Summary
This review focuses on recent structural insights into regulation and nucleic acid binding of Superfamily 2 (SF2)-type helicases as they relate to chromatin remodelers. We review structural features of the Chd1 chromatin remodeler regarding regulation of the ATPase motor, and discuss related strategies observed for other SF2 ATPases. Since no SWI2/SNF2 ATPases have yet been captured bound to DNA in a state competent for ATP-hydrolysis, we turn to structural examples from the DEAD-box RNA helicase family, and suggest that SWI2/SNF2-specific inserts may be poised to alter canonical duplex DNA structure.
Keywords: Chromatin remodeling, SF2 ATPase, SWI2, SNF2, DEAD-box RNA helicase, Chd1, Rad54, RapA, eIF4AIII, eIF4A, DDX19, Dbp5, Mss116, Hef, Hda3
Introduction
Helicase-type proteins span several superfamilies that encompass functionally diverse collections of proteins involved in all aspects of nucleic acid processing, metabolism, and regulation [1]. Chromatin remodelers constitute one specialized family within helicase Superfamily 2 (SF2), and were originally named for their ability to alter the position and structure of nucleosomes. The ATPase motor, called SWI2/SNF2 after the first chromatin remodeler studied, has been found in a wide range of proteins, not all of which target the nucleosome but are aimed at disrupting distinct protein-nucleic acid complexes [2, 3]. Within the SWI2/SNF2 family, the ATPase motor is typically accompanied by multiple auxiliary domains, which presumably modify action of the ATPase through targeting and regulation.
Like all helicase Superfamily 1 (SF1) and SF2 ATPases, SWI2/SNF2 ATPases consist of two covalently linked RecA-like domains that form a bi-lobed motor (Box 1). The architecture of SF1 and SF2 ATPases has been extensively reviewed [1, 4, 5], therefore we will focus on particular elements relevant to regulation and DNA binding of the ATPase motor. As observed in many structural examples of SF1 and SF2 ATPases, unique domains or sub-domains are often found closely associated with the bi-lobed motor, and can be attached at the amino- or carboxyl-terminus, or protrude as extended loops from one or both of the RecA-like domains. These unique domains have been shown to aid the ATPase motor in binding to particular substrates, or participate in distortion of bound nucleic acids [4].
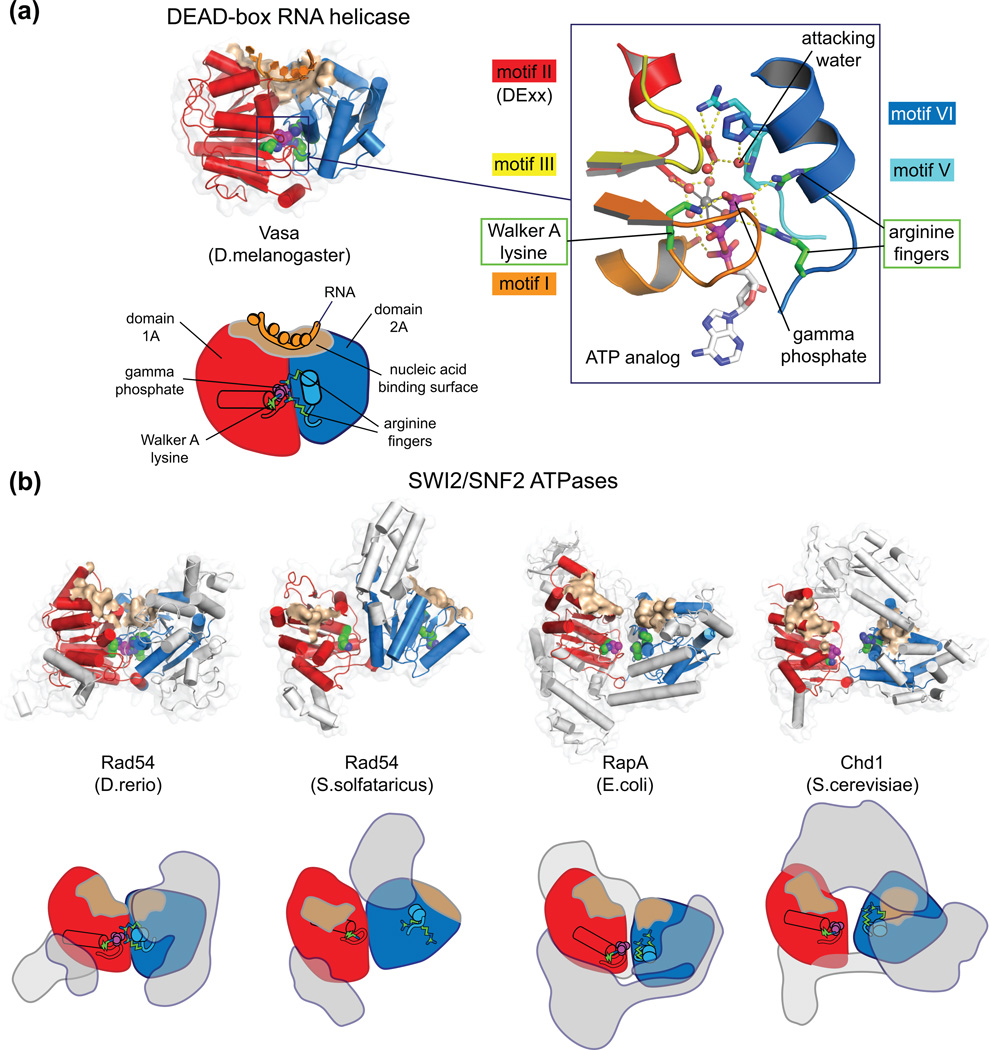
BOX 1. Core features of SF2 ATPases.
(a) As illustrated by the structure of the DEAD-box RNA helicase Vasa in complex with an RNA oligomer (2DB3 [30]), SF2 ATPase motors consist of two covalently linked RecA-like domains that together form a centrally located pocket for ATP hydrolysis [1, 4, 5]. Formation of a closed, catalytically active conformation requires an intricate clustering of conserved residues in the central ATPase cleft. These highly conserved residues reside on segments called helicase motifs [44], which couple closure of the central cleft, nucleic acid binding, and hydrolysis. When in a closed, active configuration, the nucleic acid binding surfaces on each half of the ATPase motor align to form a contiguous DNA or RNA binding groove across the central cleft. An extensive database for RNA helicase structures and sequences has been described by Jankowsky and colleagues [45].
(b) Backbone and schematic representations of four SWI2/SNF2 ATPase structures, highlighting the locations of catalytically important residues. Shown as green spheres are the Walker A lysine from motif I on domain 1A, essential for proper binding of the triphosphate tail, and two arginine finger residues on motif VI of domain 2A, required for ATP hydrolysis. When present, phosphate or sulfate ions bound in the P-loop are shown as magenta spheres. The positions of these catalytic residues, in conjunction with the location of the DNA-binding surfaces (brown), reveal how much the ATPase motors deviate from a closed, catalytically competent organization. As observed with other SF2 ATPases, SWI2/SNF2 structures display a variety of conformational states, suggesting that the interdomain linker is flexible [2]. The ATPase motor in the Danio rerio (zebrafish) Rad54 structure (1Z3I [6]) is relatively closed, whereas the two ATPase lobes for S. solfataricus Rad54 (1Z63 [7]) are flipped ~180° relatively to the active configuration. Elements outside the conserved SF2 ATPase core, including the SWI2/SNF2-specific subdomains and other subfamily-specific domains, are colored gray. As illustrated by the full-length RapA structure (3DMQ [8]) and Chd1 (3MWY [9]), non-ATPase domains can constitute a significant fraction of the polypeptide chain. Unlike Chd1, which is believed to be monomeric [46], most chromatin remodelers posses multiple, non-covalently associated subunits that can potentially influence action of the ATPase motor [47].
To date, crystal structures of only a few representatives containing the SWI2/SNF2 ATPase motor have been solved: Rad54 from zebrafish [6••] and Sulfolobus solfataricus [7••], E. coli Rap1 [8•], and S. cerevisiae Chd1 [9••]. These structures have revealed the common structural features of SWI2/SNF2 ATPases and provided several examples of how the ATPase motor interacts with auxiliary domains. We begin with a description of how the SWI2/SNF2 ATPase motor of the Chd1 remodeler is negatively regulated by a pair of chromodomains that can block the DNA-binding surface. Although no SWI2/SNF2 ATPases have yet been solved in an active, hydrolysis competent configuration, the S. solfataricus Rad54 structure in complex with duplex DNA demonstrated that the first ATPase domain (1A) contacts nucleic acid at the same location as observed in other SF2 ATPases. To extend our structural understanding of SWI2/SNF2 proteins, we will draw parallels with the well-studied DEAD-box RNA helicase family, a distinct but related family of SF2 ATPases for which many structures have been solved in various activated and inhibited states. We conclude with a structural comparison of SWI2/SNF2-specific inserts to nucleic acid disrupting elements found in other SF2 ATPases. The sequence and structural conservation of these inserts suggests that translocation by SWI2/SNF2 ATPases may be accompanied by localized distortion of the DNA duplex.
Keeping false substrates at bay: blocking the nucleic acid binding site
The crystal structure of the chromodomain-ATPase portion of Chd1 revealed an apparent autoinhibited organization, where the double chromodomains pack against both halves of the ATPase motor, spanning the central cleft (Figure 1a; [9••]). The conformation of the ATPase motor appears to be in an inactive form, which like many SF2 ATPases without nucleic acid substrates, displays an opened conformation and thus lacks the close association of helicase motifs across the central cleft that are critical for hydrolysis (Box 1). Although DNA was not present in these crystals, surfaces expected to bind to DNA were identified by comparison to other SF2 ATPases solved with nucleic acid substrates. Superposition of Chd1 with other SF2 ATPases revealed that the site where nucleic acid strands contact the second ATPase lobe (domain 2A) was occupied by a helix linking the two chromodomains (Figure 1a). This chromodomain linker helix contains a number of conserved acidic positions, providing a high density of negative charges that complements the positively charged surface expected to bind to DNA.
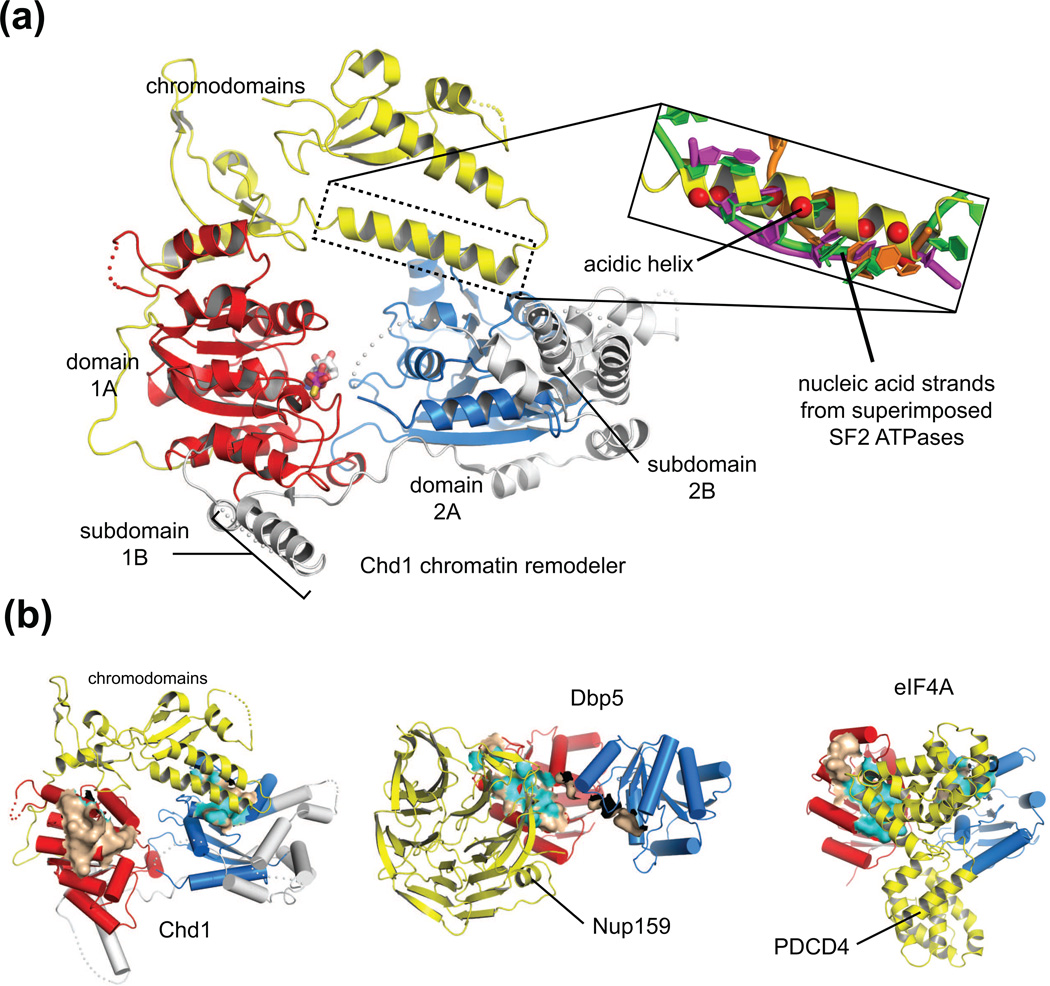
Figure 1. Blocking nucleic acid binding surfaces is a common mode of regulation for SF2 ATPases.
(a) Overview of Chd1 structure showing that an acidic helix between the chromodomains packs against a surface important for DNA binding in other SF2 ATPases. To illustrate the overlap of the Chd1 acidic helix with the DNA-binding surface, three SF2 proteins in a hydrolysis-competent state and bound to nucleic acid were superimposed on Chd1 (3MWY [9]), using domain 2A (blue) as the reference. In the inset, only the single-stranded nucleic acids are shown for Dbp5 (orange, 3FHT [11]), NS3 (magenta, 1A1V [48]), and Hel308 (green, 2P6R [49]), with the Cα positions of acidic residues on the Chd1 helix shown as red spheres.
(b) Surfaces expected to bind to nucleic acids on domains 1A (red) or 2A (blue) of several SF2 ATPase motors are occluded by non-ATPase domains (yellow). Nucleic acid binding surfaces were approximated as residues within 4 Å of the RNA bound to the Vasa helicase (2DB3 [30]), using each ATPase domain separately as a reference for superpositions. These approximated nucleic acid binding surfaces are colored brown and highlighted with cyan where the surface is within 4 Å and thus sterically blocked by a non-ATPase domain. Note that Chd1 chromodomains block ATPase domain 2A, Nup159 blocks ATPase domain 1A, and PDCD4 blocks surfaces on both domains 1A and 2A.
This interaction between the ATPase motor and the acidic helix of the chromodomains enables Chd1 to discriminate between nucleosomes and naked DNA. Normally, ATPase activity of Chd1 is preferentially stimulated by nucleosome substrates over naked DNA. However, disruption of acidic residues on this helix reduced discrimination against naked DNA, and removal of both chromodomains resulted in equal stimulation by DNA and nucleosome substrates [9••]. By physically interacting with a DNA-binding surface on the ATPase motor, the chromodomains may provide a regulatory gate: when in contact with the DNA-binding surface of the ATPase motor, the chromodomains dampen activation of the ATPase motor, whereas this inhibitory interaction can be relieved by nucleosomes, which allow the ATPase motor to productively engage with DNA.
A similar strategy for inhibiting an ATPase motor though occlusion of DNA-binding surfaces has also recently been observed for two eukaryotic DEAD-box RNA helicases. Yeast Dbp5 (called DDX19 in humans) plays an essential role in export of mRNAs from the nucleus, promoting exchange of mRNA binding proteins as they pass through the nuclear pore [10]. Dbp5 can form a complex with the nuclear pore protein Nup159 (also called Nup214), and this interaction partially occludes an RNA-binding surface on domain 1A of Dbp5 [11•, 12•, 13••] (Figure 1b). Nup159 thus effectively reduces the ability of Dbp5 to bind to RNA, and similarly to Chd1, this interaction blocks RNA-dependent ATP hydrolysis by Dbp5 [11•].
Another system where competition with nucleic acid substrates was observed is that of the initiation factor required for translation, eIF4A, which binds to the 5’ end of RNA transcripts and helps present them to the ribosome [14]. As a critical step in gene expression, translation initiation is tightly controlled, and one negative regulator of eIF4A is PDCD4 (programmed cell death protein 4) [15]. Two studies revealed how PDCD4 inhibits eIF4A by blocking the RNA binding site [16•, 17•]. PDCD4 consists of two similarly folded MA3 domains, and it was found that each of these domains could wedge itself in the central cleft of eIF4A. The interactions of PDCD4 with eIF4A are extensive and occlude the RNA-binding surfaces on both sides of the central ATPase cleft (Figure 1b). This situation differs from those seen for Chd1 and Dbp5, where the nucleic acid binding site is only blocked on one domain of the ATPase motor. Although not presently clear, the consequences of binding both nucleic acid binding surfaces may reflect the role of PDCD4 more as a true inhibitor of eIF4A action, whereas the partial blocks achieved by Chd1 chromodomains and Nup159 may be indicative of an ability to increase specificity and promote turnover.
Influencing RecA-like domain orientation as a means of regulating activity
In addition to blocking the DNA binding site, SF2 helicase activity may be influenced by preventing proper closure of the RecA-like domains. For the PDCD4-eIF4A complexes, interactions of PDCD4 with the ATPase cleft not only block nucleic acid binding surfaces, but also appear to stabilize a splayed open conformation of the helicase incompatible with hydrolysis [16•, 17•] (Figure 1b). For Chd1, the opened organization of the ATPase motor coupled with contacts made by chromodomains suggests that the chromodomains may stabilize an inactive state in the absence of nucleosomes [9]. Another example where domain organization appears to negatively regulate ATPase activity has been seen with DDX19, the human ortholog of Dbp5. In the absence of activators, a helix at the DDX19 N-terminus packs between both domains of the ATPase motor in a manner that would sterically interfere with closure of the central ATPase cleft [18•] (Figure 2a). Unlike PDCD4, the DDX19 N-terminal helix does not contact nucleic acid binding surfaces, but blocks ATP hydrolysis by wedging itself between conserved helicase motifs on either side of the cleft, thus preventing the proper organization of residues necessary for ATP hydrolysis. This inhibitory N-terminal helix was shown to make the protein dependent on RNA binding for ATPase activation, as deletion of an N-terminal segment including this helix promoted RNA-independent ATP hydrolysis [18•].
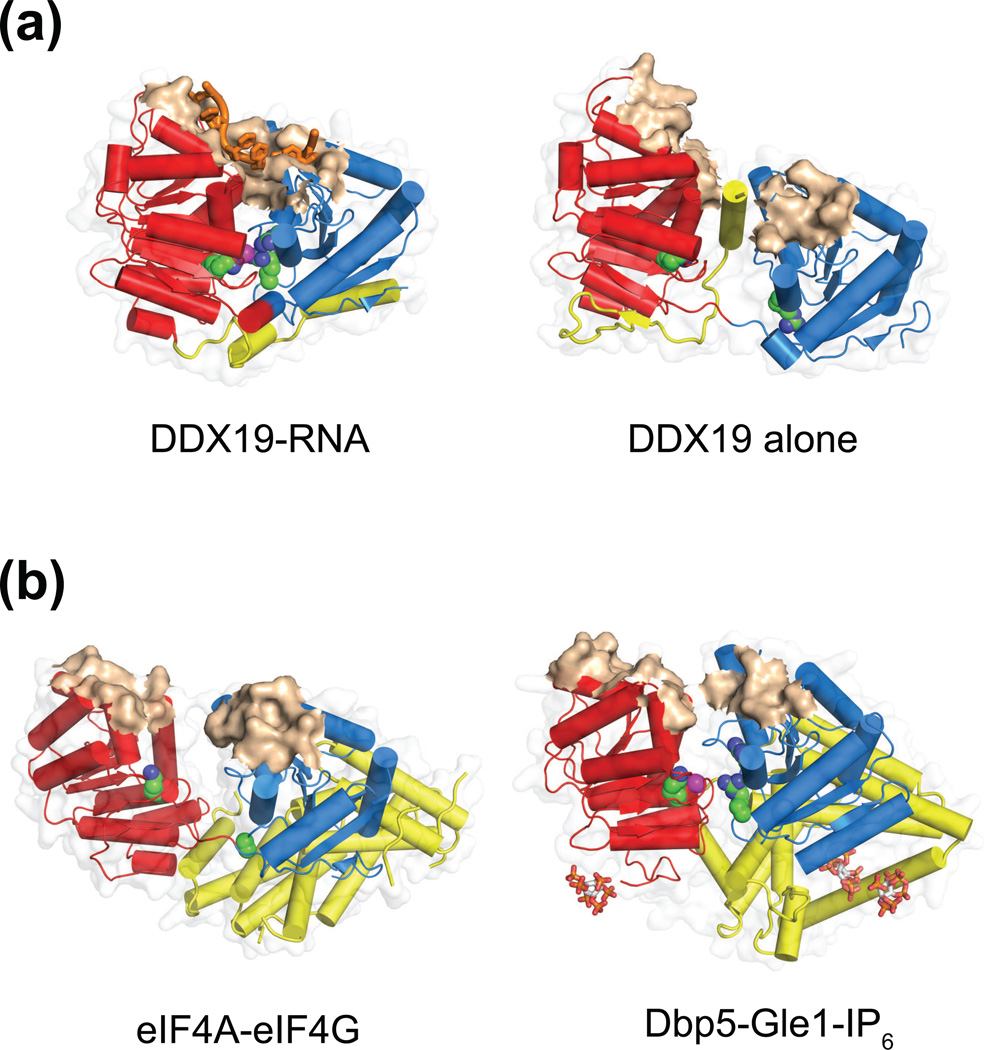
Figure 2. Auxiliary elements appear to influence domain orientations of the ATPase motor.
(a) The DDX19 ATPase is shown in an RNA-bound state (left, 3G0H [18]), with the two ATPase lobes tightly packed together and poised for hydrolysis, and an opened state (right, 3EWS [18]) where an N-terminal segment (yellow) is wedged in the central ATPase cleft. In an RNA-bound state, catalytically important residues such as the P-loop lysine and motif VI arginines (green) cluster tightly around nucleotide phosphates (magenta), whereas in the absence of RNA, communication between ATPase domains is disrupted and the nucleic acid binding surface forms two noncontiguous patches.
(b) Two DEAD-box helicases bound by activators (yellow) on surfaces that are distal to the nucleic acid binding surface (brown): eIF4A-eIF4G (left, 2VSO [19]) and Dbp5-Gle1-IP6 (right, 3RRM [13]). The activators stabilize the ATPase motors in an opened conformation, which appear to disrupt the RNA-binding surfaces and central ATP-binding pocket. These interactions promote the rate-limiting step of RNA release and thus speed up the catalytic cycle. IP6 molecules, which mediate the interaction between Dbp5 and Gle1, are shown as sticks.
Structural studies of DEAD-box helicases have revealed that in addition to inhibition, elements that influence ATPase domain organization can also be stimulatory. ATPase activity of eIF4A is stimulated by the HEAT-repeat protein eIF4G, which contacts both ATPase lobes [19•] (Figure 2b). When bound to eIF4G, the two lobes of eIF4A are positioned with the central helicase motifs roughly facing each other, yet the motor is in an opened conformation with the lobes too distantly separated to allow for ATP hydrolysis. This interaction stimulates eIF4A indirectly by increasing the dissociation rate of RNA, a rate limiting step for enzyme turnover [13••]. Another example where influencing helicase domain arrangement stimulates ATPase activity has been demonstrated for the Dbp5 helicase. In a manner similar to eIF4A, Dbp5 is activated by Gle1, a HEAT-repeat protein that stabilizes an opened organization of the ATPase domains and promotes RNA release analogously to eIF4G [13••] (Figure 2b). As an activator, Gle1 displaces the inhibitory N-terminal helix of Dbp5 that blocks domain closure, but interestingly, can bind concurrently with and override the inhibitory effects of Nup159 [13••]. Thus, as exemplified by Dbp5 and eIF4A, mechanisms for inhibition and stimulation of ATPase activity can cooperate and coordinate to tune action of SF2 ATPase motors for particular circumstances and substrates.
Both of these strategies for regulating ATPase activity – blocking DNA-binding surfaces and influencing the ATPase domain organization – fit into the concept of modular allostery, where formation of an activated structure relies upon displacement of an auxiliary element [20]. Modular allostery may be utilized for ensuring substrate specificity for SWI2/SNF2 proteins, many of which require specific protein-DNA substrates for full activation of the ATPase motor, and, like Chd1, may utilize auxiliary domains to reduce ATPase activation by improper substrates. For Rad54, for example, removal of the N-terminus permitted naked DNA to stimulate ATPase activity to the same level as the preferred Rad51+DNA substrate [21], and likewise deletion of the CSB (Cockayne Syndrome B) N-terminus increased DNA-stimulated ATPase activity several fold [22]. For Mot1, an N-terminal portion was shown to increase specificity for its substrate (in this case TATA-binding protein, TBP), and inhibition of DNA-stimulated hydrolysis was proposed to occur through electrostatic interactions of this region with the ATPase motor [23]. Though regulatory segments have not been identified for ALC1 and ISWI-type remodelers, these proteins, like Chd1, are preferentially stimulated by nucleosome substrates [24–26]. The ALC1 remodeler possesses a C-terminal macro domain that can bind to poly(ADP)-ribose (PAR), and auto-PARylation of PARP1 additionally stimulates ATPase and remodeling activities [25, 26]. While the detailed mechanisms employed by these SWI2/SNF2 proteins await further structural studies, the range of regulatory strategies observed for DEAD-box and Chd1 ATPase motors suggests that SWI2/SNF2 proteins will likely display a rich variety of mechanisms utilizing modular allostery to control ATPase activation.
Helical subdomains of SWI2/SNF2 ATPases appear positioned to distort duplex DNA structure
The ATPase motors of chromatin remodelers are distinguished by the presence of helical subdomains (1B, 2B) inserted in each of the RecA-like lobes [2, 3, 6••, 7••] (Figure 3a). When the two lobes of the ATPase motor are modeled in a hydrolysis competent closed state, these helical subdomains cluster together at one edge of the predicted DNA-binding site. As previously suggested, the locations of these subdomains may imply a direct interaction with DNA [2, 3, 6••, 7••]. Interestingly, an alpha-helical, DNA-interacting domain is inserted in the same location in the Hef helicase, an SF2 ATPase not in the SWI2/SNF2 family that can process forked DNA structures [3, 27]. Deletion of the inserted helical domain in the Hef helicase abrogates stimulation by forked DNA substrates, indicating that this domain is required for recognition and/or strand separation. Strand separation by processive 3’-5’ helicases is typically aided by a “separation pin” that destabilizes base pairing just upstream of the site of translocation [4]. Although SWI/SNF2 ATPases do not separate duplex DNA as they translocate [28, 29], these separation pins occupy the same location relative to the ATPase motor as the helical insertions of Hef and the SWI2/SNF2-specific 2B subdomain (Figure 3b).
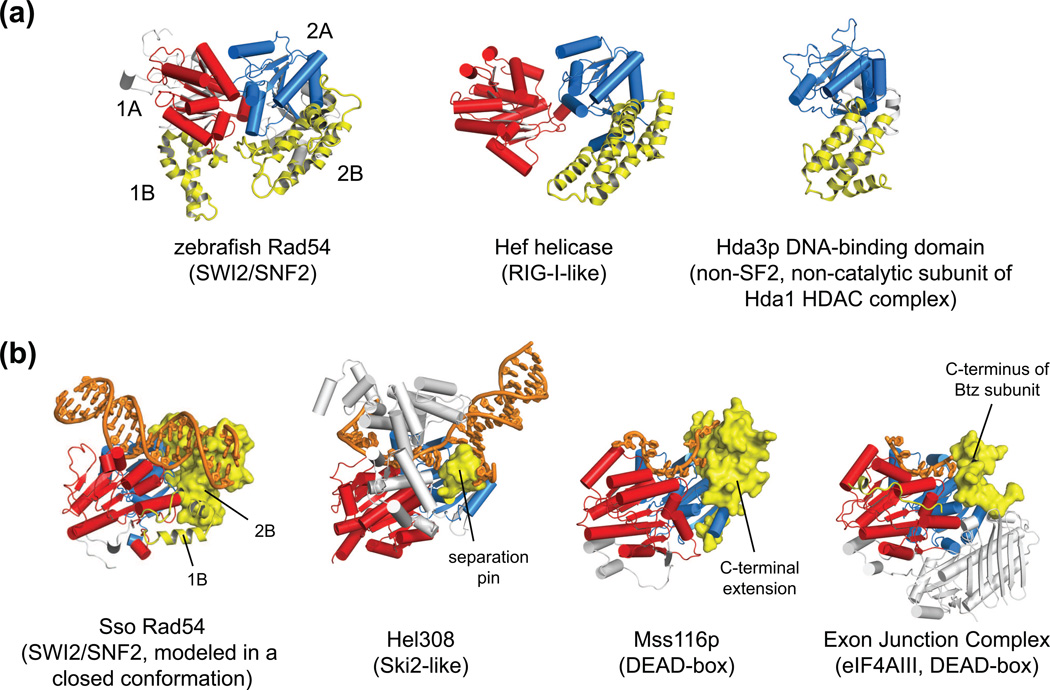
Figure 3. The SWI2/SNF2-specific subdomains are positioned on the ATPase motor similarly to nucleic acid interacting elements in other SF2 proteins.
(a) When the SWI2/SNF2 ATPase motor is organized close to a tightly packed, catalytically competent state (shown here for zebrafish Rad54, 1Z3I [6]), the helical inserts characteristic of SWI2/SNF2 proteins (subdomains 1B, 2B, highlighted in yellow) cluster together on one side of the ATPase motor. Inserted helical domains with similar topologies have also been observed at the same position within domain 2A for the non-SWI2/SNF2 proteins Hef helicase (1WP9 [3, 27]) and Hda3 (3HGT [43]). Hda3 only possesses the second ATPase lobe and does not by sequence appear to be a catalytically active ATPase.
(b) SWI2/SNF2-specific subdomains appear positioned to distort a DNA duplex. Modeling the S.solfataricus Rad54 ATPase (1Z63 [7]) in a closed form (based on DEAD-box helicases with active-site geometry) reveals a clash between the SWI2/SNF2-specific inserts on the second ATPase lobe (subdomain 2B) and duplex DNA bound to domain 1A. Elements occupying a position similar to the SWI2/SNF2 subdomain 2B (yellow) alter the paths of nucleic acid strands bound to several SF2 proteins, including Hel308 (2P6R [49]), Mss116p (3I5X [31]), and eIF4AIII of the Exon Junction Complex (2J0S [32]). The SF2-type family for each ATPase is given in parentheses [4].
Additionally, recent crystal structures of the non-translocating DEAD box RNA helicases Mss116p and eIF4AIII have revealed elements located in positions similar to the SWI2/SNF2-specific inserts that can locally distort nucleic acid structure upon binding. Members of the DEAD-box family of RNA helicases have been proposed to remodel RNA substrates by bending the nucleic acid backbone [30]. Mss116p, which is believed to act as a general RNA chaperone important for splicing, translation, and RNA processing, possesses a C-terminal helical subdomain that alters the trajectory of the bound nucleic acid strand [31••] (Figure 3b). Although inserted at a distinct place in the protein primary sequence, this helical subdomain occupies a space strikingly similar to that of the SWI2/SNF2-specific inserts. Adjacent to the canonical nucleic acid binding surface spanning the two ATPase lobes, this helical subdomain presents a steric barrier that forces a dramatic bend in the nucleic acid strand. Another example where the nucleic acid strand appears redirected has been observed for the core exon junction complex, formed by a four-protein assembly containing the DEAD-box helicase eIF4AIII [32•, 33•, 34]. In this complex, one of the core components, Barentsz (Btz) (also called MLN51, for Metastatic Lymph Node 51), packs against eIF4AIII in the same location as the SWI2/SNF2-specific subdomains and base stacks with one RNA nucleotide, blocking a helical path (Figure 3b). Thus, it seems that mechanisms for disrupting the path of a bound nucleic acid have evolved independently in several systems by targeting a similar location on the helicase motor.
Although functionally diverse, a common thread for SWI2/SNF2 proteins appears to be the ability to disrupt or remodel particular protein-DNA complexes. The SWI2/SNF2-specific subdomains, which were shown to be functionally important for ATPase activity and mutated in various human diseases, have been proposed to aid in translocation and/or processivity of the ATPase on DNA [6••, 7••]. Recently, the RSC chromatin remodeler was shown to translocate along DNA at high (>20 pN) forces when tethered to DNA, supporting the notion that translocation alone may be capable of disrupting protein-DNA contacts such as those found within the nucleosome [35•]. Indeed, translocation by the powerful and highly processive bacterial RecBCD helicase (a complex of SF1 ATPases) has been shown to be sufficient for sliding and displacing several types of DNA-bound complexes, including nucleosomes [36]. In contrast to mechanisms where DNA translocation has been proposed to be sufficient [37], other models describing action of SWI2/SNF2 ATPases such as chromatin remodelers have suggested that a localized distortion of duplex DNA is also critical for reorganization of protein-DNA complexes [37–42]. Coupled to translocation, a localized distortion of the DNA helix promoted by the SWI2/SNF2-specific subdomains may be required for breaking protein-DNA contacts during the remodeling reaction. Intriguingly, the Hda3 subunit of a nucleosome-associated complex was recently shown to possess just the second lobe of an SF2 ATPase motor along with the SWI2/SNF2-specific subdomain 2B [43]. The maintenance of this subdomain in conjunction with a non-functional helicase domain suggests that the SWI2/SNF2-specific insertions may play a role independent of or complementary to DNA translocation, such as recognizing characteristic distortions in DNA.
Concluding Remarks
Despite only a few structural examples of SWI2/SNF2 ATPases, a wealth of structural data for related SF2 ATPases, particularly those from the DEAD-box family of RNA helicases, has enabled predictions for how auxiliary domains such as the Chd1 chromodomains can influence activation of the ATPase motor. While comparisons between families such as SWI2/SNF2 and DEAD-box helicases will continue to be informative, significant progress in understanding the molecular mechanisms utilized by SWI2/SNF2 proteins will rely heavily on new structures. New structures, captured in both activated and inhibited states, will reveal additional regulatory mechanisms for directing action of the ATPase motor, likely reinforcing basic principles that are now emerging as well as potentially revealing differences specific to the SWI2/SNF2 family. Additional structures of SWI2/SNF2 proteins, in particular bound to DNA or DNA-protein substrates, will also likely be instrumental in understanding how these proteins disrupt target complexes. Discovering how the SWI2/SNF2-specific subdomains contact and/or influence the structure of duplex DNA should additionally provide important mechanistic insight towards understanding how this family of enzymes works.
HIGHLIGHTS.
SF2 ATPases can be regulated by blocking nucleic acid binding surfaces.
Domain organization of SF2 ATPase motors can be influenced by regulatory elements.
Swi2/Snf2-specific subdomains are positioned to disrupt nucleic acid structure.
Acknowledgements
We thank Ilana Nodelman and other members of the Bowman lab for critical comments and discussions. This work was supported by a grant from the National Institutes of Health (R01 GM084129).
Footnotes
Publisher's Disclaimer: This is a PDF file of an unedited manuscript that has been accepted for publication. As a service to our customers we are providing this early version of the manuscript. The manuscript will undergo copyediting, typesetting, and review of the resulting proof before it is published in its final citable form. Please note that during the production process errors may be discovered which could affect the content, and all legal disclaimers that apply to the journal pertain.
References
- 1.Singleton MR, Dillingham MS, Wigley DB. Structure and mechanism of helicases and nucleic acid translocases. Annu Rev Biochem. 2007;76:23–50. doi: 10.1146/annurev.biochem.76.052305.115300. [DOI] [PubMed] [Google Scholar]
- 2.Durr H, Flaus A, Owen-Hughes T, Hopfner KP. Snf2 family ATPases and DExx box helicases: differences and unifying concepts from high-resolution crystal structures. Nucleic Acids Res. 2006;34:4160–4167. doi: 10.1093/nar/gkl540. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Flaus A, Martin DM, Barton GJ, Owen-Hughes T. Identification of multiple distinct Snf2 subfamilies with conserved structural motifs. Nucleic Acids Res. 2006;34:2887–2905. doi: 10.1093/nar/gkl295. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Fairman-Williams ME, Guenther UP, Jankowsky E. SF1 and SF2 helicases: family matters. Curr Opin Struct Biol. 2010;20:313–324. doi: 10.1016/j.sbi.2010.03.011. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Jankowsky E. RNA helicases at work: binding and rearranging. Trends Biochem Sci. 2011;36:19–29. doi: 10.1016/j.tibs.2010.07.008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6. Thoma NH, Czyzewski BK, Alexeev AA, Mazin AV, Kowalczykowski SC, Pavletich NP. Structure of the SWI2/SNF2 chromatin-remodeling domain of eukaryotic Rad54. Nat Struct Mol Biol. 2005;12:350–356. doi: 10.1038/nsmb919. This paper, along with [••7], presents the first crystal structure of a SWI2/SNF2 family member Rad54, which revealed the structure and organization of the conserved SWI2/SNF2-specific subdomains. The authors highlight positions of disease-associated mutations from other SNF2 family members, several of which occur within the SWI2/SNF2-specific inserts.
- 7. Durr H, Korner C, Muller M, Hickmann V, Hopfner KP. X-ray structures of the Sulfolobus solfataricus SWI2/SNF2 ATPase core and its complex with DNA. Cell. 2005;121:363–373. doi: 10.1016/j.cell.2005.03.026. Along with [6••], this work presents the first SWI2/SNF2 crystal structure, and provided the first glimpse of how the ATPase motor interacts with double-stranded DNA. In the structure, domain 1A contacts one DNA strand analogously to other SF2 ATPases, reinforcing the idea that all SF2 proteins bind to nucleic acid strands across the central cleft in a similar fashion. The 2A domain in this structure shows an unconventional arrangement, flipped approximately 180° from what is expected to bring the conserved helicase motifs together.
- 8. Shaw G, Gan J, Zhou YN, Zhi H, Subburaman P, Zhang R, Joachimiak A, Jin DJ, Ji X. Structure of RapA, a Swi2/Snf2 protein that recycles RNA polymerase during transcription. Structure. 2008;16:1417–1427. doi: 10.1016/j.str.2008.06.012. This work provides the third such example of a snf2 family member and the only structure of a full-length snf2 family member. The crystal structure demonstrates the way in which auxiliary domains of a full length SWI2/SNF2 family member make contacts with the conserved ATPase core.
- 9. Hauk G, McKnight JN, Nodelman IM, Bowman GD. The chromodomains of the Chd1 chromatin remodeler regulate DNA access to the ATPase motor. Mol Cell. 2010;39:711–723. doi: 10.1016/j.molcel.2010.08.012. This paper presents the first crystal structure of a chromatin remodeler SWI2/SNF2, which shows that for the Chd1 subfamily, an N-terminal pair of chromodomains can block DNA from accessing and activating the ATPase motor domain. By reducing ATPase stimulation in the absence of nucleosomes, the chromodomains appear to provide a mechanism for discriminating among different substrates.
- 10.Tran EJ, Zhou Y, Corbett AH, Wente SR. The DEAD-box protein Dbp5 controls mRNA export by triggering specific RNA:protein remodeling events. Mol Cell. 2007;28:850–859. doi: 10.1016/j.molcel.2007.09.019. [DOI] [PubMed] [Google Scholar]
- 11. von Moeller H, Basquin C, Conti E. The mRNA export protein DBP5 binds RNA and the cytoplasmic nucleoporin NUP214 in a mutually exclusive manner. Nat Struct Mol Biol. 2009;16:247–254. doi: 10.1038/nsmb.1561. The work presented in [•11] and [•12] provide the molecular basis by which Nup214 (yeast Nup159) inhibits the SF2 helicase DDX19 (yeast Dbp5). Crystal structures of the closed form of DDX19 bound to AMPPNP and RNA along with the N-terminal domain 1A bound to Nup214 demonstrate that Nup214 binds to a RNA binding surface of DDX19. These structures provide a molecular explanation for the ability of Nup214 to reduce RNA-stimulated ATPase and RNA binding of DDX19.
- 12. Napetschnig J, Kassube SA, Debler EW, Wong RW, Blobel G, Hoelz A. Structural and functional analysis of the interaction between the nucleoporin Nup214 and the DEAD-box helicase Ddx19. Proc Natl Acad Sci U S A. 2009;106:3089–3094. doi: 10.1073/pnas.0813267106. See [•11] above.
- 13. Montpetit B, Thomsen ND, Helmke KJ, Seeliger MA, Berger JM, Weis K. A conserved mechanism of DEAD-box ATPase activation by nucleoporins and InsP6 in mRNA export. Nature. 2011;472:238–242. doi: 10.1038/nature09862. The authors present multiple structures of yeast Dbp5 (DDX19 in humans) bound to activators and inhibitors, neatly explaining how the Gle1 activator and IP6 overcome inhibition by Nup159. Gle1 binds to a surface of Dbp5 away from the nucleic acid binding surface, mediated by IP6, stabilizing an open form of Dbp5 that promotes RNA release. The authors also show that this mechanism of accelerating RNA release also applies to the eIF4A/eIF4G complex, which shares a high degree of structural similarity.
- 14.Gingras AC, Raught B, Sonenberg N. eIF4 initiation factors: effectors of mRNA recruitment to ribosomes and regulators of translation. Annu Rev Biochem. 1999;68:913–963. doi: 10.1146/annurev.biochem.68.1.913. [DOI] [PubMed] [Google Scholar]
- 15.Yang HS, Jansen AP, Komar AA, Zheng X, Merrick WC, Costes S, Lockett SJ, Sonenberg N, Colburn NH. The transformation suppressor Pdcd4 is a novel eukaryotic translation initiation factor 4A binding protein that inhibits translation. Mol Cell Biol. 2003;23:26–37. doi: 10.1128/MCB.23.1.26-37.2003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16. Chang JH, Cho YH, Sohn SY, Choi JM, Kim A, Kim YC, Jang SK, Cho Y. Crystal structure of the eIF4A-PDCD4 complex. Proc Natl Acad Sci U S A. 2009;106:3148–3153. doi: 10.1073/pnas.0808275106. This work, along with [•17], provides the physical basis by which PDCD4 to represses eIF4A. The structures reveal that PDCD4 binds to nucleic acid binding surfaces of eIF4A and prevents proper closure of domain 1A and 2A.
- 17. Loh PG, Yang HS, Walsh MA, Wang Q, Wang X, Cheng Z, Liu D, Song H. Structural basis for translational inhibition by the tumour suppressor Pdcd4. EMBO J. 2009;28:274–285. doi: 10.1038/emboj.2008.278. See [•16].
- 18. Collins R, Karlberg T, Lehtio L, Schutz P, van den Berg S, Dahlgren LG, Hammarstrom M, Weigelt J, Schuler H. The DEXD/H-box RNA helicase DDX19 is regulated by an {alpha}-helical switch. J Biol Chem. 2009;284:10296–10300. doi: 10.1074/jbc.C900018200. This work presents an interesting auto-inhibited form of DDX19, where the N-terminal helix of DDX19 is inserted between lobes 1A and 2A and thus prevents proper closure of the domains. The authors show that removal of the inhibitory N-terminal segment results in the loss of RNA-regulated ATPase stimulation.
- 19. Schutz P, Bumann M, Oberholzer AE, Bieniossek C, Trachsel H, Altmann M, Baumann U. Crystal structure of the yeast eIF4A-eIF4G complex: an RNA-helicase controlled by protein-protein interactions. Proc Natl Acad Sci U S A. 2008;105:9564–9569. doi: 10.1073/pnas.0800418105. This work presents the first crystal structure of eIF4A bound to its activator, eIF4G. Unlike previous models for activation, the structure shows eIF4A adopts a conformation in which its helicase lobes are slightly opened when bound to eIF4G.
- 20.Dueber JE, Yeh BJ, Bhattacharyya RP, Lim WA. Rewiring cell signaling: the logic and plasticity of eukaryotic protein circuitry. Curr Opin Struct Biol. 2004;14:690–699. doi: 10.1016/j.sbi.2004.10.004. [DOI] [PubMed] [Google Scholar]
- 21.Alexiadis V, Lusser A, Kadonaga JT. A conserved N-terminal motif in Rad54 is important for chromatin remodeling and homologous strand pairing. J Biol Chem. 2004;279:27824–27829. doi: 10.1074/jbc.M402648200. [DOI] [PubMed] [Google Scholar]
- 22.Lake RJ, Geyko A, Hemashettar G, Zhao Y, Fan HY. UV-induced association of the CSB remodeling protein with chromatin requires ATP-dependent relief of N-terminal autorepression. Mol Cell. 2010;37:235–246. doi: 10.1016/j.molcel.2009.10.027. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Adamkewicz JI, Mueller CG, Hansen KE, Prud'homme WA, Thorner J. Purification and enzymic properties of Mot1 ATPase, a regulator of basal transcription in the yeast Saccharomyces cerevisiae. J Biol Chem. 2000;275:21158–21168. doi: 10.1074/jbc.M002639200. [DOI] [PubMed] [Google Scholar]
- 24.Tsukiyama T, Palmer J, Landel CC, Shiloach J, Wu C. Characterization of the imitation switch subfamily of ATP-dependent chromatin-remodeling factors in Saccharomyces cerevisiae. Genes Dev. 1999;13:686–697. doi: 10.1101/gad.13.6.686. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Ahel D, Horejsi Z, Wiechens N, Polo SE, Garcia-Wilson E, Ahel I, Flynn H, Skehel M, West SC, Jackson SP, Owen-Hughes T, Boulton SJ. Poly(ADP-ribose)-dependent regulation of DNA repair by the chromatin remodeling enzyme ALC1. Science. 2009;325:1240–1243. doi: 10.1126/science.1177321. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Gottschalk AJ, Timinszky G, Kong SE, Jin J, Cai Y, Swanson SK, Washburn MP, Florens L, Ladurner AG, Conaway JW, Conaway RC. Poly(ADP-ribosyl)ation directs recruitment and activation of an ATP-dependent chromatin remodeler. Proc Natl Acad Sci U S A. 2009;106:13770–13774. doi: 10.1073/pnas.0906920106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Nishino T, Komori K, Tsuchiya D, Ishino Y, Morikawa K. Crystal structure and functional implications of Pyrococcus furiosus hef helicase domain involved in branched DNA processing. Structure. 2005;13:143–153. doi: 10.1016/j.str.2004.11.008. [DOI] [PubMed] [Google Scholar]
- 28.Cote J, Quinn J, Workman JL, Peterson CL. Stimulation of GAL4 derivative binding to nucleosomal DNA by the yeast SWI/SNF complex. Science. 1994;265:53–60. doi: 10.1126/science.8016655. [DOI] [PubMed] [Google Scholar]
- 29.Petukhova G, Stratton S, Sung P. Catalysis of homologous DNA pairing by yeast Rad51 and Rad54 proteins. Nature. 1998;393:91–94. doi: 10.1038/30037. [DOI] [PubMed] [Google Scholar]
- 30.Sengoku T, Nureki O, Nakamura A, Kobayashi S, Yokoyama S. Structural basis for RNA unwinding by the DEAD-box protein Drosophila Vasa. Cell. 2006;125:287–300. doi: 10.1016/j.cell.2006.01.054. [DOI] [PubMed] [Google Scholar]
- 31. Del Campo M, Lambowitz AM. Structure of the Yeast DEAD box protein Mss116p reveals two wedges that crimp RNA. Mol Cell. 2009;35:598–609. doi: 10.1016/j.molcel.2009.07.032. The authors present several structures of Mss116p bound to RNA. Unexpectedly, a C-terminal extension of Mss116 adopts a helical fold that drastically alters the path of the RNA substrate bound across the two helicase lobes. Although Mss116p is a non-translocating helicase, this C-terminal extension spatially aligns with separation pins found in translocating helicases.
- 32. Bono F, Ebert J, Lorentzen E, Conti E. The crystal structure of the exon junction complex reveals how it maintains a stable grip on mRNA. Cell. 2006;126:713–725. doi: 10.1016/j.cell.2006.08.006. This paper, along with [•33], presents a structure of the DEAD box helicase eIF4AIII in the context of the exon junction complex. A fragment of the Barentsz protein component of the exon junction complex, which is positioned analogously to the SWI2/SNF2-specific 2B subdomain, redirects the path of the bound RNA molecule, inducing a bend.
- 33. Andersen CB, Ballut L, Johansen JS, Chamieh H, Nielsen KH, Oliveira CL, Pedersen JS, Seraphin B, Le Hir H, Andersen GR. Structure of the exon junction core complex with a trapped DEAD-box ATPase bound to RNA. Science. 2006;313:1968–1972. doi: 10.1126/science.1131981. See [•32] above.
- 34.Le Hir H, Andersen GR. Structural insights into the exon junction complex. Curr Opin Struct Biol. 2008;18:112–119. doi: 10.1016/j.sbi.2007.11.002. [DOI] [PubMed] [Google Scholar]
- 35. Sirinakis G, Clapier CR, Gao Y, Viswanathan R, Cairns BR, Zhang Y. The RSC chromatin remodelling ATPase translocates DNA with high force and small step size. EMBO J. 2011;30:2364–2372. doi: 10.1038/emboj.2011.141. In this work, optical tweezer experiments demonstrate that the motor domain of the RSC chromatin remodeler, when tethered with an exogenous DNA binding domain, is capable of translocation, even under high force. The authors suggests that translocation by the RSC motor (coupled to Arp7 and Arp9 co-activators) is capable of displacing nucleosomes by translocation.
- 36.Finkelstein IJ, Visnapuu ML, Greene EC. Single-molecule imaging reveals mechanisms of protein disruption by a DNA translocase. Nature. 2010;468:983–987. doi: 10.1038/nature09561. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Lorch Y, Maier-Davis B, Kornberg RD. Mechanism of chromatin remodeling. Proc Natl Acad Sci U S A. 2010;107:3458–3462. doi: 10.1073/pnas.1000398107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.van Holde K, Yager T. Models for chromatin remodeling: a critical comparison. Biochem Cell Biol. 2003;81:169–172. doi: 10.1139/o03-038. [DOI] [PubMed] [Google Scholar]
- 39.Langst G, Becker PB. Nucleosome remodeling: one mechanism, many phenomena? Biochim Biophys Acta. 2004;1677:58–63. doi: 10.1016/j.bbaexp.2003.10.011. [DOI] [PubMed] [Google Scholar]
- 40.Saha A, Wittmeyer J, Cairns BR. Chromatin remodelling: the industrial revolution of DNA around histones. Nat Rev Mol Cell Biol. 2006;7:437–447. doi: 10.1038/nrm1945. [DOI] [PubMed] [Google Scholar]
- 41.Zofall M, Persinger J, Kassabov SR, Bartholomew B. Chromatin remodeling by ISW2 and SWI/SNF requires DNA translocation inside the nucleosome. Nat Struct Mol Biol. 2006;13:339–346. doi: 10.1038/nsmb1071. [DOI] [PubMed] [Google Scholar]
- 42.Bowman GD. Mechanisms of ATP-dependent nucleosome sliding. Curr Opin Struct Biol. 2010;20:73–81. doi: 10.1016/j.sbi.2009.12.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Lee JH, Maskos K, Huber R. Structural and functional studies of the yeast class II Hda1 histone deacetylase complex. J Mol Biol. 2009;391:744–757. doi: 10.1016/j.jmb.2009.06.059. [DOI] [PubMed] [Google Scholar]
- 44.Gorbalenya AE, Koonin EV. Helicases: amino acid sequence comparisons and structure-function relationships. Curr Opin Struct Biol. 1993;3:419–429. [Google Scholar]
- 45.Jankowsky A, Guenther UP, Jankowsky E. The RNA helicase database. Nucleic Acids Res. 2011;39:D338–D341. doi: 10.1093/nar/gkq1002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Lusser A, Urwin DL, Kadonaga JT. Distinct activities of CHD1 and ACF in ATP-dependent chromatin assembly. Nat Struct Mol Biol. 2005;12:160–166. doi: 10.1038/nsmb884. [DOI] [PubMed] [Google Scholar]
- 47.Gangaraju VK, Bartholomew B. Mechanisms of ATP dependent chromatin remodeling. Mutat Res. 2007;618:3–17. doi: 10.1016/j.mrfmmm.2006.08.015. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Kim JL, Morgenstern KA, Griffith JP, Dwyer MD, Thomson JA, Murcko MA, Lin C, Caron PR. Hepatitis C virus NS3 RNA helicase domain with a bound oligonucleotide: the crystal structure provides insights into the mode of unwinding. Structure. 1998;6:89–100. doi: 10.1016/s0969-2126(98)00010-0. [DOI] [PubMed] [Google Scholar]
- 49.Buttner K, Nehring S, Hopfner KP. Structural basis for DNA duplex separation by a superfamily-2 helicase. Nat Struct Mol Biol. 2007;14:647–652. doi: 10.1038/nsmb1246. [DOI] [PubMed] [Google Scholar]