Abstract
The carbapenemase gene blaKPC, which is rapidly spreading worldwide, is located on a Tn3-based transposon, Tn4401. In a transposition-conjugation assay, Tn4401 was able to mobilize blaKPC-2 gene at a frequency of 4.4 × 10−6/recipient cell. A 5-bp target site duplication was evidenced upon each insertion without target site specificity. This study demonstrated that Tn4401 is an active transposon capable of mobilizing blaKPC genes at high frequency.
TEXT
By far, the most frequent class A carbapenemases are the KPC enzymes (Klebsiella pneumoniae carbapenemases), with the KPC-2 variant being most common (15). These carbapenemases were identified first in 2001 in North Carolina (26), and until 2005, the geographical distribution of these enzymes in the Enterobacteriaceae, including Klebsiella pneumoniae, was limited to the eastern part of the United States (3, 26). KPC producers are now reported to be found worldwide, mainly in the Enterobacteriaceae but also in Pseudomonas aeruginosa (24) and Acinetobacter sp. (20). Possible explanations for this rapid dissemination are that blaKPC genes are present in an epidemic K. pneumoniae strain ST258 (4) and also that they are present on a wide variety of plasmids that vary in size, nature, and structure (4, 6, 11). In most cases, these plasmids are self-transferable at least to Escherichia coli. Studies of the genetic structure surrounding blaKPC-2 genes have identified a Tn3-based transposon, Tn4401 (13). Tn4401 is 10 kb, is delimited by two 39-bp imperfect inverted repeat sequences, and harbors transposase and resolvase genes and two insertion sequences, ISKpn6 and ISKpn7, in addition to blaKPC-2 (Fig. 1). Three isoforms of Tn4401, differing by a 100- to 200-bp sequence upstream of blaKPC-2, are currently known (isoforms a, b, and c). A few descriptions of different genetic environments of blaKPC genes have been published, reporting the presence of other insertion sequences upstream of the blaKPC gene (21, 25) but with downstream sequences similar to those of Tn4401, suggesting that these ISs have been inserted in Tn4401. Here, we investigated the functional role of Tn4401 in mobilization and diffusion of the blaKPC-2 gene.
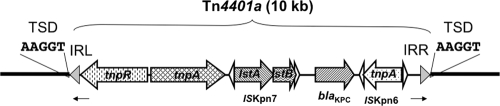
Fig. 1.
Schematic representation of Tn4401a inserted into pOX38-Gen (insertion I1 in Fig. 3). Genes and their corresponding transcription orientations are indicated by horizontal arrows. Tn4401a (isoform a [100-bp deletion as described in reference 13]) is delimited by two inverted repeat sequences, IRR and IRL (grey triangles). Small open triangles represent the inverted repeats of ISKpn6 and ISKpn7. Small arrows below the sequence represent primers used for direct sequencing of the insertion sites of Tn4401. The thick line corresponds to the pOX38-GEN DNA sequence.
Transposition experiments were performed with the natural nonconjugative plasmid pBC633 (13), which was electroporated into the recA Escherichia coli strain RZ211 harboring the plasmid pOX38-Gen for transposition experiments (2, 9). A 24-h broth was used as the donor for subsequent mating-out assays with the azide-resistant E. coli strain J53 as the recipient, as previously described (2, 9, 13). pOX38 transconjugants harboring Tn4401 insertions (pOX38-Gen::Tn4401) were selected on ticarcillin (Tic) (100 μg/ml) plus azide (Az) (100 μg/ml) as Ticr Genr Azr colonies. The plasmid content of E. coli J53(pOX38- Gen::Tn4401) was investigated by the Kieser extraction method (2, 10). The presence of the blaKPC gene in transconjugants was confirmed by PCR, and the location of the blaKPC gene was determined by Southern hybridization with a specific and internal probe (13). Transposition experiments with the natural plasmid pBC633 showed that, along with pOX38-Gen, pBC633 was also transferred to E. coli J53 Azr in mating-out assays, suggesting comobilization of pBC633 by pOX38-Gen (Fig. 2A). The presence of the two plasmids in the same transconjugants made further analysis of transposition events difficult. Nevertheless, true transposition events were also evidenced; a unique plasmid (pOX38-Gen::Tn4401) from two transconjugants could be extracted, and direct sequencing revealed insertion of Tn4401 in pOX38-Gen (Fig. 2A and 3B).
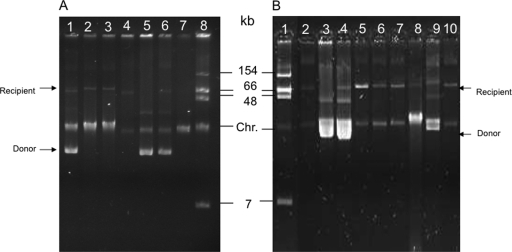
Fig. 2.
Plasmid extraction by the Kieser method. (A) transposition experiments performed with the natural plasmid pBC633. Lanes 1 to 3, transconjugants in E. coli J53 Azr after a transposition assay. E. coli J53 transconjugant 1 (lane 1) possesses both pOX38Gen and pBC633, due to cointegration events, whereas E. coli J53 transconjugants 2 and 3 (lanes 2 and 3) harbor only pOX38Gen::Tn4401, due to true transposition events. Lane 4, E. coli RZ211 harboring pOX38Gen; lane 5, K. pneumoniae KPN633 harboring the natural plasmid pBC633 (11); lane 6, E. coli RZ211 harboring both pOX38Gen and pBC633; lane 7, E. coli J53 Azr (only the chromosomal band [Chr] is visible); lane 8, E. coli 50192 harboring four plasmids (7, 48, 66, and 154 kb). (B) Transposition experiments performed with the recombinant plasmid pTn4401. Lane 1, E. coli 50192 harboring four plasmids (7, 48, 66, and 154 kb); lane 2, E. coli RZ211 harboring pOX38Gen; lane 3, E. coli DH10B harboring pTn4401; lane 4, E. coli J53 transconjugant possessing both plasmids pOX38Gen and pTn4401 due to a cointegration event; lanes 5 to 8, E. coli J53 transconjugants harboring pOX38Gen::Tn4401, selected as Cms colonies; lane 9, E. coli RZ211 harboring both pOX38Gen and pTn4401 (Cmr); lane 10, subsequent transconjugant from strain 9 to E. coli DH10B Strr.
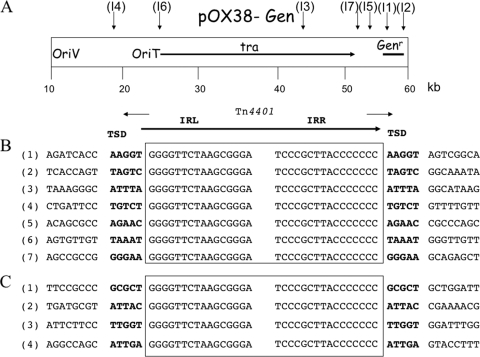
Fig. 3.
Target sites of Tn4401 insertions. (A) Map positions of Tn4401 insertions in plasmid pOX38-Gen. Insertions of the tagged transposon (I1 to I7) are indicated by vertical arrows. The origin of replication (oriV), the origin of transfer (oriT), the tra genes required for plasmid transfer, and the gentamicin resistance gene (Genr) are indicated. (B and C) Nucleotide sequence alignments of Tn4401 in pOX38-Gen. (B) Seven transconjugants were analyzed from a transposition assay with pBC633 or pTn4401. (C) Tn4401 insertions in natural plasmids. 1, pCOL from P. aeruginosa 2404 (13); 2, pBC2303 from K. pneumoniae KPN2303 (13); 3, pBC633 from K. pneumoniae KPN633 (13); 4, pYC from K. pneumoniae YC (13). Nucleotide sequences of the end regions of Tn4401 are boxed. Target site sequences duplicated after transposition are indicated by boldface letters. The small arrows above the schematic representation of Tn4401 represent primers used for direct sequencing of pOX38Gen::Tn4401 transposants.
In order to address whether the comobilization of pBC633, along with pOX38-Gen, was due to comobilization or cointegration events followed by subsequent resolution in the recipient strain, plasmid (pACYC184), which is known for not being comobilizable and which carries a chloramphenicol (Cm) resistance gene, was used to obtain a recombinant plasmid, pTn4401. The latter was constructed by inserting an 11.4-kb PsiI-HpaI fragment of pBC633 containing Tn4401 into an EcoRV-restricted pACYC184 vector. Recombinant plasmid pTn4401 was electroporated into E. coli DH10B, and transformants expressing reduced susceptibility to imipenem and resistance to chloramphenicol were retained for further analysis. Transposition experiments were performed as described for pBC633. Transconjugants were additionally screened for resistance to chloramphenicol. Screening for chloramphenicol resistance together with plasmid extraction revealed that in up to 20% of cases, pTn4401 was also transferred to E. coli J53 Azr, resulting in Ticr Genr Azr Cmr colonies (Table 1). Transfer of both plasmids, pTn4401 and pOX38-Gen, suggests cointegration events that occur during replicative transposition, the transposition mechanism used by Tn3-like transposons (23). To detect true transposition events, Cm-susceptible Ticr Genr Azr colonies were searched for (Table 1). Analysis of their plasmid content revealed a unique plasmid with a size compatible with the presence of pOX38-Gen::Tn4401 (Fig. 2B). In order to study more putative transposants and their target site insertion, E. coli J53 transconjugants harboring pTn4401 and exhibiting a band corresponding in size to insertion of pTn4401 in pOX38-Gen were used for a second mating-out experiment to transfer pOX38-Gen::Tn4401 from E. coli J53 to E. coli DH10B Strr (Fig. 2B). Transconjugants were successfully obtained, and pOX38-Gen::Tn4401 transposants were further characterized. Transposition experiments were then performed in triplicate. Results of screening for Cm resistance and calculated frequencies of transposition events are indicated in Table 1. Overall, Tn4401 was capable of true transposition events (80% of events), in addition to cointegration (20% of events), which could subsequently be resolved in the recipient cells. Transposition of Tn4401 and conjugation of pOX38-Gen::Tn4401 occurred at high frequency (4.4 × 10−6/recipient cell) (Table 1). Given that the conjugation frequency of pOX38-Gen ranged from 10−2 to 10−3/recipient cell, the true transposition frequency is around 10−3 to 10−4/recipient plasmid.
Table 1.
Results of the three independent transposition experiments performed with the recombinant plasmid pTn4401
| Expt | Frequency (%) of event/recipient cella |
||||||
|---|---|---|---|---|---|---|---|
| TcR Cms | TcR Cmr | Total for TcR | Recipient strain J53Azr | Cointegration events | Transposition events | Total | |
| 1 | 9 × 101 (82) | 2 × 101 (18) | 1.1 × 102 | 392 × 105 | 5 × 10−7 | 2.3 × 10−6 | 2.8 × 10−6 |
| 2 | 8 × 101 (80) | 2 × 101 (20) | 1.6 × 102 | 324 × 105 | 9.9 × 10−7 | 3.9 × 10−6 | 4.9 × 10−6 |
| 3 | 1 × 102 (91) | 1 × 101 (9) | 1.1 × 102 | 350 × 105 | 4.9 × 10−7 | 4.9 × 10−6 | 5.4 × 10−6 |
| Avg ± SD | 6.6 × 10−7 ± 2.8 × 10−7 | 3.7 × 10−6 ± 1.3 × 10−6 | 4.4 × 10−6 ± 1.4 × 10−6 | ||||
TcR, transconjugant; TcR Cms, transconjugants in E. coli J53 Azr Ticr Genr Azr Cms; TcR Cmr, transconjugants in E. coli J53 Azr Ticr Genr Azr Cmr.
To determine whether Tn4401 had a target site preference, several pOX38-Gen::Tn4401 plasmids isolated from independent experiments were extracted and analyzed by direct sequencing of the ends of Tn4401 using laboratory-designed primers (13). As previously shown with natural plasmids (4, 13), Tn4401 insertions occurred at different sites (Fig. 3C). Alignment of these insertion site sequences did not reveal conserved motifs. Nevertheless, a 5-bp target site duplication, consistent with a transposition event, was observed in all the pOX38-Genr::Tn4401 plasmids studied (Fig. 3B and C).
Analysis of the genetic elements associated with resistance genes give information about their means of acquisition and diffusion. In Gram-negative rods, many ß-lactamase genes have been described as gene cassettes inserted into class 1 integrons, such as blaVEB (14), blaVIM (7), blaCARB-9 (16), and blaGES (5), or surrounded by insertion sequences forming composite transposons, such as blaOXA-48 (2) and blaPER (17). Tn3-based transposons have been reported to be associated with blaTEM genes (19), which encode the most widespread ß-lactamase in the Enterobacteriaceae. Since the first description of an E. coli isolate resistant to aminopenicillins due to the production of TEM-1 ß-lactamase in the 1960s, a rapid increase in the prevalence of TEM-1-producing isolates has occurred. The corresponding bla gene was carried by Tn3 elements associated with plasmids of different incompatibility groups, which subsequently were responsible for the rapid spread of TEM variants. Moreover, Tn3 was shown to efficiently transpose the blaTEM ampicillin resistance gene marker (8, 22).
Similarly, Tn4401 is a Tn3-like transposon that is associated with blaKPC genes (13). This transposon was identified in isolates from different geographical origins, and of different sequence types (ST), in Enterobacteriaceae (1, 4, 11, 12, 13) and in P. aeruginosa (13, 18). In all cases, Tn4401 was inserted at different loci and on plasmids varying in size and incompatibility group (1, 4, 13). We have demonstrated here that Tn4401 is capable of transposition with 5-bp target site duplication and without target site specificity. This work confirms that KPC genes benefit from all the genetic tools available—active transposons, self-transferable plasmids, and efficient ST types—for their rapid spread to various bacterial species.
Acknowledgments
This work was funded by INSERM, France, and by a grant from the European Community (7th Framework program FP7/2007-2013 under grant agreement no. 241742).
Footnotes
Published ahead of print on 15 August 2011.
REFERENCES
- 1. Andrade N. L., et al. 2011. Dissemination of blaKPC-2 by the spread of Klebsiella pneumoniae clonal complex 258 clones (ST258, ST11, ST437) and plasmids (IncFII, IncN, IncL/M) among Enterobacteriaceae species in Brazil. Antimicrob. Agents Chemother. 55:3579–3583 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2. Aubert D., Naas T., Héritier C., Poirel L., Nordmann P. 2006. Functional characterization of IS1999, an IS4 family element involved in mobilization and expression of β-lactam resistance genes. J. Bacteriol. 188:6506–6514 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3. Bratu S., et al. 2005. Emergence of KPC-possessing Klebsiella pneumoniae in Brooklyn, New York: epidemiology and recommendations for detection. Antimicrob. Agents Chemother. 49:3018–3020 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4. Cuzon G., et al. 2010. Worldwide diversity of Klebsiella pneumoniae that produce ß-lactamase blaKPC-2 gene. Emerg. Infect. Dis. 16:1349–1356 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5. Dubois V., et al. 2002. Molecular characterization of a novel class 1 integron containing blaGES-1 and a fused product of aac3-Ib/aac6′-Ib′ gene cassettes in Pseudomonas aeruginosa. Antimicrob. Agents Chemother. 46:638–645 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6. Gootz T. D., et al. 2009. Genetic organization of transposase regions surrounding blaKPC carbapenemase genes on plasmids from Klebsiella strains isolated in a New York City hospital. Antimicrob. Agents Chemother. 53:1998–2004 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7. Hammami S., et al. 2010. Diversity in VIM-2-encoding class 1 integrons and occasional blaSHV2a carriage in isolates of a persistent, multidrug-resistant Pseudomonas aeruginosa clone from Tunis. Clin. Microbiol. Infect. 16:189–193 [DOI] [PubMed] [Google Scholar]
- 8. Heritage J., Hawkey P. M., Todd N., Lewis I. J. 1992. Transposition of the gene encoding a TEM-12 extended-spectrum ß-lactamase. Antimicrob. Agents Chemother. 36:1981–1986 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9. Johnson R. C., Reznikoff W. S. 1984. Copy number control of Tn5 transposition. Genetics 107:9–18 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10. Kieser T. 1984. Factors affecting the isolation of cccDNA from Streptomyces lividans and Escherichia coli. Plasmid 12:19–36 [DOI] [PubMed] [Google Scholar]
- 11. Leavitt A., Chmelnitsky I., Ofek I., Carmeli Y., Navon-Venezia S. 2010. Plasmid pKpQIL encoding KPC-3 and TEM-1 confers carbapenem resistance in an extremely drug-resistant epidemic Klebsiella pneumoniae strain. J. Antimicrob. Chemother. 65:243–248 [DOI] [PubMed] [Google Scholar]
- 12. Miriagou V., et al. 2003. Imipenem resistance in a Salmonella clinical strain due to plasmid-mediated class A carbapenemase KPC-2. Antimicrob. Agents Chemother. 47:1297–1300 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13. Naas T., et al. 2008. Genetic structures at the origin of acquisition of the ß-lactamase blaKPC gene. Antimicrob. Agents Chemother. 52:1257–1263 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14. Naas T., Poirel L., Karim A., Nordmann P. 1999. Molecular characterization of In50, a class 1 integron encoding the gene for the extended-spectrum ß-lactamase VEB-1 in Pseudomonas aeruginosa. FEMS Microbiol. Lett. 176:411–419 [DOI] [PubMed] [Google Scholar]
- 15. Nordmann P., Cuzon G., Naas T. 2009. The real threat of KPC carbapenemase-producing bacteria. Lancet Infect. Dis. 9:321–331 [DOI] [PubMed] [Google Scholar]
- 16. Petroni A., et al. 2004. CARB-9, a carbenicillinase encoded in the VCR region of Vibrio cholerae non-O1, non-O139 belongs to a family of cassette-encoded ß-lactamases. Antimicrob. Agents Chemother. 48:4042–4046 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17. Poirel L., Cabanne L., Nordmann P. 2005. Genetic environment and expression of the extended-spectrum ß-lactamase blaPER-1 gene in gram-negative bacteria. Antimicrob. Agents Chemother. 49:1708–1713 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18. Poirel L., Lagrutta E., Cleary T., Munoz-Price L. S., Nordmann P. 2010. Emergence of KPC-producing Pseudomonas aeruginosa, United States. Antimicrob. Agents Chemother. 54:3072. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19. Poirel L., Naas T., Nordmann P. 2008. Genetic support of extended-spectrum ß-lactamases. Clin. Microbiol. Infect. 14(Suppl. 1):75–81 [DOI] [PubMed] [Google Scholar]
- 20. Robledo I. E., et al. 2010. Detection of KPC in Acinetobacter spp. in Puerto Rico. Antimicrob. Agents Chemother. 54:1354–1357 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21. Shen P., et al. 2009. Novel genetic environment of the carbapenem-hydrolyzing ß-lactamase KPC-2 among Enterobacteriaceae in China. Antimicrob. Agents Chemother. 53:4333–4338 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22. Sirot D., et al. 1991. Translocation of antibiotic resistance determinants including an extended-spectrum ß-lactamase between conjugative plasmids of Klebsiella pneumoniae and Escherichia coli. Antimicrob. Agents Chemother. 35:1576–1581 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23. Turlan C., Chandler M. 2000. Playing second fiddle: second-strand processing and liberation of transposable elements from donor DNA. Trends Microbiol. 8:268–274 [DOI] [PubMed] [Google Scholar]
- 24. Villegas M. V., et al. 2007. First identification of Pseudomonas aeruginosa isolates producing a KPC-type carbapenem-hydrolyzing ß-lactamase. Antimicrob. Agents Chemother. 51:1553–1555 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25. Wolter D. J., et al. 2009. Phenotypic and enzymatic comparative analysis of the novel KPC variant KPC-5 and its evolutionary variants, KPC-2 and KPC-4. Antimicrob. Agents Chemother. 53:557–562 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26. Yigit H., et al. 2001. Novel carbapenem-hydrolyzing ß-lactamase KPC-1 from a carbapenem-resistant strain of Klebsiella pneumoniae. Antimicrob. Agents Chemother. 45:1151–1161 [DOI] [PMC free article] [PubMed] [Google Scholar]