Abstract
Munc119 (also denoted as RG4) is a mammalian ortholog of the Caenorhabditis elegans protein unc119 and is essential for vision and synaptic transmission at photoreceptor ribbon synapses by unknown molecular mechanisms. Munc119/RG4 is related to the prenyl-binding protein PrBP/δ and expressed at high levels in photoreceptor ribbon synapses. Synaptic ribbons are presynaptic specializations in the active zone of these tonically active synapses and contain RIBEYE as a unique and major component. In the present study, we identified Munc119 as a RIBEYE-interacting protein at photoreceptor ribbon synapses using five independent approaches. The PrBP/δ homology domain of Munc119 is essential for the interaction with the NADH binding region of RIBEYE(B) domain. But RIBEYE-Munc119 interaction does not depend on NADH binding. A RIBEYE point mutant (RE(B)E844Q) that no longer interacted with Munc119 still bound NADH, arguing that binding of Munc119 and NADH to RIBEYE are independent from each other. Our data indicate that Munc119 is a synaptic ribbon-associated component. We show that Munc119 can be recruited to synaptic ribbons via its interaction with RIBEYE. Our data suggest that the RIBEYE-Munc119 interaction is essential for synaptic transmission at the photoreceptor ribbon synapse.
Munc119 (also denoted as RG4, Ref. 1) is a mammalian ortholog of the Caenorhabditis elegans protein unc119 and essential for normal vision and synaptic transmission at photoreceptor synapses (1–3). Munc119/RG4 was initially identified by a differential display screen and shown to be expressed at high levels in photoreceptor synapses (1–3). Munc119 consists of an N-terminal, 77-amino acid long proline-rich region, and a 163-amino acid long C-terminal domain that shares significant sequence homology to the prenyl-binding protein PrBP/δ3 (previously also denoted as the δ-subunit of photoreceptor cGMP-dependent phosphodiesterase (PDE6D) (4–6). The C-terminal PrBP/δ homology domain of Munc119 is highly conserved between species and is essential for Munc119 function (1–3). The essential function of Munc119 for synaptic transmission at photoreceptor synapses and for vision has been demonstrated in a cone rod dystrophy patient with a premature termination codon mutation (5). This termination codon mutation resulted in a Munc119 protein that lacked the PrBP/δ domain. Consistently, a transgenic mouse model that reproduced this premature termination codon mutation of Munc119 displayed similarly strong disturbances of synaptic transmission at photoreceptor synapses and defects in vision (5, 7, 8). The mechanism of how Munc119 works in photoreceptor synapses is not clear.
Photoreceptor synapses are mainly ribbon-type synapses (for review, see Refs. 9–11). Ribbon synapses are specialized, tonically active chemical synapses. To maintain tonic exocytosis, ribbon synapses are equipped with specialized presynaptic structures, the synaptic ribbons (for review, see Refs. 9–11). Synaptic ribbons are presynaptic structures in the active zone complex of these synapses and are associated with large amounts of synaptic vesicles (for review, see Refs. 9–11). The protein RIBEYE is a major component of synaptic ribbons and exclusively localized to these structures (12–14). RIBEYE consists of a unique A-domain and a B-domain, which is largely identical to CtBP2 (12). RIBEYE(B) domain belongs to a family of d-isomer-specific 2-hydroxy acid dehydrogenases and binds NAD(H) with high affinity (12). Structural analyses of the CtBP protein family that also includes RIBEYE (for review, see Ref. 15) revealed the presence of two distinct subdomains: a central NADH binding domain (NBD) and a substrate binding domain (SBD) (16, 17). The RIBEYE-specific substrate that binds to the SBD is not yet known as well as the precise physiological function of RIBEYE in the synapse.
To better understand the physiological role and molecular composition of synaptic ribbons, we performed a YTH screen using the RIBEYE (B) domain as a bait. In this screen, we identified Munc119 as a potential RIBEYE-interacting protein.
EXPERIMENTAL PROCEDURES
Plasmids—Details on all plasmids are deposited as supplemental materials.
Yeast Two-hybrid Methods—For YTH analyses, the Gal4-based Matchmaker yeast two-hybrid system (Clontech) was used according to the manufacturer's instructions. For the YTH screening, we used a bovine retinal YTH cDNA library from the retina (18). The cDNA of the respective bait proteins were cloned in-frame with the Gal4-DNA binding domain of pGBKT7. The cDNA of the indicated prey proteins were cloned in-frame with the Gal4 activation domain of pACT2 or pGADT7. The bait and prey plasmids confer tryptophan and leucine prototrophy to the respective auxotrophic yeast strains. Yeast strains Y187 and AH109 were used that contain distinct auxotrophic marker genes: AH109 [MATa, trp1-901, leu2-3, 112, ura3-52, his3-200, Gal4Δ, gal80Δ, LYS2::GAL1UAS-GAL1TATA-HIS3, GAL2UAS-GAL2TATA-ADE2, URA3::MEL1UAS-MEL1TATA-lacZ] (19); Y187 [MATα, ura3-52, his3-200, ade2-101, trp1-901, leu2-3, 112, gal4Δ,. met, gal80Δ, URA3::GAL1UAS-GAL1TATA-lacZ] (20). Bait plasmids were electroporated into AH109 yeast, prey plasmids into Y187 yeast. Preparation of electrocompetent yeasts and electroporation of yeasts were done as described (21). For identifying transformants, yeasts were plated on the respective selective plates to identify the resulting convertants to the respective prototrophy (drop out media Clontech/QBiogene). For interaction analyses, AH109 yeasts containing the respective bait plasmid were mated with Y187 yeasts containing the respective prey plasmid. Mating was performed for 5 h at 30 °C in 1 ml of YPD medium with heavy vortexing. For assessing mating efficiency, half of the mated sample was streaked on –LW plates, the other half was plated on –ALWH selective plate with 10 mm aminotriazole (3-amino 1,2,4-triazole, ATZ) added. For the matings, pSE1111 and pSE1112 that encode irrelevant proteins (22) as well as the empty bait and prey vectors were used as negative controls. Expression of β-galactosidase (β-gal) marker gene activity was qualitatively analyzed by filter assays and quantitatively with liquid assays as described (23, 24).
Cell Culture—COS- and R28 retinal progenitor cells were cultured as previously described (12, 25, 26). COS cells were transiently transfected with the DEAE-dextran method (12) or with lipofection using the perfectin reagent (PEQLAB) according to the manufacturer's instructions.
GST Pull-down Assays from Transfected COS Cells—COS cells were transfected with the indicated eukaryotic expression constructs (empty GSTpEBG, Munc119(1–240)-GSTpEBG and RE(B)-EGFP, see supplemental materials). For the experimental assays, Munc119(1–240)-GSTpEBG was co-transfected with RE(B)-EGFP. Empty GSTpEBG co-transfected with RE(B)-EGFP served as control assays. 48 h after transfection, the cells were collected from the plates and pelleted at 6,000 rpm for 5 min at 4 °C. All subsequent steps were performed at 4 °C if not denoted otherwise. The cell pellets were washed with ice-cold phosphate-buffered saline and lysed with 500 μl of ice-cold lysis buffer (100 mm Tris-HCl, pH 7.9, 150 mm NaCl, 1 mm EDTA containing 1% Triton X-100) for 30 min. Subsequently, the samples were centrifuged at 13,000 rpm for 15 min. The lysate from experiment and control assay were incubated overnight with 10 μl of washed glutathione-Sepharose beads each (Amersham Biosciences). After incubation, the samples were centrifuged at 13,000 rpm for 1 min, and the supernatants removed. The pellets were washed with ice-cold phosphate-buffered saline three times. The final pellets were boiled in 25 μl of SDS sample buffer and subjected to 10% SDS-PAGE followed by Western blot analyses with the indicated antibodies.
Immunoprecipitation from R28 Retinal Progenitor Cells—All steps were performed at 4 °C if not denoted otherwise. Washed R28 cell pellets were lysed with lysis buffer (100 mm Tris-HCl, pH 7.9, 150 mm NaCl, 1 mm EDTA, 1% Triton X-100) for 45 min on ice. The lysate was centrifuged twice at 13,000 rpm (15 min, Eppendorf centrifuge (Biofuge Fresco, rotor 3329), and supernatants were subsequently precleared by the addition of 20 μl of washed protein A-Sepharose beads (Sigma) and 10 μl of RIBEYE preimmune serum for 1 h. Following centrifugation (13,000 rpm, 1 min), the precleared lysate was divided into two equal aliquots and incubated either with 10 μl of control IgG (U2656 preimmune serum) or with 10 μl of anti-RIBEYE (U2656 immune serum) for overnight at 4 °C. After overnight incubation, 20 μl of washed protein A-Sepharose beads were added to the samples, and incubation was continued for another 1 h. Subsequently, samples were centrifuged and washed five times with incubation buffer. The washed protein A-Sepharose pellets were boiled in 30 μl of SDS sample buffer and analyzed by Western blot analyses as described below.
Immunoprecipitation from the Bovine Retina—All steps were performed at 4 °C if not denoted otherwise. For each immunoprecipitation, a freshly isolated bovine retina was incubated with 2 ml of lysis buffer (100 mm Tris-HCl, pH 7.9, 150 mm NaCl, 1 mm EDTA) containing 1% Triton X-100 for 30 min at 4 °C. Then the sample was centrifuged at 13,000 rpm for 15 min. Samples were transferred to 2-ml syringes and forcefully ejected through 23-gauge needles to mechanically disrupt the retinal tissue. Mechanical crushing through 23-gauge needles was repeated 40–50 times. The mechanical disruption is essential to fractionate synaptic ribbons and to make them accessible for immunoprecipitation. Without mechanical treatment, no RIBEYE was observed in the respective tissue lysate. After mechanical disruption, lysis was allowed to proceed for further 30 min on ice. Samples were centrifuged twice at 13,000 rpm for 30 min. The supernatant was incubated with 10 μl of Munc119 preimmune serum and 20 μl of washed protein A-Sepharose beads for 1 h at 4 °C. Afterward, the sample was centrifuged at 13,000 rpm for 15 min, and the precleared lysate was divided into two aliquots and incubated either with 10 μl of Munc119 immune serum (Munc119 V2T2.120) or with Munc119 preimmune serum (control IgG) together with 20 μl of washed protein A-Sepharose beads (overnight). After overnight incubation, samples were centrifuged at 3,000 rpm (2 min) to pellet the protein A-Sepharose beads. The pellet was washed three times with 1 ml of lysis buffer. The final pellet was boiled with SDS loading buffer and subjected to SDS-PAGE followed by Western blotting with the indicated antibodies.
Antibodies—The following antibodies were used in the present study: mouse monoclonal anti-GST (Sigma), mouse monoclonal anti-MBP (New England Biolabs), anti-RIBEYE(B) domain (U2656, 12), mouse monoclonal anti-CtBP2 (BD Biosciences), and polyclonal anti-EGFP (T3743; gift of Dr. Thomas C. Südhof, Dallas, TX). Full-length bovine Munc119-GST was used as an antigen to generate the immune serum Munc119 V2T2. For the experiments, immune serum at the 120th day after immunization was used. The antibody specifically detects Munc119 (supplemental Fig. S6). In extracts of R28 cells and bovine retina the Munc119 immune serum V2T2.120 specifically detected Munc119 at the expected running position at 35 kDa (Figs. 4 and 5 and supplemental Fig. S6).
FIGURE 4.
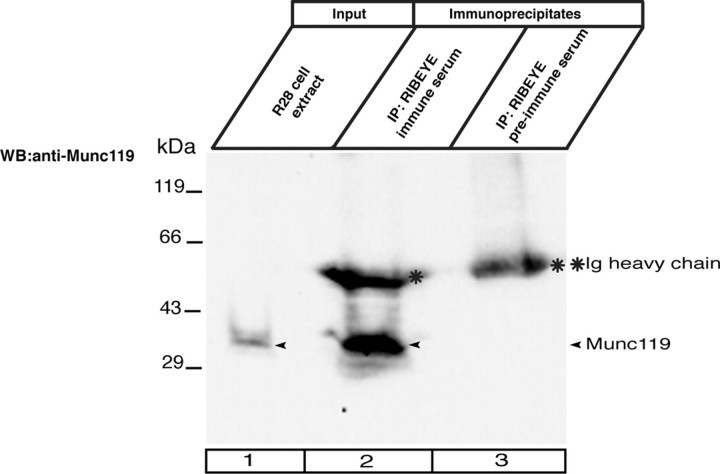
Co-immunoprecipitation of RIBEYE and Munc119 from R28 retinal precursor cells. R28 retinal progenitor cells endogenously express soluble Munc119 and RIBEYE, which can be readily solubilized from R28 cells by Triton X-100 lysis as described under “Experimental Procedures.” Munc119 was co-immunoprecipitated by antibodies against RIBEYE from extracts of R28 retinal progenitor cells (lane 2). Immunoprecipitated Munc119 is indicated by an arrowhead in lane 2. The RIBEYE preimmune serum did not co-immunoprecipitate Munc119 (lane 3) demonstrating the specificity of the co-immunoprecipitation. Lane 1 shows the input fraction (5% of total input); all of the protein A-beads with the immunoprecipitated proteins (100%) were loaded on the gel (lanes 2 and 3). Asterisks indicate the immunoglobulin heavy chains.
FIGURE 5.
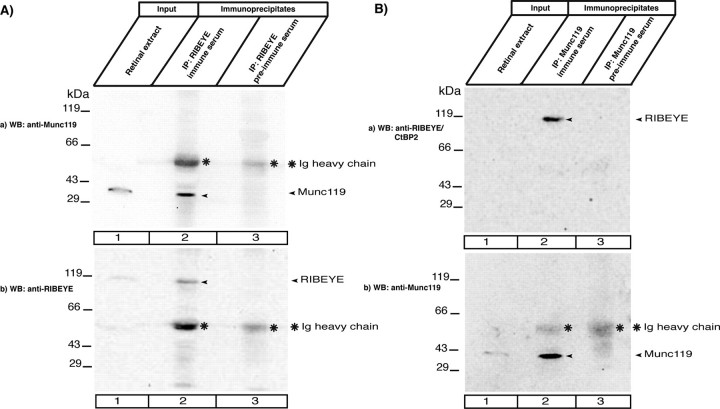
Co-immunoprecipitation of RIBEYE and Munc119 from the bovine retina. Co-immunoprecipitation of RIBEYE and Munc119 from bovine retina. In A, RIBEYE immune serum and RIBEYE preimmune serum were tested for their capability to co-immunoprecipitate Munc119. Munc119 is co-immunoprecipitated by RIBEYE immune serum (lane 2, Aa) but not by RIBEYE preimmune serum (lane 3, A). Ab, shows the same blot as in Aa but reprobed with anti-RIBEYE antibodies. This blot shows the presence of RIBEYE precipitated by the immune serum (lane 2) but not by the preimmune serum (lane 3). Asterisks indicate the immunoglobulin heavy chains. Lane 1 shows the input fraction (2% of total input). The loaded 2% input fraction corresponds roughly to 200 μg of total proteins (in a volume of ≈20 μl). Considerably more input fraction could not be loaded on the gel for volume reasons and also not to overload the gel. Furthermore, synaptic ribbons are mechanically stable, Triton X-100-insoluble structures, which can only be extracted to a certain extent from the bovine retina by the combination of mechanical and chemical lysis. Therefore, the RIBEYE immunosignal is weak in the input fractions. RIBEYE is highly enriched in the experimental immunoprecipitates (lane 2) but absent in the control immunoprecipitates (lane 3). Asterisks indicate the immunoglobulin heavy chains. 100% of the protein A-beads containing the immunoprecipitated proteins were loaded on the gel for experimental and control immunoprecipitations (lanes 2 and 3). In B, Munc119 immune serum and Munc119 preimmune serum were tested for their capability to co-immunoprecipitate RIBEYE. RIBEYE is co-immunoprecipitated by Munc119 immune serum (lane 2, Ba) but not by Munc119 preimmune serum (lane 3, Ba). Bb shows the same blot as in Ba but reprobed with anti-Munc119. This blot shows the presence of Munc119 immunoprecipitated by the immune serum but not by the preimmune serume. Asterisks indicate the immunoglobulin heavy chains.
Miscellaneous Methods—SDS-PAGE and Western blotting was performed as previously described (12). The fusion protein was expressed in BL21(DE3) as previously described (12, see also supplemental materials). Synaptic ribbons were purified as previously described (12, 27). Immunofluorescence microscopy was performed as previously described (28) using a Zeiss Axiovert 200M equipped with the respective filter sets.
RESULTS
Identification of Munc119 as a RIBEYE-interacting Protein—Using RIBEYE(B) as bait, we obtained three independent clones of Munc119 from the retinal YTH cDNA library as potential interaction partners of RIBEYE. One clone encoded full-length Munc119, the two other clones encoded truncated Munc119 proteins that started at lysine 92 (Lys-92) and isoleucine 93 (Ile-93), shortly after the beginning of the PrBP/δ-homology domain of Munc119 (Fig. 1A), suggesting that the PrBP/δ domain of Munc119 is probably responsible for the interaction. Using bait constructs that encoded for the PRD-(aa1–77)- or PrBP/δ-(aa78–240) domain of Munc119 we verified that the PrBP/δ-homology domain of Munc119 is indeed responsible for the interaction with RIBEYE (Fig. 1C). The PRD of Munc119 did not interact with RIBEYE in the YTH system. In the case of RIBEYE, the NADH binding subdomain of RIBEYE(B) domain (NBD) is mediating the interaction with Munc119. Munc119 also interacted with full-length RIBEYE indicating that the A-domain of RIBEYE is not inhibiting the interaction of the RIBEYE(B) domain with Munc119 (Fig. 1D). The interactions in the YTH were assayed by the growth of the respective mated yeast on –ALWH selective plates, indicating protein-protein interaction, as well as by qualitative and quantitative assessment of the β-galactosidase marker gene expression (Fig. 1, C and D and supplemental Fig. S1).
FIGURE 1.
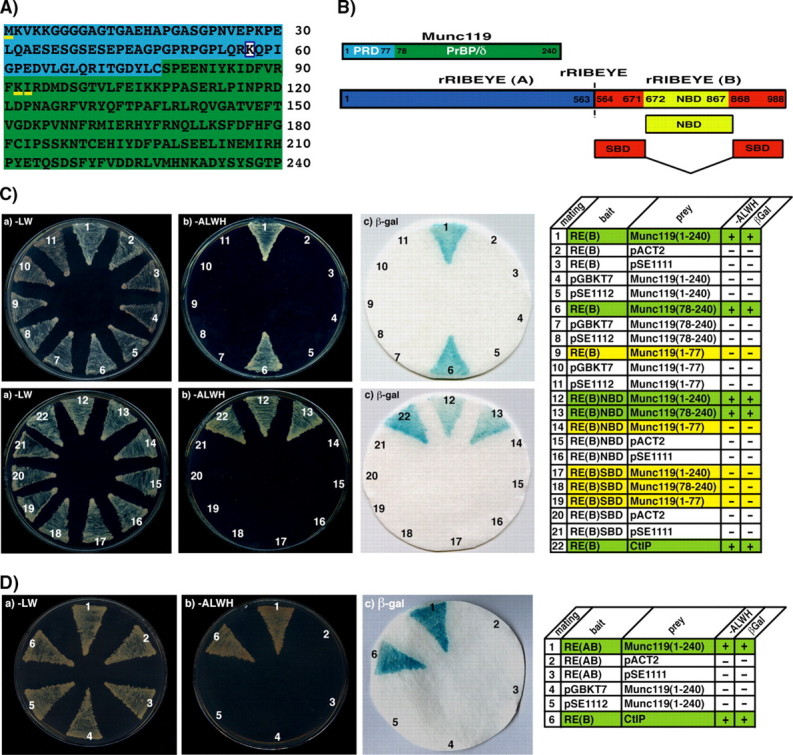
Interaction of RIBEYE(B) and RIBEYE(AB) with Munc119 in the YTH system. A, amino acid sequence of bovine Munc119. The proline-rich domain (PRD, aa1–77) is colored in blue, the PrBP/δ-homology domain of Munc119 (aa78–240) colored in green. The boxed lysine indicates the site of a premature stop mutation that causes cone rod dystrophy in a human patient (5). The amino acids methionine 1 (M1), lysine 92 (K92), and isoleucine 93 (I93), which are underlined in yellow indicate the beginning of the reading frames of three independently obtained Munc119 YTH prey clones. The amino acid sequence of Munc119 obtained in our YTH screen is identical to the Munc119 sequence deposited at GenBank™ (accession BC103449.1). B, schematic domain structures of RIBEYE and Munc119. RIBEYE contains of a large N-terminal A-domain and a C-terminal B-domain. The B-domain of RIBEYE contains the NAD(H) binding subdomain (NBD, depicted in yellow) and the substrate binding subdomain (SBD, depicted in red). C, RIBEYE(B) interacts with the PrBP/δ-homology domain of Munc119 in YTH. Summary plates of YTH analyses obtained with the indicated bait and prey plasmids. For convenience, experimental bait-prey pairs are underlayered in color (green in the case of interacting bait-prey pairs; yellow in the case of non-interacting bait-prey-pairs; control matings are non-colored). D, RIBEYE(B) and also full-length RIBEYE (RIBEYE(AB) interact with Munc119. The interaction is mediated via the NBD of RIBEYE and the PrBP/δ homology domain of Munc119, Munc119(78–240) (C; matings 6, 12–13). Mating 22 in C and mating 6 in D denote an unrelated positive control mating (CtIP). pSE1111 is an irrelevant prey vector, and pSE1112 an irrelevant bait vector (22).
Munc119 Interacts with RIBEYE in GST Pull-down Assays—We used various independent approaches to verify the Munc119-RIBEYE(B) interaction. First, we performed pull-down experiments using bacterially expressed and purified fusion protein (Fig. 2). We used GST-tagged proteins (Munc119-GST and GST) as immobilized bait proteins and MBP-tagged proteins (RIBEYE(B)-MBP and MBP) as soluble prey proteins. Munc119-GST (but not GST alone) interacted with RIBEYE(B)-MBP (but not MBP alone) as judged by protein pull-down analyses (Fig. 2). Specificity of interaction in these fusion protein pull-downs was consistently shown by SDS-PAGE and Western blot analyses (Fig. 2, A and B). Next, we analyzed whether RIBEYE(B) interacts with Munc119 in transfected COS cells. For this purpose, COS cells were co-transfected with eukaryotic expression plasmids that encoded for RIBEYE(B)-EGFP and GST-tagged Munc119 or GST alone (as control protein). Munc119-GST (but not GST alone) pulled-down RIBEYE(B)-EGFP from a crude cell extract of transfected COS cells (Fig. 3), further demonstrating a specific interaction between the RIBEYE(B) domain and Munc119.
FIGURE 2.
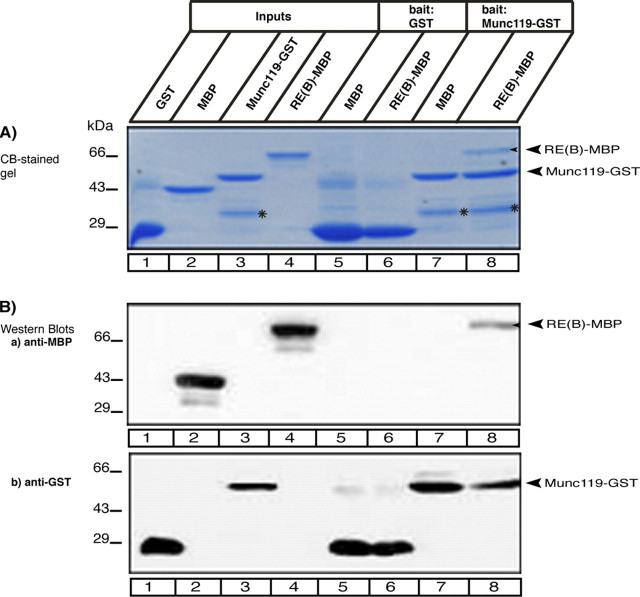
RIBEYE(B) specifically interacts with Munc119 in fusion protein pull-down assays. Pull-down analyses of RIBEYE(B)/Munc119 complexes using bacterially expressed fusion proteins. In A, pull-down experiments were analyzed by Coomassie Blue-stained polyacrylamide gel after SDS-PAGE. In B, by Western blot analyses with the indicated antibodies. A and B, lanes 1–4 show the indicated purified fusion proteins (input fractions). All input lanes, except for lane 4, represent 50% of the input fraction. Lane 4 represents 25% of the input fraction. In lanes 5–8, 100% was loaded. GST-tagged fusion proteins were used as immobilized bait proteins and MBP-tagged proteins as soluble prey proteins. Only Munc119-GST pulled-down RE(B)-MBP (lane 8) but not GST alone (lane 6). Neither GST alone nor Munc119-GST pulled-down MBP alone (lanes 5 and 7). The asterisks in lanes 3, 7, and 8 of A label a break-down product of Munc119-GST. SDS-PAGE clearly demonstrated that Munc119-GST does not pull-down MBP alone (A). To further exclude that any MBP is nonspecifically pulled-down by Munc119-GST, we also analyzed the results of the pull-down assays by Western blotting with anti-MBP antibodies. B, Western blot analyses with anti-MBP antibodies (B) clearly show that only RE(B)-MBP (lane 8) but not MBP alone (lane 7) is pulled-down by Munc119-GST. GST alone does not pull-down RE(B)-MBP as well as MBP alone as shown by Western blotting with antibodies against MBP (Ba) demonstrating the specificity of the interaction and completely confirming the results in A. In Bb, the same blot as analyzed in Ba was reprobed (after stripping) with antibodies against GST to show equal loading of the bait proteins. Abbreviations: CB, Coomassie Blue.
FIGURE 3.
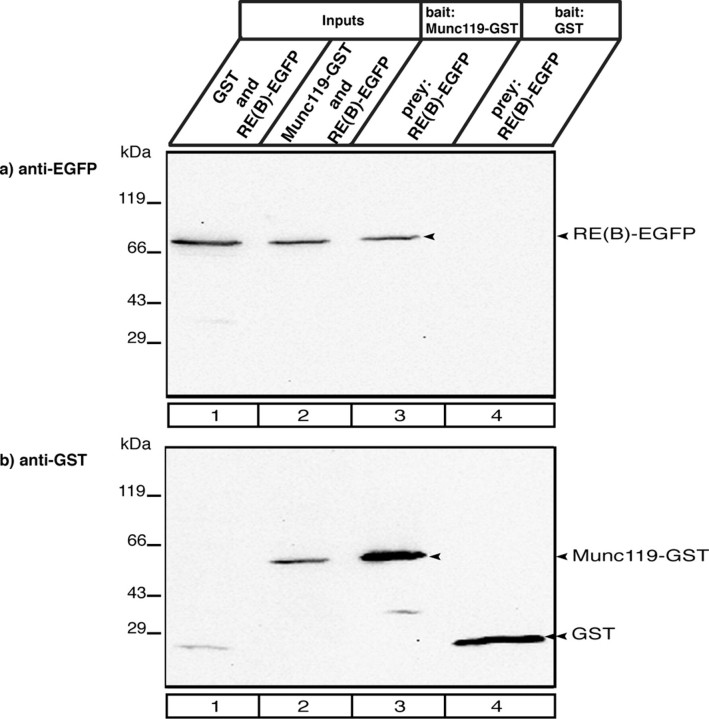
RIBEYE interacts with Munc119 in transfected COS cells. COS cells were co-transfected either with Munc119-GSTpEBG and RE(B)-EGFP (experimental assays) or with empty GSTpEBG and RE(B)-EGFP (control assays) using lipofection. Glutathione beads were added to the respective cell lysates. Proteins bound to the glutathione beads (lanes 3 and 4) were analyzed via Western blotting with antibodies against GST and EGFP. Munc119-GST pulled-down RIBEYE(B)-EGFP (lane 3) but not GST alone (lane 4) demonstrating the specific interaction between Munc119 and RIBEYE(B). Lanes 1–2 show the respective input fractions (10% of total input); Lanes 3 and 4 show 100% of the pulled-down proteins. In b, the same blot as shown in a was reprobed (after stripping) with antibodies against GST to show equal loading of the samples.
Immunoprecipitation of Endogenous Munc119 and RIBEYE from R28 Retinal Progenitor Cells and Bovine Retina—R28 is an E1A-immortalized retinal precursor cells line (25). These cells are immature, non-fully differentiated cells that express both neuronal and glial cell markers (26). R28 cells endogenously express Munc119 (Ref. 26, Fig. 4, and supplemental Fig. S6) and RIBEYE in a Triton X-100 soluble fraction (supplemental Fig. S6 and data not shown). Therefore, we used R28 cells for immunoprecipitation experiments and tested whether RIBEYE immune serum could co-immunoprecipitate Munc119 from R28 cell extracts. RIBEYE preimmune serum served as control serum. Indeed, RIBEYE immune serum co-immunoprecipitated endogenous Munc119 whereas RIBEYE preimmune serum did not (Fig. 4).
Next, we prepared extracts from bovine retina as described under “Experimental Procedures” and tested whether antibodies against RIBEYE could co-immunoprecipitate Munc119. RIBEYE immune serum (but not RIBEYE preimmune serum) co-immunoprecipitated Munc119 together with RIBEYE showing a specific interaction of these proteins also in the retina. Similarly, Munc119 immune serum (but not Munc119 preimmune serum) co-immunoprecipitated RIBEYE together with Munc119 (Fig. 5). Because RIBEYE is exclusively present at synaptic ribbons in the mature retina (12) the co-immunoprecipitation experiments suggest that Munc119 may be a component of synaptic ribbons.
Binding of Munc119 to RIBEYE(B) Is Independent of NADH Binding to RIBEYE—Previous YTH analyses demonstrated that the NAD(H) binding subdomain (NBD) of RIBEYE(B) is mediating the interaction with Munc119. Therefore, we generated point mutants of the NBD and analyzed these point mutants of the NBD for their capability to interact with Munc119 in the YTH system to further map the interaction site of Munc119 on RIBEYE(B) domain. RIBEYE(B)G730 is an essential component of the NAD(H) binding motif, and the RIBEYE(B) point mutant RIBEYE(B)G730A does not bind significant levels of NAD(H) (12).4 In contrast to the NAD(H) binding deficiency, RIBEYE(B)G730A still interacted with Munc119 in YTH analyses indicating that NAD(H) binding is not important for binding of Munc119 to RIBEYE(B) domain (Fig. 6). Similarly, the binding of Munc119 to RIBEYE(B) analyzed by biochemical pull-down analyses was not changed by the addition of either NAD+ or NADH. Increasing concentrations of both NAD+ or NADH did not significantly influence the binding of Munc119 to RIBEYE(B) (supplemental Fig. S2).
FIGURE 6.
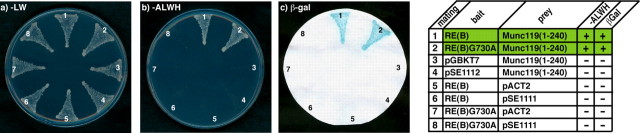
Binding of Munc119 to RIBEYE(B) is independent of NADH binding. Summary plates of YTH analyses obtained with the indicated bait and prey plasmids. For convenience, experimental bait-prey pairs are underlayered in color (green in the case of interacting bait-prey pairs; control matings are non-colored). The NADH binding deficient RIBEYE point mutant RE(B)G730A (mating 2) interacts with Munc119 in YTH indicating that NADH binding to RIBEYE is not essential for the binding of Munc119.
We generated further point mutants located on the NAD(H) binding domain of RIBEYE, namely RE(B)D758N, RE(B)E844Q, RE(B)-F848W, RE(B)K854Q, RE(B)I796A, RE(B)D820N, and RE(B)E790Q, to further map the docking site of Munc119 on the NBD of RIBEYE(B). All of these latter point mutants still interact with Munc119 except for RE(B)E844Q pointing that this amino acid is crucial for the interaction with Munc119 (supplemental Fig. S3). Glutamate E844 is located close to the NADH binding cleft of RIBEYE (12, 16, 17). As judged by NADH-dependent FRET experiments performed as described (29), this mutant still binds NADH (supplemental Fig. S4). Based on these data, the Munc119 binding region of RIBEYE appears to be topographically close to the NADH binding cleft of RIBEYE(B).
Munc119 Is Specifically Recruited to Purified Synaptic Ribbons—The co-immunoprecipitation experiments from bovine retina suggested the presence of Munc119 on synaptic ribbons (Fig. 5). Immunolabeling data clearly showed the presence of Munc119 in the presynaptic terminals at ribbon sites and also at sites close to the synaptic ribbon (Fig. 7B). Interestingly, purified synaptic ribbons isolated from bovine retina specifically recruited externally added soluble Munc119-GST fusion protein to synaptic ribbons (Fig. 7A). The binding of Munc119 to synaptic ribbons was specific because the control protein GST alone did not bind to synaptic ribbons. Because Munc119 is virtually absent from purified synaptic ribbons (Fig. 7A, lane 3), Munc119 appears to be a synaptic ribbon-associated component that can relatively easily dissociate from synaptic ribbons (see “Discussion”).
FIGURE 7.
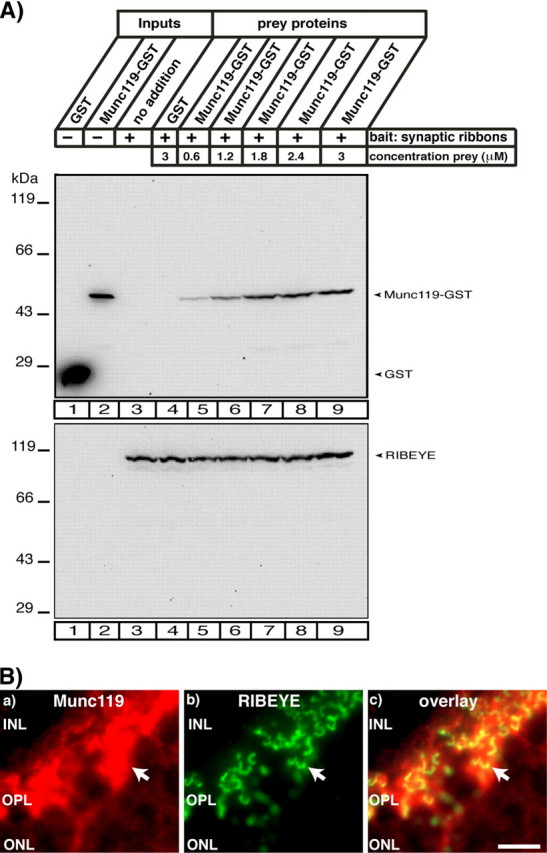
Purified synaptic ribbons specifically recruit Munc119. A, binding of Munc119 fusion protein to synaptic ribbons. 20 μg of purified synaptic ribbons were tested for their capability to bind soluble Munc119 fusion protein at the indicated concentrations. GST alone was used as control protein. Purified ribbons specifically bound Munc119-GST but not GST (A). For GST (lane 1) and Munc119-GST (lane 2) 10% were loaded as input; for the synaptic ribbon (lane 3) 100% were loaded as input. The two depicted blots show representative examples of four different experiments, which all showed the same result. The lower blot is stripped and reprobed with antibodies against RIBEYE to show equal loading of ribbons. A quantitative analysis of binding of Munc119 to synaptic ribbons is given in supplemental Fig. S5. B, Munc119 co-localizes with synaptic ribbons. Immunolabeling of the outer plexiform layer of the bovine retina that contain photoreceptor ribbon synapses with polyclonal antibodies against Munc119 and monoclonal antibodies against RIBEYE(B)/CtBP2. Strong immunosignals of Munc119 were found at synaptic ribbons and in close vicinity to synaptic ribbons. Abbreviations: ONL, outer nuclear layer; OPL, outer plexiform layer; INL, inner nuclear layer. Scale bar:10 μm.
The sequence of bovine Munc119 obtained in the present study by YTH screening with RIBEYE(B) as bait protein is identical to the bovine Munc119 sequence previously deposited at GenBank™ (Accession Number BC103449.1).
DISCUSSION
Munc119, a mammalian ortholog of the C. elegans protein unc119, is essential for synaptic transmission at the ribbon synapse and for vision (2). In the present study, we demonstrated that Munc119 interacts with the synaptic ribbon protein RIBEYE. The interaction between RIBEYE and Munc119 was consistently shown by five different independent methods, including YTH analyses, fusion protein pull-downs, interaction analyses in transfected COS cells, and immunoprecipitations from R28 retinal precursor cells and from bovine retina. The NADH binding subdomain of RIBEYE was shown to be responsible for the interaction with Munc119. Based on the analyses of RIBEYE(B) point mutants, the binding site of Munc119 appears to be close to the NADH binding site of RIBEYE; but the binding of Munc119 is independent upon NADH binding. In support of this view, RIBEYE(B)G730A that does not bind NADH still interacted with Munc119. Conversely, RIBEYE(B)E844Q that did not interact with Munc119 still bound NADH. Interestingly, the PrBP/δ-homology domain of Munc119 whose deletion caused the dramatic defects in vision and synaptic transmission in the respective patients and transgenic mouse model (5) was shown to be responsible for binding to RIBEYE. Therefore, the interaction with RIBEYE could mediate the essential physiological function of Munc119 in synaptic transmission at the photoreceptor ribbon synapse.
Although the physiological importance of Munc119 for synaptic transmission at the photoreceptor ribbon synapse has been well documented it is not yet clear how Munc119 works at the molecular level in the synapse. A key in the understanding of the function of Munc119 is probably its high homology to PrBP/δ. PrBP/δ binds and dissociates prenylated proteins from membranes (6, 30). This enzymatic activity is important for intracellular membrane and protein trafficking (6, 30). In photoreceptors, the trafficking role of PrBP/δ predominantly occurs in the inner and outer segments. We propose that Munc119 fulfills similar functions in the photoreceptor ribbon terminals. Such a trafficking role would be particularly important for the tonically active ribbon synapses, which are characterized by intensive membrane and protein trafficking. The idea that Munc119 supports similar processes as PrBP/δ but at different subcellular locations is further supported by the finding that PrBP/δ and Munc119 share common interaction partners (Arl2/3; 31, 32, 33). Additionally, Munc119 could activate Src-type signaling kinases in the photoreceptor synapse as recently observed for Munc119 in certain cells of the immune system (34, 35). A recent study demonstrated that Munc119 binds to CaBP4 (36) and thus, RIBEYE-Munc119 complexes might also be involved in the regulation of intracellular Ca2+ levels in the presynaptic ribbon terminal.
We have shown that purified ribbons could specifically recruit Munc119. Purified synaptic ribbons, which go through stringent washing steps (12, 27), contain little if any Munc119. In conclusion, Munc119 is most likely a peripherally associated component of synaptic ribbons that can relatively easy dissociate from them. In support of this suggestion, a large portion of Munc119 is soluble (Ref. 2 and data not shown). Large amounts of Munc119 are present in the presynaptic terminals in close vicinity to synaptic ribbons as judged by immunolabeling. Munc119 in presynaptic photoreceptor terminals could be recruited to synaptic ribbons. The factors that regulate the association/dissociation of Munc119 with synaptic ribbons in vivo remain to be elucidated by future analyses.
The synaptic defects observed in Munc119 transgenic mice and Munc119-deficient patients stress the role of Munc119 for vision and synaptic processing in the visual system. The fact that the PrBP/δ homology domain is crucial for Munc119 function particularly emphasizes the physiological importance of the RIBEYE-Munc119 interaction for synaptic transmission at the photoreceptor ribbon synapse.
Supplementary Material
Acknowledgments
We thank Conny Franke for excellent technical assistance.
This work was supported by the Sonderforschungsbereich 530 (SFB530) (TPC11) and by HOMFOR (to F. S.). The costs of publication of this article were defrayed in part by the payment of page charges. This article must therefore be hereby marked “advertisement” in accordance with 18 U.S.C. Section 1734 solely to indicate this fact. The nucleotide sequence(s) reported in this paper has been submitted to the GenBank™/EBI Data Bank with accession number(s) BC103449.1.
The on-line version of this article (available at http://www.jbc.org) contains supplemental materials and Figs. S1–S6.
Footnotes
The abbreviations used are: PrBP/δ, prenyl-binding protein δ; RE, RIBEYE; RE(B), RIBEYE(B) domain; NBD, NAD(H) binding domain; SBD, substrate binding domain; RE(AB), full-length RIBEYE(AB); PRD, proline-rich domain of Munc119; YTH, yeast-two-hybrid; GST, glutathione S-transferase; MBP, maltose-binding protein.
K. Schwarz and F. Schmitz, unpublished data.
References
- 1.Higashide, T., Murakami, A., McLaren, M. J., and Inana, G. (1996) J. Biol. Chem. 271 1797–1804 [DOI] [PubMed] [Google Scholar]
- 2.Higashide, T., McLaren, M. J., and Inana, G. (1998) Invest. Ophthalmol. Vis. Sci. 39 690–698 [PubMed] [Google Scholar]
- 3.Swanson, D. A., Chang, J. T., Campochiara, P. A., Zack, D. J., and Valle, D. (1998) Investig. Ophthalmol. Vis. Sci. 39 2085–2094 [PubMed] [Google Scholar]
- 4.Li, L., Florio, S. K., Pettenati, M. J., Rao, N., Beavo, J. A., and Baehr, W. (1998) Genomics 49 76–82 [DOI] [PubMed] [Google Scholar]
- 5.Kobayashi, A., Higashide, T., Hamasaki, D., Kubota, S., Sakuma, H., An, W., Fujimaki, T., McLaren, M. J., Weleber, R. G., and Inana, G. (2000) Investig. Ophthalmol. Vis. Sci. 11 3268–3277 [PubMed] [Google Scholar]
- 6.Zhang, H., Liu, X.-H., Zhang, K., Chen, C.-K., Frederick, J. M., Prestwich, G. D., and Baehr, W. (2004) J. Biol. Chem. 279 407–413 [DOI] [PubMed] [Google Scholar]
- 7.Kubota, S., Kobayashi, A., Mori, N., Higashide, T., McLaren, M. J., and Inana, G. (2002) Investig. Ophthalmol. Vis. Sci. 43 308–313 [PubMed] [Google Scholar]
- 8.Mori, N., Ishiba, Y., Kubota, S., Kobayashi, A., Higashide, T., McLaren, M. J., and Inana, G. (2006) Investig. Ophthalmol. Vis. Sci. 47 1281–1292 [DOI] [PubMed] [Google Scholar]
- 9.Sterling, P., and Matthews, G. (2005) Trends Neurosci. 28 20–29 [DOI] [PubMed] [Google Scholar]
- 10.Heidelberger, R., Thoreson, W. B., and Witkovsky, P. (2005) Prog. Ret. Eye Res. 24 682–720 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.tom Dieck, S., and Brandstätter, J. H. (2006) Cell Tiss. Res. 326 339–346 [DOI] [PubMed] [Google Scholar]
- 12.Schmitz, F., Königstorfer, A., and Südhof, T. C. (2000) Neuron 28 857–872 [DOI] [PubMed] [Google Scholar]
- 13.Zenisek, D., Horst, N. K., Merrifield, C., Sterling, P., and Matthews, G. (2004) J. Neurosci. 24 9752–9759 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Wan, L., Almers, W., and Chen, W. (2005) J. Neurosci. 25 941–949 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Chinnadurai, G. (2002) Mol. Cell 9 213–224 [DOI] [PubMed] [Google Scholar]
- 16.Kumar, V., Carlson, J. E., Ohgi, K. A., Edwards, T. A., Rose, D. W., Escalante, C. R., Rosenfeld, M. G., and Aggarwal, A. K. (2002) Mol. Cell 10 857–869 [DOI] [PubMed] [Google Scholar]
- 17.Nardini, M., Spano, S., Cericola, C., Pesce, A., Massaro, A., Millo, E., Luini, A., Corda, D., and Bolognesi, M. (2003) EMBO J. 22 3122–3130 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Tai, A. W., Chuang, J. Z., Wolfrum, U., and Sung C. H. (1999) Cell 97 877–887 [DOI] [PubMed] [Google Scholar]
- 19.James, P., Halladay, J., and Craig, E. A. (1996) Genetics 144 1425–1436 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Harper, J. W., Adami, G. R., Wei, N., Keyomarsi, K., and Elledge, S. J. (1993) Cell 75 805–816 [DOI] [PubMed] [Google Scholar]
- 21.Helmuth, M., Altrock, W., Böckers, T. M., Gundelfinger, E. D., and Kreutz, M. R. (2001) Anal. Biochem. 293 149–152 [DOI] [PubMed] [Google Scholar]
- 22.Bai, C., and Elledge, S. J. (1996) Methods Enzymol. 273 331–347 [DOI] [PubMed] [Google Scholar]
- 23.Wang, Y., Okamoto, M., Schmitz, F., Hofmann, K., and Südhof, T. C. (1997) Nature 388 593–598 [DOI] [PubMed] [Google Scholar]
- 24.Stahl, B., Diehlmann, A., and Südhof, T. C. (1999) J. Biol. Chem. 14 9141–9148 [DOI] [PubMed] [Google Scholar]
- 25.Seigel, G. M. (1996) Dev. Biol. Anim. 32 66–68 [DOI] [PubMed] [Google Scholar]
- 26.Seigel, G. M., Sun, W., Wang, J., Hershberger, D. H., Campbell, L. M., and Salvi, R. J. (2004) Curr. Eye Res. 28 257–269 [DOI] [PubMed] [Google Scholar]
- 27.Schmitz, F., Bechmann, M., and Drenckhahn, D. (1996) J. Neurosci. 16 7109–7116 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Schmitz, F., Tabares, L., Khimich, D., Strenzke, N., DeLa Villa-Polo, P., Castellano-Munoz, M., Bulankina, A., Moser, T., Fernandez-Chacon, R., and Südhof, T. C. (2006) Proc. Natl. Acad. Sci. U. S. A. 103 2925–2931 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Fjeld, C. C., Birdsong, W. T., and Goodman, R. H. (2003) Proc. Natl. Acad. Sci. U. S. A. 100 9202–9207 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Zhang, H., Li, A., Doan, T., Rieke, F., Detwiler, P. B., Frederick, J. M., and Baehr, W. (2007) Proc. Natl. Acad. Sci. U. S. A. 104 8857–8862 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.van Valkenburgh, H., Shern, J. F., Sharer, J. D., Zhu, X., and Kahn, R. A. (2001) J. Biol. Chem. 276 22826–22837 [DOI] [PubMed] [Google Scholar]
- 32.Hanzal-Bayer, M., Renault, R., Roversi, P., Wittinghofer, A., and Hillig, R. C. (2002) EMBO J. 21 2095–2106 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Kobayashi, A., Kubota, S., Mori, N., McLaren, M. J., and Inana, G. (2003) FEBS Lett. 534 26–32 [DOI] [PubMed] [Google Scholar]
- 34.Cen, O., Gorska, M. M., Stafford, S. J., Sur, S., and Alam, R. (2003) J. Biol. Chem. 278 8837–8845 [DOI] [PubMed] [Google Scholar]
- 35.Gorska, M. M., Stafford, S. J., Cen, O., Sur, S., and Alam, R. (2004) J. Exp. Med. 199 369–379 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Haeseleer, F. (2008) Investig. Ophthalmol. Vis. Sci. 49 2366–2375 [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.