Abstract
A sea urchin histone gene fragment containing inverted regions of the normal repeat has been cloned in pBR322. Restriction enzyme mapping and homoduplex analysis of this fragment indicate that the H1-H4 spacer of one repeat is situated alongside the inverted H2A-H1 spacer of another repeat. The site of the breakpoint has been sequenced and compared with homologous stretches of the normal repeat. The breakpoints in the original duplexes were found to be within 4-6 bp of the H4 mRNA cap site and 8-10 bp of the H1 mRNA cap site in the standard repeat. The breakpoints in both original duplexes contain short direct repeats and an overlap of 3 bases. As this is the first breakpoint resulting in the apposition of inverted sequence to be analyzed at the level of DNA sequence, we speculate whether structural features described here are typical of such rearrangements. The structure observed is consistent with, but does not prove, that the sequence is the endpoint of a true inversion since only one junction has been isolated and characterized.
Full text
PDF
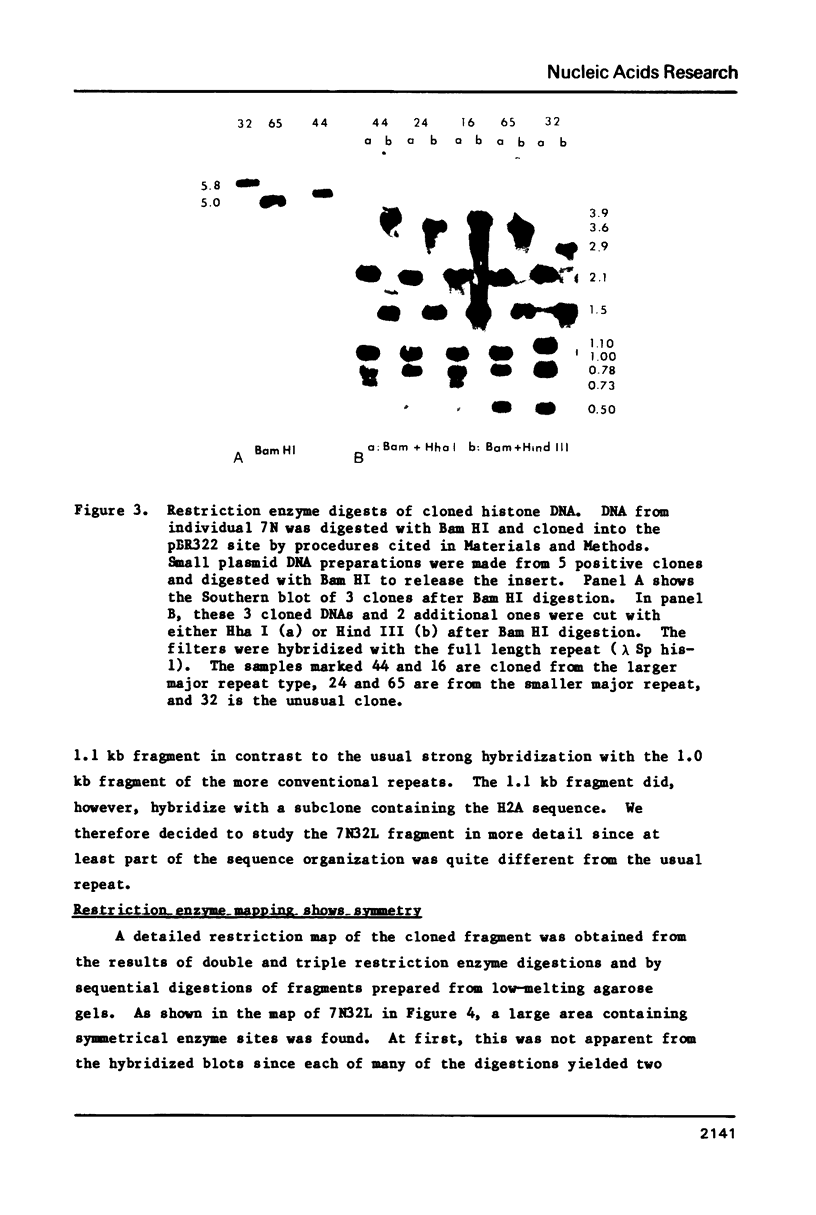
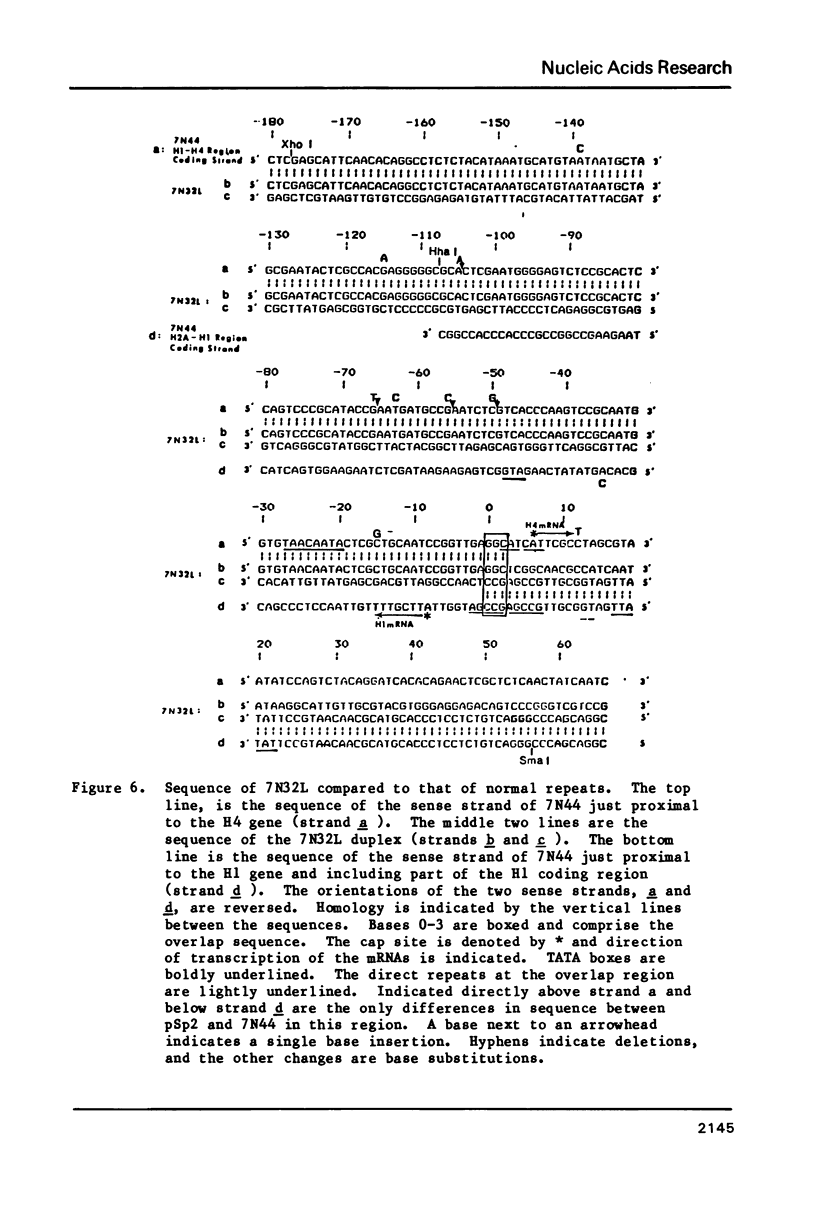


















Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Albertini A. M., Hofer M., Calos M. P., Miller J. H. On the formation of spontaneous deletions: the importance of short sequence homologies in the generation of large deletions. Cell. 1982 Jun;29(2):319–328. doi: 10.1016/0092-8674(82)90148-9. [DOI] [PubMed] [Google Scholar]
- Birnboim H. C., Doly J. A rapid alkaline extraction procedure for screening recombinant plasmid DNA. Nucleic Acids Res. 1979 Nov 24;7(6):1513–1523. doi: 10.1093/nar/7.6.1513. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Childs G., Maxson R., Cohn R. H., Kedes L. Orphons: dispersed genetic elements derived from tandem repetitive genes of eucaryotes. Cell. 1981 Mar;23(3):651–663. doi: 10.1016/0092-8674(81)90428-1. [DOI] [PubMed] [Google Scholar]
- Cohn R. H., Lowry J. C., Kedes L. H. Histone genes of the sea urchin (S. purpuratus) cloned in E coli: order, polarity, and strandedness of the five histone-coding and spacer regions. Cell. 1976 Sep;9(1):147–161. doi: 10.1016/0092-8674(76)90060-x. [DOI] [PubMed] [Google Scholar]
- Collins M., Rubin G. M. Structure of the Drosophila mutable allele, white-crimson, and its white-ivory and wild-type derivatives. Cell. 1982 Aug;30(1):71–79. doi: 10.1016/0092-8674(82)90013-7. [DOI] [PubMed] [Google Scholar]
- Dunsmuir P., Brorein W. J., Jr, Simon M. A., Rubin G. M. Insertion of the Drosophila transposable element copia generates a 5 base pair duplication. Cell. 1980 Sep;21(2):575–579. doi: 10.1016/0092-8674(80)90495-x. [DOI] [PubMed] [Google Scholar]
- Farabaugh P. J., Schmeissner U., Hofer M., Miller J. H. Genetic studies of the lac repressor. VII. On the molecular nature of spontaneous hotspots in the lacI gene of Escherichia coli. J Mol Biol. 1978 Dec 25;126(4):847–857. doi: 10.1016/0022-2836(78)90023-2. [DOI] [PubMed] [Google Scholar]
- Grindley N. D., Lauth M. R., Wells R. G., Wityk R. J., Salvo J. J., Reed R. R. Transposon-mediated site-specific recombination: identification of three binding sites for resolvase at the res sites of gamma delta and Tn3. Cell. 1982 Aug;30(1):19–27. doi: 10.1016/0092-8674(82)90007-1. [DOI] [PubMed] [Google Scholar]
- Grunstein M., Diamond K. E., Knoppel E., Grunstein J. E. Comparison of the early histone H4 gene sequence of Strongylocentrotus purpuratus with maternal, early, and late histone H4 mRNA sequences. Biochemistry. 1981 Mar 3;20(5):1216–1223. doi: 10.1021/bi00508a025. [DOI] [PubMed] [Google Scholar]
- Grunstein M. Hatching in the sea urchin Lytechinus pictus is accompanied by a shift in histone H4 gene activity. Proc Natl Acad Sci U S A. 1978 Sep;75(9):4135–4139. doi: 10.1073/pnas.75.9.4135. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Grunstein M., Hogness D. S. Colony hybridization: a method for the isolation of cloned DNAs that contain a specific gene. Proc Natl Acad Sci U S A. 1975 Oct;72(10):3961–3965. doi: 10.1073/pnas.72.10.3961. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hentschel C. C., Birnstiel M. L. The organization and expression of histone gene families. Cell. 1981 Aug;25(2):301–313. doi: 10.1016/0092-8674(81)90048-9. [DOI] [PubMed] [Google Scholar]
- Joseph D. R., Stafford D. W. Purification of sea urchin ribosomal RNA genes with a single-strand specific nuclease. Biochim Biophys Acta. 1976 Jan 19;418(2):167–174. doi: 10.1016/0005-2787(76)90066-6. [DOI] [PubMed] [Google Scholar]
- Kedes L. H., Cohn R. H., Lowry J. C., Chang A. C., Cohen S. N. The organization of sea urchin histone genes. Cell. 1975 Nov;6(3):359–369. doi: 10.1016/0092-8674(75)90185-3. [DOI] [PubMed] [Google Scholar]
- Kunkel N. S., Weinberg E. S. Histone gene transcripts in the cleavage and mesenchyme blastula embryo of the sea urchin, S. purpuratus. Cell. 1978 Jun;14(2):313–326. doi: 10.1016/0092-8674(78)90117-4. [DOI] [PubMed] [Google Scholar]
- Levis R., Collins M., Rubin G. M. FB elements are the common basis for the instability of the wDZL and wC Drosophila mutations. Cell. 1982 Sep;30(2):551–565. doi: 10.1016/0092-8674(82)90252-5. [DOI] [PubMed] [Google Scholar]
- Levis R., Rubin G. M. The unstable wDZL mutation of Drosophila is caused by a 13 kilobase insertion that is imprecisely excised in phenotypic revertants. Cell. 1982 Sep;30(2):543–550. doi: 10.1016/0092-8674(82)90251-3. [DOI] [PubMed] [Google Scholar]
- Levy S., Sures I., Kedes L. The nucleotide and amino acid coding sequence of a gene for H1 histone that interacts with euchromatin. The early embryonic H1 gene of the sea urchin Strongylocentrotus purpuratus. J Biol Chem. 1982 Aug 25;257(16):9438–9443. [PubMed] [Google Scholar]
- Lifton R. P., Goldberg M. L., Karp R. W., Hogness D. S. The organization of the histone genes in Drosophila melanogaster: functional and evolutionary implications. Cold Spring Harb Symp Quant Biol. 1978;42(Pt 2):1047–1051. doi: 10.1101/sqb.1978.042.01.105. [DOI] [PubMed] [Google Scholar]
- Maniatis T., Jeffrey A., Kleid D. G. Nucleotide sequence of the rightward operator of phage lambda. Proc Natl Acad Sci U S A. 1975 Mar;72(3):1184–1188. doi: 10.1073/pnas.72.3.1184. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Maxam A. M., Gilbert W. Sequencing end-labeled DNA with base-specific chemical cleavages. Methods Enzymol. 1980;65(1):499–560. doi: 10.1016/s0076-6879(80)65059-9. [DOI] [PubMed] [Google Scholar]
- Ohtsubo H., Ohtsubo E. Nucleotide sequence of an insertion element, IS1. Proc Natl Acad Sci U S A. 1978 Feb;75(2):615–619. doi: 10.1073/pnas.75.2.615. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Overton G. C., Weinberg E. S. Length and sequence heterogeneity of the histone gene repeat unit of the sea urchin, S. purpuratus. Cell. 1978 Jun;14(2):247–257. doi: 10.1016/0092-8674(78)90111-3. [DOI] [PubMed] [Google Scholar]
- Spradling A. C., Rubin G. M. Drosophila genome organization: conserved and dynamic aspects. Annu Rev Genet. 1981;15:219–264. doi: 10.1146/annurev.ge.15.120181.001251. [DOI] [PubMed] [Google Scholar]
- Stafford D. W., Bieber D. Concentration of DNA solutions by extraction with 2-butanol. Biochim Biophys Acta. 1975 Jan 6;378(1):18–21. doi: 10.1016/0005-2787(75)90132-x. [DOI] [PubMed] [Google Scholar]
- Sures I., Levy S., Kedes L. H. Leader sequences of Strongylocentrotus purpuratus histone mRNAs start at a unique heptanucleotide common to all five histone genes. Proc Natl Acad Sci U S A. 1980 Mar;77(3):1265–1269. doi: 10.1073/pnas.77.3.1265. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Weinberg E. S., Overton G. C., Shutt R. H., Reeder R. H. Histone gene arrangement in the sea urchin, Strongylocentrotus purpuratus. Proc Natl Acad Sci U S A. 1975 Dec;72(12):4815–4819. doi: 10.1073/pnas.72.12.4815. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zachar Z., Bingham P. M. Regulation of white locus expression: the structure of mutant alleles at the white locus of Drosophila melanogaster. Cell. 1982 Sep;30(2):529–541. doi: 10.1016/0092-8674(82)90250-1. [DOI] [PubMed] [Google Scholar]