Abstract
Thailandepsin A [systematic name: (E)-(1S,5S,6R,9S,20R)-6-[(2S)-butan-2-yl]-5-hydroxy-20-[2-(methylsulfanyl)ethyl]-2-oxa-11,12-dithia-7,19,22-triazabicyclo[7.7.6]docosa-15-ene-3,8,18,21-tetraone], C23H37N3O6S3, is a newly reported [Wang et al. (2011). J. Nat. Prod. doi:10.1021/np200324x] bicyclic depsipeptide that has potent histone deacetylase inhibitory activity and broad-spectrum antiproliferative activity. The absolute configuration of thailandepsin A has been determined from the anomalous dispersion and the stereochemistry of all chiral C atoms. Intramolecular N—H⋯O and N—H⋯S hydrogen bonds occur. Intermolecular N—H⋯O and O—H⋯O hydrogen bonds are observed in the crystal structure.
Related literature
For general background to histone deacetylase (HDAC) inhibitors as a new class of anticancer agents, see: FDA (2010 ▶); Furumai et al. (2002 ▶); Grant et al. (2010 ▶); Khan & La Thangue (2008 ▶); Mann et al. (2007 ▶); Ueda et al. (1994 ▶). For related structures, see: Shigematsu et al. (1994 ▶). For geometric data, see: Chou & Blinn (1997 ▶). For the biological activity of the title compound, see: Wang et al. (2011 ▶).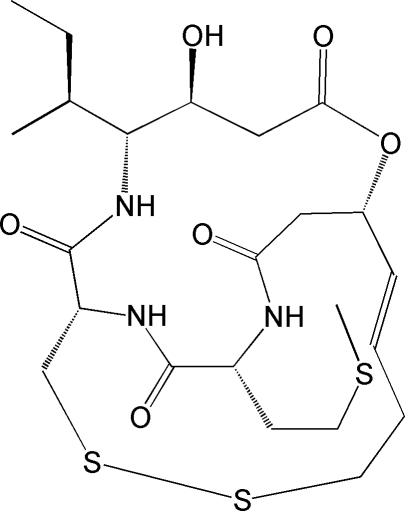
Experimental
Crystal data
C23H37N3O6S3
M r = 547.74
Orthorhombic,
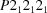
a = 12.7747 (3) Å
b = 13.2926 (3) Å
c = 15.4218 (4) Å
V = 2618.76 (11) Å3
Z = 4
Cu Kα radiation
μ = 2.96 mm−1
T = 100 K
0.45 × 0.42 × 0.38 mm
Data collection
Bruker SMART APEXII area-detector diffractometer
Absorption correction: multi-scan (SADABS; Bruker, 2007 ▶) T min = 0.352, T max = 0.403
34598 measured reflections
4990 independent reflections
4981 reflections with I > 2σ(I)
R int = 0.025
Refinement
R[F 2 > 2σ(F 2)] = 0.027
wR(F 2) = 0.071
S = 1.05
4990 reflections
326 parameters
4 restraints
H atoms treated by a mixture of independent and constrained refinement
Δρmax = 0.35 e Å−3
Δρmin = −0.36 e Å−3
Absolute structure: Flack (1983 ▶), 2102 Friedel pairs
Flack parameter: 0.000 (9)
Data collection: APEX2 (Bruker, 2007 ▶); cell refinement: SAINT (Bruker, 2007 ▶); data reduction: SAINT; program(s) used to solve structure: SHELXTL (Sheldrick, 2008 ▶); program(s) used to refine structure: SHELXTL; molecular graphics: SHELXTL and OLEX2 (Dolomanov et al., 2009 ▶); software used to prepare material for publication: SHELXTL.
Supplementary Material
Crystal structure: contains datablock(s) I, global. DOI: 10.1107/S1600536811041390/zl2411sup1.cif
Supplementary material file. DOI: 10.1107/S1600536811041390/zl2411Isup2.cdx
Structure factors: contains datablock(s) I. DOI: 10.1107/S1600536811041390/zl2411Isup3.hkl
Supplementary material file. DOI: 10.1107/S1600536811041390/zl2411Isup4.cml
Additional supplementary materials: crystallographic information; 3D view; checkCIF report
Table 1. Hydrogen-bond geometry (Å, °).
| D—H⋯A | D—H | H⋯A | D⋯A | D—H⋯A |
|---|---|---|---|---|
| O4—H4⋯O5i | 0.84 (1) | 1.90 (1) | 2.7394 (15) | 176 (2) |
| N1—H1⋯O6 | 0.88 (1) | 2.06 (1) | 2.9203 (17) | 166 (2) |
| N2—H2⋯S2 | 0.88 (1) | 2.83 (2) | 3.2491 (13) | 111 (2) |
| N3—H3⋯O4ii | 0.88 (1) | 2.33 (1) | 3.1534 (16) | 155 (2) |
Symmetry codes: (i) 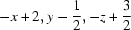 ; (ii)
; (ii) 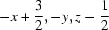 .
.
Acknowledgments
Support for this work was obtained from a Research Growth Initiative Award from the University of Wisconsin–Milwaukee and NIH/NCI grant R01 CA 152212 (both to Y-QC). The authors thank Lara C. Spencer and Ilia A. Guzei (University of Wisconsin-Madison Department of Chemistry Crystallography Facility) for collecting the crystallographic data.
supplementary crystallographic information
Comment
With the FDA approval of both SAHA (Vorinostat) and FK228 (Romidepsin) for the treatment of cutaneous T-cell lymphoma (FDA, 2010; Mann et al., 2007), histone deacetylase (HDAC) inhibitors have been in the spotlight in recent years as a new class of anticancer agents (Grant et al., 2010; Khan & La Thangue, 2008). FK228, a natural product produced by Chromobacterium violaceum No. 968 (Ueda et al., 1994), represents a family of natural products that contain a signature disulfide bond that is known or presumed to mediate a novel mode of anticancer action in which a reduced thiol group "warhead" chelates a Zn2+ in the catalytic center of Class I and Class II HDACs thereby inhibiting the enzyme activities (Furumai et al., 2002; Wang et al., 2011). The crystal structure of FK228 was reported in 1994 (Shigematsu et al., 1994).
Thailandepsin A is a natural analogue of FK228 newly discovered from Burkholderia thailandensis E264 by a genomics-guided approach; it has potent histone deacetylase inhibitory activities and broad-spectrum antiproliferative activities (Wang et al., 2011). The chemical structure of thailandepsin A was established by a combination of spectroscopic analyses, chemical derivatization and degradation. Here we report the crystal structure of thailandepsin A.
Thailandepsin A is a bicyclic depsipeptide and consists of four building blocks, D-cysteine (D-Cys), D-methionine (D-Met), 4-amino-3-hydroxy-5-methylheptanoic acid (Ahhp, derived from an isoleucine and an acetate unit) and 3-hydroxy-7-mercapto-4-heptenoic acid (Acyl, derived from a cysteine and two acetate units). The primary structure of thailandepsin A is D-Met-D-Cys-Ahhp-Acyl. X-ray crystallographic analysis indicates that the skeleton of thailandepsin A consists of a [7,7,6] 22-membered ring adopting an uncommon cage-shape that includes a 15-membered macrocyclic lactone and a 15-membered ring and a signature disulfide bond. The bridge ring is almost perpendicular to the main ring and the dihedral angle of these two least-squares planes is 77.7 (1)°. The side chains of methionine and isoleucine have less strain and can freely rotate on the single bonds. In order to obtain minimum energy positions, the alkyl groups arrange so on the molecular skeleton that they point away from each other.
The absolute configurations at C2, C8, C11 and C13 are S and the absolute configurations at C12 and 18 are R as established based on the results of anomalous dispersion. The geometric isomerism of the double bond in the Acyl component is determined as E. The backbone moiety from the carboxyl group of Acyl through methionine and cysteine to the amine group of Ahhp, (Acyl)-CO1—Met2—Cys3—NH4-(Ahhp), forms a peculiar secondary structure, a type I' β-turn, and the value of Ψ and Φ are 57.26 (17)°, 29.76 (18)°, 95.49 (16)° and -18.11 (19)° (Chou & Blinn, 1997). There are two intramolecular and two intermolecular hydrogen bonds present (Table 1, Fig. 1 and 2).
Experimental
Thailandepsin A was purified from the fermentation broth of B. thailandensis E264 as described earlier (Wang et al., 2011). Pure thailandepsin A was dissolved in methanol and block-like crystals were obtained after evaporation of the solvent at room temperature.
Refinement
All hydrogen atoms attached to the carbon atoms were placed in geometrically idealized positions (C—H = 0.98, 0.99 and 1.00 Å on the primary, secondary and tertiary aliphatic C atoms respectively, 0.95 Å on aromatic C). The H atoms were refined as riding, with isotropic displacement coefficients of Uiso(H) = 1.5Ueq(C) for methyl groups or 1.2Ueq(C) otherwise. The hydrogen atoms attached to N and O were located in difference maps and refined independently with restraints and constraints. The H atoms on N atoms were restrained to have N—H distances of 0.880 (1) Å and their Uiso values were constrained as equal to 1.2 times the Ueq of their carrier atoms. The H atom on O was restrained to have an O—H distance of 0.840 (1) Å and the Uiso value was assigned as equal to 1.5 times the Ueq of the oxygen atom.
Figures
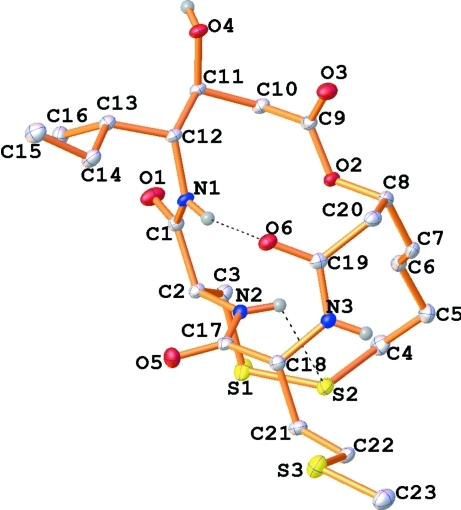
Fig. 1.
The molecular structure of thailandepsin A with displacement ellipsoids shown at the 50% probability level. For clarity, all H atoms attached to carbon atoms are omitted. Intramolecular hydrogen bonds are shown as dashed lines.
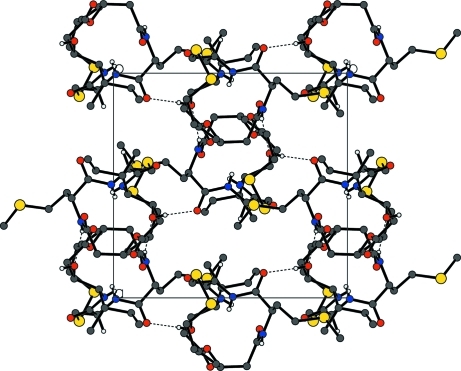
Fig. 2.
A packing diagram of thailandepsin A, viewed along the c axis. For clarity, all H atoms attached to carbon atoms are omitted. The dashed lines represent hydrogen bonds.
Crystal data
| C23H37N3O6S3 | F(000) = 1168 |
| Mr = 547.74 | Dx = 1.389 Mg m−3 |
| Orthorhombic, P212121 | Cu Kα radiation, λ = 1.54178 Å |
| Hall symbol: P 2ac 2ab | Cell parameters from 9793 reflections |
| a = 12.7747 (3) Å | θ = 3.5–71.2° |
| b = 13.2926 (3) Å | µ = 2.96 mm−1 |
| c = 15.4218 (4) Å | T = 100 K |
| V = 2618.76 (11) Å3 | Block, colourless |
| Z = 4 | 0.45 × 0.42 × 0.38 mm |
Data collection
| Bruker SMART APEXII area-detector diffractometer | 4990 independent reflections |
| Radiation source: fine-focus sealed tube | 4981 reflections with I > 2σ(I) |
| graphite | Rint = 0.025 |
| 0.50° ω and 0.5 ° φ scans | θmax = 71.7°, θmin = 4.4° |
| Absorption correction: multi-scan (SADABS; Bruker, 2007) | h = −15→14 |
| Tmin = 0.352, Tmax = 0.403 | k = −15→16 |
| 34598 measured reflections | l = −18→17 |
Refinement
| Refinement on F2 | Secondary atom site location: difference Fourier map |
| Least-squares matrix: full | Hydrogen site location: inferred from neighbouring sites |
| R[F2 > 2σ(F2)] = 0.027 | H atoms treated by a mixture of independent and constrained refinement |
| wR(F2) = 0.071 | w = 1/[σ2(Fo2) + (0.051P)2 + 0.7593P] where P = (Fo2 + 2Fc2)/3 |
| S = 1.05 | (Δ/σ)max = 0.001 |
| 4990 reflections | Δρmax = 0.35 e Å−3 |
| 326 parameters | Δρmin = −0.36 e Å−3 |
| 4 restraints | Absolute structure: Flack (1983), ???? Friedel pairs |
| Primary atom site location: structure-invariant direct methods | Flack parameter: 0.000 (9) |
Special details
| Geometry. All e.s.d.'s (except the e.s.d. in the dihedral angle between two l.s. planes) are estimated using the full covariance matrix. The cell e.s.d.'s are taken into account individually in the estimation of e.s.d.'s in distances, angles and torsion angles; correlations between e.s.d.'s in cell parameters are only used when they are defined by crystal symmetry. An approximate (isotropic) treatment of cell e.s.d.'s is used for estimating e.s.d.'s involving l.s. planes. |
| Refinement. Refinement of F2 against ALL reflections. The weighted R-factor wR and goodness of fit S are based on F2, conventional R-factors R are based on F, with F set to zero for negative F2. The threshold expression of F2 > σ(F2) is used only for calculating R-factors(gt) etc. and is not relevant to the choice of reflections for refinement. R-factors based on F2 are statistically about twice as large as those based on F, and R- factors based on ALL data will be even larger. |
Fractional atomic coordinates and isotropic or equivalent isotropic displacement parameters (Å2)
| x | y | z | Uiso*/Ueq | ||
| S1 | 1.10290 (3) | −0.14637 (3) | 0.38774 (2) | 0.02184 (10) | |
| S2 | 0.97356 (3) | −0.08216 (3) | 0.33328 (2) | 0.01709 (9) | |
| S3 | 0.91337 (3) | 0.40012 (3) | 0.40152 (3) | 0.02362 (10) | |
| O1 | 1.08885 (10) | −0.18335 (9) | 0.67591 (8) | 0.0223 (3) | |
| O2 | 0.77376 (8) | −0.09984 (8) | 0.61157 (7) | 0.0141 (2) | |
| O3 | 0.69383 (9) | −0.08629 (9) | 0.74184 (7) | 0.0190 (2) | |
| O4 | 0.85790 (9) | −0.16775 (8) | 0.88174 (7) | 0.0140 (2) | |
| H4 | 0.8699 (17) | −0.2254 (7) | 0.9016 (13) | 0.021* | |
| O5 | 1.11125 (9) | 0.14154 (8) | 0.55597 (7) | 0.0179 (2) | |
| O6 | 0.86120 (9) | 0.12479 (8) | 0.64747 (7) | 0.0158 (2) | |
| N1 | 0.99720 (10) | −0.04020 (10) | 0.70173 (8) | 0.0128 (3) | |
| H1 | 0.9652 (14) | 0.0126 (9) | 0.6798 (12) | 0.015* | |
| N2 | 1.00458 (10) | 0.00984 (9) | 0.52676 (8) | 0.0121 (2) | |
| H2 | 0.9444 (8) | −0.0082 (14) | 0.5038 (12) | 0.014* | |
| N3 | 0.83857 (10) | 0.13784 (10) | 0.50282 (8) | 0.0138 (3) | |
| H3 | 0.7955 (12) | 0.1335 (15) | 0.4584 (8) | 0.017* | |
| C1 | 1.05386 (12) | −0.10227 (12) | 0.65205 (10) | 0.0143 (3) | |
| C2 | 1.07489 (12) | −0.06840 (11) | 0.55727 (9) | 0.0131 (3) | |
| H2A | 1.1482 | −0.0423 | 0.5537 | 0.016* | |
| C3 | 1.06714 (13) | −0.16258 (12) | 0.50086 (10) | 0.0174 (3) | |
| H3A | 0.9943 | −0.1877 | 0.5034 | 0.021* | |
| H3B | 1.1128 | −0.2151 | 0.5262 | 0.021* | |
| C4 | 0.88346 (14) | −0.18723 (13) | 0.31638 (11) | 0.0202 (3) | |
| H4A | 0.8850 | −0.2082 | 0.2548 | 0.024* | |
| H4B | 0.9052 | −0.2453 | 0.3524 | 0.024* | |
| C5 | 0.77251 (12) | −0.15466 (13) | 0.34132 (11) | 0.0190 (3) | |
| H5A | 0.7542 | −0.0927 | 0.3091 | 0.023* | |
| H5B | 0.7225 | −0.2079 | 0.3240 | 0.023* | |
| C6 | 0.76184 (12) | −0.13526 (13) | 0.43705 (10) | 0.0173 (3) | |
| H6 | 0.7980 | −0.1793 | 0.4752 | 0.021* | |
| C7 | 0.70632 (12) | −0.06202 (12) | 0.47258 (10) | 0.0162 (3) | |
| H7 | 0.6720 | −0.0170 | 0.4341 | 0.019* | |
| C8 | 0.69224 (12) | −0.04303 (12) | 0.56827 (10) | 0.0154 (3) | |
| H8 | 0.6227 | −0.0704 | 0.5865 | 0.018* | |
| C9 | 0.76158 (12) | −0.12126 (11) | 0.69723 (10) | 0.0136 (3) | |
| C10 | 0.84280 (12) | −0.19799 (11) | 0.72498 (10) | 0.0138 (3) | |
| H10A | 0.8877 | −0.2133 | 0.6743 | 0.017* | |
| H10B | 0.8059 | −0.2608 | 0.7409 | 0.017* | |
| C11 | 0.91409 (12) | −0.16801 (11) | 0.80094 (9) | 0.0127 (3) | |
| H11 | 0.9724 | −0.2180 | 0.8051 | 0.015* | |
| C12 | 0.96190 (12) | −0.06200 (11) | 0.79096 (9) | 0.0125 (3) | |
| H12 | 0.9029 | −0.0143 | 0.8019 | 0.015* | |
| C13 | 1.04416 (12) | −0.03790 (11) | 0.86101 (10) | 0.0146 (3) | |
| H13 | 1.0124 | −0.0559 | 0.9182 | 0.017* | |
| C14 | 1.06664 (13) | 0.07539 (12) | 0.86315 (10) | 0.0184 (3) | |
| H14A | 1.1138 | 0.0927 | 0.8144 | 0.022* | |
| H14B | 1.0002 | 0.1125 | 0.8548 | 0.022* | |
| C15 | 1.11702 (14) | 0.10927 (13) | 0.94810 (11) | 0.0222 (3) | |
| H15A | 1.0677 | 0.0988 | 0.9960 | 0.033* | |
| H15B | 1.1350 | 0.1808 | 0.9443 | 0.033* | |
| H15C | 1.1807 | 0.0700 | 0.9585 | 0.033* | |
| C16 | 1.14602 (12) | −0.09806 (13) | 0.85283 (10) | 0.0184 (3) | |
| H16A | 1.1848 | −0.0752 | 0.8017 | 0.028* | |
| H16B | 1.1296 | −0.1697 | 0.8467 | 0.028* | |
| H16C | 1.1888 | −0.0878 | 0.9048 | 0.028* | |
| C17 | 1.02739 (12) | 0.10824 (11) | 0.52881 (9) | 0.0134 (3) | |
| C18 | 0.94399 (12) | 0.17890 (11) | 0.49209 (10) | 0.0138 (3) | |
| H18 | 0.9480 | 0.2442 | 0.5241 | 0.017* | |
| C19 | 0.80478 (12) | 0.11430 (11) | 0.58346 (10) | 0.0141 (3) | |
| C20 | 0.69604 (12) | 0.06994 (12) | 0.59060 (10) | 0.0160 (3) | |
| H20A | 0.6486 | 0.1068 | 0.5510 | 0.019* | |
| H20B | 0.6701 | 0.0797 | 0.6505 | 0.019* | |
| C21 | 0.96752 (13) | 0.19916 (11) | 0.39575 (10) | 0.0158 (3) | |
| H21A | 1.0409 | 0.2224 | 0.3904 | 0.019* | |
| H21B | 0.9613 | 0.1351 | 0.3634 | 0.019* | |
| C22 | 0.89580 (13) | 0.27705 (12) | 0.35353 (10) | 0.0168 (3) | |
| H22A | 0.8220 | 0.2558 | 0.3605 | 0.020* | |
| H22B | 0.9112 | 0.2807 | 0.2907 | 0.020* | |
| C23 | 0.82077 (15) | 0.46899 (14) | 0.33643 (12) | 0.0250 (4) | |
| H23A | 0.8190 | 0.5393 | 0.3554 | 0.037* | |
| H23B | 0.7510 | 0.4392 | 0.3431 | 0.037* | |
| H23C | 0.8419 | 0.4659 | 0.2754 | 0.037* |
Atomic displacement parameters (Å2)
| U11 | U22 | U33 | U12 | U13 | U23 | |
| S1 | 0.01829 (19) | 0.0340 (2) | 0.01321 (18) | 0.00771 (17) | −0.00022 (14) | −0.00696 (15) |
| S2 | 0.01714 (18) | 0.02111 (19) | 0.01302 (16) | 0.00009 (15) | −0.00063 (14) | −0.00078 (14) |
| S3 | 0.0279 (2) | 0.0207 (2) | 0.0222 (2) | 0.00109 (16) | −0.00648 (16) | 0.00032 (16) |
| O1 | 0.0286 (6) | 0.0210 (6) | 0.0172 (6) | 0.0118 (5) | 0.0048 (5) | 0.0045 (5) |
| O2 | 0.0140 (5) | 0.0171 (5) | 0.0112 (5) | 0.0017 (4) | 0.0001 (4) | 0.0034 (4) |
| O3 | 0.0199 (5) | 0.0216 (6) | 0.0156 (5) | 0.0040 (5) | 0.0037 (4) | 0.0017 (5) |
| O4 | 0.0190 (5) | 0.0131 (5) | 0.0098 (5) | 0.0019 (4) | 0.0019 (4) | 0.0018 (4) |
| O5 | 0.0167 (5) | 0.0166 (5) | 0.0205 (5) | −0.0020 (4) | −0.0032 (5) | −0.0018 (5) |
| O6 | 0.0197 (5) | 0.0159 (5) | 0.0119 (5) | 0.0022 (4) | 0.0004 (4) | −0.0001 (4) |
| N1 | 0.0161 (6) | 0.0129 (6) | 0.0093 (6) | 0.0026 (5) | −0.0002 (5) | 0.0029 (5) |
| N2 | 0.0122 (6) | 0.0137 (6) | 0.0103 (6) | −0.0004 (5) | −0.0003 (4) | −0.0003 (5) |
| N3 | 0.0139 (6) | 0.0155 (6) | 0.0121 (6) | 0.0005 (5) | −0.0008 (5) | 0.0010 (5) |
| C1 | 0.0145 (7) | 0.0156 (7) | 0.0129 (7) | 0.0005 (6) | 0.0002 (5) | 0.0013 (6) |
| C2 | 0.0146 (7) | 0.0142 (7) | 0.0104 (6) | 0.0017 (6) | 0.0001 (5) | −0.0004 (6) |
| C3 | 0.0227 (7) | 0.0153 (7) | 0.0142 (7) | 0.0037 (6) | −0.0015 (6) | −0.0014 (6) |
| C4 | 0.0243 (8) | 0.0191 (7) | 0.0170 (8) | −0.0014 (7) | −0.0020 (7) | −0.0043 (6) |
| C5 | 0.0184 (8) | 0.0213 (8) | 0.0172 (8) | −0.0030 (6) | −0.0031 (6) | −0.0015 (6) |
| C6 | 0.0167 (7) | 0.0196 (7) | 0.0156 (7) | −0.0045 (6) | −0.0035 (6) | 0.0037 (6) |
| C7 | 0.0150 (7) | 0.0194 (7) | 0.0143 (7) | −0.0019 (6) | −0.0042 (6) | 0.0040 (6) |
| C8 | 0.0126 (7) | 0.0182 (7) | 0.0154 (7) | 0.0001 (6) | −0.0011 (6) | 0.0038 (6) |
| C9 | 0.0165 (7) | 0.0135 (7) | 0.0107 (6) | −0.0032 (6) | −0.0007 (6) | 0.0002 (5) |
| C10 | 0.0183 (7) | 0.0118 (7) | 0.0111 (7) | 0.0006 (6) | −0.0002 (6) | −0.0001 (5) |
| C11 | 0.0157 (7) | 0.0136 (7) | 0.0087 (6) | 0.0015 (6) | 0.0001 (6) | 0.0010 (5) |
| C12 | 0.0152 (7) | 0.0133 (6) | 0.0089 (6) | 0.0014 (6) | 0.0010 (6) | 0.0009 (5) |
| C13 | 0.0176 (7) | 0.0156 (7) | 0.0106 (7) | 0.0009 (6) | −0.0008 (6) | 0.0000 (5) |
| C14 | 0.0249 (8) | 0.0155 (7) | 0.0148 (7) | −0.0014 (6) | −0.0034 (6) | 0.0017 (6) |
| C15 | 0.0287 (9) | 0.0189 (8) | 0.0190 (8) | −0.0025 (7) | −0.0021 (7) | −0.0013 (6) |
| C16 | 0.0161 (7) | 0.0208 (8) | 0.0182 (8) | 0.0011 (6) | −0.0020 (6) | 0.0011 (6) |
| C17 | 0.0164 (7) | 0.0158 (7) | 0.0079 (6) | −0.0003 (6) | 0.0022 (6) | −0.0004 (5) |
| C18 | 0.0153 (7) | 0.0133 (7) | 0.0127 (7) | −0.0014 (6) | 0.0000 (6) | 0.0000 (6) |
| C19 | 0.0160 (7) | 0.0119 (7) | 0.0144 (7) | 0.0037 (6) | 0.0017 (6) | 0.0005 (6) |
| C20 | 0.0154 (7) | 0.0184 (7) | 0.0142 (7) | 0.0036 (6) | 0.0020 (6) | 0.0026 (6) |
| C21 | 0.0192 (7) | 0.0154 (7) | 0.0128 (7) | −0.0013 (6) | 0.0012 (6) | 0.0013 (6) |
| C22 | 0.0210 (8) | 0.0171 (7) | 0.0122 (7) | −0.0029 (6) | −0.0026 (6) | 0.0015 (5) |
| C23 | 0.0259 (8) | 0.0281 (9) | 0.0209 (8) | 0.0081 (7) | 0.0006 (7) | 0.0028 (7) |
Geometric parameters (Å, °)
| S1—C3 | 1.8162 (17) | C8—C20 | 1.541 (2) |
| S1—S2 | 2.0406 (6) | C8—H8 | 1.0000 |
| S2—C4 | 1.8285 (17) | C9—C10 | 1.517 (2) |
| S3—C23 | 1.8014 (18) | C10—C11 | 1.536 (2) |
| S3—C22 | 1.8095 (16) | C10—H10A | 0.9900 |
| O1—C1 | 1.223 (2) | C10—H10B | 0.9900 |
| O2—C9 | 1.3602 (18) | C11—C12 | 1.543 (2) |
| O2—C8 | 1.4494 (18) | C11—H11 | 1.0000 |
| O3—C9 | 1.199 (2) | C12—C13 | 1.541 (2) |
| O4—C11 | 1.4380 (17) | C12—H12 | 1.0000 |
| O4—H4 | 0.8399 (10) | C13—C16 | 1.533 (2) |
| O5—C17 | 1.233 (2) | C13—C14 | 1.533 (2) |
| O6—C19 | 1.230 (2) | C13—H13 | 1.0000 |
| N1—C1 | 1.338 (2) | C14—C15 | 1.528 (2) |
| N1—C12 | 1.4769 (18) | C14—H14A | 0.9900 |
| N1—H1 | 0.8797 (10) | C14—H14B | 0.9900 |
| N2—C17 | 1.340 (2) | C15—H15A | 0.9800 |
| N2—C2 | 1.4524 (19) | C15—H15B | 0.9800 |
| N2—H2 | 0.8798 (10) | C15—H15C | 0.9800 |
| N3—C19 | 1.353 (2) | C16—H16A | 0.9800 |
| N3—C18 | 1.4624 (18) | C16—H16B | 0.9800 |
| N3—H3 | 0.8799 (10) | C16—H16C | 0.9800 |
| C1—C2 | 1.553 (2) | C17—C18 | 1.529 (2) |
| C2—C3 | 1.528 (2) | C18—C21 | 1.540 (2) |
| C2—H2A | 1.0000 | C18—H18 | 1.0000 |
| C3—H3A | 0.9900 | C19—C20 | 1.513 (2) |
| C3—H3B | 0.9900 | C20—H20A | 0.9900 |
| C4—C5 | 1.531 (2) | C20—H20B | 0.9900 |
| C4—H4A | 0.9900 | C21—C22 | 1.528 (2) |
| C4—H4B | 0.9900 | C21—H21A | 0.9900 |
| C5—C6 | 1.505 (2) | C21—H21B | 0.9900 |
| C5—H5A | 0.9900 | C22—H22A | 0.9900 |
| C5—H5B | 0.9900 | C22—H22B | 0.9900 |
| C6—C7 | 1.323 (2) | C23—H23A | 0.9800 |
| C6—H6 | 0.9500 | C23—H23B | 0.9800 |
| C7—C8 | 1.508 (2) | C23—H23C | 0.9800 |
| C7—H7 | 0.9500 | ||
| C3—S1—S2 | 103.96 (6) | C12—C11—H11 | 108.5 |
| C4—S2—S1 | 104.41 (6) | N1—C12—C13 | 113.84 (12) |
| C23—S3—C22 | 98.63 (8) | N1—C12—C11 | 113.14 (12) |
| C9—O2—C8 | 118.31 (12) | C13—C12—C11 | 112.95 (12) |
| C11—O4—H4 | 102.9 (15) | N1—C12—H12 | 105.3 |
| C1—N1—C12 | 125.27 (13) | C13—C12—H12 | 105.3 |
| C1—N1—H1 | 121.5 (13) | C11—C12—H12 | 105.3 |
| C12—N1—H1 | 111.9 (13) | C16—C13—C14 | 110.81 (13) |
| C17—N2—C2 | 123.82 (13) | C16—C13—C12 | 114.39 (13) |
| C17—N2—H2 | 117.7 (13) | C14—C13—C12 | 110.31 (12) |
| C2—N2—H2 | 118.4 (13) | C16—C13—H13 | 107.0 |
| C19—N3—C18 | 118.95 (13) | C14—C13—H13 | 107.0 |
| C19—N3—H3 | 120.1 (13) | C12—C13—H13 | 107.0 |
| C18—N3—H3 | 120.8 (13) | C15—C14—C13 | 112.76 (13) |
| O1—C1—N1 | 124.65 (14) | C15—C14—H14A | 109.0 |
| O1—C1—C2 | 118.38 (14) | C13—C14—H14A | 109.0 |
| N1—C1—C2 | 116.97 (13) | C15—C14—H14B | 109.0 |
| N2—C2—C3 | 111.24 (12) | C13—C14—H14B | 109.0 |
| N2—C2—C1 | 113.92 (12) | H14A—C14—H14B | 107.8 |
| C3—C2—C1 | 106.69 (12) | C14—C15—H15A | 109.5 |
| N2—C2—H2A | 108.3 | C14—C15—H15B | 109.5 |
| C3—C2—H2A | 108.3 | H15A—C15—H15B | 109.5 |
| C1—C2—H2A | 108.3 | C14—C15—H15C | 109.5 |
| C2—C3—S1 | 115.69 (11) | H15A—C15—H15C | 109.5 |
| C2—C3—H3A | 108.4 | H15B—C15—H15C | 109.5 |
| S1—C3—H3A | 108.4 | C13—C16—H16A | 109.5 |
| C2—C3—H3B | 108.4 | C13—C16—H16B | 109.5 |
| S1—C3—H3B | 108.4 | H16A—C16—H16B | 109.5 |
| H3A—C3—H3B | 107.4 | C13—C16—H16C | 109.5 |
| C5—C4—S2 | 109.33 (11) | H16A—C16—H16C | 109.5 |
| C5—C4—H4A | 109.8 | H16B—C16—H16C | 109.5 |
| S2—C4—H4A | 109.8 | O5—C17—N2 | 123.19 (14) |
| C5—C4—H4B | 109.8 | O5—C17—C18 | 120.72 (13) |
| S2—C4—H4B | 109.8 | N2—C17—C18 | 116.05 (13) |
| H4A—C4—H4B | 108.3 | N3—C18—C17 | 111.75 (12) |
| C6—C5—C4 | 112.26 (13) | N3—C18—C21 | 110.75 (12) |
| C6—C5—H5A | 109.2 | C17—C18—C21 | 109.20 (12) |
| C4—C5—H5A | 109.2 | N3—C18—H18 | 108.4 |
| C6—C5—H5B | 109.2 | C17—C18—H18 | 108.4 |
| C4—C5—H5B | 109.2 | C21—C18—H18 | 108.4 |
| H5A—C5—H5B | 107.9 | O6—C19—N3 | 121.62 (14) |
| C7—C6—C5 | 125.51 (15) | O6—C19—C20 | 121.58 (14) |
| C7—C6—H6 | 117.2 | N3—C19—C20 | 116.73 (14) |
| C5—C6—H6 | 117.2 | C19—C20—C8 | 113.10 (13) |
| C6—C7—C8 | 126.32 (14) | C19—C20—H20A | 109.0 |
| C6—C7—H7 | 116.8 | C8—C20—H20A | 109.0 |
| C8—C7—H7 | 116.8 | C19—C20—H20B | 109.0 |
| O2—C8—C7 | 106.14 (12) | C8—C20—H20B | 109.0 |
| O2—C8—C20 | 112.45 (12) | H20A—C20—H20B | 107.8 |
| C7—C8—C20 | 112.21 (13) | C22—C21—C18 | 114.37 (13) |
| O2—C8—H8 | 108.6 | C22—C21—H21A | 108.7 |
| C7—C8—H8 | 108.6 | C18—C21—H21A | 108.7 |
| C20—C8—H8 | 108.6 | C22—C21—H21B | 108.7 |
| O3—C9—O2 | 123.95 (14) | C18—C21—H21B | 108.7 |
| O3—C9—C10 | 126.34 (14) | H21A—C21—H21B | 107.6 |
| O2—C9—C10 | 109.66 (12) | C21—C22—S3 | 111.34 (11) |
| C9—C10—C11 | 116.50 (12) | C21—C22—H22A | 109.4 |
| C9—C10—H10A | 108.2 | S3—C22—H22A | 109.4 |
| C11—C10—H10A | 108.2 | C21—C22—H22B | 109.4 |
| C9—C10—H10B | 108.2 | S3—C22—H22B | 109.4 |
| C11—C10—H10B | 108.2 | H22A—C22—H22B | 108.0 |
| H10A—C10—H10B | 107.3 | S3—C23—H23A | 109.5 |
| O4—C11—C10 | 111.44 (12) | S3—C23—H23B | 109.5 |
| O4—C11—C12 | 106.37 (11) | H23A—C23—H23B | 109.5 |
| C10—C11—C12 | 113.29 (12) | S3—C23—H23C | 109.5 |
| O4—C11—H11 | 108.5 | H23A—C23—H23C | 109.5 |
| C10—C11—H11 | 108.5 | H23B—C23—H23C | 109.5 |
| C3—S1—S2—C4 | −79.45 (8) | O4—C11—C12—N1 | −164.12 (12) |
| C12—N1—C1—O1 | −4.2 (2) | C10—C11—C12—N1 | −41.37 (17) |
| C12—N1—C1—C2 | 175.04 (13) | O4—C11—C12—C13 | 64.71 (15) |
| C17—N2—C2—C3 | −143.88 (14) | C10—C11—C12—C13 | −172.53 (12) |
| C17—N2—C2—C1 | 95.49 (16) | N1—C12—C13—C16 | −61.27 (17) |
| O1—C1—C2—N2 | 161.20 (14) | C11—C12—C13—C16 | 69.54 (16) |
| N1—C1—C2—N2 | −18.11 (19) | N1—C12—C13—C14 | 64.44 (16) |
| O1—C1—C2—C3 | 38.06 (19) | C11—C12—C13—C14 | −164.75 (12) |
| N1—C1—C2—C3 | −141.25 (14) | C16—C13—C14—C15 | −71.70 (17) |
| N2—C2—C3—S1 | 61.75 (15) | C12—C13—C14—C15 | 160.60 (13) |
| C1—C2—C3—S1 | −173.45 (10) | C2—N2—C17—O5 | 0.9 (2) |
| S2—S1—C3—C2 | −79.42 (12) | C2—N2—C17—C18 | 178.86 (12) |
| S1—S2—C4—C5 | 138.27 (10) | C19—N3—C18—C17 | 57.26 (17) |
| S2—C4—C5—C6 | −67.15 (16) | C19—N3—C18—C21 | 179.24 (13) |
| C4—C5—C6—C7 | 141.33 (16) | O5—C17—C18—N3 | −152.25 (13) |
| C5—C6—C7—C8 | 178.20 (15) | N2—C17—C18—N3 | 29.76 (18) |
| C9—O2—C8—C7 | −160.50 (12) | O5—C17—C18—C21 | 84.87 (16) |
| C9—O2—C8—C20 | 76.45 (16) | N2—C17—C18—C21 | −93.11 (15) |
| C6—C7—C8—O2 | 16.4 (2) | C18—N3—C19—O6 | −1.4 (2) |
| C6—C7—C8—C20 | 139.65 (16) | C18—N3—C19—C20 | −178.68 (12) |
| C8—O2—C9—O3 | −9.1 (2) | O6—C19—C20—C8 | −97.53 (17) |
| C8—O2—C9—C10 | 168.50 (12) | N3—C19—C20—C8 | 79.73 (16) |
| O3—C9—C10—C11 | −58.7 (2) | O2—C8—C20—C19 | 47.62 (17) |
| O2—C9—C10—C11 | 123.75 (13) | C7—C8—C20—C19 | −71.95 (17) |
| C9—C10—C11—O4 | 71.69 (16) | N3—C18—C21—C22 | 62.21 (16) |
| C9—C10—C11—C12 | −48.21 (17) | C17—C18—C21—C22 | −174.33 (13) |
| C1—N1—C12—C13 | 84.41 (18) | C18—C21—C22—S3 | 64.71 (15) |
| C1—N1—C12—C11 | −46.31 (19) | C23—S3—C22—C21 | 179.27 (12) |
Hydrogen-bond geometry (Å, °)
| D—H···A | D—H | H···A | D···A | D—H···A |
| O4—H4···O5i | 0.84 (1) | 1.90 (1) | 2.7394 (15) | 176 (2) |
| N1—H1···O6 | 0.88 (1) | 2.06 (1) | 2.9203 (17) | 166.(2) |
| N2—H2···S2 | 0.88 (1) | 2.83 (2) | 3.2491 (13) | 111.(2) |
| N3—H3···O4ii | 0.88 (1) | 2.33 (1) | 3.1534 (16) | 155.(2) |
Symmetry codes: (i) −x+2, y−1/2, −z+3/2; (ii) −x+3/2, −y, z−1/2.
Footnotes
Supplementary data and figures for this paper are available from the IUCr electronic archives (Reference: ZL2411).
References
- Bruker (2007). APEX2, SADABS and SAINT Bruker AXS Inc., Madison, Wisconsin, USA.
- Chou, K. C. & Blinn, J. R. (1997). J. Protein Chem. 16, 575–595. [DOI] [PubMed]
- Dolomanov, O. V., Bourhis, L. J., Gildea, R. J., Howard, J. A. K. & Puschmann, H. (2009). J. Appl. Cryst. 42, 339–341.
- FDA (2010). J. Natl Cancer Inst. 102, 219.
- Flack, H. D. (1983). Acta Cryst. A39, 876–881.
- Furumai, R., Matsuyama, A., Kobashi, N., Lee, K. H., Nishiyama, M., Nakajima, H., Tanaka, A., Komatsu, Y., Nishino, N., Yoshida, M. & Horinouchi, S. (2002). Cancer Res. 62, 4916–4921. [PubMed]
- Grant, C., Rahman, F., Piekarz, R., Peer, C., Frye, R., Robey, R. W., Gardner, E. R., Figg, W. D. & Bates, S. E. (2010). Expert Rev. Anticancer Ther. 10, 997–1008. [DOI] [PMC free article] [PubMed]
- Khan, O. & La Thangue, N. B. (2008). Nat. Clin. Pract. Oncol. 5, 714–726. [DOI] [PubMed]
- Mann, B. S., Johnson, J. R., Cohen, M. H., Justice, R. & Pazdur, R. (2007). The Oncologist, 12, 1247–1252. [DOI] [PubMed]
- Sheldrick, G. M. (2008). Acta Cryst. A64, 112–122. [DOI] [PubMed]
- Shigematsu, N., Ueda, H., Takase, S., Tanaka, H., Yamamoto, K. & Tada, T. (1994). J. Antibiot. (Tokyo), 47, 311–314. [DOI] [PubMed]
- Ueda, H., Nakajima, H., Hori, Y., Fujita, T., Nishimura, M., Goto, T. & Okuhara, M. (1994). J. Antibiot. (Tokyo), 47, 301–310. [DOI] [PubMed]
- Wang, C., Henkes, L. M., Doughty, L. B., He, M., Wang, D., Meyer-Almes, F. J. & Cheng, Y. Q. (2011). J. Nat. Prod. doi:10.1021/np200324x. [DOI] [PMC free article] [PubMed]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Crystal structure: contains datablock(s) I, global. DOI: 10.1107/S1600536811041390/zl2411sup1.cif
Supplementary material file. DOI: 10.1107/S1600536811041390/zl2411Isup2.cdx
Structure factors: contains datablock(s) I. DOI: 10.1107/S1600536811041390/zl2411Isup3.hkl
Supplementary material file. DOI: 10.1107/S1600536811041390/zl2411Isup4.cml
Additional supplementary materials: crystallographic information; 3D view; checkCIF report