Abstract
In the title compound, C12H11N3O2, the benzotriazole ring system is approximately planar [maximum deviation = 0.008 (1) Å] and its mean plane is oriented at a dihedral angle of 24.05 (4)° with respect to the furan ring. In the crystal, O—H⋯N hydrogen bonds link the molecules into chains along the ac diagonal. π–π stacking between the furan rings, between the triazole and benzene rings, and between the benzene rings [centroid–centroid distances = 3.724 (1), 3.786 (1) and 3.8623 (9) Å] are also observed.
Related literature
For general background to the biological activity of benzotriazole derivatives, see: Hirokawa et al. (1998 ▶); Yu et al. (2003 ▶); Kopanska et al. (2004 ▶). For related structures, see: Caira et al. (2004 ▶); Katritzky et al. (2001 ▶); Özel Güven et al. (2008 ▶, 2010 ▶, 2011 ▶); Nanjunda Swamy et al. (2006 ▶).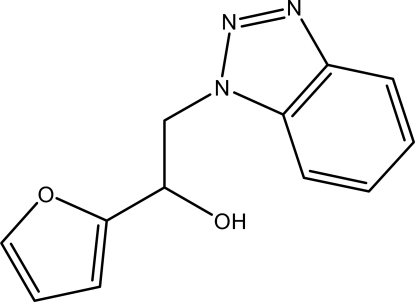
Experimental
Crystal data
C12H11N3O2
M r = 229.24
Monoclinic,
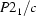
a = 11.3606 (4) Å
b = 11.1034 (4) Å
c = 8.7860 (2) Å
β = 96.938 (2)°
V = 1100.16 (6) Å3
Z = 4
Mo Kα radiation
μ = 0.10 mm−1
T = 120 K
0.50 × 0.50 × 0.20 mm
Data collection
Bruker–Nonius KappaCCD diffractometer
Absorption correction: multi-scan (SADABS; Sheldrick, 2007 ▶) T min = 0.953, T max = 0.981
12372 measured reflections
2531 independent reflections
2166 reflections with I > 2σ(I)
R int = 0.037
Refinement
R[F 2 > 2σ(F 2)] = 0.054
wR(F 2) = 0.139
S = 1.11
2531 reflections
155 parameters
H-atom parameters constrained
Δρmax = 0.58 e Å−3
Δρmin = −0.55 e Å−3
Data collection: COLLECT (Nonius, 1998 ▶); cell refinement: DENZO (Otwinowski & Minor, 1997 ▶) and COLLECT; data reduction: DENZO and COLLECT; program(s) used to solve structure: SHELXS97 (Sheldrick, 2008 ▶); program(s) used to refine structure: SHELXL97 (Sheldrick, 2008 ▶); molecular graphics: ORTEP-3 for Windows (Farrugia, 1997 ▶); software used to prepare material for publication: WinGX (Farrugia, 1999 ▶) and PLATON (Spek, 2009 ▶).
Supplementary Material
Crystal structure: contains datablock(s) I, global. DOI: 10.1107/S1600536811051798/xu5402sup1.cif
Structure factors: contains datablock(s) I. DOI: 10.1107/S1600536811051798/xu5402Isup2.hkl
Supplementary material file. DOI: 10.1107/S1600536811051798/xu5402Isup3.cml
Additional supplementary materials: crystallographic information; 3D view; checkCIF report
Table 1. Hydrogen-bond geometry (Å, °).
| D—H⋯A | D—H | H⋯A | D⋯A | D—H⋯A |
|---|---|---|---|---|
| O1—H1⋯N3i | 0.82 | 2.26 | 2.7968 (18) | 123 |
Symmetry code: (i) 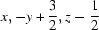 .
.
Acknowledgments
The authors acknowledge the Zonguldak Karaelmas University Research Fund (project No. 2010-13-02-05).
supplementary crystallographic information
Comment
Azole compounds have important biological activities. Benzotriazol derivatives also exhibit a good degree of analgesic, anti-inflammatory, diuretic, antiviral and antihypertensive activities (Kopanska et al., 2004; Yu et al., 2003; Hirokawa et al., 1998). Crystal structures of similar compounds like 1-phenyl-2-(1H-1,2,4-triazol-1-yl)ethanol (Özel Güven et al., 2008), 2-(1H-benzotriazol-1-yl)-1-phenylethanol (Özel Güven et al., 2010), 2-(1H-benzotriazol-1-yl)-3-(2,6-dichlorophenyl)-1-phenylpropan-1-ol (Özel Güven et al., 2011), fluconazole (Caira et al., 2004), and other benzotriazole ring possesing compounds (Katritzky et al., 2001; Nanjunda Swamy et al., 2006) have been reported before. Now, we report herein the crystal structure of the title alcohol, (I).
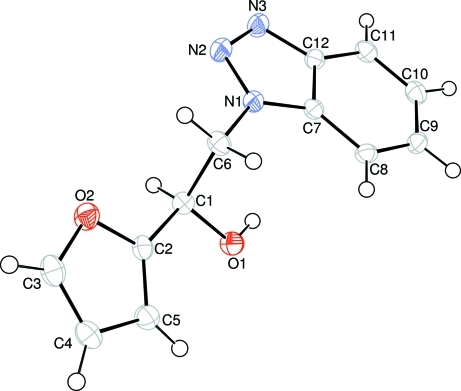
In the molecule of the title compound (Fig. 1), the bond lengths and angles are generally within normal ranges. The planar benzotriazole ring [B (N1-N3/C7-C12)] is oriented with respect to the furan [A (O2/C2-C5)] ring at a dihedral angle of A/B = 24.05 (4)°. Atom C6 is 0.043 (2) Å away from the plane of the benzotriazole ring and atoms C1 and O1 are 0.010 (2) and 0.043 (1) Å away from the plane of the furan ring, respectively.
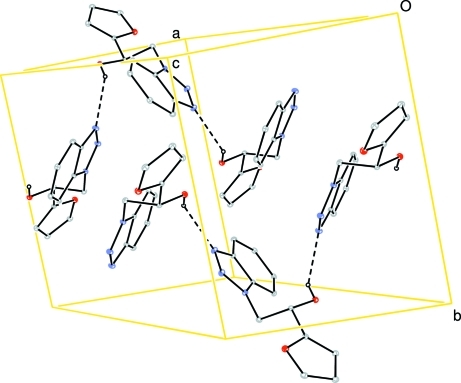
In the crystal, O—H···N hydrogen bonds (table 1) link the molecules into chains (Fig. 2). There also exist π···π contacts between the furan rings, between the triazole and benzene rings and between the benzene rings, Cg1—Cg1i, Cg2—Cg3ii and Cg3—Cg3ii, may further stabilize the structure [centroid-centroid distances = 3.724 (1), 3.786 (1) and 3.8623 (9) Å; symmetry codes: (i) 1 - x, 1 - y, 1 - z; (ii) -x, 1 - y, 1 - z; Cg1, Cg2 and Cg3 are the centroids of the rings A (O2/C2-C5), C (N1-N3/C7/C12) and D (C7-C12), respectively].
Experimental
The title compound, (I), was synthesized by reduction of 2-(1H-benzotriazol-1-yl)-1-(furan-2-yl)ethanone with sodiumborohydrate. A mixture of 2-(1H-benzotriazol-1-yl)-1-(furan-2-yl)ethanone (1010 mg, 4.44 mmol) and sodium borohydrate (561 mg, 8.89 mmol) in ethanol (50 ml) was refluxed for 4 h. After evaporation of the solvent, the mixture was neutralized with dilute HCl, and then refluxed for 30 min. After the mixture was cooled, the solution was alkalinized with dilute NaOH and the resulting precipitate was filtered. The filtrate was extracted with chloroform, then the organic phase was dried and evaporated. The residue was crystallized from 2-propanol to obtain colorless crystals suitable for X-ray analysis (yield; 634 mg, 62%).
Refinement
H atoms were positioned geometrically with O—H = 0.82 Å (for OH group), C—H = 0.98, 0.93 and 0.97 Å for methine, aromatic and methylene H, respectively, and constrained to ride on their parent atoms, with Uiso(H) = k × Ueq(C,O), where k = 1.5 for OH H-atom and k = 1.2 for all other H-atoms.
Figures
Fig. 1.
The molecular structure of the title compound with the atom-numbering scheme. Displacement ellipsoids are drawn at the 50% probability level.
Fig. 2.
A partial packing diagram. Hydrogen bonds are shown as dashed lines. Hydrogen atoms not involved in hydrogen bonding have been omitted for clarity.
Crystal data
| C12H11N3O2 | F(000) = 480 |
| Mr = 229.24 | Dx = 1.384 Mg m−3 |
| Monoclinic, P21/c | Mo Kα radiation, λ = 0.71073 Å |
| Hall symbol: -P 2ybc | Cell parameters from 6399 reflections |
| a = 11.3606 (4) Å | θ = 2.9–27.5° |
| b = 11.1034 (4) Å | µ = 0.10 mm−1 |
| c = 8.7860 (2) Å | T = 120 K |
| β = 96.938 (2)° | Block, colorless |
| V = 1100.16 (6) Å3 | 0.50 × 0.50 × 0.20 mm |
| Z = 4 |
Data collection
| Bruker–Nonius KappaCCD diffractometer | 2531 independent reflections |
| Radiation source: fine-focus sealed tube | 2166 reflections with I > 2σ(I) |
| graphite | Rint = 0.037 |
| φ and ω scans | θmax = 27.5°, θmin = 3.0° |
| Absorption correction: multi-scan (SADABS; Sheldrick, 2007) | h = −14→14 |
| Tmin = 0.953, Tmax = 0.981 | k = −14→14 |
| 12372 measured reflections | l = −10→11 |
Refinement
| Refinement on F2 | Secondary atom site location: difference Fourier map |
| Least-squares matrix: full | Hydrogen site location: inferred from neighbouring sites |
| R[F2 > 2σ(F2)] = 0.054 | H-atom parameters constrained |
| wR(F2) = 0.139 | w = 1/[σ2(Fo2) + (0.0748P)2 + 0.4779P] where P = (Fo2 + 2Fc2)/3 |
| S = 1.11 | (Δ/σ)max < 0.001 |
| 2531 reflections | Δρmax = 0.58 e Å−3 |
| 155 parameters | Δρmin = −0.55 e Å−3 |
| 0 restraints | Extinction correction: SHELXL97 (Sheldrick, 2008), Fc*=kFc[1+0.001xFc2λ3/sin(2θ)]-1/4 |
| Primary atom site location: structure-invariant direct methods | Extinction coefficient: 0.144 (12) |
Special details
| Geometry. All esds (except the esd in the dihedral angle between two l.s. planes) are estimated using the full covariance matrix. The cell esds are taken into account individually in the estimation of esds in distances, angles and torsion angles; correlations between esds in cell parameters are only used when they are defined by crystal symmetry. An approximate (isotropic) treatment of cell esds is used for estimating esds involving l.s. planes. |
| Refinement. Refinement of F2 against ALL reflections. The weighted R-factor wR and goodness of fit S are based on F2, conventional R-factors R are based on F, with F set to zero for negative F2. The threshold expression of F2 > 2sigma(F2) is used only for calculating R-factors(gt) etc. and is not relevant to the choice of reflections for refinement. R-factors based on F2 are statistically about twice as large as those based on F, and R- factors based on ALL data will be even larger. |
Fractional atomic coordinates and isotropic or equivalent isotropic displacement parameters (Å2)
| x | y | z | Uiso*/Ueq | ||
| O1 | 0.20677 (10) | 0.98199 (10) | 0.08354 (12) | 0.0223 (3) | |
| H1 | 0.1559 | 0.9346 | 0.1048 | 0.033* | |
| O2 | 0.50236 (10) | 1.06715 (11) | 0.24705 (13) | 0.0247 (3) | |
| N1 | 0.18510 (11) | 0.91870 (12) | 0.39959 (14) | 0.0196 (3) | |
| N2 | 0.22565 (12) | 0.82297 (13) | 0.48488 (16) | 0.0249 (3) | |
| N3 | 0.13604 (12) | 0.75376 (13) | 0.50674 (16) | 0.0248 (3) | |
| C1 | 0.30653 (14) | 0.97510 (14) | 0.19621 (17) | 0.0193 (3) | |
| H1A | 0.3392 | 0.8933 | 0.1989 | 0.023* | |
| C2 | 0.39723 (13) | 1.06237 (14) | 0.15318 (17) | 0.0194 (3) | |
| C3 | 0.57056 (15) | 1.15175 (15) | 0.18424 (19) | 0.0257 (4) | |
| H3 | 0.6471 | 1.1732 | 0.2247 | 0.031* | |
| C4 | 0.51146 (15) | 1.19920 (15) | 0.05628 (19) | 0.0252 (4) | |
| H4 | 0.5385 | 1.2581 | −0.0062 | 0.030* | |
| C5 | 0.39773 (14) | 1.14020 (15) | 0.03582 (18) | 0.0238 (4) | |
| H5 | 0.3367 | 1.1533 | −0.0429 | 0.029* | |
| C6 | 0.26911 (14) | 1.00691 (14) | 0.35398 (17) | 0.0214 (3) | |
| H6A | 0.3386 | 1.0091 | 0.4298 | 0.026* | |
| H6B | 0.2330 | 1.0862 | 0.3496 | 0.026* | |
| C7 | 0.06500 (13) | 0.91156 (13) | 0.36343 (16) | 0.0178 (3) | |
| C8 | −0.01985 (14) | 0.98551 (14) | 0.27982 (17) | 0.0202 (3) | |
| H8 | 0.0010 | 1.0560 | 0.2326 | 0.024* | |
| C9 | −0.13571 (14) | 0.94734 (15) | 0.27193 (17) | 0.0226 (4) | |
| H9 | −0.1949 | 0.9939 | 0.2182 | 0.027* | |
| C10 | −0.16792 (14) | 0.83958 (15) | 0.34283 (17) | 0.0232 (4) | |
| H10 | −0.2474 | 0.8174 | 0.3339 | 0.028* | |
| C11 | −0.08469 (14) | 0.76679 (14) | 0.42472 (18) | 0.0222 (4) | |
| H11 | −0.1059 | 0.6962 | 0.4713 | 0.027* | |
| C12 | 0.03398 (13) | 0.80499 (13) | 0.43403 (17) | 0.0191 (3) |
Atomic displacement parameters (Å2)
| U11 | U22 | U33 | U12 | U13 | U23 | |
| O1 | 0.0207 (6) | 0.0231 (6) | 0.0226 (6) | −0.0047 (4) | 0.0004 (4) | −0.0002 (4) |
| O2 | 0.0204 (6) | 0.0259 (6) | 0.0271 (6) | −0.0032 (4) | −0.0001 (4) | 0.0002 (4) |
| N1 | 0.0181 (6) | 0.0205 (6) | 0.0205 (6) | 0.0001 (5) | 0.0030 (5) | 0.0020 (5) |
| N2 | 0.0224 (7) | 0.0249 (7) | 0.0272 (7) | 0.0045 (5) | 0.0023 (5) | 0.0044 (5) |
| N3 | 0.0231 (7) | 0.0221 (7) | 0.0293 (7) | 0.0043 (5) | 0.0038 (5) | 0.0059 (6) |
| C1 | 0.0204 (7) | 0.0172 (7) | 0.0203 (7) | −0.0007 (6) | 0.0028 (6) | −0.0012 (5) |
| C2 | 0.0168 (7) | 0.0204 (7) | 0.0213 (7) | 0.0006 (6) | 0.0031 (6) | −0.0038 (6) |
| C3 | 0.0203 (8) | 0.0251 (8) | 0.0322 (8) | −0.0052 (6) | 0.0054 (6) | −0.0052 (6) |
| C4 | 0.0246 (8) | 0.0237 (8) | 0.0285 (8) | −0.0047 (6) | 0.0087 (6) | −0.0016 (6) |
| C5 | 0.0223 (8) | 0.0264 (8) | 0.0228 (7) | −0.0012 (6) | 0.0029 (6) | 0.0007 (6) |
| C6 | 0.0201 (7) | 0.0221 (8) | 0.0223 (7) | −0.0039 (6) | 0.0041 (6) | −0.0025 (6) |
| C7 | 0.0185 (7) | 0.0178 (7) | 0.0173 (7) | −0.0002 (6) | 0.0034 (5) | −0.0017 (5) |
| C8 | 0.0248 (8) | 0.0178 (7) | 0.0184 (7) | 0.0020 (6) | 0.0040 (6) | 0.0030 (5) |
| C9 | 0.0213 (8) | 0.0274 (8) | 0.0185 (7) | 0.0056 (6) | 0.0002 (6) | −0.0009 (6) |
| C10 | 0.0190 (7) | 0.0286 (8) | 0.0224 (7) | −0.0031 (6) | 0.0043 (6) | −0.0042 (6) |
| C11 | 0.0243 (8) | 0.0191 (8) | 0.0244 (8) | −0.0022 (6) | 0.0074 (6) | −0.0004 (6) |
| C12 | 0.0212 (8) | 0.0169 (7) | 0.0195 (7) | 0.0022 (6) | 0.0041 (6) | 0.0004 (5) |
Geometric parameters (Å, °)
| O1—C1 | 1.4139 (18) | C4—H4 | 0.9300 |
| O1—H1 | 0.8200 | C5—C4 | 1.440 (2) |
| O2—C2 | 1.3681 (19) | C5—H5 | 0.9300 |
| O2—C3 | 1.375 (2) | C6—H6A | 0.9700 |
| N1—N2 | 1.3492 (18) | C6—H6B | 0.9700 |
| N1—C6 | 1.4571 (19) | C7—C12 | 1.401 (2) |
| N1—C7 | 1.365 (2) | C8—C7 | 1.404 (2) |
| N3—N2 | 1.308 (2) | C8—C9 | 1.376 (2) |
| N3—C12 | 1.377 (2) | C8—H8 | 0.9300 |
| C1—C6 | 1.539 (2) | C9—H9 | 0.9300 |
| C1—H1A | 0.9800 | C10—C9 | 1.417 (2) |
| C2—C1 | 1.496 (2) | C10—H10 | 0.9300 |
| C2—C5 | 1.346 (2) | C11—C10 | 1.379 (2) |
| C3—C4 | 1.345 (2) | C11—C12 | 1.406 (2) |
| C3—H3 | 0.9300 | C11—H11 | 0.9300 |
| C1—O1—H1 | 109.5 | N1—C6—C1 | 110.77 (12) |
| C2—O2—C3 | 106.13 (12) | N1—C6—H6A | 109.5 |
| N2—N1—C7 | 110.36 (12) | N1—C6—H6B | 109.5 |
| N2—N1—C6 | 119.43 (12) | C1—C6—H6A | 109.5 |
| C7—N1—C6 | 130.13 (13) | C1—C6—H6B | 109.5 |
| N3—N2—N1 | 108.96 (13) | H6A—C6—H6B | 108.1 |
| N2—N3—C12 | 108.40 (13) | N1—C7—C8 | 133.73 (14) |
| O1—C1—C2 | 107.84 (12) | N1—C7—C12 | 104.11 (13) |
| O1—C1—C6 | 109.45 (12) | C12—C7—C8 | 122.15 (14) |
| O1—C1—H1A | 109.6 | C7—C8—H8 | 122.0 |
| C2—C1—C6 | 110.64 (12) | C9—C8—C7 | 115.98 (14) |
| C2—C1—H1A | 109.6 | C9—C8—H8 | 122.0 |
| C6—C1—H1A | 109.6 | C8—C9—C10 | 122.27 (15) |
| O2—C2—C1 | 116.78 (13) | C8—C9—H9 | 118.9 |
| C5—C2—O2 | 110.66 (14) | C10—C9—H9 | 118.9 |
| C5—C2—C1 | 132.56 (14) | C9—C10—H10 | 119.1 |
| O2—C3—H3 | 124.6 | C11—C10—C9 | 121.81 (15) |
| C4—C3—O2 | 110.75 (14) | C11—C10—H10 | 119.1 |
| C4—C3—H3 | 124.6 | C10—C11—C12 | 116.47 (14) |
| C3—C4—C5 | 106.01 (14) | C10—C11—H11 | 121.8 |
| C3—C4—H4 | 127.0 | C12—C11—H11 | 121.8 |
| C5—C4—H4 | 127.0 | N3—C12—C7 | 108.16 (13) |
| C2—C5—C4 | 106.45 (14) | N3—C12—C11 | 130.53 (15) |
| C2—C5—H5 | 126.8 | C7—C12—C11 | 121.31 (14) |
| C4—C5—H5 | 126.8 | ||
| C6—N1—N2—N3 | −177.41 (13) | C5—C2—C1—O1 | 0.9 (2) |
| C7—N1—N2—N3 | −0.44 (17) | C5—C2—C1—C6 | −118.73 (19) |
| N2—N1—C6—C1 | 92.27 (16) | O2—C2—C5—C4 | −0.09 (18) |
| C7—N1—C6—C1 | −84.02 (19) | C1—C2—C5—C4 | −179.58 (15) |
| N2—N1—C7—C8 | 179.68 (16) | O2—C3—C4—C5 | −0.27 (18) |
| N2—N1—C7—C12 | 0.62 (16) | C2—C5—C4—C3 | 0.21 (18) |
| C6—N1—C7—C8 | −3.8 (3) | N1—C7—C12—N3 | −0.58 (16) |
| C6—N1—C7—C12 | 177.17 (14) | N1—C7—C12—C11 | 179.06 (14) |
| C12—N3—N2—N1 | 0.05 (17) | C8—C7—C12—N3 | −179.77 (13) |
| N2—N3—C12—C7 | 0.34 (17) | C8—C7—C12—C11 | −0.1 (2) |
| N2—N3—C12—C11 | −179.25 (15) | C9—C8—C7—N1 | −178.60 (15) |
| C3—O2—C2—C1 | 179.51 (13) | C9—C8—C7—C12 | 0.3 (2) |
| C3—O2—C2—C5 | −0.07 (17) | C7—C8—C9—C10 | −0.4 (2) |
| C2—O2—C3—C4 | 0.22 (18) | C11—C10—C9—C8 | 0.3 (2) |
| O1—C1—C6—N1 | 64.08 (16) | C10—C11—C12—N3 | 179.57 (15) |
| C2—C1—C6—N1 | −177.23 (12) | C10—C11—C12—C7 | 0.0 (2) |
| O2—C2—C1—O1 | −178.53 (12) | C12—C11—C10—C9 | −0.1 (2) |
| O2—C2—C1—C6 | 61.80 (17) |
Hydrogen-bond geometry (Å, °)
| D—H···A | D—H | H···A | D···A | D—H···A |
| O1—H1···N3i | 0.82 | 2.26 | 2.7968 (18) | 123 |
Symmetry codes: (i) x, −y+3/2, z−1/2.
Footnotes
Supplementary data and figures for this paper are available from the IUCr electronic archives (Reference: XU5402).
References
- Caira, M. R., Alkhamis, K. A. & Obaidat, R. M. (2004). J. Pharm. Sci. 93, 601–611. [DOI] [PubMed]
- Farrugia, L. J. (1997). J. Appl. Cryst. 30, 565.
- Farrugia, L. J. (1999). J. Appl. Cryst. 32, 837–838.
- Hirokawa, Y., Yamazaki, H., Yoshida, N. & Kato, S. (1998). Bioorg. & Med. Chem. Lett. 8, 1973–1978. [DOI] [PubMed]
- Katritzky, A. R., Zhang, S. M., Kurz, T., Wang, M. Y. & Steel, P. J. (2001). Org. Lett. 3, 2807–2809. [DOI] [PubMed]
- Kopanska, K., Najda, A., Zebrowska, J., Chomicz, L., Piekarczyk, J., Myjak, P. & Bretner, M. (2004). Bioorg. Med. Chem. 12, 2617–2624. [DOI] [PubMed]
- Nanjunda Swamy, S., Basappa, Sarala, G., Priya, B. S., Gaonkar, S. L., Shashidhara Prasad, J. & Rangappa, K. S. (2006). Bioorg. Med. Chem. Lett. 16, 999–1004. [DOI] [PubMed]
- Nonius (1998). COLLECT Nonius BV, Delft, The Netherlands.
- Otwinowski, Z. & Minor, W. (1997). Methods in Enzymology, Vol. 276, Macromolecular Crystallography, Part A, edited by C. W. Carter Jr & R. M. Sweet, pp. 307-326. New York: Academic Press.
- Özel Güven, Ö., Bayraktar, M., Coles, S. J. & Hökelek, T. (2010). Acta Cryst. E66, o959. [DOI] [PMC free article] [PubMed]
- Özel Güven, Ö., Çapanlar, S., Coles, S. J. & Hökelek, T. (2011). Acta Cryst. E67, o2510. [DOI] [PMC free article] [PubMed]
- Özel Güven, Ö., Tahtacı, H., Coles, S. J. & Hökelek, T. (2008). Acta Cryst. E64, o1254. [DOI] [PMC free article] [PubMed]
- Sheldrick, G. M. (2007). SADABS Bruker AXS Inc., Madison, Wisconsin, USA.
- Sheldrick, G. M. (2008). Acta Cryst. A64, 112–122. [DOI] [PubMed]
- Spek, A. L. (2009). Acta Cryst. D65, 148–155. [DOI] [PMC free article] [PubMed]
- Yu, K. L., Zhang, Y., Civiello, R. L., Kadow, K. F., Cianci, C., Krystal, M. & Meanwell, N. A. (2003). Bioorg. Med. Chem. Lett. 13, 2141–2144. [DOI] [PubMed]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Crystal structure: contains datablock(s) I, global. DOI: 10.1107/S1600536811051798/xu5402sup1.cif
Structure factors: contains datablock(s) I. DOI: 10.1107/S1600536811051798/xu5402Isup2.hkl
Supplementary material file. DOI: 10.1107/S1600536811051798/xu5402Isup3.cml
Additional supplementary materials: crystallographic information; 3D view; checkCIF report