Abstract
Large-scale projects are providing rapid global access to a wealth of mouse genetic resources to help discover disease genes and to manipulate their function.
It is a little known fact that Gregor Mendel, remembered for his studies of trait inheritance in pea plants, also experimented with breeding mice to understand coat color traits. Had it not been for the disapproval of Bishop Anton Ernst Schaffgotsch, who led the Augustinian monastery where Mendel studied, he might well have been credited as the father of mouse genetics [1]. Instead, CC Little started generating inbred lines of mice half a century later, driven by a desire to understand cancer biology and recognizing the importance of reproducible genetic crosses [1]. From these beginnings more than 300 strains of laboratory mice have been developed; each line has been faithfully replicated and cryo-preserved, making them a renewable genetic resource. Most are the result of the blending together of Mus musculus sub-species, including domesticus and musculus, with some contribution of castaneous and molossinus, resulting in a distinctive genetic mosaic of these progenitors in each inbred line [2].
Today's geneticists usually turn to one of these inbred mouse strains when attempting to model human disease because mice offer advantages that few species can match. Importantly, the mouse genome can be easily manipulated with greater speed, scale and sophistication than that of other mammals, and the efforts of the International Mouse Genome Sequencing Consortium has resulted in a high quality reference genome sequence that is the envy of other model organism users [3]. The future for mouse genetics promises to be even more exciting now that high-throughput sequencing of mouse strain genomes has started, and efforts are under way to systematically disrupt every gene in the mouse genome and phenotype the resulting mutant animals [4]. Here, we outline the tools and technologies that have emerged for using mice to discover and characterize disease genes, and the resources that are being developed to accelerate these discoveries.
Sequencing mouse genomes
In 2002 the International Mouse Genome Sequencing Consortium released the first draft of the genome from C57BL/6J, an inbred strain of the laboratory mouse [3], and a finished genome was released in 2009 [5]. As one of the most globally used lines, C57BL/6J was a wise choice for the reference mouse strain, but it is by no means the only strain used in research. Therefore, subsequent efforts were initiated to generate genomic sequence of other inbred strains. Firstly, four different strains of the laboratory mouse were included by Celera in a whole-genomic shotgun sequencing project: A/J, DBA/2J, 129X1/SvJ and 129S1/SvImJ [6]. This resulted in 27.4 million sequencing reads, giving a total of 5.3x coverage of the mouse genome. Secondly, more than 150,000 short insert clones were sequenced from the 129S5SvEvBrd strain covering 4.7% of the reference genome [7]. Thirdly, Perlegen Sciences used hybridization to re-sequence 15 inbred mouse strains [8]; this set included 11 classical strains and four strains derived from the wild. Unlike the other resources, Perlegen's approach did not generate sequence reads, and their hybridization sequencing technology queried only 1.49 Gigabases of the reference genome (equivalent to about 58% of the C57BL/6J sequence that is non-repetitive). Furthermore, to generate high accuracy calls, high stringency cutoffs were used, resulting in a false negative rate estimated to be as high as 50% [2]. Therefore, available sequence data lacked the coverage and breadth of strains to make it a widely used resource.
The first non-reference mouse chromosomes to be sequenced were A/J and CAST/EiJ chromosome 17, revealing significant variation at the nucleotide level and also considerable structural differences [9]. Building on that work, we commenced the Mouse Genomes Project, which has sequenced the genomes of 17 key mouse strains using next-generation sequencing on the Illumina platform (Box 1). At the last data freeze in December 2009 an average of 25x sequence coverage of each strain had been generated, and a deep catalog of variants [10]. These data provide a comprehensive insight into the genomes of the 17 strains, allowing immediate access to background genetic information for most mouse models of disease in addition to facilitating the analysis of the molecular basis of complex traits with unparalleled resolution.
Genetic manipulation of mice in the post-genomic era
Technologies for modifying the mouse genome can be split into two broad classes: those for gene-driven analyses and those for random mutagenesis.
The collection and propagation of mice harboring spontaneous mutations with striking phenotypes, such as the obese mouse, served mouse geneticists well for most of the 20th century. When it became clear that the rate of random germline mutation can be significantly increased by exposure to radiation or to chemical mutagens such as N-ethyl-N-nitrosourea (ENU) [11], large-scale mutagenesis programs followed, resulting in an explosion in the number of mutant lines. Phenotypic screens of these lines led to the identification of many hundreds of new mutations and candidate disease genes [12,13]. One notable example of a successful forward genetic screen, reviewed in [14], identified 89 ENU-induced mutants that influence the immune system, of which at least 69 have now been characterized at the molecular level. However, mapping random mutations and identifying the affected gene can be an arduous process, often taking years; therefore, causal mutations for only a fraction of mutant lines have been identified thus far. Screening DNA from archived mutagenized lines for mutations in a specific gene of interest is a parallel 'gene driven' strategy that has proven successful [15,16]. With the advent of new sequencing technologies it is now cost effective to sequence mutagenized mouse exomes in their entirety, enabling the rapid identification of candidate disease genes from existing resources and meaning that mutagenesis-driven approaches may return as a powerful tool for studying disease genes. Other methods of random mutagenesis include retroviruses, transposons (reviewed in [17]), and 'gene traps' [18]. These DNA-based mutagens can be easily mapped using approaches such as splinkerette PCR [19] and are discussed in more detail below.
The genome of the mouse can also be manipulated by pronuclear injection of DNA into oocytes or by modification of embryonic stem (ES) cells, which can then be injected into blastocysts to make chimeras, allowing modified alleles to be transmitted through the germline. Direct pronuclear injection results in random integration of the injected DNA [20]; consequently, transgene copy numbers and integration sites differ between lines, potentially resulting in very different phenotypes. Large genomic fragments such as bacterial artificial chromosomes (BACs) may also be injected (reviewed in [21]); these have proven particularly useful in complementation studies or 'rescue' experiments for identifying genes contributing to a genetically mapped disease trait of interest [22]. By contrast, DNA introduced into ES cells in culture can undergo site-specific, homology-directed recombination [23], thus enabling the generation of targeted gain- and loss-of-function alleles as well as the engineering of large-scale rearrangements of entire mouse chromosomes (Figure 1) [24,25]. Other recently developed techniques include transgenic small hairpin RNAs (shRNAs), which are often delivered by lentiviral transgenesis [26,27], single-stranded oligonucleotides (ssODNs; reviewed in [28]), and zinc-finger nucleases (ZFNs) [29], which can be used to generate subtle sequence-specific genomic modifications. Here we will address in more detail a few of these technologies, focusing on recent advances uniquely available to mouse geneticists.
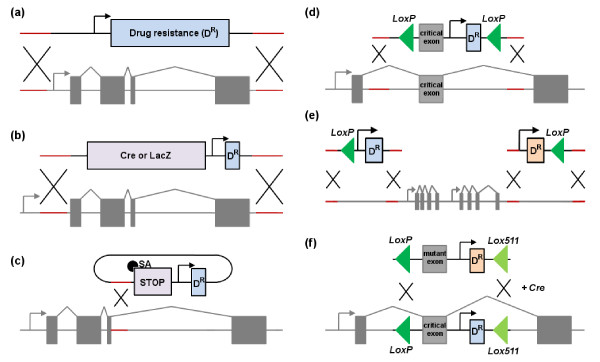
Figure 1.
Gene targeting strategies used in mouse ES cells. Targeting is achieved by recombination (black crosses) between homology arms (red lines). (a) A knockout vector replaces an entire gene with a selection cassette containing drug resistance (DR), enabling the selection of successfully targeted ES cell clones. (b) A knock-in vector allows the expression of a transgene, such as LacZ or Cre, by the promoter (gray arrow) of the targeted gene. (c) Insertion vectors can interfere with splicing by disrupting a target gene by the introduction of an exon with an early termination codon or a 5' splice acceptor site (SA). They typically target the genome with a single crossover event. (d) A conditional allele with directional DNA sequences (LoxP, green triangles) either side of a critical exon. Recombination between the sites will result in a null allele. (e) LoxP sites can also be targeted megabases apart, either side of a larger cluster of genes, enabling chromosome engineering. (f) Heterospecific Lox sites, such as LoxP and Lox511, are targeted by the site-specific recombinase Cre. Recombinase-mediated cassette exchange (RMCE) enables the efficient swapping of one targeted cassette containing incompatible target sites for another cassette flanked by an identical pair of sites. This enables the rapid generation of new alleles, such as introducing a point mutation in a critical exon.
ES cell gene-targeting
ES cell technology has been a profound advance in mouse genetics (detailed in [30]). Historically, the majority of manipulations have been performed in ES cells derived from 129 sub-strains (Table 1). Recently, robust and highly germline-competent ES cells derived from the popular C57BL/6 strains have been developed, such as JM8 and C2 (Table 1). To assist in tracking the contribution of these ES cells to chimerism, and to identify mice that have transmitted their genome through the germline, a dominant Agouti (yellow) coat color allele was engineered in JM8 cells [31]. This now enables the study of mutant alleles on a common, controlled genetic background without the need for generations of backcrossing.
Table 1.
Commonly used ES cell lines for generating genetically modified mice
| ES cell line | Genetic background | Comments | Reference |
|---|---|---|---|
| E14TG2a | 129P2OlaHsd | Feeder-independent; suitable for injections into C57BL/6 blastocysts | [104] |
| AB2.2 | 129S7/SvEvBrd-Hprtb-m2 | Feeder-dependent | [105] |
| J1 | 129SvJae | Feeder-dependent | [106] |
| Bruce4 | C57BL/6J-Thy1.1 | Have a tendency to aneuploidy | [107] |
| B6/Blu-1 | C57BL/6N | Generated by Tim Ley (Washington University, St Louis, USA) | Personal communication |
| JM8.parental | C57BL/6N | 76% GLT rate | [31] |
| JM8.F6 | C57BL/6N | Feeder-dependent JM8 subline. Suitable for injections in BALB/c or C57BL/6J-Tyrc/c blastocysts | [31] |
| JM8.N4 | C57BL/6N | Feeder-independent JM8 subline. Suitable for injections in BALB/c or C57BL/6J-Tyrc/c blastocysts | [31] |
| JM8A1.N3 | C57BL/6N | JM8.F6-derived line with a repaired Agouti locus. Feeder independent | [31] |
| C2 | C57BL/6NTac | Efficient GLT using a combination of ICR morula aggregation with outbred host embryos | [108] |
| R1 | 129X1/SvJ × 129S1 (hybrid) | Feeder-dependent | [109] |
| G4 | 129S6/SvEvTac × C57BL/6Ncr F1 hybrid | Feeder-dependent; typically used for tetraploid complementation assays | [75] |
Abbreviations: GLT, germ line transmission; ICR, Institute for Cancer Research.
Gene targeting in mouse ES cells can be achieved by homologous recombination, using replacement, insertion or knock-in vectors, all of which contain a region of homology with the locus to be targeted. In replacement vectors, crucial exons (or entire genes) are replaced by a selection cassette to generate a null knockout allele (Figure 1a). Knock-in vectors are designed such that a transgene or reporter is transcriptionally regulated by the endogenous promoter of the locus (Figure 1b; reviewed in [32]). By contrast, insertion vectors rely on gene rearrangement by interfering with splicing to disrupt a target gene (Figure 1c). Significant resources are available for obtaining suitable genomic DNA for targeting vector construction, including genome-wide end-sequenced BAC libraries for C57BL/6J-derived [33] and 129-derived strains [7]. Homology arms (the part of the vector that aligns with the genome to facilitate recombination) were typically generated by restriction digest of large DNA fragments or by PCR amplification, but increasingly 'recombineering' technologies are being used [34], which make it possible to engineer virtually any mutation into the mouse genome with base pair resolution. In addition, customized targeting vectors can be generated on a contract basis by several companies.
Gene modification with conditions
Conditional gene modification is used to enable spatial and/or temporal control over the modification of the gene of interest. To this end, site-specific recombinase (SSR) systems are used, including Cre-LoxP, Flp-FRT, φC31 integrase-attB/attP and most recently Dre-rox [35]. For a comprehensive review of the use of site-specific recombinases for manipulation of the mouse genome, see [36]. The DNA sequences that the SSRs recognize are typically directional and can either flank the target DNA for excision from the genome or be used to invert segments of DNA. SSRs can be used for the generation of single gene knockouts or rearrangements, and for chromosome engineering on a megabase scale (Figure 1d,e) [37,38].
SSRs can be expressed from endogenous promoters (as shown in Figure 1b) and in a tissue- or cell-specific manner. This is particularly useful when studying the organ-specific function of genes that are widely expressed and essential for embryonic development. For example, a conditional allele of Sox9, a gene implicated in campomelic dysplasia in humans, is necessary to study its function in cartilage in mice because germline deletion of Sox9 results in perinatal lethality [39]. For somatic mutagenesis, inducible gene-modification systems may be used. These systems allow temporal 'inducible' control of SSR expression. There are several inducible expression systems available, including tetracycline [40], LacZ [41], and the tamoxifen-inducible systems [42]. These systems have been invaluable in studying genes and neural circuits involved in learning and memory, by turning genes and cellular markers 'on' or 'off' during controlled time periods (reviewed in [43]), and in a range of other biological systems.
There are now over 500 tissue- or cell-specific Cre recombinase mice (some of which are inducible) documented in databases such as Cre-Zoo and Cre-X-mice (Table 2) [44]. However, as conditional modification technologies become increasingly sophisticated, the potential for non-specific effects, from mis-regulation of the targeted gene to incomplete recombination by the SSR, must remain a consideration [45,46]. For example, a recent study highlighted the potential for protein expression from episomal products of Cre recombinase-excised genes, particularly when deletion occurs in cells that have a low population turnover [47].
Table 2.
Resources generated from large-scale mouse genetics projects
| Resource | Web site | Description | Reference |
|---|---|---|---|
| Australian Phenomics Network | http://www.australianphenomics.org.au | Provides resources and services for producing, screening and archiving mutant mice | |
| Canadian Mouse Mutant Repository | http://www.cmmr.ca/ | Archived ES cells, sperm, ova, embryos, and DNA | |
| Cancer Models Database | http://cancermodels.nci.nih.gov | A database of mouse lines that model the genesis, progression or clinical course of human cancers | |
| Center for Animal Resources and Development | http://cardb.cc.kumamoto-u.ac.jp/transgenic | A repository of over 1,300 mutant lines | |
| Centre for Modeling Human Disease | http://www.cmhd.ca/databases | ENU mutagenesis and gene trap databases | |
| Collaborative Cross | http://mouse.ornl.gov/projects/collabcross.html | A resource for the genetic analysis of complex traits | [110] |
| Cre-X-mice | http://nagy.mshri.on.ca | A Cre-expressing transgenic mouse line database | [44] |
| CreZoo | http://bioit.fleming.gr/crezoo | A Cre-expressing transgenic mouse line database | |
| Ensembl | http://www.ensembl.org | A genome database for mouse and other eukaryotes | [111] |
| European Conditional Mouse Mutagenesis Program | http://www.eucomm.org | Aims to generate and distribute a collection of 13,000 mutated ES cell lines using conditional approaches | [112] |
| European Mouse Disease Clinic | http://www.eumodic.org | Aims to generate phenome data on 650 knockout mice generated by the EUCOMM project | [82] |
| European Mouse Mutant Archive | http://www.emmanet.org | A European repository with over 1,700 mutant strains | [113] |
| European Mouse Phenotyping Resource for Standardized Screens (EMPReSS) | http://empress.har.mrc.ac.uk | A primary screening platform with over 100 standard operating procedures validated on inbred strains | [81] |
| European Union Mouse Research for Public Health and Industrial Applications | http://www.eumorphia.org | Novel approaches in phenotyping, mutagenesis and informatics to improve the characterization of mouse models | [114] |
| EuroPhenome | http://www.europhenome.org | A database to hold phenome data from EMPReSS | [115] |
| Federation of International Mouse Resources | http://www.fimre.org | Coordinates repositories and resource centers globally | [116] |
| GenomeSpace | http://www.genomespace.org | A central workspace for genomics tools, including Galaxy, Integrative Genomics Viewer and UCSC Browser | |
| Heterogeneous Stock Phenotyping Project | http://mus.well.ox.ac.uk/mouse/HS | A searchable map of QTLs that contribute to variation in over 100 complex traits, using Heterogeneous Stock mice | [88] |
| International Gene Trap Consortium | http://www.genetrap.org | Information on >380,000 gene-trapped ES cell lines | [117] |
| International Knockout Mouse Consortium | http://www.komp.org/ikmc | Aims to minimize overlap, share resources, and improve services among the three major knockout projects | [79] |
| Knockout Mouse Project | http://www.knockoutmouse.org | Aims to target 8,500 genes and make mice available to the community | [4] |
| MouseBook | http://www.mousebook.org | MRC Harwell's mouse resources; includes a frozen embryo and sperm archive, an ENU screen and DNA archive, and standardized phenotyping procedures | [118] |
| Mouse Genome Database | http://www.informatics.jax.org | Provides integrated genetic, genomic, and biological data on laboratory mouse strains | [119] |
| Mouse Genomes Project | http://www.sanger.ac.uk/resources | Raw sequence data, SNPs and assemblies of 17 mouse genomes | |
| Mouse Phenome Project | http://phenome.jax.org | A collection of baseline phenotypic data on commonly used inbred mouse strains | [83] |
| Mouse Resources Portal | http://www.sanger.ac.uk/mouseportal | The Sanger Institute's resources; includes available BACs, gene targeting vectors, ES cells and mutant mouse lines with associated phenotypic data | [10] |
| Mutant Mouse Regional Resource Centers | http://www.mmrrc.org | A repository of mouse stocks and ES cell lines | |
| North American Conditional Mouse Mutagenesis project | http://www.norcomm.org | Aims to target >2,000 genes that have not been previously targeted or trapped | [4] |
| PB Mutagenesis Information Center | http://www.idmshanghai.cn/PBmice | A database for storing, retrieving and displaying information derived from PB transposon insertions | [120] |
| RIKEN Bioresource Center | http://www.brc.riken.jp/lab/animal/en | A Japanese repository of live and archived lines |
Recombinase-mediated cassette exchange
Using homologous recombination to introduce genetic material into a desired genetic location in the mouse genome is not always straightforward. The efficiency is often dependent on the nature of the genomic target site and on the design of the targeting vector. Therefore, the ability to efficiently introduce secondary modifications to already successfully targeted cassettes is advantageous. Recombinase-mediated cassette exchange (RMCE) is a process in which site-specific recombinases exchange one gene cassette, flanked by a pair of incompatible target sites, for another cassette flanked by an identical pair of sites (Figure 1f) [48]. Apart from the naturally occurring heterotypic SSR sites (attB and attP for φC31), several variant sites have been developed for Cre and Flp, providing the required heterospecificity crucial for RMCE (for example, LoxP/Lox511 and FRT/FRT3; see [49] for a complete list). In RMCE, typically one cassette is present in the host genome, whereas the other cassette (and the recombinase) is introduced into the host ES cell by electroporation, chemical-mediated or adenoviral-mediated gene transfer [50]. Transient expression of the recombinase will direct integration of the SSR site-flanked cassette, which can then be selected by drug resistance. RMCE-based techniques are proving to be useful in the rapid production of custom allelic series [51]: they have recently been used to compare the impact of different tumor-associated mutations in p53 [52], and to study the effect of multiple enhancer elements on the expression of a targeted cassette [53].
Transposons for mutagenesis
Unlike most of the methods described so far, which allow manipulation of the genome with base pair precision, transposable elements provide the power to molecularly tag, and therefore rapidly map, random mutagenic events. The application of transposons to the field of mouse genetics has become possible only in the past decade. So far, four distinct DNA transposons have been shown to function in mice: Tol2, Minos, Sleeping Beauty (SB) and PiggyBac (PB) (reviewed in [17]), with the latter two being the most widely used. DNA transposons use a 'cut-and-paste' transposition mechanism. When both the transposase enzyme and a transposon vector are present in the same nucleus, the transposase can mediate excision of the transposon from the donor site and integration into another target site in the host cell genome. RNA-mediated transposition, driven by a 'copy-and-paste' mechanism, has also been introduced into mice for mutagenesis [54].
Transposons can be used for germline mutagenesis in mice (reviewed in [55]). However, this technique is inefficient for genome-wide forward genetic screens, owing to the low rate of transposition (one to three de novo insertions per gamete) and the tendency for local hopping exhibited by most of the transposons; though some researchers have taken advantage of this observation to saturate smaller genomic regions [56,57]. So far, the most common use for transposons has been in the field of cancer genetics [17]. Retroviral insertional mutagenesis has traditionally been used to study the genetics of hematopoietic and mammary cancers (Box 2), but the study of other tumor types has been limited by viral tropism. Initial studies demonstrating the validity of transposon-mediated insertional mutagenesis (using SB) identified both known and novel cancer genes involved in sarcoma and lymphoma [58,59]. Since then, transposons have been engineered to produce gain-of-function mutations in epithelial cells resulting in the development of a wide variety of carcinomas [60]. In addition, Cre-inducible SB transposase alleles can restrict mutagenesis to specific tissues, permitting studies into colorectal cancer and hepatocellular carcinoma [61,62]. More recently, PB has been used for somatic mutagenesis, representing another tool for cancer gene discovery in the mouse [63].
Transposons can also be used to generate transgenic mice by loading them with genetic cargo. SB, PB and Tol2 are all efficient in delivering large transgenes, up to 70 kb in size [64]. PB has also been used together with SSR technology to generate large-scale rearrangements of the mouse genome, including duplications, deletions, and translocations [65]. Recently, transposons have been used to deliver the reprogramming factors required for generating induced pluripotent stem (iPS) cells [66,67].
Gene trap mutagenesis
Gene trapping in mouse ES cells is an efficient method for mutagenesis of the mammalian genome. Insertion of a gene trap vector can disrupt gene function and/or report gene expression, and because these vectors integrate into the genome they provide a convenient tag that facilitates the identification of their insertion site. A typical strategy involves electroporating into ES cells a vector containing a 5' splice acceptor that splices to the upstream exon of the trapped gene, and thus the endogenous promoter of the trapped gene is used to drive the expression of the reporter gene [18]. However, the vector can also be delivered by retroviral infection, or transposon-mediated insertion and identification of the trap insertion sites in the resultant ES cell clones performed by splinkerette PCR (detailed in [68]).
Recent developments in trapping technology involve the use of 'conditional traps', which enable the induced modification of trap alleles, in vitro or in vivo, using SSRs, and using RMCE to exchange trapped vectors with other functional cassettes [69]. Gene trapping strategies have also been successfully developed to screen for genes that have specific expression patterns ('enhancer traps' [70]) or are acting in specific biological pathways ('induction trapping' [71,72]). Another approach to direct gene trapping toward genes in a specific pathway is to perform a phenotypic screen in ES cells. However, most insertions will cause heterozygous mutations (which will generate detectable phenotypes only for haploinsufficient genes). One strategy to overcome this has been to use ES cells that have a deficiency in the Bloom (Blm) DNA helicase. These cells show high levels of mitotic recombination, which facilitates the generation of homozygosity in cell lines from colonies carrying heterozygous mutations [73].
'ES cell-driven' mouse production
Another advantage that the mouse has over other model organisms is in the rapid generation of mutant mice using ES cell-driven approaches. These enable the production of mice that are entirely, or almost entirely, derived from ES cells without the requirement for germline transmission. These approaches involve the injection of ES cells into eight cell embryos or a process called tetraploid complementation and allow the generation of mutant mice in weeks rather than months [74,75]. By combining these approaches with shRNA-mediated knockdown, several groups have shown that it is possible to rapidly generate knockdown mice for the analysis of somatic gene function [76,77]. Mice somatically overexpressing genes in an inducible and regulated way have also been developed using these approaches [78].
Mouse genetics on a grand scale
The success of the genome sequencing consortia over the past two decades established a model for further large-scale, collaborative projects aimed at functionally characterizing genomes (Table 2). Examples include the International Knockout Mouse Consortium (IKMC), and its constituent regional projects, which collectively aim to generate mutant alleles for every protein-coding gene in the mouse genome and to make the resources available to the scientific community [4]. Researchers can now search the IKMC website and acquire, at minimal cost, mice or ES cells that lack a gene of interest [79], thereby accelerating the path from a gene of interest to mutant mouse line. By May 2011, IKMC had over 16,000 ES cell lines with mutations in protein coding genes. Many of these alleles are 'knockout first' alleles, which are designed to introduce a LacZ expression marker into a target gene, and the allele can be tailored by using Cre and Flp to generate null and conditional alleles, respectively [80].
In parallel, a number of past and ongoing standardized phenotyping projects have documented traits in inbred strains and mutant lines for phenotypes relevant to human disease, including the Mouse Phenome Project, the European Mouse Disease Clinic (EUMODIC) and the Mouse Genetics Project (MGP; based at the Wellcome Trust Sanger Institute); see also Table 2[81-84]. The results from many screens are made available online, enabling researchers to identify potentially interesting phenotypes for detailed analysis (Figure 2). For example, primary MGP analysis of mice lacking the gene Slx4 identified a number of developmental and DNA instability phenotypes. Detailed secondary analysis revealed the mouse to phenocopy a new sub-type of the human genetic illness, Fanconi anemia [85-87].
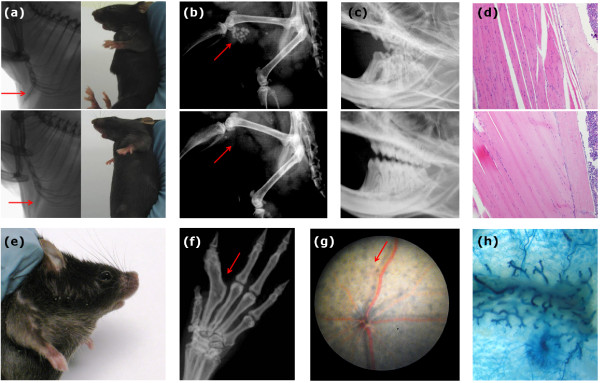
Figure 2.
Phenotypic screening of genetically modified mice. Mutant mice available to the community are systematically screened for a range of phenotypes by the Sanger Mouse Genetics Project and the data published online [10]. Some examples of observed phenotypes are shown. (a) An outwardly protruding xiphoid process in Fam73btm1a(KOMP)Wtsi homozygous mice (top), compared with wild-type controls (bottom). (b) The formation of uroliths (bladder stones) in Clnd16tm1a(KOMP)Wtsi homozygous mice (top), not present in controls (bottom). (c) Underdeveloped molars in Sparctm1a(EUCOMM)Wtsi homozygous mice (top), compared with control mice (bottom). (d) Abnormal skeletal muscle in Zfp106tm1a(KOMP)Wtsi homozygous mice (top), compared with controls (bottom). (e) A shortened, upturned snout in Smc3tm1a(EUCOMM)Wtsi heterozygous mice. (f) Metacarpophalangeal joint fusion in Dnase1l2tm1(KOMP)Wtsi homozygous mice. (g) Spots of retinal hyperpigmentation in Slc9a8tm1a(KOMP)Wtsi homozygous mice. (h) LacZ reporter gene expression in the adult mammary gland of Myh9tm1a(EUCOMM)Wtsi heterozygous mice.
In an effort to identify quantitative trait loci (QTLs), large stocks of genetically heterogeneous (HS) mice have been generated [88,89]. Individual mice have been phenotyped and genotyped to facilitate high-precision QTL mapping. The Collaborative Cross (CC) is a resource that is using a similar strategy by interbreeding eight strains of mice to generate around 300 new inbred lines [90], which, unlike HS mice, are being cryopreserved for posterity. It is estimated that the CC will capture approximately 90% of the genetic variability in laboratory mice and will allow the mapping of genetic networks that underlie complex diseases. Moreover, the progenitor strains of the CC were selected for sequencing in the Mouse Genomes Project (Box 1), which should allow the QTLs identified by phenotyping CC mice to be rapidly resolved into a list of candidate variants. When complete, the CC will mark a new era in the discovery of the molecular basis of complex traits in the mouse. Meanwhile, large-scale phenotyping of the strains developed so far is well under way. Finally, EuTRACC is a project to generate ES cells that carry a targeted tandem affinity purification tag (TAP-tag). Initially this will be several hundred transcription factor genes, but this is an effort that is likely to extend genome-wide. This resource will facilitate mass spectrometry of native protein complexes to better understand the mouse 'interactome' [91].
Towards the future
Mouse genetics has a bright future. Genome-wide association studies have identified hundreds of alleles statistically associated with human disease, which now demand detailed functional analysis. Early examples suggest the mouse will be the ideal model in moving from genetic association studies to understanding molecular mechanisms leading to complex disease [92]. The ablation of a large proportion of the coding mouse genome within the next 5 years, at least in ES cells, should rapidly accelerate these studies.
The modular design of modern gene targeting cassettes, together with SSRs and RMCE, makes for an incredibly flexible system of genetic engineering in mice. This is establishing the mouse as a leading model in scientific disciplines previously dominated by work in simpler organisms. For example, gene targeting combined with channelrhodopsin, which allows the control of neural activation using light [93], allows the visualization, and fine manipulation, of precise neural circuits in the mammalian brain that was until recently only possible in Drosophila and C. elegans [94,95].
However, there are also challenges ahead. A significant number of regions across the mouse genome, typically those containing clusters of highly homologous, tightly arrayed genes, are not amenable to efficient gene targeting. Moreover, the same loci are often difficult to sequence, with some lacking complete coverage even in the high quality reference genome [96]. Thus, as much as 5 to 10% of the functional mouse genome may fall through the cracks of the present large-scale projects unless new technologies, or clever combinations of current technologies, are developed and used to investigate these genes. Nevertheless, the mouse is likely to remain the non-human vertebrate with the most sequenced, and best studied, genome for the foreseeable future. Together, the advances described here will underpin an understanding of mouse genetics within the current decade unthinkable to CC Little when he first began generating inbred lines over a century ago [1].
Competing interests
The authors declare that they have no competing interests.
Box 1: A genome for all reasons
The 17 strains being sequenced as part of the Mouse Genomes Project were carefully selected to support other major mouse genetics resources. Three 129 strains were chosen because they serve as the background for thousands of existing gene knock-outs. The C57BL/6N strain is the origin of the highly germline-competent JM8 ES cells that are being used in large-scale gene targeting programs [31]. Nine common lab strains were chosen because of their historical utility, and also because they include the progenitors of the heterogeneous stock and Collaborative Cross mice that are used in dissecting complex traits [88,89]. Finally, four wild-derived strains have been sequenced because they represent some of the founder sub-species of many inbred laboratory lines, and are also important models of cancer and infection resistance [2].
Box 2: Exploiting viruses in mouse genetics
The first transgenic mice were generated by infecting embryos with viruses [97], and today viral vectors remain an integral part of the mouse genetics toolkit. Lentiviruses integrate their genome into the host's DNA, making them an effective transgene delivery vector. The lentiviral genome, derived from immunodeficiency viruses, has been deconstructed and distributed across multiple plasmids to minimize the potential formation of replication-competent viruses [98]. A transgene of interest may be included in a plasmid containing a viral packaging signal. This is co-transfected into a cell line (typically human embryonic kidney HEK293T cells) with other plasmids expressing proteins required for viral production, such as envelope proteins. Viruses produced in this way can be introduced into oocytes for transgenesis (reviewed in [99]). In its simplest form, this method necessitates only a few weeks between target selection and phenotypic analysis, offering a distinct advantage over other approaches. To enable pooled loss-of-function screens to identify complex genetic interactions, lentiviral short hairpin RNA (shRNA) libraries targeting most mouse genes have been generated [27]. Some groups have recently used ultrasound-guided microinjections of lentiviruses to deliver genes to organs and tissues of early mammalian embryos in utero [100].
Slow transforming retroviruses have been widely used to generate mouse models of cancer [101]. They can re-infect the same cell, randomly inserting their genome into the host DNA multiple times, resulting in an accumulation of mutations. This process of progressive mutagenesis recapitulates the multi-step progression of human tumorigenesis (reviewed in [102]). The development of next-generation sequencing technologies has dramatically enhanced the process of identifying retroviral insertion sites, and databases, such as the Retroviral Tagged Cancer Gene Database, have been developed to map insertion sites to the reference genome [103].
Contributor Information
Louise van der Weyden, Email: lvdw@sanger.ac.uk.
Jacqueline K White, Email: not@valid.com.
David J Adams, Email: da1@sanger.ac.uk.
Darren W Logan, Email: dl5@sanger.ac.uk.
Acknowledgements
We thank the Sanger Mouse Genetics Project for generating, managing and phenotyping the mice shown in Figure 2.
References
- Paigen K. One hundred years of mouse genetics: an intellectual history. I. The classical period (1902-1980). Genetics. 2003;163:1–7. doi: 10.1093/genetics/163.1.1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yang H, Bell TA, Churchill GA, Pardo-Manuel de Villena F. On the subspecific origin of the laboratory mouse. Nat Genet. 2007;39:1100–1107. doi: 10.1038/ng2087. [DOI] [PubMed] [Google Scholar]
- Waterston RH, Lindblad-Toh K, Birney E, Rogers J, Abril JF, Agarwal P, Agarwala R, Ainscough R, Alexandersson M, An P, Antonarakis SE, Attwood J, Baertsch R, Bailey J, Barlow K, Beck S, Berry E, Birren B, Bloom T, Bork P, Botcherby M, Bray N, Brent MR, Brown DG, Brown SD, Bult C, Burton J, Butler J, Campbell RD. et al. Initial sequencing and comparative analysis of the mouse genome. Nature. 2002;420:520–562. doi: 10.1038/nature01262. [DOI] [PubMed] [Google Scholar]
- Collins FS, Rossant J, Wurst W. A mouse for all reasons. Cell. 2007;128:9–13. doi: 10.1016/j.cell.2006.12.018. [DOI] [PubMed] [Google Scholar]
- Church DM, Goodstadt L, Hillier LW, Zody MC, Goldstein S, She X, Bult CJ, Agarwala R, Cherry JL, DiCuccio M, Hlavina W, Kapustin Y, Meric P, Maglott D, Birtle Z, Marques AC, Graves T, Zhou S, Teague B, Potamousis K, Churas C, Place M, Herschleb J, Runnheim R, Forrest D, Amos-Landgraf J, Schwartz DC, Cheng Z, Lindblad-Toh K, Eichler EE, Ponting CP. Lineage-specific biology revealed by a finished genome assembly of the mouse. PLoS Biol. 2009;7:e1000112. doi: 10.1371/journal.pbio.1000112. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mural RJ, Adams MD, Myers EW, Smith HO, Miklos GL, Wides R, Halpern A, Li PW, Sutton GG, Nadeau J, Salzberg SL, Holt RA, Kodira CD, Lu F, Chen L, Deng Z, Evangelista CC, Gan W, Heiman TJ, Li J, Li Z, Merkulov GV, Milshina NV, Naik AK, Qi R, Shue BC, Wang A, Wang J, Wang X, Yan X. et al. A comparison of whole-genome shotgun-derived mouse chromosome 16 and the human genome. Science. 2002;296:1661–1671. doi: 10.1126/science.1069193. [DOI] [PubMed] [Google Scholar]
- Adams DJ, Quail MA, Cox T, van der Weyden L, Gorick BD, Su Q, Chan WI, Davies R, Bonfield JK, Law F, Humphray S, Plumb B, Liu P, Rogers J, Bradley A. A genome-wide, end-sequenced 129Sv BAC library resource for targeting vector construction. Genomics. 2005;86:753–758. doi: 10.1016/j.ygeno.2005.08.003. [DOI] [PubMed] [Google Scholar]
- Frazer KA, Eskin E, Kang HM, Bogue MA, Hinds DA, Beilharz EJ, Gupta RV, Montgomery J, Morenzoni MM, Nilsen GB, Pethiyagoda CL, Stuve LL, Johnson FM, Daly MJ, Wade CM, Cox DR. A sequence-based variation map of 8.27 million SNPs in inbred mouse strains. Nature. 2007;448:1050–1053. doi: 10.1038/nature06067. [DOI] [PubMed] [Google Scholar]
- Sudbery I, Stalker J, Simpson JT, Keane T, Rust AG, Hurles ME, Walter K, Lynch D, Teboul L, Brown SD, Li H, Ning Z, Nadeau JH, Croniger CM, Durbin R, Adams DJ. Deep short-read sequencing of chromosome 17 from the mouse strains A/J and CAST/Ei identifies significant germline variation and candidate genes that regulate liver triglyceride levels. Genome Biol. 2009;10:R112. doi: 10.1186/gb-2009-10-10-r112. [DOI] [PMC free article] [PubMed] [Google Scholar]
- The Wellcome Trust Sanger Institute Mouse Portal. http://www.sanger.ac.uk/mouseportal/ [DOI] [PubMed]
- Hitotsumachi S, Carpenter DA, Russell WL. Dose-repetition increases the mutagenic effectiveness of N-ethyl-N-nitrosourea in mouse spermatogonia. Proc Natl Acad Sci USA. 1985;82:6619–6621. doi: 10.1073/pnas.82.19.6619. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hrabe de Angelis MH, Flaswinkel H, Fuchs H, Rathkolb B, Soewarto D, Marschall S, Heffner S, Pargent W, Wuensch K, Jung M, Reis A, Richter T, Alessandrini F, Jakob T, Fuchs E, Kolb H, Kremmer E, Schaeble K, Rollinski B, Roscher A, Peters C, Meitinger T, Strom T, Steckler T, Holsboer F, Klopstock T, Gekeler F, Schindewolf C, Jung T. et al. Genome-wide, large-scale production of mutant mice by ENU mutagenesis. Nat Genet. 2000;25:444–447. doi: 10.1038/78146. [DOI] [PubMed] [Google Scholar]
- Nolan PM, Peters J, Strivens M, Rogers D, Hagan J, Spurr N, Gray IC, Vizor L, Brooker D, Whitehill E, Washbourne R, Hough T, Greenaway S, Hewitt M, Liu X, McCormack S, Pickford K, Selley R, Wells C, Tymowska-Lalanne Z, Roby P, Glenister P, Thornton C, Thaung C, Stevenson JA, Arkell R, Mburu P, Hardisty R, Kiernan A, Erven A. et al. A systematic, genome-wide, phenotype-driven mutagenesis programme for gene function studies in the mouse. Nat Genet. 2000;25:440–443. doi: 10.1038/78140. [DOI] [PubMed] [Google Scholar]
- Beutler B, Du X, Xia Y. Precis on forward genetics in mice. Nat Immunol. 2007;8:659–664. doi: 10.1038/ni0707-659. [DOI] [PubMed] [Google Scholar]
- Coghill EL, Hugill A, Parkinson N, Davison C, Glenister P, Clements S, Hunter J, Cox RD, Brown SD. A gene-driven approach to the identification of ENU mutants in the mouse. Nat Genet. 2002;30:255–256. doi: 10.1038/ng847. [DOI] [PubMed] [Google Scholar]
- Quwailid MM, Hugill A, Dear N, Vizor L, Wells S, Horner E, Fuller S, Weedon J, McMath H, Woodman P, Edwards D, Campbell D, Rodger S, Carey J, Roberts A, Glenister P, Lalanne Z, Parkinson N, Coghill EL, McKeone R, Cox S, Willan J, Greenfield A, Keays D, Brady S, Spurr N, Gray I, Hunter J, Brown SD, Cox RD. A gene-driven ENU-based approach to generating an allelic series in any gene. Mamm Genome. 2004;15:585–591. doi: 10.1007/s00335-004-2379-z. [DOI] [PubMed] [Google Scholar]
- Dupuy AJ. Current applications of transposons in mouse genetics. Methods Enzymol. 2010;477:53–70. doi: 10.1016/S0076-6879(10)77004-X. [DOI] [PubMed] [Google Scholar]
- Friedrich G, Soriano P. Promoter traps in embryonic stem cells: a genetic screen to identify and mutate developmental genes in mice. Genes Dev. 1991;5:1513–1523. doi: 10.1101/gad.5.9.1513. [DOI] [PubMed] [Google Scholar]
- Uren AG, Mikkers H, Kool J, van der Weyden L, Lund AH, Wilson CH, Rance R, Jonkers J, van Lohuizen M, Berns A, Adams DJ. A high-throughput splinkerette-PCR method for the isolation and sequencing of retroviral insertion sites. Nat Protoc. 2009;4:789–798. doi: 10.1038/nprot.2009.64. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gordon JW, Scangos GA, Plotkin DJ, Barbosa JA, Ruddle FH. Genetic transformation of mouse embryos by microinjection of purified DNA. Proc Natl Acad Sci USA. 1980;77:7380–7384. doi: 10.1073/pnas.77.12.7380. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Marshall VM, Allison J, Templeton T, Foote SJ. Generation of BAC transgenic mice. Methods Mol Biol. 2004;256:159–182. doi: 10.1385/1-59259-753-X:159. [DOI] [PubMed] [Google Scholar]
- Vaingankar SM, Li Y, Corti A, Biswas N, Gayen J, O'Connor DT, Mahata SK. Long human CHGA flanking chromosome 14 sequence required for optimal BAC transgenic "rescue" of disease phenotypes in the mouse Chga knockout. Physiol Genomics. 2010;41:91–101. doi: 10.1152/physiolgenomics.00086.2009. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Thomas KR, Capecchi MR. Site-directed mutagenesis by gene targeting in mouse embryo-derived stem cells. Cell. 1987;51:503–512. doi: 10.1016/0092-8674(87)90646-5. [DOI] [PubMed] [Google Scholar]
- Zheng B, Mills AA, Bradley A. A system for rapid generation of coat color-tagged knockouts and defined chromosomal rearrangements in mice. Nucleic Acids Res. 1999;27:2354–2360. doi: 10.1093/nar/27.11.2354. [DOI] [PMC free article] [PubMed] [Google Scholar]
- van der Weyden L, Shaw-Smith C, Bradley A. Chromosome engineering in ES cells. Methods Mol Biol. 2009;530:49–77. doi: 10.1007/978-1-59745-471-1_4. [DOI] [PubMed] [Google Scholar]
- Gao X, Zhang P. Transgenic RNA interference in mice. Physiology (Bethesda) 2007;22:161–166. doi: 10.1152/physiol.00002.2007. [DOI] [PubMed] [Google Scholar]
- Moffat J, Grueneberg DA, Yang X, Kim SY, Kloepfer AM, Hinkle G, Piqani B, Eisenhaure TM, Luo B, Grenier JK, Carpenter AE, Foo SY, Stewart SA, Stockwell BR, Hacohen N, Hahn WC, Lander ES, Sabatini DM, Root DE. A lentiviral RNAi library for human and mouse genes applied to an arrayed viral high-content screen. Cell. 2006;124:1283–1298. doi: 10.1016/j.cell.2006.01.040. [DOI] [PubMed] [Google Scholar]
- Aarts M, Te Riele H. Progress and prospects: oligonucleotide-directed gene modification in mouse embryonic stem cells: a route to therapeutic application. Gene Ther. 2011;18:213–219. doi: 10.1038/gt.2010.161. [DOI] [PubMed] [Google Scholar]
- Meyer M, de Angelis MH, Wurst W, Kuhn R. Gene targeting by homologous recombination in mouse zygotes mediated by zinc-finger nucleases. Proc Natl Acad Sci USA. 2010;107:15022–15026. doi: 10.1073/pnas.1009424107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Adams DJ, van der Weyden L. Contemporary approaches for modifying the mouse genome. Physiol Genomics. 2008;34:225–238. doi: 10.1152/physiolgenomics.90242.2008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pettitt SJ, Liang Q, Rairdan XY, Moran JL, Prosser HM, Beier DR, Lloyd KC, Bradley A, Skarnes WC. Agouti C57BL/6N embryonic stem cells for mouse genetic resources. Nat Methods. 2009;6:493–495. doi: 10.1038/nmeth.1342. [DOI] [PMC free article] [PubMed] [Google Scholar]
- van der Weyden L, Adams DJ, Bradley A. Tools for targeted manipulation of the mouse genome. Physiol Genomics. 2002;11:133–164. doi: 10.1152/physiolgenomics.00074.2002. [DOI] [PubMed] [Google Scholar]
- Osoegawa K, Tateno M, Woon PY, Frengen E, Mammoser AG, Catanese JJ, Hayashizaki Y, de Jong PJ. Bacterial artificial chromosome libraries for mouse sequencing and functional analysis. Genome Res. 2000;10:116–128. [PMC free article] [PubMed] [Google Scholar]
- Copeland NG, Jenkins NA, Court DL. Recombineering: a powerful new tool for mouse functional genomics. Nat Rev Genet. 2001;2:769–779. doi: 10.1038/35093556. [DOI] [PubMed] [Google Scholar]
- Anastassiadis K, Fu J, Patsch C, Hu S, Weidlich S, Duerschke K, Buchholz F, Edenhofer F, Stewart AF. Dre recombinase, like Cre, is a highly efficient site-specific recombinase in E. coli, mammalian cells and mice. Dis Model Mech. 2009;2:508–515. doi: 10.1242/dmm.003087. [DOI] [PubMed] [Google Scholar]
- Birling MC, Gofflot F, Warot X. Site-specific recombinases for manipulation of the mouse genome. Methods Mol Biol. 2009;561:245–263. doi: 10.1007/978-1-60327-019-9_16. [DOI] [PubMed] [Google Scholar]
- Schnutgen F, Doerflinger N, Calleja C, Wendling O, Chambon P, Ghyselinck NB. A directional strategy for monitoring Cre-mediated recombination at the cellular level in the mouse. Nat Biotechnol. 2003;21:562–565. doi: 10.1038/nbt811. [DOI] [PubMed] [Google Scholar]
- Yu Y, Bradley A. Engineering chromosomal rearrangements in mice. Nat Rev Genet. 2001;2:780–790. doi: 10.1038/35093564. [DOI] [PubMed] [Google Scholar]
- Yap SP, Xing X, Kraus P, Sivakamasundari V, Chan HY, Lufkin T. Generation of mice with a novel conditional null allele of the Sox9 gene. Biotechnol Lett. 2011. [DOI] [PubMed]
- Gossen M, Bujard H. Tight control of gene expression in mammalian cells by tetracycline-responsive promoters. Proc Natl Acad Sci USA. 1992;89:5547–5551. doi: 10.1073/pnas.89.12.5547. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cronin CA, Gluba W, Scrable H. The lac operator-repressor system is functional in the mouse. Genes Dev. 2001;15:1506–1517. doi: 10.1101/gad.892001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Feil R, Brocard J, Mascrez B, LeMeur M, Metzger D, Chambon P. Ligand-activated site-specific recombination in mice. Proc Natl Acad Sci USA. 1996;93:10887–10890. doi: 10.1073/pnas.93.20.10887. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Reijmers L, Mayford M. Genetic control of active neural circuits. Front Mol Neurosci. 2009;2:27. doi: 10.3389/neuro.02.027.2009. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nagy A, Mar L, Watts G. Creation and use of a cre recombinase transgenic database. Methods Mol Biol. 2009;530:365–378. doi: 10.1007/978-1-59745-471-1_19. [DOI] [PubMed] [Google Scholar]
- Wassarman KM, Lewandoski M, Campbell K, Joyner AL, Rubenstein JL, Martinez S, Martin GR. Specification of the anterior hindbrain and establishment of a normal mid/hindbrain organizer is dependent on Gbx2 gene function. Development. 1997;124:2923–2934. doi: 10.1242/dev.124.15.2923. [DOI] [PubMed] [Google Scholar]
- Schulz TJ, Glaubitz M, Kuhlow D, Thierbach R, Birringer M, Steinberg P, Pfeiffer AF, Ristow M. Variable expression of Cre recombinase transgenes precludes reliable prediction of tissue-specific gene disruption by tail-biopsy genotyping. PLoS One. 2007;2:e1013. doi: 10.1371/journal.pone.0001013. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Turlo KA, Gallaher SD, Vora R, Laski FA, Iruela-Arispe ML. When Cre-mediated recombination in mice does not result in protein loss. Genetics. 2010;186:959–967. doi: 10.1534/genetics.110.121608. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Turan S, Galla M, Ernst E, Qiao J, Voelkel C, Schiedlmeier B, Zehe C, Bode J. Recombinase-mediated cassette exchange (RMCE): traditional concepts and current challenges. J Mol Biol. 2011;407:193–221. doi: 10.1016/j.jmb.2011.01.004. [DOI] [PubMed] [Google Scholar]
- Wirth D, Gama-Norton L, Riemer P, Sandhu U, Schucht R, Hauser H. Road to precision: recombinase-based targeting technologies for genome engineering. Curr Opin Biotechnol. 2007;18:411–419. doi: 10.1016/j.copbio.2007.07.013. [DOI] [PubMed] [Google Scholar]
- Sorrell DA, Robinson CJ, Smith JA, Kolb AF. Recombinase mediated cassette exchange into genomic targets using an adenovirus vector. Nucleic Acids Res. 2010;38:e123. doi: 10.1093/nar/gkq192. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Osterwalder M, Galli A, Rosen B, Skarnes WC, Zeller R, Lopez-Rios J. Dual RMCE for efficient re-engineering of mouse mutant alleles. Nat Methods. 2010;7:893–895. doi: 10.1038/nmeth.1521. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wei QX, Odell AF, van der Hoeven F, Hollstein M. Rapid derivation of genetically related mutants from embryonic cells harboring a recombinase-specific Trp53 platform. Cell Cycle. 2011;10:1261–1270. doi: 10.4161/cc.10.8.15303. [DOI] [PubMed] [Google Scholar]
- Chen SX, Osipovich AB, Ustione A, Potter LA, Hipkens S, Gangula R, Yuan W, Piston DW, Magnuson MA. Quantification of factors influencing fluorescent protein expression using RMCE to generate an allelic series in the ROSA26 locus in mice. Dis Model Mech. 2011. [DOI] [PMC free article] [PubMed]
- An W, Han JS, Wheelan SJ, Davis ES, Coombes CE, Ye P, Triplett C, Boeke JD. Active retrotransposition by a synthetic L1 element in mice. Proc Natl Acad Sci USA. 2006;103:18662–18667. doi: 10.1073/pnas.0605300103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chew SK, Rad R, Futreal PA, Bradley A, Liu P. Genetic screens using the piggyBac transposon. Methods. 2011;53:366–371. doi: 10.1016/j.ymeth.2010.12.022. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Keng VW, Yae K, Hayakawa T, Mizuno S, Uno Y, Yusa K, Kokubu C, Kinoshita T, Akagi K, Jenkins NA, Copeland NG, Horie K, Takeda J. Region-specific saturation germline mutagenesis in mice using the Sleeping Beauty transposon system. Nat Methods. 2005;2:763–769. doi: 10.1038/nmeth795. [DOI] [PubMed] [Google Scholar]
- Geurts AM, Collier LS, Geurts JL, Oseth LL, Bell ML, Mu D, Lucito R, Godbout SA, Green LE, Lowe SW, Hirsch BA, Leinwand LA, Largaespada DA. Gene mutations and genomic rearrangements in the mouse as a result of transposon mobilization from chromosomal concatemers. PLoS Genet. 2006;2:e156. doi: 10.1371/journal.pgen.0020156. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dupuy AJ, Akagi K, Largaespada DA, Copeland NG, Jenkins NA. Mammalian mutagenesis using a highly mobile somatic Sleeping Beauty transposon system. Nature. 2005;436:221–226. doi: 10.1038/nature03691. [DOI] [PubMed] [Google Scholar]
- Collier LS, Carlson CM, Ravimohan S, Dupuy AJ, Largaespada DA. Cancer gene discovery in solid tumours using transposon-based somatic mutagenesis in the mouse. Nature. 2005;436:272–276. doi: 10.1038/nature03681. [DOI] [PubMed] [Google Scholar]
- Dupuy AJ, Rogers LM, Kim J, Nannapaneni K, Starr TK, Liu P, Largaespada DA, Scheetz TE, Jenkins NA, Copeland NG. A modified sleeping beauty transposon system that can be used to model a wide variety of human cancers in mice. Cancer Res. 2009;69:8150–8156. doi: 10.1158/0008-5472.CAN-09-1135. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Keng VW, Villanueva A, Chiang DY, Dupuy AJ, Ryan BJ, Matise I, Silverstein KA, Sarver A, Starr TK, Akagi K, Tessarollo L, Collier LS, Powers S, Lowe SW, Jenkins NA, Copeland NG, Llovet JM, Largaespada DA. A conditional transposon-based insertional mutagenesis screen for genes associated with mouse hepatocellular carcinoma. Nat Biotechnol. 2009;27:264–274. doi: 10.1038/nbt.1526. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Starr TK, Allaei R, Silverstein KA, Staggs RA, Sarver AL, Bergemann TL, Gupta M, O'Sullivan MG, Matise I, Dupuy AJ, Collier LS, Powers S, Oberg AL, Asmann YW, Thibodeau SN, Tessarollo L, Copeland NG, Jenkins NA, Cormier RT, Largaespada DA. A transposon-based genetic screen in mice identifies genes altered in colorectal cancer. Science. 2009;323:1747–1750. doi: 10.1126/science.1163040. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rad R, Rad L, Wang W, Cadinanos J, Vassiliou G, Rice S, Campos LS, Yusa K, Banerjee R, Li MA, de la Rosa J, Strong A, Lu D, Ellis P, Conte N, Yang FT, Liu P, Bradley A. PiggyBac transposon mutagenesis: a tool for cancer gene discovery in mice. Science. 2010;330:1104–1107. doi: 10.1126/science.1193004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Suster ML, Sumiyama K, Kawakami K. Transposon-mediated BAC transgenesis in zebrafish and mice. BMC Genomics. 2009;10:477. doi: 10.1186/1471-2164-10-477. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wu S, Ying G, Wu Q, Capecchi MR. Toward simpler and faster genome-wide mutagenesis in mice. Nat Genet. 2007;39:922–930. doi: 10.1038/ng2060. [DOI] [PubMed] [Google Scholar]
- Kaji K, Norrby K, Paca A, Mileikovsky M, Mohseni P, Woltjen K. Virus-free induction of pluripotency and subsequent excision of reprogramming factors. Nature. 2009;458:771–775. doi: 10.1038/nature07864. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yusa K, Rad R, Takeda J, Bradley A. Generation of transgene-free induced pluripotent mouse stem cells by the piggyBac transposon. Nat Methods. 2009;6:363–369. doi: 10.1038/nmeth.1323. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Friedel RH, Soriano P. Gene trap mutagenesis in the mouse. Methods Enzymol. 2010;477:243–269. doi: 10.1016/S0076-6879(10)77013-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Floss T, Schnutgen F. Conditional gene trapping using the FLEx system. Methods Mol Biol. 2008;435:127–138. doi: 10.1007/978-1-59745-232-8_9. [DOI] [PubMed] [Google Scholar]
- Korn R, Schoor M, Neuhaus H, Henseling U, Soininen R, Zachgo J, Gossler A. Enhancer trap integrations in mouse embryonic stem cells give rise to staining patterns in chimaeric embryos with a high frequency and detect endogenous genes. Mech Dev. 1992;39:95–109. doi: 10.1016/0925-4773(92)90029-J. [DOI] [PubMed] [Google Scholar]
- Forrester LM, Nagy A, Sam M, Watt A, Stevenson L, Bernstein A, Joyner AL, Wurst W. An induction gene trap screen in embryonic stem cells: Identification of genes that respond to retinoic acid in vitro. Proc Natl Acad Sci USA. 1996;93:1677–1682. doi: 10.1073/pnas.93.4.1677. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hirashima M, Bernstein A, Stanford WL, Rossant J. Gene-trap expression screening to identify endothelial-specific genes. Blood. 2004;104:711–718. doi: 10.1182/blood-2004-01-0254. [DOI] [PubMed] [Google Scholar]
- Guo G, Wang W, Bradley A. Mismatch repair genes identified using genetic screens in Blm-deficient embryonic stem cells. Nature. 2004;429:891–895. doi: 10.1038/nature02653. [DOI] [PubMed] [Google Scholar]
- DeChiara TM, Poueymirou WT, Auerbach W, Frendewey D, Yancopoulos GD, Valenzuela DM. Producing fully ES cell-derived mice from eight-cell stage embryo injections. Methods Enzymol. 2010;476:285–294. doi: 10.1016/S0076-6879(10)76016-X. [DOI] [PubMed] [Google Scholar]
- George SH, Gertsenstein M, Vintersten K, Korets-Smith E, Murphy J, Stevens ME, Haigh JJ, Nagy A. Developmental and adult phenotyping directly from mutant embryonic stem cells. Proc Natl Acad Sci USA. 2007;104:4455–4460. doi: 10.1073/pnas.0609277104. [DOI] [PMC free article] [PubMed] [Google Scholar]
- McJunkin K, Mazurek A, Premsrirut PK, Zuber J, Dow LE, Simon J, Stillman B, Lowe SW. Reversible suppression of an essential gene in adult mice using transgenic RNA interference. Proc Natl Acad Sci USA. 2011;108:7113–7118. doi: 10.1073/pnas.1104097108. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Premsrirut PK, Dow LE, Kim SY, Camiolo M, Malone CD, Miething C, Scuoppo C, Zuber J, Dickins RA, Kogan SC, Shroyer KR, Sordella R, Hannon GJ, Lowe SW. A rapid and scalable system for studying gene function in mice using conditional RNA interference. Cell. 2011;145:145–158. doi: 10.1016/j.cell.2011.03.012. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hochedlinger K, Yamada Y, Beard C, Jaenisch R. Ectopic expression of Oct-4 blocks progenitor-cell differentiation and causes dysplasia in epithelial tissues. Cell. 2005;121:465–477. doi: 10.1016/j.cell.2005.02.018. [DOI] [PubMed] [Google Scholar]
- Ringwald M, Iyer V, Mason JC, Stone KR, Tadepally HD, Kadin JA, Bult CJ, Eppig JT, Oakley DJ, Briois S, Stupka E, Maselli V, Smedley D, Liu S, Hansen J, Baldock R, Hicks GG, Skarnes WC. The IKMC web portal: a central point of entry to data and resources from the International Knockout Mouse Consortium. Nucleic Acids Res. 2011;39:D849–D855. doi: 10.1093/nar/gkq879. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Testa G, Schaft J, van der Hoeven F, Glaser S, Anastassiadis K, Zhang Y, Hermann T, Stremmel W, Stewart AF. A reliable lacZ expression reporter cassette for multipurpose, knockout-first alleles. Genesis. 2004;38:151–158. doi: 10.1002/gene.20012. [DOI] [PubMed] [Google Scholar]
- Brown SD, Chambon P, de Angelis MH. EMPReSS: standardized phenotype screens for functional annotation of the mouse genome. Nat Genet. 2005;37:1155. doi: 10.1038/ng1105-1155. [DOI] [PubMed] [Google Scholar]
- Fuchs H, Gailus-Durner V, Adler T, Aguilar-Pimentel JA, Becker L, Calzada-Wack J, Da Silva-Buttkus P, Neff F, Götz A, Hans W, Hölter SM, Horsch M, Kastenmüller G, Kemter E, Lengger C, Maier H, Matloka M, Möller G, Naton B, Prehn C, Puk O, Rácz I, Rathkolb B, Römisch-Margl W, Rozman J, Wang-Sattler R, Schrewe A, Stöger C, Tost M, Adamski J, Aigner B, Beckers J. et al. Mouse phenotyping. Methods. 2011;53:120–135. doi: 10.1016/j.ymeth.2010.08.006. [DOI] [PubMed] [Google Scholar]
- Bogue MA, Grubb SC. The Mouse Phenome Project. Genetica. 2004;122:71–74. doi: 10.1007/s10709-004-1438-4. [DOI] [PubMed] [Google Scholar]
- Crawley JN, Belknap JK, Collins A, Crabbe JC, Frankel W, Henderson N, Hitzemann RJ, Maxson SC, Miner LL, Silva AJ, Wehner JM, Wynshaw-Boris A, Paylor R. Behavioral phenotypes of inbred mouse strains: implications and recommendations for molecular studies. Psychopharmacology (Berl) 1997;132:107–124. doi: 10.1007/s002130050327. [DOI] [PubMed] [Google Scholar]
- Crossan GP, van der Weyden L, Rosado IV, Langevin F, Gaillard PH, McIntyre RE, Gallagher F, Kettunen MI, Lewis DY, Brindle K, Arends MJ, Adams DJ, Patel KJ. Disruption of mouse Slx4, a regulator of structure-specific nucleases, phenocopies Fanconi anemia. Nat Genet. 2011;43:147–152. doi: 10.1038/ng.752. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kim Y, Lach FP, Desetty R, Hanenberg H, Auerbach AD, Smogorzewska A. Mutations of the SLX4 gene in Fanconi anemia. Nat Genet. 2011;43:142–146. doi: 10.1038/ng.750. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Stoepker C, Hain K, Schuster B, Hilhorst-Hofstee Y, Rooimans MA, Steltenpool J, Oostra AB, Eirich K, Korthof ET, Nieuwint AW, Jaspers NG, Bettecken T, Joenje H, Schindler D, Rouse J, de Winter JP. SLX4, a coordinator of structure-specific endonucleases, is mutated in a new Fanconi anemia subtype. Nat Genet. 2011;43:138–141. doi: 10.1038/ng.751. [DOI] [PubMed] [Google Scholar]
- Valdar W, Solberg LC, Gauguier D, Burnett S, Klenerman P, Cookson WO, Taylor MS, Rawlins JN, Mott R, Flint J. Genome-wide genetic association of complex traits in heterogeneous stock mice. Nat Genet. 2006;38:879–887. doi: 10.1038/ng1840. [DOI] [PubMed] [Google Scholar]
- Threadgill DW, Hunter KW, Williams RW. Genetic dissection of complex and quantitative traits: from fantasy to reality via a community effort. Mamm Genome. 2002;13:175–178. doi: 10.1007/s00335-001-4001-y. [DOI] [PubMed] [Google Scholar]
- Chesler EJ, Miller DR, Branstetter LR, Galloway LD, Jackson BL, Philip VM, Voy BH, Culiat CT, Threadgill DW, Williams RW, Churchill GA, Johnson DK, Manly KF. The Collaborative Cross at Oak Ridge National Laboratory: developing a powerful resource for systems genetics. Mamm Genome. 2008;19:382–389. doi: 10.1007/s00335-008-9135-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Collins MO, Choudhary JS. Mapping multiprotein complexes by affinity purification and mass spectrometry. Curr Opin Biotechnol. 2008;19:324–330. doi: 10.1016/j.copbio.2008.06.002. [DOI] [PubMed] [Google Scholar]
- Larder R, Cheung MK, Tung YC, Yeo GS, Coll AP. Where to go with FTO?. Trends Endocrinol Metab. 2011;22:53–59. doi: 10.1016/j.tem.2010.11.001. [DOI] [PubMed] [Google Scholar]
- LaLumiere RT. A new technique for controlling the brain: optogenetics and its potential for use in research and the clinic. Brain Stimul. 2011;4:1–6. doi: 10.1016/j.brs.2010.09.009. [DOI] [PubMed] [Google Scholar]
- Haubensak W, Kunwar PS, Cai H, Ciocchi S, Wall NR, Ponnusamy R, Biag J, Dong HW, Deisseroth K, Callaway EM, Fanselow MS, Lüthi A, Anderson DJ. Genetic dissection of an amygdala microcircuit that gates conditioned fear. Nature. 2010;468:270–276. doi: 10.1038/nature09553. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Aponte Y, Atasoy D, Sternson SM. AGRP neurons are sufficient to orchestrate feeding behavior rapidly and without training. Nat Neurosci. 2011;14:351–355. doi: 10.1038/nn.2739. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Logan DW, Marton TF, Stowers L. Species specificity in major urinary proteins by parallel evolution. PLoS ONE. 2008;3:e3280. doi: 10.1371/journal.pone.0003280. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jaenisch R, Mintz B. Simian virus 40 DNA sequences in DNA of healthy adult mice derived from preimplantation blastocysts injected with viral DNA. Proc Natl Acad Sci USA. 1974;71:1250–1254. doi: 10.1073/pnas.71.4.1250. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Naldini L, Blomer U, Gallay P, Ory D, Mulligan R, Gage FH, Verma IM, Trono D. In vivo gene delivery and stable transduction of nondividing cells by a lentiviral vector. Science. 1996;272:263–267. doi: 10.1126/science.272.5259.263. [DOI] [PubMed] [Google Scholar]
- Pfeifer A, Hofmann A. Lentiviral transgenesis. Methods Mol Biol. 2009;530:391–405. doi: 10.1007/978-1-59745-471-1_21. [DOI] [PubMed] [Google Scholar]
- Endo M, Zoltick PW, Peranteau WH, Radu A, Muvarak N, Ito M, Yang Z, Cotsarelis G, Flake AW. Efficient in vivo targeting of epidermal stem cells by early gestational intraamniotic injection of lentiviral vector driven by the keratin 5 promoter. Mol Ther. 2008;16:131–137. doi: 10.1038/sj.mt.6300332. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mikkers H, Berns A. Retroviral insertional mutagenesis: tagging cancer pathways. Adv Cancer Res. 2003;88:53–99. doi: 10.1016/s0065-230x(03)88304-5. [DOI] [PubMed] [Google Scholar]
- Uren AG, Kool J, Berns A, van Lohuizen M. Retroviral insertional mutagenesis: past, present and future. Oncogene. 2005;24:7656–7672. doi: 10.1038/sj.onc.1209043. [DOI] [PubMed] [Google Scholar]
- Akagi K, Suzuki T, Stephens RM, Jenkins NA, Copeland NG. RTCGD: retroviral tagged cancer gene database. Nucleic Acids Res. 2004;32:D523–D527. doi: 10.1093/nar/gkh013. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hooper M, Hardy K, Handyside A, Hunter S, Monk M. HPRT-deficient (Lesch-Nyhan) mouse embryos derived from germline colonization by cultured cells. Nature. 1987;326:292–295. doi: 10.1038/326292a0. [DOI] [PubMed] [Google Scholar]
- Ramirez-Solis R, Liu P, Bradley A. Chromosome engineering in mice. Nature. 1995;378:720–724. doi: 10.1038/378720a0. [DOI] [PubMed] [Google Scholar]
- Li E, Bestor TH, Jaenisch R. Targeted mutation of the DNA methyltransferase gene results in embryonic lethality. Cell. 1992;69:915–926. doi: 10.1016/0092-8674(92)90611-F. [DOI] [PubMed] [Google Scholar]
- Kontgen F, Suss G, Stewart C, Steinmetz M, Bluethmann H. Targeted disruption of the MHC class II Aa gene in C57BL/6 mice. Int Immunol. 1993;5:957–964. doi: 10.1093/intimm/5.8.957. [DOI] [PubMed] [Google Scholar]
- Gertsenstein M, Nutter LM, Reid T, Pereira M, Stanford WL, Rossant J, Nagy A. Efficient generation of germ line transmitting chimeras from C57BL/6N ES cells by aggregation with outbred host embryos. PLoS One. 2010;5:e11260. doi: 10.1371/journal.pone.0011260. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nagy A, Rossant J, Nagy R, Abramow-Newerly W, Roder JC. Derivation of completely cell culture-derived mice from early-passage embryonic stem cells. Proc Natl Acad Sci USA. 1993;90:8424–8428. doi: 10.1073/pnas.90.18.8424. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Churchill GA, Airey DC, Allayee H, Angel JM, Attie AD, Beatty J, Beavis WD, Belknap JK, Bennett B, Berrettini W, Bleich A, Bogue M, Broman KW, Buck KJ, Buckler E, Burmeister M, Chesler EJ, Cheverud JM, Clapcote S, Cook MN, Cox RD, Crabbe JC, Crusio WE, Darvasi A, Deschepper CF, Doerge RW, Farber CR, Forejt J, Gaile D, Garlow SJ. et al. The Collaborative Cross, a community resource for the genetic analysis of complex traits. Nat Genet. 2004;36:1133–1137. doi: 10.1038/ng1104-1133. [DOI] [PubMed] [Google Scholar]
- Flicek P, Amode MR, Barrell D, Beal K, Brent S, Chen Y, Clapham P, Coates G, Fairley S, Fitzgerald S, Gordon L, Hendrix M, Hourlier T, Johnson N, Kähäri A, Keefe D, Keenan S, Kinsella R, Kokocinski F, Kulesha E, Larsson P, Longden I, McLaren W, Overduin B, Pritchard B, Riat HS, Rios D, Ritchie GR, Ruffier M, Schuster M. et al. Ensembl 2011. Nucleic Acids Res. 2011;39:D800–806. doi: 10.1093/nar/gkq1064. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Auwerx J, Avner P, Baldock R, Ballabio A, Balling R, Barbacid M, Berns A, Bradley A, Brown S, Carmeliet P, Chambon P, Cox R, Davidson D, Davies K, Duboule D, Forejt J, Granucci F, Hastie N, de Angelis MH, Jackson I, Kioussis D, Kollias G, Lathrop M, Lendahl U, Malumbres M, von Melchner H, Müller W, Partanen J, Ricciardi-Castagnoli P, Rigby P. et al. The European dimension for the mouse genome mutagenesis program. Nat Genet. 2004;36:925–927. doi: 10.1038/ng0904-925. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wilkinson P, Sengerova J, Matteoni R, Chen CK, Soulat G, Ureta-Vidal A, Fessele S, Hagn M, Massimi M, Pickford K, Butler RH, Marschall S, Mallon AM, Pickard A, Raspa M, Scavizzi F, Fray M, Larrigaldie V, Leyritz J, Birney E, Tocchini-Valentini GP, Brown S, Herault Y, Montoliu L, de Angelis MH, Smedley D. EMMA--mouse mutant resources for the international scientific community. Nucleic Acids Res. 2010;38:D570–D576. doi: 10.1093/nar/gkp799. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mandillo S, Tucci V, Hölter SM, Meziane H, Banchaabouchi MA, Kallnik M, Lad HV, Nolan PM, Ouagazzal AM, Coghill EL, Gale K, Golini E, Jacquot S, Krezel W, Parker A, Riet F, Schneider I, Marazziti D, Auwerx J, Brown SD, Chambon P, Rosenthal N, Tocchini-Valentini G, Wurst W. Reliability, robustness, and reproducibility in mouse behavioral phenotyping: a cross-laboratory study. Physiol Genomics. 2008;34:243–255. doi: 10.1152/physiolgenomics.90207.2008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mallon AM, Blake A, Hancock JM. EuroPhenome and EMPReSS: online mouse phenotyping resource. Nucleic Acids Res. 2008;36:D715–D718. doi: 10.1093/nar/gkm728. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Davisson M. FIMRe: Federation of International Mouse Resources: global networking of resource centers. Mamm Genome. 2006;17:363–364. doi: 10.1007/s00335-006-0001-2. [DOI] [PubMed] [Google Scholar]
- Araki M, Araki K, Yamamura K. International Gene Trap Project: towards gene-driven saturation mutagenesis in mice. Curr Pharm Biotechnol. 2009;10:221–229. doi: 10.2174/138920109787315006. [DOI] [PubMed] [Google Scholar]
- Blake A, Pickford K, Greenaway S, Thomas S, Pickard A, Williamson CM, Adams NC, Walling A, Beck T, Fray M, Peters J, Weaver T, Brown SD, Hancock JM, Mallon AM. MouseBook: an integrated portal of mouse resources. Nucleic Acids Res. 2010;38:D593–D599. doi: 10.1093/nar/gkp867. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Blake JA, Bult CJ, Kadin JA, Richardson JE, Eppig JT. The Mouse Genome Database (MGD): premier model organism resource for mammalian genomics and genetics. Nucleic Acids Res. 2011;39:D842–848. doi: 10.1093/nar/gkq1008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sun LV, Jin K, Liu Y, Yang W, Xie X, Ye L, Wang L, Zhu L, Ding S, Su Y, Zhou J, Han M, Zhuang Y, Xu T, Wu X, Gu N, Zhong Y. PBmice: an integrated database system of piggyBac (PB) insertional mutations and their characterizations in mice. Nucleic Acids Res. 2008;36:D729–D734. doi: 10.1093/nar/gkm790. [DOI] [PMC free article] [PubMed] [Google Scholar]