Abstract
Background/Aims
Oxidative stress results in protein oxidation and is implicated in carcinogenesis. Sulfiredoxin (Srx) is responsible for the enzymatic reversal of inactivated peroxiredoxin (Prx). Nuclear factor E2-related factor 2 (Nrf2) binds to antioxidant responsive elements and upregulates the expression of Srx and Prx during oxidative stress. We aimed to elucidate the biological functions and potential roles of Srx in lung cancer.
Methods
To study the roles of Srx and Prx III in lung cancer, we compared the protein levels of Nrf2, Prxs, thioredoxin, and Srx in 40 surgically resected human lung cancer tissues using immunoblot and immunohistochemical analyses. Transforming growth factor-β1, tumor necrosis factor-α, and camptothecin treatment were used to examine Prx III inactivation in Mv1Lu mink lung epithelial cells and A549 lung cancer cells.
Results
Prx I and Prx III proteins were markedly overexpressed in lung cancer tissues. A significant increase in the oxidized form of a cysteine sulfhydryl at the catalytic site of Prxs was found in carcinogenic lung tissue compared to normal lung tissue. Densitometric analyses of immunoblot data revealed significant Srx expression, which was higher in squamous cell carcinoma tissue (60%, 12/20) than in adenocarcinoma (20%, 4/20). Also, Nrf2 was present in the nuclear compartment of cancer cells.
Conclusions
Srx and Prx III proteins were markedly overexpressed in human squamous cell carcinoma, suggesting that these proteins may play a protective role against oxidative injury and compensate for the high rate of mitochondrial metabolism in lung cancer.
Keywords: GA-binding protein transcription factor, Peroxiredoxins, Sulfiredoxin, Lung neoplasms
INTRODUCTION
Oxidative stress and growth factor stimulation can cause harmful changes in DNA, proteins, and lipids. Signaling by reactive oxygen species (ROS), especially hydrogen peroxide, plays a central role in a variety of cellular events such as growth, proliferation, and carcinogenesis [1]. The lungs are constantly exposed to oxidative stress and xenobiotic challenges such as environmental pollution, toxic fumes, and cigarette smoke during gas exchange, and many specialized and dedicated proteins function as antioxidants to protect pulmonary epithelial cells from the heavy oxidative stress load. These antioxidant-related proteins include glutathione, catalase, superoxide dismutase, manganese superoxide dismutase, nicotinamide adenine dinucleotide phosphate oxidases, peroxiredoxins (Prxs), and others [2].
Prxs are a ubiquitous family of peroxidases that function to protect human cells from oxidative stress. Involved in a number of cellular activities, Prxs remove hydrogen peroxide produced during normal cellular metabolism, growth, differentiation, inflammation, and proliferation [3,4].
Sulfiredoxins (Srxs) are a family of low molecular weight, sulfur-containing proteins that function to maintain cellular redox balance. Srx is responsible for enzymatically reversing inactivated Prxs, in an ATP-, Mg2+-, and Ca2+-dependent manner [5-7].
Nuclear factor E2-related factor 2 (Nrf2) is a well-known master transcription factor for antioxidant responsive element (ARE)-mediated protein expression [8,9]. The pathway involving Kelch-like ECH-associated protein 1 (Keap1), Nrf2, and ARE-mediated protein expression is critical for protecting cells from both endogenous and exogenous stress [10,11]. A recent study using Nrf2-knockout mice clearly demonstrated that hyperoxia-induced expression of Srx in the lungs is dependent on the Nrf2-signaling pathway [12].
During carcinogenesis, cancer cells take advantage of the Nrf2/ARE protective machinery to survive the oxidative stress triggered by uncontrolled growth, chemotherapy, and radiotherapy, allowing them to endure the toxic effects of the therapy [13-16]. It was recently reported that a cigarette smoke induced-Keap1 mutation caused the release of Nrf2 and the activation of ARE-mediated protein transcription, contributing to the development of squamous carcinoma in the head and neck [17,18]. However, the pathophysiological role of Nrf2 and its associations among the Prx and Srx proteins in human lung disease have not been clearly defined.
Our aim was to elucidate the role of ROS-related signal transduction and to explore the associations between Nrf2, peroxiredoxin, sulfiredoxin, and thioredoxin (Trx) in human lung cancer. Thus, we examined Nrf2-dependent, ARE-mediated expression of Srx in human lung cancer.
METHODS
Materials
RPMI 1640 medium, fetal bovine serum (FBS), trypsin-EDTA, and penicillin-streptomycin were purchased from Gibco BRL (Sparks, MD, USA). Alexa 488-conjugated goat anti-mouse and anti-rabbit immunoglobin G (IgG), Alexa 594-conjugated goat anti-rabbit IgG, Mitotracker Red CMXRos, and DAPI were purchased from Molecular Probes Inc. (Eugene, OR, USA). Antibodies against specific Prx isoforms and Trx were purchased from Young in Frontier Co. Ltd. (Seoul, Korea). Nrf2, anti-hemagglutinin (HA), and α-tubulin were obtained from Santa Cruz Biotechnology (Santa Cruz, CA, USA). Anti-Srx antibodies and vectors, pCGN and pCGN-Srx, were kindly provided by Dr. H.A. Woo (Ewha Womans University, Seoul, Korea). Camptothecin and protease inhibitor were purchased from Sigma-Aldrich (St. Louis, MO, USA).
Subjects
Forty patients who underwent lung cancer-related surgery at Ajou University Medical Center (Suwon, Korea) were enrolled in this study. Paired samples of tumor tissue and normal lung tissue were collected from each individual after a pathological evaluation. Informed consent was obtained and the study protocol was approved by the Internal Review Board of Ajou University Medical Center.
Cell culture and viability
The A549 cell line (human lung carcinoma) was obtained from the American Type Culture Collection (Manassas, VA, USA). Cells were cultured in RPMI 1640 supplemented with 10% FBS and 1% penicillin-streptomycin at 37℃ in a humidified atmosphere containing 5% CO2. Cells (1 × 105) were seeded onto six-well plates, cultured in RPMI containing 10% FBS for 12 hours, and then treated with H2O2 (200 µM), transforming growth factor (TGF)-β1 (2 or 10 µg/mL), tumor necrosis factor (TNF)-α (10 µg/mL), or camptothecin (10 µM/mL) for the indicated periods. Cells were continuously cultured as necessary by refreshing the medium every day with medium containing the same growth factors. At the indicated times, cells were harvested by trypsinization, stained with 0.2% (w/v) trypan blue (Gibco) to exclude dead cells, and counted with a hemacytometer. The population of trypan blue-stained cells was counted using an inverted microscope (Olympus, Tokyo, Japan) to evaluate cellular viability.
Transfection and depletion of siRNA
Cells were transfected with pCGN and Srx-pCGN-HA using FuGENE HD transfection reagent (Roche Diagnostics, GmbH, Mannheim, Germany). Human Srx small interfering RNA (siRNA; SMART pool M-015263-00-0005) and siCONTROL® non-targeting siRNA were obtained from Dharmacon Research (Chicago, IL, USA). Cells were seeded onto six-well plates and allowed to reach 60% confluence. The next day, the cells were transfected with 20 nM Srx-siRNA or control siRNA using Lipofectamine RNAiMax reagent (Invitrogen, Carlsbad, CA, USA).
Immunocytochemistry
Cells were grown on glass-bottom culture dishes, fixed with 4% paraformaldehyde, and permeabilized for 5 minutes with 0.2% Triton X-100. The cells were incubated in 1 × phosphate buffered saline (PBS) containing 1% bovine serum albumin for 1 hour and then with primary antibody overnight at 4℃. After washing, the cells were incubated with a secondary antibody. Immunostained cells were washed with 1 × PBS and mounted with prolong GOLD (Molecular Probes). HA-tagged Srx was detected with anti-HA monoclonal antibody (10 µg/mL) and Alexa 488-conjugated goat anti-mouse IgG (5 µg/mL). Prx III was detected with polyclonal antibodies (10 µg/mL) and Alexa 488-conjugated goat anti-rabbit IgG (5 µg/mL) or Alexa 594-conjugated goat anti-rabbit IgG (5 µg/mL). Mitochondria were stained with 0.1 µM MitoTracker Red CMXRos. Confocal fluorescence images were obtained using an LSM510 microscope (Carl Zeiss, MD, USA).
Co-immunoprecipitation and immunoblot analysis
For immunoblot analysis, lung tissues were homogenized in lysis buffer containing 50 mM Tris-HCl (pH 7.5), 150 mM NaCl, 1 mM EDTA, 1% Triton X, and a protease inhibitor cocktail (Sigma-Aldrich). Cell lysates were centrifuged at 12,000 rpm for 10 minutes at 4℃, and the samples were boiled for 5 minutes in 2 × sample buffer. For immunoprecipitation, A549 cells were lysed in lysis buffer and centrifuged at 12,000 × g for 10 minutes at 4℃. The A549 cell lysates were precleared with Protein A agarose beads (Upstate, Charlottesville, VA, USA), anti-HA antibody (2 µL) was added to the lysates, and the lysates were rotated at 4℃ for 2 hours. Protein A agarose beads (20 µL) were then added and the samples were rotated in tubes for an additional 2 hours. The samples were washed four times and then boiled for 5 minutes in 1× sample buffer. The sample proteins were separated by sodium dodecyl sulfate polyacrylamide gels (SDS-PAGE) and transferred to nitrocellulose membranes (Scheicher and Schuell BioScience Gmbh, Dassel, Germany). After blocking, each membrane was incubated with primary antibody and then with sheep anti-mouse IgG or donkey anti-rabbit IgG (Amersham Biosciences, Buckinghamshire, UK). Immune complexes were detected by enhanced chemiluminescence (Amersham Biosciences).
Two-dimensional electrophoresis
A549 cells were lysed in lysis buffer (8 M urea, 4% CHAPS, 40 mM Tris, 20 M DTT, 0.5% IPG, pH 3-10 non-linear) containing protease inhibitors and then sonicated three times with 20-s pulses. Insoluble materials were removed by centrifugation at 12,000 × g for 15 minutes, and the lysate was mixed with rehydration buffer (8 M urea, 2% CHAPS, 20 mM DTT, 0.5% IPG buffer, pH 3-10 non-linear, bromophenol blue) and loaded onto immobilized pH gradient strips (pH 3-10 non-linear; Amersham Biosciences, Piscataway, NJ, USA) for 16 hours at room temperature. Isoelectric focusing (Amersham Biosciences) and preparation of the immobilized pH gradient strips for SDS-PAGE were conducted according to the manufacturer's procedures. SDS-PAGE was performed in 12% gels using an Amersham Bioscience SE600 13 cm vertical unit, and the proteins were transferred to nitrocellulose membranes for immunoblot analysis.
Subcellular fractionation
Cytosolic and mitochondrial-enriched fractions were prepared from A549 cells using a Subcellular Proteome Extraction kit (Calbiochem, Darmstadt, Germany). Cells were washed twice in ice-cold (4℃) wash buffer for 5 minutes and then incubated with ice-cold extraction buffer I containing protease inhibitor cocktail for 10 minutes at 4℃ with gentle agitation. The supernatant was collected as the cytosolic fraction. Then, the cells were incubated with ice-cold extraction buffer II containing protease inhibitor cocktail for 30 minutes at 4℃ with gentle agitation, and this supernatant was collected as the mitochondria-enriched heavy membrane fraction.
Immunohistochemistry
Sections of paraffin wax-embedded human tissue were used for immunohistochemistry. After deparaffinization, nonspecific binding was blocked with 10% normal goat serum (Amersham Biosciences) and the sections were probed with Srx-specific polyclonal antibodies overnight at 4℃. Tissue sections were rinsed three times in PBS and incubated with streptavidin-biotin-peroxidase reagents (LSAB kit; Dako, Copenhagen, Denmark). Immunoreactivity was visualized with diaminobenzidine. The sections were counterstained with hematoxylin and eosin.
Statistical analysis
Data were analyzed at least three times using SPSS version 11.0 (SPSS Inc., Chicago, IL, USA). Values are presented as mean ± SD of triplicate determinations. Student's t test was used to compare continuous data. A p value < 0.05 was considered statistically significant.
RESULTS
Localization of Prx III and Srx in A549 cells
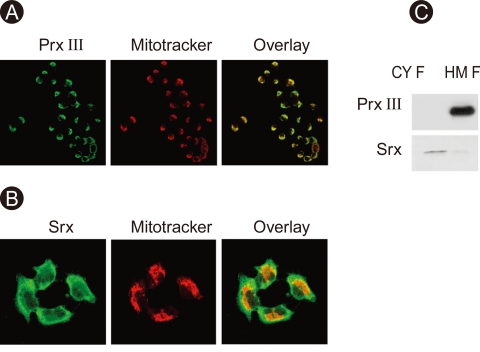
Immunocytochemical analysis followed by confocal microscopy demonstrated the subcellular localization of Prx III and Srx in A549 cells. Prx III co-localized specifically with the Mitotracker dye in the mitochondrial fraction (Fig. 1A). Srx was present in the cytosol under basal conditions (Fig. 1B). The localization of Srx and Prx III was also examined by immunoblot analysis of the soluble cytosolic and heavy membrane fractions. Srx was found predominantly in the cytosol, and Prx III was detected in the mitochondrial fraction (Fig. 1).
Figure 1.
Expression and cellular localization of peroxir-edoxin (prx) III and sulfiredoxin (srx). A549 cells were stained for Prx III (FITC), Srx (FITC), and mitochondrial fraction (Mitotracker) and then examined by confocal microscopy (A, B). A549 cells were subjected to subcellular fractionation to yield mitochondria-enriched heavy membrane (HM) and cytosolic fractions (CYF). Cell lysates were prepared and subjected to immunoblot analysis with antibodies to either Prx III or Srx (C).
Oxidation of Prx III in mink lung epithelial cells
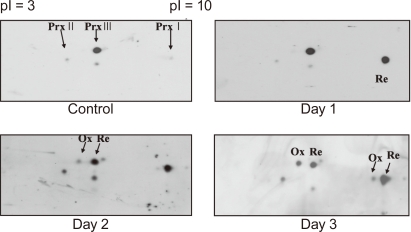
Mink lung epithelial cells (Mv1Lu) were exposed to TGF-β1 (2 ng/mL) for 72 hours, after which only the oxidized form of Prx III was identified by two-dimensional (2D) electrophoresis and immunoblot analysis (Fig. 2).
Figure 2.
Oxidation of peroxiredoxin (Prx) I, II, and III in Mv-1Lu by transforming growth factor (TGF)-β1. Mv1Lu cells were cultured in DMEM containing 10% FBS and were treated with 2 ng/mL TGF-β1 for the periods indicated. Cell lysates were analyzed with 2D gel electrophoresis, followed by immunoblot analysis with indicated antibodies. Ox, oxidation, re, reduced form.
Srx-dependent regulation of Prx III oxidation in A549 cells
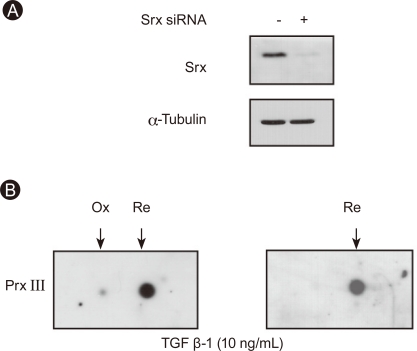
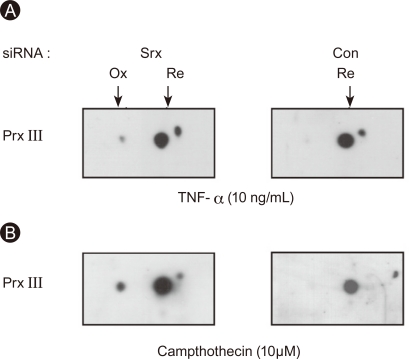
The amount of Srx was reduced by 90% in A549 cells transfected with Srx-specific siRNA (20 nM) compared to control cells (Fig. 3A). A549 cells were treated with TGF-β1 (10 ng/mL) for 72 hours, and the cell lysates were subjected to 2D electrophoresis followed by immunoblot analysis. The sulfinic forms of Prx III were found only in A549 cells depleted of Srx, and oxidized Prx III was observed in A549 cells treated with either TNF-α (10 ng/mL) or camptothecin (10 µM) for 24 hours (Fig. 4).
Figure 3.
Sulfiredoxin (Srx) controls peroxiredoxin (Prx) III oxidation in A549 by transforming growth factor (TGF)-β1. Cells were transfected with either a control siRNA or a siRNA specific for human Srx mRNA, after which cell lysates were prepared and subjected to immunoblot analysis with antibodies to Srx or α-tubulin (A). Two days after cells were transfected as in (A), cells were exposed to 10 ng/mL TGF-β1 for 3 days and cell lysates were analyzed with 2D gel electrophoresis, followed by immunoblot analysis with Prx III antibody (B). Ox, oxidation form; Re, reduced form.
Figure 4.
Sulfiredoxin (Srx) controls peroxiredoxin (Prx) III oxidation in A549 cells by tumor necrosis factor (TNF)-α and camptothecin. Cells were transfected with either control siRNA or siRNA specific for human Srx mRNA. Two days after transfection, cells were exposed to 10 ng/mL TNF-α for 24 hours (A) or 10 µM camptothecin for 24 hours (B) and cell lysates were analyzed with 2D gel electrophoresis, followed by immunoblot analysis with Prx III antibody. Ox, oxidation form; Re, reduced form.
Nrf2-dependent, ARE-mediated expression of proteins in human lung cancer
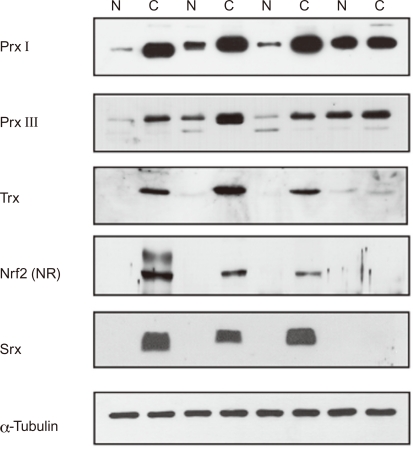
The expression levels of Nrf2, Prx, Trx, and Srx were examined by concomitant immunoblot analysis. The expression levels of Nrf2, Prxs (particularly Prx I and III), Trx, and Srx were higher in lung cancer tissues than in the paired normal lung tissues (Fig. 5). Significant Nrf2 overexpression was observed in the nucleic fraction of lung cancer tissues.
Figure 5.
Increased expression of antioxidant responsive element (ARE)-proteins in human lung cancer tissue (squamous cell carcinoma). Expression of peroxiredoxin (Prx) II, and III as well as thioredoxin (Trx) and sulfiredoxin (Srx) in both lung cancer (C) and paired normal tissue (N) was subjected to immunoblot analysis with antibodies to each, respectively. Expression of nuclear factor E2-related factor 2 (Nrf2) in both lung cancer (C) and paired normal tissue (N) were subjected to subcellular fractionation and the nucleic fraction (NR) was subjected to immunoblot analysis with Nrf2 antibody.
Srx overexpression in human lung cancer
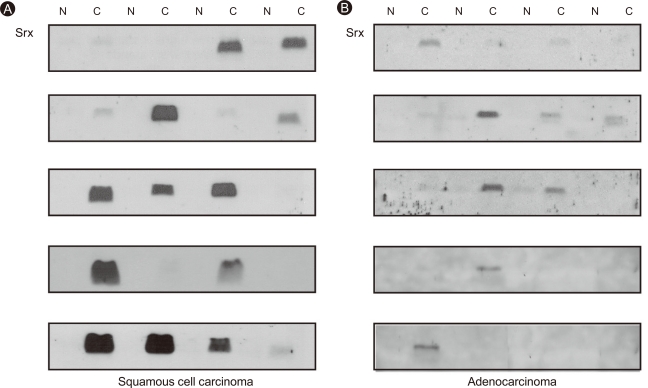
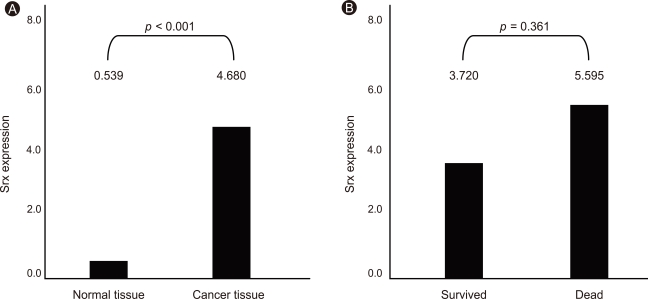
We examined Srx expression in 40 resected stage I and II lung cancers using immunoblot analysis. Markedly increased Srx expression was observed in 60% of squamous cell carcinomas compared to the paired normal lung tissues (p < 0.001, n = 20; Fig. 6A), whereas Srx overexpression occurred in only 20% of adenocarcinomas (p < 0.001, n = 20; Fig. 6B). Srx expression levels in lung cancer and paired normal tissues were quantitated by densitometry (Fig. 7A and 8A), revealing that Srx was significantly overexpressed in both squamous cell carcinoma and adenocarcinoma cell types (Fig. 7B and 8B). Srx protein was markedly overexpressed in all lung cancer subtypes (p < 0.001; Fig. 9A).
Figure 6.
Increased expression of sulfiredoxin (Srx) in human lung cancer tissue: squamous cell carcinoma (A) and adenocarcinoma (B). Expression of Srx in both lung cancer (C) and paired normal tissue (N) were subjected to immunoblot analysis with Srx antibody.
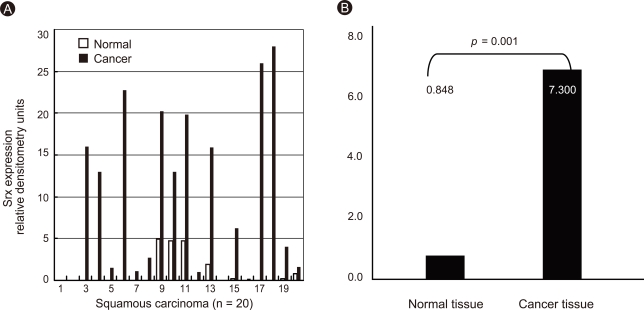
Figure 7.
Expression of sulfiredoxin (Srx) in human squamous cell carcinoma. Individual data were quantified as densitometry units (A) and expressed as relative to the corresponding value for expression of Srx in lung cancer and paired normal tissue (B).
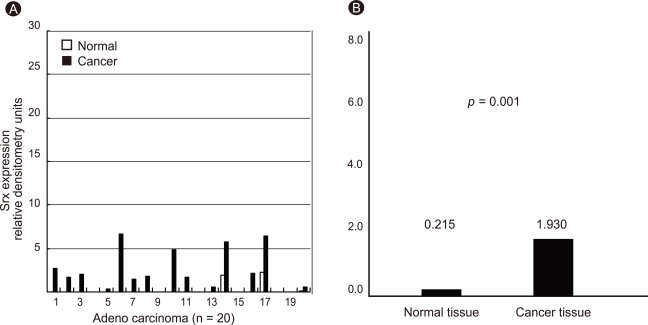
Figure 8.
Expression of sulfiredoxin (Srx) in human adenocarcinoma. Individual data were quantified as densitometry units (A) and expressed as relative to the corresponding value for expression of Srx in lung cancer and paired normal tissue (B).
Figure 9.
Expression of sulfiredoxin (Srx) and survival in all cell types of lung cancer tissue. Data are expressed as relative to the corresponding value for expression of Srx in all lung cancer cell types and paired normal tissue (A). Data are expressed as relative to the corresponding value for expression and survival of Srx in all cell types of lung cancer and paired normal tissue (B).
Srx expression and survival in lung cancer
Srx expression level negatively correlated with patient survival but was not significant (Fig. 9B).
Srx expression in lung cancer tissues
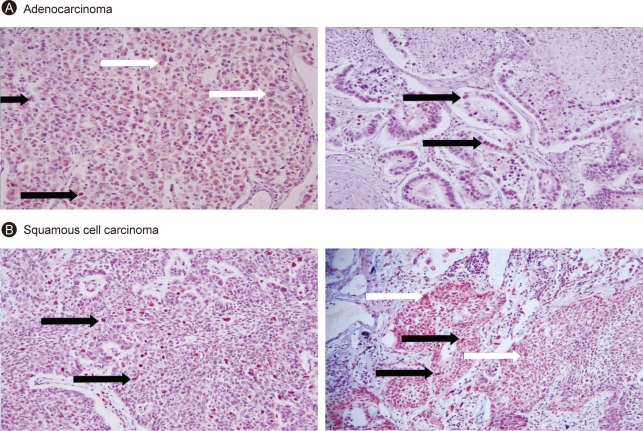
Immunohistochemical staining of paraffin-embedded tissues using an anti-Srx monoclonal antibody demonstrated that the increase in Srx protein originated from cancer tissue; the tumor tissue was much more heavily stained than the normal lung parenchyma. Srx protein was located mainly in the cytoplasm of cancer cells, but was also found in the nuclei and nucleoli (Fig. 10).
Figure 10.
Expression of sulfiredoxin (Srx) in human lung cancer tissue. Paraffin-fixed 5 M slide of lung cancer tissue was deparaffinized, incubated with Srx antibody, and visualized with DAB as the chromogen (H&E, × 400). Immunohistochemistry of adenocarcinoma (A) and squamous cell carcinoma (B) are shown. black arrow indicates nucleus and white arrow indicates cytoplasm.
DISCUSSION
The principal finding of this study is that Srx protein was markedly overexpressed in human lung squamous cell carcinoma. This suggests that Srx, in cooperation with increased levels of Prxs (particularly Prx I and III) and Trx, may protect pulmonary epithelial cells from oxidative injury. Furthermore, a significantly increased level of Nrf2 was found in the nucleic fraction of lung cancer tissues, where it may regulate ARE-mediated protein expression.
It was reported previously that Prxs are expressed abundantly in bronchial and spulmonary epithelial cells [19]. However, the connection between these elevated protein levels and underlying overexpression mechanisms was not clear. Our present study showed that the expression levels of Nrf2, Prxs (particularly Prx I and III), Trx, and Srx were higher in cancer tissues than in paired normal lung tissues. Furthermore, Nrf2 was present at the site where Srx was overexpressed in lung cancer tissues. A previous study revealed that increased expression of Srx, which regenerates inactive 2-Cys Prx, relies on the Nrf2-dependent signaling pathway [17,20-25]. Therefore, we speculated that Nrf2 activation may be a master control for upregulating ARE-mediated protein expression. Our results are consistent with a recent report indicating that oxidative stress can induce Prx6 and Srx expression through Nrf2 in the A549 cell line [2]. However, further experiments with mutated Keap1 are necessary to confirm these findings.
Nrf2 activation resulting from both acute and chronic oxidative stresses attributable to environmental carcinogens in smoke or to acute and chronic inflammation may induce subsequent ARE-mediated overexpression of proteins, initially protecting pulmonary epithelial cells from injury [26-28]. Moreover, the Keap1 mutation and increased ARE-mediated protein transcription have been identified in many patients with head and neck squamous cell carcinoma [17,18,29]. Thus, we believe that Srx overexpression in squamous carcinoma occurs in response to an increased level of the sulfinic form of mitochondrial Prx III, which is produced as a result of oxidative stress caused by carcinogens such as smoke and pollution or by the high growth and metabolic rates of tumors. The higher level of Srx could regenerate sulfinic Prx III to meet increased antioxidant demand for tumor cell survival [22]. Increased Nrf2 expression may be explained in the same way. Srx overexpression was found less frequently in adenocarcinoma, perhaps because adenocarcinoma tends to occur more often in younger, female, non-smokers, or around previous chronic inflammation, and is related to other genetic factors. Further studies are needed to elucidate the role of Srx in human lung carcinogenesis. Inhibition of Srx and inactivation of Prx III may be potential therapeutic strategies for future lung cancer treatment.
As expected, Srx immunohistochemistry revealed greater staining of Srx in cancer tissues than in normal tissues. Previous reports have demonstrated that Srx is expressed in the cytoplasm [23]; however, we also found Srx in nuclei and nucleoli. The role of Srx protein in the nuclei and nucleoli should be verified by further studies.
Oxidative stress from carcinogens increases the intracellular ROS level, which triggers Nrf2-dependent induction of Srx expression. The reduction of peroxides by Prx III results in its hyperoxidation to the sulfinic form (Prx III-SO2). The reverse reaction, which activates PRX, is catalyzed by Srx. Nrf2 expression and ARE-mediated expression of Srx, Prx I and III, and Trx are increased to meet the demands of tumor cell growth.
Our study had some limitations. First, a correlation between patient prognosis and Srx expression was not verified; the observation period and number of subjects were insufficient to reach a statistically significant conclusion. Second, when we compared Nrf2 expression between paired normal lung and cancer tissues, it was not possible to separate subcellular fractions of the tissues; therefore, we were unable to compare Srx in the tissues. Third, in a few cases, Srx, Prx, and Trx expression did not differ between squamous cell carcinoma and paired normal tissues. Because squamous cell carcinoma develops through multifactorial processes and Srx induction or overexpression may participate in more than one pathway depending on the cellular site and individual tissue, Srx, Prx, and Trx expression may not be consistent among cases. In our experience, Srx protein has such a short half-life that it might have already been depleted in some cases.
In conclusion, the Srx and Prx III proteins were markedly overexpressed in a Nrf2-dependent manner in human squamous cell carcinoma, suggesting that these proteins may play a protective role against oxidative injury and compensate for the high rate of mitochondrial metabolism in lung cancer cells.
Acknowledgments
This study was partially supported by a 2002 grant from the Graduate School of Ajou University School of Medicine, Suwon, Korea.
Footnotes
No potential conflict of interest relevant to this article was reported.
References
- 1.Rhee SG. Cell signaling: H2O2, a necessary evil for cell signaling. Science. 2006;312:1882–1883. doi: 10.1126/science.1130481. [DOI] [PubMed] [Google Scholar]
- 2.Chowdhury I, Mo Y, Gao L, Kazi A, Fisher AB, Feinstein SI. Oxidant stress stimulates expression of the human peroxiredoxin 6 gene by a transcriptional mechanism involving an antioxidant response element. Free Radic Biol Med. 2009;46:146–153. doi: 10.1016/reeradbiomed.2008.09.027. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Kang SW, Chae HZ, Seo MS, Kim K, Baines IC, Rhee SG. Mammalian peroxiredoxin isoforms can reduce hydrogen peroxide generated in response to growth factors and tumor necrosis factor-alpha. J Biol Chem. 1998;273:6297–6302. doi: 10.1074/jbc.273.11.6297. [DOI] [PubMed] [Google Scholar]
- 4.Kang SW, Rhee SG, Chang TS, Jeong W, Choi MH. 2-Cys peroxiredoxin function in intracellular signal transduction: therapeutic implications. Trends Mol Med. 2005;11:571–578. doi: 10.1016/j.molmed.2005.10.006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Chang TS, Jeong W, Woo HA, Lee SM, Park S, Rhee SG. Characterization of mammalian sulfiredoxin and its reactivation of hyperoxidized peroxiredoxin through reduction of cysteine sulfinic acid in the active site to cysteine. J Biol Chem. 2004;279:50994–51001. doi: 10.1074/jbc.M409482200. [DOI] [PubMed] [Google Scholar]
- 6.Lei K, Townsend DM, Tew KD. Protein cysteine sulfinic acid reductase (sulfiredoxin) as a regulator of cell proliferation and drug response. Oncogene. 2008;27:4877–4887. doi: 10.1038/onc.2008.132. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Rhee SG, Jeong W, Chang TS, Woo HA. Sulfiredoxin, the cysteine sulfinic acid reductase specific to 2-Cys peroxiredoxin: its discovery, mechanism of action, and biological significance. Kidney Int Suppl. 2007;(106):S3–S8. doi: 10.1038/sj.ki.5002380. [DOI] [PubMed] [Google Scholar]
- 8.Kwak MK, Wakabayashi N, Greenlaw JL, Yamamoto M, Kensler TW. Antioxidants enhance mammalian proteasome expression through the Keap1-Nrf2 signaling pathway. Mol Cell Biol. 2003;23:8786–8794. doi: 10.1128/MCB.23.23.8786-8794.2003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Lau A, Villeneuve NF, Sun Z, Wong PK, Zhang DD. Dual roles of Nrf2 in cancer. Pharmacol Res. 2008;58:262–270. doi: 10.1016/j.phrs.2008.09.003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Cho HY, Jedlicka AE, Reddy SP, et al. Role of NRF2 in protection against hyperoxic lung injury in mice. Am J Respir Cell Mol Biol. 2002;26:175–182. doi: 10.1165/ajrcmb.26.2.4501. [DOI] [PubMed] [Google Scholar]
- 11.Kensler TW, Wakabayashi N, Biswal S. Cell survival responses to environmental stresses via the Keap1-Nrf2-ARE pathway. Annu Rev Pharmacol Toxicol. 2007;47:89–116. doi: 10.1146/annurev.pharmtox.46.120604.141046. [DOI] [PubMed] [Google Scholar]
- 12.Cho HY, Reddy SP, Kleeberger SR. Nrf2 defends the lung from oxidative stress. Antioxid Redox Signal. 2006;8:76–87. doi: 10.1089/ars.2006.8.76. [DOI] [PubMed] [Google Scholar]
- 13.Colburn NH, Kensler TW. Targeting transcription factors for cancer prevention: the case of Nrf2. Cancer Prev Res (Phila) 2008;1:153–155. doi: 10.1158/1940-6207.CAPR-08-0025. [DOI] [PubMed] [Google Scholar]
- 14.Kwak MK, Itoh K, Yamamoto M, Sutter TR, Kensler TW. Role of transcription factor Nrf2 in the induction of hepatic phase 2 and antioxidative enzymes in vivo by the cancer chemoprotective agent, 3H-1, 2-dimethiole-3-thione. Mol Med. 2001;7:135–145. [PMC free article] [PubMed] [Google Scholar]
- 15.Kwak MK, Itoh K, Yamamoto M, Kensler TW. Enhanced expression of the transcription factor Nrf2 by cancer chemopreventive agents: role of antioxidant response element-like sequences in the nrf2 promoter. Mol Cell Biol. 2002;22:2883–2892. doi: 10.1128/MCB.22.9.2883-2892.2002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Osburn WO, Kensler TW. Nrf2 signaling: an adaptive response pathway for protection against environmental toxic insults. Mutat Res. 2008;659:31–39. doi: 10.1016/j.mrrev.2007.11.006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Park JJ, Chang HW, Jeong EJ, et al. Peroxiredoxin IV protects cells from radiation-induced apoptosis in head-and-neck squamous cell carcinoma. Int J Radiat Oncol Biol Phys. 2009;73:1196–1202. doi: 10.1016/j.ijrobp.2008.10.070. [DOI] [PubMed] [Google Scholar]
- 18.Stacy DR, Ely K, Massion PP, et al. Increased expression of nuclear factor E2 p45-related factor 2 (NRF2) in head and neck squamous cell carcinomas. Head Neck. 2006;28:813–818. doi: 10.1002/hed.20430. [DOI] [PubMed] [Google Scholar]
- 19.Park JH, Kim YS, Lee HL, et al. Expression of peroxiredoxin and thioredoxin in human lung cancer and paired normal lung. Respirology. 2006;11:269–275. doi: 10.1111/j.1440-1843.2006.00849.x. [DOI] [PubMed] [Google Scholar]
- 20.Bae SH, Woo HA, Sung SH, et al. Induction of sulfiredoxin via an Nrf2-dependent pathway and hyperoxidation of peroxiredoxin III in the lungs of mice exposed to hyperoxia. Antioxid Redox Signal. 2009;11:937–948. doi: 10.1089/ars.2008.2325. [DOI] [PubMed] [Google Scholar]
- 21.Findlay VJ, Tapiero H, Townsend DM. Sulfiredoxin: a potential therapeutic agent? Biomed Pharmacother. 2005;59:374–379. doi: 10.1016/j.biopha.2005.07.003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Jeong W, Park SJ, Chang TS, Lee DY, Rhee SG. Molecular mechanism of the reduction of cysteine sulfinic acid of peroxiredoxin to cysteine by mammalian sulfiredoxin. J Biol Chem. 2006;281:14400–14407. doi: 10.1074/jbc.M511082200. [DOI] [PubMed] [Google Scholar]
- 23.Noh YH, Baek JY, Jeong W, Rhee SG, Chang TS. Sulfiredoxin translocation into mitochondria plays a crucial role in reducing hyperoxidized peroxiredoxin III. J Biol Chem. 2009;284:8470–8477. doi: 10.1074/jbc.M808981200. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Soriano FX, Baxter P, Murray LM, Sporn MB, Gillingwater TH, Hardingham GE. Transcriptional regulation of the AP-1 and Nrf2 target gene sulfiredoxin. Mol Cells. 2009;27:279–282. doi: 10.1007/s10059-009-0050-y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Woo HA, Jeong W, Chang TS, et al. Reduction of cysteine sulfinic acid by sulfiredoxin is specific to 2-cys peroxiredoxins. J Biol Chem. 2005;280:3125–3128. doi: 10.1074/jbc.C400496200. [DOI] [PubMed] [Google Scholar]
- 26.Hübner RH, Schwartz JD, De Bishnu P, et al. Coordinate control of expression of Nrf2-modulated genes in the human small airway epithelium is highly responsive to cigarette smoking. Mol Med. 2009;15:203–219. doi: 10.2119/molmed.2008.00130. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Rahman I. Antioxidant therapeutic advances in COPD. Ther Adv Respir Dis. 2008;2:351–374. doi: 10.1177/1753465808098224. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Singh A, Ling G, Suhasini AN, et al. Nrf2-dependent sulfiredoxin-1 expression protects against cigarette smoke-induced oxidative stress in lungs. Free Radic Biol Med. 2009;46:376–386. doi: 10.1016/j.freeradbiomed.2008.10.026. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Shibata T, Kokubu A, Gotoh M, et al. Genetic alteration of Keap1 confers constitutive Nrf2 activation and resistance to chemotherapy in gallbladder cancer. Gastroenterology. 2008;135:1358–1368. doi: 10.1053/j.gastro.2008.06.082. [DOI] [PubMed] [Google Scholar]