Abstract
In the crystal structure of the title compound, C8H6N2O3, the molecules are linked by a pairs of intermolecular N—H⋯O hydrogen bonds, forming inversion dimers. The aldehyde group is in the same plane as the pyrimidine ring [with a maximum deviation of 0.083 (2) Å for the O atom), and the linear propargyl group [C—C—C = 178.99 (19)°] makes a dihedral angle of 74.36 (13)° with the ring.
Related literature
For applications of acyclic pyrimidine nucleosides, see: De Clercq (2009 ▶, 2010a
▶,b
▶); Fan et al. (2011 ▶).
Experimental
Crystal data
C8H6N2O3
M r = 178.15
Monoclinic,

a = 5.1756 (7) Å
b = 8.4877 (12) Å
c = 18.565 (3) Å
β = 90.611 (2)°
V = 815.5 (2) Å3
Z = 4
Mo Kα radiation
μ = 0.11 mm−1
T = 296 K
0.41 × 0.37 × 0.25 mm
Data collection
Bruker SMART CCD area-detector diffractometer
Absorption correction: multi-scan (SADABS; Bruker, 1997 ▶) T min = 0.955, T max = 0.972
5826 measured reflections
1520 independent reflections
1261 reflections with I > 2σ(I)
R int = 0.020
Refinement
R[F 2 > 2σ(F 2)] = 0.040
wR(F 2) = 0.123
S = 1.08
1520 reflections
118 parameters
H-atom parameters constrained
Δρmax = 0.14 e Å−3
Δρmin = −0.23 e Å−3
Data collection: SMART (Bruker, 1997 ▶); cell refinement: SAINT (Bruker, 1997 ▶); data reduction: SAINT; program(s) used to solve structure: SHELXS97 (Sheldrick, 2008 ▶); program(s) used to refine structure: SHELXL97 (Sheldrick, 2008 ▶); molecular graphics: SHELXTL (Sheldrick, 2008 ▶); software used to prepare material for publication: SHELXTL.
Supplementary Material
Crystal structure: contains datablock(s) I, global. DOI: 10.1107/S1600536811032272/is2760sup1.cif
Structure factors: contains datablock(s) I. DOI: 10.1107/S1600536811032272/is2760Isup2.hkl
Supplementary material file. DOI: 10.1107/S1600536811032272/is2760Isup3.cml
Additional supplementary materials: crystallographic information; 3D view; checkCIF report
Table 1. Hydrogen-bond geometry (Å, °).
| D—H⋯A | D—H | H⋯A | D⋯A | D—H⋯A |
|---|---|---|---|---|
| N2—H2⋯O1i | 0.86 | 1.98 | 2.8329 (18) | 174 |
Symmetry code: (i)  .
.
Acknowledgments
This work was supported by the National Natural Science Foundation of China (No. 20972042) and the Natural Science Foundation of Department of Education of Henan Province (No. 2008 A150013).
supplementary crystallographic information
Comment
Acyclic pyrimidine nucleosides have drawn much attention because of their insteresting structures and broad utilizations as effective drugs for the treatment of diseases caused by herpes simplex virus (HSV) and varizella zoster (VZV) (De Clercq, 2009, 2010a,b). The title compound can be used as a powerful synthon for the preparation of acyclic pyrimidine nucleoside derivatives with potential biological activities due to the rich and extensive chemistry of the aldehyde carbonyl (Fan, 2011). Herein, we report the synthesis and crystal structure of the title compound.
In the title compound, C8H6N2O3, all the atoms in the pyrimidine ring, atoms connected directly with the pyrimidine ring and atoms in the aldehyde carbonyl group in the 5-position of the pyrimidine ring are in the same plane, which means there is a big conjugated system in the molecule. The linear structure of the propynyl group is connected with the big plane at an angle of 150.3°. In the crystal structure, the molecules are linked via intermolecular N—H···O hydrogen bond.
Experimental
To a solution of K2S2O8 (16.5 mmol) and CuSO4 (3.2 mmol) in 30 ml H2O was added a CH3CN solution (25 ml) of 5-methyl-1-(prop-2-ynyl)pyrimidine-2,4(1H,3H)-dione (8 mmol) and 2,6-lutidine (3.2 ml). The mixture was stirred at 60 °C for 5 h. Upon completion, the mixture was concentrated to half of the initial volume, and the remaining solution was extracted with EtOAc. The organic layer was washed with H2O. The aqueous layers were combined and back-extracted with CHCl3. Then the organic layers were combined, dried over Na2SO4, and then concentrated. The residue was purified through silica gel column chromatography with a mixture of methylene chloride-methanol (60:1, v/v) as eluent to give 1,2,3,4-tetrahydro-2,4-dioxo-1-(prop-2-ynyl)- pyrimidine-5-carbaldehyde. Single crystals of the title compound were obtained by slow evaporation of the solvent from a methylene chloride-methanol (60:1 v/v) solution.
Refinement
H atoms were positioned geometrically and refined using riding model, with C—H = 0.93 or 0.97 Å, and N—H = 0.86 Å, and with Uiso(H) = 1.2Ueq(C, N).
Figures
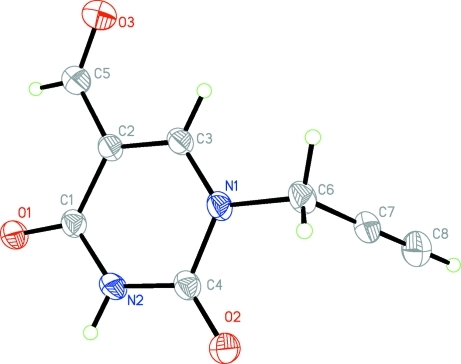
Fig. 1.
Molecular structure of the title compound, with displacement ellipsoids drawn at the 30% probability level.
Fig. 2.
Crystal packing of the title compound with view along the a axis. Intermolecular N—H···O hydrogen bonds are shown as dashed lines.
Crystal data
| C8H6N2O3 | F(000) = 368 |
| Mr = 178.15 | Dx = 1.451 Mg m−3 |
| Monoclinic, P21/n | Mo Kα radiation, λ = 0.71073 Å |
| Hall symbol: -P 2yn | Cell parameters from 2188 reflections |
| a = 5.1756 (7) Å | θ = 2.6–26.7° |
| b = 8.4877 (12) Å | µ = 0.11 mm−1 |
| c = 18.565 (3) Å | T = 296 K |
| β = 90.611 (2)° | Block, colourless |
| V = 815.5 (2) Å3 | 0.41 × 0.37 × 0.25 mm |
| Z = 4 |
Data collection
| Bruker SMART CCD area-detector diffractometer | 1520 independent reflections |
| Radiation source: fine-focus sealed tube | 1261 reflections with I > 2σ(I) |
| graphite | Rint = 0.020 |
| phi and ω scans | θmax = 25.5°, θmin = 2.6° |
| Absorption correction: multi-scan (SADABS; Bruker, 1997) | h = −6→6 |
| Tmin = 0.955, Tmax = 0.972 | k = −10→10 |
| 5826 measured reflections | l = −22→21 |
Refinement
| Refinement on F2 | Primary atom site location: structure-invariant direct methods |
| Least-squares matrix: full | Secondary atom site location: difference Fourier map |
| R[F2 > 2σ(F2)] = 0.040 | Hydrogen site location: inferred from neighbouring sites |
| wR(F2) = 0.123 | H-atom parameters constrained |
| S = 1.08 | w = 1/[σ2(Fo2) + (0.0694P)2 + 0.1695P] where P = (Fo2 + 2Fc2)/3 |
| 1520 reflections | (Δ/σ)max < 0.001 |
| 118 parameters | Δρmax = 0.14 e Å−3 |
| 0 restraints | Δρmin = −0.23 e Å−3 |
Special details
| Geometry. All e.s.d.'s (except the e.s.d. in the dihedral angle between two l.s. planes)are estimated using the full covariance matrix. The cell e.s.d.'s are takeninto account individually in the estimation of e.s.d.'s in distances, anglesand torsion angles; correlations between e.s.d.'s in cell parameters are onlyused when they are defined by crystal symmetry. An approximate (isotropic)treatment of cell e.s.d.'s is used for estimating e.s.d.'s involving l.s. planes. |
| Refinement. Refinement of F2 against ALL reflections. The weighted R-factor wR andgoodness of fit S are based on F2, conventional R-factors R are basedon F, with F set to zero for negative F2. The threshold expression ofF2 > σ(F2) is used only for calculating R-factors(gt) etc. and isnot relevant to the choice of reflections for refinement. R-factors basedon F2 are statistically about twice as large as those based on F, and R-factors based on ALL data will be even larger. |
Fractional atomic coordinates and isotropic or equivalent isotropic displacement parameters (Å2)
| x | y | z | Uiso*/Ueq | ||
| C1 | 0.7042 (3) | 0.8893 (2) | 0.44458 (8) | 0.0374 (4) | |
| C2 | 0.5046 (3) | 0.8128 (2) | 0.40196 (8) | 0.0376 (4) | |
| C3 | 0.4934 (3) | 0.84333 (19) | 0.33033 (8) | 0.0372 (4) | |
| H3 | 0.3631 | 0.7959 | 0.3031 | 0.045* | |
| C4 | 0.8652 (3) | 1.01190 (19) | 0.33360 (8) | 0.0368 (4) | |
| C5 | 0.3180 (4) | 0.7061 (2) | 0.43522 (10) | 0.0510 (5) | |
| H5 | 0.3435 | 0.6787 | 0.4833 | 0.061* | |
| C6 | 0.6377 (3) | 0.9724 (2) | 0.21870 (8) | 0.0432 (4) | |
| H6A | 0.4685 | 0.9390 | 0.2017 | 0.052* | |
| H6B | 0.6512 | 1.0851 | 0.2110 | 0.052* | |
| C7 | 0.8363 (4) | 0.8923 (2) | 0.17686 (9) | 0.0464 (5) | |
| C8 | 0.9931 (4) | 0.8282 (3) | 0.14242 (11) | 0.0612 (6) | |
| H8 | 1.1175 | 0.7773 | 0.1151 | 0.073* | |
| N1 | 0.6616 (2) | 0.93903 (17) | 0.29646 (7) | 0.0370 (4) | |
| N2 | 0.8664 (3) | 0.98582 (16) | 0.40649 (7) | 0.0398 (4) | |
| H2 | 0.9818 | 1.0356 | 0.4312 | 0.048* | |
| O1 | 0.7344 (2) | 0.87340 (16) | 0.51012 (6) | 0.0490 (4) | |
| O2 | 1.0262 (2) | 1.09061 (15) | 0.30334 (6) | 0.0472 (4) | |
| O3 | 0.1324 (3) | 0.65141 (19) | 0.40376 (8) | 0.0685 (5) |
Atomic displacement parameters (Å2)
| U11 | U22 | U33 | U12 | U13 | U23 | |
| C1 | 0.0360 (8) | 0.0454 (9) | 0.0308 (8) | 0.0027 (7) | −0.0031 (6) | −0.0023 (7) |
| C2 | 0.0353 (8) | 0.0442 (9) | 0.0333 (8) | 0.0011 (7) | −0.0018 (6) | −0.0066 (7) |
| C3 | 0.0326 (8) | 0.0439 (9) | 0.0350 (8) | 0.0021 (7) | −0.0041 (6) | −0.0089 (7) |
| C4 | 0.0365 (8) | 0.0414 (9) | 0.0323 (8) | 0.0027 (7) | −0.0036 (7) | −0.0023 (7) |
| C5 | 0.0503 (10) | 0.0600 (11) | 0.0427 (10) | −0.0103 (9) | −0.0011 (8) | −0.0042 (8) |
| C6 | 0.0451 (10) | 0.0551 (10) | 0.0294 (8) | 0.0008 (8) | −0.0085 (7) | 0.0013 (7) |
| C7 | 0.0533 (11) | 0.0548 (11) | 0.0311 (8) | −0.0078 (9) | −0.0027 (8) | −0.0019 (8) |
| C8 | 0.0625 (13) | 0.0738 (14) | 0.0474 (11) | −0.0033 (11) | 0.0076 (10) | −0.0125 (10) |
| N1 | 0.0366 (7) | 0.0471 (8) | 0.0272 (7) | 0.0016 (6) | −0.0044 (5) | −0.0029 (6) |
| N2 | 0.0402 (8) | 0.0494 (8) | 0.0297 (7) | −0.0079 (6) | −0.0084 (5) | −0.0014 (6) |
| O1 | 0.0500 (7) | 0.0683 (8) | 0.0285 (6) | −0.0120 (6) | −0.0057 (5) | 0.0017 (5) |
| O2 | 0.0470 (7) | 0.0566 (8) | 0.0378 (7) | −0.0094 (6) | −0.0019 (5) | 0.0039 (5) |
| O3 | 0.0592 (9) | 0.0805 (11) | 0.0658 (10) | −0.0216 (7) | 0.0004 (7) | −0.0139 (8) |
Geometric parameters (Å, °)
| C1—O1 | 1.2325 (19) | C5—O3 | 1.211 (2) |
| C1—N2 | 1.374 (2) | C5—H5 | 0.9300 |
| C1—C2 | 1.449 (2) | C6—C7 | 1.462 (3) |
| C2—C3 | 1.356 (2) | C6—N1 | 1.475 (2) |
| C2—C5 | 1.465 (3) | C6—H6A | 0.9700 |
| C3—N1 | 1.351 (2) | C6—H6B | 0.9700 |
| C3—H3 | 0.9300 | C7—C8 | 1.173 (3) |
| C4—O2 | 1.211 (2) | C8—H8 | 0.9300 |
| C4—N2 | 1.371 (2) | N2—H2 | 0.8600 |
| C4—N1 | 1.397 (2) | ||
| O1—C1—N2 | 120.11 (14) | C7—C6—N1 | 112.24 (14) |
| O1—C1—C2 | 124.91 (15) | C7—C6—H6A | 109.2 |
| N2—C1—C2 | 114.98 (13) | N1—C6—H6A | 109.2 |
| C3—C2—C1 | 118.22 (15) | C7—C6—H6B | 109.2 |
| C3—C2—C5 | 120.68 (15) | N1—C6—H6B | 109.2 |
| C1—C2—C5 | 121.10 (15) | H6A—C6—H6B | 107.9 |
| N1—C3—C2 | 123.42 (14) | C8—C7—C6 | 178.99 (19) |
| N1—C3—H3 | 118.3 | C7—C8—H8 | 180.0 |
| C2—C3—H3 | 118.3 | C3—N1—C4 | 121.52 (13) |
| O2—C4—N2 | 123.44 (14) | C3—N1—C6 | 121.56 (13) |
| O2—C4—N1 | 122.30 (14) | C4—N1—C6 | 116.91 (14) |
| N2—C4—N1 | 114.26 (14) | C4—N2—C1 | 127.43 (13) |
| O3—C5—C2 | 123.80 (18) | C4—N2—H2 | 116.3 |
| O3—C5—H5 | 118.1 | C1—N2—H2 | 116.3 |
| C2—C5—H5 | 118.1 | ||
| O1—C1—C2—C3 | 179.16 (16) | O2—C4—N1—C3 | 175.63 (15) |
| N2—C1—C2—C3 | −0.7 (2) | N2—C4—N1—C3 | −4.1 (2) |
| O1—C1—C2—C5 | −0.4 (3) | O2—C4—N1—C6 | −4.8 (2) |
| N2—C1—C2—C5 | 179.81 (15) | N2—C4—N1—C6 | 175.44 (14) |
| C1—C2—C3—N1 | 1.3 (2) | C7—C6—N1—C3 | −106.92 (18) |
| C5—C2—C3—N1 | −179.19 (15) | C7—C6—N1—C4 | 73.53 (19) |
| C3—C2—C5—O3 | −7.1 (3) | O2—C4—N2—C1 | −174.72 (16) |
| C1—C2—C5—O3 | 172.38 (18) | N1—C4—N2—C1 | 5.0 (2) |
| C2—C3—N1—C4 | 1.3 (2) | O1—C1—N2—C4 | 177.47 (15) |
| C2—C3—N1—C6 | −178.27 (15) | C2—C1—N2—C4 | −2.7 (2) |
Hydrogen-bond geometry (Å, °)
| D—H···A | D—H | H···A | D···A | D—H···A |
| N2—H2···O1i | 0.86 | 1.98 | 2.8329 (18) | 174. |
Symmetry codes: (i) −x+2, −y+2, −z+1.
Footnotes
Supplementary data and figures for this paper are available from the IUCr electronic archives (Reference: IS2760).
References
- Bruker (1997). SADABS, SMART and SAINT Bruker AXS Inc., Madison, Wisconsin, USA.
- De Clercq, E. (2009). Rev. Med. Virol. 19, 287–299. [DOI] [PubMed]
- De Clercq, E. (2010a). J. Med. Chem. 53, 1438–1450. [DOI] [PubMed]
- De Clercq, E. (2010b). Antiviral Res. 85, 19–24. [DOI] [PubMed]
- Fan, X.-S., Wang, Y.-Y., Qu, Y.-Y., Xu, H.-Y., He, Y., Zhang, X.-Y. & Wang, J.-J. (2011). J. Org. Chem. 76, 982–985. [DOI] [PubMed]
- Sheldrick, G. M. (2008). Acta Cryst. A64, 112–122. [DOI] [PubMed]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Crystal structure: contains datablock(s) I, global. DOI: 10.1107/S1600536811032272/is2760sup1.cif
Structure factors: contains datablock(s) I. DOI: 10.1107/S1600536811032272/is2760Isup2.hkl
Supplementary material file. DOI: 10.1107/S1600536811032272/is2760Isup3.cml
Additional supplementary materials: crystallographic information; 3D view; checkCIF report