Abstract
The VERNALIZATION1 (VRN1) gene of temperate cereals is transcriptionally activated by prolonged cold during winter (vernalization) to promote flowering. To investigate the mechanisms controlling induction of VRN1 by prolonged cold, different regions of the VRN1 gene were fused to the GREEN FLUORESCENT PROTEIN (GFP) reporter and expression of the resulting gene constructs was assayed in transgenic barley (Hordeum vulgare). A 2 kb segment of the promoter of VRN1 was sufficient for GFP expression in the leaves and shoot apex of transgenic barley plants. Fluorescence increased at the shoot apex prior to inflorescence initiation and was subsequently maintained in the developing inflorescence. The promoter was also sufficient for low-temperature induction of GFP expression. A naturally occurring insertion in the proximal promoter, which is associated with elevated VRN1 expression and early flowering in some spring wheats, did not abolish induction of VRN1 transcription by prolonged cold, however. A translational fusion of the promoter and transcribed regions of VRN1 to GFP, VRN1::GFP, was localised to nuclei of cells at the shoot apex of transgenic barley plants. The distribution of VRN1::GFP at the shoot apex was similar to the expression pattern of the VRN1 promoter-GFP reporter gene. Fluorescence from the VRN1::GFP fusion protein increased in the developing leaves after prolonged cold treatment. These observations suggest that the promoter of VRN1 is targeted by mechanisms that trigger vernalization-induced flowering in economically important temperate cereal crops.
Introduction
Plants growing in temperate regions time flowering to coincide with favourable seasonal conditions. Winter frost can damage cold sensitive reproductive organs, while heat and water stress during summer can reduce fertility, so in temperate regions many plants flower in spring when conditions are optimal. One cue that promotes spring flowering is prolonged exposure to winter cold - vernalization [1]. Vernalization-induced flowering is a feature of many plants, including Arabidopsis and economically important temperate cereal crops, such as wheat (Triticum spp.) and barley (Hordeum vulgare), although different genes mediate this seasonal flowering response in these distantly related angiosperms (see [2]).
The VERNALIZATION1 gene (VRN1) is a central regulator of vernalization-induced flowering in temperate cereals (see [2]–[5]). VRN1 encodes a MADS box transcription factor that promotes flowering [6]–[9]. VRN1 is transcribed at low basal levels but transcript abundance increases with prolonged cold treatment [6]–[9]. The response of VRN1 expression to cold is quantitative, with longer cold treatments inducing higher transcript levels [6], [7], [10], [11]. This parallels the degree to which flowering is accelerated [11]. Mutants that lack VRN1, and flanking genes, are unable to flower [12], [13]. VRN1-like genes are likely to play similar roles in other temperate grasses, including economically important species such as Lolium perenne and Phleum pratense [14], [15].
Vernalization activates expression of VRN1 in the leaf and shoot apex. At the shoot apex expression of VRN1 promotes the transition to reproductive development [11], [16], [17]. Expression of VRN1 in leaves unlocks the long-day flowering response, allowing long daylengths to further accelerate reproductive development post-vernalization [3], [16]. Molecular analyses have identified potential regulatory targets of VRN1. These include the grass/cereal specific transcription factors VRN2 and ODDSOC2 [18]–[21].
Alleles of VRN1 that are expressed without cold treatment allow flowering without vernalization [6]–[8]. These “active alleles” have been used to breed wheats and barleys that flower without vernalization (spring types), which are grown where vernalization does not occur. Some active alleles of the wheat VRN1 gene have mutations near the transcriptional start site, which might disrupt promoter sequences required to repress transcription prior to winter [8], [22], [23]. Additionally, the first intron of VRN1 contains a broad region that is required to maintain repression of VRN1 prior to winter [10], [24]–[27]. Alleles lacking large sections of the first intron are actively expressed and are associated with early flowering without vernalization, whereas some alleles lack smaller segments of the first intron and are associated with moderate increases in VRN1 activity and weaker promotion of flowering [26], [27]. An insertion of a mobile genetic element at the 5′ end of the first intron is also associated with active expression of VRN1 without vernalization [28], so the repressive action of the first intron is not related simply to size.
The state of chromatin at the VRN1 locus appears to be an important determinant of activity [29]. Without vernalization, the chromatin at VRN1 has high levels of the repressive histone modification histone 3 lysine 27 tri-methylation (H3K27Me3), which is typically associated with an inactive chromatin state [29]. This modification is found within the first intron of the VRN1 gene and at the start point of transcription, sites that are critical for repression [29]. The presence of H3K27Me3 at these sites might contribute to repression of VRN1 prior to winter [29].
The mechanisms that activate expression of VRN1 in response to prolonged cold are unclear. Although the floral repressor VRN2 is required to delay flowering prior to vernalization [18], cold induction of VRN1 takes place in conditions where VRN2 is not actively expressed and can occur in the absence of the VRN2 gene [11], [16], [19]. So VRN2 seems unlikely to mediate cold-induction of VRN1. The hypothesis that the VEGETATIVE TO REPRODUCTIVE TRANSITION 2 gene is repressed by cold to allow increased expression of VRN1 [30] also seems unlikely, since VRT2 expression increases at low temperatures [31]. Furthermore, although VRT2 binds to a sequence motif at the promoter of VRN1 in vitro, this motif is not critical for low-temperature induction [23]. Finally, alleles of VRN1 that lack most of the first intron, which are expressed at high basal levels, are induced by cold [27]. This suggests that cold activation might occur independently of the repressive mechanism that acts at the first intron [27].
Studies in Arabidopsis, and other plants, have demonstrated that activation of transcription by low temperatures can be mediated by cis-acting elements located in the promoters of low-temperature responsive genes [32]. In this study, we examine whether the promoter of the VRN1 gene can mediate a transcriptional response to prolonged cold, using transgenic barley as a model system.
Results
A VRN1 promoter reporter gene fusion is expressed in the leaves and shoot apex of transgenic barley plants
A 2 kb fragment from the promoter of the barley (Hordeum vulgare) VERNALIZATION1 gene (referred to hereafter as VRN1) was fused to the GFP reporter gene (Figure 1A) and the resulting construct (PVRN1:GFP) was transformed into barley (cv. Golden Promise). GFP activity was detected in the shoot apex and developing leaves of transgenic barley plants that carry the PVRN1:GFP construct (Figure 2). GFP fluorescence increased at the shoot apex at the late vegetative stage (Figure 2B) and remained active throughout the shoot apex during reproductive development, although expression was lower in the developing florets than in other parts of the inflorescence (Figure 2D). GFP fluorescence was also observed in leaves (Figure 2E and 2F).
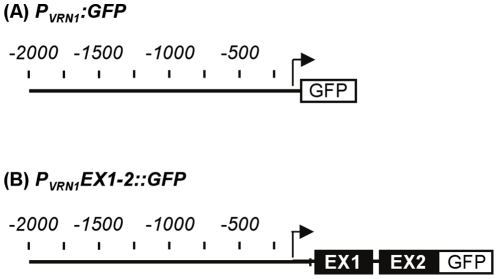
Figure 1. Schematic representation of VRN1reporter gene constructs.
(A) A 2.0 kb fragment (SpeI-NcoI) from the VRN1 promoter was fused to the GFP reporter gene with the NOPALINE SYNTHASE (NOS) terminator sequence to generate the PVRN1:GFP construct. (B) The PVRN1EX1-2:GFP construct was generated by fusing the same promoter region to exon1 and exon 2 of VRN1 (EX1 and EX2), separated by a 0.35 kb segment of intron 1, followed by the GFP-NOS cassette. Scale indicates base pairs from the translational start site of MADS box open reading frame, indicated as negative relative to the coding regions of the VRN1 locus. Arrow indicates transcriptional start site, which did not vary between control or prolonged cold treatments.
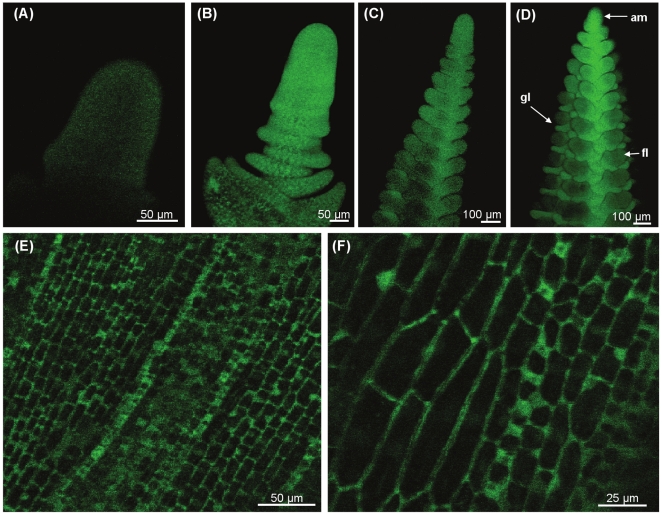
Figure 2. GFP fluorescence in the leaves and shoot apices of transgenic plants with the PVRN1:GFP construct.
(A) GFP fluorescence in the vegetative shoot apex of 12 day old transgenic plants carrying the PVRN1:GFP construct. (B) The shoot apex at the late vegetative stage of 18 day old plants. (C,D) Shoot apices at different stages of reproductive development. (E,F) Fluorescence in the cytoplasm of cells in the developing leaves of 18 day old plants. am indicates the apical meristem, fl indicates the developing florets, gl indicates glume primordia.
The promoter of VRN1 is sufficient to mediate transcriptional induction of the GFP reporter gene by prolonged cold
The levels of GFP transcript generated from the PVRN1:GFP construct were assayed by quantitative reverse transcriptase PCR (qRT-PCR) in transgenic seedlings exposed to prolonged cold treatment (28 days at 4°). Expression was assayed in three independent transgenic lines. Transcript levels were higher in seedlings at the end of prolonged cold treatment than in control seedlings grown to an identical stage of development at normal temperatures (4 days at 20°) (Figure 3A). The degree of induction by cold varied between 1.5 to 4 fold, depending on the transgenic line assayed.
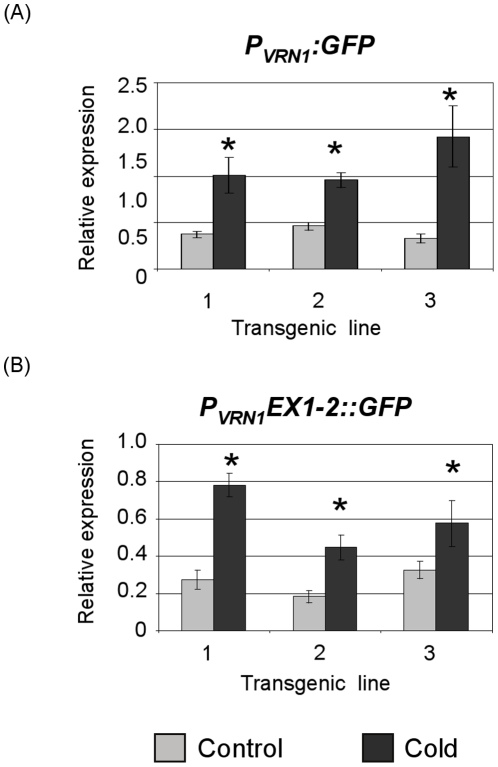
Figure 3. Expression of the PVRN1:GFP and PVRN1EX1-2::GFP constructs in control versus cold treated seedlings.
(A) GFP transcript levels in transgenic seedlings carrying the PVRN1:GFP construct. Expression was assayed by quantitative reverse transcriptase PCR in control seedlings, germinated in darkness at normal glasshouse temperatures (20 degrees for 5 days), compared to seedlings germinated and grown in darkness to an identical stage of development at low temperatures (4 degrees for 28 days). Expression levels were assayed in three independent transgenic lines. (B) Expression of the PVRN1EX1-2::GFP, assayed as outlined above, in three independent transgenic lines. Expression is shown relative to ACTIN. Error bars show standard error for a minimum of 3 biological replicates. * indicates P<0.05.
A second reporter gene fusion was constructed by fusing the same 2 kb region of the VRN1 promoter to the GFP reporter gene, with the addition of the first and second exons from the VRN1 gene, separated by a small segment (0.35 kb) of the first intron (PVRN1EX1-2::GFP, Figure 1B ). The spliced version of this translational fusion construct was expressed in transgenic barley plants, although no GFP fluorescence was detected, possibly due to poor translation or instability of the fusion protein produced. Levels of the spliced PVRN1EX1-2::GFP transcript increased in response to prolonged cold treatment (Figure 3B), similar to the PVRN1:GFP construct.
Cellular and tissue localisation of a VRN1::GFP translational fusion in transgenic barley
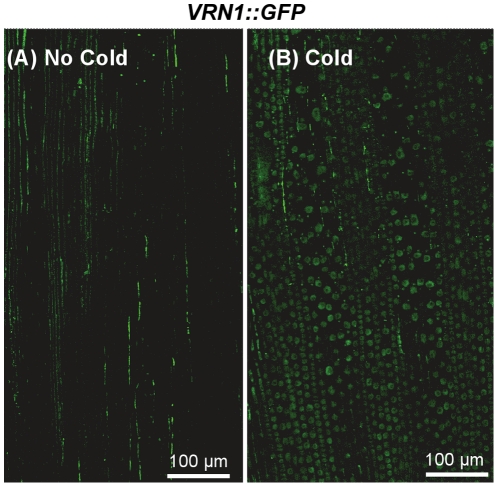
The VRN1 gene, the promoter and transcribed regions minus the majority of the large first intron (1.1 kb of the 10.7 kb intron was present in the construct), was translationally fused to the GFP reporter gene followed by the 3′ UTR of the VRN1 gene. The resulting VRN1::GFP construct (Figure 4A) was transformed into barley. GFP fluorescence signal was observed in the nuclei of cells (Figure 4). The nuclear VRN1::GFP fluorescence increased during vegetative shoot apex development and was higher in the pre-double ridge and double ridge apices than in the early vegetative shoot apex (Figure 4). At later stages of development, the VRN1::GFP signal was detected throughout the shoot apex, although expression was lower in the developing florets than in other parts of the inflorescence, similar to the PVRN1:GFP construct (Figure 4H). VRN1::GFP transcript levels increased in seedlings during prolonged cold treatment, although induction was weaker than that observed in the PVRN1:GFP lines (Figure S1). Whereas GFP signal was weak in the developing leaves of seedlings grown without cold treatment, strong fluorescence was observed in the nuclei in cells from the developing leaves of seedlings that had experienced prolonged cold (Figure 5).
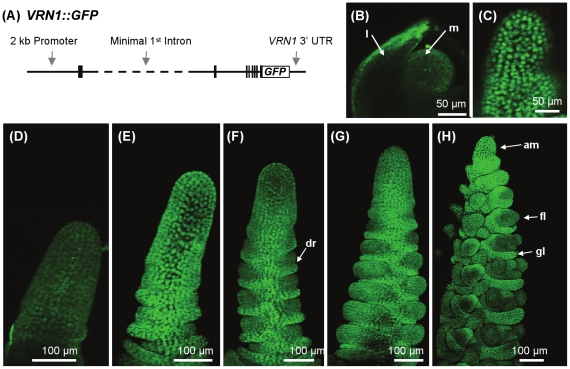
Figure 4. Activity of a VRN1::GFP fusion protein in transgenic barley plants.
(A) Schematic representation of a VRN1::GFP fusion construct that fuses the entire VRN1 gene, minus most of the first intron, to the GFP reporter gene followed by the 3′ UTR and terminator sequence of the VRN1 gene. Vertical bars represent exons, dotted line represents the missing segment from intron 1, relative to the wildtype version of VRN1 from vernalization responsive barleys (B) Low signal from the VRN1::GFP fusion in the vegetative shoot apical meristem (m) and developing leaf (l) of 7 day old seedlings. (C) Nuclear localisation of the VRN1::GFP fusion protein in the meristem of a reproductive shoot apex. (D–H) Time course of shoot apex development showing vegetative shoot apices (D,E), a shoot apex at the transition to reproductive development (F), indicated by double ridges (dr), and reproductive shoot apices (G,H). am indicates the apical meristem, fl indicates the developing florets, gl indicates glume primordia. Images B–H were taken on identical settings to allow direct comparison of fluorescence levels. (note: image C is a section from image H, 2 fold expanded).
Figure 5. Activity of a VRN1::GFP fusion protein in the developing leaves of transgenic barley plants.
(A) Low signal from the VRN1::GFP fusion in the developing leaves of plants grown at normal glasshouse temperatures. (B) Nuclear localisation of the VRN1::GFP fusion protein in cells within the developing leaves of seedlings germinated and grown at low temperatures to an identical stage of development. Images were taken with identical settings to allow comparison of fluorescence levels. Similar results were seen in two independent transgenic lines. Some background signals, “green lines” caused by reflection, are present in both images.
Cold response motifs and a small upstream open reading frame are found in the promoter and 5′ untranslated region of the VERNALIZATION1 gene
The promoter of VRN1 was scanned for potential cis-acting elements, by searching for sequence motifs similar to known transcription factor binding sites. This identified putative binding sites for C-REPEAT BINDING FACTOR (CBF) and INDUCER OF CBF1 (ICE1) transcription factors within the 2 kb of sequence upstream of the predicted translational start site at the VRN1 gene (Figure 6A, Figure S2). These transcription factors have been shown to activate the expression of cold-responsive genes in Arabidopsis, and other plants [32]. Binding sites for B-ZIP and ethylene responsive transcription factors were also identified.
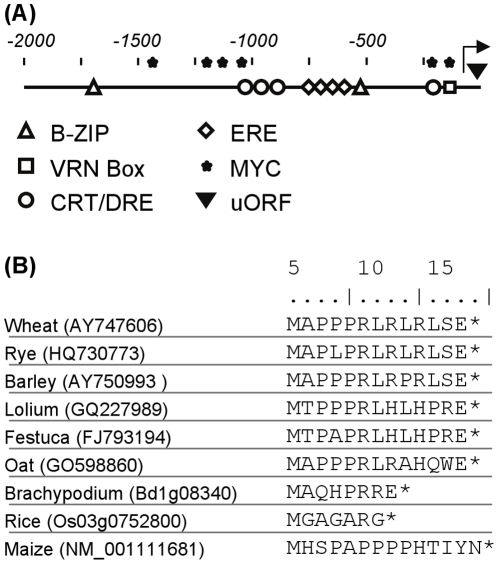
Figure 6. Potential regulatory motifs at the promoter of the VRN1 locus.
(A) Schematic representation of potential transcription factor binding sites at the promoter of VRN1. ERE indicates potential ethylene response element (GCCGCC), CRT/DRE: C-repeat transcription factor core binding site (CCGAC), MYC: MYC transcription factor binding site (CANNTG), B-ZIP: B-ZIP transcription factor binding site (ACGT). VRN Box: indicates the position of the putative vernalization regulatory motif suggested by Pidal et al. [23]. uORF denotes the position of the small upstream open reading frame. Scale indicates bp from transcriptional start site, which is indicated by an arrow. (B) Alignment of the predicted amino acid sequences of the upstream open reading frame from the 5′ untranslated regions of VRN1 genes from temperate cereals (wheat, rye, barley, oats), temperate grasses (Lolium, Festuca, Brachypodium) and VRN1 orthologues from warm climate cereals (rice and maize). Scale indicates amino acid residues from initiation codon.
Another potential regulatory element identified at the VRN1 locus is a small upstream open reading frame (uORF) encoding 13 amino acids (MAPPPRLRPRLSE) in the predicted 5′ leader sequence of the VRN1 mRNA (Figure 6B). As predicted, the uORF is found in the 5′ end of VRN1 cDNA sequences in GENBANK, and this was further confirmed by 5′ RACE (Rapid Amplification of cDNA ends). The transcriptional start site, which did not vary between vernalized versus non-vernalized plants is annotated in Figure S2. The presence of a proline rich small open reading frame is a conserved feature of the VRN1 genes of barley and wheat. uORFs are also a feature of VRN1-like genes from other cereals and grasses, upstream of the main open reading frame that encodes the AP1/FRUITFULL-like MADS box transcription factor (Figure 6B). Small uORFs can influence translation from downstream ORFs by decreasing translational initiation rates or by stalling ribosomes and causing non-sense mediated message decay (see [33]).
An insertion in the VRN box does not abolish induction of VRN1 by prolonged cold
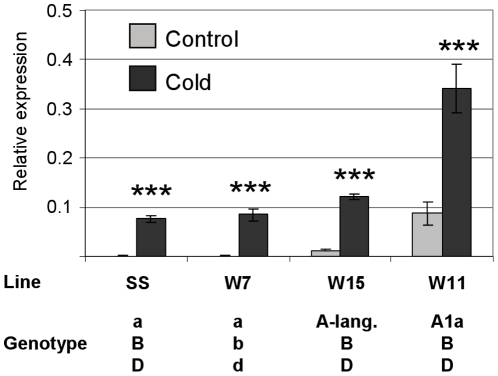
Mutations in a proximal promoter region defined as the VRN box are associated with elevated VRN1 expression and early flowering in wheat (Pidal et al. 2009). To examine whether mutations in the VRN box influence the capacity for low-temperature induction of VRN1, transcriptional induction of VRN1 by prolonged low-temperature treatment was examined in near-isogenic wheat lines with different alleles of the VRN1 gene: a wildtype allele on the A genome (normal VRN box, full length first intron, parental line) versus alleles with either a 7,222 bp deletion within the first intron (line W15) or a 231 bp insertion in the VRN box on the A genome (line W11). Basal expression from the VRN1 gene on the A genome (VRN-A1) was highest in the W11 line carrying the promoter insertion allele (Figure 7), and was also elevated in the W15 line carrying the intron deletion allele (Students T-tests: P<0.05, P<0.01 versus the parental line with the wildtype allele, for the promoter insertion or intron deletion alleles respectively). Low-temperature induction of the A genome copy of VRN1 occurred irrespective of VRN1 genotype (Figure 7). Thus, mutations that are associated with increased basal expression of the VRN1 gene copy on the A genome (homeoallele) do not prevent low-temperature induction.
Figure 7. Transcript levels for the VRN1 gene (A genome) of wheat near-isogenic lines with different VRN1 genotypes.
Transcript levels of the A genome copy of VRN1 were assayed in hexaploid bread wheat (Triticum aestivum) near-isogenic lines that vary for VRN1 genotype. Expression was assayed by quantitative reverse transcriptase PCR, using A genome specific primers, in seedlings germinated in darkness at 20° for 5 days (control) or 4° for 6 weeks (cold). The wheat lines used carry different combinations of wildtype VRN1 alleles on each of the three genomes (a, b or d) or alleles with high basal activity (A1a, A-lang., B or D). The A1a allele (line W11) has a promoter insertion in the VRN box of the VRN1 gene on the A genome, but a full-length first intron [22]. The A-lang. allele (line W15), first identified in the Langdon cultivar of the tetraploid wheat Triticum durum [24], has a 7.2 kb deletion within the first intron, but has the wildtype promoter sequence. Similarly, the B and D alleles have deletions within the first intron of the VRN1 genes on the B and D genomes respectively [24]. Expression is shown relative to ACTIN, error bars show standard error for four biological replicates, *** indicates P<0.001.
Discussion
The 2 kb of DNA sequence upstream of the primary translational start site (MADS box open reading frame) of the VRN1 gene is sufficient to drive expression of the GFP reporter gene in the shoot apex and leaves; organs where VRN1 expression has been detected using quantitative RT-PCR [8], [11], [17]. The distribution of PVRN1:GFP expression observed in shoot apices and leaves of barley (Figure 2) is similar to the expression patterns of the VRN1 genes of wheat (T. aestivum, T. monococcum) and oats (Avena sativa) detected by in situ hybridisation [17], [30]. Similarly, at later stages of barley inflorescence development (Figure 2D), the expression pattern of the PVRN1:GFP construct is consistent with the results of in situ hybridisation analysis of VRN1 expression, which show expression of VRN1 in meristems and organ primordia of the developing barley spike [34].
Expression of the PVRN1:GFP and VRN1::GFP reporter genes increased at the shoot apex shortly before the transition to reproductive development (inflorescence initiation). This is consistent with the observation that VRN1 transcript levels increase at the shoot apex of this barley cultivar prior to inflorescence initiation, when assayed by quantitative RT-PCR [31]. Similarly, when the promoter of the maize (Zea mays) orthologue of VRN1 (ZmMADS4) was fused to the β-GLUCURONIDASE reporter gene an increase in reporter activity occurs at the equivalent stage of development in transgenic maize plants [35]. This expression pattern was confirmed by in situ hybridisation [35]. Thus, the expression patterns of the VRN1 promoter reporter genes used in this study are similar to those of the endogenous VRN1 genes from a range of cereals.
The increase in VRN1 expression prior to inflorescence initiation and the distribution of VRN1 expression in the developing inflorescence are consistent with a role for VRN1 in promoting inflorescence meristem or organ identity. The lower VRN1 activity seen in the developing florets (Figure 2D) suggests that less VRN1 activity is required for the differentiation of floral organs. Other VRN1-like genes, Barley MADS3 (BM3) and BM8, might contribute to the development of these organs, compensating for lower VRN1 activity [34].
In addition to driving basal expression of a reporter gene in the shoot apex and developing leaves, a 2 kb segment from the promoter of VRN1 was sufficient for induction of reporter gene expression by prolonged cold. Based on these observations, and those of previous studies [26]–[29], we suggest that the vernalization response of cereals depends largely upon two antagonistic regulatory mechanisms that control transcriptional activity of VRN1: activation of transcription by the promoter, which has a low basal activity but can be activated by cold, counteracted by constitutive repression through the first intron of the gene (Figure 8).
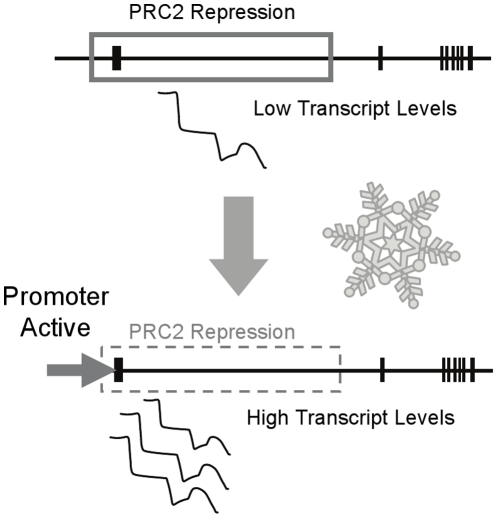
Figure 8. A model for transcriptional regulation of VRN1.
Prior to winter, chromatin at the VRN1 locus is maintained in an inactive state by histone modifications deposited by a plant Polycomb Repressor Complex 2 (PRC2), resulting in low transcript levels. When plants are exposed to prolonged cold (snowflake) the promoter of VRN1 becomes more active, leading to increased transcription and higher steady state transcript levels. This triggers a change in the state of chromatin at the VRN1 locus, with a shift towards an active state.
A key question is what mechanisms act at the promoter to mediate cold induction? The barley cultivar used in this study lacks VRN2, ruling out involvement of this gene. Similarly, the first intron is not present in the PVRN1:GFP reporter gene fusion, which is low-temperature responsive (Figure 3A), consistent with previous suggestions that the intron is not critical for cold induction [27], [31]. The VRN1 promoter contains the putative VRN box, a region mutated in some active VRN1 alleles of wheat [23]. An insertion in this promoter element (the Vrn-A1a allele described by Fu et al. [24]) did not disrupt low-temperature induction of VRN1 (Figure 7), however, suggesting that while this region can influence basal expression of VRN1 it is not critical for low-temperature induction. Presumably mutations in this region affect a transcriptional repression mechanism that acts separately to any mechanisms that mediate low-temperature induction.
The promoter of VRN1 contains sequence motifs that might be targeted by low-temperature responsive transcription factors, including putative CBF and myc recognition sites (Figure 6). Binding of low-temperature responsive transcription factors, such as CBFs or the myc INDUCER OF CBF EXPRESSION1 (ICE1), to these putative binding sites might play a role in inducing expression of VRN1 during vernalization. The promoter of the barley VRN1 gene also contains binding sites for B-ZIP transcription factors that might mediate long-day activation of VRN1 in vernalized plants or in some early flowering genotypes [36]. The potential role of these putative regulatory elements will be examined in future studies.
Another potential regulatory mechanism identified in the 5′ region of VRN1 is a small upstream open reading frame (uORF). uORFs are widespread in mammalian genes and typically reduce protein expression from larger downstream open reading frames by 30–80% [37]. In humans, the presence of a uORFs in a gene can vary between individuals, causing variation in protein expression from downstream open reading frames and this is the basis for some diseases [37]. The uORF in the 5′ of the VRN1 gene is conserved amongst VRN1 orthologues in temperate cereals and related grasses (Figure 6B). The uORF was not required for the transcriptional response to prolonged cold, since it was not included in the low-temperature responsive PVRN1:GFP construct, but could potentially influence post-transcriptional regulation of the endogenous VRN1 gene. In some early flowering accessions of the wild wheat T. timopheevi the uORF is deleted (Figure S3, [38]), although it is unclear whether this contributes to natural variation in flowering behaviour of this wild wheat.
In summary, we have shown that the promoter of VRN1 is able to drive basal expression of the GFP reporter gene in the leaves and shoot apex of transgenic barley plants and is sufficient for induction of reporter gene expression by prolonged cold. Understanding the mechanisms that act at the promoter of VRN1 to mediate the transcriptional response to prolonged cold of this central regulator of flowering can provide further insights into how vernalization induces spring flowering of economically important cereal crops.
Materials and Methods
Construction of VRN1-reporter gene fusion constructs
The PVRN1:GFP construct was generated by sub-cloning a 2 kb SpeI-NcoI VRN1 promoter fragment from the Morex BAC clone 631P8 and ligating this to a GFP-nopaline synthase terminator cassette. The resulting promoter-reporter gene cassette was inserted into the pVEC8 T-DNA vector [39]. The PVRN1EX1-2::GFP construct was generated by first amplifying the promoter, first exon and a small segment of the first intron (5′-CACACCGCTCTACTAGTTC-3′ and 5′-GGCGCGCCCTGCAGGGAGCGCCAGCTCCGCC-3′) of VRN1 from a winter barley (cv. Sonja). This was ligated to a second fragment from inside the first intron into the second exon (5′-GGCGCGCCCGATTATCAAAATGTACCA-3′ and 5′-CCCGGGTCACTTGAAACGAGAACC-3′), to generate a construct with 2 kb of promoter sequence, with the first and second (partial) exons separated by a 0.35 kb segment of the first intron. The resulting fragment was ligated to the GFP-nopaline synthase terminator, and inserted into pVEC8. The VRN1::GFP fusion was generated by fusing the 3′ UTR of VRN1, amplified from the Morex BAC 631P8 (5′-TTTCTAGACAGCCCATGTAAGCGTACTATTCAG-3′ and 5′-TTACTAGTAATGGCAGGTGTTCTGTTTGTTTATG-3′), to GFP. The GFP-UTR fragment was then ligated to a fragment amplified from the Morex BAC 631P8, from within the first intron to the last exon, minus the stop codon (5′-TTTCTAGACAGCCCATGTAAGCGTACTATTCAG-3′ and 5′-TTACTAGTAATGGCAGGTGTTCTGTTTGTTTATG-3′), with the open reading frames of VRN1 and GFP in frame. Primers included compatible restriction sites to facilitate cloning. The resulting VRN1-GFP-UTR fragment was then ligated to the SpeI-NotI promoter fragment and the resulting reporter gene fusion was inserted into the pVEC8 T-DNA vector. The resulting VRN1::GFP fusion lacks most of the first intron (bp 2557 to 12214 relative to the Strider VRN1 sequence) similar to the HvVRN1-3 allele (bp 2968 to 11687).
Barley Transformation
Barley transformation was performed by Agrobacterium transformation of excised embryos [40], [41], with cv. Golden Promise, which flowers without vernalization and is daylength insensitive (genotype HvVRN1-1, ΔHvVRN2, ppd-H1). Expression analysis was performed with plants from the T1 generation. Plants were screened for the presence of transgenes using primers that amplify the hygromycin selectable marker gene (5′-AAAAGCCTGAACTCACCGC-3′ and 5′-TCGTCCATCACAGTTTGCC-3′).
Gene expression analysis
Seeds were germinated in soil and grown in darkness in foil covered pots at 20 degrees or 4 degrees. RNA was extracted from seedlings with Spectrum Plant Total RNA Kit (Sigma Aldrich, www.sigma-aldrich.com) following manufacturer's instructions. RNA was treated with 10 units of RQ1 RNase-free DNase (Promega, Madison, WI) for 15 min at 37°C. Total RNA (5 µg) was reverse-transcribed with Super Script III reverse transcriptase (Invitrogen, www.invitrogen.com), according to manufacturer instructions. qRT-PCR was performed in a Rotorgene Q real-time PCR cycler (Qiagen; www.qiagen.com) as described previously [20].
The following primer sets were used for quantitative RT-PCR: HvVRN1: 5′-GGAAACTGAAGGCGAAGGTTGA and 5′-TGGTTCTTCCTGGCTCTGATATGTT-3′, PVRN1:GFP 5′-TACAACTACAACAGCCACAAC-3′ and 5′-GCTTCTCGTTGGGGTCTTTG-3′, PVRN1EX1-2::GFP 5′-AGAACAAGATCAACCGCCA-3′ and 5′-ATCGCCCTCGCCCTCGCCGG-3′, VRN1::GFP 5′-CGCAGATACCAGCAATCACCCAG-3′ and 5′-ATCGCCCTCGCCCTCGCCGG-3′. For quantification of expression of the A genome copy of the VRN1 gene from bread wheat the homeoallele specific primers 5′-CAGCCTGGTGTATGTTGCGGTTGC-3′ and 5′-ATTACTCGTACAGCCATCTCAGCC-3′ were used. The expression of each gene was normalized to ACTIN using the Rotorgene software package (Qiagen, www.qiagen.com), which takes amplification efficiency of each primer set into account for quantification calculations. ACTIN primers have been described previously [19]. All data presented are the average of mRNA levels from three biological repeats, unless stated otherwise, with error bars representing the standard error of the mean.
Rapid Amplification of cDNA ends (RACE)
5′ RACE was carried out with the FirstChoice RLM-RACE kit from Ambion (Ambion; www.invitrogen.com/ambion) following manufacturer′s instructions with total RNA from Golden Promise seedlings grown under control conditions or Golden Promise seedlings grown to an identical stage of development at 4 degrees (28 days). Nested PCR was performed with the 5′RACE Outer Primer provided by the kit and the gene specific VRN1-R Outer primer (5′-GCATTCGTGCATAAGTTGGTTC-3′) and for the Inner 5′ RLM-RACE PCR we used the 5′RACE Inner Primer and the gene specific VRN1-R Inner primer (5′-TATTGTCTCAACCTTCGCCTTC-3′). PCR products were sequenced after ethanol precipitation.
Confocal microscopy
Barley apices and leaf primordia were dissected under a binocular dissecting microscope and mounted in water. The plant material was then observed in Leica SP2 confocal laser scanning microscope (Leica Microsystems, Sydney, Australia) equipped with a standard Ar 488 laser excitation. Excitation was at 488 nm, and emission was collected between 500 and 550 nm. Chlorophyll autofluorescence was collected between 650 and 720 nm. Nuclear localisation was confirmed by co-localization with 4′,6-diamidino-2-phenylindole (DAPI) stain (Figure S4). Spikes were fixed in 4% paraformaldehyde in 50 mM phosphate buffer, pH 7.2, overnight, then rinsed in buffer and stained with 5 ug/ml DAPI for 30 sec. After mounting in fresh buffer, GFP was detected with 488 nm excitation, and emission from 505–550 nm collected. DAPI was then detected with 405 nm excitation, and emission from 415–485 nm collected using a Leica SP2 confocal microscope in sequential scanning mode to avoid bleedthrough of DAPI fluorescence into the GFP emission channel.
Production of near-isogenic wheat lines
Wheats carrying different VRN1 alleles were crossed to the Australian wheat cultivar Sunstate (PPD-D1 insensitive allele, VRN-A1 wildtype, VRN1-B1 intron deletion and VRN-D1 intron deletion) and then backcrossed a further 4 generations before homozygous plants were selected for contrasting VRN1 genotypes. The donors of different VRN1 alleles were: AUS7374 for Langdon VRN1 A genome allele with an intron deletion, AUS 2380 “Extra Early Blackhull” for recessive B and D genome alleles or AUS1499 for the VRN-A1a promoter insertion allele.
Supporting Information
Low-temperature induction of the VRN1::GFP construct.Transcript levels for the VRN1::GFP fusion in transgenic seedlings. Expression was assayed in control seedlings, germinated at normal glasshouse temperatures (20 degrees for 4 days), and compared to seedlings germinated and grown to an identical stage of development at low temperatures (4 degrees for 28 days). Expression levels were assayed in two independent transgenic lines. Error bars show standard error. * indicates P<0.05.
(TIF)
Sequence motifs in the VRN1 promoter.The promoter sequence of the barley VRN1 gene showing potential transcription factor recognition sites and other putative regulatory sequences. The transcriptional start site, identified by 5′RACE is also indicated (TSS); this did not vary with different temperature treatments.
(DOC)
Deletion of the 5′ small open reading frame from the VRN1 gene of a wild wheat.Comparison of the A genome VRN1 gene from hexaploid wheat (VRN-A1, Genbank AY747600.1) with the sequence of the VRN1 gene from Triticum timopheevi (Genbank GQ451763). The VRN box is shaded grey. The putative CARG box is shaded black and the region that encodes the small upstream open reading frame is shown in bold text and boxed.
(DOC)
Co-localisation of DAPI staining and VRN1::GFP within cells of the developing inflorescence.Localization of DAPI staining compared to VRN1::GFP signal in the developing glume of a barley inflorescence. Double headed arrow shows DAPI and GFP signal in the same nucleus.
(TIF)
Acknowledgments
We thank Sandra Stops and Sarah Fieg for expert technical assistance. We also thank Dr Rosemary White for assistance with confocal microscopy and DAPI staining of barley tissues.
Footnotes
Competing Interests: The authors have declared that no competing interests exist.
Funding: This research was funded by a Commonwealth Scientific and Industrial Research Organisation Office of the Chief Executive postdoctoral fellowship to SNO, then MMA-P. Near-isogenic wheat lines were developed in the project CSP00131, funded by the Australian Grains Research and Development Corporation. The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
References
- 1.Chouard P. Vernalization and its relation to dormancy. Ann Rev Plant Physiol. 1960;1:191–238. [Google Scholar]
- 2.Trevaskis B. The central role of the VERNALIZATION1 gene in the vernalization response of cereals. Functional Plant Biol. 2010;37:479–487. [Google Scholar]
- 3.Trevaskis B, Hemming MN, Dennis ES, Peacock WJ. The molecular basis of vernalization-induced flowering in cereals. Trends Plant Sci. 2007;12:352–357. doi: 10.1016/j.tplants.2007.06.010. [DOI] [PubMed] [Google Scholar]
- 4.Greenup AG, Peacock WJ, Dennis, ES, Trevaskis B. The molecular biology of seasonal flowering-responses in Arabidopsis and the cereals. Ann Bot (Lond) 2009;103:1165–1172. doi: 10.1093/aob/mcp063. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Distelfeld A, Li C, Dubcovsky J. Regulation of flowering in temperate cereals. Current Opinion Plant Biol. 2009;12:178–184. doi: 10.1016/j.pbi.2008.12.010. [DOI] [PubMed] [Google Scholar]
- 6.Danyluk J, Kane NA, Breton G, Limin AE, Fowler DB, et al. TaVRT-1, a putative transcription factor associated with vegetative to reproductive transition in cereals. Plant Physiol. 2003;132:1849–1860. doi: 10.1104/pp.103.023523. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Trevaskis B, Bagnall DJ, Ellis MH, Peacock WJ, Dennis ES. MADS box genes control vernalization-induced flowering in cereals. Proc Natl Acad Sci U S A. 2003;100:13099–13104. doi: 10.1073/pnas.1635053100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Yan L, Loukoianov A, Tranquilli G, Helguera M, Fahima T, et al. Positional cloning of the wheat vernalization gene VRN1. Proc Natl Acad Sci U S A. 2003;100:6263–6268. doi: 10.1073/pnas.0937399100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Murai K, Miyamae M, Kato H, Takumi S, Ogihara Y. WAP1, a wheat APETALA1 homolog, plays a central role in the phase transition from vegetative to reproductive growth. Plant Cell Physiol. 2003;44:1255–1265. doi: 10.1093/pcp/pcg171. [DOI] [PubMed] [Google Scholar]
- 10.von Zitzewitz J, Szucs P, Dubcovsky J, Yan LL, Francia E, et al. Molecular and structural characterization of barley vernalization genes. Plant Mol Biol. 2005;59:449–467. doi: 10.1007/s11103-005-0351-2. [DOI] [PubMed] [Google Scholar]
- 11.Sasani S, Hemming MN, Oliver S, Greenup AG, Tavakkol-Afshari R, et al. The influence of vernalization and daylength cues on the expression of flowering-time genes in the leaves and shoot apex of barley (Hordeum vulgare). J Exp Botany. 2009;60:2169–2178. doi: 10.1093/jxb/erp098. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Shitsukawa N, Ikari C, Shimada S, Kitagawa S, Sakamoto K, et al. The einkorn wheat (Triticum monococcum) mutant, maintained vegetative phase, is caused by a deletion in the VRN1 gene. Genes Genet Syst. 2007;82:167–170. doi: 10.1266/ggs.82.167. [DOI] [PubMed] [Google Scholar]
- 13.Distelfeld A, Dubcovsky J. Characterization of the maintained vegetative phase deletions from diploid wheat and their effect on VRN2 and FT transcript levels. Mol Genet Genomics. 2010;283:223–232. doi: 10.1007/s00438-009-0510-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Jensen LB, Andersen JR, Frei U, Xing Y, Taylor C, et al. QTL mapping of vernalization response in perennial ryegrass (Lolium perenne L.) reveals co-location with an orthologue of wheat VRN1. Theor Appl Genet. 2005;110:527–536. doi: 10.1007/s00122-004-1865-8. [DOI] [PubMed] [Google Scholar]
- 15.Seppänen MM, Pakarinen K, Jokela V, Andersen JR, Fiil A, et al. Vernalization response of Phleum pratense and its relationships to stem lignification and floral transition. Ann Bot (Lond) 2010;106:697–707. doi: 10.1093/aob/mcq174. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Hemming MN, Peacock WJ, Dennis ES, Trevaskis B. Low-temperature and daylength cues are integrated to regulate FLOWERING LOCUS T in barley. Plant Physiol. 2008;147:355–366. doi: 10.1104/pp.108.116418. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Preston JC, Kellogg EA. Discrete developmental roles for temperate cereal grass VERNALIZATION1/FRUITFULL-like genes in flowering competency and the transition to flowering. Plant Physiol. 2008;146:265–276. doi: 10.1104/pp.107.109561. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Yan L, Loukoianov A, Blechl A, Tranquilli G, Ramakrishna W, et al. The wheat VRN2 gene is a flowering repressor down-regulated by vernalization. Science. 2004;303:1640–1644. doi: 10.1126/science.1094305. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Trevaskis B, Hemming MN, Peacock WJ, Dennis ES. HvVRN2 responds to daylength, whereas HvVRN1 is regulated by vernalization and developmental status. Plant Physiol. 2006;140:1397–1405. doi: 10.1104/pp.105.073486. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Greenup AG, Sasani S, Oliver SN, Talbot MJ, Dennis ES, et al. ODDSOC2 is a MADS box floral repressor that is down-regulated by vernalization in temperate cereals. Plant Physiol: 2010;153:1062–1073. doi: 10.1104/pp.109.152488. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Greenup AG, Sasani S, Oliver SN, Walford SA, Millar AM, et al. Transcriptome analysis of the vernalization response in barley (Hordeum vulgare) seedlings. PLoS ONE. 2011;6:17900. doi: 10.1371/journal.pone.0017900. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Yan L, Helguera M, Kato K, Fukuyama S, Sherman J, et al. Allelic variation at the VRN-1 promoter region in polyploid wheat. Theor Appl Genet. 2004;109:1677–1686. doi: 10.1007/s00122-004-1796-4. [DOI] [PubMed] [Google Scholar]
- 23.Pidal B, Yan L, Fu D, Zhang F, Tranquilli G, et al. The CARG-box located upstream from the transcriptional start of wheat vernalization gene VRN1 is not necessary for the vernalization response. Heredity. 2009;100:355–364. doi: 10.1093/jhered/esp002. [DOI] [PubMed] [Google Scholar]
- 24.Fu DL, Szucs P, Yan LL, Helguera M, Skinner JS, et al. Large deletions within the first intron in VRN-1 are associated with spring growth habit in barley and wheat. Mol Genet Genomics. 2005;273:54–65. doi: 10.1007/s00438-004-1095-4. [DOI] [PubMed] [Google Scholar]
- 25.Cockram J, Chiapparino E, Taylor SA, Stamati K, Donini P, et al. Haplotype analysis of vernalization loci in European barley germplasm reveals novel VRN-H1 alleles and a predominant winter VRN-H1/VRN-H2 multi-locus haplotype. Theor Appl Genet. 2007;115:993–1001. doi: 10.1007/s00122-007-0626-x. [DOI] [PubMed] [Google Scholar]
- 26.Szucs P, Skinner JS, Karsai I, Cuesta-Marcos A, Haggard KG, et al. Validation of the VRN-H2/VRN-H1 epistatic model in barley reveals that intron length variation in VRN-H1 may account for a continuum of vernalization sensitivity. Mol Genet Genomics. 2007;277:249–261. doi: 10.1007/s00438-006-0195-8. [DOI] [PubMed] [Google Scholar]
- 27.Hemming MN, Fieg S, Peacock WJ, Dennis ES, Trevaskis B. Regions associated with repression of the barley (Hordeum vulgare) VERNALIZATION1 gene are not required for cold induction. Mol Genet Genomics. 2009;282:107–117. doi: 10.1007/s00438-009-0449-3. [DOI] [PubMed] [Google Scholar]
- 28.Stockinger EJ, Skinner JS, Gardner KG, Francia E, Pecchioni N. Expression levels of barley Cbf genes at the Frost resistance-H2 locus are dependent upon alleles at Fr-H1 and Fr-H2. Plant J. 2007;51:308–321. doi: 10.1111/j.1365-313X.2007.0141.x. [DOI] [PubMed] [Google Scholar]
- 29.Oliver SN, Finnegan EJ, Dennis ES, Peacock WJ, Trevaskis B. Vernalization-induced flowering in cereals is associated with changes in histone methylation at the VERNALIZATION1 gene. Proc Natl Acad Sci U S A. 2009;106:8386–8391. doi: 10.1073/pnas.0903566106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Kane NA, Agharbaoui Z, Diallo AO, Adam H, Tominaga Y, et al. TaVRT2 represses transcription of the wheat vernalization gene TaVRN1. Plant J. 2007;51:670–680. doi: 10.1111/j.1365-313X.2007.03172.x. [DOI] [PubMed] [Google Scholar]
- 31.Trevaskis B, Tadege M, Hemming MN, Peacock WJ, Dennis ES, et al. Short Vegetative Phase-like MADS-box genes inhibit floral meristem identity in barley. Plant Physiol. 2007;143:225–235. doi: 10.1104/pp.106.090860. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Thomashaw M. Molecular basis of plant cold acclimation: insights gained from studying the CBF cold response pathway. Plant Physiol. 2010;154:571–577. doi: 10.1104/pp.110.161794. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Hood HM, Neafsey DE, Galagan J, Sachs MS. Evolutionary roles of upstream open reading frames in mediating gene regulation in fungi. Ann Rev Microbiol. 2009;63:385–409. doi: 10.1146/annurev.micro.62.081307.162835. [DOI] [PubMed] [Google Scholar]
- 34.Schmitz J, Franzen R, Ngyuen TH, Garcia-Maroto F, Pozzi C, et al. Cloning, mapping and expression analysis of barley MADS-box genes. Plant Mol Biol. 2000;42:899–913. doi: 10.1023/a:1006425619953. [DOI] [PubMed] [Google Scholar]
- 35.Danilevskaya ON, Meng X, Selinger DA, Deschamps S, Hermon P, et al. Involvement of the MADS-box gene ZMM4 in floral induction and inflorescence development in maize. Plant Physiol. 2008;147:2054–2069. doi: 10.1104/pp.107.115261. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Li CX, Dubcovsky J. Wheat FT protein regulates VRN1 transcription through interactions with FDL2. Plant J. 2008;55:543–554. doi: 10.1111/j.1365-313X.2008.03526.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Calvo SE, Pagliarini DJ, Mootha VK. Upstream open reading frames cause widespread reduction of protein expression and are polymorphic among humans. Proc Natl Acad Sci U S A. 2009;106:7507–7512. doi: 10.1073/pnas.0810916106. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Golovnina KA, Kondratenko EY, Blinov AG, Goncharov NP. Molecular characterization of vernalization loci VRN1 in wild and cultivated wheats. BMC Plant Biol. 2010;11:168. doi: 10.1186/1471-2229-10-168. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Wang M, Li Z, Matthews PR, Upadhyaya NM, Waterhouse PM. Improved vectors for Agrobacterium tumefaciens-mediated transformation of monocot plants. Acta Horticulture. 1998;461:401–408. [Google Scholar]
- 40.Tingay S, McElroy E, Kalla R, Fieg S, Wang M, et al. Agrobacterium mediated barley transformation. Plant J. 1997;11:1369–1376. [Google Scholar]
- 41.Mathews PR, Wang MB, Waterhouse PM, Thornton S, Fieg S, et al. Marker gene elimination from transgenic barley, using co-transformation with adjacent ‘‘twin T-DNAs’’ on a standard Agrobacterium transformation vector. Mol Breed. 2001;7:195–202. [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Low-temperature induction of the VRN1::GFP construct.Transcript levels for the VRN1::GFP fusion in transgenic seedlings. Expression was assayed in control seedlings, germinated at normal glasshouse temperatures (20 degrees for 4 days), and compared to seedlings germinated and grown to an identical stage of development at low temperatures (4 degrees for 28 days). Expression levels were assayed in two independent transgenic lines. Error bars show standard error. * indicates P<0.05.
(TIF)
Sequence motifs in the VRN1 promoter.The promoter sequence of the barley VRN1 gene showing potential transcription factor recognition sites and other putative regulatory sequences. The transcriptional start site, identified by 5′RACE is also indicated (TSS); this did not vary with different temperature treatments.
(DOC)
Deletion of the 5′ small open reading frame from the VRN1 gene of a wild wheat.Comparison of the A genome VRN1 gene from hexaploid wheat (VRN-A1, Genbank AY747600.1) with the sequence of the VRN1 gene from Triticum timopheevi (Genbank GQ451763). The VRN box is shaded grey. The putative CARG box is shaded black and the region that encodes the small upstream open reading frame is shown in bold text and boxed.
(DOC)
Co-localisation of DAPI staining and VRN1::GFP within cells of the developing inflorescence.Localization of DAPI staining compared to VRN1::GFP signal in the developing glume of a barley inflorescence. Double headed arrow shows DAPI and GFP signal in the same nucleus.
(TIF)