Abstract
Background
Glucocorticoids such as prednisolone and dexamethasone are critical drugs used in multi-agent chemotherapy protocols used to treat acute lymphoblastic leukemia (ALL), and response to glucocorticoids is highly predictive of outcome. The NOD/SCID xenograft mouse model of ALL is a clinically relevant model in which the mice develop a systemic leukemia which retains the fundamental biological characteristics of the original disease. Here we report a study evaluating the NOD/SCID xenograft mouse model to investigate glucocorticoid-induced gene expression. Cells from a glucocorticoid-sensitive xenograft derived from a child with B-cell precursor ALL were inoculated into NOD/SCID mice. When highly engrafted the mice were randomized into groups of 4 to receive dexamethasone 15 mg/kg by intraperitoneal injection or vehicle control. Leukemia cells were harvested from mice spleens at 0, 8, 24 or 48 hours thereafter, and gene expression analyzed on Illumina WG-6_V3 chips, comparing all groups to time 0 hours.
Results
The 8 hour dexamethasone-treated timepoint had the highest number of significantly differentially expressed genes, with fewer observed at the 24 and 48 hour timepoints, and with minimal changes seen across the time-matched controls. When compared to publicly available datasets of glucocorticoid-induced gene expression from an in vitro cell line study and from an in vivo study of patients with ALL, at the level of pathways, expression changes in the 8 hour xenograft samples showed a similar response to patients treated with glucocorticoids. Replicate analysis revealed that at the 8 hour timepoint, a dataset with high signal and differential expression, using data from 3 replicates instead of 4 resulted in excellent recovery scores of > 0.9. However at other timepoints with less signal very poor recovery scores were obtained with 3 replicates.
Conclusions
The NOD/SCID xenograft mouse model provides a reproducible experimental system in which to investigate clinically-relevant mechanisms of drug-induced gene regulation in ALL; the 8 hour timepoint provides the highest number of significantly differentially expressed genes; time-matched controls are redundant and excellent recovery scores can be obtained with 3 replicates.
Background
Glucocorticoids such as prednisolone and dexamethasone are critical components of multi-agent chemotherapy protocols used in the treatment of acute lymphoblastic leukemia (ALL) [1] due to their ability to induce apoptosis in lymphoid cells. Despite their use for over 50 years their mechanism of action is not completely understood. Glucocorticoids are steroid hormones that act on target cells through interaction with a specific glucocorticoid receptor (GR) [2]. The GR is held in a cytosolic complex by a number of co-chaperone molecules including HSP-90 and HSP-70 [3], and on ligand binding dissociates from the co-chaperone complex, dimerizes and is transported to the nucleus where it binds to palindromic DNA sequences known as glucocorticoid response elements (GREs) found in the promoter regions of target genes [4]. This leads to the activation of transcription of primary target genes, repression of transcription through interaction with negative GREs [5] or of gene activation through transcription factors such as AP-1 and NF-ΚB [6]. In lymphoid cells, this results in repression of cell cycle progression through cyclin-D3 and C-MYC [7], and cell death through the activation of apoptosis. Glucocorticoids also induce other non-apoptotic mechanisms of programmed cell death including autophagy [8] and mediate a number of pathways involved in the metabolism of carbohydrates, lipids and proteins.
A number of studies have published microarray data of glucocorticoid-induced genes in lymphoid cells, but comparison of the data is complicated by technical differences in platform and chip type. Previous studies of glucocorticoid-induced genes in ALL have been carried out using in vitro cell-line models [9-15] and patient-derived cells, both in vivo [16] and in vitro [17]. Cell lines are extensively used in the study of ALL but in the process of immortalization acquire multiple genetic defects, particularly in the p53 pathway [18], and mechanisms demonstrated in cell lines are often not replicated in more clinically relevant models. Primary patient cells have a finite supply and rarely survive ex vivo for more than a few days. The non-obese diabetic/severe combined immunodeficient (NOD/SCID) xenograft mouse model is widely used to study ALL. In this model, human leukemia cells obtained from diagnostic bone marrow biopsies are inoculated into NOD/SCID mice, and on engraftment establish a systemic leukemia which retains the fundamental biological characteristics of the original disease [19]. It has also been shown that the in vivo responses to chemotherapeutic agents, including dexamethasone, correlates with patient outcome [20], and thus the NOD/SCID xenograft mouse model provides a stable, reproducible and clinically relevant model with which to study ALL. Here we report the first study investigating glucocorticoid-induced gene expression in ALL using the NOD/SCID xenograft model, the optimal experimental design, and a comparison of our microarray data to publicly available datasets of glucocorticoid-induced genes in other experimental models.
Methods
NOD/SCID xenograft mouse model
All experimental studies were approved by the Human Research Ethics Committee and the Animal Care and Ethics Committee of the University of New South Wales. ALL-3, a glucocorticoid-sensitive xenograft derived from a 12 year old girl with mixed lineage leukemia (MLL)-rearranged BCP-ALL, was chosen for this study. Although MLL-rearranged ALL is associated with a poor prednisolone response and an inferior outcome [21], this patient is currently a long-term survivor. ALL-3 demonstrates in vitro glucocorticoid sensitivity, with an IC50 of 9.4 nM on exposure to dexamethasone. In the in vivo NOD/SCID xenograft mouse model, ALL-3 is highly responsive to 4 weeks of treatment with single agent dexamethasone, with rapid clearance of leukemic blasts from the peripheral blood and recurrence of leukemia delayed by 63.4 days compared to vehicle-treated controls [20].
Cells from ALL-3 were inoculated by tail-vein injection into 28 NOD/SCID mice. The mice were bled weekly and the samples stained with fluorescein isothiocyanate (FITC)-conjugated anti-murine CD45 and allophycocyanin (APC)-conjugated anti-human CD45 (BioLegend, San Diego, CA). Following lysis of erythrocytes with FACS lysing solution (BD Biosciences, San Jose, CA), samples were analyzed by multiparametric flow cytometry on a FACSCanto cytometer (BD Biosciences, San Jose, CA). Engraftment was calculated as the proportion of human versus total CD45+ cells.
When high level (> 70%) engraftment was achieved in the peripheral blood, between 8 and 10 weeks post-transplantation, the mice were randomized and split into groups of 4 to receive either dexamethasone 15 mg/kg (Sigma-Aldrich, St Louis, MO) or vehicle control by intraperitoneal injection. Mice were culled by CO2 asphyxiation at 0 hours (pre-treatment, group 1), 8 hours (groups 2 and 3), 24 hours (groups 4 and 5) or 48 hours (groups 6 and 7) following treatment. The mice in groups 6 and 7 received a second dose of dexamethasone or vehicle control at 24 hours. Two mice succumbed early to thymoma, a well-recognized complication in NOD/SCID mice, resulting in 3 mice in each of groups 6 and 7. Cell suspensions of spleens were prepared and mononuclear cells enriched and purified to > 97% human by density gradient centrifugation using LymphoPrep (Axis-Shield, Norway), and cell viability assessed by trypan blue exclusion. RNA was extracted using the RNeasy mini kit (Qiagen, Hilden, Germany) and the RNA integrity verified (Agilent Bioanalyzer, Santa Clara, CA). The RNA was amplified using the Illumina TotalPrep RNA amplification kit (Ambion, Austin, TX) and hybridized onto Illumina WG-6_V3 chips (Illumina, San Diego, CA). The chips were scanned on the Illumina Bead Array Reader (Illumina, San Diego, CA) and gene expression analyzed. The data have been deposited in NCBI's Gene Expression Omnibus [22] and are accessible through GEO Series accession number GSE30392 http://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE30392.
Gene expression and functional analysis
The sample probe profiles with no normalization or background correction were exported from BeadStudio (version 3.0.14, Illumina, San Diego, CA). The data were pre-processed using variance stabilizing transformation [23] and robust spline normalization in lumi [24] which takes advantage of each probe being represented by > 25 beads. Differential gene expression was determined using limma [25] by comparing all treated groups to time 0 hours, with the positive False Discovery Rate correction for multiple testing [26]. Complete linkage hierarchical clustering using Euclidian distance was used to compare groups to each other. Functional analysis was performed using gene set enrichment analysis (GSEA) version 2.04 [27], comparing the limma moderated t-statistic for each probe in a pre-ranked file, against the c2_all collection of gene sets from the Molecular Signatures Database [27] version 2.5 with 1000 permutations. The similarity of the top 100 up- and down-regulated genesets was assessed using meta-GSEA (Cowley et al, manuscript in preparation).
Comparison of models
The molecular response to glucocorticoids in xenografts was compared to publicly available microarray data [13,16] using parametric analysis of gene set enrichment [28] implemented in the PGSEA package (version 1.20.1, Furge and Dykema) from the Bioconductor project [29], with some modifications to the algorithm to assess significance of the genes that are in the geneset and represented on the microarray, and to allow more control over control sample specification (available upon request). Expression levels of each gene in each sample were converted to expression ratios relative to patient matched controls before glucocorticoid treatment (Schmidt et al), time-matched controls (Rainer et al), or time 0 hours (xenografts). Within each dataset, these gene-level ratios were summarized into geneset-level Z-scores, using PGSEA with genesets from the c2_all collection [27]. The Z-scores from each sample from the 3 studies were combined and then compared by hierarchical clustering of the top 100 gene sets demonstrating the greatest variance across the combined studies.
Replicate analysis
The stability of results when reducing the number of replicates was assessed using the Recovery Score method [30] from the GeneSelector package (version 1.4.0) of the Bioconductor project [29].
Results and Discussion
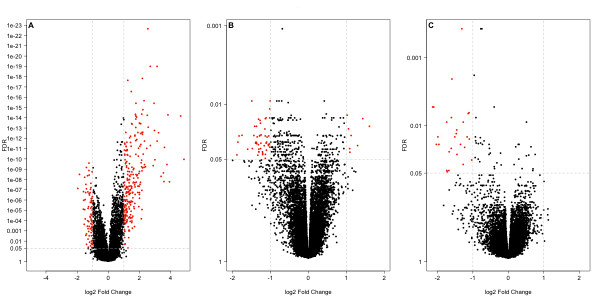
It has been demonstrated that changes in gene expression can be detected as early as 6 hours after treatment of ALL with glucocorticoids in vivo [16] and in vitro [11], although earlier timepoints show few, if any, significantly differentially expressed genes [17]. In this study the 8 hour dexamethasone-treated timepoint demonstrated the highest number of differentially expressed genes compared to baseline control, with far fewer observed at the 24 and 48 hour dexamethasone-treated timepoints (Tables 1 and 2, and Figure 1). Whilst a similar proportion of up- and down-regulated genes were identified at the 8 hour dexamethasone-treated timepoint (1158 vs 1072 respectively, FDR < 0.05), of those with large fold changes (FC > 2 or FC < 0.5, red dots in Figure 1A), 75% were up regulated (199 vs 65 respectively), consistent with the predominant role of glucocorticoids as transcriptional activators. The large numbers of statistically differentially expressed genes (FDR < 0.05) with small fold changes (0.5 < FC < 2) are indicative of both small measurement error across replicates, and thus the high reproducibility of the xenograft model, and good experimental power resulting from using 4 replicates. There was minimal significant differential gene expression across the time-matched controls (Tables 1 and 2). This demonstrates that in the xenograft mouse model, the 8 hour timepoint provides the greatest information, and that these changes are not sustained over later timepoints. The handling of the mice and intraperitoneal vehicle control injections had minimal effect on gene expression, and thus time-matched controls are redundant.
Table 1.
Number of differentially expressed genes by False Discovery Rate (FDR), compared to time 0 hours.
| Timepoint (hours) | FDR < 0.25 | FDR < 0.1 | FDR < 0.05 | |||
|---|---|---|---|---|---|---|
| + | - | + | - | + | - | |
| Dex 8 | 2313 | 2434 | 1470 | 1423 | 1158 | 1072 |
| Dex 24 | 970 | 1087 | 273 | 421 | 75 | 195 |
| Dex 48 | 321 | 327 | 41 | 95 | 12 | 44 |
| Con 8 | 0 | 0 | 0 | 0 | 0 | 0 |
| Con 24 | 0 | 1 | 0 | 1 | 0 | 1 |
| Con 48 | 0 | 1 | 0 | 1 | 0 | 1 |
+ upregulated; - downregulated; Dex, dexamethasone-treated; and Con, control
Table 2.
Number of differentially expressed genes by Fold Change (FC), compared to time 0 hours.
| Timepoint (hours) | FC > 1.5 | FC > 2 | FC > 4 | |||
|---|---|---|---|---|---|---|
| + | - | + | - | + | - | |
| Dex 8 | 501 | 429 | 201 | 68 | 38 | 0 |
| Dex 24 | 137 | 341 | 15 | 90 | 0 | 0 |
| Dex 48 | 79 | 234 | 5 | 69 | 0 | 3 |
| Con 8 | 1 | 37 | 1 | 2 | 0 | 0 |
| Con 24 | 1 | 5 | 0 | 0 | 0 | 0 |
| Con 48 | 7 | 34 | 0 | 2 | 0 | 0 |
+ upregulated; - downregulated; Dex, dexamethasone-treated; and Con, control
Figure 1.
Volcano plots of significantly differentially expressed genes following treatment with dexamethasone at 8 hours (A), 24 hours (B), 48 hours (C). Significance was defined as log2 Fold Change > 1 or < -1 with False Discovery Rate (FDR) < 0.05. Each dot represents a single gene, and significant genes indicated by red dots.
At the 8 hour timepoint, there were 173 genes upregulated with a t-statistic (the ratio of fold change to standard error) > 10 and 25 genes downregulated with a t-statistic < -10 (corresponding to P < 1.74 × 10-9 and FDR < 2.95 × 10-7, table 3). None of these genes had sustained expression changes at 24 or 48 hours, and although this could potentially reflect the early death of sensitive cells, there was no significant difference in the total number of cells harvested from the spleens at any timepoint compared to the time-matched controls (data not shown), and all harvests had a cell viability of ≥ 96%.
Table 3.
Genes regulated 8 hours following dexamethasone treatment.
| ProbeSet ID | Gene | t | P | FDR | Definition |
|---|---|---|---|---|---|
| Upregulated | |||||
| ILMN_5080450 | ZBTB16 | 83.77 | < 2.2E-16 | < 2.2E-16 | zinc finger and BTB domain containing 16 |
| ILMN_3800088 | MMP7 | 53.22 | < 2.2E-16 | < 2.2E-16 | matrix metallopeptidase 7 |
| ILMN_1770593 | CH25H | 53.14 | < 2.2E-16 | < 2.2E-16 | cholesterol 25-hydroxylase |
| ILMN_6560328 | C6orf85 | 44.60 | < 2.2E-16 | < 2.2E-16 | chromosome 6 open reading frame 85 |
| ILMN_7570561 | TSC22D3 | 39.16 | < 2.2E-16 | < 2.2E-16 | TSC22 domain family, member 3 |
| ILMN_580187 | PDE8B | 33.88 | < 2.2E-16 | 3.90E-16 | phosphodiesterase 8B |
| ILMN_5130066 | C8orf61 | 33.82 | < 2.2E-16 | 3.90E-16 | chromosome 8 open reading frame 61 |
| ILMN_4120431 | TMEM100 | 31.38 | < 2.2E-16 | 1.64E-15 | transmembrane protein 100 |
| ILMN_650553 | BIN1 | 29.76 | < 2.2E-16 | 4.43E-15 | bridging integrator 1 |
| ILMN_1400373 | SLA | 29.57 | < 2.2E-16 | 4.63E-15 | Src-like-adaptor |
| ILMN_6330593 | PTHR1 | 29.28 | < 2.2E-16 | 5.22E-15 | parathyroid hormone receptor 1 |
| ILMN_6110037 | LILRA3 | 29.04 | < 2.2E-16 | 5.75E-15 | leukocyte immunoglobulin-like receptor subfamily A, member 3 |
| ILMN_4150477 | LOXL4 | 28.67 | < 2.2E-16 | 6.66E-15 | lysyl oxidase-like 4 |
| ILMN_2680079 | OGFRL1 | 28.65 | < 2.2E-16 | 6.66E-15 | opioid growth factor receptor-like 1 |
| ILMN_4210411 | NDRG2 | 28.20 | < 2.2E-16 | 8.62E-15 | NDRG family member 2 |
| ILMN_3780093 | LILRA1 | 27.86 | < 2.2E-16 | 1.05E-14 | leukocyte immunoglobulin-like receptor subfamily A, member 1 |
| ILMN_240441 | IL1R2 | 27.46 | < 2.2E-16 | 1.33E-14 | interleukin 1 receptor, type II |
| ILMN_4730315 | MERTK | 26.14 | < 2.2E-16 | 3.31E-14 | c-mer proto-oncogene tyrosine kinase |
| ILMN_3800538 | ACPL2 | 25.90 | < 2.2E-16 | 3.72E-14 | acid phosphatase-like 2 |
| ILMN_6860392 | UGT2B17 | 25.83 | < 2.2E-16 | 3.72E-14 | UDP glucuronosyltransferase 2 family, polypeptide B17 |
| ILMN_4730541 | SLC44A1 | 25.82 | < 2.2E-16 | 3.72E-14 | solute carrier family 44, member 1 |
| ILMN_4860546 | CTHRC1 | 25.64 | < 2.2E-16 | 4.10E-14 | collagen triple helix repeat containing 1 |
| ILMN_3460270 | ZHX3 | 24.56 | < 2.2E-16 | 8.79E-14 | zinc fingers and homeoboxes 3 |
| ILMN_10639 | RASSF4 | 23.21 | < 2.2E-16 | 2.57E-13 | Ras association (RalGDS/AF-6) domain family 4 |
| ILMN_1190064 | UGT2B7 | 23.13 | < 2.2E-16 | 2.67E-13 | UDP glucuronosyltransferase 2 family, polypeptide B7 |
| ILMN_6400603 | MGC2463 | 23.06 | < 2.2E-16 | 2.71E-13 | poliovirus receptor related immunoglobulin domain containing |
| ILMN_3450187 | IRGM | 23.04 | < 2.2E-16 | 2.71E-13 | immunity-related GTPase family, M |
| ILMN_6620528 | MT1X | 22.95 | 2.40E-16 | 2.85E-13 | metallothionein 1X |
| ILMN_1260341 | IL13RA1 | 22.47 | 3.67E-16 | 4.13E-13 | interleukin 13 receptor, alpha 1 |
| ILMN_2650112 | SLC16A2 | 22.25 | 4.48E-16 | 4.91E-13 | solute carrier family 16, member 2 |
| ILMN_5570170 | PNMT | 22.01 | 5.59E-16 | 5.95E-13 | phenylethanolamine N-methyltransferase |
| ILMN_870376 | C9orf152 | 21.93 | 6.02E-16 | 6.25E-13 | chromosome 9 open reading frame 152 |
| ILMN_3190379 | TGFBR3 | 21.52 | 8.78E-16 | 8.89E-13 | transforming growth factor, beta receptor III |
| ILMN_1780142 | DSCR1 | 21.08 | 1.33E-15 | 1.31E-12 | Down syndrome critical region gene 1 |
| ILMN_2640341 | FKBP5 | 20.63 | 2.05E-15 | 1.89E-12 | FK506 binding protein 5 |
| ILMN_7610136 | LOC652626 | 20.43 | 2.48E-15 | 2.23E-12 | Leukocyte immunoglobulin-like receptor subfamily B member 2 |
| ILMN_1410609 | CORO2A | 20.34 | 2.72E-15 | 2.35E-12 | coronin, actin binding protein, 2A |
| ILMN_1780088 | TBXA2R | 20.29 | 2.84E-15 | 2.40E-12 | thromboxane A2 receptor |
| ILMN_270431 | BAALC | 20.23 | 3.02E-15 | 2.50E-12 | brain and acute leukemia, cytoplasmic |
| ILMN_6280176 | GBE1 | 20.02 | 3.72E-15 | 3.01E-12 | glucan (1,4-alpha-), branching enzyme 1 |
| ILMN_6060113 | TBX15 | 19.81 | 4.62E-15 | 3.67E-12 | T-box 15 |
| ILMN_4890743 | IQSEC1 | 19.71 | 5.09E-15 | 3.97E-12 | IQ motif and Sec7 domain 1 |
| ILMN_150056 | DPEP1 | 19.65 | 5.41E-15 | 4.13E-12 | dipeptidase 1 |
| ILMN_2060364 | BTNL9 | 19.26 | 8.04E-15 | 5.91E-12 | butyrophilin-like 9 |
| ILMN_3830735 | UPB1 | 19.23 | 8.30E-15 | 5.91E-12 | ureidopropionase, beta |
| ILMN_5670377 | STYK1 | 19.15 | 9.09E-15 | 6.35E-12 | serine/threonine/tyrosine kinase 1 |
| ILMN_4390630 | STAG3 | 18.72 | 1.42E-14 | 9.39E-12 | stromal antigen 3 |
| ILMN_4070048 | NPHP4 | 18.44 | 1.91E-14 | 1.25E-11 | nephronophthisis 4 |
| ILMN_4220474 | C6orf81 | 18.31 | 2.16E-14 | 1.39E-11 | chromosome 6 open reading frame 81 |
| ILMN_1470746 | PTPN3 | 18.30 | 2.23E-14 | 1.41E-11 | protein tyrosine phosphatase, non-receptor type 3 |
| ILMN_5860576 | C20orf133 | 18.25 | 2.36E-14 | 1.47E-11 | MACRO domain containing 2 |
| ILMN_6020468 | PPP1R14A | 18.18 | 2.52E-14 | 1.55E-11 | protein phosphatase 1, regulatory (inhibitor) subunit 14A |
| ILMN_1400634 | MT1M | 18.10 | 2.76E-14 | 1.64E-11 | metallothionein 1M |
| ILMN_4250315 | ITGA9 | 17.90 | 3.46E-14 | 2.03E-11 | integrin, alpha 9 |
| ILMN_5080471 | MAP3K6 | 17.40 | 6.02E-14 | 3.44E-11 | mitogen-activated protein kinase 6 |
| ILMN_5360242 | FLJ42461 | 17.36 | 6.28E-14 | 3.53E-11 | smoothelin-like 2 |
| ILMN_6620402 | NUDT16 | 17.33 | 6.50E-14 | 3.60E-11 | nudix (nucleoside diphosphate linked moiety X)-type motif 16 |
| ILMN_3360112 | TMEM2 | 17.26 | 7.04E-14 | 3.85E-11 | transmembrane protein 2 |
| ILMN_6840743 | PER1 | 17.22 | 7.41E-14 | 3.99E-11 | period homolog 1 |
| ILMN_4220347 | LRRC1 | 17.12 | 8.29E-14 | 4.33E-11 | leucine rich repeat containing 1 |
| ILMN_4850592 | P2RY14 | 17.11 | 8.35E-14 | 4.33E-11 | purinergic receptor P2Y, G-protein coupled, 14 |
| ILMN_6560300 | SLC31A2 | 16.91 | 1.05E-13 | 5.39E-11 | solute carrier family 31 member 2 |
| ILMN_4060091 | DKFZ | 16.87 | 1.11E-13 | 5.62E-11 | DKFZp451A211 protein |
| ILMN_6770370 | LOC92196 | 16.28 | 2.23E-13 | 1.11E-10 | death associated protein-like 1 |
| ILMN_580487 | IL9R | 16.21 | 2.40E-13 | 1.18E-10 | interleukin 9 receptor |
| ILMN_1990300 | SOCS1 | 16.18 | 2.49E-13 | 1.21E-10 | suppressor of cytokine signaling 1 |
| ILMN_5720424 | NRP1 | 16.17 | 2.54E-13 | 1.22E-10 | neuropilin 1 |
| ILMN_4180427 | CIB4 | 16.11 | 2.74E-13 | 1.30E-10 | calcium and integrin binding family member 4 |
| ILMN_4180544 | ROPN1L | 16.08 | 2.81E-13 | 1.32E-10 | ropporin 1-like |
| ILMN_4250167 | SOX13 | 16.04 | 2.95E-13 | 1.37E-10 | SRY (sex determining region Y)-box 13 |
| ILMN_6330170 | CHKA | 15.81 | 3.94E-13 | 1.81E-10 | choline kinase alpha, 3 |
| ILMN_4560192 | SFXN5 | 15.62 | 4.95E-13 | 2.25E-10 | sideroflexin 5 |
| ILMN_2810136 | CAPN11 | 15.56 | 5.33E-13 | 2.40E-10 | calpain 11 |
| ILMN_2690709 | VIPR1 | 15.38 | 6.68E-13 | 2.91E-10 | vasoactive intestinal peptide receptor 1 |
| ILMN_630091 | NCOA7 | 15.38 | 6.69E-13 | 2.91E-10 | nuclear receptor coactivator 7 |
| ILMN_5390730 | MGC17330 | 15.21 | 8.25E-13 | 3.55E-10 | phosphoinositide-3-kinase interacting protein 1 |
| ILMN_130364 | MST150 | 15.19 | 8.49E-13 | 3.62E-10 | MSTP150 |
| ILMN_3450241 | KIAA0774 | 14.95 | 1.16E-12 | 4.77E-10 | KIAA0774 |
| ILMN_2230678 | ACACB | 14.80 | 1.41E-12 | 5.76E-10 | acetyl-Coenzyme A carboxylase beta |
| ILMN_5870307 | LOC440359 | 14.78 | 1.44E-12 | 5.83E-10 | similar to muscle Y-box protein YB2 |
| ILMN_3840554 | SPOCK2 | 14.76 | 1.49E-12 | 5.95E-10 | sparc/osteonectin, cwcv and kazal-like domains 2 |
| ILMN_5810600 | MAP3K5 | 14.69 | 1.63E-12 | 6.47E-10 | mitogen-activated protein kinase 5 |
| ILMN_2360719 | IRAK3 | 14.65 | 1.71E-12 | 6.65E-10 | interleukin-1 receptor-associated kinase 3 |
| ILMN_1510121 | MTSS1 | 14.64 | 1.73E-12 | 6.66E-10 | metastasis suppressor 1 |
| ILMN_540671 | LILRB2 | 14.54 | 1.98E-12 | 7.41E-10 | leukocyte immunoglobulin-like receptor subfamily B, member 2 |
| ILMN_6980377 | MTMR15 | 14.44 | 2.26E-12 | 8.39E-10 | myotubularin related protein 15 |
| ILMN_6220288 | PRDM1 | 14.43 | 2.28E-12 | 8.39E-10 | PR domain containing 1, with ZNF domain |
| ILMN_7330739 | NDRG4 | 14.42 | 2.30E-12 | 8.39E-10 | NDRG family member 4 |
| ILMN_2600470 | WDR60 | 14.20 | 3.10E-12 | 1.12E-09 | WD repeat domain 60 |
| ILMN_4050441 | SH3MD4 | 14.16 | 3.27E-12 | 1.17E-09 | SH3 multiple domains 4 |
| ILMN_6760546 | TIPARP | 13.89 | 4.74E-12 | 1.64E-09 | TCDD-inducible poly(ADP-ribose) polymerase |
| ILMN_2760537 | MTE | 13.89 | 4.75E-12 | 1.64E-09 | metallothionein E |
| ILMN_160019 | SORT1 | 13.79 | 5.44E-12 | 1.83E-09 | sortilin 1 |
| ILMN_6330132 | ISG20 | 13.60 | 7.00E-12 | 2.32E-09 | interferon stimulated exonuclease gene 20 kDa |
| ILMN_1510685 | DOK4 | 13.52 | 7.86E-12 | 2.58E-09 | docking protein 4 |
| ILMN_1240228 | PAG1 | 13.47 | 8.50E-12 | 2.77E-09 | phosphoprotein associated glycosphingolipid microdomains 1 |
| ILMN_580592 | CPNE8 | 13.32 | 1.04E-11 | 3.31E-09 | copine VIII |
| ILMN_5870301 | KIAA0513 | 13.32 | 1.05E-11 | 3.31E-09 | KIAA0513 |
| ILMN_20129 | CD52 | 13.32 | 1.05E-11 | 3.31E-09 | CD52 molecule |
| ILMN_1820386 | PARVB | 13.31 | 1.06E-11 | 3.31E-09 | parvin, beta |
| ILMN_6200402 | MT1A | 13.24 | 1.17E-11 | 3.64E-09 | metallothionein 1A |
| ILMN_290661 | CLN8 | 13.10 | 1.43E-11 | 4.36E-09 | ceroid-lipofuscinosis, neuronal 8 |
| ILMN_670082 | GNA12 | 13.08 | 1.47E-11 | 4.43E-09 | guanine nucleotide binding protein (G protein) alpha 12 |
| ILMN_5570286 | TACC2 | 12.99 | 1.67E-11 | 5.00E-09 | transforming, acidic coiled-coil containing protein 2 |
| ILMN_3190411 | STARD13 | 12.93 | 1.81E-11 | 5.32E-09 | START domain containing 13 |
| ILMN_4540138 | NGB | 12.92 | 1.85E-11 | 5.39E-09 | neuroglobin |
| ILMN_2000646 | B4GALT4 | 12.83 | 2.10E-11 | 6.07E-09 | UDP-galactosyltransferase, polypeptide 4 |
| ILMN_7100731 | CYGB | 12.81 | 2.17E-11 | 6.17E-09 | cytoglobin |
| ILMN_7050113 | NTRK1 | 12.71 | 2.52E-11 | 7.09E-09 | neurotrophic tyrosine kinase receptor, type 1 |
| ILMN_2490670 | GNPTAB | 12.66 | 2.71E-11 | 7.52E-09 | N-acetylglucosamine-1-phosphate transferase, alpha and beta |
| ILMN_20170 | ZNF385 | 12.48 | 3.55E-11 | 9.72E-09 | zinc finger protein 385 |
| ILMN_2630687 | CHPT1 | 12.43 | 3.80E-11 | 1.02E-08 | choline phosphotransferase 1 |
| ILMN_4120215 | WASF2 | 12.43 | 3.81E-11 | 1.02E-08 | WAS protein family, member 2 |
| ILMN_5260494 | TMLHE | 12.39 | 4.06E-11 | 1.08E-08 | trimethyllysine hydroxylase, epsilon |
| ILMN_5220333 | C14orf139 | 12.31 | 4.54E-11 | 1.20E-08 | chromosome 14 open reading frame 139 |
| ILMN_3850440 | FCER1G | 12.12 | 6.07E-11 | 1.60E-08 | Fc fragment of IgE, receptor for; gamma polypeptide |
| ILMN_1030008 | TGFB3 | 12.11 | 6.21E-11 | 1.63E-08 | transforming growth factor, beta 3 |
| ILMN_1450468 | MYT1 | 12.02 | 7.04E-11 | 1.81E-08 | myelin transcription factor 1 |
| ILMN_7560541 | SLC2A5 | 12.01 | 7.19E-11 | 1.83E-08 | solute carrier family 2 member 5 |
| ILMN_2030438 | GBA2 | 12.01 | 7.21E-11 | 1.83E-08 | glucosidase, beta (bile acid) 2 |
| ILMN_6840328 | SMAD3 | 12.00 | 7.35E-11 | 1.86E-08 | SMAD family member 3 |
| ILMN_3930390 | SMAP1L | 11.91 | 8.40E-11 | 2.11E-08 | stromal membrane-associated protein 1-like |
| ILMN_7570196 | TSPAN9 | 11.90 | 8.54E-11 | 2.12E-08 | tetraspanin 9 |
| ILMN_6980546 | CACNA1I | 11.90 | 8.56E-11 | 2.12E-08 | calcium channel, voltage-dependent, T type, alpha 1I subunit |
| ILMN_1710364 | LCN6 | 11.89 | 8.72E-11 | 2.15E-08 | lipocalin 6 |
| ILMN_5360424 | RPS6KA2 | 11.77 | 1.04E-10 | 2.54E-08 | ribosomal protein S6 kinase, 90 kDa, polypeptide 2 |
| ILMN_5890193 | MS4A4A | 11.72 | 1.14E-10 | 2.75E-08 | membrane-spanning 4-domains, subfamily A, member 4 |
| ILMN_3390292 | KLF9 | 11.66 | 1.24E-10 | 2.98E-08 | Kruppel-like factor 9 |
| ILMN_5720059 | GFOD1 | 11.65 | 1.26E-10 | 3.02E-08 | glucose-fructose oxidoreductase domain containing 1 |
| ILMN_7650523 | TMEM46 | 11.57 | 1.43E-10 | 3.39E-08 | transmembrane protein 46 |
| ILMN_5700392 | LOC654000 | 11.46 | 1.70E-10 | 3.95E-08 | ribosome biogenesis protein BMS1 homolog 2 |
| ILMN_4810348 | C1orf188 | 11.40 | 1.88E-10 | 4.33E-08 | chromosome 1 open reading frame 188 |
| ILMN_4280180 | CHRNA3 | 11.39 | 1.91E-10 | 4.37E-08 | cholinergic receptor, nicotinic, alpha 3 |
| ILMN_270458 | CRISPLD1 | 11.37 | 1.96E-10 | 4.45E-08 | cysteine-rich secretory protein LCCL domain containing 1 |
| ILMN_450615 | MT2A | 11.37 | 1.97E-10 | 4.46E-08 | metallothionein 2A |
| ILMN_20470 | GRASP | 11.35 | 2.02E-10 | 4.51E-08 | GRP1-associated scaffold protein |
| ILMN_3370594 | LILRA2 | 11.35 | 2.03E-10 | 4.51E-08 | leukocyte immunoglobulin-like receptor subfamily A, member 2 |
| ILMN_5220397 | RREB1 | 11.34 | 2.05E-10 | 4.53E-08 | ras responsive element binding protein 1 |
| ILMN_1410192 | TDRD9 | 11.34 | 2.07E-10 | 4.56E-08 | tudor domain containing 9 |
| ILMN_4070259 | LOC653133 | 11.27 | 2.30E-10 | 4.99E-08 | guanine nucleotide binding protein (G protein) alpha 12 |
| ILMN_5960682 | RBPMS2 | 11.24 | 2.41E-10 | 5.21E-08 | RNA binding protein with multiple splicing 2 |
| ILMN_1440300 | SLC27A3 | 11.22 | 2.50E-10 | 5.37E-08 | solute carrier family 27, member 3 |
| ILMN_5050768 | LONRF1 | 11.20 | 2.58E-10 | 5.53E-08 | LON peptidase N-terminal domain and ring finger 1 |
| ILMN_6270273 | KHDRBS3 | 11.18 | 2.67E-10 | 5.68E-08 | KH domain, RNA binding, signal transduction associated 3 |
| ILMN_7100603 | KCNK3 | 11.17 | 2.70E-10 | 5.72E-08 | potassium channel, subfamily K, member 3 |
| ILMN_2320129 | CSDA | 11.03 | 3.38E-10 | 7.08E-08 | cold shock domain protein A |
| ILMN_3930022 | LOC644739 | 10.99 | 3.63E-10 | 7.54E-08 | Wiskott-Aldrich syndrome protein family member 4 |
| ILMN_7400133 | CUGBP2 | 10.90 | 4.20E-10 | 8.63E-08 | CUG triplet repeat, RNA binding protein 2 |
| ILMN_3290301 | FZD8 | 10.88 | 4.33E-10 | 8.76E-08 | frizzled homolog 8 |
| ILMN_7320520 | MTUS1 | 10.88 | 4.33E-10 | 8.76E-08 | mitochondrial tumor suppressor 1 |
| ILMN_3780053 | PALLD | 10.82 | 4.79E-10 | 9.60E-08 | palladin, cytoskeletal associated protein |
| ILMN_6860162 | LOC441019 | 10.74 | 5.49E-10 | 1.09E-07 | hypothetical LOC441019 |
| ILMN_5810154 | ALOX15B | 10.74 | 5.50E-10 | 1.09E-07 | arachidonate 15-lipoxygenase, type B |
| ILMN_3930736 | CHST3 | 10.73 | 5.59E-10 | 1.09E-07 | carbohydrate (chondroitin 6) sulfotransferase 3 |
| ILMN_60470 | STX11 | 10.72 | 5.68E-10 | 1.10E-07 | syntaxin 11 |
| ILMN_3390484 | SERINC2 | 10.69 | 5.95E-10 | 1.15E-07 | serine incorporator 2 |
| ILMN_1430647 | TAX1BP3 | 10.61 | 6.82E-10 | 1.31E-07 | Tax1 (human T-cell leukemia virus type I) binding protein 3 |
| ILMN_5960440 | VDR | 10.60 | 6.99E-10 | 1.34E-07 | vitamin D (1,25-dihydroxyvitamin D3) receptor |
| ILMN_6290735 | EPHB3 | 10.51 | 8.10E-10 | 1.53E-07 | EPH receptor B3 |
| ILMN_2680372 | SH2D4A | 10.46 | 8.78E-10 | 1.64E-07 | SH2 domain containing 4A |
| ILMN_2480050 | SOX7 | 10.44 | 9.13E-10 | 1.69E-07 | SRY (sex determining region Y)-box 7 |
| ILMN_130128 | LOC285016 | 10.41 | 9.61E-10 | 1.76E-07 | hypothetical protein LOC285016 |
| ILMN_4890451 | GRAMD3 | 10.39 | 9.87E-10 | 1.80E-07 | GRAM domain containing 3 |
| ILMN_770161 | C10orf73 | 10.39 | 9.92E-10 | 1.81E-07 | chromosome 10 open reading frame 73 |
| ILMN_2450202 | KIF3C | 10.35 | 1.05E-09 | 1.88E-07 | kinesin family member 3C |
| ILMN_6840468 | HAL | 10.35 | 1.06E-09 | 1.89E-07 | histidine ammonia-lyase |
| ILMN_2470070 | TBL1X | 10.30 | 1.15E-09 | 2.04E-07 | transducin (beta)-like 1X-linked |
| ILMN_2320114 | KLF13 | 10.27 | 1.22E-09 | 2.15E-07 | Kruppel-like factor 13 |
| ILMN_6380112 | DIP | 10.23 | 1.31E-09 | 2.27E-07 | death-inducing-protein |
| ILMN_2470358 | IFNGR1 | 10.22 | 1.32E-09 | 2.30E-07 | interferon gamma receptor 1 |
| ILMN_4250735 | IL27RA | 10.07 | 1.70E-09 | 2.91E-07 | interleukin 27 receptor, alpha |
| ILMN_1470215 | MAP3K8 | 10.07 | 1.72E-09 | 2.91E-07 | mitogen-activated protein kinase 8 |
| ILMN_2940373 | TACC1 | 10.06 | 1.74E-09 | 2.94E-07 | transforming, acidic coiled-coil containing protein 1 |
| Downregulated | |||||
| ILMN_770538 | LYSMD2 | -15.49 | 5.81E-13 | 2.58E-10 | LysM, putative peptidoglycan-binding, domain containing 2 |
| ILMN_7150059 | STAMBPL1 | -14.61 | 1.79E-12 | 6.84E-10 | STAM binding protein-like 1 |
| ILMN_5340692 | STRBP | -14.56 | 1.93E-12 | 7.31E-10 | spermatid perinuclear RNA binding protein |
| ILMN_4210397 | GLDC | -14.05 | 3.80E-12 | 1.34E-09 | glycine dehydrogenase |
| ILMN_6980327 | DKC1 | -13.79 | 5.44E-12 | 1.83E-09 | dyskeratosis congenita 1, dyskerin |
| ILMN_50086 | TCF12 | -13.23 | 1.19E-11 | 3.69E-09 | transcription factor 12 |
| ILMN_4860356 | BYSL | -12.81 | 2.17E-11 | 6.17E-09 | bystin-like |
| ILMN_4280228 | IVNS1ABP | -12.70 | 2.55E-11 | 7.12E-09 | influenza virus NS1A binding protein |
| ILMN_1990379 | SLFN11 | -11.82 | 9.63E-11 | 2.36E-08 | schlafen family member 11 |
| ILMN_5220338 | MPEG1 | -11.64 | 1.27E-10 | 3.03E-08 | macrophage expressed gene 1 |
| ILMN_450168 | SFRS7 | -11.50 | 1.60E-10 | 3.74E-08 | splicing factor, arginine/serine-rich 7, 35 kDa |
| ILMN_3460687 | KIAA0690 | -11.42 | 1.81E-10 | 4.19E-08 | ribosomal RNA processing 12 homolog |
| ILMN_3400360 | MAPRE2 | -11.36 | 1.99E-10 | 4.48E-08 | microtubule-associated protein, RP/EB family, member 2 |
| ILMN_4010414 | PPFIBP1 | -11.12 | 2.92E-10 | 6.16E-08 | PTPRF interacting protein, binding protein 1 (liprin beta 1) |
| ILMN_1190139 | UGT3A2 | -10.99 | 3.61E-10 | 7.54E-08 | UDP glycosyltransferase 3 family, polypeptide A2 |
| ILMN_4150201 | BCL2 | -10.93 | 3.99E-10 | 8.24E-08 | B-cell CLL/lymphoma 2 |
| ILMN_780240 | C12orf24 | -10.85 | 4.53E-10 | 9.13E-08 | chromosome 12 open reading frame 24 |
| ILMN_6760167 | MARCH3 | -10.73 | 5.60E-10 | 1.09E-07 | membrane-associated ring finger (C3HC4) 3 |
| ILMN_3940615 | PUS7 | -10.52 | 7.99E-10 | 1.52E-07 | pseudouridylate synthase 7 homolog |
| ILMN_20544 | GART | -10.41 | 9.53E-10 | 1.76E-07 | phosphoribosylglycinamide formyltransferase |
| ILMN_2480326 | HSP90B1 | -10.36 | 1.05E-09 | 1.88E-07 | heat shock protein 90 kDa beta (Grp94), member 1 |
| ILMN_5270367 | CTSC | -10.25 | 1.26E-09 | 2.20E-07 | cathepsin C |
| ILMN_5420095 | MYC | -10.21 | 1.36E-09 | 2.34E-07 | v-myc myelocytomatosis viral oncogene homolog |
| ILMN_4610180 | PIK3C2B | -10.20 | 1.38E-09 | 2.37E-07 | phosphoinositide-3-kinase, class 2, beta polypeptide |
| ILMN_6450300 | GEMIN4 | -10.00 | 1.95E-09 | 3.27E-07 | gem (nuclear organelle) associated protein 4 |
t, t-statistic; and FDR, false discovery rate
The most significantly differentially expressed gene at the 8 hour dexamethasone-treated timepoint was ZBTB16, a known transcriptional repressor and glucocorticoid response gene, which has been shown to modulate glucocorticoid sensitivity in CEM T-ALL cells [31]. Other known glucocorticoid response genes upregulated included TSC22D3 [32] and SOCS1 [15], both downstream targets of the glucocorticoid receptor, FKBP5 [33], a co-chaperone of the glucocorticoid receptor, and MAP kinases 5, 6 and 8 [34]. Downregulated genes at 8 hours included BCL-2 [35] and C-MYC [36], both previously described, but also HSP90B1, a glucocorticoid receptor co-chaperone and regulator of apoptosis. The only pro-apoptotic gene consistently upregulated across multiple microarray analyses is the BH3-only BCL-2 family member BIM, and it has been shown that BIM has a critical role in glucocorticoid sensitivity and resistance [37], although in this current study BIM was only induced 1.3 fold. Thus these genes identified are consistent with previous reports of glucocorticoid-induced genes in ALL. Within these experimental systems however there are significant potential differences in glucocorticoid exposure between in vitro and in vivo models - a crucial one is that cells in vitro are continuously exposed to glucocorticoid whereas in in vivo models the glucocorticoid is subject to pharmacokinetic and pharmacodynamic changes which more accurately reflect changes in patients.
At the later timepoints, significant differential gene expression was much less marked and predominantly downregulated. At 24 hours 5 genes were upregulated (t-statistic > 6) and 10 genes downregulated (t-statistic < -6, table 4), and at 48 hours 1 gene was upregulated (t-statistic > 6) and 15 genes downregulated (t-statistic < -6, table 5). At 24 hours, upregulated genes included NFKBIA, an inhibitor of NF-ΚB, and TRIM74, which was sustained at 48 hours, the significance of which is uncertain. Downregulated genes were those involved in cell cycle progression, including CCNF at 24 hours, and CCNF, CDC20 and AURKA at 48 hours, consistent with growth arrest.
Table 4.
Genes regulated 24 hours following dexamethasone treatment.
| ProbeSet ID | Gene | t | P | FDR | Definition |
|---|---|---|---|---|---|
| Upregulated | |||||
| ILMN_3930687 | FAM112A | 6.67 | 1.32E-06 | 0.0091 | family with sequence similarity 112, member A |
| ILMN_6620255 | TRIM74 | 6.29 | 3.06E-06 | 0.0132 | tripartite motif-containing 74 |
| ILMN_4280113 | NFKBIA | 6.23 | 3.48E-06 | 0.0138 | nuclear factor kappa B inhibitor, alpha |
| ILMN_2140136 | EMR2 | 6.10 | 4.65E-06 | 0.0149 | egf-like containing, mucin-like, hormone receptor-like 2 |
| ILMN_7000397 | ANKRD15 | 6.08 | 4.91E-06 | 0.0149 | ankyrin repeat domain 15 |
| Downregulated | |||||
| ILMN_870524 | HOXB8 | -8.60 | 2.53E-08 | 0.0011 | homeo box B8 |
| ILMN_4830520 | LOC144501 | -6.72 | 1.19E-06 | 0.0091 | hypothetical protein LOC144501 |
| ILMN_6110332 | ARHGAP19 | -6.70 | 1.24E-06 | 0.0091 | Rho GTPase activating protein 19 |
| ILMN_2970619 | ESPL1 | -6.65 | 1.38E-06 | 0.0091 | extra spindle pole bodies homolog 1 |
| ILMN_3130541 | CCNF | -6.64 | 1.43E-06 | 0.0091 | cyclin F |
| ILMN_4760577 | CENPA | -6.62 | 1.46E-06 | 0.0091 | centromere protein A |
| ILMN_4810646 | PIF1 | -6.54 | 1.76E-06 | 0.0095 | PIF1 5'-to-3' DNA helicase homolog |
| ILMN_1070762 | PSRC1 | -6.40 | 2.38E-06 | 0.0114 | proline/serine-rich coiled-coil 1 |
| ILMN_4860703 | LOC648695 | -6.19 | 3.82E-06 | 0.0138 | retinoblastoma binding protein 4 |
| ILMN_1110538 | INCENP | -6.05 | 5.19E-06 | 0.0149 | inner centromere protein antigens 135/155 kDa |
t, t-statistic; and FDR, false discovery rate
Table 5.
Genes regulated 48 hours following dexamethasone treatment.
| ProbeSet ID | Gene | t | P | FDR | Definition |
|---|---|---|---|---|---|
| Upregulated | |||||
| ILMN_6620255 | TRIM74 | 6.30 | 3.01E-06 | 0.0089 | tripartite motif-containing 74 |
| Downregulated | |||||
| ILMN_4810646 | PIF1 | -8.85 | 1.58E-08 | 0.0004 | PIF1 5'-to-3' DNA helicase homolog |
| ILMN_870524 | HOXB8 | -8.66 | 2.26E-08 | 0.0004 | homeo box B8 |
| ILMN_1450193 | LGALS1 | -8.57 | 2.66E-08 | 0.0004 | lectin, galactoside-binding, soluble, 1 (galectin 1) |
| ILMN_4760577 | CENPA | -7.64 | 1.71E-07 | 0.0018 | centromere protein A |
| ILMN_4730605 | AURKA | -7.47 | 2.42E-07 | 0.0021 | aurora kinase A |
| ILMN_1500010 | CDC20 | -6.84 | 9.09E-07 | 0.0053 | CDC20 cell division cycle 20 homolog |
| ILMN_4060064 | HMMR | -6.82 | 9.61E-07 | 0.0053 | hyaluronan-mediated motility receptor |
| ILMN_2070408 | MID1 | -6.80 | 9.97E-07 | 0.0053 | midline 1 (Opitz/BBB syndrome) |
| ILMN_2070288 | MT1E | -6.66 | 1.36E-06 | 0.0065 | metallothionein 1E |
| ILMN_1070762 | PSRC1 | -6.60 | 1.55E-06 | 0.0067 | proline/serine-rich coiled-coil 1 |
| ILMN_150543 | C20orf129 | -6.46 | 2.12E-06 | 0.0077 | chromosome 20 open reading frame 129 |
| ILMN_5870193 | FAM64A | -6.45 | 2.14E-06 | 0.0077 | family with sequence similarity 64, member A |
| ILMN_2810201 | KIF14 | -6.34 | 2.77E-06 | 0.0089 | kinesin family member 14 |
| ILMN_1050195 | KIF20A | -6.28 | 3.11E-06 | 0.0089 | kinesin family member 20A |
| ILMN_3130541 | CCNF | -6.05 | 5.21E-06 | 0.0131 | cyclin F |
t, t-statistic; and FDR, false discovery rate
Functional analysis using GSEA and meta-GSEA on the expression profiles obtained at 8 hours and 24 hours after dexamethasone treatment (additional files 1 and 2), revealed a significant upregulation of metabolic pathways, particularly adipogenesis at 8 hours, and a marked effect on pathways associated with cell cycling and proliferation, particularly downregulation of C-MYC at 8 hours and NF-ΚB at 24 hours, and upregulation of apoptotic pathways at 24 hours. Glucocorticoids are known to have effects on multiple cellular metabolic pathways, including glucose and carbohydrate metabolism, and have pro-adipogenic effects [38]. Suppression of C-MYC is a critical step prior to the initiation of apoptosis by dexamethasone in ALL [39] and suppression of NF-ΚB has been described [40].
To determine whether the molecular response to glucocorticoids in this xenograft model of ALL mimicked the effects seen in either glucocorticoid-treated patients with ALL [16] or cell-line models of ALL [13], we applied parametric gene set enrichment analysis (PGSEA) [28]. Comparing gene expression profiles from multiple experiments is notoriously difficult and typically any true similarities are swamped by technical differences in microarray vendor, normalization strategies and analytical approach. By summarizing genes at the gene set level (such as genes in the same pathway), these technical differences are mitigated, allowing comparison of samples from multiple studies.
We performed PGSEA on the 6-8 hour samples from the 3 studies, and then two-dimensional hierarchical clustering to identify the relationships between the different ALL models (Figure 2 and annotated in additional file 3). This revealed considerable heterogeneity in the molecular response to glucocorticoids in patients into at least 2, and possibly 4 different groups (green bars, Figure 2), which may represent different modes of response to glucocorticoids in patients. Relative to this inter-patient heterogeneity, both cell lines (purple bars, Figure 2) and xenografts (black bars, Figure 2) are remarkably reproducible; we anticipate that adding additional xenograft models of ALL from distinct patients will mirror the heterogeneity of the patient from whom they were derived. It is also evident that overall, glucocorticoid-treated xenografts co-cluster with a group of 3 patients (B-ALL-37, -38, and -40), all of whom had BCP-ALL and a good early prednisolone response, with varied cytogenetics (hyperploidy, t(12;21), and normal respectively). At more relaxed clustering thresholds, the glucocorticoid-treated xenografts cluster with 4 other patients with BCP-ALL (B-ALL-24, -31, -33, and 43) and the cell lines.
Figure 2.
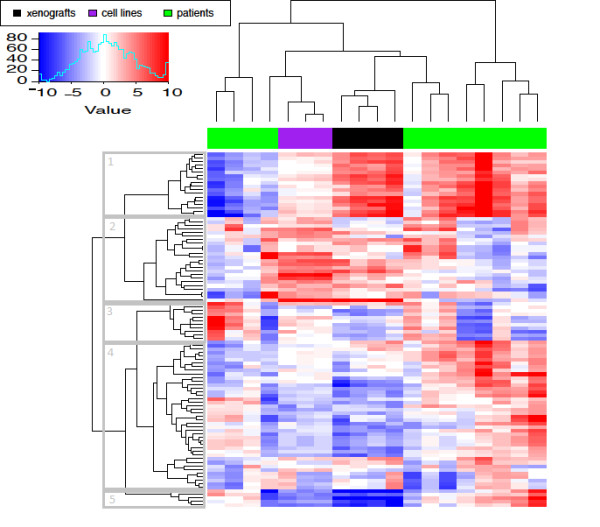
Parametric GSEA of combined top 100 glucocorticoid-induced gene sets with greatest variance from xenograft, patient and cell line models. Hierarchical clustering with gene sets in rows, samples in columns (xenografts - black, patient - green, cell line - purple). Each colour of each cell represents the Z-score (see legend). Boxes 1-5 represent defined clusters.
We identified 5 clusters of gene sets with distinct expression profiles, each behaving differently in the 3 models of ALL. Cluster 1 demonstrated the markedly heterogeneous patterns seen in patient samples, with the xenograft samples showing a pattern similar to 8 of the patients; cluster 2 showed genesets that showed strong enrichment in the cell line study, and included a number of genesets associated with cell proliferation; cluster 3 did not show any striking differences across the three ALL models; cluster 4 showed genesets downregulated in both xenografts and cell lines compared to the patient samples, and included a number genesets associated with cell cycle progression, DNA/RNA replication and MYC; cluster 5 showed genesets strongly downregulated in the xenograft and cell line models, and included genesets associated with MYC and metabolic processes, particularly catabolism and energy production. In this limited comparison, it is clear that glucocorticoid-induced gene expression patterns seen in ALL are dependent on the experimental model, and that the patterns derived from the xenograft model show a greater similarity to patient-derived data than to cell lines.
A search of the TRANSFAC database v8.3 [41] of CoMoDis [42] identified GRE motifs (within 100 kb either side of the gene) in only 25 (14.5%) of the top 173 upregulated genes at the 8 hour timepoint in this study, and no GRE motifs were identified in the upregulated genes at 24 or 48 hours. This supports accumulating evidence that glucocorticoids exert long-range effects through very distal steroid receptor binding sites [43]. Analysis of significantly differentially expressed glucocorticoid-induced genes in an in vitro cell line study [13] revealed a similar number of early response genes after 6 hours of exposure (60 upregulated (t-stat > 10) and 27 downregulated (t-stat < -10)) but a significantly greater number of genes after 24 hours (593 upregulated (t-stat > 10) and 782 downregulated (t-stat < -10)). Interestingly, all but 2 of the genes upregulated at 6 hours remained significantly upregulated at 24 hours, and 17 of the downregulated genes at 6 hours remained downregulated at 24 hours. GRE motifs were identified in 15 (25.0%) of the top 60 upregulated genes at 6 hours, and 87 (14.6%) of the top 593 genes at 24 hours. The observed difference between the studies in gene expression at later timepoints is consistent with continuous rather than physiological glucocorticoid exposure. In addition, in the cell line study, the GR (NR3C1) undergoes highly significant early and sustained autoupregulation, which in the continuous presence of ligand drives downstream gene expression. In contrast, in the xenograft model minimal GR upregulation is seen at the early timepoint but no significant change in GR expression is seen at either of the later timepoints.
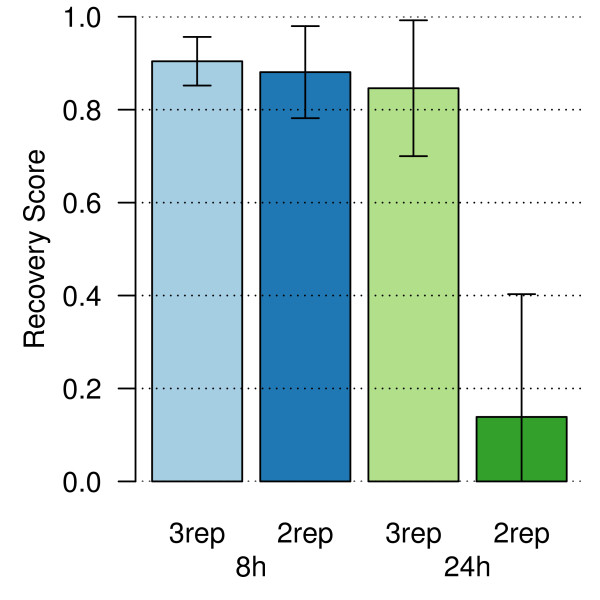
Given the good statistical power observed in Figure 1A, we proceeded to determine whether we could use fewer replicates and still identify a majority of the differentially expressed genes. Replicate analysis (Figure 3) revealed that at the 8 hour dexamethasone-treated timepoint, a dataset with high signal and differential expression, using data from any 3 randomly chosen biological replicates instead of 4 resulted in excellent recovery scores of > 0.9. That is, on average, 90% of the differentially expressed genes identified from all 4 samples were also identified in any combination of 3 arrays. At 24 hours, a timepoint with less signal, the average recovery score was 0.85 with 3 replicates, but was more variable than at 8 hours. Using just 2 biological replicates recovered 88% of the list of differentially expressed genes at 8 hours, which dropped to 14% at 24 hours. This confirms that the 8 hour time point has the strongest signal, which is reproducible across different subsets of biological replicates. We recommend using a minimum of 3 biological replicates, since fewer replicates destabilized our ability to identify differentially expressed genes. This has important considerations for experimental design, and has significant implications on cost and animal numbers.
Figure 3.
Recovery scores at 8 hours and 24 hours when randomly selecting all combinations of 3 replicates (3rep) or 2 replicates (2rep) from the set of 4 biological replicates. The Recovery Score represents the proportion of differentially expressed genes from all 4 replicates recovered when using fewer replicates.
Conclusions
We conclude that the NOD/SCID ALL xenograft mouse model provides biologically relevant insights into glucocorticoid-induced gene expression, in a consistent, reproducible and clinically relevant model system. We have demonstrated that the 8 hour timepoint provides the highest number of significantly differentially expressed genes, that time-matched controls are redundant and excellent recovery scores can be obtained with 3 replicates. We have thus established the optimal experimental design, with subsequent important implications for costs and animal numbers.
Competing interests
The authors declare that they have no competing interests.
Authors' contributions
VAB performed all experimental work and wrote the paper, VAB and MJC analyzed the data, TNT provided critical appraisal of the paper and WK and RBL designed the study. All authors read and approved the final manuscript.
Supplementary Material
metaGSEA of genesets 8 hours after treatment with dexamethasone. metaGSEA of top 100 up- and down-regulated genesets identified by Gene Set Enrichment Analysis (GSEA) 8 hours after treatment with dexamethasone.
metaGSEA of genesets 24 hours after treatment with dexamethasone. metaGSEA of top 100 up- and down-regulated genesets identified by Gene Set Enrichment Analysis (GSEA) 24 hours after treatment with dexamethasone.
Annotated pGSEA comparing glucocorticoid-induced genesets in xenograft, cell line and patient datasets. Hierarchical cluster by parametric Gene Set Enrichment Analysis (PGSEA) of the top 100 genesets with the greatest variance across three models (xenograft in vivo, cell line in vitro, patient in vivo) of glucocorticoid-induced gene expression in ALL, with annotation of the gene sets.
Contributor Information
Vivek A Bhadri, Email: vbhadri@ccia.unsw.edu.au.
Mark J Cowley, Email: m.cowley@garvan.org.au.
Warren Kaplan, Email: w.kaplan@garvan.org.au.
Toby N Trahair, Email: toby.trahair@sesiahs.health.nsw.gov.au.
Richard B Lock, Email: rlock@ccia.unsw.edu.au.
Acknowledgements of Research Support
This research was supported by Children's Cancer Institute Australia for Medical Research (CCIA) and by a grant from the National Health and Medical Research Council (NHMRC). VAB was supported by fellowships from the Leukaemia Foundation and the Steven Walter Foundation. TNT was supported by fellowships from the Cancer Institute NSW and the NHMRC. RBL was supported by a fellowship from the NHMRC. MJC and WK were supported by the Cancer Institute NSW. CCIA is affiliated to Sydney Children's Hospital and the University of New South Wales.
References
- Gaynon PS, Carrel AL. Glucocorticosteroid therapy in childhood acute lymphoblastic leukemia. Adv Exp Med Biol. 1999;457:593–605. doi: 10.1007/978-1-4615-4811-9_66. [DOI] [PubMed] [Google Scholar]
- Baxter JD, Harris AW, Tomkins GM, Cohn M. Glucocorticoid receptors in lymphoma cells in culture: relationship to glucocorticoid killing activity. Science. 1971;171(967):189–191. doi: 10.1126/science.171.3967.189. [DOI] [PubMed] [Google Scholar]
- Pratt WB, Toft DO. Steroid receptor interactions with heat shock protein and immunophilin chaperones. Endocr Rev. 1997;18(3):306–360. doi: 10.1210/er.18.3.306. [DOI] [PubMed] [Google Scholar]
- Yamamoto KR, Alberts BM. Steroid receptors: elements for modulation of eukaryotic transcription. Annu Rev Biochem. 1976;45:721–746. doi: 10.1146/annurev.bi.45.070176.003445. [DOI] [PubMed] [Google Scholar]
- Drouin J, Charron J, Gagner JP, Jeannotte L, Nemer M, Plante RK, Wrange O. Pro-opiomelanocortin gene: a model for negative regulation of transcription by glucocorticoids. J Cell Biochem. 1987;35(4):293–304. doi: 10.1002/jcb.240350404. [DOI] [PubMed] [Google Scholar]
- Ray A, Prefontaine KE. Physical association and functional antagonism between the p65 subunit of transcription factor NF-kappa B and the glucocorticoid receptor. Proc Natl Acad Sci USA. 1994;91(2):752–756. doi: 10.1073/pnas.91.2.752. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ausserlechner MJ, Obexer P, Bock G, Geley S, Kofler R. Cyclin D3 and c-MYC control glucocorticoid-induced cell cycle arrest but not apoptosis in lymphoblastic leukemia cells. Cell Death Differ. 2004;11(2):165–174. doi: 10.1038/sj.cdd.4401328. [DOI] [PubMed] [Google Scholar]
- Laane E, Tamm KP, Buentke E, Ito K, Khahariza P, Oscarsson J, Corcoran M, Bjorklund AC, Hultenby K, Lundin J. et al. Cell death induced by dexamethasone in lymphoid leukemia is mediated through initiation of autophagy. Cell Death Differ. 2009;16(7):1018–1029. doi: 10.1038/cdd.2009.46. [DOI] [PubMed] [Google Scholar]
- Webb MS, Miller AL, Johnson BH, Fofanov Y, Li T, Wood TG, Thompson EB. Gene networks in glucocorticoid-evoked apoptosis of leukemic cells. J Steroid Biochem Mol Biol. 2003;85(2-5):183–193. doi: 10.1016/S0960-0760(03)00194-8. [DOI] [PubMed] [Google Scholar]
- Tonko M, Ausserlechner MJ, Bernhard D, Helmberg A, Kofler R. Gene expression profiles of proliferating vs. G1/G0 arrested human leukemia cells suggest a mechanism for glucocorticoid-induced apoptosis. Faseb J. 2001;15(3):693–699. doi: 10.1096/fj.00-0327com. [DOI] [PubMed] [Google Scholar]
- Wang Z, Malone MH, He H, McColl KS, Distelhorst CW. Microarray analysis uncovers the induction of the proapoptotic BH3-only protein Bim in multiple models of glucocorticoid-induced apoptosis. J Biol Chem. 2003;278(26):23861–23867. doi: 10.1074/jbc.M301843200. [DOI] [PubMed] [Google Scholar]
- Medh RD, Webb MS, Miller AL, Johnson BH, Fofanov Y, Li T, Wood TG, Luxon BA, Thompson EB. Gene expression profile of human lymphoid CEM cells sensitive and resistant to glucocorticoid-evoked apoptosis. Genomics. 2003;81(6):543–555. doi: 10.1016/S0888-7543(03)00045-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rainer J, Ploner C, Jesacher S, Ploner A, Eduardoff M, Mansha M, Wasim M, Panzer-Grumayer R, Trajanoski Z, Niederegger H. et al. Glucocorticoid-regulated microRNAs and mirtrons in acute lymphoblastic leukemia. Leukemia. 2009;23(4):746–752. doi: 10.1038/leu.2008.370. [DOI] [PubMed] [Google Scholar]
- Obexer P, Certa U, Kofler R, Helmberg A. Expression profiling of glucocorticoid-treated T-ALL cell lines: rapid repression of multiple genes involved in RNA-, protein- and nucleotide synthesis. Oncogene. 2001;20(32):4324–4336. doi: 10.1038/sj.onc.1204573. [DOI] [PubMed] [Google Scholar]
- Yoshida NL, Miyashita T, U M, Yamada M, Reed JC, Sugita Y, Oshida T. Analysis of gene expression patterns during glucocorticoid-induced apoptosis using oligonucleotide arrays. Biochem Biophys Res Commun. 2002;293(4):1254–1261. doi: 10.1016/S0006-291X(02)00361-3. [DOI] [PubMed] [Google Scholar]
- Schmidt S, Rainer J, Riml S, Ploner C, Jesacher S, Achmuller C, Presul E, Skvortsov S, Crazzolara R, Fiegl M. et al. Identification of glucocorticoid-response genes in children with acute lymphoblastic leukemia. Blood. 2006;107(5):2061–2069. doi: 10.1182/blood-2005-07-2853. [DOI] [PubMed] [Google Scholar]
- Tissing WJ, den Boer ML, Meijerink JP, Menezes RX, Swagemakers S, van der Spek PJ, Sallan SE, Armstrong SA, Pieters R. Genomewide identification of prednisolone-responsive genes in acute lymphoblastic leukemia cells. Blood. 2007;109(9):3929–3935. doi: 10.1182/blood-2006-11-056366. [DOI] [PubMed] [Google Scholar]
- Drexler HG, Fombonne S, Matsuo Y, Hu ZB, Hamaguchi H, Uphoff CC. p53 alterations in human leukemia-lymphoma cell lines: in vitroartifact or prerequisite for cell immortalization? Leukemia. 2000;14(1):198–206. doi: 10.1038/sj.leu.2401604. [DOI] [PubMed] [Google Scholar]
- Lock RB, Liem N, Farnsworth ML, Milross CG, Xue C, Tajbakhsh M, Haber M, Norris MD, Marshall GM, Rice AM. The nonobese diabetic/severe combined immunodeficient (NOD/SCID) mouse model of childhood acute lymphoblastic leukemia reveals intrinsic differences in biologic characteristics at diagnosis and relapse. Blood. 2002;99(11):4100–4108. doi: 10.1182/blood.V99.11.4100. [DOI] [PubMed] [Google Scholar]
- Liem NL, Papa RA, Milross CG, Schmid MA, Tajbakhsh M, Choi S, Ramirez CD, Rice AM, Haber M, Norris MD. et al. Characterization of childhood acute lymphoblastic leukemia xenograft models for the preclinical evaluation of new therapies. Blood. 2004;103(10):3905–3914. doi: 10.1182/blood-2003-08-2911. [DOI] [PubMed] [Google Scholar]
- Dordelmann M, Reiter A, Borkhardt A, Ludwig WD, Gotz N, Viehmann S, Gadner H, Riehm H, Schrappe M. Prednisone response is the strongest predictor of treatment outcome in infant acute lymphoblastic leukemia. Blood. 1999;94(4):1209–1217. [PubMed] [Google Scholar]
- Edgar R, Domrachev M, Lash AE. Gene Expression Omnibus: NCBI gene expression and hybridization array data repository. Nucleic Acids Res. 2002;30(1):207–210. doi: 10.1093/nar/30.1.207. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lin SM, Du P, Huber W, Kibbe WA. Model-based variance-stabilizing transformation for Illumina microarray data. Nucleic Acids Res. 2008;36(2):e11. doi: 10.1093/nar/gkm1075. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Du P, Kibbe WA, Lin SM. lumi: a pipeline for processing Illumina microarray. Bioinformatics. 2008;24(13):1547–1548. doi: 10.1093/bioinformatics/btn224. [DOI] [PubMed] [Google Scholar]
- Smyth GK. Linear models and empirical bayes methods for assessing differential expression in microarray experiments. Stat Appl Genet Mol Biol. 2004;3:Article3. doi: 10.2202/1544-6115.1027. [DOI] [PubMed] [Google Scholar]
- Storey JD, Tibshirani R. Statistical significance for genomewide studies. Proc Natl Acad Sci USA. 2003;100(16):9440–9445. doi: 10.1073/pnas.1530509100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Subramanian A, Tamayo P, Mootha VK, Mukherjee S, Ebert BL, Gillette MA, Paulovich A, Pomeroy SL, Golub TR, Lander ES. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc Natl Acad Sci USA. 2005;102(43):15545–15550. doi: 10.1073/pnas.0506580102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kim SY, Volsky DJ. PAGE: parametric analysis of gene set enrichment. BMC Bioinformatics. 2005;6:144. doi: 10.1186/1471-2105-6-144. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gentleman RC, Carey VJ, Bates DM, Bolstad B, Dettling M, Dudoit S, Ellis B, Gautier L, Ge Y, Gentry J. et al. Bioconductor: open software development for computational biology and bioinformatics. Genome Biol. 2004;5(10):R80. doi: 10.1186/gb-2004-5-10-r80. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pavlidis P, Li Q, Noble WS. The effect of replication on gene expression microarray experiments. Bioinformatics. 2003;19(13):1620–1627. doi: 10.1093/bioinformatics/btg227. [DOI] [PubMed] [Google Scholar]
- Wasim M, Carlet M, Mansha M, Greil R, Ploner C, Trockenbacher A, Rainer J, Kofler R. PLZF/ZBTB16, a glucocorticoid response gene in acute lymphoblastic leukemia, interferes with glucocorticoid-induced apoptosis. J Steroid Biochem Mol Biol. 2010;120(4-5):218–227. doi: 10.1016/j.jsbmb.2010.04.019. [DOI] [PMC free article] [PubMed] [Google Scholar]
- D'Adamio F, Zollo O, Moraca R, Ayroldi E, Bruscoli S, Bartoli A, Cannarile L, Migliorati G, Riccardi C. A new dexamethasone-induced gene of the leucine zipper family protects T lymphocytes from TCR/CD3-activated cell death. Immunity. 1997;7(6):803–812. doi: 10.1016/S1074-7613(00)80398-2. [DOI] [PubMed] [Google Scholar]
- Vermeer H, Hendriks-Stegeman BI, van der Burg B, van Buul-Offers SC, Jansen M. Glucocorticoid-induced increase in lymphocytic FKBP51 messenger ribonucleic acid expression: a potential marker for glucocorticoid sensitivity, potency, and bioavailability. J Clin Endocrinol Metab. 2003;88(1):277–284. doi: 10.1210/jc.2002-020354. [DOI] [PubMed] [Google Scholar]
- Lu J, Quearry B, Harada H. p38-MAP kinase activation followed by BIM induction is essential for glucocorticoid-induced apoptosis in lymphoblastic leukemia cells. FEBS Lett. 2006;580(14):3539–3544. doi: 10.1016/j.febslet.2006.05.031. [DOI] [PubMed] [Google Scholar]
- Laane E, Panaretakis T, Pokrovskaja K, Buentke E, Corcoran M, Soderhall S, Heyman M, Mazur J, Zhivotovsky B, Porwit A. et al. Dexamethasone-induced apoptosis in acute lymphoblastic leukemia involves differential regulation of Bcl-2 family members. Haematologica. 2007;92(11):1460–1469. doi: 10.3324/haematol.10543. [DOI] [PubMed] [Google Scholar]
- Thulasi R, Harbour DV, Thompson EB. Suppression of c-myc is a critical step in glucocorticoid-induced human leukemic cell lysis. J Biol Chem. 1993;268(24):18306–18312. [PubMed] [Google Scholar]
- Bachmann PS, Gorman R, Mackenzie KL, Lutze-Mann L, Lock RB. Dexamethasone resistance in B-cell precursor childhood acute lymphoblastic leukemia occurs downstream of ligand-induced nuclear translocation of the glucocorticoid receptor. Blood. 2005;105(6):2519–2526. doi: 10.1182/blood-2004-05-2023. [DOI] [PubMed] [Google Scholar]
- Pantoja C, Huff JT, Yamamoto KR. Glucocorticoid signaling defines a novel commitment state during adipogenesis in vitro. Mol Biol Cell. 2008;19(10):4032–4041. doi: 10.1091/mbc.E08-04-0420. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Medh RD, Wang A, Zhou F, Thompson EB. Constitutive expression of ectopic c-Myc delays glucocorticoid-evoked apoptosis of human leukemic CEM-C7 cells. Oncogene. 2001;20(34):4629–4639. doi: 10.1038/sj.onc.1204680. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Auphan N, DiDonato JA, Rosette C, Helmberg A, Karin M. Immunosuppression by glucocorticoids: inhibition of NF-kappa B activity through induction of I kappa B synthesis. Science. 1995;270(5234):286–290. doi: 10.1126/science.270.5234.286. [DOI] [PubMed] [Google Scholar]
- Matys V, Fricke E, Geffers R, Gossling E, Haubrock M, Hehl R, Hornischer K, Karas D, Kel AE, Kel-Margoulis OV. et al. TRANSFAC: transcriptional regulation, from patterns to profiles. Nucleic Acids Res. 2003;31(1):374–378. doi: 10.1093/nar/gkg108. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Donaldson IJ, Gottgens B. CoMoDis: composite motif discovery in mammalian genomes. Nucleic Acids Res. 2007;35(1):e1. doi: 10.1093/nar/gkl839. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Paakinaho V, Makkonen H, Jaaskelainen T, Palvimo JJ. Glucocorticoid receptor activates poised FKBP51 locus through long-distance interactions. Mol Endocrinol. 2010;24(3):511–525. doi: 10.1210/me.2009-0443. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
metaGSEA of genesets 8 hours after treatment with dexamethasone. metaGSEA of top 100 up- and down-regulated genesets identified by Gene Set Enrichment Analysis (GSEA) 8 hours after treatment with dexamethasone.
metaGSEA of genesets 24 hours after treatment with dexamethasone. metaGSEA of top 100 up- and down-regulated genesets identified by Gene Set Enrichment Analysis (GSEA) 24 hours after treatment with dexamethasone.
Annotated pGSEA comparing glucocorticoid-induced genesets in xenograft, cell line and patient datasets. Hierarchical cluster by parametric Gene Set Enrichment Analysis (PGSEA) of the top 100 genesets with the greatest variance across three models (xenograft in vivo, cell line in vitro, patient in vivo) of glucocorticoid-induced gene expression in ALL, with annotation of the gene sets.