Abstract
Background
Many studies have been published outlining the global effects of 17β-estradiol (E2) on gene expression in human epithelial breast cancer derived MCF-7 cells. These studies show large variation in results, reporting between ~100 and ~1500 genes regulated by E2, with poor overlap.
Results
We performed a meta-analysis of these expression studies, using the Rank product method to obtain a more accurate and stable list of the differentially expressed genes, and of pathways regulated by E2. We analyzed 9 time-series data sets, concentrating on response at 3-4 hrs (early) and at 24 hrs (late). We found >1000 statistically significant probe sets after correction for multiple testing at 3-4 hrs, and >2000 significant probe sets at 24 hrs. Differentially expressed genes were examined by pathway analysis. This revealed 15 early response pathways, mostly related to cell signaling and proliferation, and 20 late response pathways, mostly related to breast cancer, cell division, DNA repair and recombination.
Conclusions
Our results confirm that meta-analysis identified more differentially expressed genes than the individual studies, and that these genes act together in networks. These results provide new insight into E2 regulated mechanisms, especially in the context of breast cancer.
Keywords: microarray, meta-analysis, estrogen, breast cancer, pathways
Background
Estrogens are fundamental to the function of the female reproductive system, and have also been shown to regulate diverse cellular processes in the central nervous system, the cardiovascular system, and bone metabolism [1]. Estrogens play a key role in proliferation and differentiation of healthy breast epithelium, but also contribute to the progression of breast cancer by promoting the growth of transformed cells [2]. The predominant effect of estrogen is mediated through its interaction with two intracellular estrogen receptors, ERα and ERβ. ERα notably is strongly expressed in 80% of breast cancers [3]. Once estrogen is bound to ERα, the receptor dimerizes and associates with chromatin. ERα homodimers bind either directly to a DNA sequence motif, the estrogen response element [4-6], or indirectly via tethering to other transcription factors such as specificity protein 1 (Sp1) [7,8], activating protein 1 (AP-1) [9,10], or nuclear factor kappa b (NF-κB) [11].
17β-Estradiol (E2) is the most prevalent intracellular estrogen. Tiled or cDNA microarrays have been abundantly used for the global analysis of 17β-Estradiol (E2) responsive genes. Several studies have profiled the transcriptional response to E2 in cultured cells, particularly MCF-7 cells, and have uncovered comprehensive features of E2 regulated gene expression. At early time points following E2 induction a greater proportion of genes are up regulated, while at later time points more genes are down regulated [12]. It has been hypothesized that early ERα-mediated down regulation might be due to squelching, while the increase in the number of down regulated genes at later time points might depend on the up regulation of the corepressor nuclear receptor interacting protein 1 (NRIP1), which mediates the repression of ERα target genes [5]. Interestingly, only about 20% to 30% of the E2-regulated genes represent direct targets [13], as defined by treatment with the protein translation inhibitor cyclohexamide.
There is considerable variation in the results generated by different laboratories, even for the same cell type and hormone treatment[14]. Estimations of the number of E2-regulated genes range from ≈100 to ≈1500[14]. As noted by Cheung et al., [15] the differences between studies can be due to the origin and growth conditions of the cell lines, the length of hormone treatment, the array platforms used, or the bioinformatics methods used for data analysis.
Meta-analysis allows combining the results of several studies, to provide a global picture, with statistical support, and hopefully better power and specificity than each of the original studies. Several meta-analysis applications for microarrays have been proposed [16-18]. Two methods have been most frequently applied: one is to combine p-values, and the other is to combine effect sizes. Combining p-values has been very useful in obtaining more accurate estimates of significance. Choi et al [16]have shown that combining effect sizes can take into account inter-study variation. Here we have used the Rank product [19] method to detect differentially expressed genes in response to E2 treatment in MCF-7 cell lines, by integrating multiple array datasets from two different platforms across laboratories. This method detects genes that are consistently highly ranked in several replicated experiments, independently of their numerical intensities. It generates a single significance measurement for each gene in the combined study.
Results
Data set selection and processing
Microarray Data was downloaded from GEO (http://www.ncbi.nlm.nih.gov/geo/) [20]. Nine time course studies were used for the meta-analysis. Two common time points among the different datasets were selected for further analysis: 3 to 4 hours of treatment (early) and 24 hours of treatment (late). In total we had 4 datasets that were used for the early time point, and 7 datasets that were used for the late time point (tables 1 and 2).
Table 1.
Studies used for meta-analysis
Table 2.
Conditions of treatment and expression measure of each study
Eight of the 9 studies were provided with data normalized by the Robust Multi-Array Average (RMA)[21] method, or the related Guanine Cytosine Robust Multi-Array Analysis (GCRMA)[22] method. For one study raw scaled signal count values were provided. Rank product method does not require the recalculation of the normalized expression values. Thus we use the data processed according to what was considered best practice in each lab.
For the meta-analysis, the probesets found in common across the Affymetrix U133 GeneChip family was used. There are 22,277 common probesets between the two U133a and U133 plus 2.0, which map to 13,186 genes.
Meta-Analysis of gene expression data
The Rank product method was chosen, as non-parametric analyses are more robust in general, which is an important concern in comparing experiments done in different laboratories. The Rank product method has been shown to give good results on microarray data [18]. Out of 22,277 probe sets, we identified 1206 at the early time point and 4193 at the late time point, with a false discovery rate (called percentage of false positive predictions or pfp, see methods) ≤ 0.05. The high proportion of genes with low p-values indicates that many more genes are found to be differentially expressed than expected by chance. By mapping the probe sets to genes, we have identified 991 unique genes differentially expressed at the early time point and 3234 unique genes that are differentially expressed at the late time point. All results are available in Additional File 1.
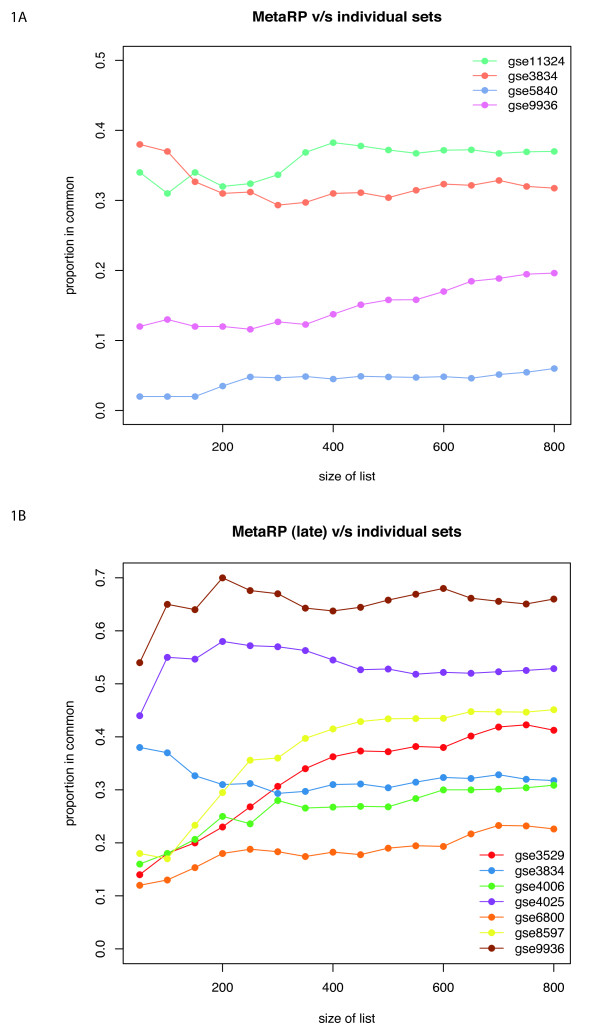
We compared the proportion of top genes identified in individual datasets with the top genes from the meta-analysis, using Correspondence At the Top (CAT) plots [17]. CAT plots quantify the consensus of two lists, based on ranks. For example if two lists have 40 genes out of the first 100 in common, the their consensus at rank 100 will be 40%. In Figure 1 the CAT plots are shown for the top 800 probes (400 up-regulated and 400-down regulated) at each time point. These plots show that no single study determines or influences the final meta-analysis gene list. Table 3 also shows that the meta-analysis is able to identify more genes at the same pfp level, demonstrating an increased statistical power.
Figure 1.
CAT plots. Concordance at the top plots comparing ranked probe set lists from of early time point (3-4 hrs) (A), and for late time point (24 hrs) (B).
Table 3.
Average number of genes identified at different pfp levels and integration driven discovery rate (IDD) in meta-analysis(shown in parenthesis)
| Pfp Value | Early Single | Early Meta | Late Single | Late Meta |
|---|---|---|---|---|
| 0.00001 | 101 | 147(0.32) | 680 | 1271(0.46) |
| 0.0001 | 112 | 210(0.47) | 979 | 1673(0.41) |
| 0.001 | 234 | 350(0.33) | 1795 | 2223(0.2) |
| 0.005 | 470 | 521(0.10) | 2834 | 2803 |
"Single" reports the union of all genes recovered by any single study at a given pfp value.
Top Genes
Tables 4 and 5 show the top 10 unique genes from the meta-analysis for the early time point and their ranks in the individual datasets. The tables also show the ranks of the union of all top 10 genes from individual datasets and meta-dataset, to illustrate the specificity of the meta-analysis relative to individual experiments.
Table 4.
Comparison of top 10 early up regulated genes
| Genes | GSE11324 | GSE3834 | GSE5840 | GSE9936 | Meta |
|---|---|---|---|---|---|
| RET | 4 | 6 | 19 | 17 | 1 |
| GREB1 | 1 | 1 | * | 26 | 2 |
| CXCL12 | 12 | 4 | * | 33 | 3 |
| MYBL1 | 3 | 5 | * | 85 | 4 |
| CA12 | 15 | 18 | 48 | 19 | 5 |
| IGFBP4 | 42 | * | 30 | 7 | 6 |
| PMAIP1 | 20 | 27 | * | 14 | 7 |
| RAPGEFL1 | 163 | * | 1 | * | 8 |
| WWC1 | 171 | 32 | 8 | 102 | 9 |
| SGK3 | 2 | 19 | * | * | 10 |
| TUBB2B | * | * | 3 | * | 13 |
| STC2 | 10 | * | * | 15 | 14 |
| EGR3 | 143 | 2 | * | 215 | 15 |
| NRIP1 | 9 | * | * | 44 | 18 |
| SGK1 | 5 | * | * | 347 | 19 |
| AREG | 138 | 9 | 24 | 61 | 23 |
| MYC | 19 | 7 | * | 104 | 24 |
| HSPB8 | * | 3 | 56 | 62 | 25 |
| PGR | 153 | * | 2 | * | 29 |
| FADS1 | * | * | * | 5 | 30 |
| IL17RB | * | * | * | 4 | 39 |
| PLOD2 | 6 | * | * | 131 | 40 |
| AMD1 | 7 | * | * | * | 41 |
| RASGRP1 | 11 | 10 | * | * | 42 |
| TNP2 | * | * | 7 | * | 49 |
| MYB | 8 | * | * | 181 | 57 |
| HOXC4 | * | * | * | 9 | 65 |
| CHST8 | * | * | 6 | 128 | 91 |
| SERPINB9 | * | * | * | 3 | 113 |
| GP5 | * | * | 10 | * | 123 |
| FCGR2B | * | * | 4 | * | 141 |
| TGM2 | * | * | 5 | * | 153 |
| TFF1 | 313 | * | * | 8 | 210 |
| TFF2 | * | * | 9 | * | 217 |
| HOXC6 | * | * | * | 1 | 282 |
| PLAC1 | * | * | * | 2 | 290 |
The column gene includes a union of top 10 regulated genes from individual studies and meta-analysis. The ranks of these genes are indicated in the individual study and in the meta analysis studies. The asterisk * indicates that the gene was not included in the significant list (pfp ≤ 0.05).
Table 5.
Comparison of top 10 late down regulated genes
| Gene | M | gse3529 | gse3834 | gse4006 | gse4025 | gse6800 | gse8597 | gse9936 |
|---|---|---|---|---|---|---|---|---|
| IL17RB | 1 | * | * | 8 | 5 | * | 12 | 2 |
| TFF1 | 2 | * | * | 92 | 19 | * | 655 | 1 |
| CDK1 | 3 | * | 59 | 28 | 13 | * | 57 | 5 |
| MYBL1 | 4 | 1 | 6 | 31 | 174 | * | 1 | 162 |
| CDC20 | 5 | 43 | 3 | * | 15 | * | 310 | 3 |
| CXCL12 | 7 | 15 | 8 | 135 | 68 | * | 17 | 57 |
| PBK | 8 | 39 | 16 | * | 18 | * | 23 | 33 |
| RAB31 | 9 | 29 | * | 14 | 4 | * | 93 | 25 |
| KIF2C | 10 | * | 63 | * | 7 | * | 226 | 4 |
| PDZK1 | 419 | 2 | 25 | 57 | * | * | 414 | * |
| GREB1 | 19 | 3 | 1 | 1 | 194 | 1 | 62 | 323 |
| EGR3 | 184 | 4 | 42 | 41 | * | 4 | 19 | * |
| AREG | * | 5 | 7 | 38 | * | 21 | 46 | 719 |
| NPY1R | 57 | 6 | * | 13 | * | * | 187 | 96 |
| MSMB | 1650 | 7 | * | * | * | * | * | * |
| RASGRP1 | * | 8 | * | * | * | * | 1020 | * |
| FABP5 | * | 9 | * | * | * | 23 | 98 | * |
| CELSR2 | 136 | 10 | 33 | 89 | * | 26 | 313 | 402 |
| DHRS2 | 294 | * | 2 | 21 | * | * | 639 | 542 |
| PTTG1 | 29 | * | 4 | * | 77 | * | 346 | 14 |
| KIF20A | 18 | * | 5 | * | 17 | * | 304 | 7 |
| PLK1 | 74 | * | 9 | * | 45 | * | 389 | 44 |
| CENPA | 15 | 71 | 10 | * | 39 | * | 270 | 9 |
| GJA1 | * | * | * | 2 | * | * | * | * |
| WISP2 | 549 | * | 67 | 3 | * | * | 342 | * |
| ASCL1 | 704 | * | * | 4 | * | * | * | 400 |
| OLFML3 | 65 | * | * | 5 | 114 | * | * | 29 |
| EPB41L3 | 605 | * | * | 6 | * | * | * | 357 |
| PEG10 | 567 | * | * | 7 | * | * | * | 505 |
| PGR | 628 | * | * | 10 | * | * | 94 | * |
| PRIM1 | 12 | * | * | * | 1 | * | 56 | 15 |
| SLC7A5 | 107 | * | * | * | 2 | 24 | * | 132 |
| PLAC1 | 17 | * | * | 132 | 3 | * | 13 | 53 |
| TPD52L1 | 14 | 32 | * | 97 | 6 | 19 | 363 | 39 |
| KIF4A | 21 | * | * | * | 8 | * | 148 | 8 |
| TPX2 | 36 | * | 40 | * | 10 | * | 244 | 34 |
| SLC26A2 | 293 | * | 66 | 109 | * | 2 | 290 | * |
| PRSS23 | 101 | 21 | * | 61 | 61 | 3 | 675 | 246 |
| PTP4A1 | 709 | * | * | * | * | 5 | * | 966 |
| CA12 | 30 | 55 | * | 65 | 44 | 6 | 249 | 60 |
| HSPB8 | 1739 | 40 | 32 | * | * | 7 | * | * |
| TSKU | * | * | * | * | * | 9 | * | * |
| SGK3 | 93 | 22 | * | 23 | * | * | 2 | 778 |
| UGT2B15 | 112 | * | * | 26 | * | * | 3 | 227 |
| TMPRSS3 | 220 | 16 | * | 56 | * | 22 | 4 | * |
| PCP4 | 255 | * | * | * | * | * | 5 | 867 |
| MCM10 | 92 | 41 | * | * | 200 | * | 6 | 243 |
| DSCC1 | 132 | 62 | * | * | * | * | 7 | 453 |
| EXO1 | 338 | * | * | * | * | * | 8 | * |
| DTL | 53 | * | * | 79 | 261 | * | 9 | 95 |
| CENPI | 463 | * | * | * | * | * | 10 | * |
| DLGAP5 | 25 | * | 17 | * | 54 | * | 159 | 10 |
The column gene includes a union of top 10 regulated genes from individual studies and meta-analysis. The ranks of these genes are indicated in the individual study and in the meta analysis studies. The asterisk * indicates that the gene was not included in the significant list (pfp ≤ 0.05).
The top early genes from the meta-analysis include many well known direct targets of ERα and ERβ [3]. The most significant ones include TFF1, CCND1, IGFBP4, C3, ADORA1, GREB1, and MYC, which have been shown by candidate gene analysis to be estrogen regulated genes in breast cancer cell lines.
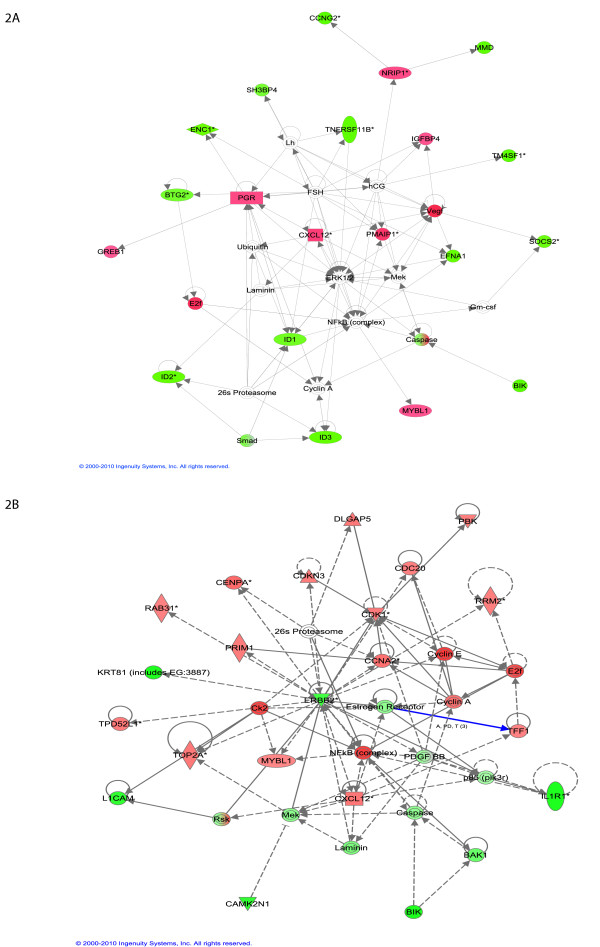
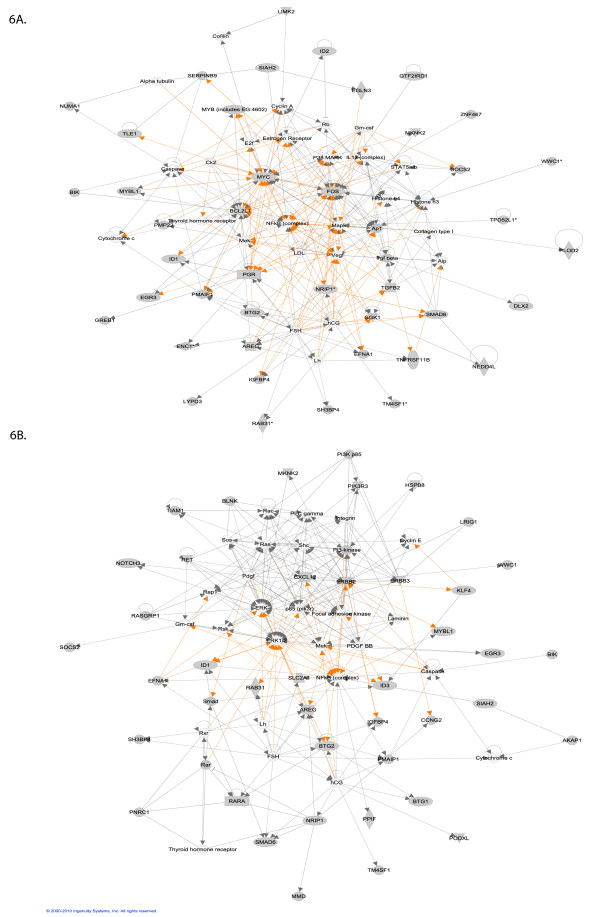
Moreover, the top genes from both the early and late time points form highly connected networks (Figure 2). These interactions show the functional relevance of our results, and point to the importance of networks in interpreting E2 signaling in MCF-7. We can already see in these data important differences between the early and late regulated genes, with the top early genes involved in cellular growth and proliferation. The early network notably shows up-regulation of the nuclear repressor protein NRIP1/RIP140[5], which is then involved in the down-regulation of cyclin G2 (CCNG2) and of monocyte-to-macrophage differentiation-associated protein (MMD) (Figure 2A). The late gene network is centered on ERBB2, a member of the epidermal growth factor (EGF) family of receptor tyrosine kinases, and one of the major molecular prognostic and predictive markers in breast cancer (Figure 2B). ERBB2 is known for its role in cell proliferation, growth, and apoptosis. This network also includes NF-kappaB, which plays a key role in the aggressiveness of breast cancer cells [23,24].
Figure 2.
Highly connected network of top 30 ranked genes at early time point (A). and late time point (B) as identified with IPA. Green node color indicates down regulation and red node color indicates up regulation. White color indicates nodes that are not included in the dataset but were assimilated into network by interaction with other molecules. Direct and indirect interactions between genes are denoted with solid and dashed lines respectively. The style of the arrows indicates specific molecular relationships and the directionality of the interaction (A acts on B). The shapes are indicative or the molecular class (i.e. protein family). Horizontal oval = transcription regulator, vertical diamond = enzyme, and circle = other.
Pathway analysis: Early time point
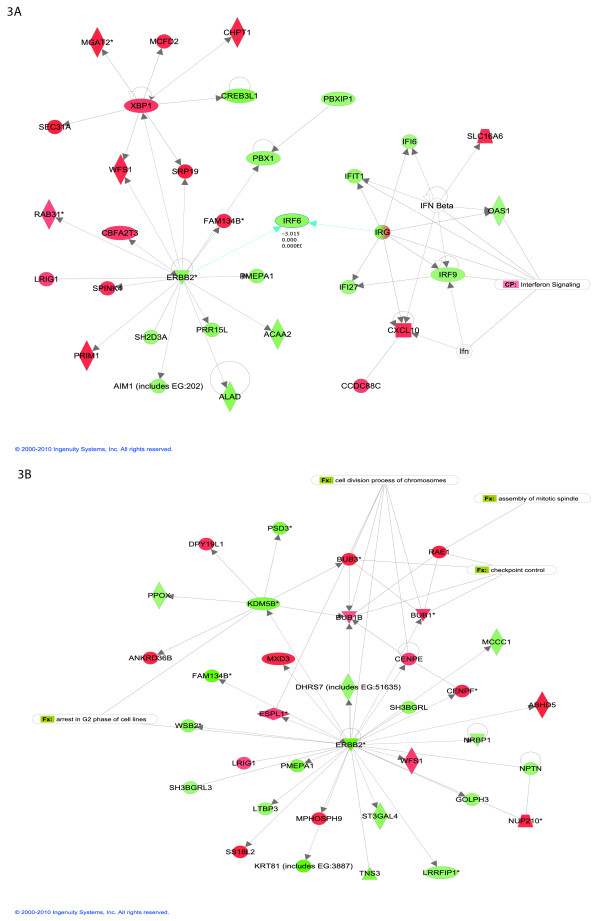
We have found 20 biological networks with a significant involvement of the genes differentially expressed at the early time point. Figure 3 shows the top network, which is consistent with the network found using the top 30 genes only: again, ERRB2 plays a central role. This network is annotated "infection mechanism, cellular growth and proliferation, genetic disorder". The interferon signaling pathway, which is involved in negative regulation of cellular proliferation and induction of cellular apoptosis, is down regulated. Of note, IRF6 is suggested to regulate epithelial cell differentiation, and several IRF family members are known to harbor tumor suppressive functions. We find IRF members IR6 and IRF9 to be down regulated in MCF-7, which adds more evidence to the hypothesis that down-regulation of the interferon signaling pathway might be important for uncontrolled cellular proliferation[25].
Figure 3.
Top networks with IPA analysis at early time point (A). and late time point (B). Green node color indicates down regulation, and red node color indicates up regulation. White color indicates nodes that are not included in the dataset but were assimilated into the network by interaction with other molecules. Direct and indirect interactions between genes are denoted with solid and dashed lines respectively. The style of the arrows indicates specific molecular relationships and the directionality of the interaction (A acts on B). The shapes are indicative or the molecular class (i.e. protein family). Horizontal oval = transcription regulator, vertical diamond = enzyme and circle = other.
The other top scoring networks at the early time point contain genes involved in cell growth, proliferation, DNA replication, protein amino acid metabolism, RNA and protein trafficking, cellular movement, and lipid metabolism. The merged image of the top three networks indicates three nodes centered on ERBB, cMYC and FOS, all key regulators of proliferative responses (see Additional File 2). In addition, several proteins involved in protein translation and metabolism are up regulated, including seven mitochondrial ribosomal proteins.
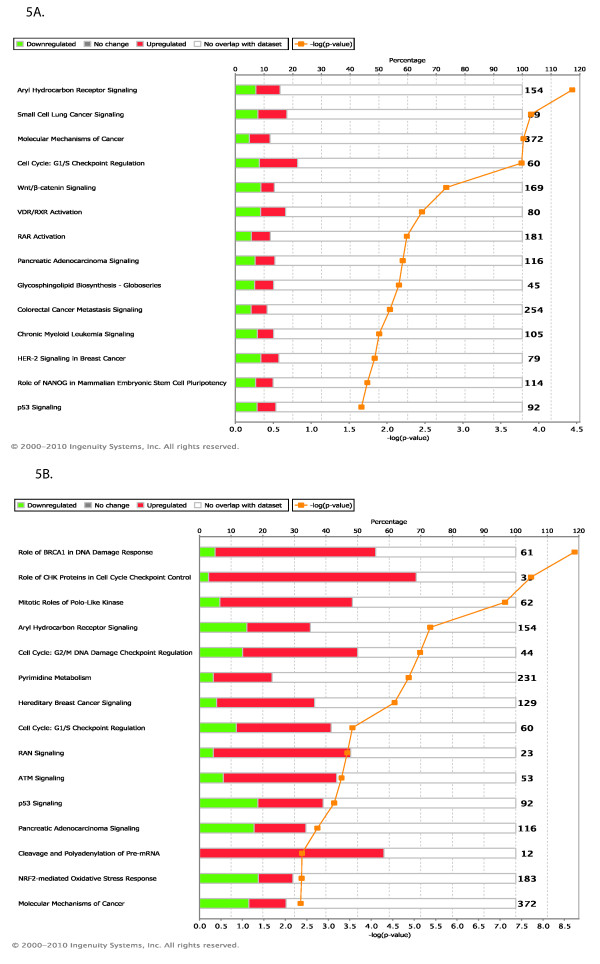
In addition to searching for relations amongst the differentially regulated genes, we can map them to very well established ("canonical") pathways. In this view, the early genes appear involved in signaling pathways such as wnt/b-catenin signaling, RAR activation and VDR activation (Figure 4A).
Figure 4.
Top Canonical pathways regulated. (A) Early time point (B) Late time point. Green color denotes the percentage of genes in the pathway that are down regulated and red color denotes the percentage of genes that are up regulated.
Pathway analysis: Late time point
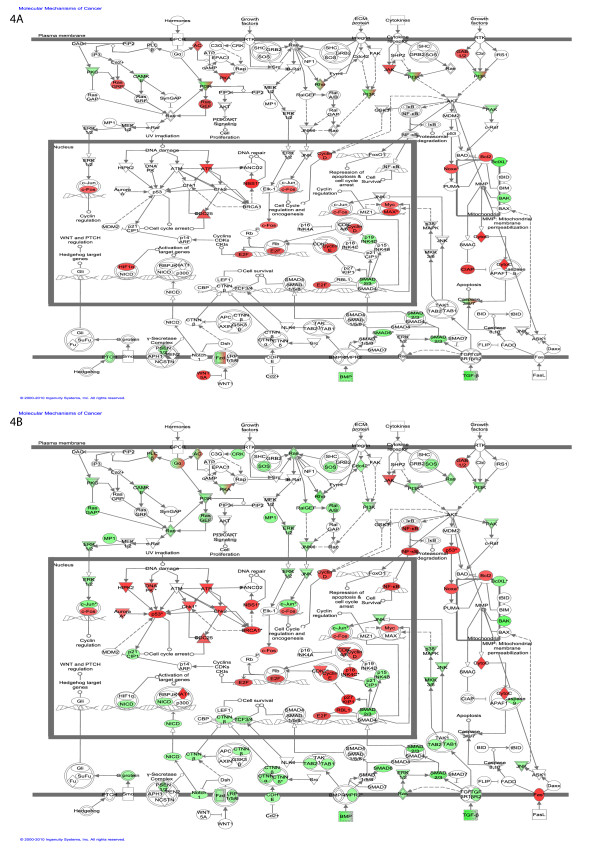
At the late time point the top network is annotated "cellular assembly and organization, cell cycle, DNA replication, recombination and repair". Figure 4 shows that the ERBB2 network is now involved in interaction with genes involved in cell division and cell cycle. The 5 top scoring gene networks share several common genes regulated at both time points, although they sometimes change in opposite directions. This similarity between time points indicates that most of the major gene expression changes start to take place as early as 3 hrs. This is clear comparing the early and late response of the genes that are involved in molecular mechanisms of cancer (Figure 5).
Figure 5.
Molecular mechanism of Cancer. (A) The genes regulated at the early time point. (B) The genes regulated at the late time point. Green node color indicates down regulation, and red node color indicates up regulation. White color indicates nodes that are not included in the dataset but were assimilated into network by interaction with other molecules. Direct and indirect interactions between genes are denoted with solid and dashed lines respectively. The style of the arrows indicates specific molecular relationships and the directionality of the interaction (A acts on B). The shapes are indicative or the molecular class (i.e. protein family). Horizontal oval = transcription regulator, vertical diamond = enzyme and circle = other.
Genes that are down regulated at the late time point also include a large proportion of structural and cell adhesion genes, such as collagens, keratins, and other intermediate filament proteins. Other down regulated genes that are potentially involved in the development of cancer include superoxide dismutase 3 (SOD3), cyclin D2 (ccnd2), transthyretin (Ttr), bone morphogenetic protein 2 (BMP2) and matrix metalloproteinase 2 (Mmp2).
Finally, the late genes can be mapped to canonical pathways involved in breast cancer, and cell cycle and division (Figure 4B).
Comparison with the GEMS dataset
A previous meta-analysis of E2 treated MCF cell lines was done by Ochsner et al. [26]. They used a parametric (weighted t-stats) method to obtain a gene expression meta-signature (GEMS) for E2-regulated gene expression in MCF-7 cells. We compared the pathway analysis for the top 50 down regulated and top 50 up regulated genes of the early time point of each study (Figure 6). Table 6 shows the list of central genes in each resulting network. Curiously, the GEMS derived network for early genes resembles more the networks derived from late genes by the Rank product method. Notably, it is centered on ERBB2 and ERK. Another difference is that the Rank product produces more highly connected genes which have been shown in the literature to be regulated by ERα, than GEMS. This includes ERα itself, which is not a top gene in the GEMS analysis. Also notable is the vascular endothelial growth factor (VEGF), which is central in the network derived from Rank product, but is not detected significantly in any individual study, nor by the GEMS analysis. VEGF expression is strongly induced as an immediate early response to E2 induction in vivo (uterine epithelial cells), mediated by ERα and hypoxia-inducible factor (HIF1α) [27].
Figure 6.
Merged network created from the top 100 (50 down regulated and 50 upregulated) differentially expressed genes using (A) Rank product algorithm, (B) the GEMS list.
Table 6.
Genes with most connections in networks from figure 6 and their number of connections
| Early probesets from Rankproduct | Early Probeset from GEMS | ||
|---|---|---|---|
| Gene | # of Connections | Gene | # of Connections |
| MYC | 31 | ERBB2 | 25 |
| FOS | 29 | ERK | 21 |
| BCL2L1 | 18 | CXCL2 | 16 |
| Mapk | 15 | NFkB complex | 19 |
| NFkB complex | 16 | Focal adhesion kinase | 11 |
| PGR | 11 | ||
| Estrogen Receptor | 8 | ||
| AP1 | 8 | ||
| NR1P1 | 7 | ||
Thus it seems that, while the GEMS analysis does detect relevant E2 regulated genes, it provides a less complete picture than the Rank product method does.
Discussion
We have identified 1206 probe sets and more than 10 pathways whose expression is significantly different between control and E2 treated MCF-7 cells 3-4 hrs after treatment, and 4193 probe sets and more than 20 pathways 24 hrs after treatment.
Although very little overlap exists on the gene level between the nine expression profiling studies, the meta analysis finds common functional processes and pathways. Table 3 shows that most of the genes identified in individual studies were also identified in the meta-analysis. The rank product algorithm returns a robust ranking for all the top genes, leading to higher reproducibility and increased specificity.
It should be noted that all the datasets we used were based on the Affymetrix platform. A recently available dataset [28], using Illumina Beadchip, provides very similar results to our analysis. Another very recent dataset [29], using GRO-seq, although produced with a different focus on transient early response to E2, also provides consistent results. Thus it does not seem that our results are platform-dependent.
We have used pathway analysis tools to understand the relationship between the regulated genes. Known targets of ERα, such as MYC, ERBB2, and ESRα, are central genes at early and late time points. This shows that the statistically significant genes are biologically relevant, and that our analysis captures key aspects of the underlying physiology.
Aryl hydrocarbon receptor (AhR) is a ligand activated transcription factor involved in xenobiotic metabolism and in mediating the toxic effects of dioxin-like compounds. Crosstalk has been observed between AhR and ER, specifically with respect to ER signaling [30]. ERα has been reported to have a positive role in AhR signaling. We find this pathway highly regulated at both time points.
The top canonical pathways regulated early are involved in cellular growth, development and proliferation, whereas those regulated late are involved in DNA replication, recombination and repair, cell cycle and cell death. These canonical pathways were not evident from any of the individual studies. Another difference between early and late response to E2 is early regulation of transcription regulators, vs. late regulation of kinases and transporters.
At the late time point we also see several genes common with the SV40T/t-antigen cancer signature identified by Deeb et al.[31] for human breast, prostate and lung carcinomas. These include genes encoding 10 centromere proteins, 8 cyclins, 15 cell division cyle proteins, 7 kinesin-like family proteins, 7 multiple minichromosome maintenance-deficient proteins, and other proliferation-related proteins and signal transduction proteins.
Conclusions
It is important to aggregate as much high-quality data as possible to minimize sources of bias influencing gene expression studies. The rank product methodology favors genes that are consistently top ranked among replicates. Therefore in the future other high quality datasets can be integrated into this work, with minimum effort. An interesting perspective will be to explore the predictive power for clinical outcome [32] of breast cancer, of the gene set from meta-analysis, to the gene sets from individual studies.
Methods
Meta-Analysis
The human genome U133A platform contain >22,000 probe sets and the human genome platform U133 Plus 2.0 array contains an additional >31,000 new probe sets, giving a total of >54,000 probe sets. For the meta-analysis, the probesets found in common across the Affymetrix U133 GeneChip family was used. There are 22,277 common probesets between the two U133a and U133 plus 2.0, that map to 13,186 genes.
The statistical significance of the results was evaluated by the non-parametric algorithm 'Rank products' [33], available as the 'RankProd' package at Bioconductor (http://www.bioconductor.org) [19]. This method detects genes that are consistently highly ranked in several replicated experiments, independently of their numerical intensities. This method ranks each feature within an experiment based on that features' score (e.g., Log expression values), and then combines these ranks, instead of combining the data or p-values. The results are provided in the form of P values defined as the probability that a given gene is ranked in the observed position by chance. A list of differentially regulated probe sets were selected based on the estimated percentage of false positive predictions (pfp)[33], which is equivalent to a false discovery rate. The pfp is calculated using a permutation based procedure (100,000 permutations were conducted). Genes with a pfp of less than 0.05 were selected for further investigation. As the rank-based procedure is non-parametric, it does not make assumptions about the model and the parameters from which the data came.
Evaluation of meta-analysis
We evaluated the performance of the Rank product algorithm using Correspondence At the Top (CAT) plots [17], which determine the proportion of genes in common between experiments as a function of list size. To generate CAT plots we use the lists of top genes for each of the study including the meta studies and plot proportion in common against the list size. We also used integration-drive discovery (IDD) to measure the number of extra genes identified by meta-analysis, compared with the union set of all individual studies at the same pfp threshold value.
Functional analysis
Functional classification analysis was done using the Ingenuity Pathway Analysis software (version 8.5). The "Core Analysis" was used, with a focus on canonical signaling pathways. Core Analysis allows for a rapid assessment and interpretation of large and small datasets in the context of biological processes, pathways, and molecular networks. In short, for a given function or pathway, the statistical significance of pathway enrichment is calculated using a right tailed Fisher's exact test based on the number of genes annotated, the number of genes represented in the input dataset, and the total number of genes being assessed in the experiment. A pathway was deemed significant if the p-value of enrichment was ≤ 0.01. The p-value is adjusted for multiple comparisons using FDR correction.
Abbreviations
ERα: Estrogen Receptor alpha; pfp: Percentage of False positive Predictions; IPA: Ingenuity Pathway Analysis; CAT: Correspondence At the Top.
Authors' contributions
VJ performed data analysis and writing the manuscript. MRR contributed to research design, writing and discussion of the results. Both authors read and approved the final manuscript.
Supplementary Material
Results of the meta-analysis. Ranking of genes following the meta-analysis, with meta-analysis scores and p-values.
Supplementary Figure: merged networks for early genes. Merged top three networks with IPA analysis at early time point. Legend as Figure 3.
Contributor Information
Vidhya Jagannathan, Email: vidhya.jagannathan@unil.ch.
Marc Robinson-Rechavi, Email: marc.robinson-rechavi@unil.ch.
Acknowledgements
We acknowledge funding from European program Crescendo and Etat de Vaud.
References
- Katzenellenbogen BS. Estrogen receptors: bioactivities and interactions with cell signaling pathways. Biol Reprod. 1996;54(2):287–293. doi: 10.1095/biolreprod54.2.287. [DOI] [PubMed] [Google Scholar]
- Anderson E. The role of oestrogen and progesterone receptors in human mammary development and tumorigenesis. Breast Cancer Res. 2002;4(5):197–201. doi: 10.1186/bcr452. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Welboren WJ, Sweep FC, Span PN, Stunnenberg HG. Genomic actions of estrogen receptor alpha: what are the targets and how are they regulated? Endocr Relat Cancer. 2009;16(4):1073–1089. doi: 10.1677/ERC-09-0086. [DOI] [PubMed] [Google Scholar]
- Welboren WJ, van Driel MA, Janssen-Megens EM, van Heeringen SJ, Sweep FC, Span PN, Stunnenberg HG. ChIP-Seq of ERalpha and RNA polymerase II defines genes differentially responding to ligands. EMBO J. 2009;28(10):1418–1428. doi: 10.1038/emboj.2009.88. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Carroll JS, Meyer CA, Song J, Li W, Geistlinger TR, Eeckhoute J, Brodsky AS, Keeton EK, Fertuck KC, Hall GF. et al. Genome-wide analysis of estrogen receptor binding sites. Nat Genet. 2006;38(11):1289–1297. doi: 10.1038/ng1901. [DOI] [PubMed] [Google Scholar]
- Lin CY, Vega VB, Thomsen JS, Zhang T, Kong SL, Xie M, Chiu KP, Lipovich L, Barnett DH, Stossi F. et al. Whole-genome cartography of estrogen receptor alpha binding sites. PLoS Genet. 2007;3(6):e87. doi: 10.1371/journal.pgen.0030087. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vyhlidal C, Samudio I, Kladde MP, Safe S. Transcriptional activation of transforming growth factor alpha by estradiol: requirement for both a GC-rich site and an estrogen response element half-site. J Mol Endocrinol. 2000;24(3):329–338. doi: 10.1677/jme.0.0240329. [DOI] [PubMed] [Google Scholar]
- Petz LN, Nardulli AM. Sp1 binding sites and an estrogen response element half-site are involved in regulation of the human progesterone receptor A promoter. Mol Endocrinol. 2000;14(7):972–985. doi: 10.1210/me.14.7.972. [DOI] [PubMed] [Google Scholar]
- DeNardo DG, Kim HT, Hilsenbeck S, Cuba V, Tsimelzon A, Brown PH. Global gene expression analysis of estrogen receptor transcription factor cross talk in breast cancer: identification of estrogen-induced/activator protein-1-dependent genes. Mol Endocrinol. 2005;19(2):362–378. doi: 10.1210/me.2004-0267. [DOI] [PubMed] [Google Scholar]
- Glidewell-Kenney C, Weiss J, Lee EJ, Pillai S, Ishikawa T, Ariazi EA, Jameson JL. ERE-independent ERalpha target genes differentially expressed in human breast tumors. Mol Cell Endocrinol. 2005;245(1-2):53–59. doi: 10.1016/j.mce.2005.10.003. [DOI] [PubMed] [Google Scholar]
- Kalaitzidis D, Gilmore TD. Transcription factor cross-talk: the estrogen receptor and NF-kappaB. Trends Endocrinol Metab. 2005;16(2):46–52. doi: 10.1016/j.tem.2005.01.004. [DOI] [PubMed] [Google Scholar]
- Frasor J, Danes JM, Komm B, Chang KC, Lyttle CR, Katzenellenbogen BS. Profiling of estrogen up- and down-regulated gene expression in human breast cancer cells: insights into gene networks and pathways underlying estrogenic control of proliferation and cell phenotype. Endocrinology. 2003;144(10):4562–4574. doi: 10.1210/en.2003-0567. [DOI] [PubMed] [Google Scholar]
- Lin CY, Strom A, Vega VB, Kong SL, Yeo AL, Thomsen JS, Chan WC, Doray B, Bangarusamy DK, Ramasamy A. et al. Discovery of estrogen receptor alpha target genes and response elements in breast tumor cells. Genome Biol. 2004;5(9):R66. doi: 10.1186/gb-2004-5-9-r66. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kininis M, Kraus WL. A global view of transcriptional regulation by nuclear receptors: gene expression, factor localization, and DNA sequence analysis. Nucl Recept Signal. 2008;6:e005. doi: 10.1621/nrs.06005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cheung E, Kraus WL. Genomic analyses of hormone signaling and gene regulation. Annu Rev Physiol. pp. 191–218. [DOI] [PubMed]
- Choi JK, Yu U, Kim S, Yoo OJ. Combining multiple microarray studies and modeling interstudy variation. Bioinformatics. 2003;19(Suppl 1):i84–90. doi: 10.1093/bioinformatics/btg1010. [DOI] [PubMed] [Google Scholar]
- Irizarry RA, Warren D, Spencer F, Kim IF, Biswal S, Frank BC, Gabrielson E, Garcia JG, Geoghegan J, Germino G. et al. Multiple-laboratory comparison of microarray platforms. Nat Methods. 2005;2(5):345–350. doi: 10.1038/nmeth756. [DOI] [PubMed] [Google Scholar]
- Hong F, Breitling R. A comparison of meta-analysis methods for detecting differentially expressed genes in microarray experiments. Bioinformatics. 2008;24(3):374–382. doi: 10.1093/bioinformatics/btm620. [DOI] [PubMed] [Google Scholar]
- Hong F, Breitling R, McEntee CW, Wittner BS, Nemhauser JL, Chory J. RankProd: a bioconductor package for detecting differentially expressed genes in meta-analysis. Bioinformatics. 2006;22(22):2825–2827. doi: 10.1093/bioinformatics/btl476. [DOI] [PubMed] [Google Scholar]
- Barrett T, Troup DB, Wilhite SE, Ledoux P, Rudnev D, Evangelista C, Kim IF, Soboleva A, Tomashevsky M, Edgar R. NCBI GEO: mining tens of millions of expression profiles--database and tools update. Nucleic Acids Res. 2007;35(Database issue):D760–765. doi: 10.1093/nar/gkl887. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Irizarry RA, Hobbs B, Collin F, Beazer-Barclay YD, Antonellis KJ, Scherf U, Speed TP. Exploration, normalization, and summaries of high density oligonucleotide array probe level data. Biostatistics. 2003;4(2):249–264. doi: 10.1093/biostatistics/4.2.249. [DOI] [PubMed] [Google Scholar]
- Wu Z, Irizarry R, Gentleman R, Martinez-Murillo F, Spencer F. A Model-Based Background Adjustment for Oligonucleotide Expression Arrays. Journal of the American Statistical Association. p. 909.
- Naugler WE, Karin M. NF-kappaB and cancer-identifying targets and mechanisms. Curr Opin Genet Dev. 2008;18(1):19–26. doi: 10.1016/j.gde.2008.01.020. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chua HL, Bhat-Nakshatri P, Clare SE, Morimiya A, Badve S, Nakshatri H. NF-kappaB represses E-cadherin expression and enhances epithelial to mesenchymal transition of mammary epithelial cells: potential involvement of ZEB-1 and ZEB-2. Oncogene. 2007;26(5):711–724. doi: 10.1038/sj.onc.1209808. [DOI] [PubMed] [Google Scholar]
- Bailey CM, Hendrix MJ. IRF6 in development and disease: a mediator of quiescence and differentiation. Cell Cycle. 2008;7(13):1925–1930. doi: 10.4161/cc.7.13.6221. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ochsner SA, Steffen DL, Hilsenbeck SG, Chen ES, Watkins C, McKenna NJ. GEMS (Gene Expression MetaSignatures), a Web resource for querying meta-analysis of expression microarray datasets: 17beta-estradiol in MCF-7 cells. Cancer Res. 2009;69(1):23–26. doi: 10.1158/0008-5472.CAN-08-3492. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Molitoris KH, Kazi AA, Koos RD. Inhibition of oxygen-induced hypoxia-inducible factor-1alpha degradation unmasks estradiol induction of vascular endothelial growth factor expression in ECC-1 cancer cells in vitro. Endocrinology. 2009;150(12):5405–5414. doi: 10.1210/en.2009-0884. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cicatiello L, Mutarelli M, Grober OMV, Paris O, Ferraro L, Ravo M, Tarallo R, Luo S, Schroth GP, Seifert M. et al. Estrogen Receptor [alpha] Controls a Gene Network in Luminal-Like Breast Cancer Cells Comprising Multiple Transcription Factors and MicroRNAs. The American Journal of Pathology. 2010;176(5):2113–2130. doi: 10.2353/ajpath.2010.090837. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hah N, Danko CG, Core L, Waterfall JJ, Siepel A, Lis JT, Kraus WL. A Rapid, Extensive, and Transient Transcriptional Response to Estrogen Signaling in Breast Cancer Cells. Cell. 2011;145(4):622–634. doi: 10.1016/j.cell.2011.03.042. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Matthews J, Gustafsson JA. Estrogen receptor and aryl hydrocarbon receptor signaling pathways. Nucl Recept Signal. 2006;4:e016. doi: 10.1621/nrs.04016. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Deeb KK, Michalowska AM, Yoon CY, Krummey SM, Hoenerhoff MJ, Kavanaugh C, Li MC, Demayo FJ, Linnoila I, Deng CX. et al. Identification of an integrated SV40 T/t-antigen cancer signature in aggressive human breast, prostate, and lung carcinomas with poor prognosis. Cancer Res. 2007;67(17):8065–8080. doi: 10.1158/0008-5472.CAN-07-1515. [DOI] [PubMed] [Google Scholar]
- Vanneschi L, Farinaccio A, Mauri G, Antoniotti M, Provero P, Giacobini M. A Comparison of Machine Learning Techniques for Survival Prediction in Breast Cancer. BioData Min. p. 12. [DOI] [PMC free article] [PubMed]
- Breitling R, Armengaud P, Amtmann A, Herzyk P. Rank products: a simple, yet powerful, new method to detect differentially regulated genes in replicated microarray experiments. FEBS Lett. 2004;573(1-3):83–92. doi: 10.1016/j.febslet.2004.07.055. [DOI] [PubMed] [Google Scholar]
- Creighton CJ, Cordero KE, Larios JM, Miller RS, Johnson MD, Chinnaiyan AM, Lippman ME, Rae JM. Genes regulated by estrogen in breast tumor cells in vitro are similarly regulated in vivo in tumor xenografts and human breast tumors. Genome Biol. 2006;7(4):R28. doi: 10.1186/gb-2006-7-4-r28. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chang EC, Charn TH, Park SH, Helferich WG, Komm B, Katzenellenbogen JA, Katzenellenbogen BS. Estrogen Receptors alpha and beta as determinants of gene expression: influence of ligand, dose, and chromatin binding. Mol Endocrinol. 2008;22(5):1032–1043. doi: 10.1210/me.2007-0356. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chang EC, Frasor J, Komm B, Katzenellenbogen BS. Impact of estrogen receptor beta on gene networks regulated by estrogen receptor alpha in breast cancer cells. Endocrinology. 2006;147(10):4831–4842. doi: 10.1210/en.2006-0563. [DOI] [PubMed] [Google Scholar]
- Gaube F, Wolfl S, Pusch L, Kroll TC, Hamburger M. Gene expression profiling reveals effects of Cimicifuga racemosa (L.) NUTT. (black cohosh) on the estrogen receptor positive human breast cancer cell line MCF-7. BMC Pharmacol. 2007;7:11. doi: 10.1186/1471-2210-7-11. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rae JM, Johnson MD, Scheys JO, Cordero KE, Larios JM, Lippman ME. GREB 1 is a critical regulator of hormone dependent breast cancer growth. Breast Cancer Res Treat. 2005;92(2):141–149. doi: 10.1007/s10549-005-1483-4. [DOI] [PubMed] [Google Scholar]
- Frasor J, Chang EC, Komm B, Lin CY, Vega VB, Liu ET, Miller LD, Smeds J, Bergh J, Katzenellenbogen BS. Gene expression preferentially regulated by tamoxifen in breast cancer cells and correlations with clinical outcome. Cancer Res. 2006;66(14):7334–7340. doi: 10.1158/0008-5472.CAN-05-4269. [DOI] [PubMed] [Google Scholar]
- Bourdeau V, Deschenes J, Laperriere D, Aid M, White JH, Mader S. Mechanisms of primary and secondary estrogen target gene regulation in breast cancer cells. Nucleic Acids Res. 2008;36(1):76–93. doi: 10.1093/nar/gkn369. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Results of the meta-analysis. Ranking of genes following the meta-analysis, with meta-analysis scores and p-values.
Supplementary Figure: merged networks for early genes. Merged top three networks with IPA analysis at early time point. Legend as Figure 3.