Abstract
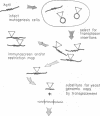
A "transplason" mutagenesis procedure was developed for the dual purposes of low resolution mapping of antigenic coding regions (using transposons) and constructing insertion mutations in yeast genes (by transplacement). Mini-Tn10 transposon derivatives containing both Escherichia coli and yeast selectable markers have been constructed. These elements are used to mutagenize lambda gt11 clones that express foreign antigens in E. coli. The transposition events are first selected in E. coli, and the effect of these insertions on antigen expression is used to locate the antigenic coding regions on the cloned DNA. Insertion mutations located within a desired yeast sequence are then substituted for the genomic copies by one-step gene transplacement. This provides a powerful method for rapidly mapping antigenic coding sequences of cloned genes and inactivating these genes in yeast to help determine their function. Several examples using this technique are presented.
Full text
PDF
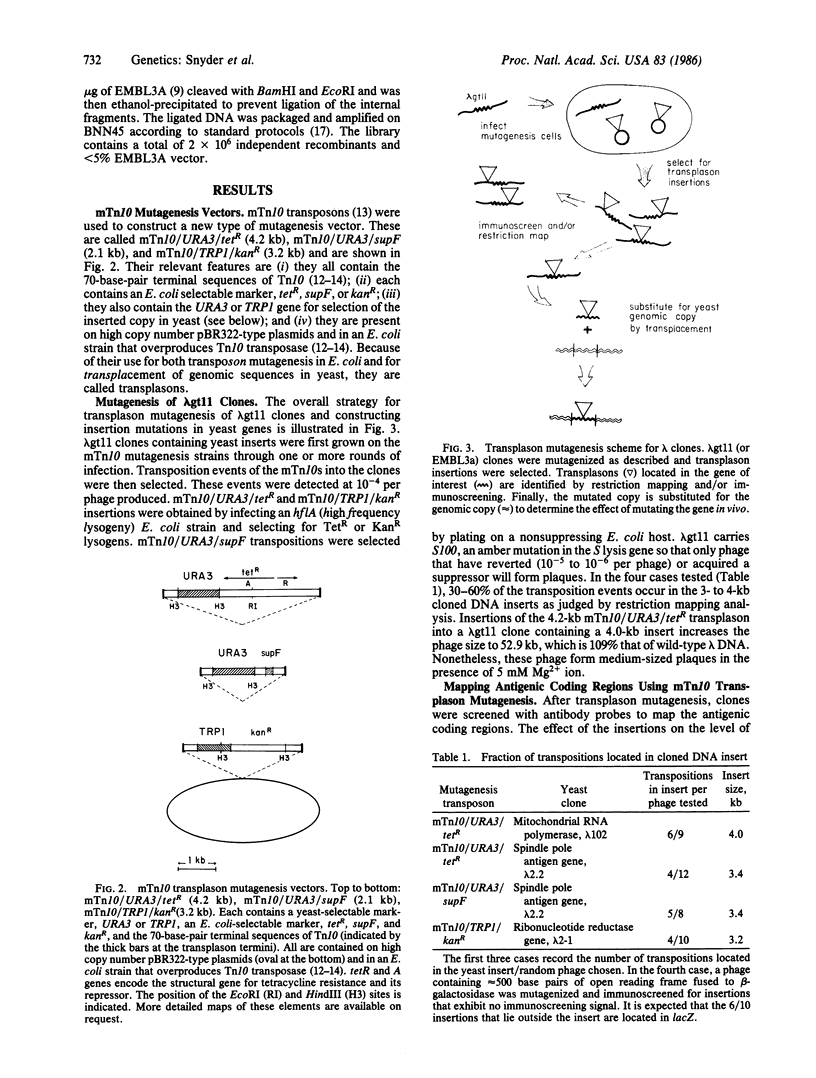




Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Botstein D., Falco S. C., Stewart S. E., Brennan M., Scherer S., Stinchcomb D. T., Struhl K., Davis R. W. Sterile host yeasts (SHY): a eukaryotic system of biological containment for recombinant DNA experiments. Gene. 1979 Dec;8(1):17–24. doi: 10.1016/0378-1119(79)90004-0. [DOI] [PubMed] [Google Scholar]
- Elledge S. J., Walker G. C. Phasmid vectors for identification of genes by complementation of Escherichia coli mutants. J Bacteriol. 1985 May;162(2):777–783. doi: 10.1128/jb.162.2.777-783.1985. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Foster T. J., Davis M. A., Roberts D. E., Takeshita K., Kleckner N. Genetic organization of transposon Tn10. Cell. 1981 Jan;23(1):201–213. doi: 10.1016/0092-8674(81)90285-3. [DOI] [PubMed] [Google Scholar]
- Frischauf A. M., Lehrach H., Poustka A., Murray N. Lambda replacement vectors carrying polylinker sequences. J Mol Biol. 1983 Nov 15;170(4):827–842. doi: 10.1016/s0022-2836(83)80190-9. [DOI] [PubMed] [Google Scholar]
- Goto T., Wang J. C. Yeast DNA topoisomerase II is encoded by a single-copy, essential gene. Cell. 1984 Apr;36(4):1073–1080. doi: 10.1016/0092-8674(84)90057-6. [DOI] [PubMed] [Google Scholar]
- Hieter P., Mann C., Snyder M., Davis R. W. Mitotic stability of yeast chromosomes: a colony color assay that measures nondisjunction and chromosome loss. Cell. 1985 Feb;40(2):381–392. doi: 10.1016/0092-8674(85)90152-7. [DOI] [PubMed] [Google Scholar]
- Ito H., Fukuda Y., Murata K., Kimura A. Transformation of intact yeast cells treated with alkali cations. J Bacteriol. 1983 Jan;153(1):163–168. doi: 10.1128/jb.153.1.163-168.1983. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Johnson L. M., Snyder M., Chang L. M., Davis R. W., Campbell J. L. Isolation of the gene encoding yeast DNA polymerase I. Cell. 1985 Nov;43(1):369–377. doi: 10.1016/0092-8674(85)90042-x. [DOI] [PubMed] [Google Scholar]
- Jorgensen R. A., Rothstein S. J., Reznikoff W. S. A restriction enzyme cleavage map of Tn5 and location of a region encoding neomycin resistance. Mol Gen Genet. 1979;177(1):65–72. doi: 10.1007/BF00267254. [DOI] [PubMed] [Google Scholar]
- Maurer R., Osmond B. C., Shekhtman E., Wong A., Botstein D. Functional interchangeability of DNA replication genes in Salmonella typhimurium and Escherichia coli demonstrated by a general complementation procedure. Genetics. 1984 Sep;108(1):1–23. doi: 10.1093/genetics/108.1.1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ozkaynak E., Finley D., Varshavsky A. The yeast ubiquitin gene: head-to-tail repeats encoding a polyubiquitin precursor protein. Nature. 1984 Dec 13;312(5995):663–666. doi: 10.1038/312663a0. [DOI] [PubMed] [Google Scholar]
- Rothstein R. J. One-step gene disruption in yeast. Methods Enzymol. 1983;101:202–211. doi: 10.1016/0076-6879(83)01015-0. [DOI] [PubMed] [Google Scholar]
- Scherer S., Davis R. W. Replacement of chromosome segments with altered DNA sequences constructed in vitro. Proc Natl Acad Sci U S A. 1979 Oct;76(10):4951–4955. doi: 10.1073/pnas.76.10.4951. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Seed B. Purification of genomic sequences from bacteriophage libraries by recombination and selection in vivo. Nucleic Acids Res. 1983 Apr 25;11(8):2427–2445. doi: 10.1093/nar/11.8.2427. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shortle D., Botstein D. Directed mutagenesis with sodium bisulfite. Methods Enzymol. 1983;100:457–468. doi: 10.1016/0076-6879(83)00073-7. [DOI] [PubMed] [Google Scholar]
- Way J. C., Davis M. A., Morisato D., Roberts D. E., Kleckner N. New Tn10 derivatives for transposon mutagenesis and for construction of lacZ operon fusions by transposition. Gene. 1984 Dec;32(3):369–379. doi: 10.1016/0378-1119(84)90012-x. [DOI] [PubMed] [Google Scholar]
- Young R. A., Bloom B. R., Grosskinsky C. M., Ivanyi J., Thomas D., Davis R. W. Dissection of Mycobacterium tuberculosis antigens using recombinant DNA. Proc Natl Acad Sci U S A. 1985 May;82(9):2583–2587. doi: 10.1073/pnas.82.9.2583. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Young R. A., Davis R. W. Efficient isolation of genes by using antibody probes. Proc Natl Acad Sci U S A. 1983 Mar;80(5):1194–1198. doi: 10.1073/pnas.80.5.1194. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Young R. A., Davis R. W. Yeast RNA polymerase II genes: isolation with antibody probes. Science. 1983 Nov 18;222(4625):778–782. doi: 10.1126/science.6356359. [DOI] [PubMed] [Google Scholar]