Abstract
High concentration of NaCl increases DNA breaks both in cell culture and in vivo. The breaks remain elevated as long as NaCl concentration remains high and are rapidly repaired when the concentration is lowered. The exact nature of the breaks, and their location, has not been entirely clear, and it has not been evident how cells survive, replicate, and maintain genome integrity in environments like the renal inner medulla in which cells are constantly exposed to high NaCl concentration. Repair of the breaks after NaCl is reduced is accompanied by formation of foci containing phosphorylated H2AX (γH2AX), which occurs around DNA double-strand breaks and contributes to their repair. Here, we confirm by specific comet assay and pulsed-field electrophoresis that cells adapted to high NaCl have increased levels of double-strand breaks. Importantly, γH2AX foci that occur during repair of the breaks are nonrandomly distributed in the mouse genome. By chromatin immunoprecipitation using anti-γH2AX antibody, followed by massive parallel sequencing (ChIP-Seq), we find that during repair of double-strand breaks induced by high NaCl, γH2AX is predominantly localized to regions of the genome devoid of genes (“gene deserts”), indicating that the high NaCl-induced double-strand breaks are located there. Localization to gene deserts helps explain why the DNA breaks are less harmful than are the random breaks induced by genotoxic agents such as UV radiation, ionizing radiation, and oxidants. We propose that the universal presence of NaCl around animal cells has directly influenced the evolution of the structure of their genomes.
Keywords: hypertonicity, salt, DNA damage, kidney, mIMCD3 cells
High extracellular NaCl increases the number of DNA breaks in mammalian cells in tissue culture (1, 2), mouse renal inner medullary cells in vivo (1), cells of the soil nematode Caenorhabditis elegans (3), and marine invertebrates (4). Acute elevation of NaCl in cell culture increases the number of DNA breaks (2, 5) and transiently arrests cells in all phases of the cell cycle (6, 7). After several hours, the cells begin proliferating again, despite the continued presence of high NaCl (7). However, even after cells adapt to high NaCl and reenter the cell cycle, numerous DNA breaks persist (1). Excessive elevation of NaCl causes apoptosis (7). However, the increased DNA breaks that occur at levels of NaCl that cells survive and to which they adapt differs from the chromatin fragmentation that occurs during apoptotic cell death. Thus, high NaCl increases DNA breaks in viable cells without the activation of caspases, nuclear condensation, or formation of apoptotic bodies characteristic of apoptosis (8, 9).
The increase of DNA breaks caused by high NaCl is not limited to proliferating cells in culture. High NaCl also induces DNA breaks in normal cells in animal tissues in vivo. Thus, numerous DNA breaks are normally present in the mouse renal inner medulla (1), where high interstitial NaCl provides the driving force for concentration of the urine (10). The excess breaks in the inner medulla disappear quickly when the high intercellular NaCl concentration in the renal medulla is lowered by the diuretic furosemide (1). The soil nematode, C. elegans is able to adapt to and live in a high NaCl environment (11), and adaptation of C. elegans to high NaCl is accompanied by increased DNA breaks (3). Finally, according to some estimates, ≈80% of all Earth's life lives in the ocean, which has a high osmolality of ≈1,000 mosmol/kg, the dominant solute being NaCl. Many marine invertebrates are osmoconformers, i.e., the NaCl in their extracellular fluids is as high as in seawater (12). Cells in tissues of osmoconforming marine invertebrates have many DNA breaks that disappear if the seawater in which they are immersed is gradually diluted to 300 mosmol/kg (4). Thus, increased DNA breaks in cells exposed to high NaCl is an evolutionarily conserved phenomenon. However, the nature of the DNA breaks, their location, and the mechanism of their induction has not been entirely clear.
A striking feature of adaptation to high NaCl is that despite increased DNA breaks, the cells do not activate the DNA damage response (1, 5, 8). However, the DNA damage response is activated quickly when NaCl is lowered. Thus, reducing NaCl to total osmolality of 300 mosmol/kg (the level normally maintained in mammalian blood and body fluids by osmoregulatory mechanisms) results in rapid repair of the DNA breaks (1, 5). This repair is accompanied by rapid phosphorylation of histone H2AX (called formation of γH2AX) (1, 5), the histone modification that normally accompanies repair of double-strand breaks (13, 14).
In the present studies, we find that the high NaCl-induced double-strand DNA breaks are not randomly distributed in the mouse genome, but are predominantly located in gene deserts, which are regions of the genome devoid of genes. Our findings are summarized on Fig. S1.
Results
High NaCl Induces Double-Strand Breaks That Are Rapidly Repaired When the NaCl Is Lowered.
We added 100 mM NaCl (which elevates the osmolality to ≈500 mosmol/kg) for 22 h to the medium bathing mIMCD3 cells. This addition of NaCl causes immediate G2/M arrest that lasts ≈6 h (6). Then, the cells begin proliferating again. By 22 h, the cell cycle distribution, appearance of the cells, and their rate of proliferation return to the condition before salt was elevated (ref. 1 and Fig. S2).
It has remained an open question what kind of DNA breaks are present in cells exposed to high NaCl. The breaks induced immediately by acute elevation of high NaCl for 1 h were originally characterized as double-strand DNA breaks (DSBs) by pulsed field gel electrophoresis (PFGE), which detects DSBs but not single-strand breaks (SSBs) (2). However, the methods used to study DNA breaks after cells adapt to high NaCl and resume proliferation were not specific to DSBs. Those methods included a version of the neutral comet assay originally developed by Ostling and Johanson (15), alkaline comet assay (1), and in vitro labeling of their 3′-OH ends with biotinylated deoxynucleotides in a reaction catalyzed by terminal deoxynucleotidyl transferase (1, 3, 4). All these methods do not distinguish between DSBs and SSBs (16–18). In the present studies, to clarify whether DNA breaks that remain induced by high NaCl in adapted cells include DSBs, we used a modification of the neutral comet assay that is specific for DSBs as opposed to SSBs (18, 19).
To confirm specificity of the neutral comet assay (19) that we used here for DSBs, we exposed cells to hydrogen peroxide (H2O2), known to induce only single-strand breaks (20). DNA breaks after H2O2 are not detected by the modified neutral comet assay but are detected by the alkaline comet assay, consistent with their single-stranded nature (Fig. 1A). We also tested mIMCD3 cells exposed to the topoisomerase inhibitor Etoposide, known to induce both double-strand and single-strand breaks in a proportion similar to ionizing radiation (20). The DNA breaks induced by Etoposide are detected by both the modified neutral and alkaline comet assay, consistent with presence of both double- and single-strand breaks (Fig. 1A). The breaks induced by high NaCl are detected by the modified neutral comet assay (Fig. 1A), indicating that they are double-stranded. The DSBs induced by high NaCl are repaired within several hours after the NaCl concentration is reduced to a total osmolality of 300 mosmol/kg (Fig. 1 B and C). Thus, an increased level of DSBs persists after cells have adapted to high NaCl and appear otherwise normal. In addition, there may also be more SSBs, but the effect of high NaCl on SSBs has not been specifically tested.
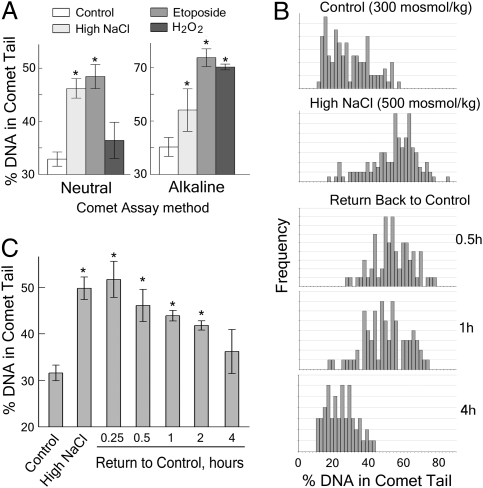
Fig. 1.
High NaCl (to 500 mosmol/kg) induces double-strand breaks (DSBs) in mIMCD3 cells that are repaired within several hours after lowering NaCl to control level (300 mosmol/kg). (A) Verification of specificity for DSBs of the Neutral Comet method used to detect DNA breaks. DNA breaks were induced by H2O2 and Etoposide. NaCl was elevated for 22 h. H2O2 induces only SSBs and oxidative damage to nucleotides. Etoposide induces both DSBs and SSBs. The % DNA in Comet Tails was used as a measure of the DNA breaks. Data are plotted as mean ± SEM (n = 5–7; *P < 0.05, t test relative to control). Neutral Comet Assay detects only the DSBs induced by Etoposide and NaCl, but not the SSBs induced by H2O2. Alkaline Comet Assay detects DNA breaks in cells treated with Etoposide, NaCl, and H2O2. Conclusion: high NaCl induces DSBs. (B and C) DSBs that are induced by high NaCl are repaired after NaCl is lowered. Cells were treated with medium in which NaCl was elevated to total osmolality 500 mosmol/kg for 22 h, and then the high NaCl medium was replaced with medium at 300 mosmol/kg. DSBs were measured by the neutral comet assay. (B) Representative distributions of % DNA in comet tails. (C) Analysis of mean % DNA in comet tails (mean ± SEM, n = 3). *P < 0.05, t test relative to control.
ATM/ATR-Dependent DNA Double-Strand Break Repair Response Is Activated During Repair of High NaCl-Induced DNA Breaks.
Common DNA damage repair responses include transient cell cycle arrest, during which the DNA repair occurs. We tested to see whether that happens during the disappearance of the DNA breaks after reduction of high NaCl (Fig. 2). Indeed, phosphorylation of checkpoint kinase 1 (Chk1), which contributes to all defined cell cycle checkpoints (21), occurs rapidly, and the phosphorylation is reduced by caffeine, which is an ATM/ATR inhibitor (22) (Fig. 2A). Similarly, G2/M cell cycle arrest activates rapidly when NaCl decreases, and this arrest is abrogated by caffeine (Fig. 2B). Repair of the breaks after NaCl is reduced is accompanied by formation of foci containing phosphorylated H2AX (γH2AX) (1, 5). This histone modification occurs around DNA double-strand breaks and contributes to their repair (13, 14). γH2AX is induced to a maximal level within 15 min after lowering NaCl, then gradually decreases (Fig. 2C) accompanying repair of the DSBs (Fig. 1C). The γH2AX induction is also sensitive to inhibition of ATM/ATR by caffeine (Fig. 2C). These results indicate that, when elevated NaCl is lowered, a classical ATM/ATR-dependent DNA damage response becomes activated, and they further confirm that DNA DSBs increase upon exposure to high NaCl.
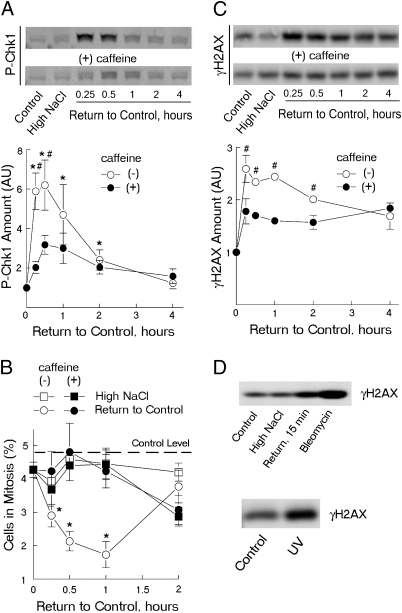
Fig. 2.
Repair of high NaCl-induced DNA breaks following lowering of NaCl is accompanied by the caffeine-sensitive chk1 phosphorylation, G2/M cell cycle arrest and γH2AX induction. NaCl bathing mIMCD3 cells was elevated by adding NaCl to total osmolality 500 mosmol/kg for 22 h, then the additional NaCl was eliminated, lowering the osmolality to 300 mosmol/kg in the presence or absence of 2 mM caffeine. At indicated time points after lowering NaCl phosphorylated Chk1 (“P-Chk1”), γH2AX and the number of cells in mitosis were determined. (A and B) Chk1 becomes phosphorylated and G2/M arrest is activated when NaCl is lowered. Caffeine decreases the chk1 phosphorylation and abrogates the cell cycle arrest. (A) Western blot analysis of P-Chk1. (A Upper) Representative Western blot. (A Lower) Densitometry (mean ± SEM, n = 3, *P < 0.05 relative to 0 time; #P < 0.05 relative to caffeine, t test). (B) Analysis of % cells in mitosis by staining with anti–P-histone H3 (mean ± SEM, n = 4; *P < 0.02, t test relative to 0 time). Dashed line shows percent in mitosis of cells in control medium before NaCl is elevated (4.8% ± 0.2, n = 16). (C and D) Western blot analysis of γH2AX changes in response to high NaCl, bleomycin, and UV irradiation. (C) γH2AX increases after NaCl is lowered, and this increase is prevented by caffeine. γH2AX is maximal 15 min after NaCl is reduced and gradually decreases as DNA breaks decrease (Fig. 1). (C Upper) Representative Western blot. (C Lower) Densitometry (mean ± SEM, n = 3; #P < 0.05, t test relative to caffeine). (D) mIMCD3 cells were exposed to bleomycin (5 μg/mL for 30 min) or 15 J/m2 of UV light as described in Materials and Methods. The increase of γH2AX 15 min after return from high NaCl is comparable to that after exposure to UV radiation and bleomycin.
During Repair of High NaCl-Induced DNA Breaks, γH2AX Is Mainly Located in Gene Deserts, Whereas Bleomycin and UV Induce γH2AX at Random Locations Throughout Genome.
Given that high NaCl induces DSBs that persist as long as the level of NaCl stays high, it has not been clear how cells survive and function in the continued presence of those breaks, why the cell cycle does not remain arrested, how DNA transcription and replication can proceed despite the breaks, and how mutations and genomic instability are prevented. To begin answering those questions, we have determined the genomic location of the breaks. Our strategy is based on the fact that during repair of DSBs, γH2AX is induced in distinct foci (Fig. 3). γH2AX-containing foci occur at DSBs (13, 14), and the number of γ-H2AX foci approximates the number of DSBs induced (ref. 14 and reviewed in ref. 23), making foci of γH2AX consistent and quantitative markers of DSBs. Therefore, to detect locations of DSBs, we performed chromatin immunoprecipitation (ChIP) by using anti-γH2AX antibody, followed by massive parallel sequencing of isolated DNA fragments (γH2AX ChIP-Seq) (24). Given that the maximal intensity of γH2AX foci occurs 15 min after NaCl is lowered and that the intensity of the foci already decreases by 30 min (Fig. 3B), we determined the genomic locations of the γH2AX foci that are present 15 min after lowering NaCl. In addition, we performed ChIP-Seq to determine the genomic locations of the γH2AX induced by bleomycin and UV radiation. Both bleomycin and UV increase γH2AX (Fig. 2D). However, pattern of γH2AX immunostaining is different (Fig. 3A) because of the different nature of damage induced by bleomycin and UV. Thus, bleomycin induces DSBs directly (25) and produces distinct γH2AX foci (Fig. 3A), whereas, after UV radiation, DSBs arise only indirectly as a result of the action of repair or degradation of arrested replication forks (26). After UV radiation, γH2AX is increased both in foci and in more diffused staining (Fig. 3A) that might not be related to DSBs but to some other types of DNA damage (27). Thus, by γH2AX ChIP-Seq we analyzed genomic locations of γH2AX induced by bleomycin, UV, and during repair of high NaCl-induced DNA breaks. We aligned sequence reads (tags) to the mouse genome and analyzed the tag density in the University of California, Santa Cruz (UCSC) genome browser (Fig. 4 and Fig. S5).
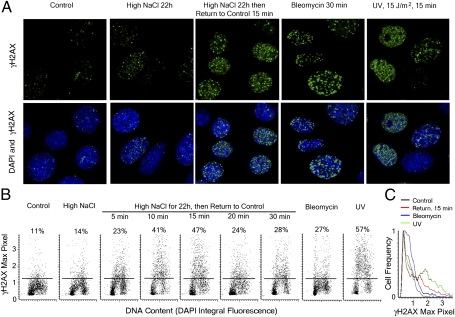
Fig. 3.
γH2AX induction after lowering of NaCl occurs in distinct foci, similar to those from other causes of DSBs, and the brightness of the foci is maximal 15 min after lowering NaCl. NaCl bathing mIMCD3 cells was elevated by adding NaCl to total osmolality of 500 mosmol/kg for 22 h, then the additional NaCl was eliminated, lowering the osmolality to 300 mosmol/kg. Cells were exposed to bleomycin (5 μg/mL for 30 min) or 15 J/m2 of UV light. (A) Immunocytochemical staining for γH2AX demonstrating γH2AX foci. (B and C) Analysis of brightness of γH2AX foci by laser scanning cytometry (LSC). (B) Cytograms (LSC), plotting Maximal Pixel Fluorescence (Max Pixel) of γH2AX in nuclei (a measure of γH2AX brightness of foci) vs. Integral DAPI (blue) fluorescence (a measure of nuclear DNA content). Numbers on cytogramms show % of cells in which Max Pixel of γH2AX fluorescence is above the level shown by horizontal lane. Brightness of γH2AX fluorescence and proportion of cells in which it increases are maximal 15 min after NaCl is lowered. Bleomycin and UV increase γH2AX as expected. (C) Cell distributions based on γH2AX fluorescence intensity. γH2AX Fluorescence intensity increases in a fraction of cells after treatment with bleomycin and UV and 15 min after lowering NaCl.
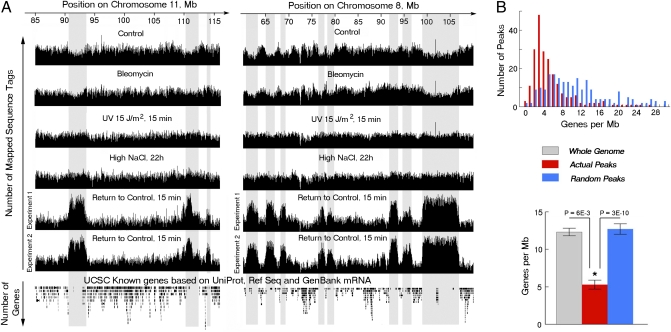
Fig. 4.
During repair of high NaCl-induced DNA breaks, γH2AX is mainly located in gene deserts. ChIP-Seq analysis of genomic locations of DNA fragments immunoprecipitated by anti-γH2AX. Chromatin was digested with MNase to mononucleosome size. Mononucleosomes containing γH2AX were immunoprecipitated with anti-γH2AX antibody and purified DNA fragments were sequenced by using a Solexa 2G Genome Analyzer. (A) UCSC genome browser view of representative examples of sequence tags mapped to the mouse genome. The lowest track shows location of genes (UCSC genome browser). ChIP-seq was performed by using mIMCD3 cells treated, as follows. Control cells maintained at 300 mosmol/kg. Bleomycin cells were exposed to 5 μg/mL bleomycin for 30 min. “UV” cells at 300 mosmol/kg exposed to UV. “High NaCl” cells maintained at 500 mosmol/kg (NaCl added) for 22 h. “Return to Control” cells: NaCl was elevated to total osmolality of 500 mosmol/kg for 22 h, then NaCl was lowered to a total osmolality of 300 mosmol/kg for 15 min. Note the distinct peaks of increased sequence tag density in Return to Control samples indicating that the γH2AX foci occur at specific locations within genome. Note further that such peaks do not occur in DNA from cells under any of the other conditions. The gray projections of the peaks in Return to Control demonstrate locations in areas of genome containing few genes (gene deserts). The lack of peaks in conditions other than return to control indicates that the γH2AX foci are located randomly in the genome under those conditions. Additional examples are shown in Fig. S5. (B) Relation between the observed γH2AX peaks and gene density throughout the mouse genome. (B Upper) Calculated number of genes per megabase within each peak of DNA immunoprecipitated by anti-γH2AX antibody during repair of high NaCl-induced DSBs. The number of peaks containing each gene densities (Genes per MB) is plotted (Actual Peaks, red), compared with the gene densities in randomly generated peaks of the same size and number in each chromosome (Random Peaks, blue). Evidently, gene density is low in γH2AX peaks throughout the genome. (B Lower) The gene density in the γH2AX peaks is also significantly below the average gene density throughout the genome.
There are clusters of increased density of mapped tags (γH2AX peaks) in cells after reduction of high NaCl (Fig. 4A and Fig. S5). The peaks occur mainly in large intergenic regions (gene deserts). Thus, H2AX becomes phosphorylated at specific locations within gene deserts during repair of the high NaCl-induced DSBs, showing that the high NaCl-induced DSBs are not randomly distributed throughout genome but occur within gene deserts. The peaks do not occur in DNA from cells maintained continuously at 300 or 500 mosmol/kg (Fig. 4A, “Control” and “High NaCl, 22h”). Bleomycin and UV also increase γH2AX (Figs. 2D and 3), but peaks do not occur in DNA from cells in which it has been damaged by Bleomycin or UV irradiation (Fig. 4A, “Bleomycin” and “UV”). Thus, the DNA damage caused by Bleomycin and UV occurs randomly throughout the genome, in marked contrast to the localized DSBs caused by high NaCl.
The localization of peaks to gene deserts is apparent by visual examination of tag density in the genome browser. Representative regions are shown in Fig. 4 and Fig. S5. To apply this observation to the whole genome, we extracted information about peak coordinates from the UCSC genome browser and analyzed gene density over the regions of identified peaks in comparison with artificial peaks of the same width and number that were randomly generated throughout genome (Fig. 4B, Upper). We identified 215 peaks of mean width of 2.4 Mb (95% confidence interval between 2.2 Mb and 2.7 Mb). Gene density over the peak regions is greatly reduced compared with random peaks or average gene density in mouse genome (Fig. 4B, Lower).
PFGE Identifies Increased DNA Fragmentation After Exposure of mIMCD3 Cells to High NaCl, and the Distribution of the Lengths of the Fragments Is Consistent with Spacing of the γH2AX Peaks.
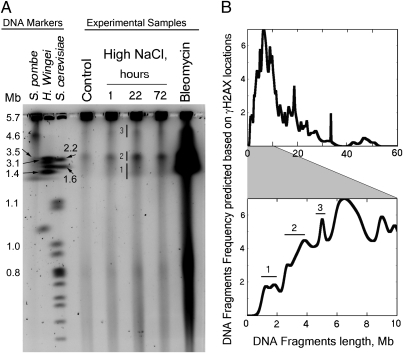
We used PFGE, which separates large DNA fragments according to size (28), to further test whether high NaCl induces double-strand breaks and whether the breaks are not randomly distributed (Fig. 5). Exposure of cells to high NaCl increases the number of DNA fragments that enter the gel and the size of the fragments is not random, consistent with a nonrandom location of the DNA breaks (Fig. 5A). Given the locations of the γH2AX peaks and assuming that there is one DNA break in each peak, we calculated a size distribution of the predicted DNA fragments (Fig. 5B and SI Materials and Methods). The calculated distribution of fragment sizes closely resembles the actual distribution (Fig. 5), which supports the conclusion that high NaCl-induced double-strand breaks occur predominantly in gene deserts.
Fig. 5.
(A) High NaCl increases DNA fragmentation in mIMCD3 cells (pulsed-field gel electrophoresis, PFGE). NaCl was elevated to a total osmolality of 500 mosmol/kg for the indicated times or 5μg/mL bleomycin was added for 30 min. Cells kept in high NaCl for 72 h were split once to maintain logarithmic growth (representative of three independent experiments). High NaCl increases DNA fragmentation, and there are size peaks of the DNA fragments, consistent with nonrandom location of the DNA breaks. (B) The distribution of DNA fragment lengths is consistent with that predicted from the distance between γH2AX peaks. We assumed one break per γH2AX peak, present at a maximal probability at the middle of the peak. Upper shows the probability distribution for fragments of lengths from 0 to 60 Mb. Because the limit of detection for PFGE is ≈6 Mb, Lower is restricted to predicted sizes <10 Mb. Note that the peaks of predicted fragment sizes (labeled 1, 2, and 3) are consistent with the peaks seen experimentally in A.
Discussion
Why there are gene deserts has remained a mystery. They were discovered when whole genome sequencing showed that genes are not evenly distributed. A substantial fraction of mammalian genomes contains gene deserts, defined as long regions (>500 kb) containing no protein-coding sequences. Gene deserts occupy ≈38% of human, 34% of mouse, 23% of rat, and 20% of dog genome (29). It is extremely unlikely that gene deserts reached their observed maximal size of 5.1 Mb with 545 deserts larger than 640 kb by chance (30), which raises the question of what selective pressure might be acting.
Animal cells are universally exposed to NaCl, and the level of NaCl may be high in animals exposed to marine or desiccated terrestrial environments. During evolution of mammals, osmoregulatory mechanisms developed that maintain osmolality of most extracellular fluids close to 300 mosmol/kg. Nevertheless, even in mammals, NaCl concentration is constantly very high in some tissues, particularly the renal medulla. Given our finding that DNA breaks induced by high NaCl are concentrated in gene deserts, we suggest that, as the size of genomes has increased, newly formed regions are susceptible (for unknown reasons) to high NaCl-induced DNA breaks and evolve to contain fewer genes, thus limiting mutations and preventing genomic instability. This suggestion is supported by several observations.
i) The neutral mutation rate (30) and the rate of genome rearrangements associated with appearances of new centromeres (31) are both higher in gene deserts than in regions containing genes.
ii) Before the evolution of vertebrates, the sizes of genomes grew in proportion to the number of genes. However, gene deserts began appearing in fish and increased in size to occupy ≈38% of the genome in humans (29). Over the same period, osmoregulatory mechanisms developed that maintain systemic osmolality close to 300 mosmol/kg. Estimates from molecular clocks of the rates of evolution show that the rates decreased significantly in vertebrates before the origin of Osteichthyes (32). That could have been due to a combination of decreased rate of mutations in protein coding regions owing to more precise osmoregulation and the low abundance of functional genes in gene deserts where they would be susceptible to NaCl-induced breaks.
iii) Recently, a model was proposed relating the rate of molecular evolution and the maximal size of genomes (33). The theory assumes that for an organism to be viable, essential genes must be functional. Further, it predicts that populations become extinct because of lethal mutagenesis when the mutation rate exceeds approximately six mutations per replication in essential parts of the genome in mesophilic organisms and one or two mutations in thermophilic ones. The theory therefore predicts that mutation rate limits essential genome size; in other words, the higher the mutation rate, the smaller the sustainable size of the genome. This theory implies that increasing the size of the genome required that genes not evolve in regions, like the present gene deserts, that are more susceptible to DNA breaks and mutations.
Our finding that high NaCl-induced DSBs are located in gene deserts is an example of nonrandom induction of DNA breaks in higher organisms. Although we are uncertain why high NaCl breaks DNA, the gene deserts apparently have properties that render them more susceptible. Limitation of high NaCl-induced DNA breaks to gene deserts helps explain why they apparently are less harmful than are the random breaks induced by genotoxic agents like UV radiation, ionizing radiation, and oxidants. Further, our finding suggests a possible role of high NaCl in evolution of the structure of the animal genome.
Perspective.
More studies are required to decipher why double-strand breaks occur predominantly in gene deserts during exposure to high NaCl. Possibilities that we are considering include decreased DNA repair in gene deserts similar to that in heterochromatin (34), presence of specific target sequences for nucleases activated by high NaCl, and high NaCl-induced alterations of chromatin in gene deserts that makes the DNA there more susceptible to damaging agents.
Materials and Methods
Methods were published for Western blot (1), exposure of cells to UV radiation (5), treatment with H2O2 (20), analysis of cells in mitosis (6), immunostaining, and analysis of brightness of γH2AX foci by laser-scanning cytometry (LSC) (5). More details are included in SI Materials and Methods, as are details of doses and timing of drug application, analysis of gene density at genomic locations enriched with γH2AX-immunoprecipitated sequence tags, and analysis of expected distribution of DNA fragment sizes, based on genomic locations of γH2AX ChIP-Seq peaks.
Cell Culture.
mIMCD3 cells (35) were grown in medium containing 45% DME Low Glucose (Invitrogen), 45% F12 Coon's Modification (No. F6636; Sigma), and 10% FBS (HyClone). Osmolality of control medium was 300–320 mosmol/kg. High NaCl medium was prepared by adding NaCl to the total osmolality of 500 mosmol/kg. All of the experiments were performed on logarithmically growing cells at ≈80% confluence. To elevate NaCl, control medium was replaced by the high NaCl media.
Analysis of Double-Strand and Single-Strand DNA Breaks by Comet Assay.
Two different assays were used: neutral comet assay modified for detection of double-strand breaks and alkaline comet assay, which detects DNA SSBs, double-strand breaks, and alkali-labile sites. Those assays were performed as described (19) with minor modifications. See SI Materials and Methods and Fig. S4 for details.
ChIP and Illumina Library Construction for Sequencing.
ChIP was performed by using Enzymatic Chromatin IP kit (No. 9003; Cell Signaling Technology). Conversion of the ChIP-enriched DNA into libraries suitable for sequencing using the Illumina Genome Analyzer was performed by using the published protocol (36). See SI Materials and Methods and Fig. S4 for the detailed ChIP-Seq protocol.
Solexa Pipeline Analysis.
Sequence tags were obtained and mapped to the mouse genome by using the Solexa Analysis Pipeline as described (37). The unique reads were retained and converted to browser extensible data (BED) files for viewing the data in the UCSC genome browser. The read number and genomic coordinates were summarized in 300-bp windows.
Analysis of DNA Fragmentation by PFGE.
Agarose embedded DNA was prepared by using the CHEF Mammalian Genomic DNA Plug Kit (No. 170–3591; Bio-Rad). Briefly, cells were rinsed with PBS, scraped off the dish, resuspended in 1% CleanCut Agarose from the kit at a final concentration of 12 million cells/mL. The agarose/cell suspension was solidified at 4 °C for 10 min in a casting mold, followed by incubation of the agarose plugs in Proteinase K solution at 50 °C for 3 d to digest proteins. PFGE was performed as described (28) by using the CHEF-DR II system (Bio-Rad) and the following parameters: 1% Megabase Agarose (No. 161–3108; Bio-Rad), 0.5× TBE running buffer, 120° reorientation angle, 6 V⋅cm−1, and 14 °C. Gels were run for 16 h with switch time of 16 s, followed by 30-h run with switch time of 80 s DNA Size Markers were as follows: Schizosaccharomyes pombe, Saccharomyces cerevisiae, and Hansenula wingei chromosomes (Bio-Rad). Gels were stained with SYBR Gold (Invitrogen).
Supplementary Material
Acknowledgments
We thank Drs. Chris Combs and Daniela Malide at the National Heart, Lung, and Blood Institute (NHLBI) Light Microscopy Core Facility for help with microscopy and images processing and Dr. Iouri Chepelev at the NHLBI Laboratory of Molecular Immunology for assistance with sequencing data analysis. This research was supported by the Intramural Research Programs of the National Institutes of Health, NHLBI.
Footnotes
The authors declare no conflict of interest.
Data deposition: The data reported in this paper have been deposited in the Gene Expression Omnibus (GEO) database, www.ncbi.nlm.nih.gov/geo (accession no. GSE32882).
See Commentary on page 20281.
This article contains supporting information online at www.pnas.org/lookup/suppl/doi:10.1073/pnas.1114677108/-/DCSupplemental.
References
- 1.Dmitrieva NI, Cai Q, Burg MB. Cells adapted to high NaCl have many DNA breaks and impaired DNA repair both in cell culture and in vivo. Proc Natl Acad Sci USA. 2004;101:2317–2322. doi: 10.1073/pnas.0308463100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Kültz D, Chakravarty D. Hyperosmolality in the form of elevated NaCl but not urea causes DNA damage in murine kidney cells. Proc Natl Acad Sci USA. 2001;98:1999–2004. doi: 10.1073/pnas.98.4.1999. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Dmitrieva NI, Celeste A, Nussenzweig A, Burg MB. Ku86 preserves chromatin integrity in cells adapted to high NaCl. Proc Natl Acad Sci USA. 2005;102:10730–10735. doi: 10.1073/pnas.0504870102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Dmitrieva NI, Ferraris JD, Norenburg JL, Burg MB. The saltiness of the sea breaks DNA in marine invertebrates: Possible implications for animal evolution. Cell Cycle. 2006;5:1320–1323. doi: 10.4161/cc.5.12.2867. [DOI] [PubMed] [Google Scholar]
- 5.Dmitrieva NI, Bulavin DV, Burg MB. High NaCl causes Mre11 to leave the nucleus, disrupting DNA damage signaling and repair. Am J Physiol Renal Physiol. 2003;285:F266–F274. doi: 10.1152/ajprenal.00060.2003. [DOI] [PubMed] [Google Scholar]
- 6.Dmitrieva NI, Bulavin DV, Fornace AJ, Jr, Burg MB. Rapid activation of G2/M checkpoint after hypertonic stress in renal inner medullary epithelial (IME) cells is protective and requires p38 kinase. Proc Natl Acad Sci USA. 2002;99:184–189. doi: 10.1073/pnas.231623498. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Michea L, et al. Cell cycle delay and apoptosis are induced by high salt and urea in renal medullary cells. Am J Physiol Renal Physiol. 2000;278:F209–F218. doi: 10.1152/ajprenal.2000.278.2.F209. [DOI] [PubMed] [Google Scholar]
- 8.Dmitrieva NI, Burg MB. Living with DNA breaks is an everyday reality for cells adapted to high NaCl. Cell Cycle. 2004;3:561–563. [PubMed] [Google Scholar]
- 9.Dmitrieva NI, Burg MB. Analysis of DNA breaks, DNA damage response, and apoptosis produced by high NaCl. Am J Physiol Renal Physiol. 2008;295:F1678–F1688. doi: 10.1152/ajprenal.90424.2008. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Bankir L. Urea and the kidney. In: Brenner BM, editor. The Kidney. Vol 5. Philadelphia: WB Saunders Company; 1996. pp. 571–606. [Google Scholar]
- 11.Lamitina ST, Morrison R, Moeckel GW, Strange K. Adaptation of the nematode Caenorhabditis elegans to extreme osmotic stress. Am J Physiol Cell Physiol. 2004;286:C785–C791. doi: 10.1152/ajpcell.00381.2003. [DOI] [PubMed] [Google Scholar]
- 12.Sherwood L, Klandorf H, Yancey P. Animal Physiology. From Genes to Organisms. Belmont, CA: Thomson Learning; 2005. [Google Scholar]
- 13.Rogakou EP, Pilch DR, Orr AH, Ivanova VS, Bonner WM. DNA double-stranded breaks induce histone H2AX phosphorylation on serine 139. J Biol Chem. 1998;273:5858–5868. doi: 10.1074/jbc.273.10.5858. [DOI] [PubMed] [Google Scholar]
- 14.Sedelnikova OA, Rogakou EP, Panyutin IG, Bonner WM. Quantitative detection of (125)IdU-induced DNA double-strand breaks with gamma-H2AX antibody. Radiat Res. 2002;158:486–492. doi: 10.1667/0033-7587(2002)158[0486:qdoiid]2.0.co;2. [DOI] [PubMed] [Google Scholar]
- 15.Ostling O, Johanson KJ. Microelectrophoretic study of radiation-induced DNA damages in individual mammalian cells. Biochem Biophys Res Commun. 1984;123:291–298. doi: 10.1016/0006-291x(84)90411-x. [DOI] [PubMed] [Google Scholar]
- 16.Dmitrieva NI, Burg MB. Osmotic stress and DNA damage. Methods Enzymol. 2007;428:241–252. doi: 10.1016/S0076-6879(07)28013-9. [DOI] [PubMed] [Google Scholar]
- 17.Collins AR, Dobson VL, Dusinská M, Kennedy G, Stĕtina R. The comet assay: What can it really tell us? Mutat Res. 1997;375:183–193. doi: 10.1016/s0027-5107(97)00013-4. [DOI] [PubMed] [Google Scholar]
- 18.Collins AR. The comet assay for DNA damage and repair: Principles, applications, and limitations. Mol Biotechnol. 2004;26:249–261. doi: 10.1385/MB:26:3:249. [DOI] [PubMed] [Google Scholar]
- 19.Olive PL, Banáth JP. The comet assay: A method to measure DNA damage in individual cells. Nat Protoc. 2006;1:23–29. doi: 10.1038/nprot.2006.5. [DOI] [PubMed] [Google Scholar]
- 20.Olive PL, Johnston PJ. DNA damage from oxidants: Influence of lesion complexity and chromatin organization. Oncol Res. 1997;9:287–294. [PubMed] [Google Scholar]
- 21.Dai Y, Grant S. New insights into checkpoint kinase 1 in the DNA damage response signaling network. Clin Cancer Res. 2010;16:376–383. doi: 10.1158/1078-0432.CCR-09-1029. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Zhou BB, et al. Caffeine abolishes the mammalian G(2)/M DNA damage checkpoint by inhibiting ataxia-telangiectasia-mutated kinase activity. J Biol Chem. 2000;275:10342–10348. doi: 10.1074/jbc.275.14.10342. [DOI] [PubMed] [Google Scholar]
- 23.Kinner A, Wu W, Staudt C, Iliakis G. Gamma-H2AX in recognition and signaling of DNA double-strand breaks in the context of chromatin. Nucleic Acids Res. 2008;36:5678–5694. doi: 10.1093/nar/gkn550. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Wold B, Myers RM. Sequence census methods for functional genomics. Nat Methods. 2008;5:19–21. doi: 10.1038/nmeth1157. [DOI] [PubMed] [Google Scholar]
- 25.Hecht SM. Bleomycin: New perspectives on the mechanism of action. J Nat Prod. 2000;63:158–168. doi: 10.1021/np990549f. [DOI] [PubMed] [Google Scholar]
- 26.Limoli CL, Giedzinski E, Bonner WM, Cleaver JE. UV-induced replication arrest in the xeroderma pigmentosum variant leads to DNA double-strand breaks, gamma -H2AX formation, and Mre11 relocalization. Proc Natl Acad Sci USA. 2002;99:233–238. doi: 10.1073/pnas.231611798. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.de Feraudy S, Revet I, Bezrookove V, Feeney L, Cleaver JE. A minority of foci or pan-nuclear apoptotic staining of gammaH2AX in the S phase after UV damage contain DNA double-strand breaks. Proc Natl Acad Sci USA. 2010;107:6870–6875. doi: 10.1073/pnas.1002175107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Herschleb J, Ananiev G, Schwartz DC. Pulsed-field gel electrophoresis. Nat Protoc. 2007;2:677–684. doi: 10.1038/nprot.2007.94. [DOI] [PubMed] [Google Scholar]
- 29.Salzburger W, Steinke D, Braasch I, Meyer A. Genome desertification in eutherians: Can gene deserts explain the uneven distribution of genes in placental mammalian genomes? J Mol Evol. 2009;69:207–216. doi: 10.1007/s00239-009-9251-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Ovcharenko I, et al. Evolution and functional classification of vertebrate gene deserts. Genome Res. 2005;15:137–145. doi: 10.1101/gr.3015505. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Lomiento M, Jiang Z, D'Addabbo P, Eichler EE, Rocchi M. Evolutionary-new centromeres preferentially emerge within gene deserts. Genome Biol. 2008;9:R173. doi: 10.1186/gb-2008-9-12-r173. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Peterson KJ, et al. Estimating metazoan divergence times with a molecular clock. Proc Natl Acad Sci USA. 2004;101:6536–6541. doi: 10.1073/pnas.0401670101. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Zeldovich KB, Chen P, Shakhnovich EI. Protein stability imposes limits on organism complexity and speed of molecular evolution. Proc Natl Acad Sci USA. 2007;104:16152–16157. doi: 10.1073/pnas.0705366104. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Goodarzi AA, Jeggo P, Lobrich M. The influence of heterochromatin on DNA double strand break repair: Getting the strong, silent type to relax. DNA Repair (Amst) 2010;9:1273–1282. doi: 10.1016/j.dnarep.2010.09.013. [DOI] [PubMed] [Google Scholar]
- 35.Rauchman MI, Nigam SK, Delpire E, Gullans SR. An osmotically tolerant inner medullary collecting duct cell line from an SV40 transgenic mouse. Am J Physiol. 1993;265:F416–F424. doi: 10.1152/ajprenal.1993.265.3.F416. [DOI] [PubMed] [Google Scholar]
- 36.Schmidt D, et al. ChIP-seq: Using high-throughput sequencing to discover protein-DNA interactions. Methods. 2009;48:240–248. doi: 10.1016/j.ymeth.2009.03.001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Barski A, et al. High-resolution profiling of histone methylations in the human genome. Cell. 2007;129:823–837. doi: 10.1016/j.cell.2007.05.009. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.