Abstract
Cofactors are intimately involved in steroid-regulated gene expression. Two critical questions are (1) the steps at which cofactors exert their biological activities and (2) the nature of that activity. Here we show that a new mathematical theory of steroid hormone action can be used to deduce the kinetic properties and reaction sequence position for the functioning of any two cofactors relative to a concentration limiting step (CLS) and to each other. The predictions of the theory, which can be applied using graphical methods similar to those of enzyme kinetics, are validated by obtaining internally consistent data for pair-wise analyses of three cofactors (TIF2, sSMRT, and NCoR) in U2OS cells. The analysis of TIF2 and sSMRT actions on GR-induction of an endogenous gene gave results identical to those with an exogenous reporter. Thus new tools to determine previously unobtainable information about the nature and position of cofactor action in any process displaying first-order Hill plot kinetics are now available.
Introduction
Ligand-regulated gene induction is a productive experimental system for examining the mechanisms of gene expression. Steroid-regulated gene induction is an extensively studied paradigm with numerous well-defined events. The initially formed, intracellular receptor-steroid complex binds to specific enhancer-like DNA elements (called hormone response elements, or HREs) to eventually modify the rates of transcription of target genes. Other steps include recruitment of chromatin remodeling factors and cofactors that increase or decrease the rates of transcription [1]–[3].
One approach to defining steroid receptor actions at the molecular level has been to pair ChIP assays with genome-wide sequencing to identify all DNA binding sites of selected cofactors in the cellular genome [4]. Nonetheless, limitations to this approach remain. Not all factor binding sites are functionally active [5], [6]. The HREs may not regulate the closest gene [7], which is the default assignment. Methods to identify new cofactors are not generally available and the kinetic mechanism of most factors and cofactors is unknown. Because virtually all possible responses to steroid hormones have been observed with endogenous genes [8], no general mode of action exists. Finally, one can determine the temporal ordering of cofactor binding to DNA but no method exists to elucidate the temporal ordering of biological function. Cofactor binding to DNA is not equivalent to cofactor action. For example, paused RNA polymerase II is often present 50 bp downstream of the start of transcription but is not engaged in transcription [9]–[11]. Thus there is an unmet need for methods that can discern the precise nature (e.g., activator, non-competitive inhibitor) and temporal order of cofactor function.
Here we describe a method that establishes the functional mechanism and order of action of any active factor/cofactor relative to what we call a “concentration limiting step” (CLS). The method is derived from a recently developed theory of gene expression that is applicable to receptor-mediated transcriptional events that display a first-order Hill plot dose-response curve (FHDC) [12] for the gene product in experiments that reach equilibrium or steady state. The CLS is analogous to the rate-limiting step for a closed system and provides a reference point for the actions of all other cofactors. The method is based on analyzing graphs constructed from the maximal activity (Amax) and potency (EC50) of the FHDC. Importantly, two cofactors can be assayed simultaneously and the graphical method determines the functional nature and order of both cofactors relative to each other and the CLS. This approach is amenable to screening cofactors because prior knowledge about their action is not required.
Results
Application of the theory to analyzing the actions of cofactors
The graphical method is derived from our steroid-mediated gene induction theory, which can generate a parametric model (i.e. formula) for the dose-response curve of the final protein product with respect to the steroid concentration in the presence of an arbitrary number of cofactors [12] (see Text S1 for the derivation). The theory considers a sequence of reaction steps, each with the form
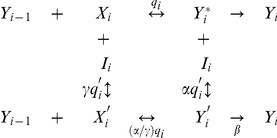 |
where Y
i is the reaction product of step i, X
i is an activating cofactor or activator, and I
i is an inhibiting cofactor or inhibitor. The labels on the reactions represent association constants for reversible reactions and reaction rates for nonreversible reactions. As in enzyme kinetics, we denote the case of 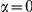 to be competitive inhibition,
to be competitive inhibition, 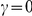 to be uncompetitive inhibition,
to be uncompetitive inhibition,  to be noncompetitive inhibition, and
to be noncompetitive inhibition, and  and
and  both nonzero to be mixed inhibition. The case of
both nonzero to be mixed inhibition. The case of 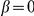 is called linear inhibition, and
is called linear inhibition, and 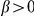 is called partial inhibition. In general, computing the dose-response curve for such a reaction sequence would be analytically intractable. However, imposing the experimentally observed constraint that the dose-response curve has a Hill-coefficient of one yields a closed-form expression for the dose-response curve in terms of the parameters of all the reactions.
is called partial inhibition. In general, computing the dose-response curve for such a reaction sequence would be analytically intractable. However, imposing the experimentally observed constraint that the dose-response curve has a Hill-coefficient of one yields a closed-form expression for the dose-response curve in terms of the parameters of all the reactions.
The theory identifies a CLS, which is a step in the reaction sequence in which products of the reactions following it (post-CLS steps) are so small that the amounts of cofactor bound are negligible compared to their free concentrations [12]. A cofactor can act before, at, or after the CLS. It can be an activator or one of the four types of inhibitor and each inhibitor can be linear or partial. Although there are eight possible types of inhibitor (e.g. partial uncompetitive, linear noncompetitive, etc.), some combinations are not possible (e.g. a competitor cannot be partial). Inhibitors refer to their action in their particular reaction and not to the final product. For example, a partial inhibitor can give a higher response if it diverts the output to a higher yielding pathway. In fact, the action of a partial uncompetitive inhibitor is similar to an activator acting in a post-CLS step. For two cofactors, there are at most 285 possible cases accounting for where they act with respect to each other and the CLS, with each cofactor being either an activator or one of eight possible types of inhibitors. It should be noted that not all types of inhibitors are physically viable or can act in all locations. For example, among all types of inhibition, only competitive inhibition occurs after the CLS and partial competitive inhibition cannot exist. Each viable case is represented by an explicit parametric model that can be compared to the data (see Text S1).
We previously deduced that Ubc9 was an activator acting after the CLS by fitting parametric models directly to the data [12]. However, in that case there were only three models to test. For the more general case with multiple cofactors, directly fitting models to the data is unwieldy. However, the models have very different qualitative behaviors for different types of cofactors and their positions of action. In particular, the EC50 (steroid concentration required for half-maximal activity) and Amax (maximal activity) behave very differently with changing cofactor concentration. The graphical method exploits these differences to predict mechanism and position from the properties of graphs of functions of EC50 and Amax versus cofactor concentration. Hence, the cofactor mechanism and position of action is inferred from the qualitative behavior of how the dose-response curve changes and does not require making direct estimates of the parameter values.
Graphical analysis of single cofactor actions
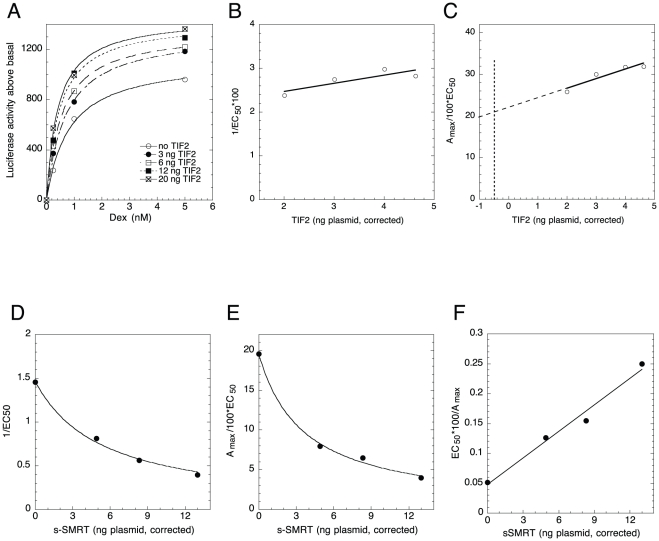
We applied our method to two well-known cofactors for glucocorticoid receptor (GR, also called NR3C1) transactivation: the coactivator TIF2 and the corepressor SMRT [13], [14]. To obtain easily quantifiable data for graphical analysis, the induction of a synthetic reporter gene (GREtkLUC) by GRs is followed in transiently transfected cells with different cofactor concentrations and triplicate sub-saturating concentrations of the glucocorticoid dexamethasone (Dex). The Amax and EC50 are abstracted from fits of the data points to a first order Hill function ( Fig. 1A ). Graphs of 1/EC50 vs. concentration and Amax/EC50 are then plotted.
Figure 1. Analysis of actions of excess TIF2 or sSMRT.
(A) Determination of Amax and EC50 by exact fit of gene induction response. Luciferase activity in transiently transfected U2OS cells (100 ng GREtkLUC reporter, 0.5 ng of GR, and the indicated amounts of TIF2 plasmid) is plotted against Dex concentration. The Amax and EC50 were determined from the best-fit curve to a first order Hill plot as described in Materials and Methods. Plots of 1/EC50 (B) and Amax/EC50 (C) vs. TIF2, corrected for non-linear protein expression as described in the Text S1. The dashed line is the extrapolation of the linear best-fit of the data. The dotted line equals the position of the y-axis at “true zero” of no TIF2 in cells. Plots of 1/EC50 (D) and Amax/EC50 (E) vs. sSMRT, corrected for non-linear protein expression. The Amax and EC50 for Dex induction of Luciferase activity in transiently transfected U2OS cells (100 ng GREtkLUC reporter, 0.5 ng of GR, 3 ng of TIF2 and the indicated amounts of sSMRT plasmid) was determined as in A. (F) Amax/EC50 vs. corrected sSMRT approaches zero at infinite sSMRT. A linear plot of Amax/EC50 vs. corrected sSMRT is diagnostic of an asymptote value equal to zero at infinite sSMRT. A positive asymptote value would give a non-linear, downward curving plot that can be linearized by subtracting the estimated asymptote from each of the values for Amax/EC50 in panel E and then plotting the reciprocal (1/[(Amax/EC50) – asymptote]). The combined results from this one representative experiment were seen in three other independent experiments.
Figure 1B shows the plot of 1/EC50 vs. transfected TIF2 plasmid (after correction for frequent non-linear expression of plasmid-encoded protein [see Text S1 and Fig. S1]) for an experiment with four TIF2 concentrations. The curve is seen to be increasing and according to the theory, the only possible parametric models are a straight line with a positive slope or a nonlinear rational function. The data are seen to be consistent with being linear. Although it is true that a nonlinear function is possible it is far less parsimonious. We thus first assume that the curves are linear and check for logical consistency with the other predictions (see Text S1 for full algorithm to determine shape of the curves). If linearity fails to give a logically consistent prediction then we can test the hypothesis that the curves are nonlinear. According to Table 1 , linear with positive slope is characteristic of TIF2 acting in one of three manners. (Note: all below graphs are for corrected non-linear protein expression.) The interpretation of the linear plot of Amax/EC50 vs. TIF2 ( Fig. 1C ) depends upon the y-axis intercept when the total TIF2 (endogenous plus added) equals zero. The relative amount of endogenous TIF2 was determined by Western blotting to be equivalent to 0.97 ng of transfected TIF2 plasmid (not shown). Experiments with fluorescent-tagged proteins indicated that ∼50% of the cells are transfected. Thus, endogenous TIF2 in transfected cells equals about 0.49 ng TIF2 plasmid. The average x-axis intercept from 4 experiments like Fig. 1C is −5.6±3.6 (S.D.) ng of TIF2 plasmid. Therefore the Amax/EC50 plots intersect the x-axis at less than “true zero” (i.e., less than no endogenous TIF2). Consequently the y-axis intercept is >0. From Table 1A, we now conclude that TIF2 is an activator after the CLS or the mathematically indistinguishable case of a partial uncompetitive inhibitor before the CLS.
Table 1. Algorithms for single factor plots for factor F.
| Plot parameters | Plot properties | Mechanistic conclusions |
| 1/EC50 vs. F | linear with zero slope (i.e. does not change with F) | 1) F = A at CLS or 2) F = PN before CLS |
| linear with positive slope | 1) F = A not at CLS; or 2) F = U before or at CLS, or 3) F = PU before CLS | |
| nonlinear decreasing curve (concave-up) | 1) F = C, 2) F = M before or at CLS | |
| nonlinear increasing curve (concave-down) | 1) F = LM or PM, before CLS; or 2) F = M at CLS | |
| Amax/EC50 vs. F | linear; y-axis intercept = 0 | 1) F = A before or at CLS |
| linear; y-axis intercept >0 | 1) F = A after CLS; or 2) F = PU before or at CLS | |
| nonlinear decreasing curve that approaches zero for large F | F = C before or at CLS | |
| nonlinear decreasing curve that approaches positive value for large F | 1) F = PM or PN, before or at CLS; or 2) F = C after CLS | |
| nonlinear increasing curve | 1) F = PM or PN, before or at CLS | |
| EC50/Amax vs. F | linear with positive slope | 1) F = C before or at CLS |
The originally described form of SMRT, sSMRT [15], lacks about 1000 amino-terminal residues [16]. Both sSMRT and full length SMRTs are corepressors of steroid receptor action. Graphical analysis of sSMRT as above with a constant, low amount of exogenous TIF2 (to increase the starting signal), yields a decreasing curve for 1/EC50 vs. sSMRT ( Fig. 1D ). The only allowable form of this curve by the theory is a decreasing nonlinear first-order decay plot. The plot of Amax/EC50 is also first-order decay ( Fig. 1E ). The important characteristic of decaying curves is whether they decay to zero or a positive value asymptotically. This can be determined graphically by plotting the reciprocal function (EC50/Amax) and seeing if it is linear, which is confirmed in Fig. 1F . The only interpretation in Table 1 compatible with Figs. 1D–F is that sSMRT is a competitive inhibitor before or at the CLS.
Graphical analysis of dual cofactor actions (TIF2 vs. sSMRT)
Our method can simultaneously analyze two cofactors and determine the site of each cofactor's action relative to the CLS, and usually relative to each other (see Table S1). Four plots (1/EC50 and Amax/EC50 for each cofactor), plus information on plasmid expression efficiency, are needed for maximal information, often with quantitation of endogenous cofactor. More than one classification of each plot may appear possible. However, by eliminating incompatible mechanistic consequences, one almost always reaches a single, internally logically consistent mechanistic description. Favorable conditions are with two cofactors of opposite activities, thereby yielding the greatest changes. As TIF2 and sSMRT fit these criteria, we analyzed their actions using four concentrations of each cofactor in all combinations, plus a control of no added cofactors.
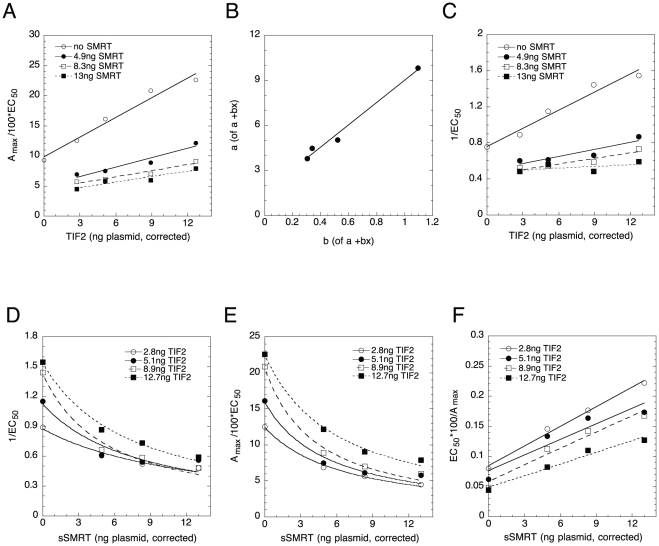
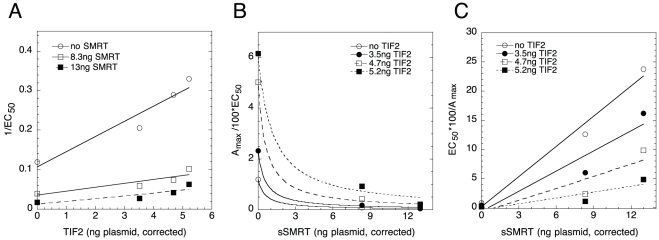
Figure 2A shows that plots of Amax/EC50 vs. TIF2 plasmid for increasing sSMRT consists of lines with progressively lower positive slopes. Western blots indicated that endogenous TIF2 in these experiments equaled 4.1 ng of TIF2 plasmid (data not shown). Entries 23,25, 28, and 29 of Table S1 are consistent with these graphs. Deciding between these entries for Fig. 2A depends upon the x- and y-axis coordinates of the intersection point of the lines. The x-axis value was determined, from what we call an “a vs. b plot” ( Fig. 2B ; see Methods), to be more negative (Ave. = −8.6±1.4, S.D., n = 4) than the endogenous TIF2 (−4.1), while the y-axis value was 2.3±2.2. This restricts the possible graphical interpretations to entries 25 and 29. Coupled with the unique assignment of entry 7 to 1/EC50 vs. TIF2 plots ( Fig. 2C ; x-axis intercept = −8.9±3.0)), the possible behavior of TIF2 and sSMRT reduces to: TIF2 is an activator acting after the CLS, sSMRT is a competitive inhibitor acting at the CLS and before TIF2. The other plots of 1/EC50 vs. sSMRT ( Fig. 2D ), Amax/EC50 vs. sSMRT ( Fig. 2E ), and EC50/Amax vs. sSMRT ( Fig. 2F ) can each be described by entries 18, 32, and 37 respectively with the same unique mechanistic interpretation as above.
Figure 2. Analysis of combined actions of TIF2 and sSMRT.
(A) Graph of Amax/EC50 vs. TIF2. The results of one representative experiment (n = 4), conducted as in Fig. 1, are shown for the indicated amounts of transfected TIF2 plasmid, corrected for non-linear protein expression. (B). Determination of intersection coordinates. The coefficients “a” and “b” of the linear plots in A, described by y = a+bx, are graphed. The negative value of the slope, and y-axis intercept, in this graph give the x- and y-axis the values respectively for intersection point of the lines in A, as described in the text and Text S1. Data for one representative graph each (n = 4) of 1/EC50 vs. TIF2 (C) or sSMRT (D), of Amax/EC50 vs. sSMRT (E) and of EC50/Amax vs. sSMRT (F) were acquired as in A and plotted against the indicated amounts of transfected plasmids, each of which have been corrected for non-linear protein expression.
Graphical analysis of TIF2 and NCoR competition
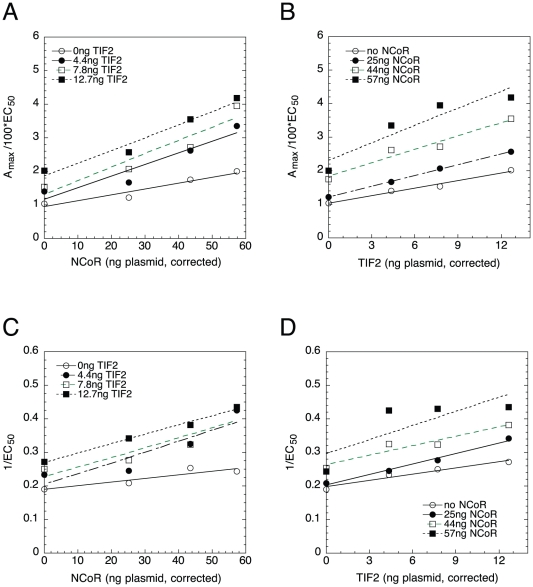
NCoR is another well-documented corepressor that is thought to act like sSMRT [13], [14]. We investigated the actions of NCoR competing with TIF2 under the conditions of Fig. 2. Western blots were used to correct for non-linear expression and to determine relative endogenous TIF2 (4.1 ng TIF2 plasmid) and NCoR (8.3 ng NCoR plasmid) levels. Given these values, only entries 26 or 27 can describe the graph of Amax/EC50 vs. NCoR ( Fig. 3A ), with calculated x- and y-intersection points of −30.0±26.7 and 1.2±1.0 (S.D., n = 6) respectively. Thus NCoR is an activator acting after the CLS and TIF2 is an activator acting after the CLS. Each of the other graphs has more than one potential classification due to insufficient precision in the graphical intersection points: Amax/EC50 vs. TIF2 ( Fig. 3B ; entries 24 and 27), 1/EC50 vs. NCoR ( Fig. 3C ; 6, 7, and 11), and 1/EC50 vs. TIF2 ( Fig. 3D ; entries 5–7 and 11). However, most graphical options are incompatible with both factors being activators after the CLS and can be eliminated to yield: NCoR is an activator acting after the CLS, TIF2 is an activator after the CLS, and TIF2 and NCoR do not act at the same step. While these data are not yet able to determine whether TIF2 is an activator after or before NCoR, the conclusion that TIF2 is an activator after the CLS is identical to the above results with TIF2 vs. sSMRT.
Figure 3. Analysis of combined actions of TIF2 and NCoR.
Graphs of Amax/EC50 vs. NCoR (A) and TIF2 (B) and of 1/EC50 vs. NCoR (C) and TIF2 (D). The results of one representative experiment (n = 6), conducted as in Fig. 1, are shown for the indicated amounts of transfected TIF2 and NCoR plasmids, corrected for non-linear protein expression.
Graphical analysis of sSMRT and NCoR competition
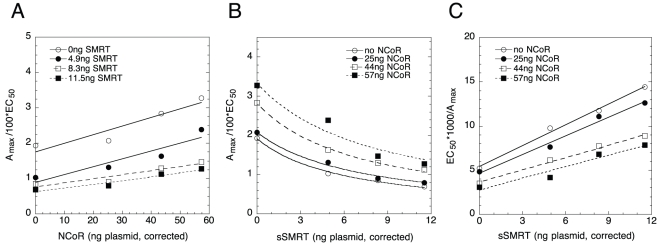
To confirm the above different actions of sSMRT and NCoR, we directly competed the activities of both cofactors under conditions of Fig. 2. Here the key graphs again involve Amax/EC50. For NCoR ( Fig. 4A ), the calculated x-axis intersection of −74±31 is clearly less than the −8.3 of endogenous NCoR while the y-axis intersection is −0.04±0.4 (S.D., n = 3). These properties are uniquely defined by entry 25. Deciding whether Amax/EC50 vs. sSMRT ( Fig. 4B ) is described by entry 32 or 33 is resolved by the linear plots of EC50/Amax ( Fig. 4C ). This characteristic of entry 37 is obtained only when nonlinear decreasing Amax/EC50 plots approach zero with infinite F2. At this point, we can conclude that: NCoR is an activator after the CLS, sSMRT is a competitive inhibitor before or at the CLS, and NCoR acts after sSMRT. The 1/EC50 graphs with either factor are consistent with several interpretations (entries 8–10 and 12 for NCoR and 18 and 19 for sSMRT; Fig. S2), one of which is, for each factor, the same as that derived from the other graphs. It is not possible with these data, though, to determine whether sSMRT acts before or at the CLS.
Figure 4. Analysis of combined actions of NCoR and sSMRT.
Graphs of (A) Amax/EC50 vs. NCoR, (B) Amax/EC50 vs. sSMRT, and (C) EC50/Amax vs. sSMRT. The results of one representative experiment of a total of three independent experiments, conducted as in Fig. 1, are shown for the indicated amounts of exogenous NCoR and sSMRT plasmids, after correction for non-linear protein expression.
Graphical analysis of TIF2 and sSMRT competition for induction of an endogenous gene
The relevance of the above competition of TIF2 and sSMRT, and the applicability of the graphical analysis to normal cellular biology, was next examined in the context of the GR-inducible IGFBP1 gene in U2OS cells [17]–[19]. qRT-PCR quantitation of IGFBP1 mRNA induction employed SyberGreen, which gives relative total activities. Therefore, the Amax in these experiments is actually the closely related fold-induction above basal level. 1/EC50 vs. TIF2 graphs ( Fig. 5A ) are exclusively described by entry 9 because the “a vs. b plots” give an x-axis intersection point (−3.5±0.6, S.D., n = 4) that is much more negative than the endogenous TIF2 in these experiments of −0.49. The non-linear Amax/EC50 ( Fig. 5B ), with the linear EC50/Amax ( Fig. 5C ), graphs vs. sSMRT implicate entries 32 and 37 respectively, which define the factors as: TIF2 is an activator after the CLS, sSMRT is a competitive inhibitor at the CLS, and TIF2 acts after sSMRT. The predictions from the other graphs (Amax/EC50 vs. TIF2 and 1/EC50 vs sSMRT; Fig. S3) are entirely consistent with this interpretation. Gratifyingly, this conclusion is the same as seen above for these factors with an exogenous reporter gene.
Figure 5. Analysis of combined actions of TIF2 and sSMRT for induction of the endogenous gene, IGFBP1.
Plots of (A) 1/EC50 vs. TIF2, (B) Amax/100×EC50 vs. sSMRT, and (C) EC50/Amax vs. sSMRT. The results of one representative experiment of a total of four independent experiments, conducted similarly as in Fig. 1, are shown for the indicated amounts of transfected NCoR and sSMRT plasmids, after correction for non-linear protein expression. The differences are that IGFBP1 mRNA (instead of luciferase) was determined by qRT-PCR and fold-induction is used in place of Amax (see text).
Discussion
This report describes how a new mathematical theory of steroid-induced gene transcription [12] gives previously unobtainable mechanistic information about GR- (NR3C1-) regulated gene induction. This information could be obtained from direct curve fitting of predictions from our theory but it would be extremely computationally intensive. Instead, we carefully examined the underlying equations of the theory. This analysis unexpectedly revealed that plots used in enzyme kinetics can be adapted to analyze the role of cofactors in GR-mediated gene transactivation. These graphical methods give information regarding the specific cofactor function and also the location in the reaction scheme where the cofactor is acting relative to a CLS. A major advantage of these graphical methods is that they are readily employed without the needs of elaborate and extensive mathematical calculations. Our method works with a single added cofactor but is more informative when two cofactors are examined together, in which case the nature and location of action of both cofactors can be determined simultaneously. Even if a precise mechanistic interpretation is not possible, differences in the graphical analyses can reveal mechanistic non-equivalence of two cofactors that otherwise may be thought to share a common mode of action.
Several approaches were taken to validate our graphical analysis using GR-regulated gene induction as the model system. The p160 family member, TIF2, is a well-documented coactivator of steroid receptors [13], [20], [21]. In three different systems, TIF2 was always a activator acting after the CLS. Likewise, the corepressor sSMRT [13], [20], [21] was always found to be a competitive inhibitor, acting at the CLS in two systems and at or before the CLS in the third. To our knowledge, this is the first identification regarding either the kinetics of mechanism, or placement of action, of TIF2 or sSMRT.
A major test of our graphical methods came upon examining the actions of three cofactors (TIF2, sSMRT, and NCoR) in all possible pair-wise combinations. The mechanism and site of action of each cofactor was the same in competition assays with both of the other two cofactors. This internal logical consistency of cofactor mechanism and site of action strongly validates our new method. It also suggests that there is just one CLS, which does not change with assay conditions. This is important in ascertaining whether the CLS is invariant and a common marker step when comparing the actions of different cofactors.
Our extended model and associated graphical analysis also works with endogenous genes. The fact that the mechanisms of TIF2 and sSMRT in IGFBP1 induction involve identical sites of action as seen with the transiently transfected GREtkLUC reporter is further support for both the utility of our method and for the invariance of the CLS. Also, it has long been assumed that most steps of GR-regulated gene induction are the same for exogenous and endogenous genes. We recently demonstrated that the modulatory activity of TIF2 is very similar with both types of reporters [22]. The present data suggest that the mode and site of TIF2 action may also be the same with both classes of reporters.
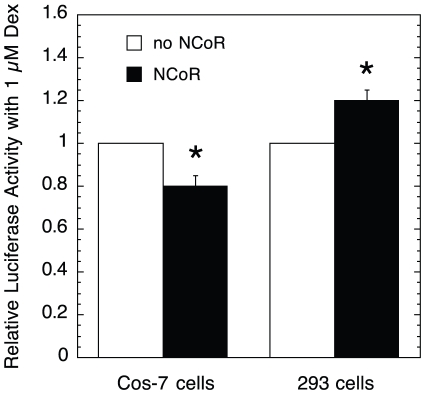
The different manners of NCoR and sSMRT action was unexpected. NCoR and sSMRT are generally thought to act as corepressors by binding to nearly the same site as coactivators, thereby excluding coactivator binding [23], [24]. Not only does our graphical analysis indicate different mechanisms of action at different steps for NCoR and sSMRT but also NCoR, being kinetically defined as an activator, could be called a coactivator because it increases the Amax in U2OS cells. However, this property of NCoR is cell-dependent, with NCoR decreasing the Amax from GREtkLUC in Cos-7 cells and increasing the Amax in 293 cells ( Fig. 6 ). Such cell-selective differences are not unique and have been seen not only for sSMRT with estrogen receptors in HeLa cells [25], and GR in CV-1 vs. 1407.2 cells and progesterone receptors in 1470.2 cells [26], but also with ZAC1b [27] and CIA with estrogen receptors [28], and with SRAP activation of androgen receptors and repression of VP16 transactivation [29]. In a recent report, 19 of 25 cofactors examined have dual activity and can increase and decrease the activity of agonist steroids with the androgen receptor in a gene-dependent manner [30]. It will be interesting to use our graphical analysis to determine the precise kinetic properties of the context-dependent activities of these versatile cofactors.
Figure 6. Cell-selective activity of NCoR.
Cos-7 or 293 cells were transiently transfected with the same amount of plasmids for glucocorticoid receptor (0.5 ng), NCoR (90 ng) or equimolar amount of control plasmid (51.6 ng of human serum albumin in the pCMX vector), and GREtkLUC reporter and induced with EtOH ±1 µM Dex as described in Materials and Methods. The total Luciferase activity with 1 µM Dex with each treatment was expressed relative to that with no added NCoR (± S.D., n = 3). P values: *≤0.021.
Our theory and methods are not without limitations. Any process not displaying FHDC kinetics of induction or repression cannot be analyzed. Conversely, our approach is applicable to any process yielding FHDC kinetics such as hormones in general [31], G-protein mediated responses [32], [33], the developmental-specific genes of Drosophila embryogenesis [34], and glucocorticoid-induced apoptosis of thymocytes [35]. While we have yet to discover a straightforward method to determine the graphical consequences of every mechanistic combination, the most common mechanisms listed in Table S1 cover over 74% of the possible cases. Thus, some graphical patterns may emerge that have not yet been linked to a specific mechanism. Nonetheless, the current range of analyzable mechanisms offers a powerful method for determining two aspects of steroid hormone action for which no alternative method presently exists: the kinetic actions of cofactors and the relative ordering of cofactor activity in the overall pathway. With this last capability, it is now theoretically possible to construct an ordered sequence based on the biological function of cofactors, much as in the ordering of pathways by epistasis analysis, even if the biochemical properties of the cofactors are not known. As a start, the present experiments in U2OS cells reveal that TIF2 and NCoR act at different steps after the CLS and after sSMRT, which acts at the CLS. This type of information should be useful in identifying downstream steps for therapeutic intervention, thereby reducing the number of side-effects that accompany the inhibition of upstream steps in steroid hormone action.
Materials and Methods
Unless otherwise indicated, all cell growth was at 37°C and all other operations were performed at r.t.
Chemicals
Dexamethasone (Dex) was purchased from Sigma (St. Louis, MO). Dex-21-mesylate (DM) was from Steraloids (Newport, RI). Anti-TIF2 mouse monoclonal antibody (No. 610984; BD Biosciences, San Jose, CA), anti-β actin monoclonal antibody (Sigma, St. Louis, MO), and goat anti-mouse horseradish peroxidase (Santa Cruz Biotechnology, Inc., Santa Cruz, CA) are commercially available. Rabbit anti-NCoR antibody was a gift from Goeffrey Rosenfeld (UC San Diego, CA).
Plasmids
The GREtkLUC reporter is a synthetic plasmid with a tandem repeat of the second glucocorticoid response element (GRE) of the rat tyrosine aminotransferase gene fused upstream of the thymidine kinase (tk) promoter driving the firefly luciferase (LUC) gene [36]. Renilla TS is a gift from Nasreldin M. Ibrahim, Otto Fröhlich, and S. Russ Price (Emory University School of Medicine), pSG5/TIF2 is from Heinrich Gronemeyer (IGBMC, Strasbourg, France), pCMX/sSMRT is from Ronald Evans (Salk Institute, La Jolla, CA), NCoR/Flag is from Geoff Rosenfeld (University of California-San Diego, San Diego, CA), and rat GR/pSG5, pSG5/human serum albumin, pCMX/human serum albumin, and pBSK have been previously described [37]–[40]. The Renilla null luciferase reporter is from Promega (Madison, WI).
Cells and Growth Conditions
U2OS human osteosarcoma cells (from ATTC, #HTB-96) were grown in high glucose DMEM (Invitrogen, Carlsbad, CA) with 10% fetal bovine serum as previously described for U2OS cells with stably transfected GR (17). Cells were split at 3- to 4-day intervals and used in experiments at approximately 60–90% confluence. For steroid treatments, Dex and DM solutions were prepared in 100% ethanol and diluted ≥1∶100 into growth medium. Growth medium for the cells was then replaced with ethanol- or steroid-medium for the indicated incubation time.
Transient Transfection and Reporter Analysis
U2OS cells (20,000 cells per well) were seeded 24 h before transfection in 24-well plates. GREtkLUC reporter plasmid (100 ng/well) and other plasmids (total DNA adjusted to 300 ng/well with pBluescriptII SK+ [Stratagene, Santa Clara, CA]) were transiently transfected with Lipofectamine 2000 (Invitrogen, Carlsbad, CA) according to the manufacturer's instructions. Cells were treated 24 h post-transfection with steroids for 16–20 h and assayed for luciferase activity using a dual-luciferase reporter assay (Promega, Madison, WI) on a Centro XS3 LB 960 luminometer (Berthold Technologies, Oak Ridge, TN). Luciferase activity was normalized to Renilla TS as an internal control.
Quantitative Real-Time PCR (qRT-PCR)
U2OS cells (150,000–200,000 cells per well) were seeded 24 h before transfection in 6-well plates. Plasmids (total DNA = 1500 ng/well) were transiently transfected with Lipofectamine 2000 (Invitrogen) according to the manufacturer's instructions in the same ratios as used for 24-well dishes. Cells were treated 24 h post-transfection with various concentrations of Dex and DM for 20 h. Total RNA was extracted using TriZol reagent (Invitrogen) and cDNA was synthesized using SuperScript III, First-Strand Synthesis Kit (Invitrogen) per manufacturer's recommendations. The relative expression level of the corresponding cDNA was quantified using SyberGreen in an ABI 7900HT real-time PCR system. The quantification was normalized against β-Actin using the 2−ΔΔCT method. For all qRT-PCR reactions, primer efficiencies were 100% (±10%), appropriate no-RT and template free controls were used, and primer melting curves were assessed to ensure specificity of the PCR products.
Western Blotting
U2OS cells were transiently transfected with TIF2 expression plasmid under conditions identical to those above. After 48 h, cells were lysed using Cytobuster Protein Extraction Reagent (EMD Chemicals, Gibbstown, NJ) and equal amounts of lysate run on 4–12% Bis-Tris NuPAGE gels (Invitrogen) per manufacturer's instructions. Western blots were prepared, probed with mouse anti-TIF2 or mouse anti-β actin antibody followed by goat anti-mouse horseradish peroxidase. Bands were visualized with enhanced chemiluminescence detection reagents as described by the manufacturer (GE Healthcare, Piscataway, NJ). The relative concentration of target proteins was determined by densitometric analysis of the exposed film.
Data analysis
Statistical significance for Luciferase activity is assessed by the two-tailed Student's t test using InStat 2.03 (GraphPad Software, San Diego, CA). Each average of triplicates is treated as one value of the n experiments. When the difference between the SDs of two populations is significantly different, the t test is invalid so the Mann-Whitney or Alternate Welch t-test is used. A nonparametric test is used if the distribution of SD values is non-Gaussian.
The maximum induced activity (Amax) was obtained with saturating concentrations of agonist steroid, which was the lower of ≥100-fold higher than the EC50 or 10 µM. One dose-response curve yields one value of EC50 via a curve-fitting program (KaleidaGraph; Synergy Software, Reading, PA) following a first order Hill plot for increase or decay (R2 almost always ≥0.95). Alternatively, we can determine Amax and EC50 directly by fitting the curve
where V = Amax/EC50 and W = 1/EC50.
Graphical analysis
After obtaining Amax/EC50 and 1/EC50 from the dose response curve, graphs of Amax/EC50 vs. C and 1/EC50 vs. C, where C is the concentration of a cofactor, were constructed. In experiments where two cofactors were assayed, graphs of Amax/EC50 vs. C and 1/EC50 vs. C for each cofactor for varying concentrations of the other cofactor would be made. Thus, if an experiment used n concentrations for each cofactor, then there would be a total of four to six graphs, each with n separate curves, that are analyzed as described in the Text S1 and Fig. S4. The shape of the curves and how they change with the other cofactor are then compared to Table 1 for a single cofactor and Table S1 for two cofactors to determine the mechanism and order of action. Some entries in the table require knowledge of the endogenous concentration in the cell because the value at zero total concentration is required. This can be obtained from Western blots.
Many of the entries in Table S1 require an estimate of the intersection point of a set of linear regression fits to the graphs. We used the following method to make this estimate. Consider a family of lines  obtained from the linear regression fits for each graph labeled by i. The intersection point occurs if there exists a point (x,y) that simultaneously satisfies this system of equations. In general, because of noise and measurement error, the lines will not intersect exactly. Solving the system simultaneously for the intersection point is not viable because the system will generally be singular. However, rearranging the linear system yields
obtained from the linear regression fits for each graph labeled by i. The intersection point occurs if there exists a point (x,y) that simultaneously satisfies this system of equations. In general, because of noise and measurement error, the lines will not intersect exactly. Solving the system simultaneously for the intersection point is not viable because the system will generally be singular. However, rearranging the linear system yields 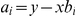 and a linear regression on the graph of a vs b, will give the least squares maximum likelihood estimate for the intersection points x and y. The a-intercept of the linear regression fit corresponds to the y value, and the negative slope corresponds to the x value, of the intersection point. The errors in the a vs b linear regression give an indication of the likelihood that the lines do in fact intersect.
and a linear regression on the graph of a vs b, will give the least squares maximum likelihood estimate for the intersection points x and y. The a-intercept of the linear regression fit corresponds to the y value, and the negative slope corresponds to the x value, of the intersection point. The errors in the a vs b linear regression give an indication of the likelihood that the lines do in fact intersect.
Supporting Information
Linearization of transfected plasmid concentrations. (A) Plot of normalized OD of Western blots (above endogenous TIF2) for different amounts of transfected TIF2 plasmid (but same amount of cell lysate protein) vs. OD of maximum amount of transfected TIF2 (20 ng). (B) Plot of normalized ODs of transfected TIF2 (above endogenous TIF2) vs. “linearized plasmid”. Amounts (ng) of linearized TIF2 plasmid were calculated as described in the Supplementary Material and then plotted against the normalized OD values of Fig. S1A. Error bars = S.D. (n = 2, each in duplicate).
(EPS)
Plots of 1/EC50 vs. cofactor for experiment of Fig. 4 . (A) 1/EC50 vs. sSMRT plasmid (corrected for non-linear expression) with different amounts of competing NCoR plasmid. (B) 1/EC50 vs. NCoR plasmid (corrected for non-linear expression) with different amounts of competing sSMRT plasmid.
(EPS)
Plots of EC50 and Amax vs. cofactor for experiment of Fig. 5 with endogenous IGFBP1 gene. (A) 1/EC50 vs. sSMRT plasmid (corrected for non-linear expression) with different amounts of competing TIF2 plasmid. (B) Amax/EC50 vs. TIF2 plasmid (corrected for non-linear expression) with different amounts of competing sSMRT plasmid.
(EPS)
Cartoon depicting process of restricting mechanistic scenarios of the six graphs used in analyzing the competition assays. Each circle represents one type of graph (labeled on perimeter) and is composed of different mechanistic scenarios (e.g., Scenario A for 1/EC50 vs. F1) for that graph (see Table 1). The area of overlap for all graphs (depicted by the filled space in the cartoon) represents the common, uniquely identified mechanism for that specific competition assay. This 4- to 6-fold requirement of “overlap” is a stringent test for determining the mechanistic explanation of any given competition assay.
(EPS)
Algorithms for two factor plots for factors F1 and F2.
(DOCX)
Derivation of the graphical method for analyzing the competitive action of factors and Correction for non-linear protein expression from transfected plasmids.
(DOCX)
Acknowledgments
We thank Sankar Adhya (NCI/NIH), Karen Ong (LBM) and members of the Steroid Hormones Section for helpful comments.
Footnotes
Competing Interests: The authors have declared that no competing interests exist.
Funding: Funding was provided by the Intramural Research Program of the NIH/NIDDK. The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
References
- 1.Metivier R, Reid G, Gannon F. Transcription in four dimensions: nuclear receptor-directed initiation of gene expression. EMBO Rep. 2006;7:161–167. doi: 10.1038/sj.embor.7400626. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Lonard DM, O'Malley BW. Nuclear receptor coregulators: judges, juries, and executioners of cellular regulation. Mol Cell. 2007;27:691–700. doi: 10.1016/j.molcel.2007.08.012. [DOI] [PubMed] [Google Scholar]
- 3.Wu SC, Zhang Y. Minireview: role of protein methylation and demethylation in nuclear hormone signaling. Mol Endocrinol. 2009;23:1323–1334. doi: 10.1210/me.2009-0131. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.John S, Sabo PJ, Johnson TA, Sung MH, Biddie SC, et al. Interaction of the glucocorticoid receptor with the chromatin landscape. Mol Cell. 2008;29:611–624. doi: 10.1016/j.molcel.2008.02.010. [DOI] [PubMed] [Google Scholar]
- 5.Carroll JS, Meyer CA, Song J, Li W, Geistlinger TR, et al. Genome-wide analysis of estrogen receptor binding sites. Nat Genet. 2006;38:1289–1297. doi: 10.1038/ng1901. [DOI] [PubMed] [Google Scholar]
- 6.Vivar OI, Zhao X, Saunier EF, Griffin C, Mayba OS, et al. Estrogen receptor beta binds to and regulates three distinct classes of target genes. J Biol Chem. 2010;285:22059–22066. doi: 10.1074/jbc.M110.114116. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Hakim O, John S, Ling JQ, Biddie SC, Hoffman AR, et al. Glucocorticoid receptor activation of the ciz1-lcn2 locus by long range interactions. J Biol Chem. 2009;284:6048–6052. doi: 10.1074/jbc.C800212200. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.John S, Johnson TA, Sung MH, Biddie SC, Trump S, et al. Kinetic complexity of the global response to glucocorticoid receptor action. Endocrinology. 2009;150:1766–1774. doi: 10.1210/en.2008-0863. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Gilchrist DA, Nechaev S, Lee C, Ghosh SK, Collins JB, et al. NELF-mediated stalling of Pol II can enhance gene expression by blocking promoter-proximal nucleosome assembly. Genes Dev. 2008;22:1921–1933. doi: 10.1101/gad.1643208. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Price DH. Poised polymerases: on your mark.get set.go! Mol Cell. 2008;30:7–10. doi: 10.1016/j.molcel.2008.03.001. [DOI] [PubMed] [Google Scholar]
- 11.Fuda NJ, Ardehali MB, Lis JT. Defining mechanisms that regulate RNA polymerase II transcription in vivo. Nature. 2009;461:186–192. doi: 10.1038/nature08449. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Ong KM, Blackford JA, Jr, Kagan BL, Simons SS, Jr, Chow CC. A new theoretical framework for gene induction and experimental comparisons. Proc Natl Acad Sci U S A. 2010;107:7107–7112. doi: 10.1073/pnas.0911095107. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.McKenna NJ, Lanz RB, O'Malley BW. Nuclear receptor coregulators: cellular and molecular biology. Endocrine Reviews. 1999;20:321–344. doi: 10.1210/edrv.20.3.0366. [DOI] [PubMed] [Google Scholar]
- 14.Xu L, Glass CK, Rosenfeld MG. Coactivator and corepressor complexes in nuclear receptor function. Current Opin Cell Biol. 1999;9:140–147. doi: 10.1016/S0959-437X(99)80021-5. [DOI] [PubMed] [Google Scholar]
- 15.Chen JD, Evans RM. A transcriptional co-repressor that interacts with nuclear hormone receptors. Nature. 1995;377:454–457. doi: 10.1038/377454a0. [DOI] [PubMed] [Google Scholar]
- 16.Ordentlich P, Downes M, Xie W, Genin A, Spinner NB, et al. Unique forms of human and mouse nuclear receptor corepressor SMRT. Proc Natl Acad Sci USA. 1999;96:2639–2644. doi: 10.1073/pnas.96.6.2639. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Rogatsky I, Wang JC, Derynck MK, Nonaka DF, Khodabakhsh DB, et al. Target-specific utilization of transcriptional regulatory surfaces by the glucocorticoid receptor. Proc Natl Acad Sci U S A. 2003;100:13845–13850. doi: 10.1073/pnas.2336092100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.He Y, Simons SS., Jr STAMP: a novel predicted factor assisting TIF2 actions in glucocorticoid receptor-mediated induction and repression. Mol Cell Biol. 2007;27:1467–1485. doi: 10.1128/MCB.01360-06. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Chen W, Dang T, Blind RD, Wang Z, Cavasotto CN, et al. Glucocorticoid receptor phosphorylation differentially affects target gene expression. Mol Endocrinol. 2008;22:1754–1766. doi: 10.1210/me.2007-0219. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Rosenfeld MG, Lunyak VV, Glass CK. Sensors and signals: a coactivator/corepressor/epigenetic code for integrating signal-dependent programs of transcriptional response. Genes Dev. 2006;20:1405–1428. doi: 10.1101/gad.1424806. [DOI] [PubMed] [Google Scholar]
- 21.Simons SS., Jr What goes on behind closed doors: physiological versus pharmacological steroid hormone actions. Bioessays. 2008;30:744–756. doi: 10.1002/bies.20792. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Luo M, Simons SS., Jr Modulation of glucocorticoid receptor induction properties by cofactors in peripheral blood mononuclear cells. Human Immunology. 2009;70:785–789. doi: 10.1016/j.humimm.2009.07.029. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Glass CK, Rosenfeld MG. The coregulator exchange in transcription functions of nuclear receptors. Genes and Develop. 2000;14:121–141. [PubMed] [Google Scholar]
- 24.Nagy L, Schwabe JW. Mechanism of the nuclear receptor molecular switch. Trends Biochem Sci. 2004;29:317–324. doi: 10.1016/j.tibs.2004.04.006. [DOI] [PubMed] [Google Scholar]
- 25.Karmakar S, Gao T, Pace MC, Oesterreich S, Smith CL. Cooperative activation of cyclin D1 and progesterone receptor gene expression by the SRC-3 coactivator and SMRT corepressor. Mol Endocrinol. 2010;24:1187–1202. doi: 10.1210/me.2009-0480. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Song L-N, Huse B, Rusconi S, Simons SS., Jr Transactivation specificity of glucocorticoid vs. progesterone receptors: role of functionally different interactions of transcription factors with amino- and carboxyl-terminal receptor domains. J Biol Chem. 2001;276:24806–24816. doi: 10.1074/jbc.M102610200. [DOI] [PubMed] [Google Scholar]
- 27.Huang S-M, Stallcup MR. Mouse Zac1, a transcriptional coactivator and repressor for nuclear receptors. Mol Cell Biol. 2000;20:1855–1867. doi: 10.1128/mcb.20.5.1855-1867.2000. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Sauve F, McBroom LD, Gallant J, Moraitis AN, Labrie F, et al. CIA, a novel estrogen receptor coactivator with a bifunctional nuclear receptor interacting determinant. Mol Cell Biol. 2001;21:343–353. doi: 10.1128/MCB.21.1.343-353.2001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Chooniedass-Kothari S, Hamedani MK, Auge C, Wang X, Carascossa S, et al. The steroid receptor RNA activator protein is recruited to promoter regions and acts as a transcriptional repressor. FEBS Lett. 2010;584:2218–2224. doi: 10.1016/j.febslet.2010.04.022. [DOI] [PubMed] [Google Scholar]
- 30.Babur O, Demir E, Gonen M, Sander C, Dogrusoz U. Discovering modulators of gene expression. Nucleic Acids Res. 2010;38:5648–5656. doi: 10.1093/nar/gkq287. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Loeb JN, Strickland S. Hormone binding and coupled response relationships in systems dependent on the generation of secondary mediators. Mol Endocrinol. 1987;1:75–82. doi: 10.1210/mend-1-1-75. [DOI] [PubMed] [Google Scholar]
- 32.Kenakin T. Efficacy in drug receptor theory: outdated concept or under-valued tool? Trends Pharmacol Sci. 1999;20:400–405. doi: 10.1016/s0165-6147(99)01361-9. [DOI] [PubMed] [Google Scholar]
- 33.Newton CL, Whay AM, McArdle CA, Zhang M, van Koppen CJ, et al. Rescue of expression and signaling of human luteinizing hormone G protein-coupled receptor mutants with an allosterically binding small-molecule agonist. Proc Natl Acad Sci U S A. 2011;108:7172–7176. doi: 10.1073/pnas.1015723108. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Karim FD, Thummel CS. Temporal coordination of regulatory gene expression by the steroid hormone ecdysone. EMBO J. 1992;11:4083–4093. doi: 10.1002/j.1460-2075.1992.tb05501.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Reichardt HM, Umland T, Bauer A, Kretz O, Schutz G. Mice with an increased glucocorticoid receptor gene dosage show enhanced resistance to stress and endotoxic shock. Mol Cell Biol. 2000;20:9009–9017. doi: 10.1128/mcb.20.23.9009-9017.2000. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Sarlis NJ, Bayly SF, Szapary D, Simons SS., Jr Quantity of partial agonist activity for antiglucocorticoids complexed with mutant glucocorticoid receptors is constant in two different transactivation assays but not predictable from steroid structure. J Steroid Biochem Molec Biol. 1999;68:89–102. doi: 10.1016/s0960-0760(99)00021-7. [DOI] [PubMed] [Google Scholar]
- 37.Szapary D, Xu M, Simons SS., Jr Induction properties of a transiently transfected glucocorticoid-responsive gene vary with glucocorticoid receptor concentration. J Biol Chem. 1996;271:30576–30582. doi: 10.1074/jbc.271.48.30576. [DOI] [PubMed] [Google Scholar]
- 38.He Y, Szapary D, Simons SS., Jr Modulation of induction properties of glucocorticoid receptor-agonist and -antagonist complexes by coactivators involves binding to receptors but is independent of ability of coactivators to augment transactivation. J Biol Chem. 2002;277:49256–49266. doi: 10.1074/jbc.M205536200. [DOI] [PubMed] [Google Scholar]
- 39.Kaul S, Blackford JA, Jr, Cho S, Simons SS., Jr Ubc9 is a novel modulator of the induction properties of glucocorticoid receptors. J Biol Chem. 2002;277:12541–12549. doi: 10.1074/jbc.M112330200. [DOI] [PubMed] [Google Scholar]
- 40.Wang Q, Richter WF, Anzick SL, Meltzer PS, Simons SS., Jr Modulation of transcriptional sensitivity of mineralocorticoid and estrogen receptors. J Steroid Biochem Molec Biol. 2004;91:197–210. doi: 10.1016/j.jsbmb.2004.04.007. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Linearization of transfected plasmid concentrations. (A) Plot of normalized OD of Western blots (above endogenous TIF2) for different amounts of transfected TIF2 plasmid (but same amount of cell lysate protein) vs. OD of maximum amount of transfected TIF2 (20 ng). (B) Plot of normalized ODs of transfected TIF2 (above endogenous TIF2) vs. “linearized plasmid”. Amounts (ng) of linearized TIF2 plasmid were calculated as described in the Supplementary Material and then plotted against the normalized OD values of Fig. S1A. Error bars = S.D. (n = 2, each in duplicate).
(EPS)
Plots of 1/EC50 vs. cofactor for experiment of Fig. 4 . (A) 1/EC50 vs. sSMRT plasmid (corrected for non-linear expression) with different amounts of competing NCoR plasmid. (B) 1/EC50 vs. NCoR plasmid (corrected for non-linear expression) with different amounts of competing sSMRT plasmid.
(EPS)
Plots of EC50 and Amax vs. cofactor for experiment of Fig. 5 with endogenous IGFBP1 gene. (A) 1/EC50 vs. sSMRT plasmid (corrected for non-linear expression) with different amounts of competing TIF2 plasmid. (B) Amax/EC50 vs. TIF2 plasmid (corrected for non-linear expression) with different amounts of competing sSMRT plasmid.
(EPS)
Cartoon depicting process of restricting mechanistic scenarios of the six graphs used in analyzing the competition assays. Each circle represents one type of graph (labeled on perimeter) and is composed of different mechanistic scenarios (e.g., Scenario A for 1/EC50 vs. F1) for that graph (see Table 1). The area of overlap for all graphs (depicted by the filled space in the cartoon) represents the common, uniquely identified mechanism for that specific competition assay. This 4- to 6-fold requirement of “overlap” is a stringent test for determining the mechanistic explanation of any given competition assay.
(EPS)
Algorithms for two factor plots for factors F1 and F2.
(DOCX)
Derivation of the graphical method for analyzing the competitive action of factors and Correction for non-linear protein expression from transfected plasmids.
(DOCX)