Abstract
Classic therapeutics for ischemic heart disease are less effective in individuals with the metabolic syndrome. As the prevalence of the metabolic syndrome is increasing, better understanding of cardiac metabolism is needed to identify potential new targets for therapeutic intervention. Thioredoxin-interacting protein (Txnip) is a regulator of metabolism and an inhibitor of the antioxidant thioredoxins, but little is known about its roles in the myocardium. We examined hearts from Txnip-KO mice by polony multiplex analysis of gene expression and an independent proteomic approach; both methods indicated suppression of genes and proteins participating in mitochondrial metabolism. Consistently, Txnip-KO mitochondria were functionally and structurally altered, showing reduced oxygen consumption and ultrastructural derangements. Given the central role that mitochondria play during hypoxia, we hypothesized that Txnip deletion would enhance ischemia-reperfusion damage. Surprisingly, Txnip-KO hearts had greater recovery of cardiac function after an ischemia-reperfusion insult. Similarly, cardiomyocyte-specific Txnip deletion reduced infarct size after reversible coronary ligation. Coordinated with reduced mitochondrial function, deletion of Txnip enhanced anaerobic glycolysis. Whereas mitochondrial ATP synthesis was minimally decreased by Txnip deletion, cellular ATP content and lactate formation were higher in Txnip-KO hearts after ischemia-reperfusion injury. Pharmacologic inhibition of glycolytic metabolism completely abolished the protection afforded the heart by Txnip deficiency under hypoxic conditions. Thus, although Txnip deletion suppresses mitochondrial function, protection from myocardial ischemia is enhanced as a result of a coordinated shift to enhanced anaerobic metabolism, which provides an energy source outside of mitochondria.
Introduction
The clinical profile of ischemic heart disease has changed profoundly over the last decade, with a growing prevalence of systemic metabolic disorders. As the metabolic syndrome can decrease the effectiveness of classic therapeutics for ischemic heart disease (1), a better understanding of cardiac metabolism will be increasingly important for the treatment of the ischemic heart. Cellular metabolic regulation is largely dependent on mitochondrial respiration, which plays an important role in energy homeostasis by metabolizing nutrients and producing ATP and heat (2). ROS represent evanescent signaling molecules that are byproducts of the electron flux through mitochondrial oxidative phosphorylation (OXPHOS). The continuation of mitochondrial aerobic respiration in the absence of oxygen results in excessive ROS production, which can lead to cellular damage (3). The link between mitochondria-derived ROS and tissue damage is well established in ischemia-reperfusion injury of the myocardium (4).
An emerging link between metabolism regulation and cellular redox balance is the ubiquitous antioxidant thioredoxin and its endogenous inhibitor, thioredoxin-interacting protein (Txnip). Thioredoxins are critical intracellular antioxidants, operating primarily by reducing oxidized protein thiols (5). Cytosolic/nuclear thioredoxin-1 (Trx1) and mitochondrial Trx2 are 2 major isoforms of thioredoxin encoded by 2 distinct genes; both function as NADPH-dependent protein disulfide oxidoreductases. Txnip, a member of the α-arrestin protein superfamily (6), is a ubiquitously expressed protein found in the nucleus, mitochondria, and cytoplasm of cells from normal tissues (7). Txnip binds to both thioredoxin isoforms (7, 8); it both inhibits thioredoxin’s ability to scavenge ROS and interferes with its binding to other signaling molecules (8, 9). Although it is unclear whether the function of Txnip is mediated solely by inhibition of thioredoxin, recent evidence has established Txnip as a potent metabolic control protein (10, 11). Txnip expression is highly glucose responsive as a result of a carbohydrate response element in its promoter (12, 13) and its association with transcription factors max like protein X (Mlx) and MondoA (14). Txnip expression is elevated in skeletal muscle of humans with type 2 diabetes; furthermore, TXNIP overexpression blunts adipocyte glucose uptake, whereas TXNIP deletion augments glucose uptake in skeletal muscle and adipocytes (15, 16).
We previously reported that Txnip-KO mice exhibit a robust increase in myocardial glucose uptake and glycogen storage (17). To gain more insight into Txnip’s physiological role in the myocardium, we performed nonbiased global expression analyses using both a highly sensitive mRNA profiling technology and a proteomic approach. These comprehensive profiles and subsequent functional analyses revealed a central role played by the mitochondria in Txnip’s function as a regulator of cardiac metabolism. Despite clear evidence that Txnip deletion results in impaired mitochondrial respiration, Txnip-null mouse hearts were potently protected from ischemic injury, revealing the importance for nonmitochondrial metabolism in ischemic protection.
Results
Comprehensive gene profiles identify transcriptional changes associated with energy metabolism in Txnip-KO hearts.
Multiple lines of evidence support the concept that Txnip is a critical regulator of cellular and global metabolism, but little is known about how Txnip operates in the heart. To gain an unbiased perspective into Txnip’s function, we characterized its effects on global myocardial transcriptional regulation using the recently described technique of polony multiplex analysis of gene expression (PMAGE). PMAGE provides large-scale transcript identification with great sensitivity for quantification of rare transcripts (18). We constructed 2 PMAGE libraries from cardiac tissues of 8-week-old male WT (n = 4) and Txnip-KO (n = 5) mice. Tag counts were normalized for 2 million tags per library and filtered for nonredundant transcripts in the sense orientation. A significance threshold of P < 0.0023 was set to identify differentially expressed genes, which reflected a P value cutoff for a false discovery rate of 0.05 by Benjamini Hochberg (19). Among 17,105 total genes surveyed, 786 were identified as differentially expressed in the Txnip-KO LV compared with that of littermate WT controls (Supplemental Table 1; supplemental material available online with this article; doi: 10.1172/JCI44927DS1). When a fold change greater than +1.5 or less than –1.5 was used as a cutoff value, 390 filtered genes met this cutoff, and the vast majority were downregulated (306 genes). Ontology analysis based on molecular function demonstrated that the widespread changes in the differentially regulated genes encoded enzymatic proteins, including dehydrogenases and oxidoreductases (Figure 1A). Ontological pathway analysis revealed that the altered gene expression primarily affected components of energy metabolism networks, specifically, those related to electron transport chain, OXPHOS, or fatty acid/carbohydrate metabolism (Figure 1B). Network reconstruction by Ingenuity pathway analysis revealed that the networks contain genes such as retinoid X receptor α, peroxisome proliferator–activated receptor γ coactivator 1-β, and other signaling molecules (Supplemental Figure 1).
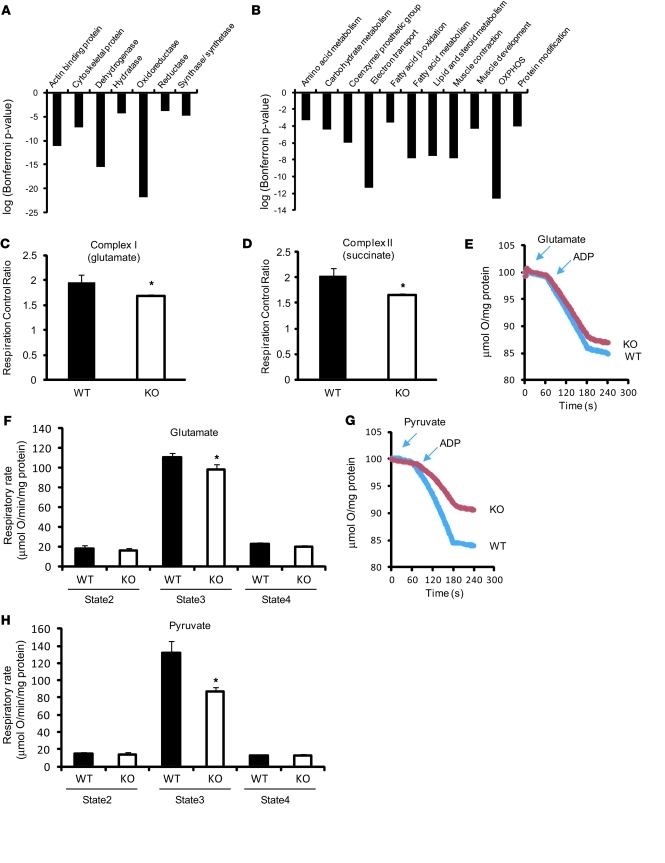
Figure 1. Deletion of Txnip in the myocardium results in mitochondrial dysfunction with decreased expression of transcripts encoding mitochondrial metabolism.
(A and B) PMAGE demonstrated transcriptional changes in mitochondrial metabolism–related pathways in Txnip-KO hearts. Gene ontology analysis was performed using the software PANTHER that classified downregulated genes by their molecular functions (A) and biological processes (B) based on published evidence. Biological process (B) covers the biological systems to which a protein contributes. Each group refers to a group of proteins with common function or process. (C and D) Respiration control ratio was obtained from permeabilized cardiac fibers incubated with indicated substrates in 11 WT and 7 Txnip-KO mice. (E–H) Representative tracing of oxygen consumption in isolated mitochondria (E and G). Mitochondria (200 μg) were incubated with 5 mM glutamate and 2 mM malate (E and F; n = 14 [WT] 13 [Txnip-KO]) or with 10 mM pyruvate and 2 mM malate (G and H; n = 4 each). State 2 and 3 respirations were measured by a Clark-type electrode in the presence of respiratory substrates before and after the addition of 1 mM ADP. State 4 respiration was measured on depletion of ADP. Values are mean ± SEM. *P < 0.05 versus WT.
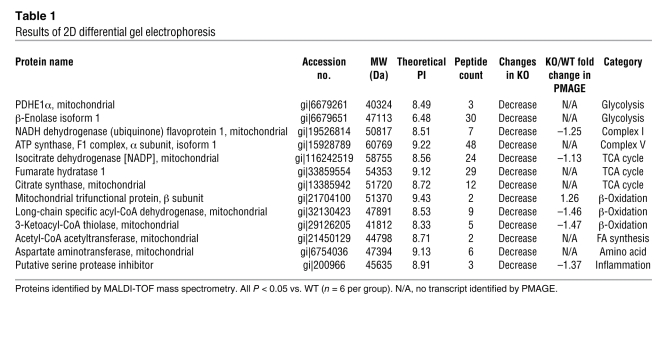
In a parallel independent study, we performed a proteomic analysis by 2-dimensional (2D) gel electrophoresis using Txnip-KO hearts compared with WT controls. We identified 21 protein spots as being differentially expressed, with each showing decreased expression in the Txnip-KO cohort relative to WT (all P < 0.05). From these spots, 13 proteins were identified by mass spectrometry (Table 1). Expression of 5 of these 13 proteins correlated well with the levels of the corresponding mRNA, according to PMAGE expression data. However, there were 8 proteins whose abundance was significantly affected by Txnip, without changes of transcription of the corresponding genes (based on the gene expression profile). This result was not unexpected, given the intrinsic technical differences between the 2 experimental methods. It has previously been shown that there are differences between parallel profiles of transcripts and proteins as a result of differential splicing, posttranslational modifications, and data integration (20, 21). Similar to the PMAGE results, 12 of the 13 proteins identified were associated with mitochondrial metabolism, including NADH dehydrogenate flavoprotein 1 (complex I), ATP synthase, F1 complex, α subunit, isoform 1 (complex V), and pyruvate dehydrogenase, E1 component, subunit α (PDHE1α). Thus, in the absence of a preformulated hypothesis, both mRNA and proteomic profiles identified that loss of Txnip in the myocardium resulted in decreased levels of multiple molecules participating in several central metabolic pathways, with a primary emphasis on perturbed mitochondrial metabolism.
Table 1 .
Results of 2D differential gel electrophoresis
Targeted deletion of Txnip decreases mitochondrial function.
As the PMAGE and proteomic data suggested that Txnip deletion could result in impaired mitochondrial function in the heart, we directly tested this hypothesis by assaying substrate-driven oxygen consumption in saponin-skinned cardiac fibers. Polarography was performed using precursor substrates specific to entry points of the tricarboxylic acid cycle or the individual complexes of electron transport flux as follows: glutamate/malate for complex I, succinate/rotenone for complex II, and pyruvate/malate for the carbohydrate-dependent flux. Glutamate/malate and succinate/rotenone state 3 (ADP-dependent) respiration, but not state 4 (ADP-independent) respiration, was reduced in Txnip-KO compared with WT cardiac fibers (Figure 1, C and D), reflecting a mild but significantly repressed respiration control ratio (i.e., state 3/state 4). This indicated that mitochondrial function was reduced distal to complex I and II in the Txnip-KO heart, but was not attributable to excessive mitochondrial uncoupling. To confirm the decreased respiratory capacity in Txnip-KO skinned fibers, we isolated purified mitochondria from mouse hearts. Consistent with the skinned fiber studies, isolated mitochondrial respiration from the Txnip-KO heart was mildly reduced during state 3 with complex I–linked substrates (glutamate/malate) compared with WT mitochondrial respiration (89% ± 5% of WT; Figure 1, E and F). Complex III and IV activities, reflecting the transfer and the initial rate of oxidation of cytochrome c, were comparable between the genotypes in the isolated mitochondria (Supplemental Figure 2, A and B). However, using the precursor substrate pyruvate resulted in an approximately 35% reduction in state 3 respiration in isolated mitochondria from Txnip-KO compared with WT hearts (Figure 1, G and H). This indicated that Txnip-KO hearts are limited in the ability to use carbohydrate precursor effectively in mitochondrial respiration. Immunoblot analysis for OXPHOS complexes indicated that there was no difference in expression levels of mitochondrial OXPHOS components between groups (Supplemental Figure 2, C and D). Blue-native gel electrophoresis in the first dimension also showed no difference in expression levels of mitochondrial OXPHOS or complexes I, III, and V (Supplemental Figure 2D). Hence, the decreased respiratory control ratio was likely caused not by global loss of respiratory complexes, but by functional loss of ADP-stimulated respiration, in Txnip-KO mitochondria.
We next measured the sensitivity of mitochondrial permeability transition pore using swelling assays in isolated mitochondria, since ROS induces opening of the mitochondrial permeability transition pore, and pore opening can result in mitochondrial dysfunction with uncoupled OXPHOS (22). No difference was observed in the mitochondrial permeability transition pore assay between the genotypes (Supplemental Figure 2, E and F). Collectively, these results support the concept that downregulation of transcripts encoding mitochondrial enzymes by deletion of Txnip is associated with reduced mitochondrial function, without changing mitochondrial permeability transition pore opening in the heart.
Txnip-KO hearts exhibit morphological changes in mitochondria.
A reduction in total cellular mitochondrial mass is one explanation that would account for the observed global decrease in mitochondrial gene expression and mitochondrial function. In particular, blunted mitochondrial respiration might reflect a reduced rate of mitochondrial proliferation and biogenesis, an increased rate of mitochondrial autophagy, or both (23). To address this, we determined mitochondrial mass by measuring mitochondrial DNA content and visualized mitochondrial morphology by transmission electron microscopy. Mitochondria copy number was determined by Southern blot analysis using specific probes for mitochondrial DNA (mtDNA) (COX II) and nuclear DNA (COX IV) and by quantitative real-time PCR (qPCR) for mtDNA (COX I) and nuclear DNA (NDUFV1). Both techniques showed that, rather than being reduced, the ratio of Txnip-KO heart mtDNA relative to nuclear DNA was slightly increased compared with that of WT controls (Figure 2, A and B).
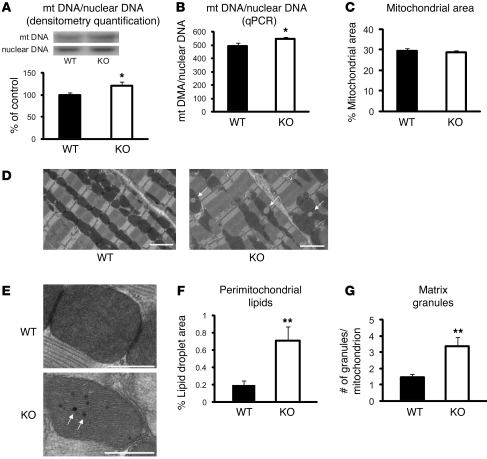
Figure 2. Ultrastructural and quantitative analyses of mitochondria.
(A) Relative mtDNA copy number in the heart, calculated as the ratio of COX II (mitochondria) to COX IV (nuclear) genes as determined by Southern blotting. The ratio, quantified by densitometry, is shown as percent of WT control (n = 6 each). (B) The COX I gene of the mtDNA and the NDUFV1 nuclear DNA gene were amplified by qPCR. Amplification curves were used to determine the relative mtDNA/nuclear DNA ratio in each sample (n = 6). (C–G) Electron micrographs from WT and Txnip-KO hearts showed morphological differences in mitochondria. Arrows denote lipid droplets (D) and mitochondrial matrix granules (E). Scale bars: 2 μm (D); 500 nm (E). Values are mean ± SEM. *P < 0.05, **P < 0.01 versus WT.
Myocardial samples were next qualitatively assessed by transmission electron microscopy for structural mitochondrial biogenesis or evidence of mitochondrial autophagy. Similar to the WT hearts, which displayed uniform mitochondria in longitudinal sections, Txnip-KO hearts showed no change in mitochondrial structure, with clear cristae and equivalent matrix density (Figure 2, D and E). No evidence of mitochondrial autophagy was observed. Quantitative image analysis confirmed no difference in mitochondrial area as a percentage of total cellular area (on 10–15 different fields per mouse) between Txnip-KO (n = 4) and WT (n = 3) mice (Figure 2C). Together, these observations suggest that a reduction in Txnip-KO mitochondrial respiration is not attributable to a global decrease in mitochondrial mass.
Several distinct morphological features were evident in the Txnip-KO mitochondria by electron microscopy. Large perimitochondrial lipid droplets were observed abutting the Txnip-KO mitochondria (Figure 2, D and F). This was particularly interesting given previous characterizations of the Txnip-KO phenotype that were consistent with enhanced de novo lipogenesis (16). An increased presence of mitochondrial matrix granules, precipitates made of phospholipids, glycoprotein, and calcium-precipitable lipoprotein, was also observed in mitochondria from Txnip-KO compared with WT hearts (Figure 2, E and G). A previous report suggested that matrix granules serve to increase calcium concentration in the mitochondrial matrix, enhancing the activity of dehydrogenases to control energy metabolism (24). It is thus possible that decreased mitochondrial respiration might be partially compensated by these morphological adaptations in Txnip-KO mice.
Targeted deletion of Txnip improves myocardial tolerance to ischemia and reperfusion.
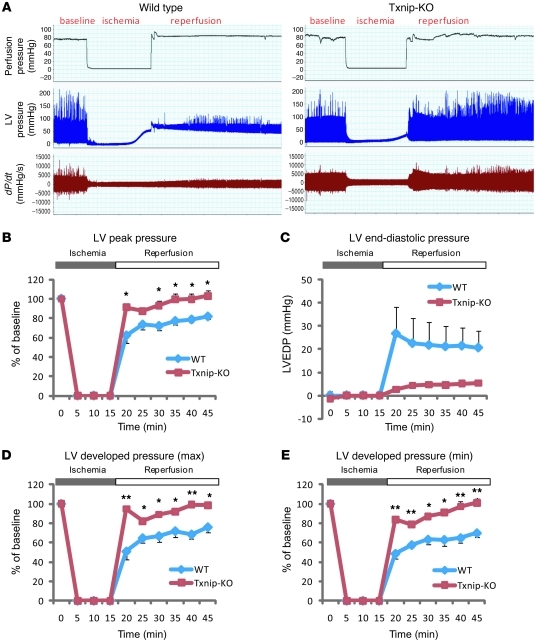
Mitochondrial respiration is an important biological process for ATP generation, redox signaling, Ca2+ handling, and cell death regulation in cardiomyocytes, all of which are crucial in hypoxic hearts for maintaining cellular homeostasis and myocardial integrity (25). Given the central role that mitochondria serve during hypoxia, we hypothesized that mitochondrial dysfunction by Txnip deletion amplifies deleterious effects during ischemia-reperfusion injury. We first tested this hypothesis using ex vivo–isolated perfused mouse hearts from total Txnip-KO mice subjected to a simulated period of ischemia followed by reperfusion: myocardial performance was measured as a function of LV developed pressure. Myocardial function during the aerobic period at baseline was not significantly different between WT (n = 4) and Txnip-KO (n = 3) hearts (LV peak pressure, WT, 107 ± 6.3 mmHg; Txnip-KO, 108 ± 5.2 mmHg). A 15-minute period of ischemia followed by 30 minutes of reperfusion substantially impaired cardiac function in WT hearts (Figure 3A), as expected. Surprisingly, Txnip-KO hearts had significantly improved postischemic recovery of LV peak pressure (5 minutes after reperfusion, KO, 91% ± 4%; WT, 62% ± 8%; P < 0.05; Figure 3B). At the end of reperfusion, LV end-diastolic pressure in Txnip-KO hearts was decreased to 5.3 ± 0.2 mmHg compared with 21 ± 7 mmHg in WT hearts (Figure 3C). Upon reperfusion, the rate of contraction (dP/dtmax) and rate of relaxation (dP/dtmin) were also markedly more preserved in Txnip-KO than in WT hearts (25 minutes after reperfusuion, KO, 91% ± 3% and 97 ± 6%, respectively; WT, 68% ± 5% and 64% ± 5%, respectively; P < 0.01; Figure 3, D and E). These results demonstrated that, despite reduced mitochondrial function, deletion of Txnip improves LV functional recovery after ischemia-reperfusion injury.
Figure 3. Txnip-KO hearts have improved functional recovery after ischemia-reperfusion injury.
(A) Representative tracing of LV pressure in a WT mouse heart (n = 3) and a Txnip-KO mouse heart (n = 4) with a Langendorff perfusion system. (B–E) LV peak pressure (B), LV end-diastolic pressure (LVEDP; C), and maximal (D) and minimal (E) LV developed pressure were measured during 15 minutes ischemia and 30 minutes reperfusion. Values are mean ± SEM. *P < 0.05, **P < 0.01 versus WT.
Mice with inducible cardiac-specific Txnip deletion have cardiometabolic phenotypes similar to total Txnip-KO mice.
To test whether the observed phenotypes were direct results of Txnip deficiency in the heart or were confounded by compensatory changes in systemic metabolites, a time-course study was performed using inducible cardiomyocyte-specific Txnip-KO mice. Temporal induction of cardiomyocyte-specific Txnip deletion was achieved using αMHC-MerCreMer/Txnipfl/fl mice injected with 4-hydroxy-tamoxifen. Gender-matched littermates of αMHC-MerCreMer/Txnipfl/fl mice treated with vehicle (i.e., lacking 4-hydroxy-tamoxifen) were used as controls. It has previously been shown that the combination of Cre and tamoxifen induces a transient cardiomyopathy, which recovers within weeks (26); thus, cardiac function can be assessed without these side effects at 4 weeks after tamoxifen injection (27). Decreased Txnip protein expression in the hearts of αMHC-MerCreMer/Txnipfl/fl mice was previously confirmed 4 weeks after 4-hydroxy-tamoxifen administration (17). Thus, we quantified the mRNA expression levels from cardiac tissues in the animals at 4 and 15 weeks after 4-hydroxy-tamoxifen administration as early and late phases, respectively. Pathway-focused gene expression profiling using qPCR revealed that Txnip-KO cardiomyocytes began changes at the transcriptional level at the early phase that persisted at the late phase (Supplemental Table 2). For instance, transcripts of the metabolic regulators pyruvate carboxylase and mitochondrial isocitrate dehydrogenase 2, both of which were downregulated in total Txnip-KO mice by PMAGE, were consistently downregulated in cardiac-specific Txnip-KO hearts compared with control hearts at the early and late phases. These transcriptional changes were not detected in WT animals (Txnip+/+) treated with 4-hydroxy-tamoxifen (Supplemental Table 2), which suggests that the effects were not caused by administration of 4-hydroxy-tamoxifen.
To confirm that these transcriptional alterations are accompanied by physiological metabolic changes, we assessed mitochondrial respiration in saponin-skinned cardiac fibers from cardiomyocyte-specific Txnip-KO mice. Using the precursor substrate pyruvate resulted in a 26%–47% reduction in respiration control ratio in cardiac fibers of cardiac-specific Txnip-KO compared with control mice (53% ± 5% of control at the early phase; 74% ± 5% of control at the late phase; P < 0.05; Figure 4, A and B). Txnip+/+ mice similarly treated with 4-hydroxy-tamoxifen did not yield a significant decrease in respiration control ratio (vehicle, 3.3 ± 0.2; 4-hydroxy-tamoxifen, 4.7 ± 0.8; n = 5; P = NS), which confirmed that the decrease in mitochondrial respiration was not caused by 4-hydroxy-tamoxifen administration. As cardiometabolic phenotypes in cardiomyocyte-specific Txnip-KO mice were similar to those of systemic Txnip-KO mice, we tested whether cardiomyocyte-specific deletion of Txnip also improves myocardial mechanical recovery after ischemia-reperfusion injury using ex vivo isolated perfused mouse hearts. A 20-minute period of ischemia followed by a 10-minute reperfusion period substantially impaired cardiac function in control hearts, as expected, at both early and late phases. Consistent with ischemic tolerance in total Txnip-KO mice, cardiomyocyte-specific Txnip-KO hearts showed significantly improved postischemic recovery of LV developed pressure, both at the early phase (5 minutes after reperfusion, dP/dtmax, 46% ± 4% increase vs. control; dP/dtmin, 98% ± 20% increase vs. control; Figure 4C) and at the late phase (5 minutes after reperfusion, dP/dtmax, 70% ± 16% increase vs. control; dP/dtmin, 49% ± 17% increase vs. control; Figure 4D). These results demonstrated that cardiometabolic phenotypes and myocardial tolerance to ischemia are direct results of Txnip deficiency, rather than a result of changes in systemic metabolic factors.
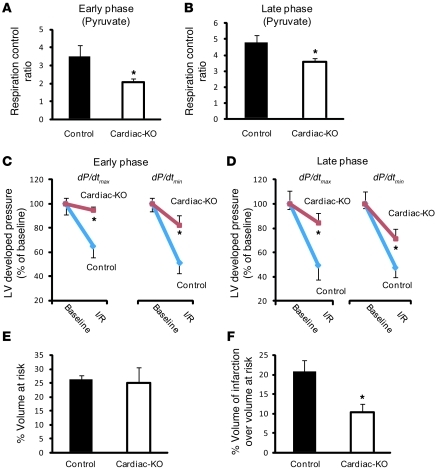
Figure 4. Inducible cardiac-specific Txnip-KO mice have cardiometabolic phenotypes similar to those of systemic Txnip-KO mice.
(A and B) Respiratory parameters were obtained from permeabilized cardiac fibers incubated with 10 mM pyruvate and 5 mM malate in cardiomyocyte-specific Txnip-KO mice (Cardiac-KO). Early and late phases represent time points at 4 (n = 6) and 15 (n = 7) weeks, respectively, after deletion of Txnip by 4-hydroxy-tamoxifen. (C and D) Cardiac-specific Txnip-KO hearts had significantly improved functional recovery after ischemia-reperfusion injury. LV developed pressure (dP/dtmax and dP/dtmin) were measured with a Langendorff perfusion system at baseline and after ischemia-reperfusuion injury (I/R) — at 5 minutes reperfusion after 20 minutes ischemia. Values are mean ± SEM. n = 3–6 per group. (E and F) LV volume at risk (E; P = NS) and infarct volume over volume at risk (F; P < 0.05) in cardiac-specific Txnip-KO mice (n = 9) and their controls (n = 5), with transient (30 minutes) ischemia followed by reperfusion (24 hours) using a reversible ligation of the distal portion of left anterior descending artery. Values are mean ± SEM. *P < 0.05 versus control.
To corroborate the unexpected ischemic tolerance of Txnip-KO hearts in vivo, cardiomyocyte-specific Txnip-KO mice and their littermate controls were subjected to 30 minutes of ischemia and 24 hours of reperfusion using a reversible ligation of the left coronary artery at the early phase. As demonstrated previously (17), cardiac-restricted deletion did not lead to the perturbations in systemic metabolism that are well-described for systemic Txnip deletion. No significant differences in blood glucose levels or body weight were observed before (17) or 24 hours after reperfusion between cardiac-specific Txnip-KO (121 ± 20 mg/dl, 25 ± 1 g) and control mice (148 ± 31 mg/dl, 25 ± 1 g; P = NS). Upon reperfusion, the measured volume of myocardium at risk was similar in cardiac-specific Txnip-KO and control mice (Figure 4E), which indicates that placement of the ligature was similar between the groups. However, infarct volume was significantly decreased by 50% ± 6% in cardiac-specific Txnip-KO mice (n = 9) compared with controls (n = 5). The infarct volume normalized by volume at risk was significantly lower in cardiac-specific Txnip-KO mice (10% ± 2%) than in control mice (21% ± 3%; P < 0.05; Figure 4F). These results demonstrate that, in spite of reduced mitochondrial function, deletion of Txnip decreased the extent of myocardial infarction and improved LV functional recovery after ischemia-reperfusion injury. Limitations of this study include small infarct size after reperfusion compared with previous reports, because we ligated a relatively distal portion of the coronary artery to avoid selection bias from a high mortality rate.
Targeted deletion of Txnip increases mitochondrial Trx2 activity after ischemia-reperfusion injury.
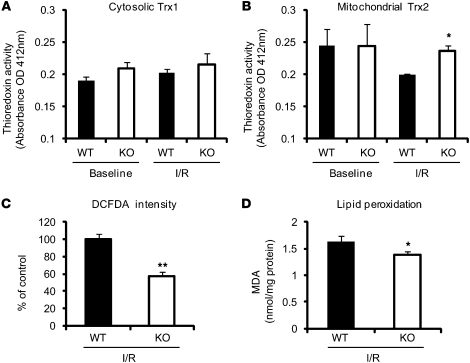
Ischemia-reperfusion injury results in ROS generation, ultimately leading to oxidative stress in cardiomyocytes (4). While thioredoxins, binding partners of Txnip, are potent antioxidants in cardiomyocytes (28, 29), dysfunctional mitochondria can generate increased ROS levels, which are an inevitable byproduct of OXPHOS (30). To determine whether altered myocardial ROS levels are associated with the ischemic tolerance in mice with Txnip deletion, we measured cytoplasmic and mitochondrial thioredoxin activities and ROS levels in Txnip-KO and WT hearts. No differences in myocardial activities of cytosolic Trx1 were seen at baseline or after 15 minutes ischemia and 30 minutes reperfusion between Txnip-KO (n = 4) and WT (n = 3) mice (Figure 5A). Although mitochondrial Trx2 activities were comparable between the groups at baseline, ablation of Txnip significantly attenuated the decrease in Trx2 activity after ischemia-reperfusion injury (Figure 5B), which indicates that Txnip deficiency is associated with enhanced mitochondrial Trx2 activity in injured hearts. Myocardial ROS generation, as assessed by staining of the green fluorescence dye dichlorodihydrofluorescein diacetate (DCFDA), was decreased in Txnip-KO hearts after ischemia-reperfusion injury (Figure 5C). In addition, the levels of cellular lipid peroxide, an indicator of oxidative stress estimated as malondialdehyde (MDA), were also decreased in the myocardium from Txnip-KO mice compared with WT mice after ischemia-reperfusion (Figure 5D). Thus, the ischemic protective effects in Txnip-KO hearts were associated with decreased levels of ROS in the myocardium, with preserved activity of mitochondrial Trx2 after ischemia-reperfusion injury.
Figure 5. Effect of Txnip deletion on thioredoxin activities and oxidative stress in the myocardium after 15 minutes ischemia and 30 minutes reperfusion.
(A and B) Thioredoxin activity was measured using an insulin disulfide reduction assay in the cytosolic (A) and mitochondria-enriched (B) fractions. (C) Frozen sections were stained with DCFDA (4 μmol/l) to assess the tissue levels of ROS, and DCFDA intensities in the myocardium were calculated. (D) Tissue levels of MDA were measured in whole heart homogenates. Values are mean ± SEM. *P < 0.05, **P < 0.01 versus WT.
Targeted deletion of Txnip enhances anaerobic glycolysis.
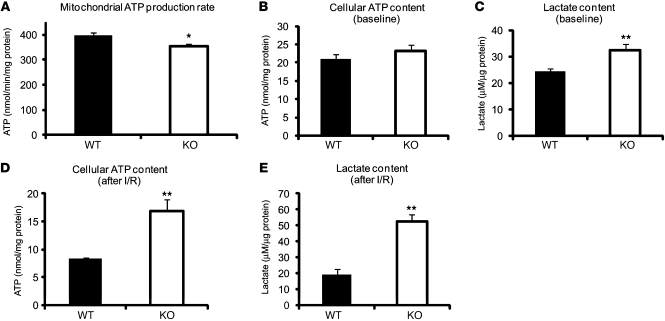
Inadequate mitochondrial respiration also brings the risk of diminished ATP production and futile ATP hydrolysis. Therefore, we measured ATP content in isolated mitochondria from Txnip-KO and WT hearts. Consistent with the reduced state 3 (ADP-dependent) respiration in Txnip-KO hearts (Figure 1F), mitochondrial ATP production rate during state 3 was slightly decreased by 11% ± 3% in Txnip-KO versus WT hearts (P < 0.05; Figure 6A). However, no significant difference in total cellular ATP content was seen at baseline between the genotypes (Figure 6B). Cellular lactate levels were increased in Txnip-KO hearts at baseline (32% ± 10% increase vs. WT; P < 0.01; Figure 6C), reflecting a dependence on glycolysis for ATP production, rather than on mitochondrial metabolism. Thus, ATP produced by enhanced glycolysis might compensate for the small decrease in aerobic ATP production in Txnip-KO cardiomyocytes.
Figure 6. Deletion of Txnip enhances basal and ischemia-reperfusion anaerobic glycolysis.
(A) Mitochondrial ATP production rate was measured kinetically in isolated mitochondria after the addition of 1 mM ADP in the presence of the respiratory substrates 5 mM glutamate and 2 mM malate, using a luciferase/luciferin assay. (B–E) Cellular ATP (B and D) and lactate (C and E) levels were measured either (B and C) at baseline from excised hearts in WT or Txnip-KO mice or (D and E) from isolated hearts in an ex vivo ischemia (15 minutes) and reperfusion (30 minutes) injury model. Values are mean ± SEM. n = 6–12 per group. *P < 0.05, **P < 0.01 versus WT.
After 15 minutes ischemia and 30 minutes reperfusion, total cellular ATP content was 2.0 ± 0.2 times higher in Txnip-KO than WT hearts (P < 0.01; Figure 6D), with a corresponding parallel increase in cytosolic lactate production (2.7 ± 0.2 times higher than WT; P < 0.01; Figure 6E). As aerobic metabolism (ATP production by mitochondria) is normally blocked during ischemia, the higher cellular ATP content in Txnip-KO hearts is likely caused by enhanced anaerobic glycolysis. These results support the concept that the basal redirection of myocardial fuel use toward anaerobic glycolytic metabolism is further enhanced under ischemia to maintain a higher energy source outside of mitochondria. Thus, the beneficial effects in Txnip-KO hearts might be associated with their greater capacity for anaerobic metabolism after ischemia-reperfusion injury.
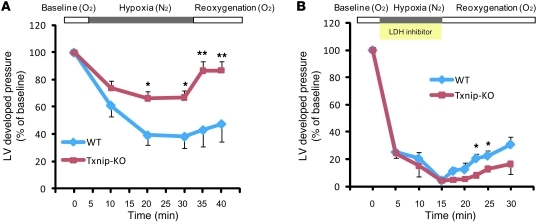
Induction of myocardial hypoxia by cessation of coronary perfusion does not permit a steady supply of anaerobic substrate to the myocardium. Thus, we sought to confirm whether increased anaerobic metabolism resulting from deletion of Txnip contributes to cardioprotection during anaerobic perfusion. Isolated mouse hearts were perfused with a hypoxic solution (95% N2, 5% CO2) and reoxygenized with a normoxic solution (95% O2, 5% CO2). Exposure to hypoxic perfusion fluid in the presence of glucose and fatty acids resulted in impaired LV mechanical performance (Figure 7A). The LV developed pressure fell progressively, up to 38% ± 9% of baseline, throughout the 30 minutes of hypoxia in WT hearts (n = 4). However, Txnip-KO hearts (n = 6) maintained better mechanical performance (LV developed pressure 67% ± 5% of baseline; P < 0.05). Furthermore, during the reoxygenation period, the recoverability of LV developed pressure after exposure to hypoxia was better preserved in Txnip-KO than in WT hearts (86% ± 7% vs. 43% ± 12% of baseline at 5 minutes of reoxygenation; P < 0.01). These results support the hypothesis that Txnip deletion redirects metabolic flux through glycolytic pathways, which provides a functional advantage to the hypoxic myocardium.
Figure 7. Txnip-KO hearts increase anaerobic metabolism under hypoxia.
(A) Txnip-KO hearts had significantly improved mechanical function during hypoxia (95% N2, 5% CO2) and recovery (95% O2, 5% CO2). Hearts were perfused in a Langendorff heart model system in the presence of glucose (5.5 mM), mixed long-chain fatty acids (0.4 mM bound to 1% albumin), DL-β-hydroxybutyrate (0.38 mM), lactate (1 mM), and insulin (50 μU/ml). (B) To inhibit cellular glycolysis, hearts were perfused with oxamate (50 mM), a competitive inhibitor of LDH. n = 3 per group. Values are mean ± SEM. *P < 0.05, **P < 0.01 versus WT.
To further challenge this concept, we then tested whether inhibition of glycolysis would preferentially affect the myocardial tolerance to hypoxia seen in Txnip-KO hearts. Cellular glycolytic metabolism was inhibited using 50 mM oxamate, a competitive inhibitor of lactate dehydrogenase (LDH), which catalyzes the reduction of pyruvate to lactate (31). It should be noted that oxamate can inhibit both cytosolic and mitochondrial LDH; hence, aerobic metabolism is also affected during aerobic perfusion. Therefore, we perfused hearts with oxamate only during anaerobic perfusion. Strikingly, the beneficial effects seen during hypoxia in Txnip-KO hearts were totally abolished by blocking the glycolytic energy biotransformation of pyruvate into lactate. The recovery of LV developed pressure deteriorated at 7.5 minutes reperfusion in Txnip-KO compared with WT hearts (39% ± 7% of WT; n = 3; P < 0.05; Figure 7B). These results indicate that enhanced glycolysis is a key mechanism for cardioprotective effects by deletion of Txnip.
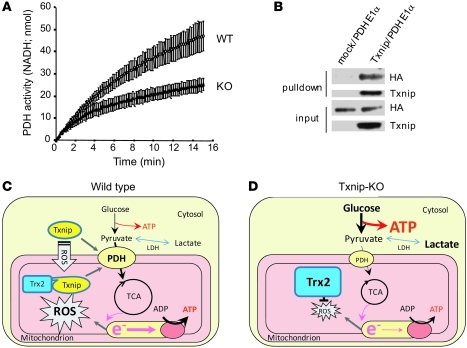
The cellular regulation of glycolytic flux relative to mitochondrial respiration is tightly controlled by a key facilitator, pyruvate dehydrogenase (PDH) complex. PDH complex is a multisubunit enzyme in the mitochondrial matrix that has previously been described as the gatekeeper for entry of glucose-derived pyruvate into mitochondria (32). Since PDHE1α was one of the mitochondrial enzymatic subunits that was decreased in Txnip-KO mice (Table 1), we measured the quantity and enzymatic activity of PDH in the mitochondria. ELISA using 2 PDH-specific antibodies against the E2 component and E3-binding protein indicated that there was no difference in PDH quantity as a complex between the groups (WT, 2.0 ± 0.1 OD/h/mg; KO, 2.1 ± 0.1 OD/h/mg; P = NS). However, PDH activity was significantly repressed in Txnip-KO hearts (54% ± 6% of WT; n = 4; P < 0.05; Figure 8A), suggestive of redirection of glycolytically derived pyruvate away from transport into the mitochondria and toward cytosolic lactate production and ATP generation in the Txnip-KO heart. Next, to investigate how Txnip controls pyruvate flux in mitochondria, we performed protein interaction pulldown assays between Txnip and a protein regulating pyruvate flux or recycling using magnetic Streptactin beads. We tested PDHE1α and pyruvate carboxylase, as both were differentially expressed in Txnip-KO hearts. Interestingly, the assay confirmed an interaction of Txnip with PDHE1α, but not with pyruvate carboxylase (Figure 8B). Recently, it has been reported that phosphorylation levels of PDHE1α are markedly increased in soleus muscle from fasting Txnip-KO mice (33). Thus, these data together suggest that Txnip is involved directly or indirectly in a key switch from aerobic to anaerobic metabolism by regulating mitochondrial respiration and PDH activity.
Figure 8. Txnip is involved in a key switch from aerobic to anaerobic metabolism.
(A) PDH activity was measured in isolated mitochondria as the rate of NADH production at 340 nm. (B) The interaction between Txnip and PDHE1α was assessed using a Txnip pulldown assay. HA-tagged PDHE1α was pulled down with Txnip, indicative of an interaction between these proteins. (C and D) Mechanism by which Txnip deletion may achieve cardioprotection in ischemia-reperfusion injury. (C) In WT hearts, upon stimulation with ROS, Txnip translocates into mitochondria to bind to and deactivate mitochondrial Trx2. (D) In Txnip-KO hearts, Trx2 is not inhibited by Txnip, thereby scavenging ROS efficiently. Txnip deficiency also reprograms glucose metabolism to more anaerobic ATP production in association with PDH, maintaining energy homeostasis in cardiomyocytes under ischemia-reperfusion injury.
Discussion
Cellular metabolism and oxidative stress are interrelated processes in the mitochondria implicated in a wide range of pathological conditions in the heart, including ischemia-reperfusion injury. The ubiquitous antioxidant thioredoxin and its endogenous inhibitor, Txnip, are emerging as a key link between mitochondrial function regulation and cellular redox balance. A previous study has shown that targeted degradation of Txnip mRNA by injection of a small nucleotide-based catalytic enzyme directly into ischemic myocardium results in enhanced cardiomyocyte survival and reduced LV remodeling (34). However, it remains unclear how Txnip mediated this adaptive response to ischemic injury. In the present study, we found that Txnip deficiency led to a substantial metabolic switch at the transcriptional level and repressed mitochondrial respiration, directing cardiomyocytes toward enhanced anaerobic glycolysis. The concept that defects in mitochondrial metabolism produce enhanced anaerobic metabolism is supported by our finding that the corresponding expression levels of the key enzymes controlling the aerobic metabolic pattern were downregulated. The RNA levels of 36 dehydrogenases and 71 oxidoreductases controlling OXPHOS (22 enzymes) and electron transport (36 enzymes) were decreased in Txnip-KO compared with WT hearts (Supplemental Table 1). Comparative analysis of the protein levels of key enzymes regulating the glycolytic and aerobic (TCA cycle) pathways supported the gene array data. For instance, the protein level of isocitrate dehydrogenase 2 participating in the TCA cycle (aerobic respiration) was significantly decreased in Txnip-KO cells. Furthermore, our experimental data confirmed that deletion of Txnip led to metabolic behavior that made substantial use of anaerobic glycolytic metabolism in association with the key metabolic switch controller PDH. Thus, defects in mitochondrial metabolism by deletion of Txnip may play a role in determining intracellular bioenergetic flux (aerobic or anaerobic). This metabolic shift was further accelerated by ischemia and was associated with ischemic tolerance in both ex vivo controlled conditions and in vivo physiological conditions. Given the central role that mitochondria serve in ischemia-reperfusion injury, we initially hypothesized that Txnip-KO hearts would manifest more severe cellular injury, reduced intracellular ATP levels, and cardiac dysfunction after ischemia and reperfusion. Surprisingly, deletion of Txnip afforded a protective advantage to the ischemic heart.
Adaptation to hypoxia requires a dramatic reduction in cellular oxygen consumption (32) while maintaining an energy source to regenerate its ATP stores. This response involves a shift in cellular fuel utilization from mitochondrial respiration, which consumes oxygen, to generation of ATP outside of mitochondria by metabolizing glucose without the use of oxygen (anaerobic glycolysis) (32). Anaerobic glycolysis appears to be the preferred means of replenishing ATP stores in hypoxic organs, and it requires almost 20 times more glucose to generate 1 mole of ATP by anaerobic glycolysis than by OXPHOS. Because lipids are not used as major precursors for anaerobic metabolism, glucose is the primary substrate to support anaerobic metabolism. Thus, anaerobic metabolism with concomitant increased use of glucose, as fostered by Txnip deletion, plays a significant role in preserving myocardial function and structure and in promoting recoverability of the hypoxic heart (35).
Among antioxidant defense systems against mitochondrial ROS, Trx2 is the major contributor for respiration-dependent H2O2 removal in mitochondria (36). Trx2 prevents ischemia-induced ROS production in a femoral artery ligation model (37). Overexpression of Trx2-dependent peroxidase (peroxiredoxin-3) improves survival after myocardial infarction with an attenuation of mitochondrial oxidative stress (38). Recently, Saxena et al. discovered a critically important interaction between mitochondrial Trx2 and Txnip (8). Using a different technique, we also confirmed the direct interaction between Txnip and Trx2 (Supplemental Figure 2G). Under physiological conditions, Saxena et al. observed that Txnip was localized primarily in the nucleus of cells, but oxidative stress led to Txnip shuttling into mitochondria, where Txnip bound to and inhibited mitochondria Trx2 (8). In the present study, we found that cytosolic Trx1 activity was not significantly affected; however, mitochondrial Trx2 activity was decreased by ischemia-reperfusion in WT hearts. In a mechanism analogous to that reported by Saxena et al., we suspect that Trx2 is inhibited by Txnip that translocates into mitochondria by ischemia-reperfusion–induced ROS formation. However, Txnip-KO hearts exhibited unperturbed Trx2 activity, even after ischemia and reperfusion. Taken together, these results suggest that a redox-sensitive interaction between Txnip and Trx2 contributes to the redox balance in mitochondria (Figure 8, C and D).
Stimuli that uncouple mitochondrial respiration prior to an ischemic insult have been suggested to underlie cardioprotection mediated by ischemic preconditioning (39). The continuation of mitochondrial aerobic respiration in the absence of oxygen results in excessive or unneutralized ROS production, mitochondrial calcium overload, and the onset of mitochondrial permeability transition, which collectively promote mitochondrial-driven cardiomyocyte death (3). Pharmacological inhibition of complex I and II has been found to be cardioprotective by limiting ROS production, resulting in reduced infarct size after reperfusion in a rat model of ischemia-reperfusion injury (40). However, the hypoxic cell requires an energy source to regenerate its ATP stores. A previous report demonstrated that inclusion of glucose in the perfusate of isolated perfused rat heart preparations results in marked improvement in electrical and mechanical performance of the heart subjected to anoxia and in enhanced recovery during the subsequent period of reoxygenation (35). While lactate production was 5-fold greater in the glucose-supported anoxic heart than in the anoxic heart without glucose, morphologic changes of mitochondria during anoxia were averted by inclusion of glucose in the perfusion fluid (35). Thus, anaerobic metabolism, with concomitant increased use of glucose, can preserve myocardial function and structure and promote ischemic heart recovery. These cardioprotective mechanisms can be feasibly achieved by deletion of Txnip, which (a) limits myocardial ROS levels effectively by mitochondrial Trx2, (b) reduces oxygen consumption in mitochondria, and (c) greatly favors cellular glucose uptake and glycogen storage in the heart while simultaneously enhancing glucose flux toward anaerobic glycolytic metabolism. As a consequence, a high-flux glycolytic state coupled with the residual respiration provided sufficient ATP for survival and cellular protection in Txnip-KO hearts. As previously observed, these data reflect the physiology of tumor-genesis and the “Warburg effect” — physiology by which cancerous cells preferentially rely upon glycolytic metabolism, despite the availability of oxygen to support mitochondrial respiration (41). Txnip acts as a mediator to integrate redox balance and energetic needs with cellular glucose supply: this may imply that the Warburg effect can be protective against ischemic insults in cardiomyocytes when ROS balance is secured by a potent antioxidant, such as mitochondrial Trx2.
We here identified Txnip as a regulator of mitochondrial function and demonstrated the role that Txnip serves during ischemia-reperfusion injury in the heart. However, it remains unclear how Txnip participates in the transcriptional regulatory circuits that control mitochondrial respiration. Mitochondrial Trx2 interacts with components of the mitochondrial respiratory chain and plays a role in the regulation of the mitochondrial membrane potential (42). Trx2 deficiency is embryonic lethal at gestational day 10.5 in mice, which coincides with the transition period from anaerobic to aerobic metabolism in the embryo (43). Thus, Txnip may regulate mitochondrial metabolism through mitochondrial Trx2.
To further identify a potential mechanistic role of Txnip by protein-protein interaction, we performed a proteomic screening using glutathione S-transferase (GST) tag protein interaction pulldown assays (Supplemental Results). Proteins from HEK 293F cells overexpressing Txnip with GST or GST alone (as a negative control) were pulled down from cellular lysates using magnetic glutathione beads, subjected to SDS-PAGE, and then analyzed by mass spectrometry. This proteomic screening revealed a putative interaction with several proteins involved in metabolism as candidates for binding targets of Txnip (Supplemental Table 3). These potential interactions of Txnip with other proteins may also be involved in a mechanistic role of Txnip as a controller of energy metabolism. Further studies will be necessary to define these roles.
In conclusion, we identified Txnip as a regulator of mitochondrial function and demonstrated that Txnip deficiency led to a substantial metabolic switch, directing cardiomyocytes toward enhanced glycolytic metabolism. Several potential pathways might be simultaneously involved in Txnip-KO mice; therefore, no single cause of mitochondrial dysfunction could be defined. Nonetheless, the present study provides insights into the role of myocardial Txnip in anaerobic metabolism that contributes to acute cardioprotection during ischemia-reperfusion injury.
Methods
Animals.
Total Txnip-KO mice and cardiac-specific Txnip-KO mice were described previously (17). For total Txnip-KO mice, gender-matched WT littermates were used as controls. Temporally inducible cardiomyocyte-specific Txnip deletion was achieved using αMHC-MerCreMer/Txnipfl/fl mice injected with 0.5 mg 4-hydroxy-tamoxifen per day for 2 weeks. For the cardiac-specific Txnip-KO mice, gender-matched littermates of αMHC-MerCreMer/Txnipfl/fl mice treated with vehicle (i.e., lacking 4-hydroxy-tamoxifen) were designated as controls, in which Txnip expression was identical to that of WT mice (17). Although tissue-specific KOs prevent the confounding compensatory influence of metabolic disarray from systemic Txnip deletion, Cre transgene and tamoxifen administration may potentially lead to unwanted side effects (26, 44). Mice with total ablation of Txnip are free from potential side effects of Cre transgene or 4-hydroxy-tamoxifen. As both mouse models have these advantages and disadvantages, we confirmed the cardiometabolic phenotypes in both total and inducible cardiomyocyte-specific Txnip-KO animals.
Gene expression analysis and 2D gel electrophoresis.
Global gene expression was analyzed by PMAGE as described previously (18). Polony sequence by ligation was used to provide an accurate multiplexed platform for high-throughput DNA tag sequencing, and digital quantification of tag data accommodated statistical analysis over a broad dynamic range of mRNA expression, including low-abundance mRNAs encoding transcriptional factors. Gene ontology analysis was performed using the software PANTHER (SRI International), which classifies genes by their functions using published evidence. Pathway-focused gene expression analysis was performed using RT2 Profiler PCR Array based on the SYBR Green system (PAMM-006A; QIAGEN). 2D gels were run, fixed in a 25% methanol/10% acetic acid solution, and stained overnight with the Blue Gel Stain Kit (Proteome Systems) as previously described (45). Spots that exhibited significant changes of intensities between genotypes (P < 0.05) were selected for identification by MALDI-TOF mass spectra Voyager-DE STR (Applied Biosystems Inc.). The mass spectrometry analyses for the 2D gel experiment were conducted in the UTHSCSA Institutional Mass Spectrometry Laboratory.
Mitochondrial respiration.
Mitochondrial oxygen consumption was assessed polarographically in saponin-skinned fibers or isolated mitochondria from mouse hearts by a Clark-type electrode and 5300A Biological Oxygen Monitor (YSI Inc.). To prepare skinned fibers, ventricular fiber bundles (5–10 mg) were mechanically separated and permeabilized with saponin (52.5 mg/l). Different substrates were used as follows: 5 mM glutamate plus 2 mM malate as complex I substrates; 5 mM succinate plus 2.2 μM rotenone as complex II substrates with inhibition of complex I by rotenone. Carbohydrate substrates were 10 mM pyruvate plus 5 mM malate for skinned fibers and 10 mM pyruvate plus 2 mM malate for isolated mitochondria. State 3 respiration was measured in the presence of respiratory substrates after the addition of 1 mM ADP. In isolated mitochondria, state 4 respiration was measured upon depletion of ADP. In skinned fibers, state 4 respiration was determined after the addition of 60 μM atractyloside (a potent inhibitor of the ATP/ADP carrier), considered the control state of respiration, as described previously (46). The respiration control ratio was calculated by dividing state 3 by state 4 respiration rates. The mitochondria-enriched fraction and cytosolic fraction were prepared by differential centrifugation from the heart, as described previously (47).
Ultrastructural and quantitative analyses of mitochondria.
Ultrastructure of mitochondria was assessed on samples perfusion fixed in 2.5% glutaraldehyde/formaldehyde (Electron Microscopy Sciences) in 0.1 M cacodylate buffer (pH 7.2) and postfixed in 1% osmium tetroxide. The tissue blocks were embedded in a Spurr/Quetol epoxy resin formulation. Thin sections were stained with uracyl acetate and lead citrate and examined by electron microscopy (JEM 1010; JEOL Inc.). Quantitative morphometric assessments of mitochondria were carried out with ImagePro (Media Cybernetics Inc.). mtDNA content was assessed by Southern blot analysis as described previously (48). 10 μg total cellular DNA was digested with NcoI and subjected to Southern analysis using a cDNA for COX II as a probe for mtDNA. The blot was stripped and rehybridized to a cDNA for COX IV, a nuclear DNA–encoded gene, as an input control. The mtDNA copy number was also determined by qPCR analysis, as described previously (49). Amplification curves for the COX I gene of the mtDNA and the NDUFV1 nuclear DNA gene were obtained to measure the relative mtDNA/nuclear DNA ratio.
Thioredoxin activity, ROS, and biochemical assays.
Thioredoxin-reducing activity was measured using the insulin disulfide reduction assay (50, 51). Recombinant human Trx1 and Trx2 proteins, produced and purified as GST fusion proteins, were used as controls for the assay. Tissue levels of lipid peroxides — as determined by MDA levels — were estimated in whole heart homogenates as described previously (17). To evaluate tissue levels of ROS, cryosections were stained with DCFDA (Invitrogen), and positive DCFDA staining in the myocardium was quantified in Matlab (Mathworks Inc.) (17). ATP content was determined using a luminometer (SpectraMax M5; Molecular Devices Inc.), using an assay that included luciferase, which generates a stable luminescent signal proportional to the amount of ATP present (52). ATP production rate was measured kinetically in isolated mitochondria after the addition of 1 mM ADP in the presence of the respiratory substrates glutamate (5 mM) and malate (2 mM), as described previously (53). Cellular lactate was measured with Lactate Assay Kit (Eton Bioscience Inc.). PDH activity was measured in isolated mitochondria as the rate of NADH production at 340 nm, as described previously (54). PDH quantity was determined by a sandwich ELISA using MitoProfile Rapid Microplate Assay Kit (MitoSciences Inc.).
Isolated perfused heart experiments.
The heart was excised from mice and perfused in a Langendorff heart model system paced at 7 Hz with a constant heart rate of 420 bpm and constant pressure of 80 mmHg (55). A water-filled balloon was inserted into the LV to record isovolumic ventricular function. The perfusate contained 118 mM NaCl, 25 mM NaHCO3, 5.3 mM KCl, 2 mM CaCl2, 1.2 mM MgSO4, 0.5 mM EDTA, 5.5 mM glucose, 0.4 mM mixed long-chain fatty acids (bound to 1% albumin), 0.38 mM DL-β-hydroxybutyrate, 1 mM lactate, and 50 μU/ml insulin, equilibrated with 95% O2 and 5% CO2 (pH 7.4). After an equilibration period, the heart was subjected to no-flow global ischemia and reperfused with the same perfusate. For hypoxia experiments, the heart was perfused with the solution gassed with 95% N2 and 5% CO2. Sodium oxamate (50 mM; Sigma-Aldrich), a competitive inhibitor of LDH, was used to inhibit cellular glycolysis during anaerobic perfusion (31).
Coronary artery ligation.
Mice of both genders, 18–20 weeks of age, were anesthetized with pentobarbital sodium (90 mg/kg i.p.) and underwent left anterior descending artery ligation, as described previously (56). Briefly, after opening the thoracic cavity, ligation of the left anterior descending coronary artery was performed with a 7-0 silk suture of a slip knot. The LV was reperfused after 30 minutes, and the ligature was kept in place for later determination of LV area at risk. After a 24-hour recovery period, mice were anesthetized again, and the heart was excised to mount on an isolated heart perfusion apparatus. To measure LV infarct size, the heart was perfused with 2,3,5-triphenyltetrazolium chloride solution. The coronary artery was then reoccluded with the suture left in place at the time of reperfusion and perfused with 0.3% phthalocyanine blue. The LV was sliced into 6 cross-sections, and each section was weighed and photographed. Nonjeopardized, ischemic but viable, and necrotic LV tissues were identified by blue, red, and white coloration, respectively. Planimetry image analysis with ImagePro was used to quantitate the 3 regions in each slice, and the areas of 3 regions from all slices were summed to calculate their respective volumes. All surgeries and subsequent analyses were performed in a blinded randomized fashion with respect to genotype.
Txnip pulldown assay.
Human Txnip coding sequence was subcloned into pCDH-CMV-MCS-EF1-GFP-T2A-Puro (System Biosciences) with a dual Strep/FLAG tag (57). PDHE1α and pyruvate carboxylase sequences were subcloned from commercially available cDNA plasmids (Open Biosystems) into the above-mentioned vector with a HA tag and cotransfected with Txnip or empty control vector into HEK293TN cells using PureFection transfection reagent (System Biosciences). Cells were lysed in 0.5% Triton X-100, 500 mM NaCl, 50 mM Tris, 1 mM phenylmethanesulfonylfluoride, and protease inhibitors (Sigma-Aldrich), pH 7.8. Txnip pulldown was performed using magnetic Streptactin beads (IBA) according to the manufacturer’s instructions. Western analysis of pulled-down proteins was performed with anti-Txnip antibody JY2 (available from MBL International) and anti-HA.11 antibody 16B12 (Covance).
Statistics.
All data are presented as mean ± SEM. Statistical analysis was performed with the unpaired 2-tailed t test between the groups. In PMAGE, P values were calculated by the Audic and Clavarie method, and q values were calculated by the Benjamini-Hochberg or Storey method, as described previously (18). In gene ontology analysis, P values were adjusted for multiple testing by Bonferroni correction. A P value less than 0.05 was considered significant.
Study approval.
Studies in mice were approved by, and animals were maintained in accordance with, the Institutional Animal Care and Use Committees of Harvard Medical School (Harvard Center for Comparative Medicine, Boston, Massachusetts, USA).
Supplementary Material
Acknowledgments
We thank Hayden Huang (Columbia University) and Kenichi Imahashi (National Institute of Environmental Health Sciences) for their technical advice. We also thank Joseph Gannon and Qiuxia Dai for their technical assistance. This work was supported by American Heart Association Scientist Development Grant 0835484N (to J. Yoshioka); Deutsche Forschungsgemeinschaft LE 2728/1-1 (to S. Lee); and NIH grants HL088977 (to W.A. Chutkow), NHLBI268201000036C-0-0-1 for the UTHSCSA Cardiovascular Proteomics Center (to M.L. Lindsey), and HL048743 and HL103582 (to R.T. Lee).
Footnotes
Conflict of interest: The authors have declared that no conflict of interest exists.
Citation for this article: J Clin Invest. 2012;122(1):267–279. doi:10.1172/JCI44927.
Rong Tian’s present address is: University of Washington, Seattle, Washington, USA.
See the related Commentary beginning on page 30.
References
- 1.Smith SC., Jr Multiple risk factors for cardiovascular disease and diabetes mellitus. Am J Med. 2007;120(3 suppl 1):S3–S11. doi: 10.1016/j.amjmed.2007.01.002. [DOI] [PubMed] [Google Scholar]
- 2.Kim JA, Wei Y, Sowers JR. Role of mitochondrial dysfunction in insulin resistance. Circ Res. 2008;102(4):401–414. doi: 10.1161/CIRCRESAHA.107.165472. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Chen Q, Camara AK, Stowe DF, Hoppel CL, Lesnefsky EJ. Modulation of electron transport protects cardiac mitochondria and decreases myocardial injury during ischemia and reperfusion. Am J Physiol Cell Physiol. 2007;292(1):C137–C147. doi: 10.1152/ajpcell.00270.2006. [DOI] [PubMed] [Google Scholar]
- 4.Yellon DM, Hausenloy DJ. Myocardial reperfusion injury. N Engl J Med. 2007;357(11):1121–1135. doi: 10.1056/NEJMra071667. [DOI] [PubMed] [Google Scholar]
- 5.Yoshioka J, Schreiter ER, Lee RT. Role of thioredoxin in cell growth through interactions with signaling molecules. Antioxid Redox Signal. 2006;8(11–12):2143–2151. doi: 10.1089/ars.2006.8.2143. [DOI] [PubMed] [Google Scholar]
- 6.Alvarez CE. On the origins of arrestin and rhodopsin. BMC Evol Biol. 2008;8:222. doi: 10.1186/1471-2148-8-222. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Dutta KK, et al. Two distinct mechanisms for loss of thioredoxin-binding protein-2 in oxidative stress-induced renal carcinogenesis. Lab Invest. 2005;85(6):798–807. doi: 10.1038/labinvest.3700280. [DOI] [PubMed] [Google Scholar]
- 8.Saxena G, Chen J, Shalev A. Intracellular shuttling and mitochondrial function of thioredoxin-interacting protein. J Biol Chem. 2010;285(6):3997–4005. doi: 10.1074/jbc.M109.034421. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Yamanaka H, et al. A possible interaction of thioredoxin with VDUP1 in HeLa cells detected in a yeast two-hybrid system. Biochem Biophys Res Commun. 2000;271(3):796–800. doi: 10.1006/bbrc.2000.2699. [DOI] [PubMed] [Google Scholar]
- 10.Bodnar JS, et al. Positional cloning of the combined hyperlipidemia gene Hyplip1. Nat Genet. 2002;30(1):110–116. doi: 10.1038/ng811. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Oka S, et al. Impaired fatty acid utilization in thioredoxin binding protein-2 (TBP-2)-deficient mice: a unique animal model of Reye syndrome. Faseb J. 2006;20(1):121–123. doi: 10.1096/fj.05-4439fje. [DOI] [PubMed] [Google Scholar]
- 12.Minn AH, Hafele C, Shalev A. Thioredoxin-interacting protein is stimulated by glucose through a carbohydrate response element and induces beta-cell apoptosis. Endocrinology. 2005;146(5):2397–2405. doi: 10.1210/en.2004-1378. [DOI] [PubMed] [Google Scholar]
- 13.Cha-Molstad H, Saxena G, Chen J, Shalev A. Glucose-stimulated expression of TXNIP is mediated by CHREBP, p300 and histone H4 acetylation in pancreatic beta cells. J Biol Chem. 2009;284(25):16898–16905. doi: 10.1074/jbc.M109.010504. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Stoltzman CA, Peterson CW, Breen KT, Muoio DM, Billin AN, Ayer DE. Glucose sensing by MondoA:Mlx complexes: A role for hexokinases and direct regulation of thioredoxin-interacting protein expression. Proc Natl Acad Sci U S A. 2008;105(19):6912–6917. doi: 10.1073/pnas.0712199105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Parikh H, et al. TXNIP regulates peripheral glucose metabolism in humans. PLoS Med. 2007;4(5):e158. doi: 10.1371/journal.pmed.0040158. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Chutkow WA, et al. Deletion of the alpha-arrestin protein Txnip in mice promotes adiposity and adipogenesis while preserving insulin sensitivity. Diabetes. 2010;59(6):1424–1434. doi: 10.2337/db09-1212. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Yoshioka J, et al. Targeted deletion of thioredoxin-interacting protein regulates cardiac dysfunction in response to pressure overload. Circ Res. 2007;101(12):1328–1338. doi: 10.1161/CIRCRESAHA.106.160515. [DOI] [PubMed] [Google Scholar]
- 18.Kim JB, et al. Polony multiplex analysis of gene expression (PMAGE) in mouse hypertrophic cardiomyopathy. Science. 2007;316(5830):1481–1484. doi: 10.1126/science.1137325. [DOI] [PubMed] [Google Scholar]
- 19.Audic S, Claverie JM. The significance of digital gene expression profiles. Genome Res. 1997;7(10):986–995. doi: 10.1101/gr.7.10.986. [DOI] [PubMed] [Google Scholar]
- 20.Hegde PS, White IR, Debouck C. Interplay of transcriptomics and proteomics. Curr Opin Biotechnol. 2003;14(6):647–651. doi: 10.1016/j.copbio.2003.10.006. [DOI] [PubMed] [Google Scholar]
- 21.Nunez C, et al. DNA microarray and proteomic analyses of the RpoS regulon in Geobacter sulfurreducens. J Bacteriol. 2006;188(8):2792–2800. doi: 10.1128/JB.188.8.2792-2800.2006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Javadov S, Karmazyn M, Escobales N. Mitochondrial permeability transition pore opening as a promising therapeutic target in cardiac diseases. . J Pharmacol Exp Ther. 2009;330(3):670–678. doi: 10.1124/jpet.109.153213. [DOI] [PubMed] [Google Scholar]
- 23.Wu Z, et al. Mechanisms controlling mitochondrial biogenesis and respiration through the thermogenic coactivator PGC-1. Cell. 1999;98(1):115–124. doi: 10.1016/S0092-8674(00)80611-X. [DOI] [PubMed] [Google Scholar]
- 24.Jacob WA, Bakker A, Hertsens RC, Biermans W. Mitochondrial matrix granules: their behavior during changing metabolic situations and their relationship to contact sites between inner and outer mitochondrial membranes. Microsc Res Tech. 1994;27(4):307–318. doi: 10.1002/jemt.1070270406. [DOI] [PubMed] [Google Scholar]
- 25.Burwell LS, Brookes PS. Mitochondria as a target for the cardioprotective effects of nitric oxide in ischemia-reperfusion injury. Antioxid Redox Signal. 2008;10(3):579–599. doi: 10.1089/ars.2007.1845. [DOI] [PubMed] [Google Scholar]
- 26.Koitabashi N, et al. Avoidance of transient cardiomyopathy in cardiomyocyte-targeted tamoxifen-induced MerCreMer gene deletion models. Circ Res. 2009;105(1):12–15. doi: 10.1161/CIRCRESAHA.109.198416. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Koitabashi N, et al. Pivotal role of cardiomyocyte TGF-beta signaling in the murine pathological response to sustained pressure overload. J Clin Invest. 2011;121(6):2301–2312. doi: 10.1172/JCI44824. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Yamamoto M, et al. Inhibition of endogenous thioredoxin in the heart increases oxidative stress and cardiac hypertrophy. J Clin Invest. 2003;112(9):1395–1406. doi: 10.1172/JCI17700. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Widder JD, et al. Attenuation of angiotensin II-induced vascular dysfunction and hypertension by overexpression of Thioredoxin 2. Hypertension. 2009;54(2):338–344. doi: 10.1161/HYPERTENSIONAHA.108.127928. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Korge P, Ping P, Weiss JN. Reactive oxygen species production in energized cardiac mitochondria during hypoxia/reoxygenation: modulation by nitric oxide. Circ Res. 2008;103(8):873–880. doi: 10.1161/CIRCRESAHA.108.180869. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Papaconstantinou J, Colowick SP. The role of glycolysis in the growth of tumor cells. II. The effect of oxamic acid on the growth of HeLa cells in tissue culture. J Biol Chem. 1961;236:285–288. [PubMed] [Google Scholar]
- 32.Kelly DP. Hypoxic reprogramming. Nat Genet. 2008;40(2):132–134. doi: 10.1038/ng0208-132. [DOI] [PubMed] [Google Scholar]
- 33.Andres AM, Ratliff EP, Sachithanantham S, Hui ST. Diminished AMPK signaling response to fasting in thioredoxin-interacting protein knockout mice. FEBS Lett. 2011;585(8):1223–1230. doi: 10.1016/j.febslet.2011.03.042. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Xiang G, et al. Catalytic degradation of vitamin D up-regulated protein 1 mRNA enhances cardiomyocyte survival and prevents left ventricular remodeling after myocardial ischemia. J Biol Chem. 2005;280(47):39394–39402. doi: 10.1074/jbc.M502966200. [DOI] [PubMed] [Google Scholar]
- 35.Weissler AM, Kruger FA, Baba N, Scarpelli DG, Leighton RF, Gallimore JK. Role of anaerobic metabolism in the preservation of functional capacity and structure of anoxic myocardium. J Clin Invest. 1968;47(2):403–416. doi: 10.1172/JCI105737. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Drechsel DA, Patel M. Respiration-dependent H2O2 removal in brain mitochondria via the thioredoxin/peroxiredoxin system. J Biol Chem. 2010;285(36):27850–27858. doi: 10.1074/jbc.M110.101196. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Dai S, et al. Endothelial-specific expression of mitochondrial thioredoxin promotes ischemia-mediated arteriogenesis and angiogenesis. Arterioscler Thromb Vasc Biol. 2009;29(4):495–502. doi: 10.1161/ATVBAHA.108.180349. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Matsushima S, et al. Overexpression of mitochondrial peroxiredoxin-3 prevents left ventricular remodeling and failure after myocardial infarction in mice. Circulation. 2006;113(14):1779–1786. doi: 10.1161/CIRCULATIONAHA.105.582239. [DOI] [PubMed] [Google Scholar]
- 39.Nadtochiy SM, Tompkins AJ, Brookes PS. Different mechanisms of mitochondrial proton leak in ischaemia/reperfusion injury and preconditioning: implications for pathology and cardioprotection. Biochem J. 2006;395(3):611–618. doi: 10.1042/BJ20051927. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Turan N, et al. The role of peroxynitrite in chemical preconditioning with 3-nitropropionic acid in rat hearts. Cardiovasc Res. 2006;70(2):384–390. doi: 10.1016/j.cardiores.2005.12.012. [DOI] [PubMed] [Google Scholar]
- 41.Warburg O. On respiratory impairment in cancer cells. Science. 1956;124(3215):269–270. [PubMed] [Google Scholar]
- 42.Damdimopoulos AE, Miranda-Vizuete A, Pelto-Huikko M, Gustafsson JA, Spyrou G. Human mitochondrial thioredoxin. Involvement in mitochondrial membrane potential and cell death. J Biol Chem. 2002;277(36):33249–33257. doi: 10.1074/jbc.M203036200. [DOI] [PubMed] [Google Scholar]
- 43.He M, et al. Identification of thioredoxin-2 as a regulator of the mitochondrial permeability transition. Toxicol Sci. 2008;105(1):44–50. doi: 10.1093/toxsci/kfn116. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Hameyer D, et al. Toxicity of ligand-dependent Cre recombinases and generation of a conditional Cre deleter mouse allowing mosaic recombination in peripheral tissues. Physiol Genomics. 2007;31(1):32–41. doi: 10.1152/physiolgenomics.00019.2007. [DOI] [PubMed] [Google Scholar]
- 45.Dai Q, Escobar GP, Hakala KW, Lambert JM, Weintraub ST, Lindsey ML. The left ventricle proteome differentiates middle-aged and old left ventricles in mice. J Proteome Res. 2008;7(2):756–765. doi: 10.1021/pr700685e. [DOI] [PubMed] [Google Scholar]
- 46.Bonnard C, et al. Mitochondrial dysfunction results from oxidative stress in the skeletal muscle of diet-induced insulin-resistant mice. J Clin Invest. 2008;118(2):789–800. doi: 10.1172/JCI32601. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Imahashi K, Schneider MD, Steenbergen C, Murphy E. Transgenic expression of Bcl-2 modulates energy metabolism, prevents cytosolic acidification during ischemia, and reduces ischemia/reperfusion injury. Circ Res. 2004;95(7):734–741. doi: 10.1161/01.RES.0000143898.67182.4c. [DOI] [PubMed] [Google Scholar]
- 48.Miura S, et al. Overexpression of peroxisome proliferator-activated receptor gamma co-activator-1alpha leads to muscle atrophy with depletion of ATP. Am J Pathol. 2006;169(4):1129–1139. doi: 10.2353/ajpath.2006.060034. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Guo W, Jiang L, Bhasin S, Khan SM, Swerdlow RH. DNA extraction procedures meaningfully influence qPCR-based mtDNA copy number determination. Mitochondrion. 2009;9(4):261–265. doi: 10.1016/j.mito.2009.03.003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Patwari P, Higgins LJ, Chutkow WA, Yoshioka J, Lee RT. The interaction of thioredoxin with Txnip. Evidence for formation of a mixed disulfide by disulfide exchange. J Biol Chem. 2006;281(31):21884–21891. doi: 10.1074/jbc.M600427200. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Miranda-Vizuete A, Damdimopoulos AE, Gustafsson J, Spyrou G. Cloning, expression, and characterization of a novel Escherichia coli thioredoxin. . J Biol Chem. 1997;272(49):30841–30847. doi: 10.1074/jbc.272.49.30841. [DOI] [PubMed] [Google Scholar]
- 52.Drew B, et al. Effects of aging and caloric restriction on mitochondrial energy production in gastrocnemius muscle and heart. Am J Physiol Regul Integr Comp Physiol. 2003;284(2):R474–R480. doi: 10.1152/ajpregu.00455.2002. [DOI] [PubMed] [Google Scholar]
- 53.Manfredi G, Yang L, Gajewski CD, Mattiazzi M. Measurements of ATP in mammalian cells. Methods. 2002;26(4):317–326. doi: 10.1016/S1046-2023(02)00037-3. [DOI] [PubMed] [Google Scholar]
- 54.Churchill EN, Murriel CL, Chen CH, Mochly-Rosen D, Szweda LI. Reperfusion-induced translocation of deltaPKC to cardiac mitochondria prevents pyruvate dehydrogenase reactivation. Circ Res. 2005;97(1):78–85. doi: 10.1161/01.RES.0000173896.32522.6e. [DOI] [PubMed] [Google Scholar]
- 55.Luptak I, Yan J, Cui L, Jain M, Liao R, Tian R. Long-term effects of increased glucose entry on mouse hearts during normal aging and ischemic stress. Circulation. 2007;116(8):901–909. doi: 10.1161/CIRCULATIONAHA.107.691253. [DOI] [PubMed] [Google Scholar]
- 56.Weinberg EO, et al. Rosuvastatin reduces experimental left ventricular infarct size after ischemia-reperfusion injury but not total coronary occlusion. Am J Physiol Heart Circ Physiol. 2005;288(4):H1802–H1809. doi: 10.1152/ajpheart.00962.2004. [DOI] [PubMed] [Google Scholar]
- 57.Gloeckner CJ, Boldt K, Schumacher A, Roepman R, Ueffing M. A novel tandem affinity purification strategy for the efficient isolation and characterisation of native protein complexes. Proteomics. 2007;7(23):4228–4234. doi: 10.1002/pmic.200700038. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.