Abstract
The NF-κB family of transcription factors plays a crucial role in cell activation, survival and proliferation. Its aberrant activity results in cancer, immunodeficiency or autoimmune disorders. Over the past two decades, tremendous progress has been made in our understanding of the signals that regulate NF-κB activation, especially how scaffold proteins link different receptors to the NF-κB-activating complex, the IκB kinase complex. The growing number of these scaffolds underscores the complexity of the signaling networks in different cell types. In this review, we discuss the role of scaffold molecules in signaling cascades induced by stimulation of antigen receptors, G-protein-coupled receptors and C-type Lectin receptors, resulting in NF-κB activation. Especially, we focus on the family of Caspase recruitment domain (CARD)-containing proteins known as CARMA and their function in activation of NF-κB, as well as the link of these scaffolds to the development of various neoplastic diseases through regulation of NF-κB.
Keywords: CARMA1, CARMA2, CARMA3, CARD9, Bcl10, NF-κB, IKK, NEMO
Introduction
Scaffold proteins are defined as molecules that bind to at least two other signaling proteins 1. Scaffold proteins typically do not posses any enzymatic or transcriptional activity, but they have the ability to assemble various combinations of multi-protein complexes, necessary for integration of signals and selective transmission of information from the surface receptors 1, 2. In most cases, scaffolds help to localize signaling molecules to specific parts of cell. They also serve as platforms for assembling enzymes and their substrates, restraining the nonspecific access of enzymes to unwanted substrates and protecting from undesirable cellular effects 3. Some scaffold proteins can have other functions, like coordination of the positive and negative feedback signals or protection of activated proteins from inactivation 1. Interestingly, scaffold proteins may exhibit distinct functions under different physiological conditions 4. Also, multiple scaffold/receptor complexes may exist simultaneously, directing both overlapping and distinct cellular events 2.
To date, a large number of scaffolds have been shown to play a crucial role in activation of the nuclear factor κB (NF-κB) following stimulation of different receptors 5, 6, 7, 8, 9. NF-κB is a family of transcription factors that control cell activation, proliferation and survival. Its activity is tightly regulated by interaction with inhibitory proteins, IκBs, which mask the nuclear localization sequence of NF-κB subunits, thereby sequestering NF-κB in cytoplasm. Upon stimulation, IκB is phosphorylated by the IκB kinase (IKK) complex, followed with ubiquitination-dependent degradation by the 26S proteasome complex. Therefore, NF-κB can be translocated into the nucleus and initiate specific target gene transcription 5, 10. In spite of the tremendous progress in the NF-κB signaling field, the signaling cascades connecting different receptors to the IKK complex remain to be fully determined (Figure 1). Previous studies have revealed the tissue-specific and stimulus-specific roles of many scaffold proteins in regulating NF-κB signaling networks 7, 8, 11, 12. Among these scaffold proteins, a family of Caspase recruitment domain (CARD)-containing scaffold proteins, known as CARD- and membrane-associated guanylate kinase-like domain-containing protein (CARMA), plays critical roles in recruitment and activation of IKK 12, 13. In this article, we discuss the role of CARMA family members and other related CARD-containing scaffold proteins in regulation of NF-κB activation in response to signals induced by the various cell surface receptors.
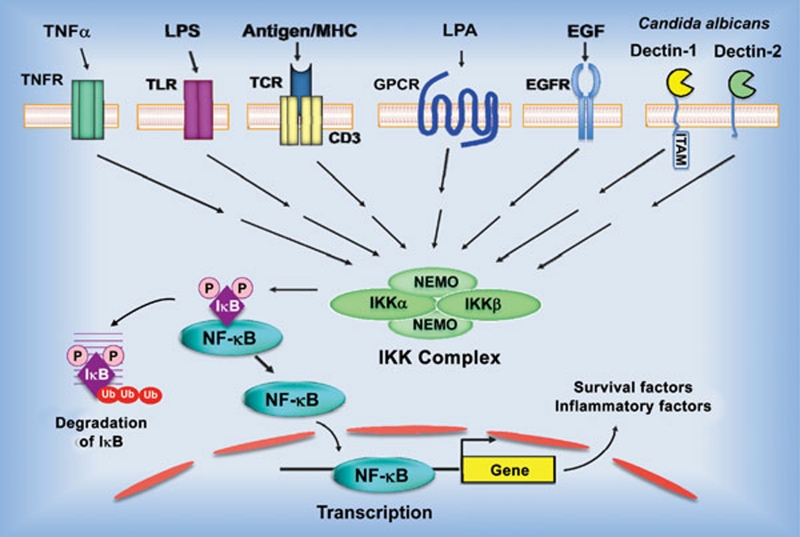
Figure 1.
Model of NF-κB activation by the canonical pathway. Stimulation of the surface receptors by different inducers initiates several proximal signaling events resulting in activation of the IκB kinase (IKK) complex, composed of two kinases, IKKα and IKKβ, and the regulatory subunit NF-κB-essential modulator (NEMO). IKK phosphorylates inhibitor of κB (IκB), which leads to its ubiquitination and subsequent degradation. NF-κB is then translocated into nuclei and initiates the target gene transcription. TNFR – tumor necrosis factor receptor; TLR – toll like receptor, LPS – lipopolysaccharide; TCR – T-cell receptor; GPCR – G protein-coupled receptors; LPA - lysophosphatidic acid; EGFR - epidermal growth factor receptor; ITAM - immunoreceptor tyrosine-based activation motif; Ub - ubiquitin.
CARMA family of scaffold proteins
The CARMA family is conserved among species and has three members, CARMA1, CARMA2, and CARMA3 that are encoded by three different genes 14, 15, 16. They all contain an N-terminal CARD domain, followed with a coiled-coil domain (C-C), a PDZ domain, an SH3 domain, and a Guanylate Kinase-like (GUK) domain in the C-terminus. The structural module of PDZ-SH3-GUK is also called Membrane-associated GUK (MAGUK) domain. CARMA family members were initially identified based on their CARD domain by bioinformatics approaches and named CARD11 (known as CARMA1 or Bimp3), CARD14 (known as CARMA2 or Bimp2), and CARD10 (known as CARMA3 or Bimp1) 14, 16, 17. CARMA1, CARMA2, and CARMA3 share high degree of sequence and structural homology (Figure 2), but they exhibit a distinct tissue distribution pattern. Original studies suggest that CARMA1 is primary expressed in hematopoietic tissues such as spleen, thymus, and peripheral blood leukocyte; CARMA2 is expressed in placenta; and CARMA3 is expressed in a broad range of tissues but not in hematopoietic cells 14, 15, 16, 17. We have analyzed the mRNA microarray data, which were generated from 353 human tissue samples 18 and deposited in the databank (Oncomine 4.4; www.oncomine.org). This analysis revealed that CARMA2 is mainly expressed in mucosa tissues (Figure 3), although the expression pattern of CARMA1 and CARMA3 is consistent with the initial studies 14, 15, 16, 17. This distinct tissue distribution suggests that CARMA family members may have the same function to activate downstream signaling events and play similar roles in different cell types. Indeed, ectopic expression of CARMA family members induces potent activation of NF-κB in most of cell lines, as well as primary cells 6, 15, 19. The overexpressed CARMA1 forms a complex with two downstream signaling molecules, Bcl10 (B-cell lymphoma protein 10), another CARD-containing scaffold protein, and MALT1 (Mucosa-associated lymphoid tissue lymphoma translocation protein 1), a caspase-like protein 20. Previous studies demonstrate that signal-dependent formation of the CARMA1-Bcl10-MALT1 complex (commonly known as the CBM complex) recruits downstream signaling components, leading to the activation of NF-κB 20, 21, 22.
Figure 2.
Structures of CARMA1-3, CARD9, Bcl10 and their tissue distribution. CARMA - caspase recruitment domain (CARD)- and membrane-associated guanylate kinase-like domain-containing protein; Bcl10 - B-cell lymphoma 10; CARD - caspase-recruitment domain; C-C - coiled-coil domain; MAGUK - membrane-associated guanylate kinase (GuK)-like domain; S/T rich – Ser/Thr rich domain.
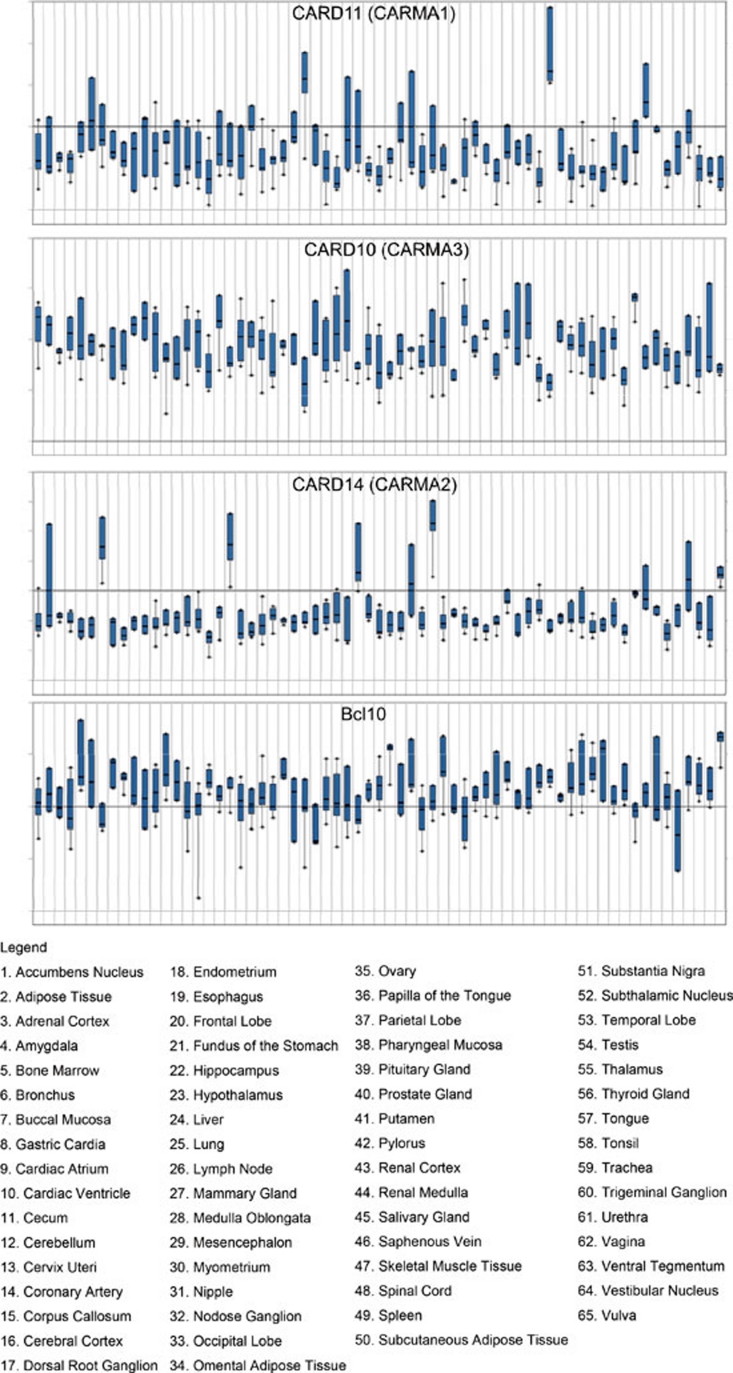
Figure 3.
Expression patterns of CARMA1, CARMA2, CARMA3 and Bcl10 in 65 different human tissues. Microarray data from the public database were analysed using Oncomine 4.4 tools (www.oncomine.org). Log2 median intensity is shown on Y-axis.
CBM proteins in antigen receptor signaling
By inducing somatic mutations in Jurkat T cells, our lab obtained a cell line that lacks CARMA1 protein expression. Using this CARMA1-deficient cells, we demonstrated that CARMA1 is required for the T cell receptor (TCR)-induced NF-κB activity 23. Independent studies using dominant negative mutants of CARMA1 24 or small interfering RNA targeting CARMA1 25 also revealed the crucial role of CARMA1 in NF-κB activation following TCR ligation. Finally, the gene-targeting experiments in mouse have further confirmed that CARMA1 is essential for antigen receptor-induced NF-κB and JNK activation, but not ERK or p38 activation 26, 27, 28. Although the development and survival of mature B and T cells are not significantly affected by CARMA1 deficiency, the signal-induced proliferation of mature cells is severely impaired 26, 27. Similar defects are observed in Bcl10-deficient lymphocytes 29.
Bcl10 was identified by functional cloning from mucosa-associated lymphoid tissue (MALT) lymphoma cells 30, 31 and by bioinformatics approaches as a CARD-containing protein 32, 33, 34. Genetic studies using Bcl10-deficient mice have revealed that Bcl10 is an essential component in the T cell receptor (TCR)- and B cell receptor (BCR)-induced NF-κB activation, and functions downstream of PKC 29, 35. Bcl10 contains an N-terminal CARD domain and a C-terminal Ser/Thr-rich domain (Figure 2). The CARD domain of Bcl10 is responsible for its association with CARMA1 and TCR-induced oligomerization 36, 37. Bcl10 oligomers can function as scaffolds for the IKK and Jun N-terminal kinase (JNK) pathways by recruiting and assembling signaling complexes containing kinases and their substrates 12. In this case, oligomerized Bcl10 associates with JNK2 and its upstream kinases, MKK7 and TAK1, in stimulated Jurkat T cells 38. The high molecular weight complex of oligomerized Bcl10 was also found to regulate TCR-induced actin polymerization 37. Interestingly, Bcl10-dependent actin polymerization has a significant impact on phagocytosis in monocytes, but CARMA1 and MALT1 are not involved in this process 39, 40.
Similarly to Bcl10, MALT1 (also known as Paracaspase) was first identified by genetic cloning from MALT lymphoma patient samples 41 and bioinformatic approach 42. Malt1 gene is localized in a break point of chromosome 18q21 and t(11;18)(q21;q21) generates API2-MALT1 fusion protein, whereas t(14;18)(q32;q21) juxtaposes Malt1 gene to the immunoglobulin locus and upregulates its expression 41, 43, 44. Although transient transfection of wild type MALT1 does not significantly activate NF-κB, overexpression of its oncogenic form potently activates NF-κB in vitro 42, 45. Consistent with these results, transgenic mice expressing Eμ-API2-MALT1 have elevated NF-κB activity 46. On the other hand, genetic inactivation of Malt1 gene in mice impairs TCR-induced NF-κB activation 47, 48. However, there are some discrepancies about the role of MALT1 in BCR-induced NF-κB activation in two mouse models. One study suggests that total NF-κB activity is significantly reduced in MALT1-deficient B cells 47, whereas another gene-targeting study shows that NF-κB activation is almost not affected by MALT1 deficiency upon BCR stimulation 48. Surprisingly, further work has revealed that the activation of c-Rel isoform of NF-κB is more severely impaired than other isoforms, such as RelA (known as p65) 49. Thus, it remains to be determined why MALT1 deficiency has more significant impact for the activation of c-Rel than other isoforms of NF-κB in B cells.
The initial characterization of MALT1 did not reveal its protease activity 42, 45, and MALT1 has been suggested to serve as an E3 ligase for the regulatory subunit of the IKK complex 50. Later, it has been shown that MALT1, through its paracaspase domain but in a protease-independent manner, controls caspase-8 (CASP8) activation, leading to activation of NF-κB and production of IL-2 51. However, an independent study indicates that MALT1 has an arginine-directed protease activity that is induced following TCR stimulation 52. The activated MALT1 can process Bcl10 and regulates TCR-induced cell adhesion to fibronectin 52. This study suggests that MALT1 protease activity is essential for the optimal activation of NF-κB, although it remains to be determined how this activity links to NF-κB. The involvement of MALT1-dependent CASP8 activation in TCR-induced NF-κB activity is consistent with the previous finding that CASP8 is involved in positive regulation of NF-κB activity 53, 54. In addition, CASP8 and c-FLIP are recruited into lipid rafts following with rapid cleavage of c-FLIP at the CASP8 cleavage site 55. Together, these studies suggest that an uncharacterized signaling cascade may connect MALT1, CASP8, and c-FLIP to NF-κB activation. Further work is required to address this possible connection.
The CBM complex in natural killer and mast cells
Besides its critical role in the TCR and BCR signaling, the CBM complex mediates NF-κB activation induced by the multiple immunoreceptor tyrosine-based activation motif (ITAM)-containing receptors 8, 56, 57. In natural killer (NK) cells, activation of the ITAM-coupled receptors leads to CARMA1-, Bcl10- and MALT1-dependent induction of NF-κB and production of proinflammatory cytokines 56, 57. However, the CBM complex is dispensable for NK cell–mediated target cell killing 8, 57. Furthermore, activation of the Fc epsilon receptor I (FcɛRI) on mast cells also engages Bcl10 and MALT1 to activate NF-κB 58, 59. The Bcl10-Malt1 complex promotes IL-6 and tumor necrosis factor (TNF)-α release, which are independent of degranulation and leukotriene secretion 58. However, it remains to be determined whether CARMA1 is also involved in FcɛR-induced signaling events. Together, these studies indicate that CBM proteins play a critical role in multiple receptor signaling pathways.
The mechanism of T cell receptor-induced CARMA1 activation
Although the molecular mechanism is not fully understand, CARMA1 is recruited into the immunological synapse (also known as lipid raft microdomain) upon TCR stimulation 24, 60, 61. This recruitment seems to be dependent on at least two steps. First, CARMA1 needs to be localized to the cytoplasmic membrane 24 and the point mutation in the MAGUK domain of CARMA1 (Leu808 replaced with Pro) impairs its membrane localization and recruitment to the immunological synapse 61. This result is consistent with the function of MAGUK domain that is believed to link proteins to the cytoplasmic membrane 62. The second step for CARMA1 recruitment to the immunological synapse seems to be dependent on its inducible interaction with an adaptor protein known as ADAP (adhesion- and degranulation-promoting adapter protein). Upon TCR engagement, ADAP and CARMA1 are recruited into the immunological synapse, and ADAP deficiency results in impaired CARMA1 translocation leading to reduced NF-κB activation 63. However, ADAP is not a membrane protein and it is unlikely that ADAP anchors the MAGUK domain of CARMA1 to the cytoplasmic membrane. Therefore, another unknown protein may be required for anchoring CARMA1 to the cytoplasmic membrane.
Upon TCR ligation, CARMA1 physically associates with protein kinase C theta (PKCθ), and this association is dependent on the linker region (residues 432–671) between the C-C and PDZ domains of CARMA1 (Figure 2) 61. The linker region is phosphorylated by PKCθ (or PKCβ in B cells) in a signal-dependent manner (Figure 4A) 19, 64 and two putative PKC phosphorylation sites, Ser552 and Ser645 (in mouse: Ser564 and Ser657), were identified and confirmed in the linker region of human CARMA1 19, 64, 65. The inducible phosphorylation of CARMA1 by PKC results in conformational changes that enable CARMA1 to associate with its downstream signaling components 19, 64, 66.
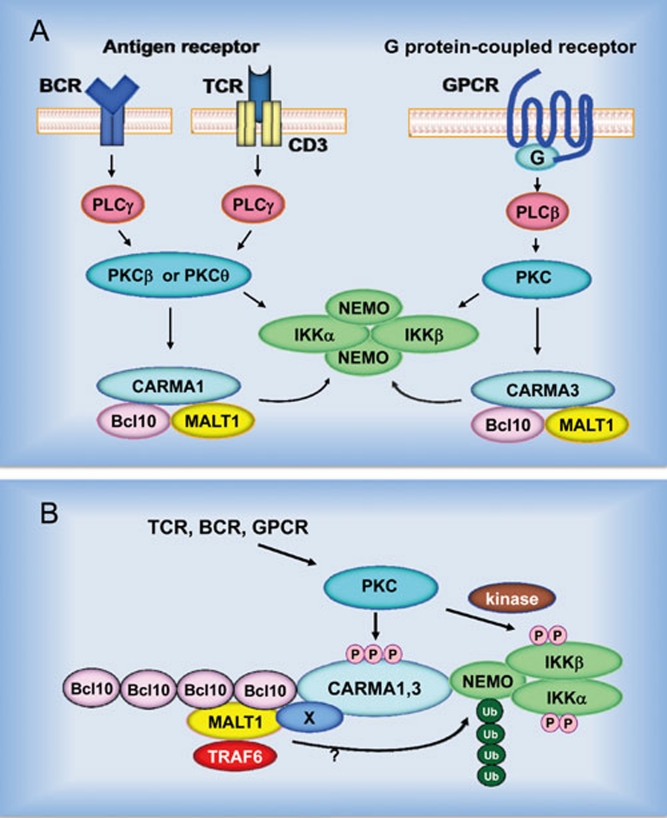
Figure 4.
Schematic model of the B-cell receptor (BCR)-, T-cell receptor (TCR) and G protein-coupled receptor-induced NF-κB activation. (A) Stimulation of the surface receptor initiates several proximal signaling events that lead to activation of phospholipase C (PLC) and protein kinase C (PKC). The activated PKC phosphorylates CARMA and enables CARMA to associate with the downstream signaling components, Bcl10 and MALT1. Formation of the CBM complex leads to activation of the IKK complex. (B) The CBM complex cooperates with tumor necrosis factor receptor-associated factor 6 (TRAF6) and possibly some unidentified protein (X) to mediate ubiquitination of NEMO. The IKK complex is also phosphorylated in a PKC-dependent manner and both modifications are essential for the IKK kinase activity.
The linker region of CARMA1 also contains multiple non-PKC sites and other kinases may be involved in its phosphorylation and regulation. Indeed, PDK1, AKT, TAK1, IKKβ, CK1α, and HPK1 have been shown to associate with or may phosphorylate CARMA1 67, 68, 69, 70, 71. Moreover, some kinases may contribute to CARMA1 phosphorylation outside the linker region. Consistent with this notion, calmodulin-dependent protein kinase II (CaMKII) is recruited to the immunological synapse following TCR stimulation, and phosphorylates CARMA1 on Ser109 72. Mutation of Ser109 to Ala residue in CARMA1 impairs its biological function 72. Once activated, CARMA1 recruits the IKK complex 60, 61 and other signaling molecules (Table 1) 63, 67, 68, 69, 73, including Bcl10 and its pre-associated partner MALT1 60, 61, 74. TCR-induced formation of the CARMA1-Bcl10-MALT1 complex is critical for IKK activation. It is conceivable that CBM functions as a molecular platform to assemble IKK with its direct activators 68, 75.
Table 1. CARMA1-interacting proteins.
| INTERACTING PROTEIN | BIOLOGICAL FUNCTION | REFERENCE |
|---|---|---|
| ADAP | adaptor protein, signal transduction | Medeiros, 2007 (63) |
| AKT | kinase | Narayan, 2006 (69) |
| β-arrestins | adaptor protein, signal transduction | Sun, 2008 (6) |
| Bcl10 | scaffold, signal transduction | Bertin, 2001 (14); Gaide, 2001 (15) |
| CaMKII | kinase | Ishiguro, 2006 (72) |
| Cbl-b | E3 ligase | Kojo, 2009 (104) |
| CK1α | kinase | Bidere, 2009 (70) |
| COP9 (CNS5) | adaptor protein, signal transduction | Welteke, 2009 (106) |
| HPK1 | kinase | Brenner, 2009 (71) |
| IKKβ | kinase | Wegener, 2006 (21); Shinohara, 2007 (65) |
| MALT1 | paracaspase, protease | Che, 2004 (74); Wegener, 2006 (21) |
| NEMO | regulatory protein | Stilo, 2004 (92); Shambharkar, 2007 (79) |
| PDK1 | kinase | Lee, 2005 (67) |
| PKCθ | kinase | Wang, 2004 (61); Matsumoto, 2005 (19) |
| TAK1 | kinase | Shinohara, 2005 (68) |
The mechanism of CARMA1-dependent activation of the IKK complex
NF-κB is activated by at least two signaling pathways: the canonical pathway and the non-canonical pathway. These two pathways utilize different IKK complexes to activate downstream signaling. The canonical pathway is activated by the IKK complex containing two catalytic subunits, IKKα and IKKβ, and the regulatory subunit, NF-κB-essential Modulator (NEMO, also known as IKKγ) 76, 77, 78, which phosphorylates IκBs and induces their degradation. In contrast, the non-canonical pathway is activated by the IKK complex containing IKKα homodimer, which phosphorylates p100 leading to proteolytic processing of p100 into p52 10. Our previous study indicates that CARMA1 is involved in regulating the canonical pathway 79. Although there is no report implicating a role of CARMA1 or Bcl10 in the non-canonical pathway, one study suggests that MALT1 may be required for BAFF receptor-induced p100 processing 80.
The activation of the IKK complex is dependent on the signal-induced phosphorylation of IKKα and IKKβ on two Ser residues (Ser176/180 for IKKα and Ser177/181 for IKKβ) within the activation loop of the kinase domain 81, 82. Although many kinases were shown to phosphorylate IKKs when overexpressed in cells, the genetic evidence indicates that two MAP3K kinases, TAK1 (TGFβ−activated kinase 1) 83, 84, 85, 86 and MEKK3 (Mitogen-activated Protein Kinase Kinase Kinase 3) 87, 88, are involved in phosphorylation of these two Ser residues in IKKα and IKKβ. Accumulating evidence indicates that TAK1 83, 84, 85, 86 and MEKK3 88, 89, 90, 91 are involved in TNFR-, IL-1βR-, TLR-, and TCR-induced IKK phosphorylation and activation. However, CARMA1 does not control this signal-induced phosphorylation of IKK, instead, CARMA1 directly associates with NEMO 92, and modulates the polyubiquitination of NEMO upon TCR stimulation (Figure 4B) 79.
Although NEMO has no catalytic activity, it is required for the kinase activity of the IKK complex in the canonical NF-κB pathway. It has been shown that NEMO can specifically recognize Lysine 63 (K63)-linked polyubiquitin chains 93, 94, 95, and becomes polyubiquitinated upon activation of NF-κB signaling cascades 50, 96. These two properties of NEMO seem to be required for activation of the IKK complex. It is also possible that K63-linked NEMO polyubiquitination allows NEMO oligomerization through cross-recognition by its own ubiquitin-binding domain and leads to generation of the high molecular weight IKK complex 97. Interestingly, several studies suggest that K63-linked polyubiquitination of NF-κB signaling components, such as RIP1 93, 98, IRAK 99, Bcl10 95, and MALT1 100, might be recognized by NEMO following stimulation of different receptors. Indeed, it has been shown that TCR ligation leads to K63-linked polyubiquitination of Bcl10 at Lys31 and Lys63 in the CARD domain, and this polyubiquitin chain can be recognized by NEMO 95. Thus, polyubiquitinated Bcl10 may be involved in a signal-dependent IKK redistribution and possibly activation 36, 95. Also, all these results suggest that NEMO might be responsible for the recruitment of the IKK complex to the specific subcellular location and/or to the specific signaling complex. However, how NEMO is involved in activation of the IKK complex still remains to be determined.
The CBM complex most likely serves as a molecular platform to recruit signaling components responsible for the K63-linked polyubiquitination of NEMO 50. Interestingly, although IKKβ phosphorylation is not defective in stimulated CARMA1- or Bcl10-deficient cells, the kinase activity of IKK is completely abolished 79, suggesting that the phosphorylation of IKK is not sufficient to induce its kinase activity. Indeed, activation of the IKK complex is not only dependent on IKK phosphorylation but also on CARMA1-dependent NEMO modification 79. Initially, the physical interaction of NEMO with CARMA family members, was identified by the yeast two-hybrid screening and confirmed by co-precipitation experiments in mammalian cells 92. Our later study has revealed that NEMO is polyubiquitinated in a CARMA1-dependent manner upon TCR engagement 79, although the mechanism by which CARMA1 regulates NEMO polyubiquitination is not fully defined. One study suggests that the CBM complex recruits TRAF6, and TRAF6 induces K63-linked polyubiquitination of NEMO upon TCR stimulation 75. Alternatively, MALT1 may induce NEMO ubiquitination at the Lys399 (Lys392 in mice) residue, because NEMO variant with K399R mutation has been shown to interfere with NF-κB activation 50. However, mice expressing the NEMO-K392R mutant are not defective in antigen receptor-induced responses 101. Therefore, it remains to be determined whether MALT1-dependent polyubiquitination of NEMO is functionally important for TCR-induced responses. In addition, although TRAF6 has been suggested to mediate TCR-induced polyubiquitination of the IKK complex 75, T cell-specific deletion of TRAF6 does not impair TCR-induced NF-κB activation 102, suggesting that either there is a redundant mechanism for TRAF6-mediated activation of IKK or TRAF6 is not involved in the activation of IKK (Figure 4B). Because CARMA1-dependent NEMO polyubiquitination is required for IKK activation, revealing the mechanism by which CARMA1 regulates NEMO polyubiquitination should provide further insight about the regulation of the IKK complex.
Negative regulation of CARMA1 and Bcl10 proteins
Posttranslational modifications contribute to the regulation of the expression level of CARMA1. Recent studies indicate that phosphorylation of some residues in CARMA1 may suppress CARMA1 function 70, 103. The Ser637 residue seems to be phosphorylated by PKC isoforms other than PKCθ or PKCβ, and mutation of Ser637 to Ala enhances CARMA1-induced NF-κB activation 103, suggesting that Ser637 phosphorylation negatively regulates CARMA1 function. Another study demonstrates that CARMA1 phosphorylation by casein kinase 1α (CK1α) leads to the attenuation of CBM-mediated NF-κB activity 70. The Ser608 residue within the CARMA1 linker region has been identified as a CK1α phosphorylation site 70, however, the mechanism by which Ser608 phosphorylation suppresses CARMA1 activity remains unclear.
Recent studies indicate that ubiquitination may also regulate the function of CBM proteins. Formation of the CBM complex appears to be negatively regulated by the E3 ligase Cbl-b 73. It has been shown that Cbl-b mediates monoubiquitination of CARMA1, which disrupts the CARMA1-Bcl10 interaction without affecting CARMA1 protein stability 104. On the other hand, K48-linked polyubiquitination leads to the proteasome-mediated degradation of CARMA1 105. The C-terminal MAGUK region of CARMA1 seems to be involved in this regulation, since SH3 and GUK domains contain the ubiquitin acceptor sites. In vitro experiments demonstrate that the cellular inhibitor of apoptosis (cIAP) might be an E3 ligase for CARMA1 105. Finally, one study shows that CARMA1 interacts with members of the COP9 signalosome, CNS2 and CNS5 106. This work suggests that COP9 regulates IKK activity by maintaining stability of the CBM complex and protecting Bcl10 from degradation 106.
Posttranslational modifications also regulate the level of Bcl10. Although the function of the C-terminal Ser/Thr rich domain of Bcl10 is not fully determined, several studies suggest that signal-dependent phosphorylation of these Ser and Thr residues may mediate degradation of Bcl10 21, 107, 108, 109, thereby terminating NF-κB activation 109. Indeed, Bcl10-deficient T cells reconstituted with the Bcl10-S138A mutant have prolonged NF-κB activation and enhanced IL-2 production 108. However, the mechanism by which Bcl10 stability is regulated remains to be defined.
The role of CBM proteins in lymphoma
Previous studies suggest that CBM proteins are involved in lymphoma pathogenesis 30, 31, 41, 110. Chromosomal translocations, which lead to the overexpression of Bcl10 and MALT1 or generation of API2-MALT1 fusion protein, were found in MALT lymphoma 30, 41, 43, and the activation of NF-κB by these oncogenic proteins is believed to be one of the hallmarks of MALT lymphoma. Consistent with this concept, transgenic mice expressing Eμ-Bcl10 have splenic B-cell expansion and develop marginal zone B-cell lymphoma 111.
Although the CARMA1 gene is not commonly rearranged in B- or T-cell lymphomas, elevated CARMA1 expression was found in adult T cell leukemia 112, primary gastric B cell lymphoma 113 and diffuse large B cell lymphoma (DLBCL) 110. Importantly, CARMA1 overexpression leads to its oligomerization through the C-C domain and activation of the downstream signaling cascades 114. Recently, pathogenic oligomerization of CARMA1 with subsequent activation of the CBM complex has been found in the activated B-cell (ABC) subtype of DLBCL 115. This pathogenic CARMA1 oligomerization results from mutations within exons encoding the C-C domain 115. Screening of patient samples performed independently by three groups has revealed that CARMA1 is mutated in about 10% of systemic ABC-DLBCL and 16% of primary central nervous system DLBCL 115, 116, 117. The oncogenic mutant of CARMA1 constitutively recruits downstream signaling components 118, and likely induces proteolytic activity of MALT1 119, 120, leading to activation of NF-κB 115.
Previous studies have shown that NF-κB activity is critical for the survival of malignant cells in ABC-DLBCL 121, and IKK inhibitors 122 or CARMA1 shRNA 110 are toxic for these cells. Therefore, CARMA1 is considered to be an attractive target for the development of specific anti-lymphoma drugs. Although the detailed mechanism of oncogenic function of CARMA1 remains to be fully elucidated, several lines of evidence are consistent with the hypothesis that mutations and/or overexpression of CARMA1 contribute to the lymphoma cell survival. Further studies are needed to determine whether mutation of CARMA1 alone is sufficient to initiate lymphoma and contributes to the malignant phenotype, such as dissemination, of lymphoma.
CARMA3-mediated NF-κB signaling pathways
CARMA3 is expressed widely in non-hematopoietic cells and has recently been described as a link between G protein-coupled receptors (GPCRs) and NF-κB 123. GPCR is the largest class of transmembrane receptors in the human genome involved in regulation of proliferation, differentiation, and immune response through the wide variety of its ligands 124. One of them is lysophosphatidic acid (LPA), a bioactive phospholipid that is a component of normal plasma and biological fluids, such as saliva and bronchoalveolar fluid 125, 126. LPA is capable of inducing diverse cellular responses by inducing activity of several transcription factors, including NF-κB and AP-1 127.
Genetic deletion of CARMA3 results in diminished LPA-induced NF-κB activation and subsequent IL-8 production in mouse embryonic fibroblasts (MEF) 123. Similar defect is observed following stimulation with other GPCR ligands, such as endothelin-1 and angiotensin-II, in the absence of CARMA3 123, 128, 129. Importantly, CARMA3 is specifically required for GPCR-induced IKK activity because CARMA3 deficiency does not affect IKK activation by other stimuli such as TNFα and lipopolysaccharide (LPS) 123.
Recent study also demonstrates that inhibition of CARMA3 in airway epithelial cells reduces LPA-mediated NF-κB activity and the production of NF-κB-dependent cytokines, TSLP and CCL20 130. Both cytokines are produced by airway epithelial cells and play important role in initiating allergic inflammation 131, 132. Furthermore, forced expression of a dominant-negative CARMA3 mutant (CARD truncation) or treatment of cells with siRNA specifically targeting CARMA3 abrogates LPA-induced signaling in ovarian cancer cells 133.
Similarly to CARMA3, Bcl10- and MALT1-deficient cells have defective NF-κB activation following LPA treatment 134, 135. These results indicate that the CARMA3-Bcl10-MALT1 complex may play an analogues role in non-hematopoietic cells as the CARMA1-Bcl10-MALT1 complex in the TCR and BCR pathways in hematopoietic cells (Figure 4A). Consistent with this possibility, GPCR-induced ubiquitination of NEMO and activation of the IKK complex is completely defective in the absence of CARMA3 but IKK phosphorylation is intact in these cells 123. Thus, similarly to CARMA1, CARMA3 mediates the signal-induced polyubiquitination of NEMO (Figure 4B). Interestingly, TRAF6 deficiency also abrogates GPCR-induced NF-κB activation 123. Therefore, TRAF6 might function as an E3 ligase to induce K63-linked poly-ubiquitination of NEMO leading to activation of the IKK complex. However, further experimental evidence is needed to prove this hypothesis.
GPCR-mediated signaling leads to phosphorylation and activation of different PKC isoforms and pretreatment with PKC inhibitors suppresses GPCR-induced NF-κB activation 6, 123. To date, it is not clear whether any specific PKC isoform is responsible for phosphorylation of CARMA3. It has been reported that PKCδ mediates NF-κB activation and IL-8 secretion in response to LPA stimulation in bronchial epithelial cells 136. Moreover, LPA activates PKCα and induces RAS–PKCα interaction, causing NF-κB activation via the CARMA3-BCL10-MALT1 signaling complex in ovarian cancer cells 133. Therefore, an outstanding question is whether any isoform of PKCs directly phosphorylates CARMA3 and, if it does, which residue of CARMA3 is phosphorylated by PKC. Interestingly, our previous study demonstrates that ectopic expression of CARMA3 in CARMA1-deficient T cells restores TCR-induced NF-κB activation 19, and mutation of Ser520 in CARMA3 (an analogue to Ser552 in CARMA1) to the Ala residue diminished CARMA3's ability to rescue CARMA1 deficiency in T cells 19. Although Ser520 might be an important phosphorylation site, it is possible that multiple residues in CARMA3 are phosphorylated by PKC, and other kinases may also contribute to CARMA3 activation in the GPCR pathways.
Our previous study has investigated how CARMA3 is linked to the upstream signaling components in the GPCR-induced cascades and found that CARMA3 interacts with β-arrestin-2 6. Upon stimulation, GPCR recruits and associates with the multifunctional scaffold molecules, β-arrestins 2. These proteins were initially considered as components to desensitize GPCR activation but more recent studies indicate that β-arrestins mediate the signal transduction to NF-κB 137. Although there are four members of the arrestin family in the human genome, only β-arrestin 1 and β-arrestin 2 are ubiquitously expressed in most tissues and function downstream of GPCRs 137. The study from our laboratory demonstrates that β-arrestin 2, but not β-arrestin 1, is required for LPA-induced NF-κB activation and subsequent IL-6 expression 6. Mechanistically, β-arrestin 2 associates with CARMA3 and most likely recruits CARMA3 into the receptor complex. Similar to CARMA3-deficient cells, GPCR-induced IKK kinase activity is completely defective in β-arrestin 2 KO mouse embryonic fibroblasts 6.
CBM proteins in the receptor tyrosine kinase pathways
Several growth factors, including Epidermal Growth Factor (EGF) 138, 139, 140, Insulin-like Growth Factor (IGF) 141, 142, 143, Platelet-Derived Growth Factor (PDGF) 144, 145, and Fibroblast Growth Factor (bFGF) 146, 147, can also induce weak, but notable NF-κB activation through their receptors that belong to a family of receptor tyrosine kinases (RTKs). Although the signaling pathways induced by this family of receptors have been intensively studied, the mechanism by which RTKs activate NF-κB is not fully defined, and the functional significance of RTK-induced NF-κB activation in cell proliferation and survival has not been fully appreciated. Our recent studies indicate that CARMA3 and Bcl10 are required for EGFR-induced NF-κB activation and cancer progression (Jiang and Lin, unpublished data). Therefore, further studies are required to determine how CARMA3 and Bcl10 are involved in the signaling pathways induced by EGFR or other RTKs.
CARD9-mediated NF-κB signaling pathways
CARD9 is another CARD-containing protein and has some similarity to CARMA family members. It was identified through a database search for CARD-containing proteins and shown to interact with Bcl10 148. Its expression seems to be restricted to myeloid cells, mainly macrophages, dendritic cells and neutrophils 148, 149, 150. Although CARD9 is structurally related to the CARMA family, it lacks the C-terminal MAGUK domain (Figure 2) that determinates plasma membrane localization. Therefore, CARD9 may localize in cytosol and be recruited to the receptor complex. Recently, three groups independently generated CARD9-deficient mice and revealed its essential role in the control of innate immunity 7, 149, 150.
Initial characterization of CARD9-deficient mice indicates that CARD9 is required for anti-fungal immune responses 149. This study shows that CARD9 is required for NF-κB activation induced by zymosan, a β-glucan component from yeast cell wall 149. Because zymosan can activate the signaling pathways induced by Dectin-1, a C-type lectin receptor, it has been proposed that CARD9 mediates Dectin-1-induced NF-κB activation 149. However, recent studies from our laboratory have surprisingly found that zymosan can still effectively activate NF-κB in macrophages from an independently generated CARD9-deficient mouse strain 151, suggesting that Dectin-1-induced NF-κB activation may also be induced through a CARD9-independent pathway 151. Therefore, the requirement of CARD9 in Dectin-1-induced NF-κB activation needs to be further investigated. Besides the Dectin-1 pathway, it has been demonstrated that CARD9 also mediates the signaling induced by other C-type lectin receptors such as Dectin-2 151, 152 and Mincle 153. These receptors have been implicated to act as the pattern recognition receptors for fungi 154, 155.
It has been suggested that CARD9 functions downstream of the tyrosine kinase Syk and couples Bcl10/MALT1 to activate classical NF-κB 7, 151, 152, 156, 157. Similarly to CARMA1 and CARMA3, CARD9 is required for activation of the IKK complex 151. Consistent with this model, CARD9-deficient macrophages and dendritic cells have impaired expression of TNF-α, IL-6, and IL-12 in response to the stimulation with fungal particles 149, 151, 152, 157 and CARD9-deficient mice are more susceptible to infection with the fungus Candida albicans 149, 151, 152, 156, 158.
Published studies suggest that Dectin-1 preferentially binds the yeast-like, unicellular form of C. albicans, whereas the Dectin-2 signaling is induced by the hyphal form 152, 158. The proximal signaling cascade leads to a sequential activation of Syk, CARD9, and IKK following stimulation of these C-type lectin receptors (Figure 5) 13. However, the latest work argues that although Syk is required for both Dectin-1- and Dectin-2-induced NF-κB activation, CARD9 is only required for Dectin-2-induced NF-κB activation 151. In addition, Syk and CARD9 do not seem to function in a linear cascade as Syk mediates IKK phosphorylation, whereas CARD9 controls NEMO polyubiquitination 151.
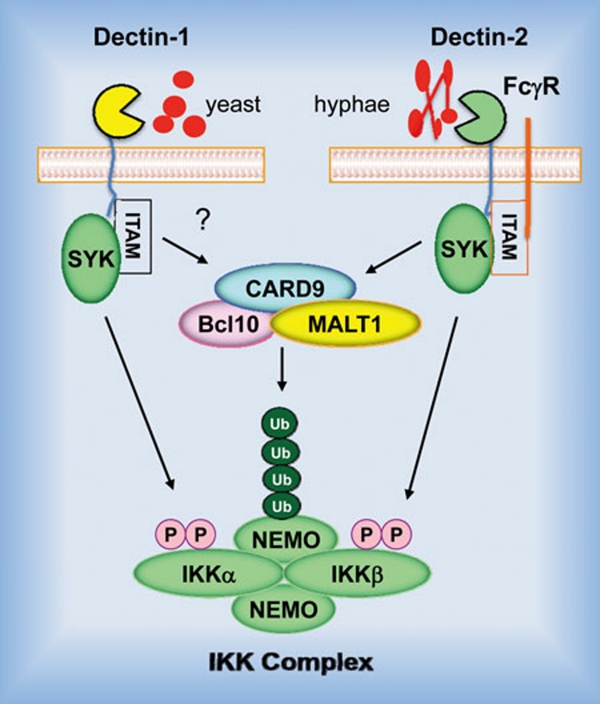
Figure 5.
CARD9 in the C-type lectin receptor signaling. Activation of the Dectin-1 receptor by the yeast-like, unicellular form of C. albicans or the Dectin-2 receptor by the hyphal form of C. albicans leads to a sequential activation of tyrosine kinase Syk, the CARD9-Bcl10-MALT1 complex and IKK. In this signaling cascade Syk mediates IKK phosphorylation, whereas CARD9 controls NEMO polyubiquitination.
Recent studies also suggest that CARD9 mediates the NF-κB activation induced by several ITAM-associated receptors, including those associating with FcRγ and DAP12 in myeloid cells 7. Interestingly, it has been proposed that the CARD9-containing complex mediates NF-κB activation induced by the ITAM-associated receptors in myeloid cells, whereas the CARMA1-containing complex mediates NF-κB activation induced by ITAM-associated receptors in lymphoid lineage cells 8. However, it remains to be determined why ITAM-associated receptors in myeloid cells utilize the CARD9-dependent, but not CARMA1-dependent, pathway to activate NF-κB, given that CARMA1 is also expressed in myeloid cells.
Other functions of CARD9
Several studies also demonstrate that CARD9 is involved in immunity to intracellular bacteria Listeria monocytogenes7, 150, 159 and Mycobacteruim tuberculosis 160. The latest work suggests that CARD9 controls the production of reactive oxygen species (ROS) by regulating the LyGDI-Rac1 complex following the phagocytosis of microorganisms by macrophages 159. CARD9 is also found to be involved in anti-viral responses 150, 161. The CARD9-Bcl10 module appears to be an essential component of the RNA helicase RIG-I-dependent proinflammatory response to certain RNA viruses that leads to IL-1β production 161.
Another study links CARD9 to kidney cancer 162. It has been shown that tumor suppressor VHL associates with CARD9 and promotes inhibitory phosphorylation of CARD9 by casein kinase 2 (CK2). Inactivation of the VHL gene is often observed in renal cell carcinoma and may lead to increased NF-κB activity. Ectopic expression of CARD9 mutants that can not be phosphorylated leads to increased NF-κB activity and decreased apoptosis in VHL-defective renal carcinoma cells, whereas knockdown of CARD9 suppresses NF-κB activity in these cells 162. Although the role of CARD9 phosphorylation by CK2 is not clear, the authors speculate that this modification may promote an inhibitory intra- or inter-molecular interaction or prevent the binding of CARD9 to NEMO or other proteins required for its activity 162. One discrepancy of this regulatory mechanism of posttranslational CARD9 modification with previous results is that CARD9 is mainly expressed in myeloid cells and the expression level of CARD9 in renal cells is very low. Nevertheless, future studies are needed to investigate the role of CARD9 in non-myeloid cells.
Concluding remarks
Significant progress towards understanding the function of the CARMA family of scaffold proteins in the NF-κB signaling pathway has been made during the past several years. However, more research is clearly required to define the CARMA-mediated signaling and the role of these proteins in disease, especially in cancer. Still, the open question is the precise molecular mechanism by which CBM activates the IKK complex. Although CARMA/CARD9-dependent NEMO polyubiquitination seems to be important for IKK activity, it is still possible that CBM mediates some additional modifications of the IKK complex members. Moreover, although previous studies indicate that CARMA2 is exclusively expressed in placenta and mucosal cells, the role of CARMA2 is completely unknown. Therefore, further studies are needed to reveal the role of CARMA2 in these tissues.
Acknowledgments
This work was supported by grants from the National Institutes of Health (RO1GM065899, RO1GM079451, RO1AI050848) to XL. MB is a Special Fellow of The Leukemia and Lymphoma Society.
References
- Shaw AS, Filbert EL. Scaffold proteins and immune-cell signalling. Nat Rev Immunol. 2009;9:47–56. doi: 10.1038/nri2473. [DOI] [PubMed] [Google Scholar]
- Defea KA.Beta-arrestins as regulators of signal termination and transduction: How do they determine what to scaffold Cell Signal 2010. Oct 12. doi: 10.1016/j.cellsig.2010.10.004 [DOI] [PubMed]
- Burack WR, Cheng AM, Shaw AS. Scaffolds, adaptors and linkers of TCR signaling: theory and practice. Curr Opin Immunol. 2002;14:312–316. doi: 10.1016/s0952-7915(02)00347-3. [DOI] [PubMed] [Google Scholar]
- Locasale JW, Shaw AS, Chakraborty AK. Scaffold proteins confer diverse regulatory properties to protein kinase cascades. Proc Natl Acad Sci USA. 2007;104:13307–13312. doi: 10.1073/pnas.0706311104. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hayden MS, Ghosh S. Shared principles in NF-kappaB signaling. Cell. 2008;132:344–362. doi: 10.1016/j.cell.2008.01.020. [DOI] [PubMed] [Google Scholar]
- Sun J, Lin X. Beta-arrestin 2 is required for lysophosphatidic acid-induced NF-kappaB activation. Proc Natl Acad Sci USA. 2008;105:17085–17090. doi: 10.1073/pnas.0802701105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hara H, Ishihara C, Takeuchi A, et al. The adaptor protein CARD9 is essential for the activation of myeloid cells through ITAM-associated and Toll-like receptors. Nat Immunol. 2007;8:619–629. doi: 10.1038/ni1466. [DOI] [PubMed] [Google Scholar]
- Hara H, Ishihara C, Takeuchi A, et al. Cell type-specific regulation of ITAM-mediated NF-kappaB activation by the adaptors, CARMA1 and CARD9. J Immunol. 2008;181:918–930. doi: 10.4049/jimmunol.181.2.918. [DOI] [PubMed] [Google Scholar]
- Sun SC, Ley SC. New insights into NF-kappaB regulation and function. Trends Immunol. 2008;29:469–478. doi: 10.1016/j.it.2008.07.003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hacker H, Karin M. Regulation and function of IKK and IKK-related kinases. Sci STKE. 2006;2006:re13. doi: 10.1126/stke.3572006re13. [DOI] [PubMed] [Google Scholar]
- Simeoni L, Kliche S, Lindquist J, Schraven B. Adaptors and linkers in T and B cells. Curr Opin Immunol. 2004;16:304–313. doi: 10.1016/j.coi.2004.03.001. [DOI] [PubMed] [Google Scholar]
- Blonska M, Lin X. CARMA1-mediated NF-kappaB and JNK activation in lymphocytes. Immunol Rev. 2009;228:199–211. doi: 10.1111/j.1600-065X.2008.00749.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hara H, Saito T. CARD9 versus CARMA1 in innate and adaptive immunity. Trends Immunol. 2009;30:234–242. doi: 10.1016/j.it.2009.03.002. [DOI] [PubMed] [Google Scholar]
- Bertin J, Wang L, Guo Y, et al. CARD11 and CARD14 are novel caspase recruitment domain (CARD)/membrane-associated guanylate kinase (MAGUK) family members that interact with BCL10 and activate NF-kappa B. J Biol Chem. 2001;276:11877–11882. doi: 10.1074/jbc.M010512200. [DOI] [PubMed] [Google Scholar]
- Gaide O, Martinon F, Micheau O, et al. Carma1, a CARD-containing binding partner of Bcl10, induces Bcl10 phosphorylation and NF-kappaB activation. FEBS Lett. 2001;496:121–127. doi: 10.1016/s0014-5793(01)02414-0. [DOI] [PubMed] [Google Scholar]
- Wang L, Guo Y, Huang WJ, et al. Card10 is a novel caspase recruitment domain/membrane-associated guanylate kinase family member that interacts with BCL10 and activates NF-kappa B. J Biol Chem. 2001;276:21405–21409. doi: 10.1074/jbc.M102488200. [DOI] [PubMed] [Google Scholar]
- McAllister-Lucas LM, Inohara N, Lucas PC, et al. Bimp1, a MAGUK family member linking protein kinase C activation to Bcl10-mediated NF-kappaB induction. J Biol Chem. 2001;276:30589–30597. doi: 10.1074/jbc.M103824200. [DOI] [PubMed] [Google Scholar]
- Roth RB, Hevezi P, Lee J, et al. Gene expression analyses reveal molecular relationships among 20 regions of the human CNS. Neurogenetics. 2006;7:67–80. doi: 10.1007/s10048-006-0032-6. [DOI] [PubMed] [Google Scholar]
- Matsumoto R, Wang D, Blonska M, et al. Phosphorylation of CARMA1 plays a critical role in T Cell receptor-mediated NF-kappaB activation. Immunity. 2005;23:575–585. doi: 10.1016/j.immuni.2005.10.007. [DOI] [PubMed] [Google Scholar]
- Lin X, Wang D. The roles of CARMA1, Bcl10, and MALT1 in antigen receptor signaling. Semin Immunol. 2004;16:429–435. doi: 10.1016/j.smim.2004.08.022. [DOI] [PubMed] [Google Scholar]
- Wegener E, Oeckinghaus A, Papadopoulou N, et al. Essential role for IkappaB kinase beta in remodeling Carma1-Bcl10-Malt1 complexes upon T cell activation. Mol Cell. 2006;23:13–23. doi: 10.1016/j.molcel.2006.05.027. [DOI] [PubMed] [Google Scholar]
- Wegener E, Krappmann D. CARD-Bcl10-Malt1 signalosomes: missing link to NF-kappaB. Sci STKE. 2007;2007:pe21. doi: 10.1126/stke.3842007pe21. [DOI] [PubMed] [Google Scholar]
- Wang D, You Y, Case SM, et al. A requirement for CARMA1 in TCR-induced NF-kappa B activation. Nat Immunol. 2002;3:830–835. doi: 10.1038/ni824. [DOI] [PubMed] [Google Scholar]
- Gaide O, Favier B, Legler DF, et al. CARMA1 is a critical lipid raft-associated regulator of TCR-induced NF-kappa B activation. Nat Immunol. 2002;3:836–843. doi: 10.1038/ni830. [DOI] [PubMed] [Google Scholar]
- Pomerantz JL, Denny EM, Baltimore D. CARD11 mediates factor-specific activation of NF-kappaB by the T cell receptor complex. EMBO J. 2002;21:5184–5194. doi: 10.1093/emboj/cdf505. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hara H, Wada T, Bakal C, et al. The MAGUK family protein CARD11 is essential for lymphocyte activation. Immunity. 2003;18:763–775. doi: 10.1016/s1074-7613(03)00148-1. [DOI] [PubMed] [Google Scholar]
- Egawa T, Albrecht B, Favier B, et al. Requirement for CARMA1 in antigen receptor-induced NF-kappa B activation and lymphocyte proliferation. Curr Biol. 2003;13:1252–1258. doi: 10.1016/s0960-9822(03)00491-3. [DOI] [PubMed] [Google Scholar]
- Newton K, Dixit V. Mice lacking the CARD of CARMA1 exhibit defective B lymphocyte development and impaired proliferation of their B and T lymphocytes. Curr Biol. 2003;13:1247–1251. doi: 10.1016/s0960-9822(03)00458-5. [DOI] [PubMed] [Google Scholar]
- Ruland J, Duncan GS, Elia A, et al. Bcl10 is a positive regulator of antigen receptor-induced activation of NF-kappaB and neural tube closure. Cell. 2001;104:33–42. doi: 10.1016/s0092-8674(01)00189-1. [DOI] [PubMed] [Google Scholar]
- Willis TG, Jadayel DM, Du MQ, et al. Bcl10 is involved in t(1;14)(p22;q32) of MALT B cell lymphoma and mutated in multiple tumor types. Cell. 1999;96:35–45. doi: 10.1016/s0092-8674(00)80957-5. [DOI] [PubMed] [Google Scholar]
- Zhang Q, Siebert R, Yan M, et al. Inactivating mutations and overexpression of BCL10, a caspase recruitment domain-containing gene, in MALT lymphoma with t(1;14)(p22;q32) Nat Genet. 1999;22:63–68. doi: 10.1038/8767. [DOI] [PubMed] [Google Scholar]
- Yan M, Lee J, Schilbach S, Goddard A, Dixit V. mE10, a novel caspase recruitment domain-containing proapoptotic molecule. J Biol Chem. 1999;274:10287–10292. doi: 10.1074/jbc.274.15.10287. [DOI] [PubMed] [Google Scholar]
- Thome M, Martinon F, Hofmann K, et al. Equine herpesvirus-2 E10 gene product, but not its cellular homologue, activates NF-kappaB transcription factor and c-Jun N-terminal kinase. J Biol Chem. 1999;274:9962–9968. doi: 10.1074/jbc.274.15.9962. [DOI] [PubMed] [Google Scholar]
- Koseki T, Inohara N, Chen S, et al. CIPER, a novel NF kappaB-activating protein containing a caspase recruitment domain with homology to Herpesvirus-2 protein E10. J Biol Chem. 1999;274:9955–9961. doi: 10.1074/jbc.274.15.9955. [DOI] [PubMed] [Google Scholar]
- Xue L, Morris SW, Orihuela C, et al. Defective development and function of Bcl10-deficient follicular, marginal zone and B1 B cells. Nat Immunol. 2003;4:857–865. doi: 10.1038/ni963. [DOI] [PubMed] [Google Scholar]
- Schaefer BC, Kappler JW, Kupfer A, Marrack P. Complex and dynamic redistribution of NF-kappaB signaling intermediates in response to T cell receptor stimulation. Proc Natl Acad Sci USA. 2004;101:1004–1009. doi: 10.1073/pnas.0307858100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rossman JS, Stoicheva NG, Langel FD, et al. POLKADOTS are foci of functional interactions in T-Cell receptor-mediated signaling to NF-kappaB. Mol Biol Cell. 2006;17:2166–2176. doi: 10.1091/mbc.E05-10-0985. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Blonska M, Pappu BP, Matsumoto R, et al. The CARMA1-Bcl10 signaling complex selectively regulates JNK2 kinase in the T cell receptor-signaling pathway. Immunity. 2007;26:55–66. doi: 10.1016/j.immuni.2006.11.008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rueda D, Gaide O, Ho L, et al. Bcl10 controls TCR- and FcgammaR-induced actin polymerization. J Immunol. 2007;178:4373–4384. doi: 10.4049/jimmunol.178.7.4373. [DOI] [PubMed] [Google Scholar]
- Thome M, Charton JE, Pelzer C, Hailfinger S. Antigen receptor signaling to NF-kappaB via CARMA1, BCL10, and MALT1. Cold Spring Harb Perspect Biol. 2010;2:a003004. doi: 10.1101/cshperspect.a003004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Akagi T, Motegi M, Tamura A, et al. A novel gene, MALT1 at 18q21, is involved in t(11;18) (q21;q21) found in low-grade B-cell lymphoma of mucosa-associated lymphoid tissue. Oncogene. 1999;18:5785–5794. doi: 10.1038/sj.onc.1203018. [DOI] [PubMed] [Google Scholar]
- Uren AG, O'Rourke K, Aravind LA, et al. Identification of paracaspases and metacaspases: two ancient families of caspase-like proteins, one of which plays a key role in MALT lymphoma. Mol Cell. 2000;6:961–967. doi: 10.1016/s1097-2765(00)00094-0. [DOI] [PubMed] [Google Scholar]
- Ye H, Gong L, Liu H, et al. MALT lymphoma with t(14;18)(q32;q21)/IGH-MALT1 is characterized by strong cytoplasmic MALT1 and BCL10 expression. J Pathol. 2005;205:293–301. doi: 10.1002/path.1715. [DOI] [PubMed] [Google Scholar]
- Du MQ. MALT lymphoma: recent advances in aetiology and molecular genetics. J Clin Exp Hematop. 2007;47:31–42. doi: 10.3960/jslrt.47.31. [DOI] [PubMed] [Google Scholar]
- Lucas PC, Yonezumi M, Inohara N, et al. Bcl10 and MALT1, independent targets of chromosomal translocation in malt lymphoma, cooperate in a novel NF-kappa B signaling pathway. J Biol Chem. 2001;276:19012–19019. doi: 10.1074/jbc.M009984200. [DOI] [PubMed] [Google Scholar]
- Baens M, Fevery S, Sagaert X, et al. Selective expansion of marginal zone B cells in Emicro-API2-MALT1 mice is linked to enhanced IkappaB kinase gamma polyubiquitination. Cancer Res. 2006;66:5270–5277. doi: 10.1158/0008-5472.CAN-05-4590. [DOI] [PubMed] [Google Scholar]
- Ruefli-Brasse AA, French DM, Dixit VM. Regulation of NF-kappaB-dependent lymphocyte activation and development by paracaspase. Science. 2003;302:1581–1584. doi: 10.1126/science.1090769. [DOI] [PubMed] [Google Scholar]
- Ruland J, Duncan GS, Wakeham A, Mak TW. Differential requirement for Malt1 in T and B cell antigen receptor signaling. Immunity. 2003;19:749–758. doi: 10.1016/s1074-7613(03)00293-0. [DOI] [PubMed] [Google Scholar]
- Ferch U, zum Buschenfelde CM, Gewies A, et al. MALT1 directs B cell receptor-induced canonical nuclear factor-kappaB signaling selectively to the c-Rel subunit. Nat Immunol. 2007;8:984–991. doi: 10.1038/ni1493. [DOI] [PubMed] [Google Scholar]
- Zhou H, Wertz I, O'Rourke K, et al. Bcl10 activates the NF-kappaB pathway through ubiquitination of NEMO. Nature. 2004;427:167–171. doi: 10.1038/nature02273. [DOI] [PubMed] [Google Scholar]
- Kawadler H, Gantz MA, Riley JL, Yang X. The paracaspase MALT1 controls caspase-8 activation during lymphocyte proliferation. Mol Cell. 2008;31:415–421. doi: 10.1016/j.molcel.2008.06.008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rebeaud F, Hailfinger S, Posevitz-Fejfar A, et al. The proteolytic activity of the paracaspase MALT1 is key in T cell activation. Nat Immunol. 2008;9:272–281. doi: 10.1038/ni1568. [DOI] [PubMed] [Google Scholar]
- Su H, Bidere N, Zheng L, et al. Requirement for caspase-8 in NF-kappaB activation by antigen receptor. Science. 2005;307:1465–1468. doi: 10.1126/science.1104765. [DOI] [PubMed] [Google Scholar]
- Bidere N, Snow AL, Sakai K, Zheng L, Lenardo MJ. Caspase-8 regulation by direct interaction with TRAF6 in T cell receptor-induced NF-kappaB activation. Curr Biol. 2006;16:1666–1671. doi: 10.1016/j.cub.2006.06.062. [DOI] [PubMed] [Google Scholar]
- Misra RS, Russell JQ, Koenig A, et al. Caspase-8 and c-FLIPL associate in lipid rafts with NF-kappaB adaptors during T cell activation. J Biol Chem. 2007;282:19365–19374. doi: 10.1074/jbc.M610610200. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Malarkannan S, Regunathan J, Chu H, et al. Bcl10 plays a divergent role in NK cell-mediated cytotoxicity and cytokine generation. J Immunol. 2007;179:3752–3762. doi: 10.4049/jimmunol.179.6.3752. [DOI] [PubMed] [Google Scholar]
- Gross O, Grupp C, Steinberg C, et al. Multiple ITAM-coupled NK-cell receptors engage the Bcl10/Malt1 complex via Carma1 for NF-kappaB and MAPK activation to selectively control cytokine production. Blood. 2008;112:2421–2428. doi: 10.1182/blood-2007-11-123513. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Klemm S, Gutermuth J, Hultner L, et al. The Bcl10-Malt1 complex segregates Fc epsilon RI-mediated nuclear factor kappa B activation and cytokine production from mast cell degranulation. J Exp Med. 2006;203:337–347. doi: 10.1084/jem.20051982. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chen Y, Pappu BP, Zeng H, et al. B cell lymphoma 10 is essential for FcepsilonR-mediated degranulation and IL-6 production in mast cells. J Immunol. 2007;178:49–57. doi: 10.4049/jimmunol.178.1.49. [DOI] [PubMed] [Google Scholar]
- Hara H, Bakal C, Wada T, et al. The molecular adapter Carma1 controls entry of IkappaB kinase into the central immune synapse. J Exp Med. 2004;200:1167–1177. doi: 10.1084/jem.20032246. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang D, Matsumoto R, You Y, et al. CD3/CD28 costimulation-induced NF-kappaB activation is mediated by recruitment of protein kinase C-theta, Bcl10, and IkappaB kinase beta to the immunological synapse through CARMA1. Mol Cell Biol. 2004;24:164–171. doi: 10.1128/MCB.24.1.164-171.2003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dimitratos SD, Woods DF, Stathakis DG, Bryant PJ. Signaling pathways are focused at specialized regions of the plasma membrane by scaffolding proteins of the MAGUK family. Bioessays. 1999;21:912–921. doi: 10.1002/(SICI)1521-1878(199911)21:11<912::AID-BIES3>3.0.CO;2-Z. [DOI] [PubMed] [Google Scholar]
- Medeiros RB, Burbach BJ, Mueller KL, et al. Regulation of NF-kappaB activation in T cells via association of the adapter proteins ADAP and CARMA1. Science. 2007;316:754–758. doi: 10.1126/science.1137895. [DOI] [PubMed] [Google Scholar]
- Sommer K, Guo B, Pomerantz JL, et al. Phosphorylation of the CARMA1 linker controls NF-kappaB activation. Immunity. 2005;23:561–574. doi: 10.1016/j.immuni.2005.09.014. [DOI] [PubMed] [Google Scholar]
- Shinohara H, Maeda S, Watarai H, Kurosaki T. IkappaB kinase beta-induced phosphorylation of CARMA1 contributes to CARMA1 Bcl10 MALT1 complex formation in B cells. J Exp Med. 2007;204:3285–3293. doi: 10.1084/jem.20070379. [DOI] [PMC free article] [PubMed] [Google Scholar]
- McCully RR, Pomerantz JL. The protein kinase C-responsive inhibitory domain of CARD11 functions in NF-kappaB activation to regulate the association of multiple signaling cofactors that differentially depend on Bcl10 and MALT1 for association. Mol Cell Biol. 2008;28:5668–5686. doi: 10.1128/MCB.00418-08. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lee KY, D'Acquisto F, Hayden MS, Shim JH, Ghosh S. PDK1 nucleates T cell receptor-induced signaling complex for NF-kappaB activation. Science. 2005;308:114–118. doi: 10.1126/science.1107107. [DOI] [PubMed] [Google Scholar]
- Shinohara H, Yasuda T, Aiba Y, et al. PKC beta regulates BCR-mediated IKK activation by facilitating the interaction between TAK1 and CARMA1. J Exp Med. 2005;202:1423–1431. doi: 10.1084/jem.20051591. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Narayan P, Holt B, Tosti R, Kane LP. CARMA1 is required for Akt-mediated NF-kappaB activation in T cells. Mol Cell Biol. 2006;26:2327–2336. doi: 10.1128/MCB.26.6.2327-2336.2006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bidere N, Ngo VN, Lee J, et al. Casein kinase 1alpha governs antigen-receptor-induced NF-kappaB activation and human lymphoma cell survival. Nature. 2009;458:92–96. doi: 10.1038/nature07613. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Brenner D, Brechmann M, Rohling S, et al. Phosphorylation of CARMA1 by HPK1 is critical for NF-kappaB activation in T cells. Proc Natl Acad Sci USA. 2009;106:14508–14513. doi: 10.1073/pnas.0900457106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ishiguro K, Green T, Rapley J, et al. Ca2+/calmodulin-dependent protein kinase II is a modulator of CARMA1-mediated NF-kappaB activation. Mol Cell Biol. 2006;26:5497–5508. doi: 10.1128/MCB.02469-05. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Qiao G, Li Z, Molinero L, et al. T-cell receptor-induced NF-kappaB activation is negatively regulated by E3 ubiquitin ligase Cbl-b. Mol Cell Biol. 2008;28:2470–2480. doi: 10.1128/MCB.01505-07. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Che T, You Y, Wang D, et al. MALT1/paracaspase is a signaling component downstream of CARMA1 and mediates T cell receptor-induced NF-kappaB activation. J Biol Chem. 2004;279:15870–15876. doi: 10.1074/jbc.M310599200. [DOI] [PubMed] [Google Scholar]
- Sun L, Deng L, Ea CK, Xia ZP, Chen ZJ. The TRAF6 ubiquitin ligase and TAK1 kinase mediate IKK activation by BCL10 and MALT1 in T lymphocytes. Mol Cell. 2004;14:289–301. doi: 10.1016/s1097-2765(04)00236-9. [DOI] [PubMed] [Google Scholar]
- Zandi E, Rothwarf DM, Delhase M, Hayakawa M, Karin M. The IkappaB kinase complex (IKK) contains two kinase subunits, IKKalpha and IKKbeta, necessary for IkappaB phosphorylation and NF-kappaB activation. Cell. 1997;91:243–252. doi: 10.1016/s0092-8674(00)80406-7. [DOI] [PubMed] [Google Scholar]
- Rothwarf DM, Zandi E, Natoli G, Karin M. IKK-gamma is an essential regulatory subunit of the IkappaB kinase complex. Nature. 1998;395:297–300. doi: 10.1038/26261. [DOI] [PubMed] [Google Scholar]
- Li ZW, Chu W, Hu Y, et al. The IKKbeta subunit of IkappaB kinase (IKK) is essential for nuclear factor kappaB activation and prevention of apoptosis. J Exp Med. 1999;189:1839–1845. doi: 10.1084/jem.189.11.1839. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shambharkar PB, Blonska M, Pappu BP, et al. Phosphorylation and ubiquitination of the IkappaB kinase complex by two distinct signaling pathways. EMBO J. 2007;26:1794–1805. doi: 10.1038/sj.emboj.7601622. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tusche MW, Ward LA, Vu F, et al. Differential requirement of MALT1 for BAFF-induced outcomes in B cell subsets. J Exp Med. 2009;206:2671–2683. doi: 10.1084/jem.20091802. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Delhase M, Hayakawa M, Chen Y, Karin M. Positive and negative regulation of IkappaB kinase activity through IKKbeta subunit phosphorylation. Science. 1999;284:309–313. doi: 10.1126/science.284.5412.309. [DOI] [PubMed] [Google Scholar]
- Ling L, Cao Z, Goeddel DV. NF-kappaB-inducing kinase activates IKK-alpha by phosphorylation of Ser-176. Proc Natl Acad Sci USA. 1998;95:3792–3797. doi: 10.1073/pnas.95.7.3792. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sato S, Sanjo H, Tsujimura T, et al. TAK1 is indispensable for development of T cells and prevention of colitis by the generation of regulatory T cells. Int Immunol. 2006;18:1405–1411. doi: 10.1093/intimm/dxl082. [DOI] [PubMed] [Google Scholar]
- Shim JH, Xiao C, Paschal AE, et al. TAK1, but not TAB1 or TAB2, plays an essential role in multiple signaling pathways in vivo. Genes Dev. 2005;19:2668–2681. doi: 10.1101/gad.1360605. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wan YY, Chi H, Xie M, Schneider MD, Flavell RA. The kinase TAK1 integrates antigen and cytokine receptor signaling for T cell development, survival and function. Nat Immunol. 2006;7:851–858. doi: 10.1038/ni1355. [DOI] [PubMed] [Google Scholar]
- Liu HH, Xie M, Schneider MD, Chen ZJ. Essential role of TAK1 in thymocyte development and activation. Proc Natl Acad Sci USA. 2006;103:11677–11682. doi: 10.1073/pnas.0603089103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sun W, Li H, Yu Y, et al. MEKK3 is required for lysophosphatidic acid-induced NF-kappaB activation. Cell Signal. 2009;21:1488–1494. doi: 10.1016/j.cellsig.2009.05.007. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shinohara H, Yamasaki S, Maeda S, Saito T, Kurosaki T. Regulation of NF-kappaB-dependent T cell activation and development by MEKK3. Int Immunol. 2009;21:393–401. doi: 10.1093/intimm/dxp007. [DOI] [PubMed] [Google Scholar]
- Yang J, Lin Y, Guo Z, et al. The essential role of MEKK3 in TNF-induced NF-kappaB activation. Nat Immunol. 2001;2:620–624. doi: 10.1038/89769. [DOI] [PubMed] [Google Scholar]
- Huang Q, Yang J, Lin Y, et al. Differential regulation of interleukin 1 receptor and Toll-like receptor signaling by MEKK3. Nat Immunol. 2004;5:98–103. doi: 10.1038/ni1014. [DOI] [PubMed] [Google Scholar]
- Abbasi S, Su B, Kellems RE, Yang J, Xia Y. The essential role of MEKK3 signaling in angiotensin II-induced calcineurin/nuclear factor of activated T-cells activation. J Biol Chem. 2005;280:36737–36746. doi: 10.1074/jbc.M506493200. [DOI] [PubMed] [Google Scholar]
- Stilo R, Liguoro D, Di Jeso B, et al. Physical and functional interaction of CARMA1 and CARMA3 with Ikappa kinase gamma-NFkappaB essential modulator. J Biol Chem. 2004;279:34323–34331. doi: 10.1074/jbc.M402244200. [DOI] [PubMed] [Google Scholar]
- Ea CK, Deng L, Xia ZP, Pineda G, Chen ZJ. Activation of IKK by TNFalpha requires site-specific ubiquitination of RIP1 and polyubiquitin binding by NEMO. Mol Cell. 2006;22:245–257. doi: 10.1016/j.molcel.2006.03.026. [DOI] [PubMed] [Google Scholar]
- Wu CJ, Conze DB, Li T, Srinivasula SM, Ashwell JD. Sensing of Lys 63-linked polyubiquitination by NEMO is a key event in NF-kappaB activation. Nat Cell Biol. 2006;8:398–406. doi: 10.1038/ncb1384. [DOI] [PubMed] [Google Scholar]
- Wu CJ, Ashwell JD. NEMO recognition of ubiquitinated Bcl10 is required for T cell receptor-mediated NF-kappaB activation. Proc Natl Acad Sci USA. 2008;105:3023–3028. doi: 10.1073/pnas.0712313105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tang ED, Wang CY, Xiong Y, Guan KL. A role for NF-kappaB essential modifier/IkappaB kinase-gamma (NEMO/IKKgamma) ubiquitination in the activation of the IkappaB kinase complex by tumor necrosis factor-alpha. J Biol Chem. 2003;278:37297–37305. doi: 10.1074/jbc.M303389200. [DOI] [PubMed] [Google Scholar]
- Israel A. The IKK complex, a central regulator of NF-kappaB activation. Cold Spring Harb Perspect Biol. 2010;2:a000158. doi: 10.1101/cshperspect.a000158. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li H, Kobayashi M, Blonska M, You Y, Lin X. Ubiquitination of RIP is required for tumor necrosis factor alpha-induced NF-kappaB activation. J Biol Chem. 2006;281:13636–13643. doi: 10.1074/jbc.M600620200. [DOI] [PubMed] [Google Scholar]
- Conze DB, Wu CJ, Thomas JA, Landstrom A, Ashwell JD. Lys63-linked polyubiquitination of IRAK-1 is required for interleukin-1 receptor- and toll-like receptor-mediated NF-kappaB activation. Mol Cell Biol. 2008;28:3538–3547. doi: 10.1128/MCB.02098-07. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Oeckinghaus A, Wegener E, Welteke V, et al. Malt1 ubiquitination triggers NF-kappaB signaling upon T-cell activation. EMBO J. 2007;26:4634–4645. doi: 10.1038/sj.emboj.7601897. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ni CY, Wu ZH, Florence WC, et al. Cutting edge: K63-linked polyubiquitination of NEMO modulates TLR signaling and inflammation in vivo. J Immunol. 2008;180:7107–7111. doi: 10.4049/jimmunol.180.11.7107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- King CG, Kobayashi T, Cejas PJ, et al. TRAF6 is a T cell-intrinsic negative regulator required for the maintenance of immune homeostasis. Nat Med. 2006;12:1088–1092. doi: 10.1038/nm1449. [DOI] [PubMed] [Google Scholar]
- Moreno-Garcia ME, Sommer K, Haftmann C, et al. Serine 649 phosphorylation within the protein kinase C-regulated domain down-regulates CARMA1 activity in lymphocytes. J Immunol. 2009;183:7362–7370. doi: 10.4049/jimmunol.0902438. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kojo S, Elly C, Harada Y, et al. Mechanisms of NKT cell anergy induction involve Cbl-b-promoted monoubiquitination of CARMA1. Proc Natl Acad Sci USA. 2009;106:17847–17851. doi: 10.1073/pnas.0904078106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Moreno-Garcia ME, Sommer K, Shinohara H, et al. MAGUK-controlled ubiquitination of CARMA1 modulates lymphocyte NF-kappaB activity. Mol Cell Biol. 2010;30:922–934. doi: 10.1128/MCB.01129-09. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Welteke V, Eitelhuber A, Duwel M, et al. COP9 signalosome controls the Carma1-Bcl10-Malt1 complex upon T-cell stimulation. EMBO Rep. 2009;10:642–648. doi: 10.1038/embor.2009.64. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ishiguro K, Ando T, Goto H, Xavier R. Bcl10 is phosphorylated on Ser138 by Ca2+/calmodulin-dependent protein kinase II. Mol Immunol. 2007;44:2095–2100. doi: 10.1016/j.molimm.2006.09.012. [DOI] [PubMed] [Google Scholar]
- Zeng H, Di L, Fu G, et al. Phosphorylation of Bcl10 negatively regulates T-cell receptor-mediated NF-kappaB activation. Mol Cell Biol. 2007;27:5235–5245. doi: 10.1128/MCB.01645-06. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lobry C, Lopez T, Israel A, Weil R. Negative feedback loop in T cell activation through IkappaB kinase-induced phosphorylation and degradation of Bcl10. Proc Natl Acad Sci USA. 2007;104:908–913. doi: 10.1073/pnas.0606982104. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ngo VN, Davis RE, Lamy L, et al. A loss-of-function RNA interference screen for molecular targets in cancer. Nature. 2006;441:106–110. doi: 10.1038/nature04687. [DOI] [PubMed] [Google Scholar]
- Li Z, Wang H, Xue L, et al. Emu-BCL10 mice exhibit constitutive activation of both canonical and noncanonical NF-kappaB pathways generating marginal zone (MZ) B-cell expansion as a precursor to splenic MZ lymphoma. Blood. 2009;114:4158–4168. doi: 10.1182/blood-2008-12-192583. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Oshiro A, Tagawa H, Ohshima K, et al. Identification of subtype-specific genomic alterations in aggressive adult T-cell leukemia/lymphoma. Blood. 2006;107:4500–4507. doi: 10.1182/blood-2005-09-3801. [DOI] [PubMed] [Google Scholar]
- Nakamura S, Matsumoto T, Yada S, et al. Overexpression of caspase recruitment domain (CARD) membrane-associated guanylate kinase 1 (CARMA1) and CARD9 in primary gastric B-cell lymphoma. Cancer. 2005;104:1885–1893. doi: 10.1002/cncr.21421. [DOI] [PubMed] [Google Scholar]
- Tanner MJ, Hanel W, Gaffen SL, Lin X. CARMA1 coiled-coil domain is involved in the oligomerization and subcellular localization of CARMA1 and is required for T cell receptor-induced NF-kappaB activation. J Biol Chem. 2007;282:17141–17147. doi: 10.1074/jbc.M700169200. [DOI] [PubMed] [Google Scholar]
- Lenz G, Davis RE, Ngo VN, et al. Oncogenic CARD11 mutations in human diffuse large B cell lymphoma. Science. 2008;319:1676–1679. doi: 10.1126/science.1153629. [DOI] [PubMed] [Google Scholar]
- Compagno M, Lim WK, Grunn A, et al. Mutations of multiple genes cause deregulation of NF-kappaB in diffuse large B-cell lymphoma. Nature. 2009;459:717–721. doi: 10.1038/nature07968. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Montesinos-Rongen M, Schmitz R, Brunn A, et al. Mutations of CARD11 but not TNFAIP3 may activate the NF-kappaB pathway in primary CNS lymphoma. Acta Neuropathol. 2010;120:529–535. doi: 10.1007/s00401-010-0709-7. [DOI] [PubMed] [Google Scholar]
- Lamason RL, McCully RR, Lew SM, Pomerantz JL. Oncogenic CARD11 Mutations Induce Hyperactive Signaling by Disrupting Autoinhibition by the PKC-Responsive Inhibitory Domain. Biochemistry. 2010;49:8240–8250. doi: 10.1021/bi101052d. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hailfinger S, Lenz G, Ngo V, et al. Essential role of MALT1 protease activity in activated B cell-like diffuse large B-cell lymphoma. Proc Natl Acad Sci USA. 2009;106:19946–19951. doi: 10.1073/pnas.0907511106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ferch U, Kloo B, Gewies A, et al. Inhibition of MALT1 protease activity is selectively toxic for activated B cell-like diffuse large B cell lymphoma cells. J Exp Med. 2009;206:2313–2320. doi: 10.1084/jem.20091167. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Davis RE, Brown KD, Siebenlist U, Staudt LM. Constitutive nuclear factor kappaB activity is required for survival of activated B cell-like diffuse large B cell lymphoma cells. J Exp Med. 2001;194:1861–1874. doi: 10.1084/jem.194.12.1861. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lam LT, Davis RE, Pierce J, et al. Small molecule inhibitors of IkappaB kinase are selectively toxic for subgroups of diffuse large B-cell lymphoma defined by gene expression profiling. Clin Cancer Res. 2005;11:28–40. [PubMed] [Google Scholar]
- Grabiner BC, Blonska M, Lin PC, et al. CARMA3 deficiency abrogates G protein-coupled receptor-induced NF-{kappa}B activation. Genes Dev. 2007;21:984–996. doi: 10.1101/gad.1502507. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lefkowitz RJ. Historical review: a brief history and personal retrospective of seven-transmembrane receptors. Trends Pharmacol Sci. 2004;25:413–422. doi: 10.1016/j.tips.2004.06.006. [DOI] [PubMed] [Google Scholar]
- Mills GB, Moolenaar WH. The emerging role of lysophosphatidic acid in cancer. Nat Rev Cancer. 2003;3:582–591. doi: 10.1038/nrc1143. [DOI] [PubMed] [Google Scholar]
- Zhao Y, Tong J, He D, et al. Role of lysophosphatidic acid receptor LPA2 in the development of allergic airway inflammation in a murine model of asthma. Respir Res. 2009;10:114. doi: 10.1186/1465-9921-10-114. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ye RD. Regulation of nuclear factor kappaB activation by G-protein-coupled receptors. J Leukoc Biol. 2001;70:839–848. [PubMed] [Google Scholar]
- McAllister-Lucas LM, Ruland J, Siu K, et al. CARMA3/Bcl10/MALT1-dependent NF-kappaB activation mediates angiotensin II-responsive inflammatory signaling in nonimmune cells. Proc Natl Acad Sci USA. 2007;104:139–144. doi: 10.1073/pnas.0601947103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- McAllister-Lucas LM, Jin X, Gu S, et al. The CARMA3-Bcl10-MALT1 signalosome promotes angiotensin II-dependent vascular inflammation and atherogenesis. J Biol Chem. 2010;285:25880–25884. doi: 10.1074/jbc.C110.109421. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Medoff BD, Landry AL, Wittbold KA, et al. CARMA3 mediates lysophosphatidic acid-stimulated cytokine secretion by bronchial epithelial cells. Am J Respir Cell Mol Biol. 2009;40:286–294. doi: 10.1165/rcmb.2008-0129OC. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhao Y, Natarajan V. Lysophosphatidic acid signaling in airway epithelium: role in airway inflammation and remodeling. Cell Signal. 2009;21:367–377. doi: 10.1016/j.cellsig.2008.10.010. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Georas SN, Berdyshev E, Hubbard W, et al. Lysophosphatidic acid is detectable in human bronchoalveolar lavage fluids at baseline and increased after segmental allergen challenge. Clin Exp Allergy. 2007;37:311–322. doi: 10.1111/j.1365-2222.2006.02626.x. [DOI] [PubMed] [Google Scholar]
- Mahanivong C, Chen HM, Yee SW, et al. Protein kinase C alpha-CARMA3 signaling axis links Ras to NF-kappa B for lysophosphatidic acid-induced urokinase plasminogen activator expression in ovarian cancer cells. Oncogene. 2008;27:1273–1280. doi: 10.1038/sj.onc.1210746. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Klemm S, Zimmermann S, Peschel C, Mak TW, Ruland J. Bcl10 and Malt1 control lysophosphatidic acid-induced NF-kappaB activation and cytokine production. Proc Natl Acad Sci USA. 2007;104:134–138. doi: 10.1073/pnas.0608388103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang D, You Y, Lin PC, et al. Bcl10 plays a critical role in NF-kappaB activation induced by G protein-coupled receptors. Proc Natl Acad Sci USA. 2007;104:145–150. doi: 10.1073/pnas.0601894104. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cummings R, Zhao Y, Jacoby D, et al. Protein kinase Cdelta mediates lysophosphatidic acid-induced NF-kappaB activation and interleukin-8 secretion in human bronchial epithelial cells. J Biol Chem. 2004;279:41085–41094. doi: 10.1074/jbc.M404045200. [DOI] [PubMed] [Google Scholar]
- Lefkowitz RJ, Shenoy SK. Transduction of receptor signals by beta-arrestins. Science. 2005;308:512–517. doi: 10.1126/science.1109237. [DOI] [PubMed] [Google Scholar]
- Sun L, Carpenter G. Epidermal growth factor activation of NF-kappaB is mediated through IkappaBalpha degradation and intracellular free calcium. Oncogene. 1998;16:2095–2102. doi: 10.1038/sj.onc.1201731. [DOI] [PubMed] [Google Scholar]
- Biswas DK, Cruz AP, Gansberger E, Pardee AB. Epidermal growth factor-induced nuclear factor kappa B activation: A major pathway of cell-cycle progression in estrogen-receptor negative breast cancer cells. Proc Natl Acad Sci USA. 2000;97:8542–8547. doi: 10.1073/pnas.97.15.8542. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Biswas DK, Iglehart JD. Linkage between EGFR family receptors and nuclear factor kappaB (NF-kappaB) signaling in breast cancer. J Cell Physiol. 2006;209:645–652. doi: 10.1002/jcp.20785. [DOI] [PubMed] [Google Scholar]
- Kim HJ, Litzenburger BC, Cui X, et al. Constitutively active type I insulin-like growth factor receptor causes transformation and xenograft growth of immortalized mammary epithelial cells and is accompanied by an epithelial-to-mesenchymal transition mediated by NF-kappaB and snail. Mol Cell Biol. 2007;27:3165–3175. doi: 10.1128/MCB.01315-06. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pons S, Torres-Aleman I. Insulin-like growth factor-I stimulates dephosphorylation of ikappa B through the serine phosphatase calcineurin (protein phosphatase 2B) J Biol Chem. 2000;275:38620–38625. doi: 10.1074/jbc.M004531200. [DOI] [PubMed] [Google Scholar]
- Kaliman P, Canicio J, Testar X, Palacin M, Zorzano A. Insulin-like growth factor-II, phosphatidylinositol 3-kinase, nuclear factor-kappaB and inducible nitric-oxide synthase define a common myogenic signaling pathway. J Biol Chem. 1999;274:17437–17444. doi: 10.1074/jbc.274.25.17437. [DOI] [PubMed] [Google Scholar]
- Olashaw NE, Kowalik TF, Huang ES, Pledger WJ. Induction of NF-kappa B-like activity by platelet-derived growth factor in mouse fibroblasts. Mol Biol Cell. 1992;3:1131–1139. doi: 10.1091/mbc.3.10.1131. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Romashkova JA, Makarov SS. NF-kappaB is a target of AKT in anti-apoptotic PDGF signalling. Nature. 1999;401:86–90. doi: 10.1038/43474. [DOI] [PubMed] [Google Scholar]
- Byrd VM, Ballard DW, Miller GG, Thomas JW. Fibroblast growth factor-1 (FGF-1) enhances IL-2 production and nuclear translocation of NF-kappaB in FGF receptor-bearing Jurkat T cells. J Immunol. 1999;162:5853–5859. [PubMed] [Google Scholar]
- Bushdid PB, Chen CL, Brantley DM, et al. NF-kappaB mediates FGF signal regulation of msx-1 expression. Dev Biol. 2001;237:107–115. doi: 10.1006/dbio.2001.0356. [DOI] [PubMed] [Google Scholar]
- Bertin J, Guo Y, Wang L, et al. CARD9 is a novel caspase recruitment domain-containing protein that interacts with BCL10/CLAP and activates NF-kappa B. J Biol Chem. 2000;275:41082–41086. doi: 10.1074/jbc.C000726200. [DOI] [PubMed] [Google Scholar]
- Gross O, Gewies A, Finger K, et al. Card9 controls a non-TLR signalling pathway for innate anti-fungal immunity. Nature. 2006;442:651–656. doi: 10.1038/nature04926. [DOI] [PubMed] [Google Scholar]
- Hsu YM, Zhang Y, You Y, et al. The adaptor protein CARD9 is required for innate immune responses to intracellular pathogens. Nat Immunol. 2007;8:198–205. doi: 10.1038/ni1426. [DOI] [PubMed] [Google Scholar]
- Bi L, Gojestani S, Wu W, et al. CARD9 mediates dectin-2-induced IkappaBalpha kinase ubiquitination leading to activation of NF-kappaB in response to stimulation by the hyphal form of Candida albicans. J Biol Chem. 2010;285:25969–25977. doi: 10.1074/jbc.M110.131300. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Robinson MJ, Osorio F, Rosas M, et al. Dectin-2 is a Syk-coupled pattern recognition receptor crucial for Th17 responses to fungal infection. J Exp Med. 2009;206:2037–2051. doi: 10.1084/jem.20082818. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yamasaki S, Ishikawa E, Sakuma M, et al. Mincle is an ITAM-coupled activating receptor that senses damaged cells. Nat Immunol. 2008;9:1179–1188. doi: 10.1038/ni.1651. [DOI] [PubMed] [Google Scholar]
- Willment JA, Brown GD. C-type lectin receptors in antifungal immunity. Trends Microbiol. 2008;16:27–32. doi: 10.1016/j.tim.2007.10.012. [DOI] [PubMed] [Google Scholar]
- Graham LM, Brown GD. The Dectin-2 family of C-type lectins in immunity and homeostasis. Cytokine. 2009;48:148–155. doi: 10.1016/j.cyto.2009.07.010. [DOI] [PMC free article] [PubMed] [Google Scholar]
- LeibundGut-Landmann S, Gross O, Robinson MJ, et al. Syk- and CARD9-dependent coupling of innate immunity to the induction of T helper cells that produce interleukin 17. Nat Immunol. 2007;8:630–638. doi: 10.1038/ni1460. [DOI] [PubMed] [Google Scholar]
- Goodridge HS, Shimada T, Wolf AJ, et al. Differential use of CARD9 by dectin-1 in macrophages and dendritic cells. J Immunol. 2009;182:1146–1154. doi: 10.4049/jimmunol.182.2.1146. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Saijo S, Ikeda S, Yamabe K, et al. Dectin-2 recognition of alpha-mannans and induction of Th17 cell differentiation is essential for host defense against Candida albicans. Immunity. 2010;32:681–691. doi: 10.1016/j.immuni.2010.05.001. [DOI] [PubMed] [Google Scholar]
- Wu W, Hsu YM, Bi L, Songyang Z, Lin X. CARD9 facilitates microbe-elicited production of reactive oxygen species by regulating the LyGDI-Rac1 complex. Nat Immunol. 2009;10:1208–1214. doi: 10.1038/ni.1788. [DOI] [PubMed] [Google Scholar]
- Werninghaus K, Babiak A, Gross O, et al. Adjuvanticity of a synthetic cord factor analogue for subunit Mycobacterium tuberculosis vaccination requires FcRgamma-Syk-Card9-dependent innate immune activation. J Exp Med. 2009;206:89–97. doi: 10.1084/jem.20081445. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Poeck H, Bscheider M, Gross O, et al. Recognition of RNA virus by RIG-I results in activation of CARD9 and inflammasome signaling for interleukin 1 beta production. Nat Immunol. 2010;11:63–69. doi: 10.1038/ni.1824. [DOI] [PubMed] [Google Scholar]
- Yang H, Minamishima YA, Yan Q, et al. pVHL acts as an adaptor to promote the inhibitory phosphorylation of the NF-kappaB agonist Card9 by CK2. Mol Cell. 2007;28:15–27. doi: 10.1016/j.molcel.2007.09.010. [DOI] [PMC free article] [PubMed] [Google Scholar]