scholar]
- 9.Lin R, Nagai Y, Sladek R, et al. Expression profiling in squamous carcinoma cells reveals pleiotropic effects of vitamin D3 analog EB1089 signaling on cell proliferation, differentiation, and immune system regulation. Mol Endocrinol. 2002;16:1243–56. doi: 10.1210/mend.16.6.0874. [DOI] [PubMed] [Google Scholar]
- 10.Rhee CS, Sen M, Lu D, et al. Wnt and frizzled receptors as potential targets for immunotherapy in head and neck squamous cell carcinomas. Oncogene. 2002;21:6598–605. doi: 10.1038/sj.onc.1205920. [DOI] [PubMed] [Google Scholar]
- 11.Thomas SM, Bhola NE, Zhang Q, et al. Cross-talk between G protein-coupled receptor and epidermal growth factor receptor signaling pathways contributes to growth and invasion of head and neck squamous cell carcinoma. Cancer Res. 2006;66:11831–9. doi: 10.1158/0008-5472.CAN-06-2876. [DOI] [PubMed] [Google Scholar]
- 12.Zhang Q, Thomas SM, Lui VW, et al. Phosphorylation of TNF-alpha converting enzyme by gastrin-releasing peptide induces amphiregulin release and EGF receptor activation. Proc Natl Acad Sci U S A. 2006;103:6901–6. doi: 10.1073/pnas.0509719103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Zhang Q, Thomas SM, Xi S, et al. SRC family kinases mediate epidermal growth factor receptor ligand cleavage, proliferation, and invasion of head and neck cancer cells. Cancer Res. 2004;64:6166–73. doi: 10.1158/0008-5472.CAN-04-0504. [DOI] [PubMed] [Google Scholar]
- 14.Zhang X, Hunt JL, Landsittel DP, et al. Correlation of protease-activated receptor-1 with differentiation markers in squamous cell carcinoma of the head and neck and its implication in lymph node metastasis. Clin Cancer Res. 2004;10:8451–9. doi: 10.1158/1078-0432.CCR-04-0546. [DOI] [PubMed] [Google Scholar]
- 15.Ogawa T, Ogawa H, Ueda Y, Ito Y, Takebayashi S, Bradford CR, Carey TE. Proceedings of the American Association for Cacner Research. Vol. 2001. 2001. Expression and mutation of galanin receptor 1 in head and neck squamous cell carcinoma (HNSCC) cell lines; p. 621. [Google Scholar]
- 16.Wang S, Hashemi T, Fried S, Clemmons AL, Hawes BE. Differential intracellular signaling of the GalR1 and GalR2 galanin receptor subtypes. Biochemistry. 1998;37:6711–7. doi: 10.1021/bi9728405. [DOI] [PubMed] [Google Scholar]
- 17.Branchek TA, Smith KE, Gerald C, Walker MW. Galanin receptor subtypes. Trends in pharmacological sciences. 2000;21:109–17. doi: 10.1016/s0165-6147(00)01446-2. [DOI] [PubMed] [Google Scholar]
- 18.Gundlach AL. Galanin/GALP and galanin receptors: role in central control of feeding, body weight/obesity and reproduction? European journal of pharmacology. 2002;440:255–68. doi: 10.1016/s0014-2999(02)01433-4. [DOI] [PubMed] [Google Scholar]
- 19.Bartfai T, Fisone G, Langel U. Galanin and galanin antagonists: molecular and biochemical perspectives. Trends in pharmacological sciences. 1992;13:312–7. doi: 10.1016/0165-6147(92)90098-q. [DOI] [PubMed] [Google Scholar]
- 20.Wittau N, Grosse R, Kalkbrenner F, Gohla A, Schultz G, Gudermann T. The galanin receptor type 2 initiates multiple signaling pathways in small cell lung cancer cells by coupling to G(q), G(i) and G(12) proteins. Oncogene. 2000;19:4199–209. doi: 10.1038/sj.onc.1203777. [DOI] [PubMed] [Google Scholar]
- 21.Berger A, Lang R, Moritz K, et al. Galanin receptor subtype GalR2 mediates apoptosis in SH-SY5Y neuroblastoma cells. Endocrinology. 2004;145:500–7. doi: 10.1210/en.2003-0649. [DOI] [PubMed] [Google Scholar]
- 22.Kanazawa T, Iwashita T, Kommareddi P, et al. Galanin and galanin receptor type 1 suppress proliferation in squamous carcinoma cells: activation of the extracellular signal regulated kinase pathway and induction of cyclin-dependent kinase inhibitors. Oncogene. 2007;26:5762–71. doi: 10.1038/sj.onc.1210384. [DOI] [PubMed] [Google Scholar]
- 23.Suzuki K, Suzuki I, Leodolter A, et al. Global DNA demethylation in gastrointestinal cancer is age dependent and precedes genomic damage. Cancer cell. 2006;9:199–207. doi: 10.1016/j.ccr.2006.02.016. [DOI] [PubMed] [Google Scholar]
- 24.Iismaa TP, Fathi Z, Hort YJ, et al. Structural organization and chromosomal localization of three human galanin receptor genes. Annals of the New York Academy of Sciences. 1998;863:56–63. doi: 10.1111/j.1749-6632.1998.tb10683.x. [DOI] [PubMed] [Google Scholar]
- 25.Baylin SB, Ohm JE. Epigenetic gene silencing in cancer - a mechanism for early oncogenic pathway addiction? Nature reviews. 2006;6:107–16. doi: 10.1038/nrc1799. [DOI] [PubMed] [Google Scholar]
- 26.Verma M, Srivastava S. Epigenetics in cancer: implications for early detection and prevention. The lancet oncology. 2002;3:755–63. doi: 10.1016/s1470-2045(02)00932-4. [DOI] [PubMed] [Google Scholar]
- 27.Park NH, Min BM, Li SL, Huang MZ, Cherick HM, Doniger J. Immortalization of normal human oral keratinocytes with type 16 human papillomavirus. Carcinogenesis. 1991;12:1627–31. doi: 10.1093/carcin/12.9.1627. [DOI] [PubMed] [Google Scholar]
- 28.Herman JG, Graff JR, Myohanen S, Nelkin BD, Baylin SB. Methylation-specific PCR: a novel PCR assay for methylation status of CpG islands. Proceedings of the National Academy of Sciences of the United States of America. 1996;93:9821–6. doi: 10.1073/pnas.93.18.9821. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Kuroki T, Trapasso F, Yendamuri S, et al. Allele loss and promoter hypermethylation of VHL, RAR-beta, RASSF1A, and FHIT tumor suppressor genes on chromosome 3p in esophageal squamous cell carcinoma. Cancer research. 2003;63:3724–8. [PubMed] [Google Scholar]
- 30.Berger A, Tuechler C, Almer D, et al. Elevated expression of galanin receptors in childhood neuroblastic tumors. Neuroendocrinology. 2002;75:130–8. doi: 10.1159/000048229. [DOI] [PubMed] [Google Scholar]
- 31.Mineta H, Miura K, Takebayashi S, et al. Cyclin D1 overexpression correlates with poor prognosis in patients with tongue squamous cell carcinoma. Oral oncology. 2000;36:194–8. doi: 10.1016/s1368-8375(99)00078-0. [DOI] [PubMed] [Google Scholar]
- 32.Cheung JO, Grant ME, Jones CJ, Hoyland JA, Freemont AJ, Hillarby MC. Apoptosis of terminal hypertrophic chondrocytes in an in vitro model of endochondral ossification. J Pathol. 2003;201:496–503. doi: 10.1002/path.1462. [DOI] [PubMed] [Google Scholar]
- 33.Wada M, Yazumi S, Takaishi S, et al. Frequent loss of RUNX3 gene expression in human bile duct and pancreatic cancer cell lines. Oncogene. 2004;23:2401–7. doi: 10.1038/sj.onc.1207395. [DOI] [PubMed] [Google Scholar]
- 34.Huang Y, Peng J, Oberley LW, Domann FE. Transcriptional inhibition of manganese superoxide dismutase (SOD2) gene expression by DNA methylation of the 5′ CpG island. Free radical biology & medicine. 1997;23:314–20. doi: 10.1016/s0891-5849(97)00095-6. [DOI] [PubMed] [Google Scholar]
- 35.Clark SJ, Harrison J, Molloy PL. Sp1 binding is inhibited by (m)Cp(m)CpG methylation. Gene. 1997;195:67–71. doi: 10.1016/s0378-1119(97)00164-9. [DOI] [PubMed] [Google Scholar]
- 36.Van Dyke DL, Worsham MJ, Benninger MS, et al. Recurrent cytogenetic abnormalities in squamous cell carcinomas of the head and neck region. Genes, chromosomes & cancer. 1994;9:192–206. doi: 10.1002/gcc.2870090308. [DOI] [PubMed] [Google Scholar]
- 37.Takebayashi S, Hickson A, Ogawa T, et al. Loss of chromosome arm 18q with tumor progression in head and neck squamous cancer. Genes, chromosomes & cancer. 2004;41:145–54. doi: 10.1002/gcc.20066. [DOI] [PubMed] [Google Scholar]
- 38.Gutkind JS. Cell growth control by G protein-coupled receptors: from signal transduction to signal integration. Oncogene. 1998;17:1331–42. doi: 10.1038/sj.onc.1202186. [DOI] [PubMed] [Google Scholar]
- 39.Dote H, Toyooka S, Tsukuda K, et al. Aberrant promoter methylation in human DAB2 interactive protein (hDAB2IP) gene in breast cancer. Clin Cancer Res. 2004;10:2082–9. doi: 10.1158/1078-0432.ccr-03-0236. [DOI] [PubMed] [Google Scholar]
- 40.Toyooka S, Toyooka KO, Harada K, et al. Aberrant methylation of the CDH13 (H-cadherin) promoter region in colorectal cancers and adenomas. Cancer research. 2002;62:3382–6. [PubMed] [Google Scholar]
- 41.Williams T, Tjian R. Analysis of the DNA-binding and activation properties of the human transcription factor AP-2. Genes & development. 1991;5:670–82. doi: 10.1101/gad.5.4.670. [DOI] [PubMed] [Google Scholar]
- 42.Tellez C, Bar-Eli M. Role and regulation of the thrombin receptor (PAR-1) in human melanoma. Oncogene. 2003;22:3130–7. doi: 10.1038/sj.onc.1206453. [DOI] [PubMed] [Google Scholar]
- 43.Xu Y, Porntadavity S, St Clair DK. Transcriptional regulation of the human manganese superoxide dismutase gene: the role of specificity protein 1 (Sp1) and activating protein-2 (AP-2) The Biochemical journal. 2002;362:401–12. doi: 10.1042/0264-6021:3620401. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Comb M, Goodman HM. CpG methylation inhibits proenkephalin gene expression and binding of the transcription factor AP-2. Nucleic acids research. 1990;18:3975–82. doi: 10.1093/nar/18.13.3975. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45.Zou MX, Butcher DT, Sadikovic B, Groves TC, Yee SP, Rodenhiser DI. Characterization of functional elements in the neurofibromatosis (NF1) proximal promoter region. Oncogene. 2004;23:330–9. doi: 10.1038/sj.onc.1207053. [DOI] [PubMed] [Google Scholar]
- 46.Mancini DN, Singh SM, Archer TK, Rodenhiser DI. Site-specific DNA methylation in the neurofibromatosis (NF1) promoter interferes with binding of CREB and SP1 transcription factors. Oncogene. 1999;18:4108–19. doi: 10.1038/sj.onc.1202764. [DOI] [PubMed] [Google Scholar]
- 47.Fujita N, Watanabe S, Ichimura T, et al. MCAF mediates MBD1-dependent transcriptional repression. Molecular and cellular biology. 2003;23:2834–43. doi: 10.1128/MCB.23.8.2834-2843.2003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Nakao M. Epigenetics: interaction of DNA methylation and chromatin. Gene. 2001;278:25–31. doi: 10.1016/s0378-1119(01)00721-1. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.