Abstract
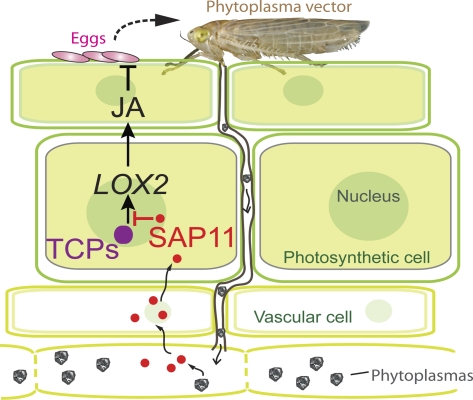
Phytoplasmas are insect-transmitted phytopathogenic bacteria that can alter plant morphology and the longevity and reproduction rates and behavior of their insect vectors. There are various examples of animal and plant parasites that alter the host phenotype to attract insect vectors, but it is unclear how these parasites accomplish this. We hypothesized that phytoplasmas produce effectors that modulate specific targets in their hosts leading to the changes in plant development and insect performance. Previously, we sequenced and mined the genome of Aster Yellows phytoplasma strain Witches’ Broom (AY-WB) and identified 56 candidate effectors. Here, we report that the secreted AY-WB protein 11 (SAP11) effector modulates plant defense responses to the advantage of the AY-WB insect vector Macrosteles quadrilineatus. SAP11 binds and destabilizes Arabidopsis CINCINNATA (CIN)-related TEOSINTE BRANCHED1, CYCLOIDEA, PROLIFERATING CELL FACTORS 1 and 2 (TCP) transcription factors, which control plant development and promote the expression of lipoxygenase (LOX) genes involved in jasmonate (JA) synthesis. Both the Arabidopsis SAP11 lines and AY-WB–infected plants produce less JA on wounding. Furthermore, the AY-WB insect vector produces more offspring on AY-WB–infected plants, SAP11 transgenic lines, and plants impaired in CIN-TCP and JA synthesis. Thus, SAP11-mediated destabilization of CIN-TCPs leads to the down-regulation of LOX2 expression and JA synthesis and an increase in M. quadrilineatus progeny. Phytoplasmas are obligate inhabitants of their plant host and insect vectors, in which the latter transmits AY-WB to a diverse range of plant species. This finding demonstrates that pathogen effectors can reach beyond the pathogen–host interface to modulate a third organism in the biological interaction.
Keywords: Arabidopsis development, insect herbivore, jasmonic acid, symbiosis, Hemiptera
Many (insect) vector-borne pathogens present a continuous hazard to human health and food production. The dispersal of these pathogens in nature largely depends on the frequency of interactions between the pathogens’ hosts and vectors, and the fitness of the pathogen is also linked to the reproductive success of its vectors. Accordingly, pathogens may manipulate their primary hosts to attract more insect vectors or to facilitate insect feeding on the host organisms. Indeed, there are spectacular examples of alterations of host phenotypic traits in response to vector-borne pathogen infection. These include malaria parasites, which make humans more attractive to mosquito vectors, particularly at the plasmodium transmissible stage (1); several vector-borne animal pathogens, which alter blood viscosity to facilitate blood ingestion by vectors (2); the fungus Puccinia monoica, which induces the formation of pseudoflowers to attract insects that enable the reproduction of the fungus (3); and phytoplasmas and viruses that improve plant host quality for their insect vectors (4, 5) and alter volatile emission of host plants to attract insect vectors (6, 7). It is well documented that pathogens produce effectors that modify specific processes in the host and alter the host–pathogen interface, often to the advantage of the pathogen (8, 9). For example, the Gram-negative plant pathogen Pseudomonas syringae secretes effectors into host cells that suppress the induction of pathogen-associated molecular pattern-triggered immune responses that act as a defense mechanism against the invading pathogen (10, 11), and Xanthomonas spp. produce transcription activator-like (TAL) effectors that target plant cell nuclei and activate the expression of host genes to facilitate their growth and possible spread in the host plant (12, 13). However, pathogen effectors that target specific processes in one host to attract insect vectors have not yet been identified.
Phytoplasmas are unculturable bacterial pathogens that belong to the class Mollicutes and are restricted to the cytoplasm of sieve cells of the plant phloem that is responsible for transport of carbohydrates and nutrients to plant sink tissues (reviewed in 14). Phytoplasmas depend on phloem-feeding insect vectors of the order Hemiptera, mostly leafhoppers, planthoppers, and psyllids, for transmission to plants (15). Thus, phytoplasmas are obligate pathogens of plants and insects that depend on both of their hosts for dispersal in nature. Nonetheless, Aster Yellows (AY) phytoplasmas, such as AY phytoplasma strain Witches’ Broom (AY-WB), have a wide plant host range affecting more than 80 dicot and monocot plant species worldwide (reviewed in 16), including numerous members of the Brassicaceae (17), such as Arabidopsis thaliana. In the United States and Canada, AY phytoplasmas are primarily vectored by Macrosteles quadrilineatus, which has a wide plant host range as well (18).
Phytoplasma infection can evoke dramatic changes in plant morphology, such as inducing the growth of multiple little leaves, the production and clustering of multiple stems and branches (witches’ brooms), retrograde metamorphosis of floral organs into leafy structures (phyllody), green coloration of nongreen flower tissues (virescence), growth of elongated stalks (bolting), and formation of bunchy fibrous secondary roots (19). These morphological changes in the plant may attract insect vectors that preferentially lay eggs in leaves and stems, and some in roots (14). Furthermore, M. quadrilineatus leafhoppers survive longer and produce more progeny on AY phytoplasma-infected plants (4), suggesting that the phytoplasmas down-regulate plant defense against the leafhopper. However, the two phytoplasma isolates (“bolt” and “severe”) involved in the study were field isolates that were not formally classified, and the mechanisms by which these bacteria impair host defenses are not yet understood.
We wished to elucidate the mechanism by which AY phytoplasma-infected plants positively affect the fecundity of M. quadrilineatus and have reasoned for some time that phytoplasmas induce phenotypic changes in plants through the production of effector proteins (14, 20–22). To identify the effectors involved in mediating these alterations in phenotype, we sequenced the AY-WB genome (20) and mined the genome for genes encoding candidate effector proteins based on the presence of an N-terminal signal peptide (22). Because phytoplasmas do not appear to possess type III secretion systems, it is likely that effector proteins are secreted through the Sec-dependent pathway similar to related Gram-positive firmicutes (23) and in line with the presence of a functional SecA-dependent secretion system in phytoplasmas (24, 25). We identified 56 secreted AY-WB proteins (SAPs) that are candidate effector proteins, including SAP11 as described here (22). The well-characterized AY phytoplasma strain AY-WB readily infects the model plant Arabidopsis. Infected plants exhibit many symptoms associated with exposure to this phytoplasma, including the production of green leaf-like flowers (phyllody) and witches’ brooms (26, 27). In a recent screen, we generated transgenic Arabidopsis plants for the majority of AY-WB effector genes leading to the identification of SAP54, which induces leafy flowers resembling the phyllody symptoms of AY-WB–infected Arabidopsis (27).
Here, we report that M. quadrilineatus produces more offspring when reared on AY-WB–infected Arabidopsis and that the AY-WB effector SAP11 is, in part, responsible for this increase in leafhopper performance. The mechanism involves SAP11-mediated destabilization of CINCINNATA (CIN)-TEOSINTE BRANCHED1, CYCLOIDEA, PROLIFERATING CELL FACTORS 1 and 2 (TCP) transcription factors, which are regulated by miR319 (28). The destabilization of CIN-TCPs results in the down-regulation of LIPOXYGENASE2 (LOX2) expression and jasmonate (JA) production, which are involved in the Arabidopsis defense response to M. quadrilineatus. Thus, SAP11 functions beyond the phytoplasma–plant interface to increase the progeny of the phytoplasma insect vector, which transmits the phytoplasma to other plants.
Results
Phytoplasma Infection of Arabidopsis Changes Plant Morphology and Increases Insect Performance.
M. quadrilineatus readily uses Arabidopsis as a feeding and reproductive host and can also inoculate it with AY-WB. Infected plants show severe symptoms, including witches’ brooms caused by an increased production of stems (Fig. 1 and SI Appendix, Fig. 1A), flowers that become leafy, and various other morphological changes indicating that AY-WB interferes with plant developmental pathways (27). Previously, it was shown that leafhopper fecundity was increased on AY bolt-infected plants but not AY severe-infected plants (4). We determined if AY-WB induces similar phenotypes in Arabidopsis. Indeed, M. quadrilineatus adults of various age ranges that were allowed access to the experimental Arabidopsis plants for a maximum of 4 d produced significantly more progeny on the AY-WB–infected plants compared with noninfected plants (Fig. 2A), with a cumulative increase of 61% in nymph numbers for all leafhopper age ranges combined [analysis of deviance (ANODE); F1, 106 = 254.17, P < 0.001]. M. quadrilineatus female survival rates did not significantly differ between AY-WB–infected and healthy plants (ANODE; F1, 107 = 0.33, P = 0.57; SI Appendix, Fig. 1B), indicating that the increased nymph numbers are likely caused by more fecund females. We also counted the number of eggs laid, hatch rate of eggs, and number of feeding/probing sites on Arabidopsis leaves. M. quadrilineatus laid more eggs on AY-WB–infected plants compared with healthy plants within each adult leafhopper age range (Fig. 2B) and for all age ranges combined (ANODE; F1, 68 = 372.06, P < 0.001), whereas egg-hatching rates did not significantly differ (ANODE; F1, 68 =1.97, P = 0.165; SI Appendix, Fig. 1C). The number of leafhopper-feeding/probing sites was slightly but significantly reduced on AY-WB–infected plants (ANODE; F1, 22 = 30.22, P < 0.001; SI Appendix, Fig. 1D). Thus, an increase in leafhopper egg-laying activity is most likely responsible for the higher leafhopper nymph production.
Fig. 1.
AY-WB infection alters Arabidopsis morphology. (A) AY-WB–infected A. thaliana Col-0 plants are smaller and bushier (Right) than similar aged healthy plants (Left). (B) Close-up view of AY-WB–infected plant in A illustrating the witches’ broom symptoms. (C) Top view of the plants shown in A. Plants were grown in 10-h/14-h light/dark at 22 °C; at 4 wk of age, they were transferred to 16-h/8-h light/dark at 23 °C/20 °C and exposed to two noncarrier (healthy plants) or AY-WB carrier (infected plants) M. quadrilineatus for 1 wk. Photographs were taken when plants were 8.5 wk old. (Scale bars = 1 cm.)
Fig. 2.
M. quadrilineatus leafhoppers are more fecund on AY-WB–infected than healthy Arabidopsis Col-0 plants. Leafhoppers produce more nymphs (A) and lay more eggs (B) on AY-WB–infected plants. AY-WB–infected and healthy plants were obtained by exposing 3-wk-old plants to three AY-WB carrier and noncarrier M. quadrilineatus males, respectively, for 1 wk. Two weeks later, each plant received eight adult females and two adult males of different age ranges for 4 d (x axes). Numbers of eggs were counted 1 d after and numbers of nymphs were counted 15 d after removal of the adult insects (y axes). Each experiment included three plants per age range per treatment, and the experiments were repeated three times (A) or two times (B). Columns show the mean numbers of nymphs (A) and eggs (B) per plant of all biological replicates for each leafhopper age range. (Scale bars = SE.) *P < 0.05 when two treatments were compared at each adult age (t probabilities calculated within GLM).
Because M. quadrilineatus can acquire AY-WB from infected plants within the 4-d exposure time, we also determined whether the cause of the nymph production increase was attributable to AY-WB infection of the plant host or leafhopper vector. Leafhoppers continuously exposed to AY-WB–infected plants before transfer to the experimental healthy plants did not show an increase in cumulative nymph production on these healthy plants for up to 6 d (ANODE; F2, 30 = 0.66, P = 0.526; SI Appendix, Fig. 1E). Leafhoppers exposed to AY-WB–infected plants for 2 d before transfer to the experimental healthy plants increased nymph production on healthy plants from 10 d on (ANODE; F2, 84 = 10.68, P < 0.001; t probability = 0.008; SI Appendix, Fig. 1E) at the time leafhoppers start to transmit AY-WB to plants. Together, these data indicate that the increased leafhopper nymph production and oviposition phenotype during the 4-d exposure of healthy insects to AY-WB–infected plants in the experiment shown in Fig. 2 are the result of changes that occur during AY-WB infection of the plant and are unlikely the result of AY-WB presence in the insect vector itself.
In summary, these results demonstrate that AY-WB infection induced various phenotypic alterations in Arabidopsis (i.e., it changes plant morphology and increases fecundity of its leafhopper vector M. quadrilineatus). Because Arabidopsis is a model organism in which plant development and defense responses against various pathogens and insects have been extensively investigated, we decided to use Arabidopsis to elucidate the molecular mechanisms by which AY-WB alters the M. quadrilineatus–plant interaction.
Expression of AY-WB Phytoplasma Effector SAP11 Changes Arabidopsis Morphology.
Previously, we identified 56 SAPs, which are candidate effector proteins (22). To reveal the function of each effector candidate, we generated stable transgenic Arabidopsis lines that express the effector candidate genes under control of the Cauliflower mosaic virus 35S promoter and identified three effector proteins that induce clear morphological changes in Arabidopsis (27). Of these, SAP11 was previously shown to target plant cell nuclei in transient expression assays and was detected beyond the phloem in nuclei of mesophyll and trichome cells in AY-WB–infected plants (22). Hence, we further investigated SAP11 action. Arabidopsis Col-0 lines expressing the SAP11 sequence (SI Appendix, Fig. 2A) corresponding to the 9-kDa mature protein (without signal peptide) had smaller rosettes (Fig. 3A), severely crinkled leaves (Fig. 3B), a bushy appearance (Fig. 3C), and crinkled siliques (Fig. 3D). The bushy phenotype of the 35S::SAP11 lines is likely derived from an increase in the number of stems compared with Col-0 (ANOVA; F3, 12 = 24.31, P < 0.001; SI Appendix, Fig. 2B), because the 35S::SAP11 lines and Col-0 did not significantly differ in the lengths of stems (ANOVA; F3, 12 = 0.64, P = 0.604; SI Appendix, Fig. 2C) and the number of branches on the main bolt (ANOVA; F3, 12 = 0.67, P = 0.5884; SI Appendix, Fig. 2D) at the time when most of the siliques had matured. The 35S::SAP11 line 5 produced lower amounts of SAP11 than the two other lines (Fig. 3E), coinciding with the least severe phenotype exhibited by line 5 (Fig. 3 A–D).
Fig. 3.
SAP11 induces crinkled leaves and siliques, and it increases stem number in Arabidopsis. (A) Rosettes featuring crinkled leaves of 6-wk-old homozygous 35S::SAP11 lines 4, 5, and 7 grown in short-day (8-h/16-h light/dark) conditions. (B) Representative leaves taken from 10-wk-old plants shown in A. (C) Homozygous 10-wk-old 35S::SAP11 lines 4, 5, and 7 produce more stems (SI Appendix, Fig. 2 B–D). Plants were grown in long-day (16-h/8-h light/dark) conditions. (D) SAP11 lines have crinkled siliques (taken from plants shown in C). (E) Western blot of protein extracts of 35S::SAP11 line 4, 5, and 7 transgenic plants probed with anti-SAP11. (Scale bars = 1 cm.) Coomassie-stained Ribulose-1,5-bisphosphate carboxylase oxygenase large subunit in a duplicated gel serves as a loading control.
To reveal how SAP11 induces the morphological alterations in Arabidopsis, a yeast two-hybrid screen was conducted at Hybrigenics Services SAS using the company's ULTImate Y2H service. This identified CIN-related TCP transcription factors 2 and 13 as SAP11 interactors (SI Appendix, Fig. 3A and Table 1). TCP transcription factors are conserved among plants (29, 30) and regulate various aspects of plant development (28, 31–33). Arabidopsis class I TCP members promote cell proliferation in leaves, whilst a subset of class II members, which are homologs of Antirrhinum CINCINNATA (31), negatively regulate leaf growth and positively regulate leaf maturation and senescence (32, 33). The balance between the two classes of TCPs determines leaf and plant morphology (32). We predicted that SAP11 may interact with multiple TCPs, because the 11 yeast two-hybrid screen prey fragments share the basic helix–loop–helix domain that is conserved among TCPs and is involved in TCP dimerization and DNA binding (29) (SI Appendix, Fig. 3A).
SAP11 Binds to and Destabilizes Arabidopsis TCP Transcription Factors.
pTRBO is a Tobacco mosaic virus-based expression vector that allows for higher levels of in planta protein production compared with the 35S constructs (34), enabling the detection of interactions between unstable (low-abundance) proteins, such as TCPs. Therefore, we used the pTRBO vectors to conduct coimmunoprecipitation (co-IP) assays of SAP11 with TCP2, TCP13, TCP4, and TCP7. TCP4 is another Arabidopsis class II CIN-TCP, whereas TCP7 belongs to class I (29, 32). Nicotiana benthamiana leaves were infiltrated with pTRBO::FLAG-SAP11 or pTRBO::FLAG-RFP alongside one of the 35S::3xmyc-TCP2, TCP13, TCP4, and TCP7 constructs. FLAG-SAP11 coimmunoprecipitated with proteins corresponding to full-length 3× myc-TCP2, TCP4, and TCP7 (SI Appendix, Fig. 3 B and C). We did not successfully coimmunoprecipitate 3× myc-TCP13 with FLAG-SAP11. This may have been caused by the complete disappearance of TCP13 in the presence of SAP11 (as discussed below). Thus, the co-IP experiments confirmed the yeast two-hybrid screen data that SAP11 interacts with TCP2 and revealed that SAP11 also interacts with another CIN-TCP, TCP4, and class I TCP7.
To investigate how SAP11 may affect the TCPs, we conducted transient coexpression analyses in N. benthamiana leaves of GFP-SAP11 and 3× myc-TCPs under control of the 35S promoter. All CIN-TCPs were absent or decreased in abundance in the presence of GFP-SAP11 compared with GFP alone, whereas the abundance of class I TCP7 was similar between the GFP-SAP11 and GFP treatments (Fig. 4). Each experiment was repeated 9 times for TCP2, TCP4, TCP13, and TCP7 and twice for the other TCPs, and we constantly observed destabilization of CIN-TCPs but not TCP7. Similar results were achieved with the pTRBO expression system, in which 3× myc-TCP2 and TCP13 were less abundant in the presence of FLAG-SAP11 compared with FLAG-RFP, whereas 3× myc-TCP7 abundance was not visibly different between FLAG-SAP11 and FLAG-RFP presence (SI Appendix, Fig. 3D). The FLAG-SAP11–mediated reductions of 3× myc-TCP4 were more variable in the pTBRO experiments, possibly because of the high expression levels of TCPs in these experiments. Nonetheless, in the 12 times this experiment was repeated, 3× myc-TCP4 was less abundant 6 times, completely absent 1 time, and equally abundant 5 times in the FLAG-SAP11 treatment compared with the FLAG-RFP control. Thus, together, the coexpression analyses indicated that SAP11 destabilizes all the class II CIN-TCPs but not class I TCP7.
Fig. 4.
SAP11 destabilizes CIN-TCP transcription factors. The abundance of the eight class II CIN-TCP members (TCP2, TCP4, TCP13, TCP3, TCP5, TCP10, TCP17, and TCP24) is reduced in the presence of GFP-SAP11 compared with GFP, whereas the abundance of class I TCP7 was not reduced in the presence of GFP-SAP11. Molecular weight markers (kDa) are indicated to the left of the blots.
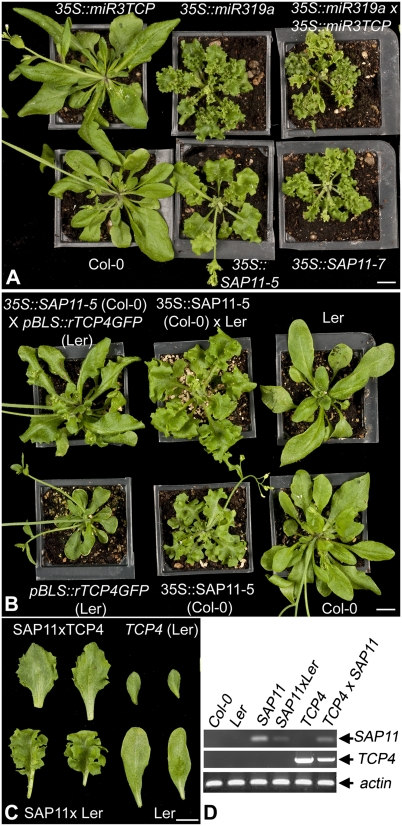
To obtain genetic evidence for TCP destabilization by SAP11, we made use of existing miR319a, miR-3TCP, and octuple cin-tcp (miR319a × miR-3TCP) Arabidopsis overexpression lines (32), in which miR319 negatively regulates the expression of five CIN-TCP genes (TCP2, TCP3, TCP4, TCP10, and TCP24) (28) and the artificial miR-3TCP negatively regulates the other three CIN-TCP genes (TCP5, TCP13, and TCP17) (32). It was previously shown that the miR-3TCP lines and Arabidopsis mutants in individual tcp genes, such as tcp2 and tcp13, have less obvious changes in leaf morphology compared with miR319a and the octuple cin-tcp lines, which exhibit deeply lobed and highly crinkled leaves, suggesting functional redundancy among the class II TCPs (32, 33). The 35S::SAP11, 35S::miR319a, and octuple cin-tcp plants had similar morphologies, and the 35S::SAP11-7 line resembled the octuple cin-tcp plants more closely than the 35S::SAP11-5 line did (Fig. 5A), consistent with higher SAP11 abundance in 35S::SAP11-7 plants compared with 35S::SAP11-5 plants (Fig. 3E). We also crossed 35S::SAP11-5 (Col-0) with a pBLS::rTCP4GFP (Ler) line, which overexpresses a miR319a-resistant TCP4 (rTCP4) fused at the C terminus to GFP under control of the postmaturation phase promoter BLS (32). The crinkled leaf phenotypes of 35S::SAP11-5 (Col-0) and 35S::SAP11-5 (Col-0) × Ler (control) plants were visibly reduced in the 35S::SAP11-5 (Col-0) × pBLS::rTCP4GFP (Ler) plants (Fig. 5 B–D), indicating that the SAP11-mediated destabilization of TCPs can be suppressed by overexpression of rTCP4GFP. Thus, the SAP11 transgenic lines phenocopy the octuple cin-tcp lines.
Fig. 5.
Genetic evidence for the SAP11-mediated stabilization of CIN-TCPs. (A) 35S::SAP11, 35S::miR319a, and 35S::miR319a × 35S::miR3TCP Arabidopsis Col-0 plants exhibit similar leaf crinkling phenotypes. (B) Overexpression of miR319-resistant TCP4GFP (rTCP4GFP) by BLS promoter reduces the crinkled leaf phenotype of 35S::SAP11 line 5. Because pBLS::rTCP4GFP has Ler background, 35S::SAP11 line 5 crossed with Ler WT is shown as a control. (C) Two representative leaves of plants shown in B. (D) PCRs demonstrating the presence of 35S::SAP11 and/or pBLS::rTCP4-GFP constructs in plants shown in B. (Scale bars = 1 cm.)
JA Synthesis Is Down-Regulated in SAP11 Transgenic Lines and AY-WB– Infected Arabidopsis.
CIN-TCPs positively regulate plant senescence, and this is partly achieved through the up-regulation of LOX2 by TCP4 binding to LOX2 promoter sites (33). LOX2 encodes a lipoxygenase that mediates the first step of the JA synthesis pathway by converting α-linolenic acid (α-LeA; 18:3) into 13-hydroperoxy-9,11,15-octadecatrienoic acid (13-HPOT) (35) and is induced by wounding and infestation of non–AY-WB carrier M. quadrilineatus (SI Appendix, Fig. 4A). We found that the expression levels of LOX2 were considerably lower in the 35S::SAP11 lines vs. WT Col-0 before wounding (nontreated) and 60 min after wounding (Fig. 6A). The 35S::SAP11 plants also produce less JA 60 min after wounding than WT Col-0 plants (ANOVA; F3, 11 = 20.42, P < 0.001; Fig. 6B). A 24-h time course of JA quantification on wounding indicated that 35S::SAP11 line 7 accumulates significantly less JA compared with Col-0 over time (Fig. 6C). Thus, the SAP11 transgenic lines have reduced LOX2 expression and JA production.
Fig. 6.
SAP11 and AY-WB–infected plants produce less JA. (A) LOX2 expression is down-regulated in nontreated and wounded 35S::SAP11 lines. Gene expression was assessed 60 min after wounding by quantitative RT-PCR in which the U-box housekeeping gene AT5G15400 transcript was used as a reference gene (58). Columns present data and SEs (bars) from three biological replicates. *P < 0.05 compared with Col-0 (Student t test). (B) SAP11 lines produce less JA. Columns show mean of HPLC results at 60 min after wounding from three biological replicates. (Scale bars = SE.) *P < 0.05 compared with Col-0 (ANOVA and Tukey's multiple comparison test). (C) 35S::SAP11 line 7 produces less JA over time. JA accumulation was measured at 0, 1.5, 6, and 24 h after wounding. The means and SEs of three biological replicates are presented. *P < 0.05 compared with corresponding Col-0 (Student t test). (D) AY-WB–infected Arabidopsis plants produce less JA. Four-week-old Arabidopsis plants were infested with AY-WB carrier or noncarrier M. quadrilineatus males for 1 wk. Insects were removed, and JA production in young and old leaves was assessed 3 wk later at 90 min after wounding (SI Appendix, Fig. 4 B and C). NT, nontreated mature leaves. Columns show means and SEs of three biological replicates. *P < 0.05 compared with corresponding healthy control (Student t test).
We also assessed JA production in AY-WB–infected plants. JA accumulation was quantified in young developing leaves and fully expanded mature (old) leaves at 1, 2, and 3 wk after removal of AY-WB carrier and noncarrier leafhoppers. AY-WB symptoms, such as elongated leaf shape with long petioles and light green color, started to appear 1 wk after removal of the AY-WB carrier insects in young leaves, and these symptoms became more severe over time (27). JA accumulation was reduced in symptomatic young leaves of AY-WB–infected plants compared with healthy plants with ∼27%, 50%, and 58% less JA accumulation at 1, 2, and 3 wk after insect removal, respectively (Fig. 6D and SI Appendix, Fig. 4 B and C). JA accumulation in old leaves was consistently lower than in young leaves. JA levels in old leaves did not differ between infected and noninfected plants at weeks 1 and 3 but were slightly increased in infected vs. noninfected plants at 2 wk on insect removal (Fig. 6D and SI Appendix, Fig. 4 B and C). Thus, AY-WB–infected plants produce overall less JA predominantly in young leaves.
M. quadrilineatus Produces More Progeny on SAP11 Lines and cin-tcp and lox2-Silenced Plants.
In Nicotiana attenuata, NaLOX3 is involved in JA production in response to wounding and insect feeding (36) and NaLOX3-silenced N. attenuata plants become hosts for Empoasca leafhoppers when planted in their natural habitat (37). We showed that leafhoppers produce more progeny on AY-WB–infected plants (Fig. 2A). Hence, we determined if M. quadrilineatus has increased nymph production and oviposition on 35S::SAP11, CIN-TCPs, and LOX2-silenced Arabidopsis lines. Nymph production on the 35S::SAP11-5 and 35S::SAP11-7 lines was higher compared with Col-0 for all adult leafhopper age ranges (Fig. 7A), with cumulative 34% and 40% increases, respectively, in nymph numbers for all age ranges combined (ANODE; F2, 117 = 41.64, P < 0.001) and no significant difference in nymph numbers between the two 35S:: SAP11 lines (ANODE; F1, 79 = 1.31, P = 0.256). Leafhoppers also laid more eggs on the two 35S::SAP11 lines (ANODE; F2, 106 = 76.09, P < 0.001; SI Appendix, Fig. 5A). The hatch rates of the eggs were not significantly different (ANODE; F2, 107 = 0.89, P = 0.414; SI Appendix, Fig. 5B), whereas the numbers of feeding/probing sites were reduced slightly in the two 35S::SAP11 lines compared with Col-0 (ANODE; F2, 25 = 14.1, P < 0.001; SI Appendix, Fig. 5C). Although the M. quadrilineatus adult female survival rates were not significantly different for the majority of leafhopper age ranges (SI Appendix, Fig. 5D), the cumulative survival rates for all age ranges combined were significantly different between Col-0 and the SAP11 lines (ANODE; F2, 118 = 18.69, P < 0.001). However, the female survival rates did not statistically influence leafhopper fecundity (ANODE; F1, 117 = 1.3, P = 0.256) or oviposition (ANODE; F1, 106 = 1.11, P = 0.295), indicating that an increase in leafhopper egg-laying activity is most likely responsible for the higher leafhopper oviposition and subsequent nymph production.
Fig. 7.
M. quadrilineatus produces more progeny on SAP11 transgenic plants, TCP-silenced plants, and JA-deficient plants. (A–D) Seven-week-old plants were exposed to eight adult females and two adult males of different age ranges for 4 d (x axes). Numbers of nymphs on each plant were counted 15 d after removal of the adult insects. Each experiment included three plants per age range per treatment, and the experiments were repeated two times. (A, C, and D) Columns show the mean numbers of nymphs per plant of all biological replicates for each leafhopper age range. *P < 0.05 when lines were compared at each adult age (t probabilities calculated within GLM). (B) Columns show the mean numbers of nymphs per plant produced by adult leafhoppers of four different age ranges (2–6 d, 6–10 d, 10–14 d, and 14–18 d) obtained from two independent experiments. Different letters above the columns (a–c) indicate statistical difference (P < 0.05) shown by t probabilities calculated within the GLM. (Scale bars = SE.)
M. quadrilineatus also produced more nymphs on the 35S::miR319a, 35S::miR3TCP, and octuple cin-tcp lines (Fig. 7B). The leafhopper fecundity increase was lower on the 35S::miR319a and miR-3TCP plants than on the octuple cin-tcp line. Furthermore, the level of fecundity increase on the octuple cin-tcp line (36%) was comparable to that of the SAP11 lines (34–40%). This is consistent with our finding that SAP11 destabilizes all eight class II CIN-TCPs.
Leafhopper adults produced 78% more progeny on the LOX2-silenced line (35S::LOX2, overexpression of LOX2 induces silencing of the endogenous LOX2) (38) compared with the control line, in which LOX2 is not silenced (antisense strand of LOX2 is expressed) (38) (ANODE; F1, 23 = 188.7, P < 0.001; Fig. 7C) and laid more eggs on the LOX2-silenced line (ANODE; F1, 23 = 186.22, P < 0.001; SI Appendix, Fig. 6A), whereas the egg-hatching rate increased only slightly (SI Appendix, Fig. 6B) on 35S::LOX2 plants (ANODE; F1, 24 = 21.91, P < 0.001). There were no significant differences in number of feeding/probing sites (ANODE; F1, 14 = 0.87, P = 0.366; SI Appendix, Fig. 6C) and the cumulative M. quadrilineatus adult female survival rates (ANODE; F1, 24 = 0.04, P = 0.853; SI Appendix, Fig. 6D) between 35S::LOX2 and the control lines. The quantification of LOX2 expression and JA production of 35S::LOX2 and the control line after wounding confirmed silencing of LOX2 in the 35S::LOX2 line (SI Appendix, Fig. 6 E and F).
LOX2 mediates the first step of the JA synthesis pathway. Hence, we determined if leafhopper reproduction is also affected on the Arabidopsis Col-0 JASMONATE RESISTANT 1 (jar1) mutant, because JAR1 acts further downstream of LOX2; it conjugates isoleucine (Ile) to JA producing JA-Ile, which is the active signaling molecule of the JA signaling pathway (39). Nymph numbers were increased at a cumulative 33% for all adult age ranges combined on Arabidopsis jar1 mutants compared with Col-0 (ANODE; F1, 21 = 56.76, P < 0.001; Fig. 7D). Although the overall M. quadrilineatus adult female survival rate was slightly but significantly increased in jar1 compared with Col-0 (ANODE; F1, 22 = 16.89, P < 0.001; SI Appendix, Fig. 6G), it did not statistically influence the increase of fecundity observed (ANODE; F1, 21 = 0.67, P = 0.422).
Altogether, our results indicate that the SAP11-mediated destabilization of CIN-TCPs leads to the down-regulation of LOX2 expression and JA synthesis and an increase in M. quadrilineatus progeny.
Discussion
AY-WB was originally isolated from infected lettuce fields in North America, and it induces characteristic symptoms, such as witches’ broom, in this host and in China aster (40). Beanland et al. (4) previously demonstrated that the leafhopper M. quadrilineatus survives longer and produces more progeny on aster plants infected with the phytoplasma isolate bolt; however, this field isolate was not formally classified within Candidatus (Ca.) phytoplasma based on 16S rDNA sequence information, nor was a mechanism identified by which the phytoplasmas might positively influence leafhopper fitness. In this work, we report that Arabidopsis Col-0 plants infected with the well-characterized strain AY-WB show witches’ broom symptoms and support the increased nymph production of M. quadrilineatus. In addition, M. quadrilineatus laid more eggs on AY-WB–infected plants than on noninfected plants, whereas egg-hatching rates and female survival rates were not affected, indicating that increased oviposition is predominantly responsible for the higher M. quadrilineatus progeny yield from infected plants. Numbers of M. quadrilineatus feeding/probing sites were reduced on AY-WB–infected plants compared with healthy plants. The leafhoppers may spend relatively more time feeding at the same location or overall spend less time feeding while they are laying eggs. Based on these data, we determined that the AY-WB Arabidopsis experimental system is an appropriate platform to enable further investigations into the mechanism(s) by which AY-WB effectors interfere with plant host development and leafhopper resistance.
An increase in leafhopper progeny on AY-WB–infected plants is likely to enhance AY-WB fitness in nature (SI Appendix, Fig. 7). M. quadrilineatus nymph production increased by 61% on AY-WB–infected vs. healthy plants (Fig. 2A), and this increased fecundity starts at the time of or shortly after AY-WB transmission to plants (SI Appendix, Fig. 1E). Following oviposition by the female, it takes ∼10 d for the nymphs to hatch from the eggs, and this is also the approximate time required for AY-WB to spread systemically throughout the plant host (SI Appendix, Fig. 7). Hence, the newly hatched nymphs have a high chance of acquiring the phytoplasmas immediately on feeding. We have observed that M. quadrilineatus can acquire AY-WB during a 2-d acquisition period on infected plants and can transmit AY-WB to healthy plants 10 d later (SI Appendix, Fig. 1E). Thus, systemic AY-WB infection of the leafhoppers is achieved as the insects are either fourth or fifth instar nymphs or adults, all of which are capable of migrating to uninfected plant hosts and transmitting the phytoplasma (SI Appendix, Fig. 7). Hence, a 61% increase in nymph production on AY-WB–infected plants probably leads to a similar increase in AY-WB transmission to other plants if all leafhoppers migrate to uninfected plants (SI Appendix, Fig. 7). The latter is likely, because M. quadrilineatus is a highly mobile and opportunistic species; it migrates annually from South America to North America, including Canada (41), and both this leafhopper species and AY phytoplasmas use wide ranges of plant species as feeding and reproductive hosts (18). Based on the above, it is expected that phytoplasmas that can enhance leafhopper oviposition and nymph production rates will have a competitive advantage over those phytoplasmas that cannot. Beanland et al. (4) found that isolate bolt increased leafhopper reproduction, whereas isolate severe did not. Hence, bolt would be expected to have a competitive advantage over severe. However, lettuce plants in the field can carry multiple AY phytoplasma strains (40); in this case, severe and probably other phytoplasmas would benefit from the increase in leafhopper reproduction rates induced by bolt.
We found that nymph production rates increased by 30–40% on 35S::SAP11 transgenic Arabidopsis plants. Because nymph production is increased by 61% on AY-WB–infected plants, it is likely that other AY-WB effectors in addition to SAP11 modulate leafhopper-plant interactions. SAP11 lies within a potential mobile unit (PMU)-like region in the AY-WB chromosome (22), consistent with the observation that virulence factors are often encoded on pathogenicity islands in the genomes of plant pathogens (42). Similarly, the SAP11 PMU-like region is predicted to encode additional effector proteins, such as SAP09, SAP56, SAP66, SAP67, and SAP68 (22), which may also affect plant-insect interactions.
The observation that the phytoplasma effector SAP11 targets JA synthesis via interaction with a plant transcription factor may be similar to the description of a viral protein produced by Tomato yellow leaf curl China virus (TYLCCNV). This geminivirus produces a protein (βC1) that acts as a molecular mimic of the plant regulatory protein AYSYMMETRIC LEAVES 2 (AS2) to suppress a subset of JA-responsive genes (43). In this manner, βC1 (and thus TYLCCNV) suppresses JA signaling, which is proposed to make infected plants into better hosts for the TYLCCNV whitefly vector, a phloem-feeding insect vector that acquires the plant virus while feeding on an infected plant (43). However, this has not been experimentally proven. Although the outcome would be similar in both cases (i.e., improvement of insect vector fitness), the mechanisms are different, because SAP11-mediated destabilization of CIN-TCPs causes a reduction in JA synthesis, whereas the βC1 complex formation with AS1 leads to a decrease in JA response. The 2b protein of the aphid-transmitted Cucumber mosaic virus is also reported to inhibit the induction of JA-responsive genes; however, the mechanism of the inhibition has not been revealed (44).
We found that AY-WB SAP11 interacts with a class I and a subset of class II TCP transcription factors and destabilizes all the class II CIN-TCPs, whereas at least one class I TCP (TCP7) appears to remain intact. This is consistent with the crinkly leaf phenotype of 35S::SAP11 transgenic lines. Class II CIN-TCPs positively regulate cell maturation, whereas class I TCPs regulate cell proliferation (32, 45). The two classes of TCPs are proposed to compete to occupy overlapping promoter sequences of downstream genes and to coordinately or competitively regulate the expression of genes (45). As seen in the 35S::miR319a and octuple cin-tcp plants, down-regulation of class II TCPs induces overproduction of immature plant cells and produces large crinkly leaves (32). Interestingly, however, 35S::miR319a and octuple cin-tcp plants did not produce an increased number of stems, indicating that the bushy appearance of 35S::SAP11 lines may not be caused by the destabilization of CIN-TCPs. Whereas SAP11 does not appear to destabilize the class I TCPs (based on the unchanged abundance of TCP7 in the presence of SAP11), it is possible that SAP11 affects three other class II CYC/TB1-like TCPs (TCP1, TCP12, and TCP18). TCP12 and TCP18 arrest the development of axillary buds (46), and SAP11-mediated degradation of these TCPs would explain the increase in stem numbers observed in SAP11 plants. That SAP11 interacts with TCPs is consistent with our previous observations that SAP11 targets nuclei in AY-WB–infected plants (22), because SAP11 is likely to encounter the transcription factors in plant cell nuclei. Furthermore, the similar level of insect fecundity increase on 35S::SAP11 and the octuple cin-tcp lines supports that SAP11 contributes to an increase in insect fecundity through the destabilization of CIN-TCPs. The spatiotemporal destabilization of CIN-TCPs in the active AY-WB infection process and the mechanism by which SAP11 destabilizes class II but not class I TCPs remain to be investigated. SAP11 does not have any homology with already characterized proteins, such as proteases; therefore, we postulate that SAP11-mediated destabilization of TCPs is not attributable to direct protease activity of SAP11; rather, it requires host protein degradation systems.
LOX2 expression and JA synthesis are down-regulated in 35S::SAP11 lines (vs. Col-0) following wounding. This is in agreement with the SAP11-mediated destabilization of the class II CIN-TCP transcription factor TCP4, which positively regulates plant senescence by up-regulating LOX2 expression directly through binding to the LOX2 promoter (33). LOX2 is involved in the synthesis of JA (35, 38) because it encodes an enzyme that converts α-LeA into 13-HPOT, a substrate in the first step of the octadecanoid pathway for JA synthesis. The octadecanoid pathway plays an important role in the plant defense response to insect herbivores (47, 48) and is strongly up-regulated on wounding and insect feeding in Arabidopsis (49). LOX2 expression is also up-regulated in response to M. quadrilineatus exposure, and the leafhopper has higher oviposition and nymph production rates on 35S::SAP11- and LOX2-silenced Arabidopsis lines, indicating that the SAP11-mediated down-regulation of LOX2 is beneficial to the AY-WB insect vector.
Substrates generated by LOX2 and other 13-HPOTs also feed into the hydroperoxide lyase pathway that produces green leaf volatiles (GLVs) in Arabidopsis (50), raising the possibility that the enhanced leafhopper fecundity may be related to the impaired synthesis of GLVs. However, the observation that M. quadrilineatus fecundity is also increased on the Arabidopsis jar1 mutant indicates that disruption of JA synthesis is most likely responsible for the increase in M. quadrilineatus fecundity observed in SAP11 transgenic lines. JAR1 converts JA into JA-Ile, which is the biologically active molecule of the JA signaling pathway and acts much further downstream of LOX2 in this pathway (39). The M. quadrilineatus egg hatch rate increased on 35S::LOX2 plants, and overall female survival rate increased slightly on jar1 mutants. Thus, the JA pathway has an impact on several aspects of M. quadrilineatus fitness, and as a consequence, we might observe clear increases in insect fecundity on JA pathway-deficient mutants.
Together, our data indicate that SAP11 interferes with the plant defense response against the insect vector through the destabilization of CIN-TCPs, thereby enhancing the reproductive success of the insect vector on which AY-WB is dependent for transmission and long-distance dispersal (model in SI Appendix, Fig. 7). At this time, it is not possible to generate AY-WB mutants that lack SAP11, because phytoplasmas cannot be cultured and transformed. Therefore, we cannot measure the SAP11 contribution to AY-WB–mediated increase in insect fecundity directly. Furthermore, we do not yet understand how SAP11 expression is regulated temporally or spatially during AY-WB colonization of the plant and if one or more of the remaining effectors (56 predicted so far) affect SAP11 expression or function. Nonetheless, we observed that on wounding, young leaves accumulate less JA in AY-WB–infected than noninfected Arabidopsis plants. These data are consistent with the detection of SAP11 in young leaves of AY-WB–infected plants (22) and observations that phytoplasmas and their effectors accumulate predominantly in sink tissues (26, 51). Taken together, we propose that SAP11 promotes AY-WB fitness by increasing insect progeny numbers (SI Appendix, Fig. 7).
It is interesting that SAP11 targets members of a transcription family that is highly conserved among plant species, an observation that is consistent with the wide host range of AY-WB. The observation that SAP11 targets class II CIN-TCPs for destabilization (and not class I regulators) is also intriguing, because CIN-TCPs are important growth regulators that promote the differentiation of cells and maturation of leaves. The down-regulation of CIN-TCPs (as observed in the octuple cin-tcp plants) elicits the production of unusually large leaves that mature slowly and comprise a large population of small and immature cells (32). Thus, another consequence of SAP11 selectively destabilizing CIN-TCPs might be a delay in the maturation of developing vegetative tissues in infected plants. Because phytoplasmas are biotrophs, it is an advantage for the pathogen to reduce plant stress and delay death. Indeed, phytoplasmas are obligate inhabitants of the plant hosts and insect vectors; hence, too rapid a decline of their host would seriously impinge on phytoplasma survival and decrease their opportunity to be acquired by insect vectors that require relatively healthy plants for feeding and reproduction. Nevertheless, whereas the SAP11-CIN-TCP interaction might influence both developmental and defense pathways in infected plants, it is likely that the primary role of SAP11 comprises the down-regulation of plant defense mechanisms against insects, which involves the JA-mediated pathway, and possibly other unknown pathways regulated by CIN-TCPs, and that any developmental perturbations (e.g., witches’ broom) arise as an indirect (albeit beneficial) consequence of destabilizing TCPs.
Whereas the function of some animal and plant pathogen effectors has been elucidated, effectors that reach beyond the pathogen–host interface to affect a third organism in the biological interaction have not yet been identified and functionally characterized. Our results show that SAP11 has a phenotype beyond the organism in which it resides, affecting not only the AY-WB–infected plant host but the reproductive success of the leafhopper vector that transmits AY-WB to a diverse range of plant species and throughout North America (18, 41). Hence, SAP11 is a vivid example of the extended phenotype of the gene, a concept put forward in Richard Dawkins’ classic book (52).
Materials and Methods
AY-WB Maintenance and Insect Rearing and Fecundity Experiments.
All insect rearing and maintenance of AY-WB–infected plant materials occurred in insect rearing cages in a growth room set at a 16-h/8-h light/dark period at 26 °C/20 °C. To obtain AY-WB–infected leafhoppers, ∼50 female and male M. quadrilineatus leafhoppers were transferred to AY-WB–infected China aster (Callistephus chinensis Nees) for 2 wk to allow oviposition. Nymphs were kept on the infected plants until and during adulthood. The adults were used for experiments or were transferred to young China aster or Arabidopsis seedlings to generate new batches of AY-WB–infected plants and leafhoppers.
Insect fecundity experiments were conducted as explained in the figure legends at a 10-h/14-h light/dark period at 22 °C. All the M. quadrilineatus leafhoppers used for fecundity assay were anesthetized with CO2 gas for sexing.
Statistical Analyses.
All statistical analysis was completed in Genstat v.13 (International Ltd.). For all data points of insect fecundity and oviposition assay, we recorded the biological replicate, adult age, and plant treatment or line as explanatory variables and female survival as response variables. ANODE was used to determine differences in female survival rates, egg hatch rate, fecundity, oviposition, and number of feeding sites. All explanatory variables were described as factors and modeled to assess their contribution to the deviance in leafhopper survival, egg hatch rate, fecundity, oviposition rates, and number of feeding sites. Female survival data and egg hatch data were analyzed using a binomial distribution within a generalized linear model (GLM). Fecundity, oviposition, and feeding data were analyzed using a Poisson distribution within a GLM. When fecundity and oviposition data were analyzed, female survival in each experiment was added as a covariate. Means of nymph or egg numbers or female survival at each insect age were compared using t probabilities calculated within the GLM. For all the other experiments, datasets were first examined for normal distributions in Genstat and then compared using ANOVA or the Student t test (one-tailed).
Generation and Analyses of 35S::SAP11 Arabidopsis Lines.
A codon-optimized version of the AY-WB SAP11 sequence (GenBank accession no. gi: 85057280) without the sequence corresponding to the signal peptide and attB1 and attB2 adapters at the ends (SI Appendix, Fig. 2A) was synthesized (GenScript). This SAP11 fragment was amplified with attB1 and attB2 adapter primers (SI Appendix, Table 2) and was cloned into pDONOR207 (Invitrogen) using Clonase LR (Invitrogen). The SAP11 fragment in pDONOR207 was then Clonase LR-cloned into the Gateway-adapted vector pB7WG2 that has a 35S promoter fragment cloned at the 5′ end of the clonase compatible insertion site (53). The resulting plasmids were transformed into Agrobacterium tumefaciens strain GV3101. Arabidopsis Col-0 was transformed by floral dip as described (54).
Detection of SAP11 in Transgenic Plants.
Antibodies to SAP11 were raised against a partial peptide sequence of SAP11 (CEEGSSSKQPDDSKK) in rabbits (GenScript). Four seedlings of 10-d-old Arabidopsis seedlings were ground in 200 μL of extraction buffer [50 mM sodium phosphate (pH 7.0), 10 mM Triton X-100, 10 mM N-lauroylsarcosine, 1 mM 2-mercaptoethanol]. Twelve microliters of the extracts was mixed with 4× NuPAGE LDS sample buffer (Invitrogen), separated on 15% (wt/vol) SDS polyacrylamide gels (PAGE), and transferred to 0.45 μm of Protran BA85 nitrocellulose membranes (Whatman) following standard procedures (55). The Western blots were incubated with αSAP11 and peroxidase-conjugated anti-rabbit IgG (Sigma–Aldrich), and detection of bound antibodies was conducted with Immobilon Western Chemiluminescent HRP Substrate (Millipore).
Yeast Two-Hybrid Analysis.
Yeast two-hybrid screening was performed by Hybrigenics Services SAS. The codon-optimized sequence of AY-WB SAP11 without signal peptide (SI Appendix, Fig. 2A) was PCR-amplified and cloned into pB27 as a C-terminal fusion to LexA (N-LexA-SAP11-C) and into pB66 as a C-terminal fusion to Gal4 DNA-binding domain (N-Gal4-SAP11-C). The constructs were used as bait to screen a random-primed A. thaliana seedlings cDNA library constructed into pP6. For the LexA bait construct, 130 million clones (13-fold the complexity of the library) were screened, using a mating approach with Y187 (mat α) and L40DGal4 (mat a) yeast strains as previously described (56). For the Gal4 construct, 74 million clones (7-fold the complexity of the library) were screened using the same mating approach with Y187 and CG1945 (mat a) yeast strains.
35S::miR319a, 35S::miR3TCP, BLS::rTCP4GFP, 35S::LOX2, and 35S::LOX2 Antisense Lines.
Seed for the 35S::miR319a (Col-0), 35S::miR3TCP (Col-0), 35S::miR319a × 35S::miR3TCP (Col-0), and pBLS::rTCP4GFP (Ler) lines was kindly provided by Idan Efroni (Weizmann Institute of Science, Rehovot, Israel), and the phenotypes of the transgenic lines were as described previously (32). The BLS promoter was selected for expression of rTCP4GFP, because 35S::rTCP4 plants show severe developmental phenotypes. Activity of the BLS promoter is initiated and maintained solely in young lamina (P3–P7 stages) (57). The 35S::LOX2 (NASC ID code N3748) and corresponding control line 35S::LOX2 antisense (NASC ID code N3749) were requested from the European Arabidopsis Stock Centre and were described previously (38).
Coexpression and Pull-Down Assays.
The coexpression and pull-down assays were conducted in N. benthamiana leaves. The A. tumefaciens clones carrying the appropriate constructs were infiltrated into individual N. benthamiana leaves using a needleless syringe. After 3 d, 2 leaf disks 11 mm in diameter were collected (1 leaf disk per leaf). The two leaf disks were ground in liquid nitrogen in the presence of 50 μL of 1× NuPage Sample Buffer (Invitrogen) and boiled for 5 min. Proteins were separated on 12.5% (wt/vol) SDS/PAGE gels. Remaining leaf samples were used for pull-down assays. The leaf was cross-linked with 30 mL of 10 mM Tris(hydroxymethyl)aminomethane (Tris)⋅HCl (pH 8.0), 400 mM sucrose, 3% (wt/vol) formaldehyde, 0.1 mM PMSF, and 5 mM β-mercaptoethanol using vacuum infiltration for 10 min. The reaction was then quenched with 2.5 mL 2-M glycine and vacuum-infiltrated for a further 5 min. Samples were ground into a fine powder in the presence of 2 mL of GTEN buffer [10% (vol/vol) glycerol, 25 mM Tris⋅HCl (pH 7.5), 1 mM EDTA, 150 mM NaCl], and centrifuged at 3,000 × g for 10 min. The supernatant was transferred to a fresh tube and centrifuged at 14,000 × g for 10 min. The FLAG-tagged proteins were extracted by combining 250 μL of supernatant with 1.7 mL of GTEN buffer plus 0.1% Tween 20 (IP buffer) and 20 μL of anti-FLAG resin (Sigma–Aldrich), which was prewashed with IP buffer. The bound proteins were eluted in 40 μL of IP buffer containing 150 ng⋅μL−1 3× FLAG peptide (Sigma–Aldrich) for 30 min. On average, 15 μL of the eluates was mixed with NuPAGE LDS sample buffer (Invitrogen), boiled for at least 30 min, and then separated on 12.5% (wt/vol) SDS/PAGE gels and analyzed on Western blots as described above.
Quantitative RT-PCR Experiments.
For wounding experiments, mature leaves of 5-wk-old Arabidopsis were harvested carefully or wounded twice by forceps. Samples were harvested 60 min after wounding. For leafhopper infestation assays, mature leaves of 5-wk-old Arabidopsis Col-0 plants were singly equipped with a clip cage, each containing 10 M. quadrilineatus adults. The leafhoppers were anesthetized by CO2 to facilitate handling. As nontreated controls, Arabidopsis leaves were clip-caged without M. quadrilineatus.
Arabidopsis leaves were snap-frozen and used for RNA extraction with TRI reagent (Sigma–Aldrich) and purified using Qiagen RNeasy columns (Qiagen Ltd.). cDNA was synthesized from 0.5 μg of total RNA using M-MLV reverse transcriptase (Invitrogen), and subjected to real-time PCR using SYBR Green JumpStart Taq ReadyMix (Sigma–Aldrich) in a DNA Engine Opticon 2 (BioRad, Life Sciences) using gene-specific primers (SI Appendix, Table 2).
Extraction and Quantification of Jasmonic Acid.
Arabidopsis leaves were harvested carefully or wounded twice by forceps, and two to three leaves per treatment were collected at indicated time points and snap-frozen in liquid nitrogen. To generate AY-WB–infected plants, six 4-wk-old plants were placed in a perforated bag with 20 AY-WB carrier or noncarrier M. quadrilineatus males. The insects were removed 1 wk later, and the plants were wounded at specific time points as described. The samples were lyophilized and ground to powder with liquid nitrogen, and then extracted at 4 °C overnight with 5 mL of methanol containing 100 ng of deuterated JA (d5-JA, C/D/N isotopes) as an internal standard. The total extracts were centrifuged in a swing-bucket rotor (Eppendorf 5810R; 4,000 rpm, 173-mm rotor radius, 5 min), and the supernatants were dried under N2 gas. The pellets were dissolved in diethyl ether/methanol (60:9) and loaded onto a Strata NH2 solid phase extraction column (Phenomenex). After two washes with 0.8 mL of chloroform/isopropanol (2:1), JA was eluted twice by 1 mL of diethyl ether containing 4% (vol/vol) acetic acid. The combined eluates were dried with N2 gas, and the pellets dissolved in methanol/water (1:4). Samples (10 μL) were analyzed using an Agilent 1100 single-quadrupole liquid chromatography-MS system.
Supplementary Material
Acknowledgments
We acknowledge Dr. Yuval Eshed and Dr. Idan Efroni (Department of Plant Sciences, Weizmann Institute of Science) for providing the seed for the 35S::miR319a (Col-0), 35S::miR3TCP (Col-0), 35S::miR319a × 35S::miR3TCP (Col-0), and pBLS::rTCP4GFP (Ler) lines. The phenotypes of these transgenic lines were described previously (32). We thank Dr. Lionel Hill (The John Innes Centre) for conducting the HPLC analyses to measure JA and Prof. James Brown and Dr. Christopher Burt (The John Innes Centre) for help with the statistical analyses of leafhopper survival and fecundity datasets. We also thank Prof. Enrico Coen (The John Innes Centre) for useful discussions, and Prof. Sophien Kamoun and Dr. Silke Robatzek (The Sainsbury Laboratory, Norwich, UK) and Prof. Ian Baldwin (Max Planck Institute for Chemical Ecology) for critical reading of various versions of this manuscript. We thank Ian Bedford, Anna Jordan, and Gavin Hatt of the JIC Insectary for insect rearing, and Andrew Davis for photography. This research was funded by the Biotechnology and Biological Sciences Research Council (BBSRC) Grant BBSEJ000CA357, The John Innes Centre, and The Gatsby Charitable Foundation. H.N.K. was funded from a BBSRC studentship. The John Innes Center is grant-aided by the BBSRC.
Footnotes
The authors declare no conflict of interest.
*This Direct Submission article had a prearranged editor.
See Author Summary on page 19111.
This article contains supporting information online at www.pnas.org/lookup/suppl/doi:10.1073/pnas.1105664108/-/DCSupplemental.
References
- 1.Lacroix R, Mukabana WR, Gouagna LC, Koella JC. Malaria infection increases attractiveness of humans to mosquitoes. PLoS Biol. 2005;3:e298. doi: 10.1371/journal.pbio.0030298. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Taylor PJ, Hurd H. The influence of host haematocrit on the blood feeding success of Anopheles stephensi: Implications for enhanced malaria transmission. Parasitology. 2001;122:491–496. doi: 10.1017/s0031182001007776. [DOI] [PubMed] [Google Scholar]
- 3.Roy BA. Floral mimicry by a plant pathogen. Nature. 1993;362:56–58. [Google Scholar]
- 4.Beanland L, Hoy CW, Miller SA, Nault LR. Influence of Aster Yellows phytoplasma on the fitness of aster leafhopper (Homoptera: Cicadellidae) Ann Entomol Soc Am. 2000;93:271–276. [Google Scholar]
- 5.Eigenbrode SD, Ding H, Shiel P, Berger PH. Volatiles from potato plants infected with potato leafroll virus attract and arrest the virus vector, Myzus persicae (Homoptera: Aphididae) Proc Biol Sci. 2002;269:455–460. doi: 10.1098/rspb.2001.1909. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Mayer CJ, Vilcinskas A, Gross J. Phytopathogen lures its insect vector by altering host plant odor. J Chem Ecol. 2008;34:1045–1049. doi: 10.1007/s10886-008-9516-1. [DOI] [PubMed] [Google Scholar]
- 7.Mauck KE, De Moraes CM, Mescher MC. Deceptive chemical signals induced by a plant virus attract insect vectors to inferior hosts. Proc Natl Acad Sci USA. 2010;107:3600–3605. doi: 10.1073/pnas.0907191107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Ray K, Marteyn B, Sansonetti PJ, Tang CM. Life on the inside: The intracellular lifestyle of cytosolic bacteria. Nat Rev Microbiol. 2009;7:333–340. doi: 10.1038/nrmicro2112. [DOI] [PubMed] [Google Scholar]
- 9.Hogenhout SA, Van der Hoorn RA, Terauchi R, Kamoun S. Emerging concepts in effector biology of plant-associated organisms. Mol Plant Microbe Interact. 2009;22:115–122. doi: 10.1094/MPMI-22-2-0115. [DOI] [PubMed] [Google Scholar]
- 10.Xiang T, et al. Pseudomonas syringae effector AvrPto blocks innate immunity by targeting receptor kinases. Curr Biol. 2008;18:74–80. doi: 10.1016/j.cub.2007.12.020. [DOI] [PubMed] [Google Scholar]
- 11.Göhre V, et al. Plant pattern-recognition receptor FLS2 is directed for degradation by the bacterial ubiquitin ligase AvrPtoB. Curr Biol. 2008;18:1824–1832. doi: 10.1016/j.cub.2008.10.063. [DOI] [PubMed] [Google Scholar]
- 12.Kay S, Hahn S, Marois E, Hause G, Bonas U. A bacterial effector acts as a plant transcription factor and induces a cell size regulator. Science. 2007;318:648–651. doi: 10.1126/science.1144956. [DOI] [PubMed] [Google Scholar]
- 13.Chen LQ, et al. Sugar transporters for intercellular exchange and nutrition of pathogens. Nature. 2010;468:527–532. doi: 10.1038/nature09606. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Hogenhout SA, et al. Phytoplasmas: Bacteria that manipulate plants and insects. Mol Plant Pathol. 2008;9:403–423. doi: 10.1111/j.1364-3703.2008.00472.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Weintraub PG, Beanland L. Insect vectors of phytoplasmas. Annu Rev Entomol. 2006;51:91–111. doi: 10.1146/annurev.ento.51.110104.151039. [DOI] [PubMed] [Google Scholar]
- 16.Firrao G, Garcia-Chapa M, Marzachì C. Phytoplasmas: Genetics, diagnosis and relationships with the plant and insect host. Front Biosci. 2007;12:1353–1375. doi: 10.2741/2153. [DOI] [PubMed] [Google Scholar]
- 17.O'Mara J, Bauernfeind R, Stevens A, Gast KLB, Stevens S. Aster Yellows. Manhattan , KS: Cooperative Extension Service; 2003. pp. 1–6. [Google Scholar]
- 18.Wilson M, Turner J. Leafhopper, planthopper and psyllid vectors of plant disease. Amgueddfa Cymru-National Museum Wales. 2010. Available at. http://naturalhistory.museumwales.ac.uk/Vectors. Accessed November 4, 2010.
- 19.Bertaccini A. Phytoplasmas: Diversity, taxonomy, and epidemiology. Front Biosci. 2007;12:673–689. doi: 10.2741/2092. [DOI] [PubMed] [Google Scholar]
- 20.Bai X, et al. Living with genome instability: The adaptation of phytoplasmas to diverse environments of their insect and plant hosts. J Bacteriol. 2006;188:3682–3696. doi: 10.1128/JB.188.10.3682-3696.2006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Hogenhout SA, Loria R. Virulence mechanisms of Gram-positive plant pathogenic bacteria. Curr Opin Plant Biol. 2008;11:449–456. doi: 10.1016/j.pbi.2008.05.007. [DOI] [PubMed] [Google Scholar]
- 22.Bai X, et al. AY-WB phytoplasma secretes a protein that targets plant cell nuclei. Mol Plant Microbe Interact. 2009;22:18–30. doi: 10.1094/MPMI-22-1-0018. [DOI] [PubMed] [Google Scholar]
- 23.Rosch J, Caparon M. A microdomain for protein secretion in Gram-positive bacteria. Science. 2004;304:1513–1515. doi: 10.1126/science.1097404. [DOI] [PubMed] [Google Scholar]
- 24.Kakizawa S, et al. Cloning and expression analysis of Phytoplasma protein translocation genes. Mol Plant Microbe Interact. 2001;14:1043–1050. doi: 10.1094/MPMI.2001.14.9.1043. [DOI] [PubMed] [Google Scholar]
- 25.Kakizawa S, et al. Secretion of immunodominant membrane protein from onion yellows phytoplasma through the Sec protein-translocation system in Escherichia coli. Microbiology. 2004;150:135–142. doi: 10.1099/mic.0.26521-0. [DOI] [PubMed] [Google Scholar]
- 26.Sugio A, et al. Diverse targets of phytoplasma effectors: From plant development to defense against insects. Annu Rev Phytopathol. 2011;49:175–195. doi: 10.1146/annurev-phyto-072910-095323. [DOI] [PubMed] [Google Scholar]
- 27.Maclean AM, et al. Phytoplasma effector SAP54 induces indeterminate leaf-like flower development in Arabidopsis plants. Plant Physiol. 2011;157:831–841. doi: 10.1104/pp.111.181586. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Palatnik JF, et al. Control of leaf morphogenesis by microRNAs. Nature. 2003;425:257–263. doi: 10.1038/nature01958. [DOI] [PubMed] [Google Scholar]
- 29.Cubas P, Lauter N, Doebley J, Coen E. The TCP domain: A motif found in proteins regulating plant growth and development. Plant J. 1999;18:215–222. doi: 10.1046/j.1365-313x.1999.00444.x. [DOI] [PubMed] [Google Scholar]
- 30.Howarth DG, Donoghue MJ. Phylogenetic analysis of the “ECE” (CYC/TB1) clade reveals duplications predating the core eudicots. Proc Natl Acad Sci USA. 2006;103:9101–9106. doi: 10.1073/pnas.0602827103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Nath U, Crawford BC, Carpenter R, Coen E. Genetic control of surface curvature. Science. 2003;299:1404–1407. doi: 10.1126/science.1079354. [DOI] [PubMed] [Google Scholar]
- 32.Efroni I, Blum E, Goldshmidt A, Eshed Y. A protracted and dynamic maturation schedule underlies Arabidopsis leaf development. Plant Cell. 2008;20:2293–2306. doi: 10.1105/tpc.107.057521. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Schommer C, et al. Control of jasmonate biosynthesis and senescence by miR319 targets. PLoS Biol. 2008;6:e230. doi: 10.1371/journal.pbio.0060230. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Lindbo JA. TRBO: A high-efficiency tobacco mosaic virus RNA-based overexpression vector. Plant Physiol. 2007;145:1232–1240. doi: 10.1104/pp.107.106377. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Turner JG, Ellis C, Devoto A. The jasmonate signal pathway. Plant Cell. 2002;14(Suppl):S153–S164. doi: 10.1105/tpc.000679. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Allmann S, Halitschke R, Schuurink RC, Baldwin IT. Oxylipin channelling in Nicotiana attenuata: Lipoxygenase 2 supplies substrates for green leaf volatile production. Plant Cell Environ. 2010;33:2028–2040. doi: 10.1111/j.1365-3040.2010.02203.x. [DOI] [PubMed] [Google Scholar]
- 37.Kessler A, Halitschke R, Baldwin IT. Silencing the jasmonate cascade: Induced plant defenses and insect populations. Science. 2004;305:665–668. doi: 10.1126/science.1096931. [DOI] [PubMed] [Google Scholar]
- 38.Bell E, Creelman RA, Mullet JE. A chloroplast lipoxygenase is required for wound-induced jasmonic acid accumulation in Arabidopsis. Proc Natl Acad Sci USA. 1995;92:8675–8679. doi: 10.1073/pnas.92.19.8675. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Staswick PE, Tiryaki I. The oxylipin signal jasmonic acid is activated by an enzyme that conjugates it to isoleucine in Arabidopsis. Plant Cell. 2004;16:2117–2127. doi: 10.1105/tpc.104.023549. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Zhang J, Hogenhout SA, Nault LR, Hoy CW, Miller SA. Molecular and symptom analyses of phytoplasma strains from lettuce reveal a diverse population. Phytopathology. 2004;94:842–849. doi: 10.1094/PHYTO.2004.94.8.842. [DOI] [PubMed] [Google Scholar]
- 41.Cranshaw W. Pests of the West: Prevention and Control for Today's Garden and Small Farm. 2nd Ed. Golden, CO: Fulcrum Publishing; 1998. p. 138. [Google Scholar]
- 42.Gal-Mor O, Finlay BB. Pathogenicity islands: A molecular toolbox for bacterial virulence. Cell Microbiol. 2006;8:1707–1719. doi: 10.1111/j.1462-5822.2006.00794.x. [DOI] [PubMed] [Google Scholar]
- 43.Yang JY, et al. betaC1, the pathogenicity factor of TYLCCNV, interacts with AS1 to alter leaf development and suppress selective jasmonic acid responses. Genes Dev. 2008;22:2564–2577. doi: 10.1101/gad.1682208. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Lewsey MG, et al. Disruption of two defensive signaling pathways by a viral RNA silencing suppressor. Mol Plant Microbe Interact. 2010;23:835–845. doi: 10.1094/MPMI-23-7-0835. [DOI] [PubMed] [Google Scholar]
- 45.Martín-Trillo M, Cubas P. TCP genes: A family snapshot ten years later. Trends Plant Sci. 2010;15:31–39. doi: 10.1016/j.tplants.2009.11.003. [DOI] [PubMed] [Google Scholar]
- 46.Aguilar-Martínez JA, Poza-Carrión C, Cubas P. Arabidopsis BRANCHED1 acts as an integrator of branching signals within axillary buds. Plant Cell. 2007;19:458–472. doi: 10.1105/tpc.106.048934. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Bonaventure G, Baldwin IT. Transduction of wound and herbivory signals in plastids. Commun Integr Biol. 2010;3:313–317. doi: 10.4161/cib.3.4.11834. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Browse J. Jasmonate passes muster: A receptor and targets for the defense hormone. Annu Rev Plant Biol. 2009;60:183–205. doi: 10.1146/annurev.arplant.043008.092007. [DOI] [PubMed] [Google Scholar]
- 49.Reymond P, Weber H, Damond M, Farmer EE. Differential gene expression in response to mechanical wounding and insect feeding in Arabidopsis. Plant Cell. 2000;12:707–720. doi: 10.1105/tpc.12.5.707. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Duan H, Huang MY, Palacio K, Schuler MA. Variations in CYP74B2 (hydroperoxide lyase) gene expression differentially affect hexenal signaling in the Columbia and Landsberg erecta ecotypes of Arabidopsis. Plant Physiol. 2005;139:1529–1544. doi: 10.1104/pp.105.067249. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Hoshi A, et al. A unique virulence factor for proliferation and dwarfism in plants identified from a phytopathogenic bacterium. Proc Natl Acad Sci USA. 2009;106:6416–6421. doi: 10.1073/pnas.0813038106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Dawkins R. The Extended Phenotype: The Long Reach of the Gene. Oxford: Oxford Univ Press; 1999. [Google Scholar]
- 53.Karimi M, Inzé D, Depicker A. GATEWAY vectors for Agrobacterium-mediated plant transformation. Trends Plant Sci. 2002;7:193–195. doi: 10.1016/s1360-1385(02)02251-3. [DOI] [PubMed] [Google Scholar]
- 54.Clough SJ, Bent AF. Floral dip: A simplified method for Agrobacterium-mediated transformation of Arabidopsis thaliana. Plant J. 1998;16:735–743. doi: 10.1046/j.1365-313x.1998.00343.x. [DOI] [PubMed] [Google Scholar]
- 55.Sambrook J, Fritsch EF, Maniatis T. Molecular Cloning: A Laboratory Manual. Cold Spring Harbor, NY: Cold Spring Harbor Laboratory Press; 1989. [Google Scholar]
- 56.Fromont-Racine M, Rain JC, Legrain P. Toward a functional analysis of the yeast genome through exhaustive two-hybrid screens. Nat Genet. 1997;16:277–282. doi: 10.1038/ng0797-277. [DOI] [PubMed] [Google Scholar]
- 57.Lifschitz E, et al. The tomato FT ortholog triggers systemic signals that regulate growth and flowering and substitute for diverse environmental stimuli. Proc Natl Acad Sci USA. 2006;103:6398–6403. doi: 10.1073/pnas.0601620103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Nemhauser JL, Mockler TC, Chory J. Interdependency of brassinosteroid and auxin signaling in Arabidopsis. PLoS Biol. 2004;2:E258. doi: 10.1371/journal.pbio.0020258. [DOI] [PMC free article] [PubMed] [Google Scholar]