Abstract
The σE pathway of extracytoplasmic stress responses in Escherichia coli is activated through sequential cleavages of the anti-σE protein, RseA, by membrane proteases DegS and RseP. Without the first cleavage by DegS, RseP is unable to cleave full-length RseA. We previously showed that a PDZ-like domain in the RseP periplasmic region is essential for this negative regulation of RseP. We now isolated additional deregulated RseP mutants. Many of the mutations affected a periplasmic region that is N-terminal to the previously defined PDZ domain. We expressed these regions and determined their crystal structures. Consistent with a recent prediction, our results indicate that RseP has tandem, circularly permutated PDZ domains (PDZ-N and PDZ-C). Strikingly, almost all the strong mutations have been mapped around the ligand binding cleft region in PDZ-N. These results together with those of an in vitro reaction reproducing the two-step RseA cleavage suggest that the proteolytic function of RseP is controlled by ligand binding to PDZ-N.
In bacteria, abnormalities in cell envelope proteins are generally referred to as “extracytoplasmic stresses.” Such stress signals are transmitted across the cytoplasmic membrane to the cytoplasm by specific signal transduction systems, leading to the induced synthesis of a set of cell surface proteins that function to cope with the stresses. These responses are called extracytoplasmic stress response (1, 2).
The σE pathway of extracytoplasmic stress response involves a proteolytic cascade that activates an alternative sigma factor (σE) responsible for transcription of stress-induced genes (1-3). σE is normally kept inactive by its tight binding to the cytoplasmic domain of the anti-σE protein, RseA, which has a type II (cytoplasmic N terminus, periplasmic C terminus) membrane topology (4-6). Under stressed conditions, DegS, a cytoplasmic membrane-anchored protease, is activated to cleave the periplasmic side of RseA (7). In this process, a peptide, conserved among many outer membrane proteins in their C termini, binds to the PDZ domain of DegS and induces a conformational change to activate its proteolytic function (8-10). After the first cleavage by DegS, RseP, another membrane protease, cleaves RseA within its transmembrane segment and releases the complex of σE and the RseA cytoplasmic domain from the membrane (11-13). The RseA cytoplasmic domain is finally degraded by cytoplasmic ATP-dependent proteases, leading to the liberation of active σE (14).
RseP is a family member of the S2P intramembrane proteases (15). Our biochemical studies (16, 17) suggested that the active site of RseP is sequestered within a proteinaceous domain that is partially embedded in the membrane, in agreement with the recently elucidated crystal structure of an S2P homolog from an archaeal species, Methanocaldococcus jannaschii (16-18). For RseP to cleave RseA, RseA must receive the prior cleavage by DegS (11, 12). Thus, there must be some mechanism by which proteolytic action of RseP against full-length RseA is suppressed. This negative regulation, which prevents unwanted expression of stress proteins, is an integral and essential part of the σE pathway regulation. Several lines of evidence suggest that multiple elements are involved in this apparently negative regulation; they include the “PDZ-like” domain of RseP (19, 20), the Gln-rich sequences in the RseA periplasmic region (19), and the periplasmic RseB protein and DegS (21). The RseB function in the negative regulation is dependent on the RseP PDZ domain, whereas DegS acts independently of it (21). In this study we further explored the regulatory roles of the PDZ-like elements in the periplasmic region of RseP.
PDZ (named after three eukaryotic proteins, in which these domains were first identified: PSD-95, DLG, and ZO-1) is one of the most widespread modular protein domains involved in protein-protein interaction and is shared by many proteins in organisms from prokaryotes to eukaryotes (22, 23). PDZ domains consist of 80-100 amino acid residues and participate in a variety of signaling processes as well as degradation, transport, and localization of proteins by recognizing specific ligands, generally their C-terminal 3-5 residues (22-24). In Escherichia coli, several cell surface proteases have this protein module presumably for specific substrate recognition and regulation of the proteolytic functions. Site-directed alterations of conserved residues in the RseP PDZ-like domain as well as deletion of this domain impaired the negative regulation, enabling RseP to cleave full-length RseP in the absence of DegS (19, 20). However, it remains unclear how the PDZ-like domain works in the modulation of RseP proteolytic function.
Here we addressed the mechanism of PDZ-mediated regulation of RseP by combined genetic, biochemical, and structural approaches. Our results suggest that RseP carries, instead of the previously assigned PDZ-like domain (PDZ*), two circularly permutated PDZ domains (PDZ-N and PDZ-C) with differential functions in the regulation with the N-terminal domain (PDZ-N) playing a major role.
EXPERIMENTAL PROCEDURES
E. coli Strains and Plasmids—E. coli strains and plasmids used in this study are listed in Table 1. Construction of the individual strains and plasmids are described in the supplemental “Experimental Procedures.”
TABLE 1.
Strains and plasmids used in this study
| Strains | Relevant genotype | Reference |
|---|---|---|
| AD16 | Δ(pro-lac), thi/F′ lacIq | 11 |
| AD1840 | AD16, ΔrseA::cat, ΔrseP::kan, ΔdegS::tet | 11 |
| AD1867 | AD16, ΔdegS::cat | This study |
| AD2239 | AD16, ΔdegS::cat, degP41::kan | This study |
| AD2249 | AD16, ΔdegS::cat, degP41::kan | This study |
| CU141 | MC4100/F′ lacIq | 48 |
| KK372 | CU141, ΔrseA::cat, ΔdegS::tet | This study |
| KK374 | CU141, ΔrseA::cat, ΔrseP::kan | 13 |
| MC4100 | araD139 Δ(argF-lac)U169 rpsL150 relA1 fibB5301 deoC1 ptsF25 rbsR | 49 |
| SN140 | CU141, ompT::kan | 48 |
|
TR71 |
MC4100, γRS45[rpoHP3-lacZ] |
A gift of T. J. Silhavy. |
|
Plasmids |
Encoded protein |
Reference |
| pTWV228 | Vector | TakaraBio |
| pKK11 | RseP-HM | 19 |
| pKK49 | RseP-HM | 13 |
| pKK131 | RseP(ΔPDZ*)-HM | 19 |
| pKK135 | RseP(ΔPDZ*/D402N)-HM | 19 |
| pSTD672 | His6-DegS | This study |
| pSTD821 | His6-MBP-RseA140 | 13 |
| pSTD929 | His6-MBP-RseA+ | This study |
| pSTD997 | RseP(A115V/G214E)-HM | This study |
| pSTD999 | RseP(L151P/H22F)-HM | This study |
| pSTD1001 | RseP(A115V/L151P)-HM | This study |
| pSTD1002 | RseP(A115V/G214E)-HM | This study |
| pSTD1008 | RseP(A115V/ΔPDZ*)-HM | This study |
| pSTD1017 | HA-MBP-RseA(LacYTM1) 140 | 16 |
| pSTD1131 | RseP(V302P)-HM | This study |
| pSTD1132 | RseP(G303E)-HM | This study |
| pSTD1135 | RseP(L151P/V302P)-HM | This study |
| pSTD1136 | RseP(L151P/G303E)-HM | This study |
| pSTD1144 | RseP(ΔPDZ-C)-HM | This study |
| pSTD1163 | RseP(ΔPDZ-N)-HM | This study |
| pSTD1175 | GST-PDZ-N | This study |
| pSTD1183 | RseP(ΔPDZ-N/G303E)-HM | This study |
| pSTD1212 | GST-PDZ-C | This study |
| pSTD1213 | GST-PDZ-N/PDZ-C | This study |
| pSTD1241 | GST-PDZ-N(L169M) | This study |
| pSTD1330 | RseP(ΔPDZ-N/V302P/G303E)-HM | This study |
Isolation of Deregulated rseP Mutants—Plasmid pKK11 was propagated for several generations at 37 °C in a mutator strain XL1-Red (Stratagene) and then transformed into TR71, with selection at 30 °C on L agar plates containing 50 μg/ml ampicillin, 40 μg/ml X-gal4, and 0.5 mm phenylethyl-β-d-thiogalactoside. Blue colonies were examined whether their plasmids yielded blue transformants again upon re-introduction into TR71. Plasmid clones (named pYGF followed by specific number) that consistently gave blue transformants on X-gal plates were subjected to DNA sequencing analysis.
Crystallographic and SAXS Analyses of the PDZ-N and the PDZ-C Domains of RseP—The PDZ-N and PDZ-C domains of RseP were expressed, purified, and crystallized as described under the supplemental “Experimental Procedures.” X-ray diffraction data sets for SeMet PDZ-N(L169M) were collected on beamline BL5A at Photon Factory, KEK (Tsukuba, Japan) with a Quantum 315 CCD detector at cryogenic temperature (100 K). Those for SeMet PDZ-C were collected on beamline BL44XU at SPring-8 (Hyogo, Japan) with an image plate/CCD hybrid detector DIP6040 (Mac Science/Bruker AXS) at 100 K. All the data were indexed and scaled with HKL2000 (25) and SCALA (26). The PDZ-N and the PDZ-C crystals contained three PDZ-N molecules and one PDZ-C molecule, respectively, in an asymmetric unit. Crystallographic data are summarized in Table 2. Phase determination was carried out from single-wavelength anomalous dispersion (SAD) data collected at the selenium-peak wavelength using the programs SHELXC/D/E (27, 28) and HKL2MAP (27, 28). The crystal of SeMet PDZ-N was found to be highly twinned (twin fraction, ∼0.33), and the diffraction data were processed by the methods of Rudolph et al. (29). Model building was carried out using the programs RESOLVE (30, 31) and ARP/WARP (30, 31). The electron density maps revealed a typical PDZ-fold for either PDZ domain (note that their primary-tertiary structure relationship exhibits a circular permutation when compared with a typical PDZ domain). The structures of PDZ-N and PDZ-C were refined against the 18.55-1.70 Å and 20.75-0.98 Å intensity data, respectively, using Refmac5 (26). The structures of PDZ-N and PDZ-C have been deposited in the Protein Data Bank under PDB code 2ZPL and 2ZPM, respectively.
TABLE 2.
Data collection and structure determination
| SeMet PDZ-N (Leu169Met)a | SeMet PDZ-Ca | SeMet PDZ-Ca | |
|---|---|---|---|
| Data collection | |||
| Beamline | BL5A at KEK | BL44XU at SPring-8 | BL44XU at SPring-8 |
| Space group | P31 | C2 | C2 |
| Cell dimensions (Å) | a = 54.8, b = 54.8, c = 75.6 α = β = 90.0°, γ = 120.0° | a = 70.1, b = 26.9, c = 42.8 α = γ = 90.0°, β = 112.1° | a = 70.2, b = 26.9, c = 42.9 α = γ = 90.0°, β = 112.1° |
| Wavelength (Å) | 0.9791 | 0.6500 | 0.9788 |
| Resolution range (Å) | 18.55-1.70 (1.79-1.70) | 21.47-0.98 (1.03-0.98) | 31.70-1.28 (1.35-1.28) |
| No. of total observations | 298,168 (29,834) | 156,823 (22,490) | 69,688 (8,936) |
| No. of unique reflections | 27,607 (3,842) | 42,333 (6,128) | 19,298 (2,706) |
| Completeness (%) | 99.0 (93.9) | 99.4 (99.0) | 99.2 (95.7) |
| I/σ(I) | 35.1 (4.2) | 17.1 (2.9) | 18.8 (6.9) |
| Multiplicity | 10.8 (7.8) | 3.7 (3.7) | 3.6 (3.3) |
| Rmergeb | 0.037 (0.376) | 0.044 (0.446) | 0.043 (0.147) |
| Rmeasc | 0.041 (0.429) | 0.053 (0.525) | 0.051 (0.175) |
| Wilson B-factor (Å2) | 26.2 | 7.3 | 10.7 |
|
Estimated twin fraction |
0.34 |
||
| Refinement | |||
| Protein residues refined | |||
| A-chain | 127-218 | 224-309 | |
| B-chain | 127-218 | ||
| C-chain | 127-184, 190-218 | ||
| No. of other molecules | |||
| Ni2+ | 3 | ||
| Glycerol | 2 | ||
| Water | 122 | 151 | |
| Rworkd | 0.150 | 0.150 | |
| Rfreee | 0.176 | 0.180 | |
| Refined twin fraction | |||
| (h, k, l) | 0.642 | ||
| (−h−k, k, −l) | 0.319 | ||
| (−h, h+k, −l) | 0.019 | ||
| (−h, −k, l) | 0.020 | ||
| Root mean square deviation | |||
| Bonds (Å) | 0.019 | 0.023 | |
| Angles (°) | 1.869 | 2.141 | |
| Average B-factor (Å2) | |||
| A-chain | 26.2 | 11.2 | |
| B-chain | 26.8 | ||
| C-chain | 29.6 | ||
| Ni2+ | 31.0 | ||
| Glycerol | 39.9 | ||
| Solvent | 34.0 | 22.8 | |
| Ramachandran plot (%) | |||
| Most favored | 93.9 | 89.7 | |
| Additional allowed | 6.1 | 10.3 | |
| Generously allowed | 0 | 0 | |
| Disallowed | 0 | 0 |
The number in parentheses represent statistics in the highest resolution shell.
Rmerge = ∑∑j|<I(h)> - [I(h)j|/∑∑j|<I(h)>|, where <I(h)> is the mean intensity of symmetry-equivalent reflections.
Rmeas = ∑√(n/n - 1)∑j |I(h) - I(h)j|∑∑j |<I(h)|, the multiplicity weighted Rmerge.
Rwork = ∑(||Fp(obs) - Fp(calc)||)/∑|Fp(obs)|.
Rfree = R factor for a selected subset (5%) of the reflections that was not included in prior refinement calculations.
SAXS patterns of a tandem PDZ-N-PDZ-C construct (PDZ-NC) were recorded at RIKEN Structural Beamline I (BL45XU, SPring-8) (32) and processed as described (33, 34) to obtain scattering curves, I(S), where S = 2sinθ/l, 2θ is the scattering angle, and λ is the wavelength of the x-ray (0.9 Å). Low and high resolution models were evaluated with GASBOR (35) and BUNCH (36) packages, respectively. Further details are described under supplemental “Experimental Procedures.”
Two-step Proteolysis of the Model Substrates by His6-DegS and RseP-HM in Purified in the Vitro System—For in vitro reconstitution of two-step cleavages of His6-MBP-RseA, His6-MBP-RseA (17 μg/ml) was incubated with the indicated combinations of His6-DegS (33 μg/ml), RseP-HM (11 μg/ml), and the OMP peptide (NH2-DNRDGNVYYF-COOH) (8) (50 μm) at 37 °C in buffer containing 50 mm Tris-HCl (pH 8.1), 1100 mm KCl, 0.02% n-dodecyl-β-d-maltoside, 2.5% glycerol, 5 μm zinc acetate, 10 mm 2-mercaptoethanol, 1 mm phenylmethylsulfonyl fluoride, and 1 mm PefaBlock SC. For assays of RseP cleavage of His6-MBP-RseA140 and His6-MBP-RseA, 10 mm 2-mercaptoethanol and 100 μg/ml E. coli phospholipids (Avanti) were also included. For all the assays, samples were withdrawn at intervals, mixed with an equal volume of 2× SDS sample buffer, and analyzed by SDS-PAGE and CBB staining.
RESULTS
Isolation of Deregulated RseP Variants Capable of Cleaving the Full-length RseA Protein—Previous site-directed substitution and deletion analyses of the PDZ-like domain of RseP (PDZ* in Fig. 1A) showed that the integrity of this domain is required for the suppression of RseP action against full-length RseA (19). To identify RseP regions that are involved in the negative regulation more thoroughly, we mutagenized plasmid pKK11 carrying the resP-his6-myc gene as described under “Experimental Procedures.” Strain TR71 having rpoHP3-lacZ, which reports the σE activity as that of β-galactosidase, forms a blue colony on X-gal indicator plates when the intracellular σE activity is elevated. TR71 carrying the original plasmid pKK11 formed white colonies on X-gal plates (data not shown), indicating that overexpression of wild type RseP-HM did not itself activate σE. In contrast, the mutagenized pool of pKK11 yielded blue transformant colonies at appreciable frequencies. In the screening, we found 50 dark blue and pale blue colonies out of about 6.6 × 104 transformants. After elimination of plasmid-un-linked mutants by plasmid re-isolation and re-transformation, we obtained 23 plasmid clones which were then characterized by DNA sequencing. We, thus, identified 19 different rseP mutant clones (Figs. 1A and 2A). Most of them contained a single amino acid substitution in the central periplasmic region, whereas one (pYGF7) contained a TM3 alteration, and another (pYGF4) contained double mutations affecting TM2 and the periplasmic domain.
FIGURE 1.
Sequence features, structures and mutations of the PDZ domains of RseP. A, schematic representation of the RseP primary sequence and positions of the mutations isolated in this study. Regions corresponding to the previously assigned PDZ-like domain (PDZ*, green arrow with dotted line) and the circularly permutated PDZ-N (red-boxed arrow) and PDZ-C (blue-boxed arrow) domains are shown at the top. Positions of the mutational alterations are shown by circles with the pYGF plasmid numbers; strong mutations (more than 2-fold induction of the reporter gene) are indicated by a red color. Regions carried by the ΔN1 and ΔN2 derivatives of RseP-HM are shown at the bottom. a.a. res. no., amino acid residue number. B-D, crystal structures of the PDZ-N and PDZ-C domains. The main chain structures of the PDZ-N (B) and PDZ-C (C) from RseP and that of the third PDZ domain of PSD95 (D) are shown by ribbon diagram representations (cyan, helices; magenta, strands; salmon, loops). The α-helix (αB) and β-strand (βB) elements thought to be important for ligand binding are marked. The carboxylate binding loop is highlighted in yellow. Side chains of amino acids showing strong mutational effects are shown by sticks in B. Note that Leu-169 was replaced with SeMet for structure determination of PDZ-N. A polypeptide ligand bound to the third PDZ-domain of PSD95 (40) is shown by a light-green stick in D. E, a typical rigid-body refinement model of PDZ-NC obtained by SAXS, which is superimposed onto the low resolution molecular shape of PDZ-NC. The envelope smoothed with SITUS package (46) was visualized on VMD (47).
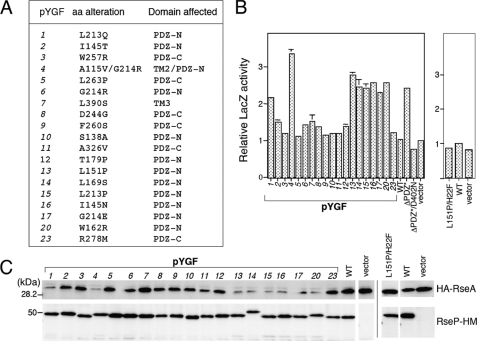
FIGURE 2.
DegS-independent proteolysis of RseA by the PDZ mutant forms of RseP-HM in vivo. A, amino acid alterations and the domains affected in the RseP-HM mutants isolated by the genetic screening. B, reporter LacZ activities induced by expression of the RseP PDZ mutants. Cells of TR71 (rpoHP3-lacZ) carrying the indicated pYGF plasmid, pKK11 (RseP-HM; WT), pKK131 (RseP(ΔPDZ*)-HM; ΔPDZ*), pKK135 (RseP(ΔPDZ*/D402N)-HM; ΔPDZ*/D402N), pSTD999 (RseP(L151P/H22F)-HM; L151P/H22F), or pTWV228 (vector) were grown at 37 °C in L medium (containing 10 g bacto-tryptone, 5 g yeast extract, and 5 g NaCl per liter; pH was adjusted to 7.2 by NaOH) and assayed for β-galactosidase activity. C, cellular levels of HA-RseA upon co-expression of the RseP PDZ mutants. pYGF plasmids, pKK11, pSTD999, and vector were introduced into AD1840/pSD691 (HA-RseA). Cells were grown at 30 °C in L medium containing 1 mm isopropyl 1-thio-β-d-galactopyranoside and 1 mm cAMP for 2 h. Portions (containing ∼6 × 107 cells) of the cultures were withdrawn and mixed with an equal volume of 10% trichloroacetic acid for subsequent SDS-PAGE and anti-HA and anti-Myc immunoblotting analyses.
Expression of these mutant RsePs caused a 1.2-3.3-fold increase in the reporter LacZ activity as compared with wild type RseP (Fig. 2B). To examine whether the mutant RsePs have an ability to cleave full-length RseA in the absence of DegS, they were co-expressed with HA-tagged RseA (HA-RseA) in a strain deleted for rseA, rseP, and degS (Fig. 2C). Immunoblotting results indicated that all the mutant forms of RseP-HM accumulated as stably as wild type RseP-HM in the cell, suggesting that the mutations did not drastically affect the overall conformation of RseP. Although cellular abundance of HA-RseA was only marginally affected upon co-expression of wild type RseP-HM (Fig. 2C, WT), it was lowered significantly upon co-expression of most of the mutant forms of RseP-HM (Fig. 2C, pYGF). Accumulation levels of HA-RseA exhibited inverse correlation with the reporter activities conferred by the RseP mutants (Figs. 2, B and C). The proteolytic function of RseP was responsible for the concurrent occurrence of the decrease in the RseA levels and of the increase in the σE activities, which was demonstrated for RseP(L151P)-HM by introducing a protease active site mutation, H22F, into it; then the double mutant, RseP(L151P/H22F)-HM, neither induced the reporter expression nor decreased the HA-RseA accumulation (Figs. 2, B and C). These results strongly suggest that the mutant RseP enzymes catalyze DegS-independent proteolysis of intact RseA.
Although the mutations could simply enhance the proteolytic activity that is intrinsic to RseP, a more interesting possibility would be that they affect the specific regulatory mechanism that prevents proteolysis of intact RseA. To address this point, the mutant proteins were expressed in the rseA, rseP, degS triple mutant cells together with a model protein HA-MBP-RseA(LacYTM1)140 (supplemental Fig. S1), which mostly lacks the periplasmic domain and, therefore, receives DegS-independent cleavage by RseP, yielding a stable cytoplasmic MBP domain as a cleavage product (13). Although about 75% of this model substrate was cleaved in cells expressing wild type RseP-HM, the cleavage efficiencies never exceeded this level in cells expressing the mutant forms of RseP. In fact, many of them were less active than wild type RseP-HM in cleaving this model substrate (the results for representative mutants are shown in supplemental Fig. S1). Similarly, other RseP mutants that we engineered and used below did not show any increased proteolytic activity against this substrate (supplemental Fig. S1 and data not shown). Thus, it is unlikely that the DegS-independent proteolysis of intact RseA by the mutant RseP enzymes was due to more efficient chemistry of catalysis.
RseP Has Two Tandemly Positioned PDZ Domains with Different Roles—Our sequencing results revealed unexpectedly that many strong mutations affect amino acid residues that are outside the previously assigned PDZ-like domain (PDZ*) (19) (Fig. 1A). They cluster in a segment N-terminal to it. These results prompted us to re-examine the amino acid sequence features of RseP. The Pfam program indicated that the region N-terminal to PDZ* has sequence characteristics that exhibit low but significant similarity to those of the PDZ domains. In agreement with this notion, Kinch et al. (15) proposed from bioinformatic analyses of the S2P family protein sequences that the central periplasmic region of RseP contains two tandemly positioned PDZ-like domains with a circularly permutated sequence arrangement. As described below, our structural analyses revealed that each of these two PDZ-like domains indeed assumes a PDZ-fold. Thus, we here-after call them PDZ-N and PDZ-C, respectively. Whereas PDZ-C has high sequence similarity to other PDZ domains (Pfam score, 2.3E-15), PDZ-N is much more limitedly related to PDZ (Pfam score, 0.21). All the strong mutations isolated in this study were found to affect residues within the PDZ-N domain (residues Ile-145, Leu-151, Trp-162, Leu-169, Leu-213, and Gly-214, shown in Fig. 1B). On the other hand, our screening yielded only weak mutations in PDZ-C.
Structural Studies of PDZ-N and PDZ-C—We determined the structures of the PDZ-N and PDZ-C regions of RseP by x-ray crystallography. We expressed these domains separately as glutathione S-transferase fusion proteins in the presence of SeMet and purified them. After proteolytic removal of the glutathione S-transferase moiety, the purified PDZ-N and PDZ-C domains were crystallized, and x-ray diffraction data were collected. We also obtained native crystals for both of the PDZ-domains, which exhibited resolutions that were comparable with SeMet-labeled crystals (data not shown). Crystal structures of PDZ-N (Fig. 1B) and PDZ-C (Fig. 1C) were solved at 1.70 and 0.98 Å resolutions, respectively, through single-wavelength anomalous dispersion experiments. Crystallographic parameters are compiled in Table 2. The overall structures of the PDZ-N and the PDZ-C domains and the configurations of their secondary structure elements are very similar to the third PDZ domain of PSD95 having a typical PDZ-fold (Fig. 1D). For the aligned residues, the root mean square deviation values between the Cα atoms of PDZ-N and PSD95 and between those of PDZ-C and PSD95 are 2.1 and 2.3 Å, respectively. It should be noted, however, that PDZ-N and PDZ-C have a circularly permutated connectivity in the primary sequence in that their C-terminal regions having two conserved β strands (βA and βB) correspond to the very N-terminal region in a typical PDZ structure, confirming the prediction by Kinch et al. (15). Thus, the PDZ domains of RseP represent a new type of circularly permutated PDZ-fold.
PDZ domains generally have a ligand binding groove that is formed by an α-helix (αB) and a β-strand (βB) as well as a carboxylate binding loop with a conserved motif (GLGF in many PDZ domains, whereas SLGI and FVGI in PDZ-N and PDZ-C, respectively) that is located at an end of the groove (Figs. 1, B and C). Our structure revealed a putative ligand binding groove formed by corresponding structural elements in each of PDZ-N and PDZ-C, although the groove of PDZ-N is much narrower and shallower than that of PDZ-C due to a small α-helix covering this groove (see below and supplemental Fig. S2A). Notably, sites of many mutations with a strong effect proved to be in the ligand binding groove region of PDZ-N; they were mapped either in the αB helix, in the carboxylate binding loop, or at the bottom of the groove (Fig. 1B). Mutations had been isolated repeatedly at the αB1 position (corresponding to Trp-162) and in the conserved carboxylate-binding loop region, which are known in other PDZs to be critical for ligand binding (23). It is conceivable that these mutations impair binding of a ligand, although its identity is unknown. It follows then that ligand binding is required for the down-regulation of the proteolytic action of RseP against full-length RseA. Our structure also showed that PDZ-N has a small helix (α′ in Fig. 1B) that partially covers the ligand binding groove. Such a structural feature could affect the putative ligand binding. On the other hand, the crystal structure of PDZ-C revealed that the C terminus of one PDZ-C molecule interacted with the ligand binding groove of the neighboring molecule in a way similar to the interaction observed between the PDZ domain of PSD95 and the C terminus of its ligand peptide (supplemental Fig. S2). It is suggested that PDZ-C also has a potential ability to bind a ligand.
To study the nature of interdomain interactions between PDZ-N and PDZ-C in solution, we carried out SAXS analysis of the PDZ-N and PDZ-C domains fused into a single polypeptide (PDZ-NC) as in the intact RseP protein (Fig. 1E, see also supplemental “Results” and Figs. S3 and S4). The results suggest that the interaction between the two tandemly positioned PDZ domains is mediated by relatively loose contacts, forming a bent-dumbbell-shaped PDZ-NC with each lobe containing a small cavity as revealed by the crystal structures of PDZ-N and PDZ-C.
Site-directed Mutational Analyses of the PDZ-N and PDZ-C Domains—The alterations of Leu-213 to Pro and Gly-214 to Glu in the conserved carboxylate binding motif of PDZ-N resulted in strong deregulation. We addressed whether mutations of the corresponding residues, Val-302 and Gly-303, in PDZ-C gave any phenotype. We changed them to Pro and Glu to mimic the regulation-impairing alterations of PDZ-N at Leu-213 and Gly-214, respectively. Expression of the RseP variant with the Val302Pro or G303E mutation alone did not increase the σE activity (Fig. 3A) nor promoted HA-RseA degradation (Fig. 3B). However, both of these mutations were found to enhance synergistically the effects of the L151P alteration in PDZ-N when combined with it (Figs. 3, A and B). These results suggest that PDZ-C has an auxiliary role that is related to the function of PDZ-N.
FIGURE 3.
Proteolytic cleavage of RseA by RseP mutants. A and B, effects of mutations in the carboxylate binding loop region of PDZ-C. C and D, effects of PDZ-N and PDZ-C deletions. E and F, enhancement of the effects of PDZ-N mutations by the A115V alteration in TM2. A, C, and E, reporter LacZ activities. Derivatives of pKK11 encoding the indicated mutant forms of RseP-HM were introduced into TR71. Cells were grown at 37 °C in L medium and assayed for β-galactosidase activity. WT, wild type. B, D, and F, cellular levels of HA-RseA. Derivatives of pKK11 encoding the indicated mutant forms of RseP-HM were introduced into AD1840/pSTD691 (HA-RseA). Cells were grown at 30 °C in L medium containing 1 mm isopropyl 1-thio-β-d-galactopyranoside and 1 mm cAMP for 2 h. Proteins were analyzed by SDS-PAGE and anti-HA and anti-Myc immunoblotting. Average values from at least two independent experiments are shown with S.D.
We then constructed RseP variants deleted for either PDZ-N or PDZ-C. A variant deleted for both of PDZ-N and PDZ-C was unstable (data not shown) and not analyzed. The PDZ-C-less variant exerted very weak effects on the σE activity and HA-RseA degradation (Figs. 3, C and D, ΔPDZ-C). Because all the strong mutations were mapped in the PDZ-N domain, we expected that deletion of PDZ-N would result in severe deregulation. Surprisingly, however, the PDZ-N-less variant only weakly promoted the σE activity and HA-RseA degradation (Figs. 3, C and D, ΔPDZ-N), although it retained significant intrinsic protease activity against HA-MBP-RseA(La-cYTM1)140 (supplemental Fig. S1). Thus, contrary to our expectation, the RseP activity against full-length RseA remains down-regulated in the absence of either of the PDZ domains. However, we found that derivatives of RseP(ΔPDZ-N)-HM with additional mutations (G303E and G303E/V302P) in the carboxylate binding loop of PDZ-C were deregulated significantly, as their expression was accompanied by increased σE activity and HA-RseA degradation. These results suggest that the PDZ-C can substitute for PDZ-N in the absence of the latter in repressing the DegS-independent cleavage of full-length RseA, presumably in a ligand binding-dependent manner (see below and “Discussion”).
TM2 Modulates the PDZ Function—Among the mutant rseP genes isolated and characterized in this work, the rseP(A115V/G214R) allele exhibited the strongest activation of σE, resulting in more than 3-fold increase in the reporter output (Figs. 2B and 3E). Whereas RseP(A115V/G214R) contains amino acid alterations in TM2 (A115V) and in PDZ-N (G214R), the RseP(G214R) single mutant (pYGF6) only weakly elevated the σE activity (Figs. 2B and 3E) and only slightly lowered the accumulation of co-expressed HA-RseA (Fig. 2C). We also constructed the A115V RseP single mutant protein, which accumulated stably in vivo (Fig. 3F). Its in vivo proteolytic activity was indistinguishable from those of wild type RseP in that it cleaved HA-MBP-RseA(LacYTM1)140 efficiently (supplemental Fig. S1) but not the full-length RseA protein (Fig. 3F) and in that it did not activate σE (Fig. 3E). Thus, the A115V and G214R mutations exerted a synergistic effect when they were combined. The A115V substitution also enhanced the mutational effects of other PDZ-N mutations, L151P and G214E, as well as the deletion of PDZ* (Figs. 3E and F). These results suggest that TM2 and PDZ-N interact functionally.
The PDZ Mutations Alter the Conformation of the RseP Periplasmic Domain—The mutations in the PDZ domains could affect the conformation of the RseP protein. To address this possibility, we examined trypsin sensitivity of the wild type and PDZ mutants of RseP-HM (Fig. 4). Cells expressing RseP-HM and its variants were converted to spheroplasts and treated with 5 μg/ml trypsin at 0 °C. Immunoblotting using anti-Myc showed that wild type RseP-HM was significantly resistant to trypsin, which did not produce any cleavage product upon incubation for up to 12 min (Fig. 4A, RseP+-HM). On the other hand, mutant proteins RseP(A115V/G214E)-HM and RseP(L151P)-HM proved to be more sensitive to trypsin, generating a major cleavage product of ∼33 kDa and several minor smaller fragments after a short (2-min) incubation (Fig. 4A, A115V/G214E and L151P). Trypsin treatment of RseP(V302P)-HM and RseP(G303E)-HM (data not shown) also resulted in the generation of a small amount of the ∼33 kDa fragment (Fig. 4A, V302P). A similar ∼33-kDa fragment was also detected by CBB staining when purified RseP(A115V/G214E)-HM was incubated with trypsin in detergent (Fig. 4B, A115V/G214E). No such fragment was detected with purified wild type RseP-HM (Fig. 4B, RseP+-HM), indicating that the mutations affected directly the trypsin sensitivity of RseP rather than its association with some other cellular proteins. The trypsin fragment generated in spheroplasts (Fig. 4C, lane 1) and that produced in a detergent extract (Fig. 4C, lane 5) exhibited essentially the same mobility on 10% SDS-PAGE. These results indicate that the PDZ-N mutations and also the PDZ-C mutations to a lesser extent induce a conformational change in the periplasmic region of RseP to allow its cleavage by trypsin at a specific site. To identify the approximate position of the cleavage, the sizes of the ∼33-kDa trypsin fragments produced from RseP(A115V/G214R)-HM in spheroplasts or in a detergent extract were compared with a series of N-terminal-truncated fragments of RseP-HM (Fig. 1A, ΔN1 and ΔN2). Their mobility on SDS-PAGE was slightly slower than that of RseP(ΔN2)-HM (Fig. 4C, lane 4). Thus, the site of trypsin cleavage lies around the interdomain region between PDZ-N and PDZ-C.
FIGURE 4.
Trypsin sensitivity of the wild type and PDZ-N and PDZ-C mutant forms of RseP-HM. A, trypsin digestion profiles of RseP-HM in spheroplasts. Cells of AD2249 carrying pKK11 (RseP-HM) or its derivatives (pSTD1002, pYGF13, and pSTD1131) having the indicated mutations were grown in L medium containing 1 mm isopropyl 1-thio-β-d-galactopyranoside and 1 mm cAMP for 2 h. Cells were then converted to spheroplasts by sucrose-lysozyme treatment and further treated with 5 μg/ml trypsin at 0 °C for the indicated time periods. Acid-denatured proteins were analyzed by SDS-PAGE and anti-Myc immunoblotting. B, trypsin digestion of the purified RseP-HM proteins. Purified preparations of RseP-HM and RseP(A115V/G214E)-HM were incubated with 5 μg/ml trypsin for the indicated time periods. After acid denaturation, proteins were analyzed by SDS-PAGE and CBB staining. C, mapping of the trypsin cleavage site. Trypsin-digested RseP(L151P)-HM (lane 1) and RseP(A115V/G214R)-HM (lane 5) (the same samples used for lane 14 of (A) and lane 9 of (B), respectively) were analyzed together with in vitro synthesized RseP-HM (lane 2), RseP(ΔN1)-HM (lane 3), and RseP(ΔN2)-HM (lane 4) by SDS-PAGE and anti-Myc immunoblotting. Open and closed arrowheads indicate intact RseP-HM and its ∼33 kDa tryptic fragment (TF). Circles indicate in vitro-synthesized RseP-HM and its N-terminal-truncated derivatives. Lane M was for molecular size markers.
In Vitro Reconstitution of PDZ-controlled Sequential RseA Cleavage by DegS and RseP—We then attempted to reproduce the RseP-catalyzed RseA proteolysis in vitro in a manner controlled by the PDZ domains. Our attempt to purify RseA having the intact N-terminal tail, to which a His6 tag was attached, was unsuccessful because such a construct tended to aggregate upon overexpression (data not shown). We then constructed His6-MBP-RseA, in which the cytoplasmic domain of RseA was replaced with His6-MBP (Fig. 5A). It was overproduced in the degS- and rseP-deleted strain and purified from detergent-solubilized membranes by Ni-NTA nickel-nitrilotriacetic acid affinity column chromatography (Fig. 5B). The resulting preparation contained the full-length protein as a major component along with minor degradation products (Fig. 5B, asterisks). We also purified His6-DegS and wild type RseP-HM (Fig. 5A, lanes 15 and 16).
FIGURE 5.
In vitro reproduction of the two-step cleavages of the model substrates by DegS and RseP. A, schematic representations of the His6-MBP-RseA and its cleavage by DegS and RseP. B, demonstration of the two-step substrate cleavages with purified components in detergent. His6-MBP-RseA was incubated with His6-DegS and RseP-HM as indicated. Lanes 15, 16, and 17 show the purified preparations of the indicated proteins individually. Proteins were analyzed by 12.5% SDS-PAGE and CBB staining. Note that the purified preparation of His6-MBP-RseA contained a minute amount of an in vivo generated degradation product (lane 9) with a size similar to the site-1-cleaved intermediate as well as to RseP-HM, and this contaminating fragment was converted to the site-2 cleavage product upon incubation with RseP in lane 14. C, effects of the PDZ mutation on the in vitro cleavage of the model substrates by RseP-HM. His6-MBP-RseA140 (5.8 μg/ml) (left panel) and His6-MBP-RseA (5 μg/ml) (right panel) were incubated with RseP-HM (13 μg/m/and 19 μg/ml for the reaction with the former and the latter substrates, respectively) and RseP (A115V/G214E)-HM (30 μg/m/and 43 μg/ml for the reaction with the former and the latter substrates, respectively). Samples were withdrawn at the indicated time points and analyzed by SDS-PAGE and CBB staining. For B and C, 140 and Full indicate His6-MBP-RseA140 and His6-MBP-RseA, respectively, whereas site-1 and site-2 indicate the DegS and the RseP cleavage products of the model substrates, respectively.
When His6-MBP-RseA was incubated with His6-DegS in the presence of an OMP peptide (YFF-COOH), it was converted to a smaller fragment (Fig. 5B, lanes 5 and 6). This conversion depended on DegS and the OMP peptide with free C terminus (lanes 1-4), representing the site-1 cleavage faithfully. We then subjected this site-1 cleavage product to the second incubation with RseP. The second incubation with RseP-HM, but not with a buffer control, produced a lower molecular mass product expected for the site-2 cleavage product (Fig. 5B, lanes 7 and 8). Purified RseP-HM was able to cleave His6-MBP-RseA140, a mimic of the DegS-cleaved intermediate, to generate a product whose electro-phoretic mobility was identical with that of the above site-2 cleavage product (data not shown and Fig. 5C, left panel). When His6-MBP-RseA was incubated simultaneously with His6-DegS and RseP-HM, it was converted to the site-2 cleavage product without appreciable accumulation of the site-1 cleavage product (lanes 11 and 12). On the other hand, His6-MBP-RseA, having the intact periplasmic region, did not receive any appreciable proteolysis by RseP-HM (lanes 13 and 14; see also Fig. 5C, right panel), indicating that the regulatory mechanism of RseP, to suppress its site-2 proteolytic activity against full-length RseA, was in operation under these in vitro conditions. Thus, our in vitro reaction using the purified and detergent-solubilized components faithfully reproduced the two-step cleavage of the “full-length” model substrate by DegS and RseP.
We then examined the PDZ mutational effects in the purified reaction system (Fig. 5C). Thus, we purified a representative RseP mutant, RseP(A115V/G214E)-HM. Consistent with the in vivo results, purified RseP(A115V/G214E)-HM was less active than wild type RseP-HM in cleaving His6-MBP-RseA140 such that 2.3 molar-fold of RseP(A115V/G214E)-HM over the wild type enzyme gave comparable kinetics (left panel). RseP(A115V/G214E)-HM degraded the full-length substrate significantly, in contrast to the wild type enzyme, which lacked the proteolytic activity against this substrate. These results suggest that the PDZ domains of RseP directly control the substrate specificity of the RseP protease.
DISCUSSION
In this work we showed that a number of mutations affecting a region N-terminal to the previously proposed PDZ domain compromised the negative regulation of RseP. We have shown unequivocally by structural determination that RseP has two PDZ domains with a new type of circular permutation in the polypeptide connectivity.
Our isolation of deregulated RseP mutants indicates that PDZ-N has a major role in the down-regulation of the RseP activity toward full-length RseA. Synergistic effects of mutations in PDZ-C suggest that PDZ-C and PDZ-N interact functionally in the regulation. On the other hand, the SAXS analysis of the covalently linked PDZ-N/PDZ-C domains suggests that their physical interaction is only moderate. It seems possible that the functional interaction between PDZ-N and PDZ-C is supported by other parts of RseP including the transmembrane domains. Alternatively, their interaction could be ligand-dependent.
Deletion of either PDZ-N or PDZ-C was found to be silent. However, the additional introduction of the mutations in the carboxylate binding loop of PDZ-C, which will impair the pre-sumptive ligand binding, into the PDZ-N-less variant of RseP-HM made it capable of cleaving full-length RseA, suggesting that PDZ-C can substitute for PDZ-N in regulating the RseP function, although PDZ-C appears to play an auxiliary role in intact RseP. In the absence of PDZ-N, PDZ-C could occupy the position at which PDZ-N is normally situated in intact RseP. It is conceivable that the positions of the PDZ domains in the RseP structure are important for their functionality in the negative regulation. In addition, the circularly permutated structures of the PDZ domains might be required for these domains to have a functional relative orientation with respect to the membrane embedded catalytic domain. The previously assigned PDZ domain (PDZ*) is actually composed of the C-terminal region of PDZ-N and a majority of PDZ-C such that its deletion resulted in generation of a single chimeric PDZ domain, most of which comes from the PDZ-N domain. This chimeric PDZ domain seems to be defective in negative regulation.
Our results suggest that PDZ-N and PDZ-C have differential roles in modulation of the RseP function. There is a precedent for functionally differentiated PDZ domains in an E. coli periplasmic protease, DegP. DegP carries tandemly positioned PDZ domains, the N-terminal one (PDZ1) of which is essential for the proteolytic activity and involved in substrate recognition, whereas the other (PDZ2) is mainly required to maintain the oligomeric structure (37, 38). Recent study showed that ligand binding to PDZ1 induces an allosteric activation of DegP (39). Similar PDZ-ligand-dependent activation has also been reported for DegS, a DegP homolog (9, 10). RseP and DegP are similar in that they have tandem PDZ domains, but the roles of these domains appear to be different between these two enzymes.
Most of the PDZ mutations of RseP lowered the intrinsic protease activity to cleave His6-MBP-RseA(LacYTM1)140 in vivo. Thus, the stimulated cleavage of intact RseA by the PDZ mutants is not ascribable to their elevated intrinsic protease activities. The recently revealed structure of an M. jannaschiic S2P protein (18) as well as our biochemical analyses of E. coli RseP based on the modifiability of engineered cysteine residues (16) showed that the proteolytic active sites of these enzymes in the conserved core membrane domains are largely sequestered from the surrounding lipidic as well as aqueous environments. Our present results indicate that mutations in the PDZ domains induce a conformational change of the periplasmic domain to expose a specific trypsin cleavage site. Also, a mutation in TM2 (A115V) enhanced the effects of the PDZ-N mutations. It is conceivable that the PDZ domains modulate the RseP function by inducing some structural changes in the periplasmic and transmembrane domains. The altered conformation might enable the full-length substrate to make access to the otherwise recessed protease active site of RseP.
We isolated a number of mutations with strong phenotypes at residues around the putative ligand binding groove of PDZ-N, including those known in some other well characterized PDZ domains to be critical for the ligand binding. We suggest that the PDZ-N function/conformation is modulated by binding of some unidentified ligand. Also, PDZ-C may have an ability to bind a ligand as well. Mutations in the carboxylate binding loop of PDZ-C caused deregulation of RseP when combined with a missense mutation or deletion of PDZ-N. Moreover, the ligand binding cleft of PDZ-C in crystals is associated with the C terminus of a neighboring molecule. On the other hand, the C-terminal portions of the βB strands in PDZ-N and PDZ-C contain an amino acid residue, Pro-217 and Pro-306, respectively. These features seem to be unfavorable for a mode of ligand binding shown for the PDZ domain of PSD95 in which an amino acid residue at the corresponding positions forms hydrogen bonds with the residue at the P(-2) position of the ligand (40). Recent studies suggest that all the four residues at positions P(-1) to P(-4) in a ligand can contribute to binding to PDZ domains (24). It would, thus, be possible that the residue at P(-2) in the putative ligands of PDZ-N and PDZ-C might be less important. A small α-helix positioned in front of PDZ-N (α′ in Fig. 1B) partially covers the ligand binding groove. It is tempting to assume that this helix is involved in modulation of the ligand binding. Consistent with this idea, our preliminary results showed that expression of an RseP derivative with a deletion of this helix led to moderate activation of σE in the absence of DegS.5 A possible role of this small helix should be addressed by further experiments.
Because the PDZ mutations that probably impair ligand binding enabled RseP to cleave full-length RseA, the ligand binding would negatively control the cleavage of full-length RseA by RseP, although the identity of ligand(s) is unknown. The PDZ-N and PDZ-C domains might cooperate in the regulation through binding of either common or different ligands. We showed that wild type RseP did not appreciably cleave the full-length model substrate, but the PDZ mutant proteins did so significantly in the in vitro reaction system using the purified enzymes and substrate. These observations suggest that the negative regulation of RseP in the cleavage of full-length RseA does not require other cellular proteins, at least stoichiometrically. In vivo experimental results showed that, whereas RseB was able to inhibit RseP-catalyzed cleavage of full-length RseA, this inhibition was significantly impaired by the absence of the intact RseP PDZ domains, suggesting that the function of RseB in this aspect is dependent on that of the RseP PDZ domains (21). RseB could be indirectly required to effectively maintain the PDZ-mediated negative regulation of RseP in vivo, for instance by affecting the ligand binding to the RseP PDZ domains. Further detailed study will be needed to elucidate the function of RseB.
One interesting possibility would be that RseA itself acts as a ligand of the RseP PDZ domain(s). Because PDZ domains generally binds a C-terminal peptide, the C terminus of RseA could be recognized by the RseP PDZ domains. This model excellently explains the release of inhibition after the DegS-dependent cleavage of the C-terminal region of RseA. However, our previous observation that several C-terminal-truncated RseA derivatives with different C-terminal sequences were still refractory to DegS-independent cleavage by RseP does not support this possibility (19). It is also known that an internal peptide can be a PDZ ligand (41, 42). The Gln-rich regions in the RseA periplasmic domain are also candidates for the ligands, as mutational alterations of these regions allow RseP to cleave RseA in a DegS-independent manner (19). In these cases, DegS-catalyzed site-1 cleavage of RseA removes the ligand-containing peptide, leading to release of the negative regulation. RseB, which is known to bind to the periplasmic region of RseA (43), might facilitate or stabilize the interaction between RseP and RseA, as suggested previously (21). It is also possible that some small peptides which had been co-purified with RseP (Fig. 5B) acted as a ligand. In this case a signal that releases the PDZ ligand-mediated negative regulation under the stressed conditions could exist separately.
Recent studies identified several novel cues, including accumulation of lipid A intermediates (44) and abnormal inner membrane proteins (45) for the σE pathway activation. Lipid A intermediates and most inner membrane proteins should lack the DegS-activating signal conserved in outer membrane proteins (C-terminal YXF peptide). They might activate the σE pathway in a DegS-independent fashion, for example, through release of the PDZ-mediated negative regulation of RseP.
Recent determination of the crystal structure of a M. jannaschiic S2P homolog has provided useful information to understand the mechanism of intramembrane proteolysis by the S2P family proteases. However, as M. jannaschiic S2P lacks a PDZ domain, the regulatory aspects involving PDZs remain to be explored. Our structures of the PDZ domains of RseP should serve as the first step toward this direction. Determination of the whole structure of RseP is a next challenge essential for our understanding of the mechanism of the PDZ domain-mediated regulation of the S2P family proteases.
Supplementary Material
Acknowledgments
We thank H. Mori and S. Chiba for discussion, T. J. Silhavy for the gift of a strain, and D. Kohda for kind help in the first crystallization trial of RseP PDZ-C with the TOPAZ screening chip. The synchrotron radiation experiments were performed at RIKEN Structural Biology Beamline I (BL45XU) at SPring-8 with the approval of RIKEN SPring-8 Center (Proposal 20080070).
The atomic coordinates and structure factors (codes 2ZPL and 2ZPM) have been deposited in the Protein Data Bank, Research Collaboratory for Structural Bioinformatics, Rutgers University, New Brunswick, NJ (http://www.rcsb.org/).
This work was supported by a grant-in-aid from the Japan Society for the Promotion of Science (to Y. A., K. Inaba, and K. Ito) and from the Ministry of Education, Culture, Sports, Science, and Technology (MEXT), Japan (to S. A. and Y. A.), by “Targeted Proteins Research Program from MEXT, Japan (to M. S.), and by Kyushu University Interdisciplinary Program in Education and Projects in Research Development (to K. Inaba). The costs of publication of this article were defrayed in part by the payment of page charges. This article must therefore be hereby marked “advertisement” in accordance with 18 U.S.C. Section 1734 solely to indicate this fact.
The on-line version of this article (available at http://www.jbc.org) contains supplemental data and Figs. S1-S4.
Footnotes
The abbreviations used are: X-gal, 5-bromo-4-chloro-3-indolyl-β-d-galactopyranoside; HA, hemagglutinin; SeMet, selenomethionine; CBB, Coomassie Brilliant Blue; LacYTM1, the first transmembrane region of LacY; RseP-HM, a C-terminal His6-Myc epitope-tagged derivative of RseP; PDZ*, the previously assigned PDZ-like domain of RseP; PDZ-N, the N-terminal PDZ domain of RseP; PDZ-NC, the linked PDZ-N and PDZ-C domains; PDZ-C, the C-terminal PDZ domain of RseP; SAXS, small angle X-ray scattering.
Y. Akiyama, unpublished results.
References
- 1.Raivio, T. L. (2005) Mol. Microbiol. 561119 -1128 [DOI] [PubMed] [Google Scholar]
- 2.Rowley, G., Spector, M., Kormanec, J., and Roberts, M. (2006) Nat. Rev. Microbiol. 4 383-394 [DOI] [PubMed] [Google Scholar]
- 3.Hayden, J. D., and Ades, S. E. (2008) PLoS ONE 3e1573. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Missiakas, D., Mayer, M. P., Lemaire, M., Georgopoulos, C., and Raina, S. (1997) Mol. Microbiol. 24 355-371 [DOI] [PubMed] [Google Scholar]
- 5.De Las Peñas, A., Connolly, L., and Gross, C. A. (1997) Mol. Microbiol. 24 373-385 [DOI] [PubMed] [Google Scholar]
- 6.Campbell, E. A., Tupy, J. L., Gruber, T. M., Wang, S., Sharp, M. M., Gross, C. A., and Darst, S. A. (2003) Mol. Cell 111067 -1078 [DOI] [PubMed] [Google Scholar]
- 7.Ades, S. E., Connolly, L. E., Alba, B. M., and Gross, C. A. (1999) Genes Dev. 132449 -2461 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Walsh, N. P., Alba, B. M., Bose, B., Gross, C. A., and Sauer, R. T. (2003) Cell 113 61-71 [DOI] [PubMed] [Google Scholar]
- 9.Hasselblatt, H., Kurzbauer, R., Wilken, C., Krojer, T., Sawa, J., Kurt, J., Kirk, R., Hasenbein, S., Ehrmann, M., and Clausen, T. (2007) Genes Dev. 212659 -2670 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Sohn, J., Grant, R. A., and Sauer, R. T. (2007) Cell 131572 -583 [DOI] [PubMed] [Google Scholar]
- 11.Kanehara, K., Ito, K., and Akiyama, Y. (2002) Genes Dev. 162147 -2155 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Alba, B. M., Leeds, J. A., Onufryk, C., Lu, C. Z., and Gross, C. A. (2002) Genes Dev. 162156 -2168 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Akiyama, Y., Kanehara, K., and Ito, K. (2004) EMBO J. 234434 -4442 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Chaba, R., Grigorova, I. L., Flynn, J. M., Baker, T. A., and Gross, C. A. (2007) Genes Dev. 21 124-136 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Kinch, L. N., Ginalski, K., and Grishin, N. V. (2006) Protein Sci. 1584 -93 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Koide, K., Maegawa, S., Ito, K., and Akiyama, Y. (2007) J. Biol. Chem. 2824553 -4560 [DOI] [PubMed] [Google Scholar]
- 17.Koide, K., Ito, K., and Akiyama, Y. (2008) J. Biol. Chem. 2839562 -9570 [DOI] [PubMed] [Google Scholar]
- 18.Feng, L., Yan, H., Wu, Z., Yan, N., Wang, Z., Jeffrey, P. D., and Shi, Y. (2007) Science 3181608 -1612 [DOI] [PubMed] [Google Scholar]
- 19.Kanehara, K., Ito, K., and Akiyama, Y. (2003) EMBO J. 226389 -6398 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Bohn, C., Collier, J., and Bouloc, P. (2004) Mol. Microbiol. 52427 -435 [DOI] [PubMed] [Google Scholar]
- 21.Grigorova, I. L., Chaba, R., Zhong, H. J., Alba, B. M., Rhodius, V., Herman, C., and Gross, C. A. (2004) Genes Dev. 182686 -2697 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Harris, B. Z., and Lim, W. A. (2001) J. Cell Sci. 1143219 -3231 [DOI] [PubMed] [Google Scholar]
- 23.Jeleñ, F., Oleksy, A., Smietana, K., and Otlewski, J. (2003) Acta Biochim. Pol. 50985 -1017 [PubMed] [Google Scholar]
- 24.Stiffler, M. A., Chen, J. R., Grantcharova, V. P., Lei, Y., Fuchs, D., Allen, J. E., Zaslavskaia, L. A., and MacBeath, G. (2007) Science 317364 -369 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Otwinowski, Z., and Minor, W. (1997) Methods Enzymol. 276307 -326 [DOI] [PubMed] [Google Scholar]
- 26.Collaborative Computational Project, N. (1994) Acta Crystallogr. 50760 -763 [Google Scholar]
- 27.Shelderick, G. (2008) Acta Crystallogr. 64112 -122 [DOI] [PubMed] [Google Scholar]
- 28.Pape, T., and Schneider, T. R. (2004) J. Appl. Crystallogr. 37843 -844 [Google Scholar]
- 29.Rudolph, M. G., Kelker, M. S., Schneider, T. R., Yeates, T. O., Oseroff, V., Heidary, D. K., Jennings, P. A., and Wilson, I. A. (2003) Acta Crystallogr. 59 290-298 [DOI] [PubMed] [Google Scholar]
- 30.Perrakis, A., Harkiolaki, M., Wilson, K. S., and Lamzin, V. S. (2001) Acta Crystallogr. 571445 -1450 [DOI] [PubMed] [Google Scholar]
- 31.Terwilliger, T. C., Adams, P. D., Moriarty, N. W., and Cohn, J. D. (2007) Acta Crystallogr. 63 101-107 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Fujisawa, T., Inoue, K., Oka, T., Iwamoto, H., Uruga, T., Kumasaka, T., Inoko, Y., Yagi, N., Yamamoto, M., and Ueki, T. (2000) J. Appl. Crystallogr. 33 797-800 [Google Scholar]
- 33.Ito, K., Kamikubo, H., Yagi, N., and Amemiya, Y. (2005) Japn. J. Applied Phys. 448684 -8691 [Google Scholar]
- 34.Akiyama, S., Nohara, A., Ito, K., and Maeda, Y. (2008) Mol. Cell 29703 -716 [DOI] [PubMed] [Google Scholar]
- 35.Svergun, D. I., Petoukhov, M. V., and Koch, M. H. J. (2001) Biophys. J. 802946 -2953 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Petoukhov, M. V., and Svergun, D. I. (2005) Biophy. J. 891237 -1250 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Spiess, C., Beil, A., and Ehrmann, M. (1999) Cell 97339 -347 [DOI] [PubMed] [Google Scholar]
- 38.Iwanczyk, J., Damjanovic, D., Kooistra, J., Leong, V., Jomaa, A., Ghirlando, R., and Ortega, J. (2007) J. Bacteriol. 1893176 -3186 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Krojer, T., Pangerl, K., Kurt, J., Sawa, J., Stingl, C., Mechtler, K., Huber, R., Ehrmann, M., and Clausen, T. (2008) Proc. Natl. Acad. Sci. U. S. A. 1057702 -7707 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Doyle, D. A., Lee, A., Lewis, J., Kim, E., Sheng, M., and MacKinnon, R. (1996) Cell 851067 -1076 [DOI] [PubMed] [Google Scholar]
- 41.Hillier, B. J., Christopherson, K. S., Prehoda, K. E., Bredt, D. S., and Lim, W. A. (1999) Science 284812 -815 [PubMed] [Google Scholar]
- 42.Penkert, R. R., DiVittorio, H. M., and Prehoda, K. E. (2004) Nat. Struct. Mol. Biol. 111122 -1127 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Cezairliyan, B. O., and Sauer, R. T. (2007) Proc. Natl. Acad. Sci. U. S. A. 1043771 -3776 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Tam, C., and Missiakas, D. (2005) Mol. Microbiol. 551403 -1412 [DOI] [PubMed] [Google Scholar]
- 45.Shimohata, N., Nagamori, S., Akiyama, Y., Kaback, H. R., and Ito, K. (2007) J. Cell Biol. 176307 -317 [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Wriggers, W., and Chacón, P. (2001) J. Appl. Crystallogr. 34 773-776 [Google Scholar]
- 47.Humphrey, W., Dalke, A., and Schulten, K. (1996) J. Mol. Graph. 1433 -38 [DOI] [PubMed] [Google Scholar]
- 48.Maegawa, S., Ito, K., and Akiyama, Y. (2005) Biochemistry 4413543 -13552 [DOI] [PubMed] [Google Scholar]
- 49.Casadaban, M. J. (1976) J. Mol. Biol. 104541 -555 [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.