Background: Incorrect end use during repair can cause chromosome rearrangements.
Results: DNA-PKcs and RAD50 limit incorrect end use, and a break downstream from an active promoter shows elevated incorrect end use; these factors and conditions have distinct effects on repair requiring end processing.
Conclusion: DNA-PKcs, RAD50, and transcription context influence correct end use.
Significance: Correct end use and end processing appear distinct processes.
Keywords: Chromosomes, DNA Repair, Genomic Instability, Homologous Recombination, Transcription, ATM, DNA-PKcs, RAD50, Double Strand Break, End Joining
Abstract
During repair of multiple chromosomal double strand breaks (DSBs), matching the correct DSB ends is essential to limit rearrangements. To investigate the maintenance of correct end use, we examined repair of two tandem noncohesive DSBs generated by endonuclease I-SceI and the 3′ nonprocessive exonuclease Trex2, which can be expressed as an I-SceI-Trex2 fusion. We examined end joining (EJ) repair that maintains correct ends (proximal-EJ) versus using incorrect ends (distal-EJ), which provides a relative measure of incorrect end use (distal end use). Previous studies showed that ATM is important to limit distal end use. Here we show that DNA-PKcs kinase activity and RAD50 are also important to limit distal end use, but that H2AX is dispensable. In contrast, we find that ATM, DNA-PKcs, and RAD50 have distinct effects on repair events requiring end processing. Furthermore, we developed reporters to examine the effects of the transcription context on DSB repair, using an inducible promoter. We find that a DSB downstream from an active promoter shows a higher frequency of distal end use, and a greater reliance on ATM for limiting incorrect end use. Conversely, DSB transcription context does not affect end processing during EJ, the frequency of homology-directed repair, or the role of RAD50 and DNA-PKcs in limiting distal end use. We suggest that RAD50, DNA-PKcs kinase activity, and transcription context are each important to limit incorrect end use during EJ repair of multiple DSBs, but that these factors and conditions have distinct roles during repair events requiring end processing.
Introduction
Recent sequencing of cancer genomes has revealed an abundance of chromosomal rearrangements in a variety of tumor types (1). Such rearrangements include abnormalities that could be observed cytogenetically, but notably the majority of intrachromosomal rearrangements involve breakpoints that are within 2 Mb (2). In addition, chromosomal rearrangements are induced by clastogenic cancer therapeutics (e.g. ionizing radiation) (3). One mechanism of such rearrangements involves end joining (EJ)3 repair of multiple simultaneous chromosomal double strand breaks (DSBs) that fails to maintain correct (proximal) end use. In addition, some cancer-associated chromosome rearrangements may involve DSB end processing to reveal short stretches of homology (microhomology) that are often observed at rearrangement junctions (2). Examining the factors and conditions that influence the maintenance of correct end use and/or DSB end processing will provide insight into the etiology of chromosomal rearrangements.
Recent evidence indicates that DNA damage response factors are important for maintaining correct end use and regulating DSB end processing during EJ. For example, during programmed rearrangements in developing lymphocytes, the ataxia telangiectasia-mutated kinase (ATM), and members of the MRE11-RAD50-NBS1 complex, are important to limit aberrant rearrangements (coding-signal hybrid junctions) of certain V(D)J recombination substrates (4–6). As well, during EJ repair of two tandem chromosomal DSBs, ATM and NBS1, are important to limit incorrect (distal) end use (7). However, ATM and NBS1 appear to show opposite effects on the degree of end processing during EJ (7, 8), which raises the notion that maintenance of correct end use and regulation of DSB end processing may be distinct aspects of repair.
In addition to ATM and NBS1, the DNA damage response factors RAD50 and DNA-PKcs are also important for chromosomal stability (9–14). RAD50 is part of the MRE11-RAD50-NBS1 complex that is recruited early to chromosomal DSBs (15); whereas DNA-PKcs is important for classical nonhomologous end joining (c-NHEJ) and V(D)J recombination (13, 14). Both factors are important for radioresistance, and hence are being examined as potential targets for causing radiosensitization during cancer therapy (16, 17). Regarding specific functions of these factors, DNA-PKcs and the MRE11-RAD50-NBS1 complex can promote DNA tethering in vitro (15, 18–22). Furthermore, the MRE11-RAD50-NBS1 complex promotes end resection, which is important for EJ events involving microhomology (alt-EJ), as well as homologous recombination (15, 23). In contrast, DNA-PKcs is important to limit homologous recombination (24, 25), yet also is important to recruit the Artemis nuclease for processing of ends during EJ (13, 26). Further characterizing the roles of DNA-PKcs and RAD50 during DSB repair will provide insight into the maintenance of genome stability and cellular radiation response.
Apart from individual DNA damage response factors, the chromosomal context of DNA damage may also influence genome stability. Namely, chromosomal regions with elevated levels of transcription have been shown to be prone to rearrangements (27–30). In contrast, previous studies have shown that the chromosomal break transcription context does not obviously affect the frequency of homology-directed repair (HDR), or the degree of end processing during EJ (31, 32).
In this study, we have examined the role of individual DNA damage response factors and chromosomal break transcription context on the frequency of incorrect end use during EJ of two tandem DSBs, as well as the frequency of repair events requiring end processing. We present evidence that inhibition of DNA-PKcs kinase activity, or depletion of RAD50, each cause an elevated frequency of incorrect (distal) end use during repair of tandem DSBs. In contrast, we find that these factors have opposing effects on a homologous repair event that involves extensive DSB end processing (single strand annealing, SSA), in that DNA-PKcs inhibits SSA, whereas RAD50 promotes this repair event. Furthermore, we find that the histone variant, H2AX, which is important for limiting end processing during EJ as well as SSA (33–35), is dispensable for maintenance of correct end use during EJ. Additionally, we find that EJ repair of a DSB downstream from an active promoter, compared with an uninduced promoter, shows a greater frequency of incorrect end use and a greater reliance on ATM to limit such EJ events. In contrast, DSB transcription context did not obviously affect the frequency of repair events requiring end processing. In summary, we suggest that the roles of factors or conditions in maintaining correct end use during EJ cannot be predicted by their influence on repair that requires DSB end processing, such that these may be distinct aspects of repair.
EXPERIMENTAL PROCEDURES
Plasmids and Cell Lines
The I-SceI-Trex2 fusion expression vectors (pCAGGS-I-SceI-Trex2 WT or H188A) were generated by mutating the stop codon of the I-SceI expression vector (pCBASce) into a BglII site (36), and subsequently inserting an amplification product from pCAGGS-Trex2 (WT or H188A) (37) downstream into BglII/XhoI sites. In this plasmid, the last three codons of I-SceI (uppercase) and the first three codons of Trex2 (uppercase) are separated by a linker sequence (lowercase): 5′-TTCCTGAAAagatcttgttcctgtgagaacatcagcATGTCTCAG. The reporter g4DRGFP was generated by inserting an amplified SceGFP gene (38) downstream of the g4 promoter in the pNEBR-X1 plasmid (New England Biolabs). The iGFP gene (38) was inserted downstream, and the two GFP genes were separated by 1.4 kb using a fragment of mouse Rb locus genomic DNA (39). Also, the lox-HYG-lox expression cassette (39) was inserted downstream of iGFP. The reporter g4EJ5GFP was generated by inserting a fragment of pim-EJ5GFP (EcoRI/NotI), containing the 5′ I-SceI site through the end of the GFP expression cassette, into pNEBR-X1.
A number of reporters were integrated into U2OS cells to generate a series of cell lines: pimDRGFP linearized with XhoI, hprtSA-GFP linearized with KpnI/SacI, EJ2-GFPpuro linearized with HpaI, and pimEJ5-GFP linearized with XhoI (36). Furthermore, the expression cassettes for the regulator proteins (Gal4-DBD-EcR and RxR-vp16) (40) were integrated into U2OS cells using the pNEBR-R1 plasmid (New England Biolabs) linearized with ScaI, to generate the U2OS-R1 cell line. The g4DRGFP reporter linearized with NotI and the g4EJ5-GFP reporter linearized with NheI were each integrated into U2OS-R1 and HEK293-A7 cells (HEK293 cells with pNEBR-R1, New England Biolabs). Linearized plasmids were introduced into cells by electroporation (0.8 ml, 710–730 V, 10 microfarads), followed by the relevant drug selection to generate individual clones (600 μg/ml of G418, 100–200 μg/ml of hygromycin B, and 1–2 μg/ml of puromycin). Intact reporters were confirmed by Southern blotting using the iGFP fragment as a probe (36). Furthermore, pimEJ5-GFP linearized with XhoI was introduced into WT (H2AXf/f) and H2AX−/− mouse ES cells (41) (kindly provided by Drs. Frederick Alt and Maria Jasin) by electroporation as described above, followed by selection in 120–150 μg/ml of hygromycin B. Targeting of the reporter within individual clones to the pim1 locus was confirmed by PCR analysis, as described previously (36).
Repair Assays
Transfection complexes were formed by mixing 3.6 μl of Lipofectamine 2000 in 200 μl of OptiMEM (Invitrogen) with 0.8 μg of expression vectors for I-SceI (pCBASce) or I-Sce-Trex2 (pCAGGS-I-SceI-Trex2). For co-expression of I-SceI and Trex2 as separate proteins, 0.4 μg of a Trex2 expression vector (pCAGGS-Trex2) was included. The day prior to transfection, 105 cells were plated onto one well of a 12-well dish, and 3 h prior to adding transfection complexes, cells were incubated in 1 ml of medium without antibiotics. Three hours after transfection complexes were added, the medium was removed and replaced with medium containing antibiotics. Except for the TxON/OFF conditions described below, samples were analyzed 3 days after transfection. The frequency of GFP+ cells was determined from a portion of the sample by fluorescence-activated cell sorting (FACS) analysis (CyAN ADP, Dako).
For EJ5-GFP experiments, at the same time as the FACS analysis, a portion of the sample was used for genomic DNA purification and proximal-EJ analysis, as described previously (7). Briefly, the region flanking an I-SceI site was amplified, the product was purified (GFX column, GE), and then digested completely with I-SceI (New England Biolabs), prior to separation by agarose gel electrophoresis. Samples from untransfected cells were analyzed in parallel to ensure complete digestion. Proximal-EJ analysis of EJ5-GFP used primers flanking the 3′ DSB: p1-EJ5PurF, 5′-agcggatcgaaattgatgat, p2-KNDRR, 5′-aagtcgtgctgcttcatgtg. For proximal-EJ analysis using g4EJGFP, the region flanking the 5′ DSB was amplified, because this DSB is adjacent to the g4 promoter: p4-KNDRF5xRE, 5′-gggccgtcagctactaccag, p5-EJ5PurR, 5′-cttttgaagcgtgcagaatg. Subsequently, distal end use was calculated for each sample by dividing the frequency of distal EJ (%GFP) by the frequency of proximal EJ (% I-SceI-resistant amplification product). These individual distal end use values were then normalized to the mean value for parallel control samples to provide a measure of relative distal end use for each experiment.
The above amplification and digestion procedures were also used to examine the I-SceI site restoration from GFP+ sorted distal EJ products following I-SceI expression in cells with the g4EJ5GFP reporter, using primers p4-KNDRF5xRE and p2-KNDRR. To examine the repair junctions of proximal EJ products, I-SceI-resistant amplification products were purified (GFX column, GE Healthcare) and cloned into TA cloning vectors (Invitrogen). Individual clones were sequenced with the M13R primer.
For the drug treatments, ATMi (KU55933, EMD Biosciences 118500) (42), PKi (NU7026, Sigma) (17), or vehicle (dimethyl sulfoxide (DMSO), Sigma) were added at the 3-h media change after adding the transfection complexes, and maintained throughout the experiments. For depletion of RAD50, 105 cells were plated onto wells containing 16 nm siRNA (siRAD50 genone smartpool or siCTRL, Thermo Scientific M-005232-01-0005 and D-001810-01-20, respectively) that had been incubated with 1.8 μl of RNAiMAX (Invitrogen) for 25 min. Subsequently (48 h), cells were transfected with expression vectors for I-SceI or I-SceI-Trex2 as described above, but also including 8 nm of the relevant siRNA in the transfection complexes.
For the reporters with the inducible g4 promoter, transcription was induced by treatment with 500 nm l-(N-(2-ethyl-3-methoxybenzolyl)-N′-3,5-dimethylbenzoyl-tert-butylhydrazine, a synthetic ecdysone receptor ligand, commercially known as RSL1 (New England Biolabs) or Genostat Ligand (Millipore) (40). For the TxON condition, L was added to medium 3 h prior to addition of the transfection complexes, and also to the medium change 3 h after the transfection reagent was removed. In contrast, at these medium changes for the TxOFF condition, vehicle (DMSO) was added instead of L. Subsequently, 3 days after transfection, L was added to both TxON and TxOFF conditions for 3 days prior to analysis of EJ frequencies as described above. Accordingly, all analysis of Tx samples was performed 6 days after transfection.
Immunoblots and Quantitative RT-PCR
To examine RAD50, 48 h after siRNA transfections as described above cells were lysed with NETN (20 mm Tris, pH 8, 100 mm NaCl, 1 mm EDTA, 0.5% IGEPAL, 1 mm DTT, and Roche Protease Inhibitor Mixture) in eight freeze/thaw cycles. Immunoblots of protein extracts were probed with anti-RAD50 antibody (Santa Cruz Biotechnology sc-74460, 1:100) and developed with HRP-conjugated secondary antibody (Santa Cruz Biotechnology sc-2005, 1:750), or probed with HRP-conjugated anti-GAPDH antibody (Abcam ab9482, 1:1500). To examine H2AX, cells were lysed in NETN for two freeze/thaw cycles, and centrifuged at 4 °C for 10 min. Histones were extracted from the pellet using 0.2 n HCl for 40 min at 4 °C. Total histones on the immunoblot were visualized with Ponceau S (Sigma P7170) followed by probing with the anti-H2AX antibody (Millipore 07–627, 1:1000), which was developed with an HRP-conjugated secondary antibody (Santa Cruz Biotechnology, sc-2004). HRP signals were visualized using ECL reagent (Amersham Biosciences).
Relative RNA abundance was examined for HEK293-A7 g4DRGFP cells treated with +L for 3 h or left untreated, by isolating RNA (RNAeasy Plus, Qiagen) for reverse transcription using random primers and MMLV-RT (Promega), which was then amplified with iQ SYBR Green Supermix (Bio-Rad), each according to the manufacturer's protocols. Transcripts downstream of the g4 promoter were amplified with 5′-cgccactatgggatcaaga and 5′-gctgaacttgtggccgttta, and normalized to parallel amplification of actin transcripts with 5′-actgggacgacatggagaag and 5′-aggaaggaaggctggaagag.
Statistical Analysis
Repair values reflect a minimum of three independent transfections. Error bars represent the mean ± S.D. Statistical analysis was performed using the unpaired t test.
RESULTS
Reporter to Examine End Use during EJ of Two Tandem DSBs, Which Involves Expressing an I-SceI-Trex2 Fusion
Chromosomal rearrangements can arise when incorrect ends are matched during repair of multiple DSBs. To examine this key aspect of genome maintenance, we used the EJ5-GFP reporter that contains two tandem I-SceI sites (Fig. 1A) (7, 37). Following I-SceI expression, distal EJ places a promoter adjacent to the rest of the GFP cassette, such that the distal EJ can be quantified as the percentage of GFP+ cells. Proximal EJ is difficult to measure with I-SceI expression alone, because proximal EJ that restores the I-SceI site cannot be differentiated from the uncut reporter. For this, we used co-expression of I-SceI with the 3′-exonuclease Trex2, which has been shown to cause a high level of I-SceI-resistant proximal EJ products, as quantified by PCR amplification across the 3′ I-SceI site, and subsequent I-SceI digestion analysis (7, 37). Notably, co-expression of I-SceI and Trex2 leads to distal EJ products that are completely I-SceI resistant (7, 37). The repair junctions following I-SceI/Trex2 co-expression mostly show very short deletions of the I-SceI 3′ overhang region, which is consistent with the low processivity of Trex2 (7, 37). We also observed a minor class of larger deletions, but importantly none of these deletions disrupt GFP expression (7, 37).
FIGURE 1.
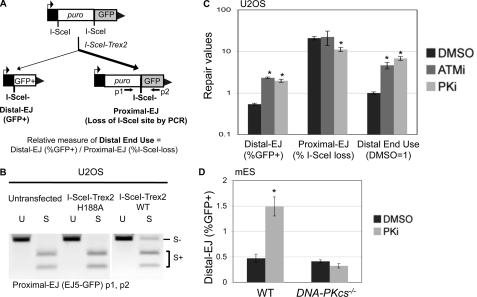
Inhibitor of DNA-PKcs kinase activity causes a marked shift to incorrect end use during repair of tandem DSBs. A, a diagram for the end use assay based on the EJ5-GFP reporter, which contains two tandem recognition sites for the I-SceI endonuclease. Expression of I-SceI and the nonprocessive 3′-exonuclease Trex2 leads to a high frequency of I-SceI-resistant EJ products between proximal DSB ends (Proximal-EJ), which can be examined by PCR amplification (primers p1 and p2) and I-SceI digestion analysis. Such expression also leads to I-SceI-resistant EJ products between distal DSB ends that are marked by restoration of a GFP+ cassette (Distal-EJ). For comparison, a relative measure of distal end use is quantified by dividing the distal EJ value by the proximal EJ value for individual samples. B, expression of I-SceI and Trex2 as a fusion protein leads to a high frequency of I-SceI-resistant proximal EJ products. U2OS cells with an integrated copy of the EJ5-GFP reporter were left untransfected, or transfected with expression vectors for I-SceI fused with either Trex2-WT or Trex2-H188A (nuclease dead). Subsequently, proximal EJ was evaluated as described in A. Shown are amplification products that are uncut (U) or digested with the I-SceI endonuclease (S). C, treating cells with kinase inhibitors of either ATM or DNA-PKcs cause a substantial shift toward distal end use. The U2OS EJ5-GFP cell line was transfected with an expression vector for the I-SceI-Trex2 fusion, and treated with a kinase inhibitor of ATM (ATMi, KU55933, 10 μm), a kinase inhibitor of DNA-PKcs (PKi, NU7026, 20 μm), or vehicle (DMSO). Shown are the frequencies of distal EJ (%GFP+), proximal EJ (%I-SceI-resistant, p1/p2), and relative distal end use (distal EJ/proximal EJ, DMSO = 1) from these transfections. Asterisks denote statistical difference from DMSO samples, p < 0.0022. D, treating cells with a kinase inhibitor of DNA-PKcs causes elevated distal EJ in WT mES cells but not DNA-PKcs−/− mES cells. Using WT and DNA-PKcs−/− mES cell lines the EJ5-GFP reporter was integrated into the pim1 locus, I-SceI was co-expressed with Trex2, and transfections were treated with PKi or DMSO as in C. Shown are distal EJ values for these transfections. The asterisk denotes a statistical difference from DMSO samples (p = 0.001).
Thus, using this reporter system, we can quantify two I-SceI-resistant EJ products from the same sample: proximal EJ by PCR amplification and I-SceI digestion analysis, and distal EJ by FACS analysis of the percentage of GFP+ cells. Accordingly, the ratio of distal EJ and proximal EJ can provide a relative measure of incorrect end use (distal end use). In considering the interpretation of these experiments, it is important to note that distal end use could arise via at least two mechanisms. For one, delayed repair could cause persistent DSBs, which would increase the probability of simultaneous tandem DSBs, which could cause an increase in distal end use. Alternatively, defects in end tethering across each DSB could lead to loss of the intervening segment, and hence distal end use. Accordingly, this reporter system does not enable a distinction between these mechanisms, which furthermore, may be interrelated. Rather, these experiments provide the relative frequency of distal end use, which can be used to define the factors and pathways important to limit incorrect end use during EJ.
To simplify this experimental approach for our current study, we developed an expression vector for a fusion protein between I-SceI and Trex2 (I-SceI-Trex2), which we then evaluated using the EJ5-GFP reporter integrated into a human osteosarcoma cell line (U2OS). We found that transfection of the I-SceI-Trex2 expression vector caused a significant increase in I-SceI-resistant proximal EJ products (Fig. 1B). In contrast, we found that transfecting an expression vector for a fusion between I-SceI and an exonuclease-deficient mutant of Trex2 (H188A) (43) did not lead to I-SceI-resistant proximal EJ products, similar to untransfected cells (Fig. 1B). Thus, expressing an I-SceI-Trex2 fusion protein causes a high frequency of proximal EJ, which is consistent with previous studies using I-SceI with Trex2 expressed as separate proteins (7, 37).
Kinase Activity of ATM and DNA-PKcs Are Each Important to Limit Distal End Use
Using the EJ5-GFP reporter system and the I-SceI/Trex2 approach, a previous study (7) showed that genetic and chemical disruption of the ATM kinase causes elevated distal end use. We postulated that another PI3K-related protein kinase, DNA-PKcs, might also play a role in maintaining correct end use, particularly because this factor can tether DNA ends in vitro (21). However, genetic disruption of DNA-PKcs, or other c-NHEJ factors (XRCC4 or XLF), each cause a loss of I-SceI-resistant proximal EJ products (7). These findings support the notion that c-NHEJ is important for repair of noncohesive ends (13, 44), such as those that arise from Trex2 processing of I-SceI-induced DSBs. Without measurable proximal EJ in the DNA-PKcs−/− cells, it is difficult to ascertain the role of DNA-PKcs during end use in this experiment. As an alternative, we considered the possibility that disruption of the kinase activity of DNA-PKcs might allow for sufficient c-NHEJ to facilitate proximal EJ, but may lead to an elevated frequency of incorrect end use.
To test this notion, we examined the effect of a chemical inhibitor of DNA-PKcs (NU7026, PKi) on repair of the EJ5-GFP reporter in U2OS cells, using the I-SceI-Trex2 approach. NU7026 shows at least 2 orders of magnitude selectivity for inhibition of DNA-PKcs versus ATM (17). For comparison, we also examined a chemical inhibitor of ATM kinase activity (KU55933, ATMi), where the selectivity of ATMi for the ATM kinase versus DNA-PKcs is also over 2 orders of magnitude (42). From these experiments (Fig. 1C), we found that treating cells with either PKi or ATMi caused a substantial increase in distal EJ (3.7- and 4.3-fold, respectively), no increase in proximal EJ (PKi caused a 1.9-fold reduction), leading to a substantial shift toward distal end use (6.8- and 4.6-fold, respectively). Using the same experimental approach, we examined the effect of PKi treatment on distal EJ in DNA-PKcs−/− and WT mouse embryonic stem (mES) cells, each with an integrated copy of the EJ5-GFP reporter. As mentioned above, proximal EJ is not detectable in DNA-PKcs−/− mES cells (7), such that we are limited to an analysis of only distal EJ frequencies. From these experiments, we found that PKi did not affect distal EJ in DNA-PKcs−/−, but caused a substantial increase in distal EJ in WT mES cells (3-fold, Fig. 1D). These findings indicate that inhibition of DNA-PKcs kinase activity leads to a greater frequency of incorrect end use during repair of multiple chromosomal DSBs.
RAD50 Is Important to Limit Distal End Use, but H2AX Is Not
We next examined the role of additional DNA damage response factors in maintaining correct end use, specifically RAD50 and H2AX, because these factors are important for chromosomal stability. RAD50 is a member of the MRE11-RAD50-NBS1 complex, which is recruited early to DSBs and can facilitate DSB end tethering and processing (15, 23, 45). Consistent with a key role in chromosomal stability, cells from a patient with an inherited genetic deficiency in RAD50 show a high frequency of spontaneous chromosomal translocations (11). H2AX is a histone variant phosphorylated at serine 139 at sites of DNA damage (46) by PI3K-related protein kinases (e.g. ATM and DNA-PKcs)(47), and appears to be important for protection of DSB ends from extensive processing during repair (33–35). Although H2AX is not required for EJ during V(D)J recombination (coding joint formation), it is important to limit the frequency of chromosomal translocations that can arise from this process (48).
To examine the role of RAD50 in maintaining correct end use during EJ, we performed RNAi depletion of RAD50 in U2OS EJ5-GFP cells (2.5-fold depletion by immunoblot, Fig. 2A), prior to expression of the I-SceI-Trex2 fusion. In comparison to parallel transfections of a nontargeting siRNA (siCTRL), we found that depletion of RAD50 caused an increase in distal EJ, but no change in proximal EJ, thereby causing a shift toward distal end use (1.8-fold, Fig. 2A). Notably, the fold-effect on distal end use caused by RAD50 depletion is similar to its fold-effect on homologous repair (shown below in Fig. 3C). Thus, RAD50 appears important for correct end use during repair of multiple DSBs, which is consistent with previous findings showing a role for NBS1 during this process in mES cells (7).
FIGURE 2.
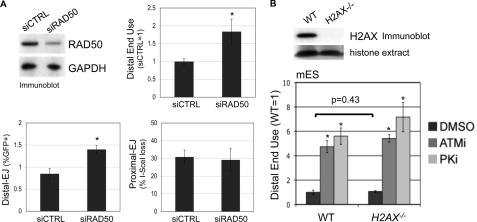
RAD50 is important to limit distal end use, whereas H2AX appears dispensable. A, RAD50 depletion causes a shift toward distal end use. RAD50 was depleted from U2OS cells by transfecting a pool of four targeting siRNAs, which are compared with transfection of a nontargeting siRNA (siCTRL). Shown are immunoblot signals for RAD50 and GAPDH from such transfections. Subsequent to this depletion, the siRNAs were co-transfected with an expression vector for the I-SceI-Trex2 fusion. Shown are Distal-EJ, proximal EJ, and relative distal end use (siCTRL = 1) values from these experiments. Asterisks denote a statistical difference from siCTRL samples (p < 0.0001). B, genetic disruption of H2AX has no clear effect on distal end use. The EJ5-GFP reporter was integrated into the pim1 locus into WT (H2AXf/f) and H2AX−/− mES cells, and these cell lines were co-transfected in parallel with expression vectors for I-SceI and Trex2, and treated with ATMi (10 μm), PKi (20 μm), or vehicle (DMSO). Shown are immunoblot signals for H2AX along with Ponceau S staining of the same blot for histone extracts from these cell lines. Shown are relative distal end use values for these experiments (WT, DMSO = 1), calculated from the ratio of distal EJ and proximal EJ frequencies for individual samples. Asterisks denote a statistical difference from DMSO-treated samples (p < 0.0001). The bracket depicts no statistical difference for the frequency of distal end use between WT and H2AX−/− samples.
FIGURE 3.
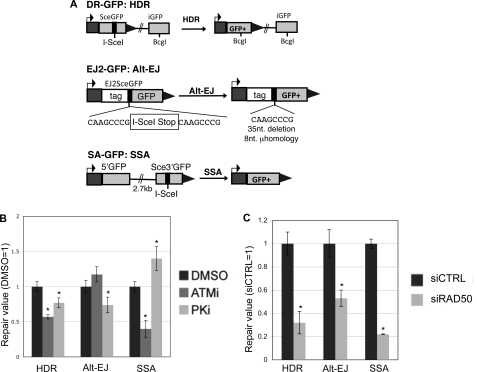
The ATM kinase inhibitor, the DNA-PKcs kinase inhibitor, and depletion of RAD50 each have distinct effects on repair events requiring DSB end processing. A, shown are diagrams of I-SceI reporters to examine repair events requiring DSB end processing: HDR (DR-GFP), alt-EJ (EJ2-GFP), and SSA (SA-GFP). Each of these reporters were individually integrated into U2OS cells to generate a panel of reporter cell lines. B, ATMi treatment causes a decrease in HDR and SSA, but not alt-EJ; whereas PKi treatment causes a decrease in HDR and alt-EJ, but an increase in SSA. Each reporter cell line described in A was transfected with I-SceI and treated with ATMi (10 μm), PKi (20 μm), or vehicle (DMSO). Subsequently, the frequency of the GFP+ repair outcome was determined for each cell line by FACS analysis. Shown are these frequencies relative to parallel transfections treated with vehicle. Asterisks denote a statistical difference from DMSO-treated samples (p < 0.0011). C, RAD50 depletion causes a decrease in HDR, alt-EJ, and SSA. RAD50 was depleted in the U2OS cell lines described in A using the siRNA pool described in Fig. 2A, prior to co-transfection of an I-SceI expression vector with the siRNA. Shown are the frequencies of each repair outcome (marked by GFP+) from these experiments, relative to parallel transfections using a nontargeting siRNA (siCTRL). Asterisks denote a statistical difference from siCTRL-treated samples (p < 0.0012).
To determine whether H2AX affects end use during repair of the EJ5-GFP reporter, we targeted this reporter to the pim1 locus into two related mES cell lines. The first is an H2AX proficient cell line (H2AXf/f, WT), which has been targeted with loxP sites flanking the H2AX genes without disrupting H2AX expression (41). The second cell line is H2AX deficient (H2AX−/−), which was generated via CRE recombinase in the H2AXf/f cells (41). After targeting EJ5-GFP to the pim1 locus of these cell lines, we re-confirmed their H2AX status by immunoblot (Fig. 2B), and then expressed I-SceI and Trex2 to analyze end use during EJ. Furthermore, we also determined the effect of ATMi and PKi treatment on end use during EJ in these cells, as described above (Fig. 1B). From these experiments, we observed no distinction between WT and H2AX−/− cells (Fig. 2B). Namely, the frequency of distal end use was not affected by H2AX loss, and ATMi or PKi caused a similar shift toward distal end use irrespective of H2AX genotype. Accordingly, these findings indicate that H2AX is dispensable for limiting distal end use. Given that H2AX appears important for protecting DSB ends from processing (33–35), we suggest that such H2AX-mediated end protection may not be essential for maintaining correct end use during EJ repair of multiple DSBs.
ATM, DNA-PKcs, and RAD50 Have Distinct Roles in Repair Pathways Requiring Access to Homology
The above findings with H2AX indicate that the roles of a genetic factor during DSB end processing may not be predictive of its requirement in maintaining correct end use during EJ. Similarly, whereas NBS1 and ATM were each previously shown to be important for correct end use in mES cells using the EJ5-GFP assay, only NBS1 appeared important to promote end processing during EJ, based on the frequency of short deletions (7). To examine this relationship further, we investigated the effect of ATM, DNA-PKcs, and RAD50 on the frequencies of a set of DSB repair events that require end processing for access to homology.
For this, we examined reporters for HDR, SSA, and alt-EJ (Fig. 3A), which have been described previously (36). DR-GFP is designed to measure homology-directed repair (HDR), which involves RAD51-mediated strand invasion and nascent DNA synthesis (49–51). DR-GFP contains a GFP cassette that has been interrupted by an I-SceI recognition site, and also contains a downstream homologous GFP fragment (iGFP). HDR of the I-SceI-induced DSB that uses iGFP as the template restores a functional GFP+ cassette. In contrast to HDR, SSA involves annealing of homologous repeats that flank a DSB, which causes a deletion between the regions of homology. SSA can be monitored by SA-GFP, which contains a 5′ GFP fragment separated by 2.7 kilobases (kb) from a 3′ GFP fragment that contains an I-SceI recognition site. SSA between these two GFP fragments that share 266 nucleotides of homology restores a functional GFP+ cassette (52). Finally, EJ2-GFP contains an I-SceI site flanked by 8 nucleotides of microhomology, which if used to bridge the DSB during alt-EJ, can restore a functional GFP+ cassette, and lead to a short 35-nucleotide deletion (36).
We integrated these reporters into U2OS cells, and examined the frequency of each repair outcome following transient transfection of an I-SceI expression vector, using chemical treatments and RNAi disruptions as described above. From these experiments, we found that ATMi treatment caused a decrease in both HDR and SSA (1.7- and 2.5-fold, respectively), but did not cause an effect on the frequency of alt-EJ (Fig. 3B). These findings are consistent with a role for ATM in promoting homologous recombination, but being dispensable for EJ events that require short end processing (8, 25, 53). In contrast, PKi caused a decrease in HDR and alt-EJ (each 1.3-fold), but an increase in SSA (1.4-fold, Fig. 3B). These results are supported by previous findings showing that chemical inhibitors of DNA-PKcs cause a decrease in HDR, but an increase in SSA (24, 54). Finally, depletion of RAD50 by RNAi caused a decrease in each repair outcome (3-fold for HDR, 1.9-fold for alt-EJ, and 4.5-fold for SSA, Fig. 3C). Thus, RAD50 is important for each of these repair events that require access to homology, similar to previous results with NBS1 and the end resection factor CtIP (36, 37). These findings indicate that each of these factors play a distinct role during repair events requiring end processing. Accordingly, we suggest that the role of an individual factor during repair requiring end processing is not predictive of its role for maintaining correct end use during EJ.
DSB Transcription Context Does Not Affect the Frequency of HDR, or End Protection during EJ
In each of the above HDR and EJ reporters, at least one of the I-SceI recognition site(s) is downstream from a strong constitutive promoter (pCAGGS: CMV enhancer, chicken β-actin promoter) (55). Such transcription could conceivably influence the mechanisms or fidelity of repair, as the transcription state of some loci appears to affect the frequency of chromosomal rearrangements (27–30). To examine HDR of a DSB under different transcription contexts, we developed a reporter similar to DR-GFP (38): g4DRGFP (Fig. 4A). In this reporter, the promoter for the SceGFP cassette in DR-GFP was replaced with the g4 promoter, which is five tandem repeats of the Gal4 DNA binding sequence followed by a TATA box. We chromosomally integrated g4DRGFP into two human cell lines, HEK293-A7 and U2OS-R1, both of which constitutively express two transcription regulator proteins: Gal4-DBD-EcR, and RxR-VP16. Gal4-DBD-EcR is a synthetic ecdysone receptor fused to a Gal4 DNA binding domain. Upon the addition of a soluble ligand (L), a conformational change allows Gal4-DBD-EcR to bind RxR-VP16 leading to VP16-mediated transcription of the g4 promoter (40). To validate the L-inducible control of transcription of g4DRGFP, we isolated RNA after 3 h in the presence or absence of L and analyzed the reverse transcripts via quantitative real-time PCR. We found that this brief L treatment caused nearly a 1000-fold increase in mRNA abundance relative to cells cultured in the absence of L (Fig. 4B).
FIGURE 4.
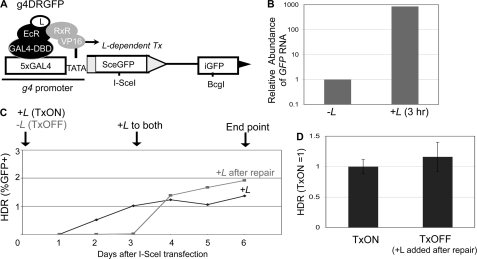
The frequency of HDR is not affected by DSB transcription context. A, schematic of g4DRGFP. The SceGFP cassette is shown downstream of GAL4 recognition sites and a minimal TATA promoter (g4). HDR of an I-SceI-induced DSB using iGFP as the template restores a GFP+ cassette, as shown in Fig. 3A. Shown are the regulator proteins EcR-GAL4-DBD and RxR-VP16, which require the presence of a soluble ligand (L) bound to EcR to induce transcription. B, addition of L leads to rapid induction of mRNA expression from the g4 promoter. The g4DRGFP reporter was integrated into two cell lines that stably express the EcR-GAL4-DBD and RxR-VP16 proteins (HEK293-A7 and U2OS-R1). Using the g4DRGFP HEK293-A7 cell line, RNA was isolated after a 3-h incubation in media containing +L or untreated (−L). Samples were reverse transcribed prior to quantitative PCR. Shown is the relative abundance of GFP RNA, based on 2ΔΔCt values normalized to actin primers and relative to the untreated condition (−L = 1). C, time course to establish treatment conditions for TxON and TxOFF samples. The g4DRGFP HEK293-A7 cell line was transfected in the presence (TxON) or absence (TxOFF) of L for 3 days, at which time L was added to all the samples for an additional 3 days of culture (6 days total). Shown is the frequency of GFP+ cells from these samples each day post-transfection. D, DSB transcription context has no effect on the frequency of HDR. An expression vector for I-SceI was transfected in the g4DRGFP U2OS-R1 cell line under the TxON and TxOFF conditions described in C. Shown is the frequency of HDR for both Tx conditions from this experiment, relative to the TxON condition (TxON = 1).
As with the parental DR-GFP reporter, downstream of SceGFP is an internal fragment of GFP (iGFP), which if used as a template for HDR of an I-SceI-induced DSB, leads to the restoration of a GFP+ cassette. Accordingly, following expression of I-SceI and completion of repair, L is added to all samples to detect the GFP+ cells. In our experience, 3 days post-transfection is sufficient for completion of repair. To confirm this notion, we analyzed GFP expression by FACS analysis in a time course following transfection of an I-SceI expression vector (Fig. 4C). To examine repair of DSBs downstream from an active promoter (TxON), L was included throughout the experiment. With such samples, 3 days was sufficient to achieve steady GFP frequencies (Fig. 4C). Accordingly, to examine repair of DSBs downstream from an uninduced promoter (TxOFF), we transfected cells with an expression vector for I-SceI in the absence of L, and maintained this culturing condition for the first 3 days post-transfection. Consistent with the requirement of L for GFP expression, the TxOFF samples did not show significant GFP+ cells at this time point (Fig. 4C). Subsequently, we added L to all the samples for an additional 3 days (6 days total), which was sufficient to achieve steady levels of GFP expression for both TxOFF and TxON samples. Thus, for each experiment below, the TxON condition includes +L, starting 3 h prior to transfection and through the entire 6-day experiment, whereas the TxOFF condition involves culturing without L through 3 days post-transfection, followed by 3 days of treatment +L (Fig. 4C).
Using the above experimental conditions for parallel transfections with the I-SceI expression vector, we found that the frequency of HDR was not distinct between the TxON and TxOFF samples (Fig. 4D). Thus, the frequency of HDR is not affected by the DSB transcription context, which is consistent with previous reports using HDR reporters with dexamethasone and doxycycline-inducible promoters (31, 32).
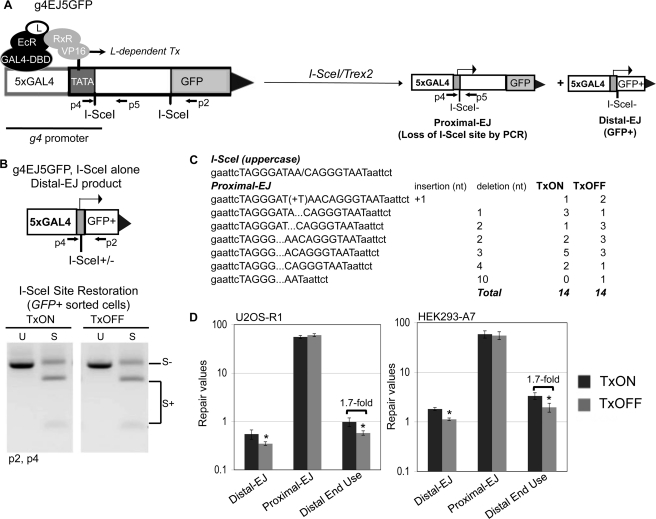
To address the effect of the transcription context on chromosomal end joining, we developed a second reporter, g4EJ5GFP (Fig. 5A), using the same inducible promoter as described above. Namely, the pCAGGS promoter upstream of the 5′ I-SceI site in EJ5-GFP was replaced with the g4 promoter. This reporter was integrated into HEK293-A7 and U2OS-R1 cells, which express transcription regulator proteins as described above. With these cell lines, we examined EJ in both TxON and TxOFF samples, using the same treatment conditions described above for the g4DRGFP reporter (see Fig. 4C).
FIGURE 5.
DSB transcription context affects maintenance of correct end use, but not the degree of end protection during EJ. A, schematic of g4EJ5GFP. Similar to EJ5-GFP, a fragment of an expression cassette including the entire GFP coding sequence is shown separated from the g4 promoter by a 1.8-kb segment flanked by two tandem I-SceI recognition sites. Distal EJ repair of the two tandem I-SceI sites places the g4 promoter directly upstream of rest of the GFP cassette. Shown are the regulator proteins that mediate L-induced transcription (Tx) from the g4 promoter, which are expressed in HEK293-A7 and U2OS-R1. The g4EJ5-GFP reporter was integrated into both of these cell lines. Also shown are diagrams of proximal EJ and distal EJ products resulting from I-SceI/Trex2 expression, both of which involve EJ of the 5′ DSB, which is directly downstream from the g4 promoter. B, the frequency of I-SceI restoration during distal EJ is not affected by transcription context. The g4EJ5-GFP HEK293-A7 cell line was transfected with an I-SceI expression vector in the TxON and TxOFF conditions, and subsequently GFP+ cells were sorted to enrich for distal EJ products that were amplified (primers p2, p4). Shown are representative products digested with I-SceI (S) or uncut (U). C, expression of the I-SceI-Trex2 fusion causes predominantly short deletions to the I-SceI overhang in proximal EJ products, irrespective of DSB transcription context. The g4EJ5GFP U2OS-R1 cell line was transfected with an expression vector for I-SceI-Trex2 fusion in the TxON and TxOFF conditions. Subsequently, I-SceI-resistant proximal EJ products (primers p4 and p5, shown in A) were isolated and cloned for sequencing. Shown are frequencies of sequence junctions for individual clones from the TxON and TxOFF conditions, along with the reference I-SceI site. D, the relative frequency of distal end use is lower in the TxOFF condition, compared with TxON. Cells with the g4EJ5GFP reporter were transfected with I-SceI/Trex2 (separate I-SceI and Trex2 plasmids for HEK293-A7, and the I-SceI-Trex2 fusion for U2OS-R1), under the TxON and TxOFF conditions. Shown are frequencies of distal EJ, proximal EJ, and distal end use from these transfections. Asterisks denote a statistical difference between the two Tx conditions (p < 0.003).
To determine whether transcription context of a DSB affects end protection during EJ, we first performed experiments expressing I-SceI alone (without Trex2). Induction of DSBs by I-SceI alone can lead to distal EJ products that use the I-SceI overhangs to restore the I-SceI site (36). Such I-SceI restoration events are promoted by c-NHEJ factors Ku70 and Xrcc4 (36, 37). To examine the effect of the transcription context on I-SceI restoration during distal EJ, we expressed I-SceI in both the TxON and TxOFF conditions. Subsequently, we sorted GFP+ cells and examined the restoration of the I-SceI site by amplifying the repair junction, followed by I-SceI digestion analysis. From this experiment, we found no difference in the frequency of I-SceI restoration between the TxON and TxOFF samples (Fig. 5B).
We also addressed whether transcription context affects end protection during EJ using the I-SceI/Trex2 approach. Namely, we transfected the U2OS-R1 g4EJ5GFP cell line with the I-SceI-Trex2 expression vector, under both TxON and TxOFF conditions. Notably, in all the experiments with g4EJ5GFP, we examined proximal EJ of the 5′ I-SceI-induced DSB, which is directly downstream from the g4 promoter (Fig. 5A). From this experiment, we cloned I-SceI-resistant proximal EJ products and sequenced individual clones. We predominantly observed partial or complete loss of the I-SceI overhang for both the TxON and TxOFF samples (13/14 and 11/14, respectively, Fig. 5C). Additionally, we observed a few clones with a 1-bp insertion (1/14 for TxON, 2/14 for TxOFF), and one clone with a slightly larger (10 bp) deletion (1/14, TxOFF). Thus, proximal EJ is rarely associated with end processing beyond the I-SceI overhang, which is consistent with previous experiments with co-expression of I-SceI and Trex2 using the EJ5-GFP reporter (37). In summary, these findings indicate that the transcription context of a DSB does not affect the degree of end processing during EJ.
Distal End Use Is More Frequent for a DSB Downstream from an Active Promoter
Using the above g4EJ5-GFP I-SceI/Trex2 experiment under TxON and TxOFF conditions, we also examined maintenance of correct end use, by quantifying the frequency of distal EJ, proximal EJ, and relative distal end use. Importantly, both the proximal EJ and distal EJ events examined in these experiments involve repair of the 5′ I-SceI-induced DSB, which is directly downstream from the g4 promoter (Fig. 5A). We found that the TxON samples, versus TxOFF, showed a higher frequency of distal EJ, but no change in proximal EJ, which leads to an increased frequency of distal end use (1.7-fold for both U2OS-R1 and HEK293-A7 cells, Fig. 5D). These findings indicate that distal end use occurs at a greater frequency when a DSB is downstream from an active promoter, compared with the uninduced promoter. Again, an increase in distal end use in this assay could be caused by greater DSB persistence that leads to a higher probability that both DSBs are cut simultaneously, and/or a reduced fidelity of end tethering. Accordingly, transcription context could be affecting either of these aspects of repair to cause elevated distal end use. Notably, because the transcription context of a DSB did not appear to affect end processing during EJ (Fig. 5, B and C), these findings are consistent with the notion that the degree of end processing during EJ is not predictive of the frequency of correct end use.
A DSB Downstream from an Active Promoter Is More Reliant on ATM for Limiting Distal End Use
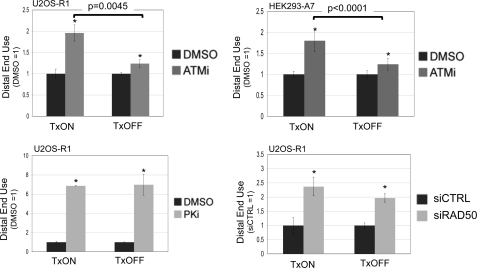
Because the frequency of distal end use is higher when a DSB is downstream from an active promoter, we considered the possibility that the relative requirement for individual factors during end use could be affected by DSB transcription context. In particular, we sought to examine the role of ATM under different DSB transcription contexts, because previous studies indicate that the chromosomal context of a DSB may affect the relative role for ATM during repair (56, 57). To test this notion, we expressed the I-SceI-Trex2 fusion in cell lines with the g4EJ5GFP reporter, using TxON and TxOFF conditions, in the presence of ATMi or vehicle (DMSO). Subsequently, we quantified the frequencies of distal EJ and proximal EJ, to calculate distal end use relative to matched DMSO-treated samples. From these experiments, we found that ATMi treatment caused a significantly greater increase in distal end use in the TxON condition, compared with the TxOFF condition, both in U2OS-R1 cells (2-fold versus 1.2-fold, respectively, Fig. 6), as well as HEK293-A7 cells (1.8- versus 1.2-fold, respectively, Fig. 6). We also determined the effect of PKi treatment and RAD50 depletion on EJ of the g4EJ5GFP reporter in the TxON and TxOFF conditions. In these experiments, PKi treatment and RAD50 depletion each caused an increase in distal end use that was not distinct between the TxON and TxOFF conditions (Fig. 6). Thus, the relative role of DNA-PKcs kinase activity or RAD50 in limiting distal end use is not obviously affected by the DSB transcription context; whereas the role of ATM kinase activity in this process is greater for a DSB downstream from an active promoter, compared with an uninduced promoter.
FIGURE 6.
A DSB downstream from an active promoter shows a greater reliance on ATM to limit distal end use. Cells with the g4EJ5GFP reporter were transfected with I-SceI/Trex2 under both the TxON and TxOFF conditions, and were treated with ATMi, PKi, vehicle (DMSO), or depleted of RAD50. Separate I-SceI and Trex2 expression plasmids were transfected into HEK293-A7 cells also treated with DMSO or 10 μm ATMi, and the I-SceI-Trex2 fusion was transfected into U2OS-R1 cells treated with DMSO, 5 μm ATMi, 20 μm PKi, or transfected with siRNA targeting RAD50 or a nontargeting sequence (siCTRL). Subsequently, the frequency of distal EJ and proximal EJ was determined for individual samples to quantify distal end use, which is shown relative to the parallel DMSO or siCTRL-treated samples for each Tx condition and cell type. Asterisks denote statistical difference from DMSO-treated samples (p < 0.002, except U2OS-R1, TxOFF, and ATMi, p = 0.013). Brackets denote a statistical difference between the fold-effect of ATMi treatment under TxON versus TxOFF conditions.
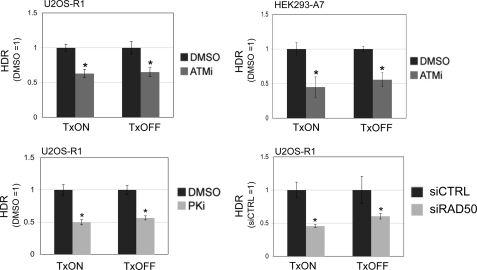
We then sought to examine whether the effect of DSB transcription context on the role of ATM during repair is limited to end use during EJ, or also includes its role in promoting HDR. For this, we transfected the I-SceI expression vector into cell lines with the g4DRGFP reporter under both TxON and TxOFF conditions, and included treatment with ATMi or vehicle (DMSO). In these experiments, we found that ATMi treatment caused a decrease in HDR that was not distinct for TxON or TxOFF samples, for both U2OS-R1 and HEK293-A7 cell lines (Fig. 7). Furthermore, using the same approach, we examined the effect of PKi treatment and RAD50 depletion on HDR in TxON and TxOFF conditions. We found that the decrease in HDR caused by PKi treatment or RAD50 depletion was not distinct for the TxON and TxOFF samples (Fig. 7). These findings indicate that the relative role of ATM kinase activity, DNA-PKcs kinase activity, and RAD50 during HDR is not affected by DSB transcription context. Accordingly, the transcription context of a DSB appears to affect the relative requirement of ATM kinase activity for maintaining correct end use during EJ, but not for promoting HDR.
FIGURE 7.
The role of ATM kinase activity, DNA-PKcs kinase activity, and RAD50 during HDR is unaffected by DSB transcription context. Cells with the g4DRGFP reporter were transfected with an I-SceI expression vector under TxON or TxOFF conditions, and treated with ATMi, PKi, DMSO, or were depleted of RAD50 via siRNA. HDR values are shown relative to parallel control samples (e.g. DMSO-treated or siCTRL-treated) from the same Tx condition. For both TxON and TxOFF conditions, shown is the frequency of HDR of ATMi-treated HEK293-A7 (5 μm) and U2OS-R1 cells (10 μm), PKi-treated U2OS-R1 cells (20 μm), and RAD50-depleted U2OS-R1 cells. Asterisks indicate a statistically significant decrease in HDR compared with parallel control samples in the same Tx condition (p < 0.0018, except siRAD50 TxOFF, p = 0.03).
DISCUSSION
Maintenance of correct end use during EJ repair of multiple DSBs is critical to limit chromosomal rearrangements. By examining a reporter system for EJ repair of two tandem chromosomal DSBs, we present evidence that a number of DNA damage response factors are important for maintaining correct end use (i.e. ATM, DNA-PKcs, and RAD50), that the transcription context of a DSB can affect the frequency of correct end use, and that a DSB downstream from an active promoter shows a greater reliance on ATM for correct end use. In contrast, individual DNA damage response factors and DSB transcription context play distinct roles during repair events that require end processing. We suggest that the influence of individual factors and DSB transcription context on maintaining correct end use during EJ cannot be predicted from their effects on repair involving end processing, such that these may be distinct aspects of DSB repair.
Correct End Use versus DSB End Processing: Distinct Roles of Individual DNA Damage Response Factors
We present evidence that DNA-PKcs kinase activity and RAD50 are each important for limiting incorrect end use during repair of two tandem DSBs (i.e. distal end use). This reporter system cannot distinguish between the contributions of DSB persistence versus incorrect end tethering to the frequency of distal end use, which furthermore, may be interrelated aspects of DSB repair. Accordingly, DNA-PKcs or RAD50 could be affecting one or more of these processes to limit distal end use.
For example, DNA-PKcs could be important for correct end tethering and/or limiting DSB persistence. In support of an end tethering function, DNA-PKcs promotes formation of a synaptic complex during EJ in vitro (21, 22). Considering a potential role in limiting DSB persistence, blocking the kinase activity of DNA-PKcs could decrease the rate of EJ, thereby increasing the probability that multiple DSBs persist simultaneously, which could lead to an elevated frequency of distal end use. Regarding this possibility, blocking autophosphorylation of DNA-PKcs has been shown to cause its persistent retention at DSBs, as well as delayed DSB repair (58, 59). Such autophosphorylation may be important to release DNA-PKcs during later steps of EJ, because a blocked synaptic complex could delay ligation (60, 61).
Along these lines, we find that genetic disruption of DNA-PKcs versus inhibition of its kinase activity (PKi treatment) cause distinct effects on EJ repair of two tandem DSBs. Namely, we find that PKi treatment causes a substantial increase in distal EJ and a moderate decrease in proximal EJ; whereas genetic disruption of DNA-PKcs fails to cause an increase in the frequency of distal EJ, but rather causes a significant loss of proximal EJ products (7). We suggest that blocking DNA-PKcs kinase activity does not cause a complete disruption of EJ, but rather alters the dynamics of EJ leading to elevated incorrect end use that could cause chromosomal rearrangements.
Notably, kinase inactivation of DNA-PKcs causes greater radiosensitivity than genetic loss of this factor (3, 54). As well, kinase inhibition of DNA-PKcs combined with ATM kinase deficiency causes a synergistic increase in chromosomal rearrangements during lymphocyte development (62). Accordingly, our findings raise the possibility that the role of DNA-PKcs kinase activity in limiting incorrect end use during EJ may be critical for radioresistance and suppression of chromosomal rearrangements.
Similar to the above models with DNA-PKcs, depletion of RAD50 could affect DSB persistence and/or end tethering. Consistent with a role in limiting DSB persistence, genetic deficiencies in the MRE11-RAD50-NBS1 complex have been shown to cause persistent coding DSB ends during repair of inverted V(D)J recombination substrates (6). Regarding end tethering, the MRE11-RAD50-NBS1 complex promotes in vitro DNA tethering and EJ via DNA ligase III (15, 18–20), and the MRE11-RAD50-XRS2 complex is important in Saccharomyces cerevisiae for maintaining spatial continuity of a broken chromosome and for suppressing chromosomal translocations (63, 64). In summary, we suggest that the functions of DNA-PKcs kinase activity and RAD50 in limiting the persistence of DSBs and/or promoting correct end tethering could contribute to their roles in limiting incorrect end use during EJ of multiple DSBs.
In contrast, we find that RAD50 and DNA-PKcs kinase activity have opposite effects on SSA: RAD50 promotes SSA, whereas DNA-PKcs suppresses this repair event. Previous studies have also shown that genetic disruption or kinase inhibition of DNA-PKcs, or genetic disruption of other c-NHEJ factors, cause an increase in the frequency of SSA (24, 52); whereas disruption of another component of the MRE11-RAD50-NBS1 complex (NBS1) causes a decrease in SSA (37, 65). As SSA requires extensive end processing to reveal the homology that flanks the DSB, these findings support the notion that c-NHEJ factors, including DNA-PKcs, are important to limit such extensive DSB end processing, whereas RAD50 and NBS1 promote this process. Considering limited end processing during EJ (EJ2-GFP, alt-EJ with a 35-bp deletion), we find that both RAD50 and DNA-PKcs kinase activity promote such repair, whereas ATM is dispensable. Furthermore, previous reports show that ATM is important to protect terminal DSB ends from processing during EJ, whereas NBS1 and DNA-PKcs promote such processing (7, 8, 37, 66). Thus, ATM, DNA-PKcs, and RAD50 each have distinct roles during repair events requiring varying degrees of end processing, whereas each of these factors is important to limit distal end use.
Consistent with these distinctions, H2AX, which is important for suppression of DSB end processing and SSA (33–35), appears dispensable for limiting distal end use. Furthermore, H2AX did not affect the relative requirement for ATM and DNA-PKcs kinase activity during this process. However, H2AX-deficient cells have been shown to exhibit elevated chromosomal rearrangements (41, 67, 68). Perhaps such rearrangements specifically occur by SSA, because this pathway has been shown to contribute to chromosomal translocations in reporter assays (69). The tandem DSB system described here only measures correct end use during EJ repair, such that examining this process during SSA of multiple DSBs may provide additional insight into the role of H2AX in promoting chromosomal stability. As another possibility, H2AX may be important for maintaining correct end use during EJ of the specific types of programmed DSBs that arise during lymphocyte development (67, 70). In summary, these findings are consistent with the notion that the role of a factor during DSB end processing is not predictive of its requirement for maintaining correct end use during EJ, such that these may be distinct processes of genome maintenance.
A DSB Downstream from an Active Promoter Is More Prone to Incorrect End Use and Is More Reliant on ATM for This Process
We have also examined the effect of DSB transcription context on repair, by developing a set of reporters that position DSBs downstream from an inducible promoter. We found that the frequency of HDR and the degree of end protection (i.e. deletion size) during EJ were not affected by DSB transcription context. Both of these findings are consistent with previous studies of the effects of DSB transcription context on repair using other inducible promoters (31, 32). In contrast, we found a relatively higher frequency of distal end use when a DSB is downstream from an active promoter. Similar to the above discussion of DNA damage response factors, the effect of transcription context on distal end use could be caused by delayed EJ and/or defects in DSB end tethering. Importantly, because DSB transcription context affects correct end use but not end protection during EJ, these findings support the notion that these are distinct aspects of repair.
Notably, our findings are consistent with previous studies indicating that transcribed loci may be particularly prone to chromosomal rearrangements (27). For instance, a chromosomal region that is highly prone to forming translocations in anaplastic large lymphoma shows aberrantly high transcription in this tumor type, even in cells that lack the translocation (29). As another example, activation of androgen receptor in prostate cells leads to an increased frequency of chromosomal translocations of its target loci (28, 30). In these studies, such transcriptional activity correlated with closer nuclear proximity between translocation partners, which may contribute to the increase in translocation frequency (28–30). In addition, transcription leads to DNA/RNA hybrids (R-loops), which have been shown to cause persistent DSBs and chromosomal instability (71, 72). We suggest that chromosomal DBSs in transcribed loci may also be prone to EJ events that fail to maintain correct end use, which could contribute to such chromosomal rearrangements.
We also examined the relative role of individual DNA damage response factors during EJ and HDR under different transcription contexts. From this analysis, we found that ATMi treatment caused a greater increase in distal end use when a DSB was downstream from an active promoter. In contrast, DSB transcription context did not affect the relative role of DNA-PKcs and RAD50 in maintaining correct end use, nor the role of ATM, DNA-PKcs, or RAD50 in promoting HDR. These findings indicate that ATM may be particularly important for correct end use during repair of DSBs that are downstream from an active promoter, which is consistent with previous studies indicating that chromosomal context may affect the role of ATM during DSB repair. For example, ATM was recently shown to play a role in the suppression of transcription caused by an array of adjacent DSBs (56). Also, chromatin remodeling factors can influence the activation of ATM during the DNA damage response (73). Along these lines, from examining cells with significant heterochromatin, ATM has been shown to be particularly important for repair of DSBs within heterochromatin, as measured by clearance of the DSB marker, γH2AX (57). We suggest that maintenance of correct end use during EJ is an additional role of ATM that appears to be affected by chromosomal context.
In conclusion, each of the DNA damage response factors analyzed here (RAD50, ATM, and DNA-PKcs) are being developed as therapeutic targets for radiosensitization during cancer therapy (16, 17, 42). Given that each of these factors appear to play a role in maintaining correct end use during EJ of multiple DSBs, but play distinct roles during repair events requiring end processing, it will be important to investigate how these distinct repair functions contribute to radioresistance. Such studies will inform the development of targeted therapeutics that disrupt DNA damage response factors to cause tumor radiosensitization.
Acknowledgments
We thank Dr. Corentin Laulier and Meilen Munoz for helpful discussions.
This work was supported by Grant RO1CA120954 from the National Cancer Institute of the NIH (to J. M. S.).
- EJ
- end joining
- ATM
- ataxia telangiectasia-mutated kinase
- ATMi
- ATM kinase inhibitor
- alt-EJ
- alternative end joining
- DSB
- double strand break
- c-NHEJ
- classical nonhomologous end joining
- g4
- GAL4 promoter
- HDR
- homology-directed repair
- L
- synthetic ecdysone receptor ligand
- PKi
- DNA-PKcs kinase inhibitor
- SSA
- single strand annealing
- Tx
- transcription
- mES
- mouse embryonic stem
- DMSO
- dimethyl sulfoxide.
REFERENCES
- 1. Stratton M. R. (2011) Science 331, 1553–1558 [DOI] [PubMed] [Google Scholar]
- 2. Stephens P. J., McBride D. J., Lin M. L., Varela I., Pleasance E. D., Simpson J. T., Stebbings L. A., Leroy C., Edkins S., Mudie L. J., Greenman C. D., Jia M., Latimer C., Teague J. W., Lau K. W., Burton J., Quail M. A., Swerdlow H., Churcher C., Natrajan R., Sieuwerts A. M., Martens J. W., Silver D. P., Langerød A., Russnes H. E., Foekens J. A., Reis-Filho J. S., van't Veer L., Richardson A. L., Børresen-Dale A. L., Campbell P. J., Futreal P. A., Stratton M. R. (2009) Nature 462, 1005–1010 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3. Mahaney B. L., Meek K., Lees-Miller S. P. (2009) Biochem. J. 417, 639–650 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4. Bredemeyer A. L., Huang C. Y., Walker L. M., Bassing C. H., Sleckman B. P. (2008) J. Immunol. 181, 2620–2625 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5. Deriano L., Stracker T. H., Baker A., Petrini J. H., Roth D. B. (2009) Mol. Cell 34, 13–25 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6. Helmink B. A., Bredemeyer A. L., Lee B. S., Huang C. Y., Sharma G. G., Walker L. M., Bednarski J. J., Lee W. L., Pandita T. K., Bassing C. H., Sleckman B. P. (2009) J. Exp. Med. 206, 669–679 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7. Bennardo N., Stark J. M. (2010) PLoS Genet. 6, e1001194. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8. Rahal E. A., Henricksen L. A., Li Y., Williams R. S., Tainer J. A., Dixon K. (2010) Cell Cycle 9, 2866–2877 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9. Shiloh Y. (2003) Nat. Rev. Cancer 3, 155–168 [DOI] [PubMed] [Google Scholar]
- 10. van der Burgt I., Chrzanowska K. H., Smeets D., Weemaes C. (1996) J. Med. Genet. 33, 153–156 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11. Waltes R., Kalb R., Gatei M., Kijas A. W., Stumm M., Sobeck A., Wieland B., Varon R., Lerenthal Y., Lavin M. F., Schindler D., Dörk T. (2009) Am. J. Hum. Genet. 84, 605–616 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12. van der Burg M., Ijspeert H., Verkaik N. S., Turul T., Wiegant W. W., Morotomi-Yano K., Mari P. O., Tezcan I., Chen D. J., Zdzienicka M. Z., van Dongen J. J., van Gent D. C. (2009) J. Clin. Invest. 119, 91–98 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13. Lieber M. R. (2010) Annu. Rev. Biochem. 79, 181–211 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14. Meek K., Dang V., Lees-Miller S. P. (2008) Adv. Immunol. 99, 33–58 [DOI] [PubMed] [Google Scholar]
- 15. Stracker T. H., Petrini J. H. (2011) Nat. Rev. Mol. Cell Biol. 12, 90–103 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16. Abuzeid W. M., Jiang X., Shi G., Wang H., Paulson D., Araki K., Jungreis D., Carney J., O'Malley B. W., Jr., Li D. (2009) J. Clin. Invest. 119, 1974–1985 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17. Veuger S. J., Curtin N. J., Richardson C. J., Smith G. C., Durkacz B. W. (2003) Cancer Res. 63, 6008–6015 [PubMed] [Google Scholar]
- 18. Della-Maria J., Zhou Y., Tsai M. S., Kuhnlein J., Carney J. P., Paull T. T., Tomkinson A. E. (2011) J. Biol. Chem. 286, 33845–33853 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19. Moreno-Herrero F., de Jager M., Dekker N. H., Kanaar R., Wyman C., Dekker C. (2005) Nature 437, 440–443 [DOI] [PubMed] [Google Scholar]
- 20. Bhaskara V., Dupré A., Lengsfeld B., Hopkins B. B., Chan A., Lee J. H., Zhang X., Gautier J., Zakian V., Paull T. T. (2007) Mol. Cell 25, 647–661 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21. DeFazio L. G., Stansel R. M., Griffith J. D., Chu G. (2002) EMBO J. 21, 3192–3200 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22. Hammel M., Yu Y., Mahaney B. L., Cai B., Ye R., Phipps B. M., Rambo R. P., Hura G. L., Pelikan M., So S., Abolfath R. M., Chen D. J., Lees-Miller S. P., Tainer J. A. (2010) J. Biol. Chem. 285, 1414–1423 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23. Paull T. T. (2010) DNA Repair 9, 1283–1291 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24. Neal J. A., Dang V., Douglas P., Wold M. S., Lees-Miller S. P., Meek K. (2011) Mol. Cell. Biol. 31, 1719–1733 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25. Shrivastav M., Miller C. A., De Haro L. P., Durant S. T., Chen B. P., Chen D. J., Nickoloff J. A. (2009) DNA Repair 8, 920–929 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26. Rooney S., Alt F. W., Lombard D., Whitlow S., Eckersdorff M., Fleming J., Fugmann S., Ferguson D. O., Schatz D. G., Sekiguchi J. (2003) J. Exp. Med. 197, 553–565 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27. Mathas S., Misteli T. (2009) Cell 139, 1047–1049 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28. Lin C., Yang L., Tanasa B., Hutt K., Ju B. G., Ohgi K., Zhang J., Rose D. W., Fu X. D., Glass C. K., Rosenfeld M. G. (2009) Cell 139, 1069–1083 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29. Mathas S., Kreher S., Meaburn K. J., Jöhrens K., Lamprecht B., Assaf C., Sterry W., Kadin M. E., Daibata M., Joos S., Hummel M., Stein H., Janz M., Anagnostopoulos I., Schrock E., Misteli T., Dörken B. (2009) Proc. Natl. Acad. Sci. U.S.A. 106, 5831–5836 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30. Mani R. S., Tomlins S. A., Callahan K., Ghosh A., Nyati M. K., Varambally S., Palanisamy N., Chinnaiyan A. M. (2009) Science 326, 1230. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31. Allen C., Miller C. A., Nickoloff J. A. (2003) DNA Repair 2, 1147–1156 [DOI] [PubMed] [Google Scholar]
- 32. Gottipati P., Cassel T. N., Savolainen L., Helleday T. (2008) Mol. Cell. Biol. 28, 154–164 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33. Xie A., Puget N., Shim I., Odate S., Jarzyna I., Bassing C. H., Alt F. W., Scully R. (2004) Mol. Cell 16, 1017–1025 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34. Helmink B. A., Tubbs A. T., Dorsett Y., Bednarski J. J., Walker L. M., Feng Z., Sharma G. G., McKinnon P. J., Zhang J., Bassing C. H., Sleckman B. P. (2011) Nature 469, 245–249 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35. Bothmer A., Robbiani D. F., Di Virgilio M., Bunting S. F., Klein I. A., Feldhahn N., Barlow J., Chen H. T., Bosque D., Callen E., Nussenzweig A., Nussenzweig M. C. (2011) Mol. Cell 42, 319–329 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36. Bennardo N., Cheng A., Huang N., Stark J. M. (2008) PLoS Genet. 4, e1000110. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37. Bennardo N., Gunn A., Cheng A., Hasty P., Stark J. M. (2009) PLoS Genet. 5, e1000683. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38. Pierce A. J., Johnson R. D., Thompson L. H., Jasin M. (1999) Genes Dev. 13, 2633–2638 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39. Stark J. M., Jasin M. (2003) Mol. Cell. Biol. 23, 733–743 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40. Palli S. R., Kapitskaya M. Z., Kumar M. B., Cress D. E. (2003) Eur. J. Biochem. 270, 1308–1315 [DOI] [PubMed] [Google Scholar]
- 41. Bassing C. H., Chua K. F., Sekiguchi J., Suh H., Whitlow S. R., Fleming J. C., Monroe B. C., Ciccone D. N., Yan C., Vlasakova K., Livingston D. M., Ferguson D. O., Scully R., Alt F. W. (2002) Proc. Natl. Acad. Sci. U.S.A. 99, 8173–8178 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42. Hickson I., Zhao Y., Richardson C. J., Green S. J., Martin N. M., Orr A. I., Reaper P. M., Jackson S. P., Curtin N. J., Smith G. C. (2004) Cancer Res. 64, 9152–9159 [DOI] [PubMed] [Google Scholar]
- 43. Chen M. J., Ma S. M., Dumitrache L. C., Hasty P. (2007) Nucleic Acids Res. 35, 2682–2694 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44. Tsai C. J., Kim S. A., Chu G. (2007) Proc. Natl. Acad. Sci. U.S.A. 104, 7851–7856 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 45. Williams G. J., Lees-Miller S. P., Tainer J. A. (2010) DNA Repair 9, 1299–1306 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46. Rogakou E. P., Boon C., Redon C., Bonner W. M. (1999) J. Cell Biol. 146, 905–916 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47. Yuan J., Adamski R., Chen J. (2010) FEBS Lett. 584, 3717–3724 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48. Yin B., Savic V., Juntilla M. M., Bredemeyer A. L., Yang-Iott K. S., Helmink B. A., Koretzky G. A., Sleckman B. P., Bassing C. H. (2009) J. Exp. Med. 206, 2625–2639 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49. Stark J. M., Hu P., Pierce A. J., Moynahan M. E., Ellis N., Jasin M. (2002) J. Biol. Chem. 277, 20185–20194 [DOI] [PubMed] [Google Scholar]
- 50. Sung P., Krejci L., Van Komen S., Sehorn M. G. (2003) J. Biol. Chem. 278, 42729–42732 [DOI] [PubMed] [Google Scholar]
- 51. Kass E. M., Jasin M. (2010) FEBS Lett. 584, 3703–3708 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52. Stark J. M., Pierce A. J., Oh J., Pastink A., Jasin M. (2004) Mol. Cell. Biol. 24, 9305–9316 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 53. Golding S. E., Rosenberg E., Khalil A., McEwen A., Holmes M., Neill S., Povirk L. F., Valerie K. (2004) J. Biol. Chem. 279, 15402–15410 [DOI] [PubMed] [Google Scholar]
- 54. Allen C., Halbrook J., Nickoloff J. A. (2003) Mol. Cancer Res. 1, 913–920 [PubMed] [Google Scholar]
- 55. Niwa H., Yamamura K., Miyazaki J. (1991) Gene 108, 193–199 [DOI] [PubMed] [Google Scholar]
- 56. Shanbhag N. M., Rafalska-Metcalf I. U., Balane-Bolivar C., Janicki S. M., Greenberg R. A. (2010) Cell 141, 970–981 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57. Goodarzi A. A., Noon A. T., Deckbar D., Ziv Y., Shiloh Y., Löbrich M., Jeggo P. A. (2008) Mol. Cell 31, 167–177 [DOI] [PubMed] [Google Scholar]
- 58. Uematsu N., Weterings E., Yano K., Morotomi-Yano K., Jakob B., Taucher-Scholz G., Mari P. O., van Gent D. C., Chen B. P., Chen D. J. (2007) J. Cell Biol. 177, 219–229 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59. Davis A. J., So S., Chen D. J. (2010) Cell Cycle 9, 2529–2536 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60. Weterings E., Verkaik N. S., Brüggenwirth H. T., Hoeijmakers J. H., van Gent D. C. (2003) Nucleic Acids Res. 31, 7238–7246 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61. Dobbs T. A., Tainer J. A., Lees-Miller S. P. (2010) DNA Repair 9, 1307–1314 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62. Gapud E. J., Sleckman B. P. (2011) Cell Cycle 10, 1928–1935 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 63. Nakai W., Westmoreland J., Yeh E., Bloom K., Resnick M. A. (2011) DNA Repair 10, 102–110 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 64. Lee K., Zhang Y., Lee S. E. (2008) Nature 454, 543–546 [DOI] [PubMed] [Google Scholar]
- 65. Yang Y. G., Saidi A., Frappart P. O., Min W., Barrucand C., Dumon-Jones V., Michelon J., Herceg Z., Wang Z. Q. (2006) EMBO J. 25, 5527–5538 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66. Fattah F., Lee E. H., Weisensel N., Wang Y., Lichter N., Hendrickson E. A. (2010) PLoS Genet. 6, e1000855. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 67. Franco S., Gostissa M., Zha S., Lombard D. B., Murphy M. M., Zarrin A. A., Yan C., Tepsuporn S., Morales J. C., Adams M. M., Lou Z., Bassing C. H., Manis J. P., Chen J., Carpenter P. B., Alt F. W. (2006) Mol. Cell 21, 201–214 [DOI] [PubMed] [Google Scholar]
- 68. Zha S., Sekiguchi J., Brush J. W., Bassing C. H., Alt F. W. (2008) Proc. Natl. Acad. Sci. U.S.A. 105, 9302–9306 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 69. Weinstock D. M., Elliott B., Jasin M. (2006) Blood 107, 777–780 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 70. Bassing C. H., Suh H., Ferguson D. O., Chua K. F., Manis J., Eckersdorff M., Gleason M., Bronson R., Lee C., Alt F. W. (2003) Cell 114, 359–370 [DOI] [PubMed] [Google Scholar]
- 71. Paulsen R. D., Soni D. V., Wollman R., Hahn A. T., Yee M. C., Guan A., Hesley J. A., Miller S. C., Cromwell E. F., Solow-Cordero D. E., Meyer T., Cimprich K. A. (2009) Mol. Cell 35, 228–239 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 72. Gan W., Guan Z., Liu J., Gui T., Shen K., Manley J. L., Li X. (2011) Genes Dev. 25, 2041–2056 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 73. Xu Y., Price B. D. (2011) Cell Cycle 10, 261–267 [DOI] [PMC free article] [PubMed] [Google Scholar]