Abstract
Angiogenesis and bone formation are intimately related processes. Hypoxia during early bone development stabilizes hypoxia-inducible factor-1α (HIF-1α) and increases angiogenic signals including vascular endothelial growth factor (VEGF). Furthermore, stabilization of HIF-1α by genetic or chemical means stimulates bone formation. On the other hand, deficiency of Runx2, a key osteogenic transcription factor, prevents vascular invasion of bone and VEGF expression. This study explores the possibility that HIF-1α and Runx2 interact to activate angiogenic signals. Runx2 over-expression in mesenchymal cells increased VEGF mRNA and protein under both normoxic and hypoxic conditions. In normoxia, Runx2 also dramatically increased HIF-1α protein. In all cases, the Runx2 response was inhibited by siRNA-mediated suppression of HIF-1α and completely blocked by the HIF-1α inhibitor, echinomycin. Similarly, treatment of preosteoblast cells with Runx2 siRNA reduced VEGF mRNA in normoxia or hypoxia. However, Runx2 is not essential for the HIF-1α response since VEGF is induced by hypoxia even in Runx2-null cells. Endogenous Runx2 and HIF-1α were colocalized to the nuclei of MC3T3-E1 preosteoblast cells. Moreover, HIF-1α and Runx2 physically interact using sites within the Runx2 RUNT domain. Chromatin immunoprecipitation also provided evidence for colocalization of Runx2 and HIF-1α on the VEGF promoter. In addition, Runx2 stimulated HIF-1α-dependent activation of an HRE-luciferase reporter gene without requiring a separate Runx2-binding enhancer. These studies indicate that Runx2 functions together with HIF-1α to stimulate angiogenic gene expression in bone cells and may in part explain the known requirement for Runx2 in bone vascularization.
Keywords: Osteoblast, vascularization, angiogenesis, transcriptional factors, hypoxia
Bone development and regeneration require an adequate blood supply. During embryogenesis, blood vessels penetrate into the avascular cartilage anlage, a necessary event for subsequent bone formation. At least two important pathways are involved in bone vascularization. The first is driven by the hypoxic environment first present in early mesenchymal condensations destined to form bone and uses the angiogenic transcription factor, hypoxia-inducible factor 1α (HIF-1α), while the second requires the master transcriptional activator of bone formation, Runx2.
HIF-1α protein levels are tightly regulated by oxygen tension via controlled proteolysis [Salceda and Caro, 1997]. Under normoxic conditions, prolyl residues near the C-terminus of HIF-1α are hydroxylated by an oxygen-dependent prolyl hydroxylase that serves as an oxygen sensor. Hydroxylated HIF-1α binds to von Hippel-Lindau protein (VHL), a target of the E3 ubiquitin ligase complex [Maxwell et al., 1999]. The formation of the HIF-1-VHL complex leads to rapid degradation of HIF-1α protein by the ubiquitin-proteosome pathway. On the other hand, in hypoxic environments, prolyl hydroxylation of HIF-1α is blocked and the protein becomes resistant to proteolysis [Huang et al., 1998; Salceda and Caro, 1997]. The stabilized HIF-1-α protein accumulates in the nucleus where it heterodimerizes with the constitutively expressed HIFβ subunit (also known as the aryl hydrocarbon receptor nuclear translocator or ARNT). This αβ-heterodimeric HIF-1 transcription factor complex binds to hypoxia-response-elements (HRE; 5′RCGTG-3′, where R is A or G, [Semenza et al., 1996] in the promotor regions of angiogenic genes such as VEGF [Forsythe et al., 1996], erythropoietin [Firth et al., 1994] and transforming growth factor-β3 [Schaffer et al., 2003].
HIF-1α activation, angiogenesis and bone formation are intimately related events. During development, initial mesenchymal condensations destined to form limbs (first seen at E10.5 in mice) exist in a hypoxic environment that induces HIF-1α. This early HIF-1α expression is necessary for formation of normal cartilage primordia and subsequent hypertrophy as well as joint development[Provot et al., 2007]. Similarly, osteoblast-specific deletion of HIF-1α impairs long bone formation and vascularization while activation of HIF-1α by osteoblast-specific deletion of VHL stimulates bone formation[Wang et al., 2007].
Like HIF-1α, Runx2 is first expressed in mesenchymal condensations of developing limbs and persists throughout skeletal development[Ducy et al., 1997]. Although Runx2 is best understood as a master regulator of skeletogenesis, it is also essential for vascular invasion of bone primordia. In addition to completely blocking the formation of osteoblasts and hypertrophic chondrocytes, Runx2 deletion prevents vascular invasion of the cartilage anlag [Komori et al., 1997; Otto et al., 1997]. This vascularization defect is related to decreased expression of the angiogenic cytokine, vascular endothelial growth factor (VEGF), which can also be induced by transfection of cells with a Runx2 expression vector [Zelzer et al., 2001]. In addition, Runx2 is transiently expressed in vascular endothelial cells and vascular smooth muscle cells of developing blood vessels, which may also be related to its role in angiogenesis [Bronckers et al., 2005; Sun et al., 2001]
The observation that both HIF-1α and Runx2 have roles in bone formation and vascularization compelled us to examine whether these two factors interact to regulate angiogenic signals. As will be shown, both Runx2 and HIF-1α can induce VEGF and these two factors complement each other for maximal VEGF induction. Furthermore, Runx2 and HIF-1α physically interact in the nucleus of osteoblasts and on the chromatin of the VEGF gene. This study expands our understanding of the relationship between angiogenesis and bone formation, possibly providing the biological basis for integrating osteogenesis and angiogenesis in future therapeutic interventions to accelerate bone healing.
Materials and Methods
Cell culture
C3H10T1/2 cells, a pluripotent murine mesenchymal cell line, and HEK293 cells, both obtained from American Tissue Culture collection, were plated at the density of 50,000 cells/cm2 in DMEM containing 10% FBS (Hyclone Laboratories, Logan, UT, USA) and 1% antibiotics. MC3T3-E1 subclone 4 and 42 cells were maintained in α-MEM supplemented with 10% FBS and 1% antibiotics. Both cell lines express osteoblast markers and mineralize after growth in ascorbic acid-containing medium[Wang et al., 1999; Xiao et al., 1997]. An mTERT (mouse telomerase reverse transcriptase)-immortalized calvarial cell line from Runx2 null mice [Bae et al., 2007] was a generous gift from Dr. Jane Lian (University of Massachusetts Medical Center, Worcester, MA). These cells were maintained in α-MEM supplemented with 5% FBS and 1% antibiotics. Antibiotics were purchased from Invitrogen (10,000 Units/mL penicillin, 10mg/ml stereptomycin). For the induction of hypoxia, cells were grown in an atmosphere containing 1% O2, 5% CO2, and 94% Nitrogen using an air-tight chamber (Billups-Rothenburg, Del Mar, CA) after changing the media. Alternatively, hypoxia mimetic conditions were induced by culturing cells in CoCl2 (100μM, Sigma), a potent inhibitor of HIF-1α degradation [Hofer et al., 2001].
Adenoviral transduction and transfection with small interfering RNAs (siRNA)
Cells were transduced with adenovirus containing Runx2 cDNA (AdRunx2) or LacZ(AdLacZ, control) at the indicated titer as previously described [Yang et al., 2003]. After 48 h, cells were cultured under normoxic/ hypoxic/ hypoxia mimetic conditions and harvested at various time points. Cells were transfected with siRNA duplexes at 50% confluency using HiPerfect (Qiagen). Specific siRNAs for mouse HIF-1α (SI 00187915), Runx2 (SI 00193032) and negative control siRNA (1027280) were purchased from Qiagen and used at a concentration of 20 nM according to the manufacturer's instructions.
Measurement of Runx2, HIF-1α and VEGF protein levels
Runx2 and HIF-1α proteins were measured by Western blotting, Briefly, cells were washed with ice cold PBS and lysed in RIPA buffer containing complete protease inhibitor cocktail (Roche Diagnostics, IN), 1mM DTT, 1mM PMSF. Lysates were fractionated by SDS-PAGE on 4-12% precast minigels (InVitrogen) and transferred to PVDF membranes. The following primary antibodies were used at a 1:500 dillution: anti-HIF-1α (NB100-105, Novus Biologicals), anti-Runx2 (D130-3, MBL) and anti-tubulin (loading standard-Sigma). Second antibody of sheep anti-mouse or donkey anti-rabbit conjugated horseradish peroxidase was used at 1:10,0000 dillution. Blots were visualized by ECL (Amersham).
VEGF in conditioned medium was measured by ELISA using a Mouse VEGF Duo-Set ELISA kit (R&D systems) that recognizes 164 and 120 amino acid residue forms of the mouse protein. The amount of secreted VEGF was normalized to the total cellular protein in each well.
Quantitation of mRNA
Real-time quantitative RT-PCR was performed as described [Zhao et al., 2005]. Briefly, total RNA was extracted by TRIzol reagent (Invitrogen) and further purified by DNase treatment (Ambion). Reverse transcription was performed using Taqman reverse transcriptase reagents (Applied Biosystems) and 2ug of total RNA. Real time Q-PCR was performed using an ABI Prism 7700 Sequence Detection system (PE Applied Biosystems, Foster City, CA, USA). TaqMan® Universal PCR Master Mix and Optimized FAM labeled probes and primers were purchased from Applied Biosystems (TaqMan® Gene Assay probes); Mouse HIF-1α (Mn00468869_m1), Runx2 (Mn00501578_m1), VEGF (Mm00437304_m1), β-ACTIN (Mm01205647_91), GAPDH (Mm99999915_g1). The real-time PCR product, mRNA expression, was calculated based on a relative standard curve and normalized to GAPDH.
Immunofluorescence localization of Runx2 and HIF-1α
Confocal microscopy was performed as described previously[Li et al., 2010]. MC3T3-E1 clone 42 cells were grown on glass cover slips under normoxic or hypoxic conditions. After 8 h, cells were fixed with 4% formaldehyde and incubated overnight with primary antibodies to Runx2 and HIF-1α (1:100 dilutions). To localize the Runx2 and HIF-1α in the cell, the secondary antibody Alexa Fluor 555-conjugated donkey anti-rabbit (red color, Invitrogen) and Alexa Fluor 488-conjugated donkey anti-mouse (green color, Invitrogen) were added for 1 hour. Finally, the slides were mounted using mounting medium containing DAPI (blue color, ProLong Gold antifade reagent, Invitrogen). Fluorescence localization was evaluated with Olympus FluoView 500 laser scanning confocal microscope system (Microscopy and Image Analysis Laboratory, University of Michigan School of Medicine). The fully maximized single image in the sequential series of Z-planes was detected by searching for the digitally captured image with brightest fluorescent emission.
Co-immunoprecipitation (IP) studies
To measure the interaction of Runx2 with endogenous HIF-1α, MC3T3-E1 Clone 42 cells were cultured in α-MEM for 48 hrs. The proteosome inhibitor, MG132 (5 μM), was added to the medium 12 hrs before harvest. Cells were then scraped into RIPA buffer containing protease inhibitor cocktail (Roche), centrifuged and subjected to IP with Runx2 or HIF-1α antibodies. SDS-PAGE and Western Blotting following standard laboratory protocols were used to visualize the results. To determine the Runx2 domain responsible for the HIF-1α interaction, various C and N-terminal deletion constructs were generated by subcloning the desired PCR products into appropriate expression vectors (below) followed by verification using DNA sequencing. Wildtype Runx2 and Runx2 with C-terminal deletions after amino acids 410, 330, 286 or 258 (410dC, 330dC, 286dC, 258dC) were subcloned into the pCMV5-FLAG vector [Thirunavukkarasu et al., 1998]. Wildtype Runx2 WT and Runx2 with N-terminal deletions to amino acid 97, 232, 242, 376 (dN97, 232, 242, 376) were subcloned into the pPGS-NEO-CITE-HA carrier DNA vector. Expression constructs were transfected into HEK293 cells together with HIF-1α expression vector using lipofectamine (Invitrogen). For each sample, 20 ul of Protein A/G plus (Santa Cruz) beads and 0.5 ∼ 1 ug of each indicated antibody were used for IP. Finally, the beads were boiled in 60 ul of 1X SDS sample buffer and 20 ul of each sample was used for SDS-PAGE analysis. Western Blotting was performed after the samples were transferred to PVDF membranes. Studies used the following antibodies: HIF-1α (NB100-105, Novus Biologicals), Runx2 (D130-3, MBL), α-tubulin (Sigma), HA (MMS-101P Covance) and FLAG (anti-M2-HRP, Sigma).
Chromatin immunoprecipitation (ChIP)
ChIP assays were performed as previously described [Li et al., 2010; Roca and Franceschi, 2008; Roca et al., 2005]. MC3T3-E1 clone 4 cells were cultured for 48 hrs and exposed to normoxic or hypoxic conditions for an additional 8 h. Cells were then fixed with 1% p-formaldehyde for 10 minutes at room temperature to make protein-DNA complexes. Chromatin was mechanically sheared by sonication to yield fragmentswith a mean size of 300 bp. Before immunoprecipitation, chromatin (10 μg DNA/assay) was pretreated with protein A/G agarose beads (Santa-Cruz Biotechnology) for 1 hour. For the immunoprecipitation, the supernatants were incubated with primary antibody (Runx2, HIF-1α, and IgG) and A/G agarose beads were used to precipitate the fragments. The input DNA and the DNA from ChIP samples was used for PCR analysis (35 cycles). Putative Runx2 and HIF-1α (hypoxia response element, HRE) binding sites in the VEGF-A promoter were analyzed using following primer sets (position on promoter is indicated): Runx2 site 5′ primer (Primer 1 = -334 ∼ -315), 5′-GCCTTCCAACCCCTACTTTC -3′; Runx2 site 3′ primer (Primer 2 = -201 ∼ -182), 5′-ATCTGTGCACCCCTTCAAAC-3′; HRE 5′ primer (Primer 3 = -952 ∼ -971), 5′-CCCAGCTGTCTCTCCTTCAG-3′; HRE 3′ primer (Primer 4 = -717 ∼ -698), 5′-ATATGTGGGGAGGGGGTTAC-3′. Primers were also designed to detect the presence of DNA within the VEGF transcribed region, which served as a control for off-target immunoprecipitation (Primer 5 = +285-304, 5′-TGTGGAAATCAGCAGACGAA-3′; Primer 6 = +491-510, 5′-GCGGTGTCTGTCTGTCTGTC-3′).
Luciferase assay
For the reporter gene assays, C3H10T1/2 cells were transiently transfected with a HRE (hypoxia responsive element)-Luc reporter plasmid composed of pGL2 vector with 68bp of Enolase 1 promoter sequence containing 2 HRE sites [Semenza et al., 1996]. Cells were plated in six well plates a density of 5 × 104 cell/cm2, grown for 12 h, and transfected with 250 ηg of HRE-luciferase reporter construct (pGL2-HRE) plus 250 ηg of pGL4-SV40 Renilla luciferase vector (Promega) for normalization. 500 ηg of RUNX2 expression vector (pCMV5-Runx2), control (pCMV5-LacZ) vector, or HIF-1α (pCEP-HIF-1a) expression plasmid were utilized. Luciferase activity was measured with the reagents and protocols from Promega and were subjected to normoxic/ hypoxic conditions for 24hrs.
Statistical Analysis
Results are presented as mean ± SE, with n = 3 per group for all comparisons. Statistical significance was assessed using a one-way ANOVA followed by Tukey's multiple-comparison test. All experiments were repeated at least twice.
Results
Runx2 over-expression increases VEGF and HIF-1α
Developmental studies suggest a possible functional interaction between Runx2 and HIF-1α in regulating angiogenesis. To determine whether increased Runx2 can stimulate angiogenetic signaling, pluripotent C3H10T1/2 mesenchymal cells, which do not express detectable Runx2, were transduced with a Runx2-expressing adenovirus (AdRunx2) and assayed for VEGF and HIF-1α protein and mRNA after exposure to normoxic, hypoxic or hypoxia mimetic conditions (CoCl2 treatment). Consistent with previous reports [Zelzer et al., 2001], Runx2 dose-dependently increased VEGF protein (Fig 1A,B) and mRNA levels (Fig 1G). The Runx2 response was most dramatic in the normoxic condition where, depending on the experiment, 2.8 to 8-fold increases in VEGF secretion were observed (compare normoxic condition in Fig 1A and B). The magnitude of Runx2 stimulation was comparable to that seen with hypoxia or hypoxia-mimetic conditions. Runx2 also increased VEGF in hypoxia and in the presence of CoCl2, although the fold stimulation was somewhat less, possibly because of the elevated VEGF levels seen under these conditions. In addition, Runx2 increased the normally low levels of HIF-1α protein seen in normoxia, but did not affect the elevated levels of HIF-1α seen in the hypoxic condition (Fig 1C,D). This response appears to be at the protein level in that Runx2 did not affect HIF-1α mRNA (Fig 1E). Taken together, these studies show that Runx2 overexpression can induce VEGF mRNA and protein and that this effect is additive with hypoxia in stimulating this angiogenic signal.
Fig 1. Runx2 regulation of VEGF and HIF-1α.
C3H10T1/2 cells were transduced with AdRunx2 at the indicated multiplicity of infection (MOI), grown for 48 hours and cultured for 18 h under normoxic / hypoxic conditions or in the presence or absence of 100μM CoCl2. VEGF in conditioned medium was analyzed by ELISA (A, B) while HIF-1α, Runx2 and α-tubulin levels were determined by Western blotting (C,D). For mRNA analysis (E-G), cells were transduced at an moi of 10 using control AdLacZ virus (-) or AdRunx2 (+) and transferred to normoxic/hypoxic conditions for 8 hours before measurement of HIF-1α (E), Runx2 (F) and VEGF (G) mRNAs by QRT-PCR. Values are means ± SD of triplicate independent samples. Statistics: a, significantly different from normoxic LacZ control; b, significantly different from hypoxic LacZ control; c, significantly different from corresponding normoxic sample (same AdRunx2 titer). For all comparisons, p < 0.05.
Depletion of endogenous Runx2 using both siRNA and genetic approaches inhibits VEGF expression in osteoblasts, but does not prevent the response to hypoxia
Because the studies in Fig. 1 used overexpression to demonstrate a role for Runx2 in VEGF synthesis, we considered it important to also determine whether endogenous Runx2 is required for VEGF expression. Two approaches were taken. In the first, siRNA was used to suppress Runx2 in MC3T3-E1 preosteoblast cells. These cells normally express Runx2 at high levels and differentiate into osteoblasts after growth in osteogenic medium[Wang et al., 1999]. Runx2 siRNA selectively reduced Runx2 protein and mRNA levels by 50-65% in both normoxic and hypoxic conditions without affecting either HIF-1α mRNA or α-tubulin (Fig 2AB). Although these cells do not express enough VEGF to be readily detected by ELISA (result not shown), exposure to hypoxia induced VEGF mRNA approximately 7-fold (Fig 2C). Treatment with Runx2 siRNA reduced VEGF mRNA by approx. 30 percent in the normoxic condition and by 40 percent in hypoxia. However, cells were still able to respond to the hypoxic stimulus, albeit at a reduced level (i.e. Hypoxia stimulated VEGF mRNA 7-fold in control cells and this induction was reduced to only 6-fold with Runx2 siRNA treatment).
Fig 2. Runx2 is necessary for maximal VEGF expression. A-C.
Runx2 siRNA inhibits hypoxic induction of VEGF mRNA. MC3T3-E1 clone 4 cells were treated with 20 nM Runx2 siRNA (+) or control siRNA (-) for 36h and cultured for an additional 8 h in normoxic or hypoxic conditions. RNA and protein were then isolated for measurement of Runx2 (A), HIF-1α (B) and VEGF mRNAs (C) by QRT-PCR or Runx2 protein (A, lower panel). D. Response of Runx2-null cells to hypoxia. An mTert-immortalized calvarial cell line from Runx2 (-/-) mice was exposed to control or hypoxic conditions with (+) or without (-) transduction with AdRunx2 (MOI = 10) before measurement of medium VEGF by ELISA. Conditions were as in Fig. 1A-D). Statistics. Panels A-C: a, significantly different from siRNA control. Panel D: a, significantly different from corresponding LacZ control; b, significantly different from corresponding normoxic sample. For all comparisons, p < 0.05.
Since the siRNA approach only partially reduced Runx2 levels, a second strategy was used to determine whether hypoxia can induce VEGF in the complete absence of Runx2. For this study, we used a calvarial cell line derived from Runx2-null mice. These cells fail to undergo osteoblast differentiation unless Runx2 is restored by transfection[Bae et al., 2007]. Exposure of Runx2-null cells to hypoxia increased VEGF protein approximately 4-fold (Fig 2D). Similar to results obtained with C3H10T1/2 cells in Fig 1, AdRunx2 caused a 3-fold increase in VEGF in the normoxic condition and a 2.5-fold increase in hypoxia. These results show that Runx2 can increase VEGF expression in normoxic and hypoxic conditions, but this factor is not essential for the basal HIF-1α-mediated response of cells to hypoxia.
Dependence of the Runx2 response on HIF-1α and effects of Runx2 on HIF-1α-dependent transcriptional activity
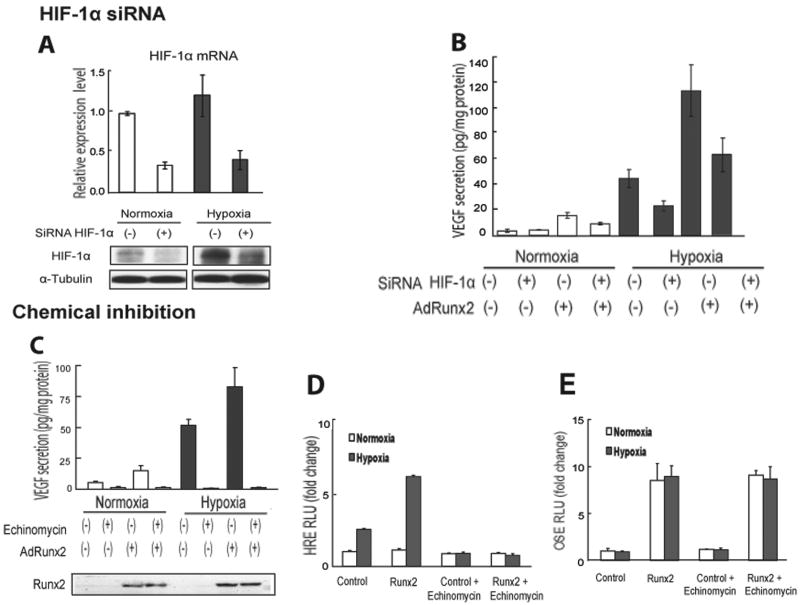
The next series of studies examined whether Runx2 induction of VEGF requires HIF-1α. Both siRNA (Fig 3AB) and chemical inhibition approaches (Fig 3C) were used to address this issue. Treatment of C3H10T1/2 cells with a specific siRNA reduced HIF-1α mRNA by 70 percent in both normoxic and hypoxic conditions (Fig 3A). Densitometric analysis of Western blots also indicated substantial reduction in HIF-1α protein. In normoxia, siRNA treatment reduced the already low HIF-1α protein to below the level of detection. However, substantial amounts of HIF-1α protein were still present in the hypoxic condition, probably as a consequence of stabilization in the low oxygen environment. Under this condition, HIF-1α siRNA reduced protein levels by about 50 percent. Effects of HIF-1α suppression were also apparent when VEGF protein levels were examined (Fig 3B). In both normoxic and hypoxic conditions, HIF-1α siRNA blocked the Runx2-dependent induction of VEGF by approximately 50%. To gain further insight into the role of HIF-1α in the Runx2 response, we also blocked HIF-1α signaling with the specific inhibitor, echinomycin. This compound selectively inhibits the ability of HIF-1α to interact with hypoxia response elements (HREs) in the regulatory regions of target genes[Kong et al., 2005]. As shown in Figure 3C, echinomycin completely blocked hypoxia and Runx2-dependent induction of VEGF without affecting Runx2 levels.
Fig. 3. Runx2 induction of VEGF requires HIF-1α.
A,B. siRNA suppression of HIF-1α inhibits VEGF secretion. C3H10T1/2 cells were transfected with control (-) or HIF-1α siRNA (+). 24 h later, cells were transduced with AdRunx2 or AdLacZ control virus and grown in normoxic/hypoxic conditions for an additional 24 h. Cells and medium were then harvested for measurement of HIF-1α mRNA and protein (A) and medium VEGF (B). C. The specific HIF-1α inhibitor, echinomycin, blocks hypoxia-induced VEGF secretion. Cells were transduced with AdlacZ or AdRunx2 (MOI = 20) and grown in normoxic or hypoxic conditions for 24 h with or without echinomycin (20 nM) as indicated. Cell layers and medium were harvested for measurement of Runx2 and VEGF, respectively. (D, E) Specificity of echinomycin inhibition and effects of Runx2 on HRE-dependent transcriptional activity. Cells were transfected with either a 6HRE-luc reporter (D) or with a 6OSE2-luc reporter (E) and grown with or without Runx2 for 24 h under normoxic or hypoxic conditions with or without echinomycin treatment before measurement of luciferase as described in Methods. Statistical analysis: a, p<0.05.
Figure 3D shows that Runx2 can also stimulate HIF-1α-dependent activation of an HRE-luc reporter gene[Semenza et al., 1996] in cells exposed to hypoxia. Since this reporter can only directly respond to HIF-1α (e.g. it does not contain Runx2 or other transcription factor binding sites), this stimulation is most likely related to Runx2 binding/stabilization of HIF-1α. In contrast, a Runx2 reporter gene, 6OSE2-luc[Ducy et al., 1997], was clearly stimulated by Runx2 in normoxic and hypoxic conditions, but hypoxia and HIF-1α stabilization did not further activate this reporter. Consistent with echinomycin having selective effects on HIF-1α-dependent transcription, it totally blocked hypoxia and Runx2-dependent induction of HRE-luc without affecting Runx2-specific induction of 6OSE2-luc.
These results taken together with the siRNA studies suggest that actions of Runx2 on VEGF expression may be dependent on HIF-1α. In contrast, HIF-1α can function in the absence of Runx2 although its activity is increased by this transcription factor.
Nuclear colocalization and physical interaction between Runx2 and HIF-1α; importance of the RUNT domain
To begin exploring the mechanism accounting for the functional interactions between Runx2 and HIF-1α demonstrated in Figures 1-3, we looked for a possible physical association between these two proteins. An initial study examined the subcellular localization of Runx2 and HIF-1α in MC3T3-E1 cells using confocal immunofluorescence microscopy (Fig. 4A). As previously reported[Javed et al., 2005]. Runx2 exhibited an exclusively nuclear distribution in both normoxic and hypoxic conditions (red channel). Although normoxic cells displayed weak HIF-1α fluorescence due to the known instability of this protein under this condition[Hirota and Semenza, 2006], the signal detected was mainly in the nucleus. Exposure of cells to hypoxia dramatically increased the overall HIF-1α signal with most of the protein again appearing in the nucleus. Interestingly, merging of the two signals indicated considerable colocalization of HIF-1α and Runx2 in the nuclear compartment (yellow color) in both normoxia and hypoxia.
Fig 4. Colocalization and physical association of Runx2 and HIF-1α.
(A) Co-localization of HIF-1α and Runx2. MC3T3-E1 clone 42 cells were grown on glass coverslips and exposed to normoxic or hypoxic conditions for 8 h., fixed and stained as described in Methods. Legend: HIF-1α (green), Runx2 (red) and DAPI (blue) for nuclear staining. Fluorescence localization was evaluated using laser scanning confocal microscopy. (B) Co-immunoprecipitation. Cells were grown for 48 hrs and MG132 was added to medium 12 hrs before harvesting. Cell lysates were immunoprecipitated with Runx2, HIF-1α antibody or IgG (control) followed by immunoblotting of Runx2 or HIF-1α antibody as indicated.
A series of co-immunoprecipitation (Co-IP) experiments were next carried out to further explore the basis for the Runx2-HIF-1α interaction. The first series of studies examined interactions between endogenous Runx2 and HIF-1α in MC3T3-E1 cells. Nuclear extracts were immunoprecipitated with either Runx2 or HIFα antibodies and the resulting precipitates were probed for Runx2 or HIF-1α on Western blots (Fig. 4B). These results indicate that Runx2 and HIF-1α form a protein complex in osteoblasts.
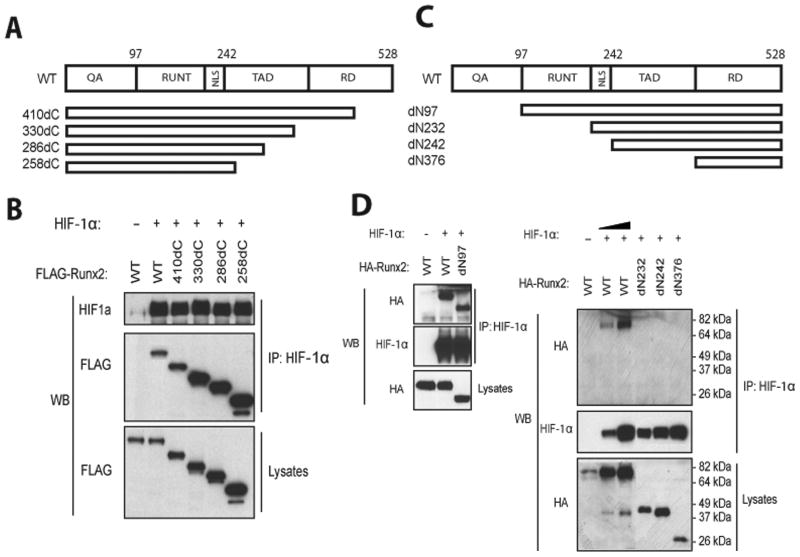
To identify the domain structures in Runx2 that are responsible for the interaction with HIF-1α, Flag-tagged C-terminal or HA-tagged N-terminal deletion constructs of Runx2 were generated and transfected into HEK293 cells together with a HIF-1α expression vector (Fig 5). Positions of deletions relative to the domain structure of Runx2 are shown in Figs 5A and C. All C-terminal Runx2 deletions available down to amino acid residue 258 could be immunoprecipitated with HIF-1α (Fig 5B). These deletions spanned a C-terminal repression domain and the proline/serine/threonine-rich activation domain. Analysis of N-terminal deletions indicated retention of binding if the first 97 N-terminal amino acid residues containing the glutamine/alanine-rich domain were deleted, but loss of binding with all further deletions starting with loss of the RUNT domain (Fig 5D). Cell lysates were also analyzed by Western Blotting to verify comparable Runx2 expression levels in different samples. Since all C-terminal deletions up to the C-terminal boundary of the RUNT domain retained binding to HIF-1α while the RUNT domain N-terminal deletion lost this binding, we conclude that the HIF-1α interacting domain is within the RUNT domain of Runx2.
Fig 5. Identification of the HIF-1α interacting region of Runx2.
The indicated FLAG-tagged C-terminal (panels A,B) and N-terminal (panels C,D) deletion mutants of Runx2 were constructed and transfected into HEK293 cells with wild type HIF-1α as described in Methods. Nuclear extracts were immunoprecipitated with anti-HIF-1α antibody and blots probed with HIF-1α or anti-FLAG antibody as indicated. Runx2 domain structure: QA, glutamine/alanine rich domain; RUNT, runt homology domain; NLS, nuclear localization signal; TAD, transactivation domains; RD, repression domain.
Runx2 and HIF-1a interact on specific sites in VEGF promoter
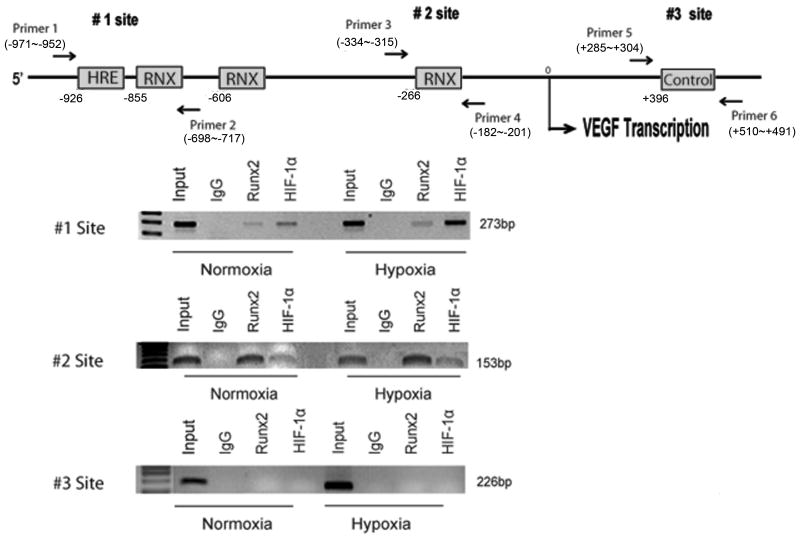
Figure 6A shows a schematic of the proximal murine Vegf-A promoter with the positions of consensus HIF-1α and Runx2 binding sites indicated. Shown are a previously characterized functional HRE at -926 bp from the transcription start site[Forsythe et al., 1996; Oosthuyse et al., 2001] and several putative Runx2 binding sites at -855, -606 and -266 bp[Peng et al., 2006]. Chromatin immunoprecipitation (ChIP) was used to examine Runx2 and HIF-1α interactions with this gene region in intact MC3T3-E1 cells. PCR primers were designed to detect immunoprecipitation of chromatin containing the 5′ HRE and adjoining Runx2 sites (site 1), the 3′ Runx2 site (site 2) as well as a control region in the transcribed portion of the Vegf-A gene that is free of either binding site (site 3). ChIP analysis with the site 1 and site 2 primer pairs detected positive signals with both Runx2 and HIF-1α antibodies while no signal was detected with isotype-matched IgG or with the control site 3 primers. As would be expected if HIF-1α were binding to the HRE at -926 bp, the site 1 primer pair detected a strong signal with HIF-1α antibody and this signal increased with hypoxia. In addition, a weaker, but still specific, signal was detected with the Runx2 antibody. Since the two 5′ consensus Runx2 sites are within 100 bp of the site 1 amplified region, it is not possible to discriminate between binding of Runx2 directly to DNA or to HIF-1α bound to its HRE. The site 2 primer pair detected a strong signal with Runx2 antibody that was not affected by oxygen status. This likely reflects binding of Runx2 to the site at -266 bp. Interestingly, a weaker signal was also detected with the HIF-1α antibody using this primer pair that was mildly increased by hypoxia. In this case, it is likely that HIF-1α associates with this chromatin region via interactions with Runx2 or some other protein rather than with an HRE. The nearest HRE at -926 bp is nearly 750 bp distant from the 3′ site 2 primer pair, too far away to be in the same amplified chromatin fragment as the Runx2 binding site at -266 bp. The observation that no Runx2 ChIP signal was detected with the site 3 primers is consistent with this interpretation since these primers amplify a DNA region that is only 470 bp away from the Runx2 site, yet failed to detect a Runx2 signal. These studies show that both HIF-1α and Runx2 are associated with the Vegf-A promoter in intact cells and provide evidence for both direct binding of each factor to its respective enhancer sequence on DNA as well as possible indirect binding of HIF-1α to Runx2-binding regions via protein-protein interactions.
Fig 6. Interaction of Runx2 and HIF-1α with VEGF chromatin.
(A) Structure of the proximal VEGF-A promoter region showing location of putative Runx2 and HIF-1α binding sites and associated PCR primer pairs. Site 1, putative Runx2 binding element; site 2, HIF-1α binding element; control site, region in the transcribed region of Vegf. (B) Chromatin immunoprecipitation. Chromatin was immunoprecipitated with the indicated antibodies and ChIP DNA analyzed with the site-specific PCR primers indicated.
Discussion
This study explores the relationship between HIF-1α and Runx2 in the control of VEGF gene expression and shows how canonical angiogenic and osteogenic transcription factors can function together to regulate an angiogenic signal. In contrast to studies that examined separate roles of HIF-1α and Runx2 in angiogenesis, the present work examined interactions between these two factors and found that HIF-1α and Runx2 physically and functionally interact to control VEGF synthesis. Although HIF-1α was able to respond to hypoxic stimuli in Runx2 null cells, the magnitude of VEGF induction in hypoxia was enhanced in the presence of Runx2. Of further interest, Runx2 strongly induced VEGF mRNA and protein expression even in normoxic conditions where HIF-1α protein levels are characteristically very low. Nevertheless, this induction required HIF-1α in that it could be strongly inhibited by HIF-1α siRNA or the HIF-1α inhibitor, echinomycin. Runx2 and HIF-1α were colocalized to discrete nuclear regions in target cells and shown to physically interact through sites in the Runx2 runt domain. ChIP analysis provided further evidence that Runx2 and HIF-1α interact on the chromatin of the Vegf-A gene.
The impetus for our work came from a number of previous studies suggesting a possible relationship between Runx2 and HIF-1α in angiogenesis and bone formation. To begin with, Runx2 and HIF-1α have similar patterns of expression during early skeletal development. In mice, mesenchymal condensations destined to form bone stain positive for HIF-1α beginning at E10.5. These regions are also hypoxic and exhibit HIF-1α-dependent transcriptional activity as detected by activation of an HRE-LacZ transgene[Provot et al., 2007]. Runx2 shares a similar time course and pattern of expression with HIF-1α, being localized to mesenchymal condensations as early as E9.5[Ducy et al., 1997; Kim et al., 1999]. Furthermore, both HIF-1α and Runx2 have clearly established roles in skeletal development and vascularization. Early conditional deletion of HIF-1α in limb bud mesenchyme using a Prx1-Cre resulted in major cartilage anomalies including defects in growth plate formation, delayed hypertrophy leading to limb shortening and joint abnormalities[Provot et al., 2007]. More selective, later deletion of HIF-1α in chondrocytes using a col II-Cre led to impaired chondrocyte survival in hypoxic areas and reduced VEGF expression[Schipani et al., 2001]. In addition, deletion of HIF-1α in osteoblasts resulted in reduced long bone cortical thickness and vascularity while activation of HIF-1α via osteoblast-specific deletion of VHL increased VEGF expression, bone formation and vascularity[Wang et al., 2007]. Similarly, Runx2 deletion was associated with defective VEGF synthesis and blockade of cartilage vascular invasion[Kim et al., 1999; Otto et al., 1997; Zelzer et al., 2001]. This vascularization defect reflects an intrinsic property of the bone rudiment rather than a defect in the surrounding vasculature since transplantation of Runx2-/- bones into a wild type host is unable to restore vascular invasion[Himeno et al., 2002]. In a related study, the human disorder, cleidocranial dysplasia, which is caused by Runx2 haploinsufficiency, was accompanied by diminished cartilage hypertrophy and a 5-fold decrease in VEGF expression[Zheng et al., 2005]. Lastly, a recent study showed that Runx2 and HIF-1α interact during trauma-induced heterotopic ossification[Lin et al., 2011]. Specifically, combined lentiviral siRNA suppression of HIF-1α and Runx2 inhibited bone formation to a greater degree than the knockdown of either factor.
Of particular interest, in the present study Runx2 stimulation of VEGF expression was observed in normoxic as well as hypoxic conditions and appeared to be dependent on HIF-1α (Figs 1-3). Effects on VEGF protein were likely due to increased Vegf-a transcription in that VEGF mRNA and protein levels generally increased in parallel. However, Runx2 stimulation of VEGF expression was not accompanied by a corresponding increase in HIF-1α mRNA. Instead, Runx2 preferentially increased HIF-1α protein levels, raising them from the low levels normally seen in high oxygen tension where HIF-1α is highly unstable due to oxygen-dependent hydroxylation and degradation. Possible explanations for this result are that Runx2 protects HIF-1α from proteosome-mediated degradation, increases HIF-1α translation or inhibits hydroxylation or VHL binding. Further studies will be required to discriminate between these possibilities. It should be pointed out that functions of HIF-1α in normoxia are not unique to our experimental system. A number of stimuli can activate HIF-1α in the presence of oxygen. These include IGF-1-dependent induction of HIF-1α and VEGF expression in human colon cancer cells[Fukuda et al., 2002], oncogenic ras up-regulation of HIF-1α and VEGF in tumor cells[Sodhi et al., 2001] and estrogen activation of HIF-1α and VEGF in the uterus[Kazi and Koos, 2007].
In addition to increasing HIF-1α protein levels, Runx2 can also increase HIF-1α-dependent transcriptional activity. This is particularly apparent in the hypoxic condition where Runx2 increased VEGF mRNA and protein without increasing HIF-1α levels. This stimulation is likely explained by interactions between HIF-1α and Runx2 on Vegf chromatin. ChIP analysis revealed that both factors bind the murine Vegf gene in intact cells. Strong HIF-1α binding was detected in the region of a well-characterized HRE at -926 bp. This site was previously shown to be essential for VEGF induction by HIF-1α in cell culture and in vivo[Forsythe et al., 1996; Oosthuyse et al., 2001]. Weaker Runx2 binding was also seen in this region either due to interactions with consensus Runx binding sites or via protein-protein interactions with HIF-1α. Stronger Runx2 binding was also detected in a more downstream portion of the promoter centered around a consensus Runx site at -266. Although the binding of Runx2 to this site has not previously been examined, the related Runt domain factor, Runx3, can bind similar sites in the human Vegf promoter. However, in this case, the effect of Runx3 was to inhibit VEGF expression[Peng et al., 2006]. Interestingly, this region also bound HIF-1α even though it does not contain any HRE consensus sites. Since the -926 bp HRE is too far away to be in this chromatin fragment, this result is likely explained by binding of HIF-1α to Runx2 or another as yet undefined protein in this region. These studies suggest that Runx2 can increase HIF-1α transcriptional activity via the formation of protein-protein interactions on chromatin.
The relative importance of Runx2 and HIF-1α to Vegf expression was assessed using a combination of siRNA knock down, genetic and chemical inhibition experiments (Figs 2 & 3). The overall conclusion of these studies was that Runx2 requires HIF-1α for activity while HIF-1α has basal activity in the absence of Runx2. This HIF-1α activity was apparent in the experiment with Runx2-null cells that retained a robust response to hypoxia. Nevertheless, Runx2 augmented HIF-1α activity in MC3T3-E1 cells since siRNA knock-down of Runx2 clearly reduced VEGF expression in normoxic and hypoxic conditions. Partial siRNA knockdown of HIF-1α also inhibited Runx2-dependent VEGF protein induction in both normoxic and hypoxic conditions. To more completely block HIF-1α activity, the chemical inhibitor, echinomycin, was used. This compound completely blocked both HIF-1α and Runx2-dependent induction of VEGF. Although echinomycin is reported to have off-target effects including the ability to interfere with myc and AP-1 transcription factors[Vlaminck et al., 2007], we do not think this explains our results. Specifically, the HRE-luc reporter used in the experiment shown in Fig. 3 is only activated by HIF-1α and does not contain binding sites for other transcription factors. Nevertheless, echinomycin completely blocked HIF-1α and Runx2-dependent induction of this reporter without affecting Runx2 induction of 6OSE2-luc. However, it is still possible that echinomycin interferes with other factors on the promoter to block Runx2 and HIF-1α induction of this gene. Further studies with HIF-1α-null cells will be required to definitively resolve whether Runx2 requires HIF-1α for its activity.
In further support of the concept that Runx2 and HIF-1α function together to regulate Vegf, we observed that these two factors could be colocalized to the nuclear compartment of MC3T3-E1 preosteoblast cells in both normoxic and hypoxic conditions. Furthermore, co-immunoprecipitation studies with C and N-terminal Runx2 deletion constructs revealed that Runx2 and HIF-1α physically interact using sequences in the runt domain of Runx2. This result is similar to a previous report with the related runt-domain family member, Runx1, which also interacts with HIF-1α through its runt domain[Peng et al., 2008]. Since there is 95 percent amino acid sequence conservation between the runt domains of Runx1 and Runx2, it is likely that a conserved sequence accounts for this interaction in both molecules. However, Runx1 inhibited the HIF-1α-dependent activation of the HRE-luciferase reporter while, in our studies, Runx2 stimulated this activity.
Although this study emphasized interactions between HIF-1α and Runx2 in bone and mesenchymal cell lines, our observations may also be relevant to other cell types such as vascular endothelial cells that express both these factors. Although not widely appreciated, Runx2 has clearly defined roles in normal and pathological vascular cell biology. Runx1 and Runx2 are both present in endothelial cells and vascular smooth muscle cells at sites of in vivo angiogenesis[Bronckers et al., 2005; Namba et al., 2000]. Furthermore, dominant-negative inhibition of Runx1 and 2 in murine and human endothelial cell lines inhibited proliferation, migration and tube formation[Namba et al., 2000; Qiao et al., 2006; Qiao et al., 2004]. It has also been proposed that the pericyte, a vascular smooth muscle-associated cell, can function as a mesenchymal stem cell/osteoprogenitor competent to differentiate into an osteogenic cell after Runx2 induction or activation (reviewed in [Towler, 2007]). Furthermore, pathologic vascular calcification is associated with up-regulation of Runx2 and bone marker genes[Byon et al., 2008]. Similarly, vascular endothelial cells are one of the classic HIF-1α targets that respond to hypoxia by expressing angiogenic factors including VEGF and by undergoing angiogenic processes including tubule/microvessel formation and extracellular matrix invasion[Manalo et al., 2005]. It is, therefore, highly likely that the Runx2-HIF-1α interaction we describe can also take place in these cells. Of further interest, HIF-1α and hypoxia have documented roles in the formation of atherosclerotic plaques that ultimately calcify through a process very similar to normal bone formation[Sluimer and Daemen, 2009].
In summary, we have shown that there is a physical and functional interaction between the major osteogenic factor, Runx2, and angiogenic master regulator, HIF-1α. This interaction stimulates VEGF gene expression in mesenchymal osteogenic cells by direct and indirect binding of these two factors to regulatory regions of the Vegf gene. These studies provide a mechanism for further clarifying the roles of Runx2 in angiogenesis and osteogenesis.
Acknowledgments
The authors wish to thank Drs. Zhaocheng Zhang and Jacques Nör for helpful discussions and use of a hypoxia chamber in the early stages of this study.
Grant support: Contract grant sponsor: NIH/NIDCR. Contract grant number: DE11723 and DE12211 (to RTF).
Contributor Information
Dr. Tae-Geon Kwon, Email: kwondk@knu.ac.kr.
Dr. Xiang Zhao, Email: xiangz@umich.edu.
Dr. Qian Yang, Email: Qian_Yang@hsdm.harvard.edu.
Dr. Yan Li, Email: yanlili@umich.edu.
Dr. Chunxi Ge, Email: chunxige@umich.edu.
Dr. Guisheng Zhao, Email: guisheng@umich.edu.
References
- Bae JS, Gutierrez S, Narla R, Pratap J, Devados R, van Wijnen AJ, Stein JL, Stein GS, Lian JB, Javed A. Reconstitution of Runx2/Cbfa1-null cells identifies a requirement for BMP2 signaling through a Runx2 functional domain during osteoblast differentiation. J Cell Biochem. 2007;100:434–49. doi: 10.1002/jcb.21039. [DOI] [PubMed] [Google Scholar]
- Bronckers AL, Sasaguri K, Cavender AC, D'Souza RN, Engelse MA. Expression of Runx2/Cbfa1/Pebp2alphaA during angiogenesis in postnatal rodent and fetal human orofacial tissues. J Bone Miner Res. 2005;20:428–37. doi: 10.1359/JBMR.041118. [DOI] [PubMed] [Google Scholar]
- Byon CH, Javed A, Dai Q, Kappes JC, Clemens TL, Darley-Usmar VM, McDonald JM, Chen Y. Oxidative stress induces vascular calcification through modulation of the osteogenic transcription factor Runx2 by AKT signaling. J Biol Chem. 2008;283:15319–27. doi: 10.1074/jbc.M800021200. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ducy P, Zhang R, Geoffroy V, Ridall AL, Karsenty G. Osf2/Cbfa1: a transcriptional activator of osteoblast differentiation. Cell. 1997;89:747–54. doi: 10.1016/s0092-8674(00)80257-3. [DOI] [PubMed] [Google Scholar]
- Firth JD, Ebert BL, Pugh CW, Ratcliffe PJ. Oxygen-regulated control elements in the phosphoglycerate kinase 1 and lactate dehydrogenase A genes: similarities with the erythropoietin 3′ enhancer. Proc Natl Acad Sci U S A. 1994;91:6496–500. doi: 10.1073/pnas.91.14.6496. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Forsythe JA, Jiang BH, Iyer NV, Agani F, Leung SW, Koos RD, Semenza GL. Activation of vascular endothelial growth factor gene transcription by hypoxia-inducible factor 1. Mol Cell Biol. 1996;16:4604–13. doi: 10.1128/mcb.16.9.4604. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fukuda R, Hirota K, Fan F, Jung YD, Ellis LM, Semenza GL. Insulin-like growth factor 1 induces hypoxia-inducible factor 1-mediated vascular endothelial growth factor expression, which is dependent on MAP kinase and phosphatidylinositol 3-kinase signaling in colon cancer cells. J Biol Chem. 2002;277:38205–11. doi: 10.1074/jbc.M203781200. [DOI] [PubMed] [Google Scholar]
- Himeno M, Enomoto H, Liu W, Ishizeki K, Nomura S, Kitamura Y, Komori T. Impaired vascular invasion of Cbfa1-deficient cartilage engrafted in the spleen. J Bone Miner Res. 2002;17:1297–305. doi: 10.1359/jbmr.2002.17.7.1297. [DOI] [PubMed] [Google Scholar]
- Hirota K, Semenza GL. Regulation of angiogenesis by hypoxia-inducible factor 1. Crit Rev Oncol Hematol. 2006;59:15–26. doi: 10.1016/j.critrevonc.2005.12.003. [DOI] [PubMed] [Google Scholar]
- Hofer T, Desbaillets I, Hopfl G, Gassmann M, Wenger RH. Dissecting hypoxia-dependent and hypoxia-independent steps in the HIF-1alpha activation cascade: implications for HIF-1alpha gene therapy. FASEB J. 2001;15:2715–7. doi: 10.1096/fj.01-0546fje. [DOI] [PubMed] [Google Scholar]
- Huang LE, Gu J, Schau M, Bunn HF. Regulation of hypoxia-inducible factor 1alpha is mediated by an O2-dependent degradation domain via the ubiquitin-proteasome pathway. Proc Natl Acad Sci U S A. 1998;95:7987–92. doi: 10.1073/pnas.95.14.7987. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Javed A, Barnes GL, Pratap J, Antkowiak T, Gerstenfeld LC, van Wijnen AJ, Stein JL, Lian JB, Stein GS. Impaired intranuclear trafficking of Runx2 (AML3/CBFA1) transcription factors in breast cancer cells inhibits osteolysis in vivo. Proc Natl Acad Sci U S A. 2005;102:1454–9. doi: 10.1073/pnas.0409121102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kazi AA, Koos RD. Estrogen-induced activation of hypoxia-inducible factor-1alpha, vascular endothelial growth factor expression, and edema in the uterus are mediated by the phosphatidylinositol 3-kinase/Akt pathway. Endocrinology. 2007;148:2363–74. doi: 10.1210/en.2006-1394. [DOI] [PubMed] [Google Scholar]
- Kim IS, Otto F, Zabel B, Mundlos S. Regulation of chondrocyte differentiation by Cbfa1. Mech Dev. 1999;80:159–70. doi: 10.1016/s0925-4773(98)00210-x. [DOI] [PubMed] [Google Scholar]
- Komori T, Yagi H, Nomura S, Yamaguchi A, Sasaki K, Deguchi K, Shimizu Y, Bronson RT, Gao YH, Inada M, Sato M, Okamoto R, Kitamura Y, Yoshiki S, Kishimoto T. Targeted disruption of Cbfa1 results in a complete lack of bone formation owing to maturational arrest of osteoblasts. Cell. 1997;89:755–64. doi: 10.1016/s0092-8674(00)80258-5. [DOI] [PubMed] [Google Scholar]
- Kong D, Park EJ, Stephen AG, Calvani M, Cardellina JH, Monks A, Fisher RJ, Shoemaker RH, Melillo G. Echinomycin, a small-molecule inhibitor of hypoxia-inducible factor-1 DNA-binding activity. Cancer Res. 2005;65:9047–55. doi: 10.1158/0008-5472.CAN-05-1235. [DOI] [PubMed] [Google Scholar]
- Li Y, Ge C, Franceschi RT. Differentiation-dependent association of phosphorylated extracellular signal-regulated kinase with the chromatin of osteoblast-related genes. J Bone Miner Res. 2010;25:154–63. doi: 10.1359/jbmr.090705. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lin L, Shen Q, Leng H, Duan X, Fu X, Yu C. Synergistic Inhibition of Endochondral Bone Formation by Silencing Hif1alpha and Runx2 in Trauma-induced Heterotopic Ossification. Mol Ther. 2011 doi: 10.1038/mt.2011.101. on line publication. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Manalo DJ, Rowan A, Lavoie T, Natarajan L, Kelly BD, Ye SQ, Garcia JG, Semenza GL. Transcriptional regulation of vascular endothelial cell responses to hypoxia by HIF-1. Blood. 2005;105:659–69. doi: 10.1182/blood-2004-07-2958. [DOI] [PubMed] [Google Scholar]
- Maxwell PH, Wiesener MS, Chang GW, Clifford SC, Vaux EC, Cockman ME, Wykoff CC, Pugh CW, Maher ER, Ratcliffe PJ. The tumour suppressor protein VHL targets hypoxia-inducible factors for oxygen-dependent proteolysis. Nature. 1999;399:271–5. doi: 10.1038/20459. [DOI] [PubMed] [Google Scholar]
- Namba K, Abe M, Saito S, Satake M, Ohmoto T, Watanabe T, Sato Y. Indispensable role of the transcription factor PEBP2/CBF in angiogenic activity of a murine endothelial cell MSS31. Oncogene. 2000;19:106–14. doi: 10.1038/sj.onc.1203257. [DOI] [PubMed] [Google Scholar]
- Oosthuyse B, Moons L, Storkebaum E, Beck H, Nuyens D, Brusselmans K, Van Dorpe J, Hellings P, Gorselink M, Heymans S, Theilmeier G, Dewerchin M, Laudenbach V, Vermylen P, Raat H, Acker T, Vleminckx V, Van Den Bosch L, Cashman N, Fujisawa H, Drost MR, Sciot R, Bruyninckx F, Hicklin DJ, Ince C, Gressens P, Lupu F, Plate KH, Robberecht W, Herbert JM, Collen D, Carmeliet P. Deletion of the hypoxia-response element in the vascular endothelial growth factor promoter causes motor neuron degeneration. Nature Genet. 2001;28:131–8. doi: 10.1038/88842. see comment. [DOI] [PubMed] [Google Scholar]
- Otto F, Thornell AP, Crompton T, Denzel A, Gilmour KC, Rosewell IR, Stamp GW, Beddington RS, Mundlos S, Olsen BR, Selby PB, Owen MJ. Cbfa1, a candidate gene for cleidocranial dysplasia syndrome, is essential for osteoblast differentiation and bone development. Cell. 1997;89:765–71. doi: 10.1016/s0092-8674(00)80259-7. [DOI] [PubMed] [Google Scholar]
- Peng Z, Wei D, Wang L, Tang H, Zhang J, Le X, Jia Z, Li Q, Xie K. RUNX3 inhibits the expression of vascular endothelial growth factor and reduces the angiogenesis, growth, and metastasis of human gastric cancer. Clin Cancer Res. 2006;12:6386–94. doi: 10.1158/1078-0432.CCR-05-2359. [DOI] [PubMed] [Google Scholar]
- Peng ZG, Zhou MY, Huang Y, Qiu JH, Wang LS, Liao SH, Dong S, Chen GQ. Physical and functional interaction of Runt-related protein 1 with hypoxia-inducible factor-1alpha. Oncogene. 2008;27:839–47. doi: 10.1038/sj.onc.1210676. [DOI] [PubMed] [Google Scholar]
- Provot S, Zinyk D, Gunes Y, Kathri R, Le Q, Kronenberg HM, Johnson RS, Longaker MT, Giaccia AJ, Schipani E. Hif-1alpha regulates differentiation of limb bud mesenchyme and joint development. J Cell Biol. 2007;177:451–64. doi: 10.1083/jcb.200612023. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Qiao M, Shapiro P, Fosbrink M, Rus H, Kumar R, Passaniti A. Cell cycle-dependent phosphorylation of the RUNX2 transcription factor by cdc2 regulates endothelial cell proliferation. J Biol Chem. 2006;281:7118–28. doi: 10.1074/jbc.M508162200. [DOI] [PubMed] [Google Scholar]
- Qiao M, Shapiro P, Kumar R, Passaniti A. Insulin-like growth factor-1 regulates endogenous RUNX2 activity in endothelial cells through a phosphatidylinositol 3-kinase/ERK-dependent and Akt-independent signaling pathway. J Biol Chem. 2004;279:42709–18. doi: 10.1074/jbc.M404480200. [DOI] [PubMed] [Google Scholar]
- Roca H, Franceschi RT. Analysis of transcription factor interactions in osteoblasts using competitive chromatin immunoprecipitation. Nucleic Acids Res. 2008;36:1723–30. doi: 10.1093/nar/gkn022. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Roca H, Phimphilai M, Gopalakrishnan R, Xiao G, Franceschi RT. Cooperative interactions between RUNX2 and homeodomain protein-binding sites are critical for the osteoblast-specific expression of the bone sialoprotein gene. J Biol Chem. 2005;280:30845–55. doi: 10.1074/jbc.M503942200. [DOI] [PubMed] [Google Scholar]
- Salceda S, Caro J. Hypoxia-inducible factor 1alpha (HIF-1alpha) protein is rapidly degraded by the ubiquitin-proteasome system under normoxic conditions. Its stabilization by hypoxia depends on redox-induced changes. J Biol Chem. 1997;272:22642–7. doi: 10.1074/jbc.272.36.22642. [DOI] [PubMed] [Google Scholar]
- Schaffer L, Scheid A, Spielmann P, Breymann C, Zimmermann R, Meuli M, Gassmann M, Marti HH, Wenger RH. Oxygen-regulated expression of TGF-beta 3, a growth factor involved in trophoblast differentiation. Placenta. 2003;24:941–50. doi: 10.1016/s0143-4004(03)00166-8. [DOI] [PubMed] [Google Scholar]
- Schipani E, Ryan HE, Didrickson S, Kobayashi T, Knight M, Johnson RS. Hypoxia in cartilage: HIF-1alpha is essential for chondrocyte growth arrest and survival. Genes Dev. 2001;15:2865–76. doi: 10.1101/gad.934301. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Semenza GL, Jiang BH, Leung SW, Passantino R, Concordet JP, Maire P, Giallongo A. Hypoxia response elements in the aldolase A, enolase 1, and lactate dehydrogenase A gene promoters contain essential binding sites for hypoxia-inducible factor 1. J Biol Chem. 1996;271:32529–37. doi: 10.1074/jbc.271.51.32529. [DOI] [PubMed] [Google Scholar]
- Sluimer JC, Daemen MJ. Novel concepts in atherogenesis: angiogenesis and hypoxia in atherosclerosis. J Pathol. 2009;218:7–29. doi: 10.1002/path.2518. [DOI] [PubMed] [Google Scholar]
- Sodhi A, Montaner S, Miyazaki H, Gutkind JS. MAPK and Akt act cooperatively but independently on hypoxia inducible factor-1alpha in rasV12 upregulation of VEGF. Biochem Biophys Res Comm. 2001;287:292–300. doi: 10.1006/bbrc.2001.5532. [DOI] [PubMed] [Google Scholar]
- Sun L, Vitolo M, Passaniti A. Runt-related gene 2 in endothelial cells: inducible expression and specific regulation of cell migration and invasion. Cancer Res. 2001;61:4994–5001. [PubMed] [Google Scholar]
- Thirunavukkarasu K, Mahajan M, McLarren KW, Stifani S, Karsenty G. Two domains unique to osteoblast-specific transcription factor Osf2/Cbfa1 contribute to its transactivation function and its inability to heterodimerize with Cbfbeta. Mol Cell Biol. 1998;18:4197–208. doi: 10.1128/mcb.18.7.4197. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Towler DA. Vascular biology and bone formation: hints from HIF. J Clin Invest. 2007;117:1477–80. doi: 10.1172/JCI32518. comment. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vlaminck B, Toffoli S, Ghislain B, Demazy C, Raes M, Michiels C. Dual effect of echinomycin on hypoxia-inducible factor-1 activity under normoxic and hypoxic conditions. FEBS J. 2007;274:5533–42. doi: 10.1111/j.1742-4658.2007.06072.x. [DOI] [PubMed] [Google Scholar]
- Wang D, Christensen K, Chawla K, Xiao G, Krebsbach PH, Franceschi RT. Isolation and characterization of MC3T3-E1 preosteoblast subclones with distinct in vitro and in vivo differentiation/mineralization potential. J Bone Miner Res. 1999;14:893–903. doi: 10.1359/jbmr.1999.14.6.893. [DOI] [PubMed] [Google Scholar]
- Wang Y, Wan C, Deng L, Liu X, Cao X, Gilbert SR, Bouxsein ML, Faugere MC, Guldberg RE, Gerstenfeld LC, Haase VH, Johnson RS, Schipani E, Clemens TL. The hypoxia-inducible factor alpha pathway couples angiogenesis to osteogenesis during skeletal development. J Clin Invest. 2007;117:1616–26. doi: 10.1172/JCI31581. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Xiao G, Cui Y, Ducy P, Karsenty G, Franceschi RT. Ascorbic acid-dependent activation of the osteocalcin promoter in MC3T3-E1 preosteoblasts: requirement for collagen matrix synthesis and the presence of an intact OSE2 sequence. Mol Endocrinol. 1997;11:1103–13. doi: 10.1210/mend.11.8.9955. [DOI] [PubMed] [Google Scholar]
- Yang S, Wei D, Wang D, Phimphilai M, Krebsbach PH, Franceschi RT. In vitro and in vivo synergistic interactions between the Runx2/Cbfa1 transcription factor and bone morphogenetic protein-2 in stimulating osteoblast differentiation. J Bone Miner Res. 2003;18:705–15. doi: 10.1359/jbmr.2003.18.4.705. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zelzer E, Glotzer DJ, Hartmann C, Thomas D, Fukai N, Soker S, Olsen BR. Tissue specific regulation of VEGF expression during bone development requires Cbfa1/Runx2. Mech Dev. 2001;106:97–106. doi: 10.1016/s0925-4773(01)00428-2. [DOI] [PubMed] [Google Scholar]
- Zhao Z, Zhao M, Xiao G, Franceschi RT. Gene transfer of the Runx2 transcription factor enhances osteogenic activity of bone marrow stromal cells in vitro and in vivo. Mol Ther. 2005;12:247–53. doi: 10.1016/j.ymthe.2005.03.009. [DOI] [PubMed] [Google Scholar]
- Zheng Q, Sebald E, Zhou G, Chen Y, Wilcox W, Lee B, Krakow D. Dysregulation of chondrogenesis in human cleidocranial dysplasia. Am J Hum Genet. 2005;77:305–12. doi: 10.1086/432261. [DOI] [PMC free article] [PubMed] [Google Scholar]