Abstract
Naturally SIV-infected sooty mangabeys (SMs) do not progress to AIDS despite high-level virus replication. We previously showed that the fraction of CD4+CCR5+ T-cells is lower in SMs compared to humans and macaques. Here we found that, after in vitro stimulation, SM CD4+ T-cells fail to up-regulate CCR5, and that this phenomenon is more pronounced in CD4+ central-memory T-cells (TCM). CD4+ T-cell activation was similarly uncoupled from CCR5 expression in SMs in vivo during (i) acute SIV infection and (ii) following antibody-mediated CD4+ T-cell depletion. Remarkably, CD4+ TCM of SMs that express low levels of CCR5 demonstrated reduced susceptibility to SIV infection both in vivo and in vitro when compared to CD4+ TCM of RMs. These data suggest that low CCR5 expression on SM CD4+ T-cells favors the preservation of CD4+ T-cell homeostasis and promotes an AIDS-free status by protecting CD4+ TCM from direct virus infection.
Introduction
Human Immunodeficiency Virus type-1 (HIV-1) infection of humans and Simian Immunodeficiency Virus (SIV) infection of rhesus macaques (RMs) lead to a progressive immune deficiency known as AIDS. In contrast, SIV infections of African “natural host” species, such as sooty mangabeys (SM) and African green monkeys (AGMs), are typically non-pathogenic despite high levels of virus replication1,2. Understanding the reasons why “natural” SIV infections are non-progressive is a key priority in contemporary AIDS research, with significant implications for HIV pathogenesis, therapy, and vaccines3.
Previous studies showed that SIVsmm infection of SMs is characterized by: (i) peak of viremia at day 10–15 post-infection, followed by a post-peak decline to set-point levels of 104–106 copies/ml of plasma4,5; (ii) a vigorous but transient immune response to the virus, with marked activation of the type-I Interferon response6,7; (iii) early depletion of mucosal CD4+ T-cells with relative preservation of Th17 cells4,8; (iv) maintenance of healthy peripheral CD4+ T-cell counts in ~80–90% of animals9; (v) short in vivo lifespan of productively infected cells10; (vi) absence of microbial translocation and mucosal immune dysfunction4,11; (vii) low levels of immune activation throughout the chronic phase of infection4,12,13.
The limited immune activation of SIV-infected SMs represents a striking difference with the generalized immune activation that characterizes pathogenic HIV and SIV infections14,15. Several hypotheses have been proposed to explain this phenotype, including SIVsmmNef-mediated down-modulation of CD3, rapid up-regulation of PD-1 in lymph nodes, lower plasmacytoid dendritic cells response to TLR ligands, and lack of microbial translocation11,16–19. Of note, the fact that SIV infection of SMs is associated with a transient phase of strong immune responses to the virus suggests the involvement of immune regulatory mechanisms, as opposed to a genetically determined inability to sense the virus6. Currently, the mechanisms by which SIV-infected SMs avoid chronic immune activation remain poorly understood, and it remains unclear to what extent this lack of immune activation represents a cause rather than a consequence of preserved immune function.
We previously reported that in natural SIV hosts the fraction of circulating and mucosal CD4+ T-cells expressing the SIV co-receptor CCR5 is markedly lower than in humans and RMs20. However, the interpretation of this observation has been complicated by the high viremia of naturally SIV-infected SMs13,21. In this study, we discovered that central memory CD4+ T-cells (TCM) of SMs are exquisitely resistant to CCR5 up-regulation upon in vivo and in vitro activation, and that this pattern of reduced CCR5 expression is associated with lower in vivo and in vitro susceptibility of CD4+ TCM to direct SIV infection. These data define a novel and potentially critical mechanism of AIDS resistance in SIV-infected SMs characterized by selective protection of CD4+ TCM from virus infection due to limited CCR5 up-regulation.
Results
The fraction of CD4+CCR5+ TCM is significantly lower in SMs than RMs
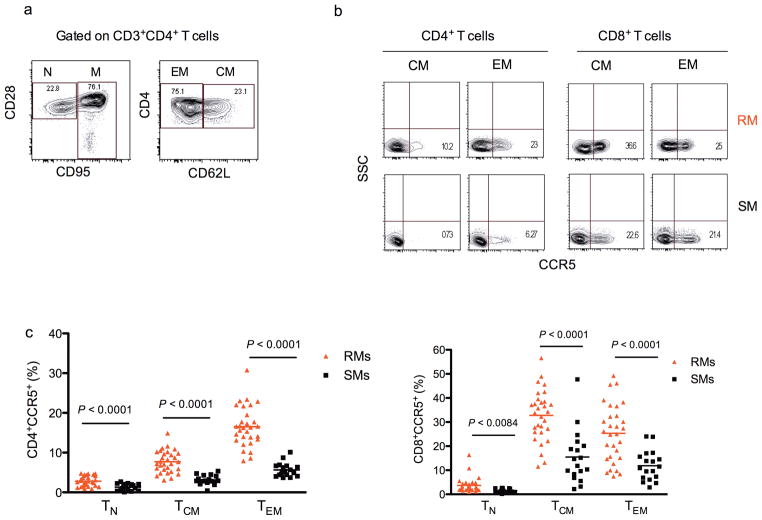
We previously showed that the fraction of CD4+CCR5+ T-cells is lower in natural SIV hosts compared to humans and RMs20. To further investigate this phenomenon, we first assessed the fraction of naive (TN, CD28+CD95−), central memory (TCM, CD95+CD62L+) and effector memory (TEM, CD95+CD62L−) CD4+ and CD8+ T-cells of uninfected RMs and SMs that express CCR5 on their surface (Fig. 1a,b)9,22,23. We found that the fraction of CCR5+ cells was significantly lower in all CD4+ T-cell subsets of SMs when compared to RMs (p<0.0001 for TN, TCM, and TEM, Fig. 1c). Of note, in both species CD4+ TEM cells expressed higher levels of CCR5 when compared to TCM (p<0.001), with the lowest CCR5 expression on TN (Fig. 1c). Lower fractions of CCR5+ cells were also observed for all subsets of CD8+ T cells of SMs as compared to RMs (p=0.0084 for TN; p<0.0001 for TCM and TEM, Figure 1c). Collectively, these data indicate that reduced expression of CCR5 is a phenomenon that involves all CD4+ T-cell subsets of SMs, including the CD4+ TCM, that, in contrast to RM CD4+ TCM, are by and large CCR5 negative.
Figure 1. The fraction of CCR5+ cells ex vivo is significantly lower in all subsets of CD4+ T-cells of SMs as compared to RMs.
The fraction of CCR5+ cells was determined in the naive (TN), central memory (TCM) and effector memory (TEM) subsets of CD4+ and CD8+ T-cells of 18 SIV uninfected sooty mangabeys (SMs) and 30 SIV uninfected rhesus macaques (RMs). (a) TN, TCM, and TEM cells were defined based on the expression of the surface markers CD28, CD95, and CD62L, as showed for CD4+ T-cells in a representative SM. (b) Staining of CCR5 on TCM and TEM CD4+ and CD8+ cells in a representative SM and a representative RM. (c) Fraction of CD4+ (left graph) or CD8+ (right graph) T-cells that express CCR5 in 18 uninfected SMs (■) and 30 uninfected RMs (
 ). Statistical analyses were performed to compare, in SMs versus RMs, the fraction of CCR5+ cells within each CD4+ and CD8+ T-cell subset.
). Statistical analyses were performed to compare, in SMs versus RMs, the fraction of CCR5+ cells within each CD4+ and CD8+ T-cell subset.
Limited CCR5 expression upon in vitro activation in CD4+ T-cells of SMs
To determine how in vitro activation affects CCR5 expression, we next measured the fraction of CD4+CCR5+ T-cells in PBMCs isolated from uninfected SMs and RMs that were stimulated with mitogens (ConA/IL-2) for eight days. CD4+ T-cells from SMs and RMs became activated and proliferated to similar extent, as determined by the expression of the proliferating marker Ki-67 (Fig. 2a) and the activation marker CD69 (data not shown). In contrast, the fraction of CD4+ T-cells expressing CCR5 was remarkably different between SMs and RMs. At baseline, the fraction of CD4+CCR5+ T-cells was higher in RMs, but a transient down-regulation of CCR5 expression in RMs resulted in similar fractions of CD4+CCR5+ T-cells by 24-hours post-stimulation. Importantly, at later time points the fraction of CD4+CCR5+ T-cells increased only in RMs, becoming significantly higher than in SMs by 120 hours post-stimulation (Fig. 2b,c). This phenomenon was not restricted to CD4+ T-cells, as a similar, albeit less dramatic difference in activation-induced CCR5 up-regulation was observed for CD8+ T-cells (Fig. S1).
Figure 2. CCR5 expression upon in vitro activation and proliferation is significantly lower in CD4+ T-cells of SMs than RMs.
Fractions of CD4+CCR5+ T-cells were determined in PBMCs from uninfected SMs (■) and RMs (
 ) after in vitro stimulations. Fraction of CD4+Ki-67+ (a) and CD4+CCR5+ (b) T-cells following stimulation with ConA/IL-2. Asterisks indicate time points when the fraction of CD4+CCR5+ T-cells was significantly lower in SMs than RMs (p values are detailed in the Result section). (c) Representative dot plots showing the fraction of CD4+CCR5+ T-cells post-stimulation in RMs (top) and SMs (bottom). (d) Flow cytometry dot plots showing Ki-67/CCR5 double staining in a representative RM and SM at 120 hours post stimulation with ConA/IL-2. (e) PBMCs isolated from RMs and SMs were labeled with CFSE prior to mitogen stimulation; levels of CCR5 were analyzed on cells expressing various levels of CFSE dilution at 120 hours post-stimulation. (f) and (g) PBMCs isolated from SMs and RMs were stimulated with recombinant IL-7, and the fraction of CD4+Ki-67+ (f) and CD4+CCR5+ (g) T-cells determined following stimulation. Asterisks indicate time points where, in RMs, the IL-7-induced increase in CD4+CCR5+ T-cells (as compared to baseline) was statistically significant (p values are detailed in the Result section). (h) In a subset of animals, levels of CCR5 mRNA (expressed as CCR5/GAPDH ratio) were determined on purified CD4+ T-cells at 0, 24, 72 and 120 hours post-stimulation with ConA/IL-2. Statistical analyses were performed to compare CCR5/GAPDH ratio between SMs and RMs.
) after in vitro stimulations. Fraction of CD4+Ki-67+ (a) and CD4+CCR5+ (b) T-cells following stimulation with ConA/IL-2. Asterisks indicate time points when the fraction of CD4+CCR5+ T-cells was significantly lower in SMs than RMs (p values are detailed in the Result section). (c) Representative dot plots showing the fraction of CD4+CCR5+ T-cells post-stimulation in RMs (top) and SMs (bottom). (d) Flow cytometry dot plots showing Ki-67/CCR5 double staining in a representative RM and SM at 120 hours post stimulation with ConA/IL-2. (e) PBMCs isolated from RMs and SMs were labeled with CFSE prior to mitogen stimulation; levels of CCR5 were analyzed on cells expressing various levels of CFSE dilution at 120 hours post-stimulation. (f) and (g) PBMCs isolated from SMs and RMs were stimulated with recombinant IL-7, and the fraction of CD4+Ki-67+ (f) and CD4+CCR5+ (g) T-cells determined following stimulation. Asterisks indicate time points where, in RMs, the IL-7-induced increase in CD4+CCR5+ T-cells (as compared to baseline) was statistically significant (p values are detailed in the Result section). (h) In a subset of animals, levels of CCR5 mRNA (expressed as CCR5/GAPDH ratio) were determined on purified CD4+ T-cells at 0, 24, 72 and 120 hours post-stimulation with ConA/IL-2. Statistical analyses were performed to compare CCR5/GAPDH ratio between SMs and RMs.
The reduced fraction of SM CD4+CCR5+ T-cells upon in vitro activation was also evident when double staining for Ki-67/CCR5 at 120 hours post-stimulation (Fig. 2d). To further define the relationship between cell proliferation and CCR5 up-regulation, we CFSE-labeled PBMCs and analyzed CCR5 expression on cells expressing various levels of CFSE dilution at 120 hours post-stimulation. SM and RM CD4+ T-cells diluted CFSE at similar levels, i.e., underwent the same number of cell divisions, but only in RM did a sizeable fraction of CD4+ T-cells that cycled once or twice up-regulate CCR5 expression (Fig. 2e). Intriguingly, most of the RM CD4+ T-cells that divided several times were CCR5-negative. This result suggests either that cycling CD4+CCR5+ T-cells proliferate more slowly than CD4+CCR5− T-cells, or that a fraction of cycling CD4+CCR5+ T-cells down-regulates CCR5 expression at later time points.
To next determine how stimuli other than mitogens affect CCR5 expression on CD4+ T-cells in SMs and RMs, we treated PBMCs with recombinant IL-7 (10ng/ml) as well as anti-CD3/CD28. Treatment with IL-7 resulted in a moderate increase of CD4+ T-cell proliferation in both SMs and RMs (Fig. 2f), but induced a 2–3-fold increase of the fraction of CD4+CCR5+ T-cells only in RMs (Fig. 2g). The IL-7-induced increase in CD4+CCR5+ T-cells observed in RMs was statistically significant at 72, 96, and 120 hours post-stimulation (p=0.0312; p=0.0156; p=0.0156, respectively) as compared to baseline. As previously reported in humans and macaques24,25, treatment with anti-CD3/CD28 resulted in reduced fraction of CD4+ T-cells expressing CCR5 in both species (data not shown).
To determine whether this low fraction of SM CD4+CCR5+ T-cells following in vitro activation results from reduced CCR5 transcription vs. abnormal trafficking to the cell surface, we longitudinally assessed the levels of CCR5 mRNA on purified CD4+ T-cells at 0, 24, 72, and 120 hours post-stimulation. We found that the levels of CCR5 mRNA were significantly lower (p=0.0159) in CD4+ T-cells isolated from SMs when compared to RMs at 72 and 120 hours post-stimulation (Fig. 2h). Interestingly, at 120 hours post-stimulation the levels of CCR5 mRNA correlated directly with the fraction of CD4+CCR5+ T-cells (data not shown). In all, these data indicate that SMs CD4+ T-cells are relatively resistant to up-regulate CCR5 expression upon in vitro activation, and suggest that reduced transcription of CCR5 contributes to this phenomenon.
Limited CCR5 expression upon in vivo activation in CD4+ T-cells of SMs
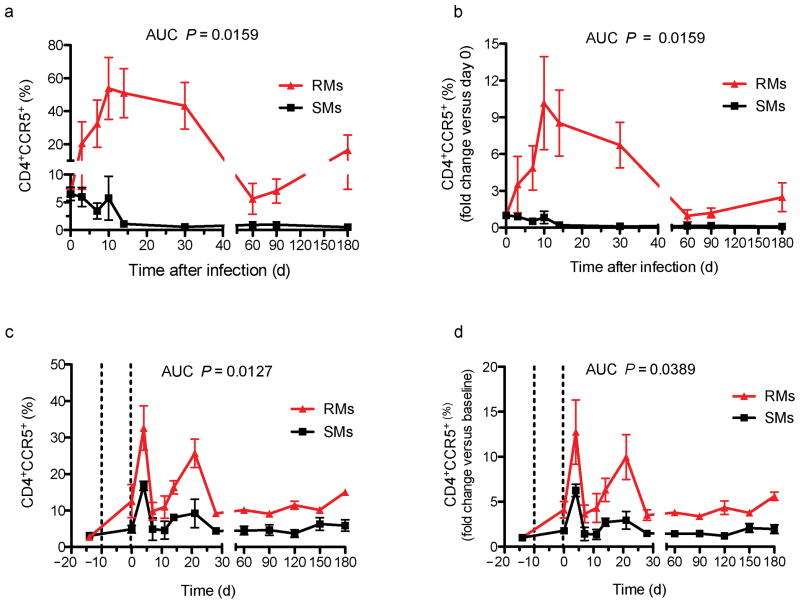
We next investigated how in vivo activation affects the fraction of CD4+CCR5+ T-cells in SMs and RMs by using two experimental conditions associated with CD4+ T-cell activation, i.e., acute SIV-infection and Ab-mediated CD4+ T-cell depletion. We first measured the levels of circulating CD4+CCR5+ T-cells during the acute phase of (i) pathogenic SIVmac239 infection of five RMs, and (ii) non-progressive SIVsmm infection of four SMs. In both species, acute SIV infection was associated with a detectable increase in T-cell activation and proliferation that lasted for 4–6 weeks4. Of note, the SIV-induced CD4+ T-cell activation was associated with increased fractions of CD4+CCR5+ T-cells in RMs, but not in SMs, with a statistically significant difference in the area under the curve (AUC) (p=0.0159) (Fig. 3). In the second experiment, monoclonal anti-CD4 antibody was administered four times over a 10-day period (dotted lines in Fig. 3c,d) to three uninfected RMs and SMs. This treatment induced a severe depletion of CD4+ T-cells followed by an increase in CD4+ T-cell proliferation that we interpreted as a homeostatic response26. Despite similar kinetics of CD4+ T-cell proliferation (data not shown), the AUC of the percentage of CD4+CCR5+ T-cells, expressed as both fraction (p=0.0127) and fold change (p=0.0389) vs. pre-depletion, was higher in RMs than SMs. The fact that CD4+ T-cells from SMs transiently up-regulated CCR5 expression early after depletion indicates that these cells are not genetically unable to express this molecule, but rather control CCR5 expression at the transcriptional and/or translational and/or post-translational level. Together with the results presented in Figure 2, these findings demonstrate that SM CD4+ T-cells are resistant to CCR5 up-regulation upon various types of in vitro and in vivo stimulation.
Figure 3. The fraction of CD4+CCR5+ T-cells upon in vivo activation is significantly lower in SMs as compared to RMs.
We determined the fraction of CD4+CCR5+ T-cells from SMs (■) and RMs (
 ) in two in vivo experimental conditions associated with activation of the CD4+ T-cell compartment, i.e., acute SIV-infection and Ab-mediated CD4+ T-cell depletion. Fraction (a) and fold change vs pre-infection, i.e. day 0 (b) of CD4+CCR5+ T-cells at different time points during (i) pathogenic SIVmac239 infection of five RMs and (ii) non-progressive experimental SIVsmm infection of four SMs. SIV-induced CD4+ T-cell activation is associated with an increased fraction of CD4+CCR5+ T-cells in RMs, but not in SMs, with a statistically significant difference in the area under the curve (AUC). Fraction (c) and fold change vs pre-depletion, i.e. day -14 (d) of CD4+CCR5+ T-cells at different time points following Ab-mediated CD4+ T-cell depletion in three uninfected RMs and three uninfected SMs. The AUC of the fraction of CD4+CCR5+ T-cells is significantly higher in RMs than SMs. The dotted lines in (c,d) indicate the 10-day period in which the anti-CD4 antibody was administered.
) in two in vivo experimental conditions associated with activation of the CD4+ T-cell compartment, i.e., acute SIV-infection and Ab-mediated CD4+ T-cell depletion. Fraction (a) and fold change vs pre-infection, i.e. day 0 (b) of CD4+CCR5+ T-cells at different time points during (i) pathogenic SIVmac239 infection of five RMs and (ii) non-progressive experimental SIVsmm infection of four SMs. SIV-induced CD4+ T-cell activation is associated with an increased fraction of CD4+CCR5+ T-cells in RMs, but not in SMs, with a statistically significant difference in the area under the curve (AUC). Fraction (c) and fold change vs pre-depletion, i.e. day -14 (d) of CD4+CCR5+ T-cells at different time points following Ab-mediated CD4+ T-cell depletion in three uninfected RMs and three uninfected SMs. The AUC of the fraction of CD4+CCR5+ T-cells is significantly higher in RMs than SMs. The dotted lines in (c,d) indicate the 10-day period in which the anti-CD4 antibody was administered.
Low fraction of CCR5+ cells after in vitro activation of SM CD4+ TCM
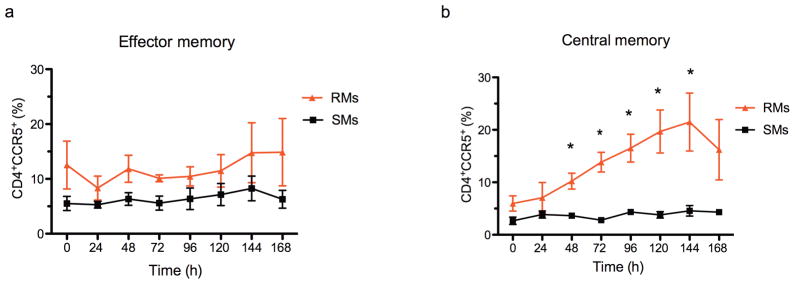
Progression to AIDS in SIVmac239-infected RMs is predicted by depletion of CD4+ TCM27–29. Here we hypothesized that reduced up-regulation of CCR5 expression on CD4+ TCM upon activation promotes the preservation of CD4+ T-cell homeostasis and the non-pathogenic outcome of SIV infection in SMs. To test this hypothesis, we longitudinally determined the fraction of CCR5+ cells in sorted CD4+ TCM and TEM after in vitro activation. While sorted CD4+ TEM from both species showed similar kinetics of CCR5 expression (Fig. 4a), only CD4+ TCM from RMs consistently up-regulated the expression of CCR5, which was significantly higher in RM than SM CD4+ TCM at 48 (p=0.0087), 72 (p=0.0043), 96 (p=0.0041), 120 (p=0.0007), and 144 (p=0.0047) hours post-stimulation (Fig. 4b). The kinetics of CCR5 expression on in vitro activated, sorted TCM and TEM of SMs and RMs for each individual monkey as measured at selected time points after activation is also presented (Fig. S2). In all, these results indicate that CD4+ TCM of SMs show a significant reduction of the activation-induced CCR5 up-regulation.
Figure 4. Lower fraction of CCR5+ cells after in vitro activation of sorted CD4+ TCM of SMs as compared to RMs.
The fractions of sorted CD4+ TCM and TEM that express CCR5 were longitudinally determined after in vitro mitogen activation in eight SIV-uninfected SMs (■) and eight SIV-uninfected RMs (
 ) TCM (CD95+CD62L+) and TEM (CD95+CD62L−) cells that were sorted and stimulated with ConA/IL-2. The graphs show the fraction of CD4+CCR5+ TEM (a) and TCM (b) cells at different time points following stimulation. Asterisks indicate time points where values are significantly higher in RMs as compared to SMs.
) TCM (CD95+CD62L+) and TEM (CD95+CD62L−) cells that were sorted and stimulated with ConA/IL-2. The graphs show the fraction of CD4+CCR5+ TEM (a) and TCM (b) cells at different time points following stimulation. Asterisks indicate time points where values are significantly higher in RMs as compared to SMs.
SM CD4+ TCM are relatively resistant to SIV infection
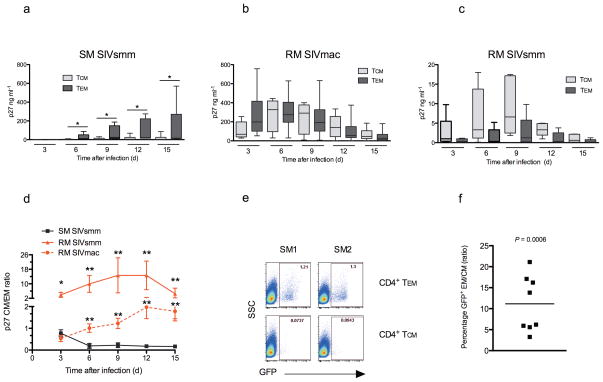
We next investigated whether the different pattern of CCR5 expression observed in CD4+ TCM and TEM of SMs and RMs is associated with different susceptibility to SIV infection. We first determined the fraction of SIV-infected cells (measured as fraction of cells expressing SIVgag DNA27) in 18 chronically SIVsmm-infected SMs and 7 chronically SIVmac239-infected RMs. We found that the fraction of SIV-infected CD4+ TCM of SMs was approximately one log lower than in RMs (p=0.0008) (Fig. 5). In contrast, no significant difference was observed between SMs and RMs in terms of SIV-infected CD4+ TEM (Fig. 5). We next sought to confirm in vitro the finding that SM CD4+ TCM are relatively resistant to SIV infection in vivo. We first purified CD4+ TCM and TEM of six uninfected SMs and infected them in vitro with a strain of SIVsmm (i.e., M949) that also replicates in primary CD4+ T-cells of RMs30. Consistent with the in vivo observation, the levels of SIV replication (i.e., p27 in the supernatant) were significantly lower in CD4+ TCM than CD4+ TEM at days 6 (p=0.0194), 9 (p=0.0304), 12 (p=0.0304), and 15 (p=0.0301) post-infection (Fig. 6a). As a control, highly purified CD4+ TCM and TEM of eleven RMs were infected in vitro using the macaque-adapted SIVmac (6 RMs) or the same virus used for SMs (SIVsmm M949; 5 RMs). In contrast to SMs, RMs showed similar (when infected with SIVmac, Fig. 6b) or even higher (when infected with SIVsmm, Fig. 6c) levels of SIV replication in CD4+ TCM than CD4+ TEM. As a result of this shift in cell targets, the TCM:TEM ratio of SIV replication was significantly higher in RMs than SMs at all time points (Fig. 6d). To further confirm the reduced susceptibility to SIV infection of SM CD4+ TCM, we infected unfractionated PBMCs from eight SMs with eGFP-expressing SIVsmm, and measured the fraction of GFP+ cells within the CD4+ TCM and TEM subsets at day 4 post-infection. We found that the fraction of GFP+ cells was ~1 log lower in SM CD4+ TCM than CD4+ TEM (p=0.0006) (Fig. 6e,f). Collectively, these findings indicate that CD4+ TCM of SMs are less susceptible, both in vivo and in vitro, to direct SIV infection as compared to both CD4+ TEM of SMs and CD4+ TCM of RMs.
Figure 5. CD4+ TCM of SMs are relatively resistant to SIV infection in vivo.
The fraction of SIV-infected CD4+ (white boxes), CD4+ TCM (light gray) and TEM (dark gray) cells, determined measuring by q-PCR the number of SIVgag DNA copies for cell equivalent, was determined in 18 naturally SIVsmm infected SMs and 7 experimentally SIVmac239 infected RMs. As showed in the graph, the fraction of SIV-infected CD4+ TCM was significantly lower in SMs than in RMs (p=0.0008), while no significant difference was observed between the two species with respect to the level of SIV-infected CD4+ TEM. Of note, in RMs the fraction of SIV-infected cells was significantly higher (p=0.03) in CD4+ TCM than TEM.
Figure 6. CD4+ TCM of SMs are relatively resistant to SIV infection in vitro.
(a–d) CD4+ TCM and TEM purified from six SMs and eleven RMs were infected in vitro with SIVsmm M949 (six SMs and five RMs), or with SIVmac (six RMs). The levels of SIV replication were determined by measuring p27 in the supernatants. In SMs (a) the levels of SIV replication were significantly lower in CD4+ TCM than CD4+ TEM at days 6, 9, 12, and 15 post-infection (as indicated by asterisks; p values are detailed in the Result section), while in RMs the levels of SIV replication in CD4+ TCM where similar to those in CD4+ TEM following infection with SIVmac (b) or higher than CD4+ TEM following infection with SIVsmm (c). As a result, the TCM vs TEM ratios of levels of SIV replication were significantly higher (as indicated by asterisks; p values are detailed in the Result section) in RMs than SMs at all tested time points (d). (e, f) Unfractionated PBMCs from eight SMs were infected with replication competent eGFP-expressing clones of SIVsmm, and the fraction of infected SM CD4+ TCM and TEM that express GFP was determined by multiparametric flow cytometry at day 4 post-infection. Figure 6e shows flow plots of GFP expression in CD4+ TCM and TEM in two representative animals. In SMs the fraction of GFP+ cells (f) is approximately 1 log lower in CD4+ TCM than CD4+ TEM (p=0.0006).
Discussion
Understanding the reasons why natural SIV hosts, such as the SMs, do not progress to AIDS despite high viremia is a key unanswered question in AIDS research, with important ramifications for HIV pathogenesis, therapy, and vaccines1,3. Here, we describe and characterize a fundamental, species-specific difference in the regulation of CCR5 expression and susceptibility to SIV infection by CD4+ TCM of SMs and RMs. We show reduced CCR5 expression in CD4+ TCM of SMs after in vitro activation, which is associated with: (i) lower levels of in vivo SIV-DNA-positive CD4+ TCM in SIV-infected SMs; and (ii) reduced permissivity to in vitro SIV infection in SM CD4+ TCM as compared to both CD4+ TEM of SMs and CD4+ TCM of RMs. Based on these results we propose that, in SIV-infected SMs, low CCR5 up-regulation upon activation protects CD4+ TCM from virus-mediated depletion, and thus favors the preservation of CD4+ T-cell homeostasis.
The association between depletion of CD4+ TCM and progression to AIDS has been elegantly shown in SIVmac-infected RMs27–29. In these studies, the Authors used a complex combination of in vivo BrdU labeling and longitudinal tissue sampling to demonstrate that while insufficient delivery of effector CD4+ T-cells to mucosal tissues is the hallmark of rapid SIV progression, depletion of CD4+ TCM during chronic infection predicts progression to AIDS better than viral load27–29. Our current data are fully consistent with a model in which a key determinant of AIDS pathogenesis is the progressive loss of CD4+ TCM homeostasis. Of note, mechanisms of CD4+ TCM loss may include not only direct virus infection, as emphasized in this study, but also bystander cell death, proliferative senescence, diminished input of naïve cells, and loss of anatomic niche14,31. Specifically, our data suggest that a fine-tuned control of CCR5 expression in SM CD4+ TCM helps protect this cell subset from SIV infection when these cells are recruited to proliferate in response to either antigenic stimulation or homeostatic stimuli.
A recent study in AGMs has shown that CD4+ T-cells of these animals down-modulate the expression of the CD4 molecule upon transition from naïve to memory cells32. In this context, our study suggests that two species of natural SIV hosts may have reached the same evolutionarily important goal, i.e., to protect CD4+ TCM from SIV infection, by down-modulating either one of the two main co-receptors for virus entry, i.e., CD4 in AGMs and CCR5 in SMs. It is tempting to speculate that this choice was dictated, in evolutionary terms, by the ability (or lack thereof) of the species-specific SIVs to use additional co-receptors, whose expression may or may not be down-regulated in CD4+ TCM. The ability of SMs to protect their CD4+ TCM cells from SIV infection may occur not only at the entry level (via regulation of CCR5 expression), but also at the level of cellular restriction factors that interfere with virus replication at the post-entry level.
The complexity of the mechanisms involved in protecting SM CD4+ TCM from SIV infection is emphasized by the observation that ~6% of SIV-infected SMs are homozygous for allelic variants of CCR5 that are not expressed on the cell surface, thus indicating that additional co-receptors are used by SIVsmm in these animals33. Whether and to what extent additional, non-CCR5-dependent mechanisms of SIV entry occur in SIV-infected SMs with wild-type CCR5 remains to be determined. In this context, it is conceivable that, in SIV-infected SMs, resistance of CD4+ TCM to viral infection and/or killing is achieved through “multiple imperfect gatekeepers” that reduce permissivity to SIV more effectively than a “single strong gatekeeper”34. A full elucidation of the molecular determinants of the species- and cell type-specific regulation of CCR5 expression on CD4+ TCM of SMs as compared to CD4+ TCM of RMs and humans is not within the scope of the current body of work. However, our data on CCR5 mRNA expression after in vitro mitogen stimulation of CD4+ T-cells of SMs and RMs suggest that this molecule is regulated, at least in part, at a transcriptional level. Whether this transcriptional regulation occurs as a result of epigenetic phenomena (i.e., DNA methylation) or is the result of different activation of specific transcription factor(s) will be object of future studies.
One of the most striking features of natural SIV infections is the presence of low levels of T-cell activation during the chronic phase of infection1,2. Based on the current set of results, we propose that the low levels of immune activation in chronically SIV-infected SMs may represent, at least in part, a consequence of the ability of these animals to reduce the level of virus infection in CD4+ TCM. In this view, the depletion of CD4+ TCM occurring during pathogenic HIV and SIV infections may trigger a vicious cycle of increased CD4+ T-cell proliferation, production of pro-inflammatory/pro-apoptotic cytokines, and virus replication. It is also conceivable that, in pathogenic infections, higher levels of virus replication in anatomic sites (i.e., lymph nodes, spleen, Peyer patches) that are both enriched in TCM and involved in the generation of innate and adaptive immune responses may contribute to the observed higher levels of chronic immune activation by dramatically increasing the local antigenic load.
While much work remains, these results strongly suggest that, in SIV-infected SMs, regulation of CCR5 expression on CD4+ TCM and protection of these cells from direct virus infection are key factors that favor CD4+ T-cell homeostasis and limit immune activation, thus identifying a novel, and potentially critical, mechanism used by these animals to avoid AIDS despite continuous virus replication.
Methods
Animals
Sixty-nine (42 females and 27 males SIV-uninfected RMs and thirty-seven (24 females and 13 males) SIV-uninfected SMs, plus twelve (all males) SIV-infected RMs and twenty-two (8 females and 14 males) SIV-infected SMs were included in this study. Animals homozygous for CCR5-null alleles were excluded from this study. All animals were housed at the Yerkes National Primate Research Center of Emory University, Atlanta, GA, and maintained in accordance with NIH guidelines. Anesthesia was used for all blood collections. All studies were approved by the University of Pennsylvania and Emory University Institutional Animal Care and Usage Committees.
Cell stimulation
Peripheral blood mononuclear cells (PBMC), isolated by density gradient centrifugation, or purified central memory (TCM) and effector memory (TEM) CD4+ T-cells were cultured at 37°C in the presence of ConA (1 μg/ml) and IL-2 (30 U/ml) at the concentration of 1×106 cells/ml. In a subset of animals, PBMC were cultured in the presence of IL-7 (10ng/ml), or with anti-CD3 plus anti-CD28. The expression of CCR5 on CD4+ and CD8+ T-cells was assessed every 24 hours by flow cytometry. To determine in vitro T-cell proliferation RM and SM PBMC were labeled with CFSE (3μM) and stimulated as described above. The fraction of CD4+ T lymphocytes that dilute CFSE and express CCR5 was measured by flow cytometry at 120 hours post-activation.
Flow cytometry
The staining for flow cytometry studies was performed according to standard procedures using monoclonal antibodies that are cross-reactive with SMs and RMs4,23. Fourteen-parameter flow cytometric analysis was performed using a LSR II flow cytometer driven by the DiVA software package (Becton Dickinson, San Jose, CA). Analysis of the acquired data was performed using FlowJo software (Tree Star, Inc., Ashland, OR).
CCR5 mRNA levels
PBMC from RMs and SMs were cultured in the presence of ConA and IL-2. CD4+ T-cells were sorted from PBMCs at 0, 24, 72, and 120 hours post-stimulation using “CD4 MicroBeads for non human primate” (Miltenyi Biotec). Total RNA was isolated from purified CD4+ T-cells using the RNAeasy extraction kit (Qiagen, Valencia, CA). The quantification of mRNAs levels for CCR5 was performed using RT-PCR on the 7900HT Real-Time PCR System (Applied Biosystems) using both SYBR-Green and TaqMan assays. Levels of CCR5 mRNAs were normalized to the expression of the housekeeping gene GAPDH. Primers that detect CCR5 expression were designed based on the available sequences in RMs and SMs. SYBR: CCR5_F 5′-AGGGCTGTGAGGCTTATCTTC-3′; CCR5_R 5′-CACCTGCATGGCTTGGTCCA-3′. TaqMan: CCR5_F 5′-GAGATTTTCAGACGTCACCA-3′; reverse: 5′-ACCCCAGATACAATCTATCCG-3′; probe: 5′-TGTACAGTCATATCAAGCTCTCTTGG-3′.
Cell sorting
Sorting of CD4+ TEM and TCM cells from RMs and SMs, both infected and uninfected, was performed on a FACSVantage SE flow cytometer (Becton Dickinson, San Jose, CA). Cells were initially gated based on light scatter, followed by positive staining for CD3 and CD4. CD4+ TEM and TCM cell subsets were gated based upon characteristic expression patterns of CD28, CD95, and CD62L (see Fig. 1a for a representative gating strategy).
Quantitative PCR for SIV gag DNA
Quantification of SIVsmm or SIVmac gag DNA was performed as previously described27. For cell number quantification, qPCR was performed simultaneously for monkey albumin gene copy number. The sequence of the forward primer for SIVsmm is 5′-GGCAGGAAAATCCCTAGCAG-3′. The reverse primer sequence is 5′-GCCCTTACTGCCTTCACTCA-3′. The probe sequence is 5′-AGTCCCTGTTCRGGCGCCAA-3′. The sequence of the forward primer for SIVmac is 5′-GTCTGCGTCAT(T/C)TGGTGCATTC-3′. The reverse primer sequence is 5′-CACTAG(C/T)TGTCTCTGCACTAT(A/G)TGTTTTG-3′. The probe sequence is 5′-CTTC(A/G)TCAGT(C/T)TGTTTCACTTTCTCTTCTGCG-3′.
In vitro SIV infections
Susceptibility of SM and RM CD4+ T-cells to in vitro SIV infection was assessed using total PBMC (in SMs) and purified CD4+ TCM and TEM (in both species). Unfractionated PBMCs from eight SMs were stimulated with ConA/IL-2 for 3 days, and then infected with a replication competent eGFP-expressing clones of SIVsmm FGb containing a IRES-eGFP fragment downstream of nef. The fraction of SM CD4+ TCM and TEM that express GFP was determined by flow cytometry at day 4 post-infection. For the experiments involving purified cell subsets, CD4+ TCM and TEM were sorted from six SMs and eleven RMs and stimulated with ConA/IL-2 for 3 days. Cells were then infected with SIVsmm M949, a strain that replicates in primary CD4+ T-cells of both species30, or SIVmac, by spin inoculation. Cells were then washed in PBS and maintained in media containing IL-2 (50 U/ml). Cell supernatants were collected before infection (day 0) and at days 3, 6, 9, 12 and 15 after infection, and replication measured by SIV Gag p27 ELISA (Advanced BioScience Laboratories).
Statistical analyses
The performed analyses for comparisons between groups include the one-way ANOVA and two-tailed t test or Mann-Whitney test. In addition, the area under the curve (AUC) has been determined in the longitudinal studies of experimental SIV infection and CD4+ T-cell depletion (Fig. 3). Significance was assessed at p<0.05 levels. All analyses were performed using the Prism 5.0 software.
Supplementary Material
Acknowledgments
The authors wish to thank Dr. Louis Picker for helpful discussions; Francois Villinger, Beth Cramer, Benton Lawson, Drs. Farida Shaheen and John “Chip” Tilton for their technical support; and Stephanie Ehnert and Tracy Meeker for their assistance with animal studies. We are grateful to the Emory CFAR Virology Core and the Penn CFAR Virology Core for their assistance. This work was supported by grant R01-AI66998 and P01-AI76074 (to G.S), R56-AI087186 (to M.P.), and P51-RR051 (to YNPRC).
Footnotes
Author Contributions
Contribution: M.P., B.C., and G.S. designed the study and wrote the paper, with contributions from the other authors as appropriate; M.P., B.C., and E.R.A. performed the immunophenotypic analyses, analyzed results, prepared the figures; L.M., A.M.O., and A. C. helped in processing the samples and analyzing the data; C.V. and J.M.B. performed the quantitative PCR for SIV gag DNA; S.N.G. provided the data on experimentally SIV-infected SMs and RMs; S.E.B. and N.F. determined CCR5 mRNA levels; P.L.H. performed the sorting experiments; T.S., M.L.C., and M.P.D. contributed to the design of the study and statistical analyses; J.E. supervised the housing and care of the animals and contributed to the design of the study and sample collection; J.M and F.K. provided the GFP-expressing SIV reporter virus; C.A., I.P., N.E.R., F.K., and R.G.C. contributed to the study design and preparation of the manuscript.
Conflict-of-interest disclosure: The authors declare no competing financial interests.
References
- 1.Paiardini MPI, Apetrei C, Silvestri G. Lessons learned from the natural hosts of HIV-related viruses. Annu Rev Med. 2009;60:485–495. doi: 10.1146/annurev.med.60.041807.123753. [DOI] [PubMed] [Google Scholar]
- 2.Pandrea I, Sodora DL, Silvestri G, Apetrei C. Into the wild: simian immunodeficiency virus (SIV) infection in natural hosts. Trends Immunol. 2008;29:419–428. doi: 10.1016/j.it.2008.05.004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Sodora DL, et al. Toward an AIDS vaccine: lessons from natural simian immunodeficiency virus infections of African nonhuman primate hosts. Nat Med. 2009;15:861–865. doi: 10.1038/nm.2013. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Gordon SN, et al. Severe depletion of mucosal CD4+ T cells in AIDS-free simian immunodeficiency virus-infected sooty mangabeys. J Immunol. 2007;179:3026–3034. doi: 10.4049/jimmunol.179.5.3026. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Silvestri G, et al. Divergent host responses during primary simian immunodeficiency virus SIVsm infection of natural sooty mangabey and nonnatural rhesus macaque hosts. J Virol. 2005;79:4043–4054. doi: 10.1128/JVI.79.7.4043-4054.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Bosinger SE, et al. Global genomic analysis reveals rapid control of a robust innate response in SIV-infected sooty mangabeys. J Clin Invest. 2009;119:3556–3572. doi: 10.1172/JCI40115. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Harris LD, et al. Downregulation of robust acute type I interferon responses distinguishes nonpathogenic simian immunodeficiency virus (SIV) infection of natural hosts from pathogenic SIV infection of rhesus macaques. J Virol. 2010;84:7886–7891. doi: 10.1128/JVI.02612-09. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Brenchley JM, et al. Differential Th17 CD4 T-cell depletion in pathogenic and nonpathogenic lentiviral infections. Blood. 2008;112:2826–2835. doi: 10.1182/blood-2008-05-159301. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Sumpter B, et al. Correlates of preserved CD4(+) T cell homeostasis during natural, nonpathogenic simian immunodeficiency virus infection of sooty mangabeys: implications for AIDS pathogenesis. J Immunol. 2007;178:1680–1691. doi: 10.4049/jimmunol.178.3.1680. [DOI] [PubMed] [Google Scholar]
- 10.Gordon SN, et al. Short-lived infected cells support virus replication in sooty mangabeys naturally infected with simian immunodeficiency virus: implications for AIDS pathogenesis. J Virol. 2008;82:3725–3735. doi: 10.1128/JVI.02408-07. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Brenchley JM, et al. Microbial translocation is a cause of systemic immune activation in chronic HIV infection. Nat Med. 2006;12:1365–1371. doi: 10.1038/nm1511. [DOI] [PubMed] [Google Scholar]
- 12.Estes JD, et al. Early resolution of acute immune activation and induction of PD-1 in SIV-infected sooty mangabeys distinguishes nonpathogenic from pathogenic infection in rhesus macaques. J Immunol. 2008;180:6798–6807. doi: 10.4049/jimmunol.180.10.6798. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Silvestri G, et al. Nonpathogenic SIV infection of sooty mangabeys is characterized by limited bystander immunopathology despite chronic high-level viremia. Immunity. 2003;18:441–452. doi: 10.1016/s1074-7613(03)00060-8. [DOI] [PubMed] [Google Scholar]
- 14.Douek DC, Roederer M, Koup RA. Emerging concepts in the immunopathogenesis of AIDS. Annu Rev Med. 2009;60:471–484. doi: 10.1146/annurev.med.60.041807.123549. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Sodora DL, Silvestri G. Immune activation and AIDS pathogenesis. AIDS. 2008;22:439–446. doi: 10.1097/QAD.0b013e3282f2dbe7. [DOI] [PubMed] [Google Scholar]
- 16.Schindler M, et al. Nef-mediated suppression of T cell activation was lost in a lentiviral lineage that gave rise to HIV-1. Cell. 2006;125:1055–1067. doi: 10.1016/j.cell.2006.04.033. [DOI] [PubMed] [Google Scholar]
- 17.Mandl JN, et al. Divergent TLR7 and TLR9 signaling and type I interferon production distinguish pathogenic and nonpathogenic AIDS virus infections. Nat Med. 2008;14:1077–1087. doi: 10.1038/nm.1871. [DOI] [PubMed] [Google Scholar]
- 18.Kornfeld C, et al. Antiinflammatory profiles during primary SIV infection in African green monkeys are associated with protection against AIDS. J Clin Invest. 2005;115:1082–1091. doi: 10.1172/JCI23006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Estes JD, et al. Early resolution of acute immune activation and induction of PD-1 in SIV-infected sooty mangabeys distinguishes nonpathogenic from pathogenic infection in rhesus macaques. J Immunol. 2008;180:6798–6807. doi: 10.4049/jimmunol.180.10.6798. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Pandrea I, et al. Paucity of CD4+CCR5+ T cells is a typical feature of natural SIV hosts. Blood. 2007;109:1069–1076. doi: 10.1182/blood-2006-05-024364. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Rey-Cuille MA, et al. Simian immunodeficiency virus replicates to high levels in sooty mangabeys without inducing disease. J Virol. 1998;72:3872–3886. doi: 10.1128/jvi.72.5.3872-3886.1998. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Pitcher CJ, et al. Development and homeostasis of T cell memory in rhesus macaque. J Immunol. 2002;168:29–43. doi: 10.4049/jimmunol.168.1.29. [DOI] [PubMed] [Google Scholar]
- 23.Paiardini M, et al. Bone marrow-based homeostatic proliferation of mature T cells in nonhuman primates: implications for AIDS pathogenesis. Blood. 2009;113:612–621. doi: 10.1182/blood-2008-06-159442. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Mummidi S, et al. Production of specific mRNA transcripts, usage of an alternate promoter, and octamer-binding transcription factors influence the surface expression levels of the HIV coreceptor CCR5 on primary T cells. J Immunol. 2007;178:5668–5681. doi: 10.4049/jimmunol.178.9.5668. [DOI] [PubMed] [Google Scholar]
- 25.Onlamoon N, et al. Optimization of in vitro expansion of macaque CD4 T cells using anti-CD3 and co-stimulation for autotransfusion therapy. J Med Primatol. 2006;35:178–193. doi: 10.1111/j.1600-0684.2006.00182.x. [DOI] [PubMed] [Google Scholar]
- 26.Engram JC, et al. Lineage-specific T-cell reconstitution following in vivo CD4+ and CD8+ lymphocyte depletion in non-human primates. Blood. 2010;116:748–758. doi: 10.1182/blood-2010-01-263814. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Okoye A, et al. Progressive CD4+ central memory T cell decline results in CD4+ effector memory insufficiency and overt disease in chronic SIV infection. J Exp Med. 2007;204:2171–2185. doi: 10.1084/jem.20070567. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Letvin NL, et al. Preserved CD4+ central memory T cells and survival in vaccinated SIV-challenged monkeys. Science. 2006;312:1530–1533. doi: 10.1126/science.1124226. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Mattapallil JJ, et al. Vaccination preserves CD4 memory T cells during acute simian immunodeficiency virus challenge. J Exp Med. 2006;203:1533–1541. doi: 10.1084/jem.20060657. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Gautam R, et al. In vitro characterization of primary SIVsmm isolates belonging to different lineages. In vitro growth on rhesus macaque cells is not predictive for in vivo replication in rhesus macaques. Virology. 2007;362:257–270. doi: 10.1016/j.virol.2006.12.037. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Zeng M, et al. Cumulative mechanisms of lymphoid tissue fibrosis and T cell depletion in HIV-1 and SIV infections. J Clin Invest. 2011;121:998–1008. doi: 10.1172/JCI45157. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Beaumier CM, et al. CD4 downregulation by memory CD4+ T cells in vivo renders African green monkeys resistant to progressive SIVagm infection. Nat Med. 2009;15:879–885. doi: 10.1038/nm.1970. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Riddick NE, et al. A Novel CCR5 Mutation Common in Sooty Mangabeys Reveals SIVsmm Infection of CCR5-Null Natural Hosts and Efficient Alternative Coreceptor Use In Vivo. PLoS Pathogens. 2010;6:e1001064. doi: 10.1371/journal.ppat.1001064. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Margolis L, Shattock R. Selective transmission of CCR5-utilizing HIV-1: the ‘gatekeeper’ problem resolved? Nat Rev Microbiol. 2006;4:312–317. doi: 10.1038/nrmicro1387. [DOI] [PubMed] [Google Scholar]
- 35.Palacios E, et al. Parallel evolution of CCR5-null phenotypes in humans and in a natural host of simian immunodeficiency virus. Curr Biol. 1998;8:943–946. doi: 10.1016/s0960-9822(07)00378-8. [DOI] [PubMed] [Google Scholar]
- 36.Novembre FJ, et al. Isolation and characterization of a neuropathogenic simian immunodeficiency virus derived from a sooty mangabey. J Virol. 1998;72:8841–8851. doi: 10.1128/jvi.72.11.8841-8851.1998. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.