Abstract
Drosophila telomeres are remarkable because they are maintained by telomere-specific retrotransposons, rather than the enzyme telomerase that maintains telomeres in almost every other eukaryotic organism. Successive transpositions of the Drosophila retrotransposons onto chromosome ends produce long head-to-tail arrays that are analogous in form and function to the long arrays of short repeats produced by telomerase in other organisms. Nevertheless, Drosophila telomere repeats are retrotransposons, complex entities three orders of magnitude longer than simple telomerase repeats. During the >40–60 My they have been coevolving with their host, these retrotransposons perforce have evolved a complex relationship with Drosophila cells to maintain populations of active elements while carrying out functions analogous to those of telomerase repeats in other organisms. Although they have assumed a vital role in maintaining the Drosophila genome, the three Drosophila telomere-specific elements are non-LTR retrotransposons, closely related to some of the best known non-telomeric elements in the Drosophila genome. Thus, these elements offer an opportunity to study ways in which retrotransposons and their host cells can coevolve cooperatively. The telomere-specific elements display several characteristics that appear important to their roles at the telomere; for example, we have recently reported that they have evolved at least two innovative mechanisms for protecting essential sequence on their 5′ends. Because every element serves as the end of the chromosome immediately after it transposes, its 5′end is subject to chromosomal erosion until it is capped by a new transposition. These two mechanisms make it possible for at least a significant fraction of elements to survive their initial time as the chromosome end without losing sequence necessary to be competent for subsequent transposition. Analysis of sequence from >90 kb of assembled telomere array shows that these mechanisms for small scale sequence protection are part of a unified set which maintains telomere length homeostasis. Here we concentrate on recently elucidated mechanisms that have evolved to provide this small scale 5′ protection.
Key words: telomeres, retrotransposons, Drosophila, chromosome evolution, end replication problem, reverse transcriptase
A Brief History of Telomeres
Early study of chromosomes revealed that chromosome ends were special bodies (telomeres) with an important job: they prevent chromosome ends from behaving like broken DNA.1–3 Telomeres prevent chromosome ends from sticking to other ends with disastrous consequences. Since then we have learned that capping the chromosome end is only one of a still growing list of telomere jobs.
Molecular studies came later, driven by the recognition that DNA polymerase could not replicate the last few nucleotides on the ends of linear DNA. This “end-replication problem”, which should lead to a continual shortening of the chromosome as cells divide, launched intensive study of the ends of linear DNA. For technical reasons, linear mammalian viruses were chosen as useful models for the daunting eukaryotic chromosome. Viruses were found to have several elegant and economical solutions to the end replication problem, like covalently attached proteins or short DNA fold-back sequences. None of the viral solutions could be found on eukaryotic chromosome ends. Instead, the clue to the end-replication problem for chromosomes came from an unlikely model, amplified extrachromosomal genes encoding ribosomal RNA in the ciliated protozoan, Tetrahymena. Blackburn found each end of this DNA was composed of a long array of simple DNA nucleotide repeats, GGGGTT. She went on to discover an enzyme, telomerase, a discovery that resolved the end-replication problem. Telomerase has an RNA template from which it reverse transcribes short repeats (GGGGTT for Tetrahymena) onto chromosome ends to compensate for lost sequence.3,4
The clue from Tetrahymena led to rapid discovery of the telomere sequences in animals, plants and single celled eukaryotes. For the majority of these organisms, telomere arrays are composed of chains of 5–10 base-pair repeats added by telomerase. Typically, vertebrates add TTAGGG, plants TTTAGGG and insects TTAGG. Single celled eukaryotes have somewhat more variable sequences but these, too, are short and G+T-rich. These repeat arrays perform the second telomere job to be recognized. They prevent incomplete DNA replication from causing eventual loss of vital genetic material. Although they solve the same problem as the ends of linear viruses, telomere arrays are orders of magnitude longer than the short terminal sequences on viral ends. Multicellular eukaryotes tend to have ten to fifty kilobases of telomere repeats on each chromosome end; even unicellular eukaryotes have a few hundred basepairs of repeats per end. Furthermore, telomere array length is regulated in species-specific and cell type-specific ways; it is dynamic and fluctuates around a set point. The set point can change with environment and genetic background, as well as physical and even psychological, health.4 Both cell aging and cancer are marked by abnormal regulation of telomere length, structure and replication. The mechanisms by which telomere length is regulated as well as the mechanisms relating length to cell physiology are still major questions in the field.
Telomeres are nucleoprotein complexes, involving both telomere-specific proteins and telomere-associated proteins.5,6 Telomere-specific proteins evolve rapidly and tend to lack strong homology to their counterparts in other species. Telomere-associated proteins interact with telomeres but are known to have other functions in the cell: DNA damage response proteins are notable examples. The complexity of telomere chromatin is not surprising given the multiple roles in cell physiology now imputed to telomeres.
Drosophila Telomeres are a Remarkable Variation on the Telomerase Theme
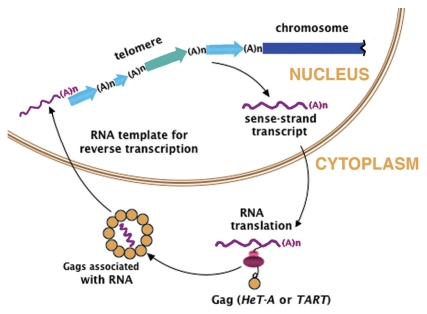
Telomeres were initially defined by study of Drosophila chromosomes, so it was a surprise to learn that Drosophila telomeres are maintained, not by telomerase, but by a dedicated set of non-LTR retrotransposons. Studies of D. melanogaster identified three elements, HeT-A,7,8 TART9 and TAHRE,10 that transpose specifically to chromosome ends where they form long head-to-tail arrays that make up telomeres in that heterochromatic region11 (Fig. 1). Although the Drosophila genome is littered with other retrotransposons related to the three telomeric elements, these other retrotransposons are not found in the telomere arrays. In fact the genomic distribution of the telomeric elements is almost the exact opposite of the distribution of other retrotransposons. The completely sequenced D. melanogaster euchromatic genome lacks even small fragments of telomeric elements although telomeric elements are found to transpose onto euchromatic DNA when that DNA is the broken end of the chromosome. Very little assembled sequence of the complex repeated DNA of heterochromatic regions is available but some of it contains fragments of telomeric elements, apparently moved into these regions by something other than active transposition.12,13
Figure 1.
Model for the extension of chromosomes by telomere-specific retrotransposons. Arrows represent the head-to-tail array of complete and 5′truncated HeT-A (blue) and TART (green) that makes up the telomere. (Actual telomeres typically have more elements than shown here). Transcription of an element in the array provides sense strand RNA (wavy magenta line) that is translated in the cytoplasm to yield Gag protein. This protein associates with the RNA and delivers it to the chromosome end where the RNA is reverse-transcribed onto the chromosome. Analogy with retroviruses suggests that reverse transcriptase is included in the Gag-RNA complex; however there is no evidence on this point. Magenta (A)n, 3′ poly(A) tail of RNA; black (A)n, oligoA at 3′end of each element in chromosomal DNA. This results from reverse transcriptase beginning DNA synthesis within the poly(A) tail of the RNA. Those oligoA terminations vary in length but are generally much shorter than their parent poly(A) tails. Modified from reference 34 by copyright permission of the Rockefeller University Press.
Analysis of telomere arrays provides strong evidence that telomeric elements transpose only to the extreme ends of intact or broken chromosomes; for broken chromosomes, transposition only to the ends is essentially incontrovertible. In telomere arrays telomeric elements are always joined head-to-tail. They are never found inserted within other elements, and there is no obvious sequence specificity to any attachment site on either 5′truncated or complete elements.12 (The 5′ termini of complete telomeric elements are also very variable). These findings are most straightforwardly explained by assuming that, because chromosome ends may be intact or variably 5′truncated, the new element is joined to whatever sequence is exposed on the end.
This retrotransposon variant actually provides a very robust system of telomere maintenance. We have identified HeT-A and TART homologues in D. virilis (separated from D. melanogaster by 40–60 My), showing that these elements have maintained telomeres since before the separation of these, and likely all, Drosophila species.14,15 Sequences of the HeT-A and TART homologues have diverged even more than sequences of typical coding regions in the two species. However both retrotransposons have conserved unusual features not found in non-telomeric retrotransposons, supporting the assumption that these features are adaptations for their roles in the telomere.
In Spite of the Obvious Differences Between Retrotransposon Telomeres and Telomerase Telomeres, these Telomeres have Fundamental Similarities
Both types of telomeres are extended by reverse-transcription of RNA templates. All non-LTR retrotransposons transpose by reverse-transcription directly onto a 3′OH of chromosomal DNA.16 HeT-A and TART specialize by reverse transcribing onto the 3′OH at the extreme end, much as telomerase adds repeats to telomeres in other organisms. In fact, telomerase and the reverse transcriptase of non-LTR elements are closely related.17 Although the Drosophila repeats (complete retrotransposons) are much longer and more complex than telomerase repeats, the length of Drosophila telomere arrays is similar to those of other multicellular eukaryotes.12,18 Length regulation is seen in both types of telomeres. It may be that RNA-templated extension evolved as the primary mechanism for telomere maintenance because it can be easily regulated and can produce rapid length change.
An increasing number of proteins are considered to be telomere-associated either because they have been found on telomeres or because mutation of the protein affects telomere structure and function. Many telomere-associated proteins in other organisms are also telomere associated in Drosophila. The list includes proteins involved with DNA damage response and repair, such as ATM, RAD50, MRE11, Ku70 and Ku80, and with chromatin structure, such as HP1 (reviewed in ref. 19). Although it seems paradoxical to have proteins involved in DNA repair also associate with telomeres, the fact is that they do in both retrotransposon- and telomerase-telomeres. This conservation suggests that the association is an essential feature of chromosome biology.
Somatic cells of Drosophila, like those of organisms with telomerase, respond to telomere loss by failure to pass cell cycle check points and, eventually, by apoptosis.20,21 Studies of Drosophila larval somatic tissues showed that the loss of a single telomere could trigger a spectrum of responses: chromosome end fusions, dicentric chromosome breakage, aneuploidy and other signs of genome instability also associated with dysfunctional telomeres in mammals and yeasts.
In contrast to the somatic cell studies, the genetic tools available in Drosophila have revealed a more complex picture in germline cells. Biessmann et al.22 showed that females with defective mu-2 genes could produce progeny carrying terminally deleted chromosomes that had no detectable telomere sequences yet did not form the end to end fusions expected of broken ends. These deletions had acquired the telomere's capping function although they could no longer replace lost terminal sequence and, therefore, progressively shortened. This was the first demonstration that the capping and end replication functions can be separated. Others found that P-element transposase, like the mu-2 mutant background, could produce similar capped terminal deletions in germline cells.23,24 In all cases the broken ends appear to be capped, however they continue to shorten at 50–100 nt per fly generation unless transposition of telomere retrotransposons rebuilds a new telomere array.
This atypical behavior of broken ends in the Drosophila germline has an intriguing resemblance to McClintock's studies of broken chromosomes in corn. She found that the broken ends she followed through repeated bridge-breakage-fusion cycles in endosperm were permanently healed when they were passed into the sporophyte.2 In contrast, chromosomes that broke while they were in sporophyte tissue did not heal. Thus, both fly and corn studies suggest that there are cell type-specific responses to telomere loss.
Retrotransposon Telomeres Must Preserve a Stock of Transposition-Competent Elements
In contrast to their structural and functional similarities, retrotransposon and telomerase telomeres differ fundamentally in the way their RNA templates are maintained. Most organisms have only a single telomerase reverse transcriptase and a single copy of its RNA template in their genomes. Both are encoded by genes located safely in the interior of a chromosome. This centralized organization produces repeat sequences that are essentially identical, not only within a genome but also across many distantly related species.
On the other hand, retrotransposon telomeres might be considered a grass roots organization. The elements transcribed to produce new transpositions are all located in telomere arrays and therefore at risk of loss in this dynamic environment.12 Telomere arrays are very long so it would appear that elements located in the more proximal regions have been there for some time without being under selection for function. Therefore, sequence decay is a second risk for transposition-competent elements. Retrotransposons appear to spread these risks by maintaining multiple replication-competent elements, rather than a single source of repeat sequences. Both HeT-A and TART have multiple subfamilies, differing noticeably in sequence, yet those found in carefully sequenced and assembled arrays appear capable of producing functional proteins.12 (TAHRE is a rare element and we do not have enough data to include it in this discussion). Our sequence analyses allow us to speculate that quality control of HeT-A and TART elements is imposed at the level of transposition; any sequence changes that do not preclude transposition will allow a mutated element to found a new subfamily.
The History of Events at Drosophila Telomeres can be Read from the Sequence of Assembled Retrotransposon Arrays
The long arrays of retrotransposons that constitute Drosophila telomeres are formed by repeated transpositions onto the chromosome end. Thus they display an ordered record of events at the end: each element is younger than its proximal neighbor. Unfortunately the complex repeated sequences in telomere arrays preclude accurate assembly from whole genome sequencing. However, we have been able to analyze sequence from a single BAC derived from the 4R telomere and also from directed sequencing of the XL telomere: both sequences were from D. melanogaster.12,25 Although neither sequence reaches the extreme end of a telomere, both are linked to subtelomere sequence of the chromosome; thus they include the most proximal, and therefore the oldest, elements in the retrotransposon arrays. The 4R sequence has >70 kb of retrotransposon array and the XL sequence has >20 kb.
These samples appear to be representative because elements are in the proportion expected from other studies of the D. melanogaster genome. HeT-A is the most abundant element, with four complete and eleven partial elements. There are two complete and three partial TARTs, and no TAHRE. (We do not include the terminal element in each array because it has been truncated by cloning). All elements have their 5′ends toward the end of the chromosome, as expected if they were reverse-transcribed onto a 3′OH thereon. Elements in the arrays are complete at their 3′ (proximal) end: partial elements have undergone variable 5′ loss that can be as much as several kb.
Our analyses of this data set show that events in telomere arrays are much more complex than the relatively regular small scale sequence loss (50–100 nt per fly generation), seen on broken chromosome ends, which has been attributed to end replication losses.22 Our quantitative analysis of small scale end erosion leads to a loss distribution with a distinct functional form that is hard to reconcile with that usually assumed for end replication losses. The erosion distribution is also very different from that for the quantitative loss distribution of 5′truncated partial elements; Thus, the process details leading to end erosion and 5′-truncation are certainly different; furthermore, the end erosion loss distribution can also include end replication losses.25
These data require us to postulate the existence of a complex process that maintains telomere length homeostasis and preserves a supply of transposition-competent elements. That process, whatever it may be, can be conceptualized as several, no doubt interacting, relatively simple mechanisms.25 The first is small scale end erosion that rarely removes all non-essential buffering 5′ sequence from the terminal element before a new transposition takes over the end to stop erosion of the element it has displaced. Elements have evolved adaptations to add enough non-essential sequence to protect complete elements (see below). Small scale erosion, including deficient end replication, removes very few nucleotides compared to those added by transposition of a new element (6–13 kb, HeT-A, TART or TAHRE) so telomere length homeostasis requires additional mechanisms to correct this imbalance. The population of 5′-truncated elements is evidence of large scale terminal loss. (There are no 3′-truncations). The length distribution of partial elements strongly suggests that complete elements are significantly over-represented. We suggest that the additional loss mechanism(s) could be due to length-selective terminal deletions (LSTD) encompassing part or all of the telomere, followed by rebuilding. LSTD could be mediated by protective telomere proteins.
The evidence that DNA turnover within intact Drosophila telomeres is more complex than the relatively regular loss seen on broken chromosomes strongly suggests that intact Drosophila telomeres, like those in other organisms, contain multiple proteins involved in modulating turnover and capping of terminal DNA. Many telomere proteins tend to evolve rapidly, in sequence and sometimes function, making it difficult to identify and compare the activity of telomere proteins even among closely related organisms (reviewed in ref. 6). Analogues of proteins implicated in modulating length regulation in other organisms have not been identified in Drosophila but it seems likely that they could be responsible for the complex turnover seen at intact telomeres. The ends of broken Drosophila chromosomes that have acquired end capping have been shown to have a few telomere proteins important for capping;26 however we probably know only a fraction of the proteins associated either with broken ends or intact telomeres in Drosophila. Further characterization of these proteins should help explain the difference in turnover of DNA at the two types of chromosome ends.
HeT-A and TART have Innovative Adaptations to Protect Essential 5′ Sequence
The typical non-LTR promoter lies within the 5′UTR immediately downstream of the transcription initiation site.27 Thus the promoter is included in the RNA and moved to the new site ensuring that the new retrotransposon can be transcribed in its new site. However, there is a risk for telomeric elements because they transpose to the end of the chromosome, exposing vital sequence in the 5′UTR. Telomeric elements have evolved at least two adaptations to protect the 5′sequence.
In the next Section and in figure 2, we describe how D. melanogaster's HeT-A (HeT-Amel) 28 and D. virilis's TART (TARTvir) 29 add non-essential sequence to the 5′end of their RNA. In contrast, TARTmel adds extra 5′ sequence by recopying part of its RNA when it is reverse-transcribed onto the chromosome.29 For at least a significant fraction of transpositions, telomere array data shows that both adaptations add sufficient buffering sequence to prevent erosion from eating into essential sequence before a new transposition arrives to cap the element. HeT-Avir is still an enigma: it has no obvious buffering sequence yet the available sequence has a good fraction of complete elements, suggesting that HeT-Avir has yet another adaptation to protect its 5′end.28
Figure 2.
Mechanisms for adding buffering 5′ sequence to transposing elements. (A) Using sequence copied from upstream neighbor. Used by HeTAmel and TARTvir. Telomere segment with a complete HeT-Amel flanked by other elements (top) and TARTvir (bottom) flanked by other elements. (Other elements are shown as gray when they could be either HeT-A or TART, but note that for these elements the immediately upstream element must be a sister element to provide a transcription start site). HeT-Amel UTRs are magenta with lighter box at 5′end of 5′UTR representing string of variably truncated Tags. TARTvir UTRs are lavender with darker box on 5′UTR representing Tag string. For both elements transcription starts at the bent arrow in the upstream element and continues through the complete downstream element. The resulting RNA (black line) has a new Tag consisting of the last nucleotides of the upstream element (short colored line and (A)n on 5′end of RNA). When the RNA is reverse transcribed onto the chromosome this new Tag becomes the 5′end of the newly transposed element, undergoes erosion, and if the element transposes again, will be internalized into the string of variably eroded Tags indicated by the 5′ box on the complete element. (B) Using sequence copied from the 3′UTR of transposing RNA. Used by TARTmel. Telomere segment with a complete TARTmel (purple UTRs) flanked by other elements (gray boxes). (A)n, 3′ oligoA in DNA; AAAAAA, 3′polyA tail on RNA; Gold arrows, PNTRs. Transcription starts at the bent arrow, producing an RNA with a very short 5′UTR. When this is reverse transcribed onto the chromosome end, the reverse transcriptase jumps back to identical sequence in the 3′ UTR and copies sequence to extend the 5′UTR, providing sacrificial DNA to buffer essential 5′ sequence.
HeT-Amel and TARTvir Share an Unusual Promoter Architecture which Adds Buffering 5′ Sequence
Promoter sequences slightly upstream of the 3′end of either of these two elements drive transcription, not of that element, but of its downstream neighbor. Transcription starts at sites within the 3′end of the upstream element.30,31 Thus the new RNA has a very short copy of 3′ sequence (plus the oligoA) from its neighbor added to its 5′end as a Tag (Fig. 2A). When the RNA is reverse-transcribed onto a chromosome end, this Tag provides expendable 5′ DNA sequence. If a new transposition caps the end before the first Tag sequence is completely eroded, the remaining sequence is left on the element as a truncated Tag, a record of sequence loss while on the chromosome end. When this element transposes again, a new Tag is added to the 5′end of the truncated Tag. Complete elements in telomere arrays have strings of several variably truncated Tags, showing that they have transposed multiple times and that either the rate of terminal erosion or the rate of new transposition (or both) is irregular.25,28
This upstream promoter requires that the element to be transcribed lie immediately downstream of another element of the same kind which can provide the promoter. (The promoting element can be a 5′truncated partial element because only a few hundred nt of the 3′UTR sequence is needed for activity).30 Thus a complete HeT-Amel, or TARTvir has a good probability of having an appropriate neighbor in a telomere array.
TARTmel Also Adds Protective 5′ Sequence but Does This by Making a Second Copy from its own 3′UTR When It Is Reverse Transcribed onto the Chromosome
Somewhat surprisingly, TARTmel does not have the promoter used by both its D. virilis homologue and its partner in the D. melanogaster telomere.29 Perhaps this mechanism is unfavorable, because TARTmel is greatly outnumbered by HeT-Amel so is less likely to have another TART as an upstream neighbor (Fig. 2B).
TARTmel has several remarkable characteristics, not seen in other telomeric elements, that allowed us to formulate a model for TARTmel 5′protection.29 There are three TARTmel subfamilies: members are randomly distributed in telomeres, suggesting that they are functionally equivalent, yet the long UTR sequences differ strikingly between subfamilies. In each subfamily the sequence of the 5′UTR is a direct repeat of sequence in the 3′UTR. We refer to these as Perfect Non-Terminal Repeats (PNTRs) because the 3′ repeat ends a few hundred nt upstream of the 3′end of the element.32 Comparison of subfamily members shows that sequence changes in one PNTR are reproduced in its partner PNTR, i.e., the repeats are evolving together.29
The 5′end of TART appears to be highly variable; the few available 5′UTR sequences range from 33 nt to 3,934 nt, and each element sequenced has a different end. Lengths do not correlate with subfamily; both the longest and the shortest 5′UTRs are TART-A. Extensive searches for transcription start sites for full length TARTmel RNA have revealed only one site.29,31 In each subfamily this site lies in the 5′UTR ∼75 nt upstream of the first coding region. This result presented a paradox because only one known TARTmel has a 5′UTR (33 nt) short enough to have been reverse-transcribed from RNA with 75 nt of 5′UTR sequence. The evidence that PNTR pairs are evolving together provides a solution to this paradox. We concluded that when the RNA is reverse-transcribed onto the chromosome, the reverse transcriptase makes a template jump from the 5′end of the RNA back to the 3′PNTR and recopies sequence to extend the 5′PNTR, adding non-essential sequence. The extreme variability in the length of the 5′UTR in genomic TARTmel elements can be explained by variable termination of reverse transcription, by terminal erosion, by terminal deletions or some combination thereof.
Concluding Remarks
This review has focused on recently discovered adaptations which reveal mechanisms used to maintain a population of complete, replicatively competent, elements in Drosophila telomeres. We want to emphasize that replicatively competent elements are not the whole story; we believe that 5′truncated elements are equally important for the telomere roles that are analogous to those of telomerase repeats. Here it is relevant that telomere elements differ from their non-telomere relatives in having very long 3′UTRs and that much of this 3′UTR sequence is strongly conserved.33 It is also the last to be lost by 5′-truncation. We believe these facts indicate that these sequences have important roles in chromosome structure.
Acknowledgments
This work was supported by National Institutes of Health grant GM50315 to M.L.P.
Abbreviations
- bp
base pairs
- Kb
kilobase
- LSTD
length-selective terminal deletion
- LTR
long terminal repeat
- My
million years
- Nt
nucleotide
- PNTR
perfect non-terminal repeat
- UTR
untranslated region
References
- 1.Muller HJ. The remaking of chromosomes. Collecting Net. 1938;13:181–195. [Google Scholar]
- 2.McClintock B. The Stability of Broken Ends of Chromosomes in Zea Mays. Genetics. 1941;26:234–282. doi: 10.1093/genetics/26.2.234. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Blackburn EH. Telomerases. Annu Rev Biochem. 1992;61:113–129. doi: 10.1146/annurev.bi.61.070192.000553. [DOI] [PubMed] [Google Scholar]
- 4.Blackburn EH. Switching and signaling at the telomere. Cell. 2001;106:661–673. doi: 10.1016/s0092-8674(01)00492-5. [DOI] [PubMed] [Google Scholar]
- 5.Martinez P, Blasco MA. Telomeric and extra-telomeric roles for telomerase and the telomere-binding proteins. Nat Rev Cancer. 2011;11:161–176. doi: 10.1038/nrc3025. [DOI] [PubMed] [Google Scholar]
- 6.Price CM, Boltz KA, Chaiken MF, Stewart JA, Beilstein MA, Shippen DE. Evolution of CST function in telomere maintenance. Cell Cycle. 2010;9:3157–3165. doi: 10.4161/cc.9.16.12547. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Traverse KL, Pardue ML. A spontaneously opened ring chromosome of Drosophila melanogaster has acquired He-T DNA sequences at both new telomeres. Proc Natl Acad Sci USA. 1988;85:8116–8120. doi: 10.1073/pnas.85.21.8116. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Biessmann H, Mason JM, Ferry K, d'Hulst M, Valgeirsdottir K, Traverse KL, et al. Addition of telomere-associated HeT DNA sequences “heals” broken chromosome ends in Drosophila. Cell. 1990;61:663–673. doi: 10.1016/0092-8674(90)90478-w. [DOI] [PubMed] [Google Scholar]
- 9.Sheen FM, Levis RW. Transposition of the LINE-like retrotransposon TART to Drosophila chromosome termini. Proc Natl Acad Sci USA. 1994;91:12510–12514. doi: 10.1073/pnas.91.26.12510. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Abad JP, De Pablos B, Osoegawa K, De Jong PJ, Martin-Gallardo A, Villasante A. TAHRE, a novel telomeric retrotransposon from Drosophila melanogaster, reveals the origin of Drosophila telomeres. Mol Biol Evol. 2004;21:1620–1624. doi: 10.1093/molbev/msh180. [DOI] [PubMed] [Google Scholar]
- 11.Pardue ML, DeBaryshe PG. Retrotransposons provide an evolutionarily robust non-telomerase mechanism to maintain telomeres. Annu Rev Genet. 2003;37:485–511. doi: 10.1146/annurev.genet.38.072902.093115. [DOI] [PubMed] [Google Scholar]
- 12.George JA, DeBaryshe PG, Traverse KL, Celniker SE, Pardue ML. Genomic organization of the Drosophila telomere retrotransposable elements. Genome Res. 2006;16:1231–1240. doi: 10.1101/gr.5348806. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Mendez-Lago M, Wild J, Whitehead SL, Tracey A, de Pablos B, Rogers J, et al. Novel sequencing strategy for repetitive DNA in a Drosophila BAC clone reveals that the centromeric region of the Y chromosome evolved from a telomere. Nucleic Acids Res. 2009;37:2264–2273. doi: 10.1093/nar/gkp085. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Casacuberta E, Pardue ML. HeT-A elements in Drosophila virilis: retrotransposon telomeres are conserved across the Drosophila genus. Proc Natl Acad Sci USA. 2003;100:14091–14096. doi: 10.1073/pnas.1936193100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Casacuberta E, Pardue ML. Transposon telomeres are widely distributed in the Drosophila genus: TART elements in the virilis group. Proc Natl Acad Sci USA. 2003;100:3363–3368. doi: 10.1073/pnas.0230353100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Luan DD, Korman MH, Jakubczak JL, Eickbush TH. Reverse transcription of R2Bm RNA is primed by a nick at the chromosomal target site: a mechanism for non-LTR retrotransposition. Cell. 1993;72:595–605. doi: 10.1016/0092-8674(93)90078-5. [DOI] [PubMed] [Google Scholar]
- 17.Nakamura TM, Morin GB, Chapman KB, Weinrich SL, Andrews WH, Lingner J, et al. Telomerase catalytic subunit homologs from fission yeast and human. Science. 1997;277:955–959. doi: 10.1126/science.277.5328.955. [DOI] [PubMed] [Google Scholar]
- 18.Abad JP, De Pablos B, Osoegawa K, De Jong PJ, Martin-Gallardo A, Villasante A. Genomic analysis of Drosophila melanogaster telomeres: full-length copies of HeT-A and TART elements at telomeres. Mol Biol Evol. 2004;21:1613–1619. doi: 10.1093/molbev/msh174. [DOI] [PubMed] [Google Scholar]
- 19.Cenci G, Ciapponi L, Gatti M. The mechanism of telomere protection: a comparison between Drosophila and humans. Chromosoma. 2005;114:135–145. doi: 10.1007/s00412-005-0005-9. [DOI] [PubMed] [Google Scholar]
- 20.Ahmad K, Golic KG. Telomere loss in somatic cells of Drosophila causes cell cycle arrest and apoptosis. Genetics. 1999;151:1041–1051. doi: 10.1093/genetics/151.3.1041. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Titen SW, Golic KG. Telomere loss provokes multiple pathways to apoptosis and produces genomic instability in Drosophila melanogaster. Genetics. 2008;180:1821–1832. doi: 10.1534/genetics.108.093625. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Biessmann H, Carter SB, Mason JM. Chromosome ends in Drosophila without telomeric DNA sequences. Proc Natl Acad Sci USA. 1990;87:1758–1761. doi: 10.1073/pnas.87.5.1758. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Levis RW. Viable deletions of a telomere from a Drosophila chromosome. Cell. 1989;58:791–801. doi: 10.1016/0092-8674(89)90112-8. [DOI] [PubMed] [Google Scholar]
- 24.Mikhailovsky S, Belenkaya T, Georgiev P. Broken chromosomal ends can be elongated by conversion in Drosophila melanogaster. Chromosoma. 1999;108:114–120. doi: 10.1007/s004120050358. [DOI] [PubMed] [Google Scholar]
- 25.DeBaryshe PG, Pardue ML. Differential maintenance of DNA sequences in telomeric and centromeric heterochromatin. Genetics. 2011;187:51–60. doi: 10.1534/genetics.110.122994. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Gao G, Walser JC, Beaucher ML, Morciano P, Wesolowska N, Chen J, et al. HipHop interacts with HOAP and HP1 to protect Drosophila telomeres in a sequence-independent manner. EMBO J. 2010;29:819–829. doi: 10.1038/emboj.2009.394. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Smale ST, Kadonaga JT. The RNA polymerase II core promoter. Annu Rev Biochem. 2003;72:449–479. doi: 10.1146/annurev.biochem.72.121801.161520. [DOI] [PubMed] [Google Scholar]
- 28.Traverse KL, George JA, Debaryshe PG, Pardue ML. Evolution of species-specific promoter-associated mechanisms for protecting chromosome ends by Drosophila Het-A telomeric transposons. Proc Natl Acad Sci USA. 2010;107:5064–5069. doi: 10.1073/pnas.1000612107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.George JA, Traverse KL, DeBaryshe PG, Kelley KJ, Pardue ML. Evolution of diverse mechanisms for protecting chromosome ends by Drosophila TART telomere retrotransposons. Proc Natl Acad Sci USA. 2010;107:21052–21057. doi: 10.1073/pnas.1015926107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Danilevskaya ON, Arkhipova IR, Traverse KL, Pardue ML. Promoting in tandem: the promoter for telomere transposon HeT-A and implications for the evolution of retroviral LTRs. Cell. 1997;88:647–655. doi: 10.1016/s0092-8674(00)81907-8. [DOI] [PubMed] [Google Scholar]
- 31.Maxwell PH, Belote JM, Levis RW. Identification of multiple transcription initiation, polyadenylation and splice sites in the Drosophila melanogaster TART family of telomeric retrotransposons. Nucleic Acids Res. 2006;34:5498–5507. doi: 10.1093/nar/gkl709. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Danilevskaya ON, Traverse KL, Hogan NC, DeBaryshe PG, Pardue ML. The two Drosophila telomeric transposable elements have very different patterns of transcription. Mol Cell Biol. 1999;19:873–881. doi: 10.1128/mcb.19.1.873. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Danilevskaya ON, Lowenhaupt K, Pardue ML. Conserved subfamilies of the Drosophila HeT-A telomere-specific retrotransposon. Genetics. 1998;148:233–242. doi: 10.1093/genetics/148.1.233. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Rashkova S, Karam SE, Kellum R, Pardue ML. Gag proteins of the two Drosophila telomeric retrotransposons are targeted to chromosome ends. J Cell Biol. 2002;159:397–402. doi: 10.1083/jcb.200205039. [DOI] [PMC free article] [PubMed] [Google Scholar]