Abstract
Dioxins are widespread environmental contaminants that induce the carcinogen-activating enzyme, cytochrome P450 1A1 (CYP1A1) through an aryl hydrocarbon receptor (AhR)-dependent mechanism. We previously demonstrated that harmine inhibits the dioxin-mediated induction of Cyp1a1 activity in murine hepatoma cells. Therefore, the aim of this study is to determine the effect of harmine and its main metabolite, harmol, on the dioxin-mediated induction of CYP1A1 in human HepG2 and murine Hepa 1c1c7 hepatoma cells. Our results showed that harmine and harmol significantly inhibited the dioxin-mediated induction of CYP1A1 at mRNA, protein, and activity levels in a concentration-dependent manner in human and murine hepatoma cells. Moreover, harmine and harmol inhibited the AhR-dependent luciferase activity and the activation and transformation of AhR using the electrophoretic mobility shift assay. In addition, harmine and harmol displaced [3H]TCDD in the competitive ligand binding assay. At posttranslational level, both harmine and harmol decreased the protein stability of CYP1A1, suggesting that posttranslational mechanism is involved. Furthermore, we demonstrated that the underlying mechanisms of the posttranslational modifications of both compounds involve ubiquitin-proteasomal pathway and direct inhibitory effects of CYP1A1 enzyme. We concluded that harmine and its metabolite, harmol, are new inhibitors of dioxin-mediated effects.
Keywords: Aryl hydrocarbon receptor, Carcinogenesis, CYP 1A1, Harmine, Harmol
1. Introduction
Exposure to environmental contaminants such as polycyclic aromatic hydrocarbons (PAH) and halogenated aromatic hydrocarbons (HAH), including dioxins, plays an important role in the development of several types of human cancers. PAH and HAH are wide spread environmental contaminants that present in cigarette smoke, coal tar, automobile exhaust and charbroiled food. The carcinogenic effects of several PAH and HAH are initiated after their binding and activation of the aryl hydrocarbon receptor (AhR) (Mandal, 2005).
AhR is a ligand-activated transcription factor that is found inactive in the cytoplasm aggregated with two 90-KDa heat-shock proteins (HSP90), the co-chaperone p23 and a 43-KDa protein termed hepatitis B virus X-associated protein 2 (XAP2). Upon ligand binding, AhR gets activated and translocated to the nucleus where it heterodimerizes with another protein called AhR nuclear translocator (ARNT). This complex then binds to its DNA consensus sequence called xenobiotic responsive element (XRE) that is found in the upstream of the CYP1A1 and other AhR-responsive genes and hence stimulates transcription of these genes (Denison and Nagy, 2003).
CYP1A1 is a phase I xenobiotic metabolizing enzyme that bioactivates several PAH and other hydrophobic environmental procarcinogens into their ultimate carcinogenic forms (Shimada and Fujii-Kuriyama, 2004). CYP1A1 metabolizes PAH into reactive intermediates that form DNA adducts, which lead to mutagenesis and carcinogenesis (Shimada and Fujii-Kuriyama, 2004). Several lines of evidence demonstrate a strong correlation between the activity of CYP1A1 and the increased risk of different human cancers such as lung, colon and rectal cancers (Shah et al., 2009; Slattery et al., 2004). Therefore, the level of CYP1A1 is considered as a useful biomarker for the exposure to several carcinogens. Furthermore, the inhibition of AhR activity and its regulated gene, CYP1A1 could result in the prevention of the toxic effects caused by the AhR ligands, including carcinogenicity (Puppala et al., 2008).
β-carbolines are a large group of natural and synthetic indole alkaloids that are widely distributed in nature, including various foods, plants, marine creatures, insects, mammalians, as well as human tissues (Cao et al., 2007). β-carbolines attracted considerable attention as they possess diverse pharmacological activities such as sedative, hypnotic, anxiolytic, anticonvulsant, antitumor, antithrombotic, antiparasitic, antimicrobial, as well as antiviral activities (Cao et al., 2007).
Harmine, 7-methoxy-1-methyl-9H-pyrido[3,4-b]indole and harmol, 1-methyl-9H-pyrido[3,4-b]indole-7-ol (Fig.1), are β-carboline compounds that naturally found in several medicinal plants including Peganum harmala (Zygophyllaceae) and Banisteriopsis Caapi (Malpighiaceae) (Herraiz et al., 2010; Samoylenko et al., 2010). Harmine possesses several pharmacological activities such as antiplatelet aggregating, antimicrobial, antioxidant and antiprotozoal activities (Arshad et al., 2008; Di Giorgio et al., 2004; Im et al., 2009; Moura et al., 2007). Harmine can interact with several enzymes and neurotransmittors including topoisomerase I, 5-HT, monoamine oxidase-A, and cycline dependent kinases (Cao et al., 2005b; Cao et al., 2007; Herraiz et al., 2010; Song et al., 2004). Moreover, harmine is highly cytotoxic to several human tumor cell lines and showed promising antitumor effect for mice bearing tumor cells (Cao et al., 2005a). We previously demonstrated that Peganum harmala extract and its main active ingredient, harmine, inhibit the dioxin-mediated induction of Cyp1a1 at the catalytic activity level. Therefore, the aim of this study is to determine the effect of harmine and its main metabolite, harmol, on dioxin-mediated induction of CYP1A1 in human hepatoma HepG2 cells and to investigate the molecular mechanisms involved..
Figure 1.

Chemical structure of harmine (7-methoxy-1-methyl-9H-pyrido[3,4-b]indole), and harmol (1-methyl-9H-pyrido[3,4-b]indole-7-ol).
2. Material and methods
2.1. Chemicals and reagents
Cycloheximide (CHX), 3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyl tetrazolium bromide (MTT), 7-ethoxyresorufin (7ER), fluorescamine, harmine hydrochloride (>98% pure), 3-methylcholanthrene (3MC), β-naphthoflavone (βNF), and rabbit anti-goat IgG secondary antibody were purchased from Sigma-Aldrich (St. Louis, MO). 2,3,7,8-Tetrachlorodibenzo-p-dioxin (TCDD), >99% pure, was obtained from Cambridge Isotope Laboratories (Woburn, MA). TRIzol and Lipofectamine 2000 reagents were purchased from Invitrogen (Carlsbad, CA). Primary anti-mouse/human CYP1A1 antibody and primary goat anti-mouse/human glyceraldehyde-3-phosphate dehydrogenase (GAPDH) were purchased from Santa Cruz (Santa Cruz, CA). Goat anti-mouse/human IgG secondary antibody was obtained from R&D systems (Minneapolis, MN). High-Capacity cDNA Reverse Transcription Kit and SYBR® Green PCR Master Mix were purchased from Applied Biosystems (Foster City, CA). Harmol hydrochloride (99% pure) was obtained from MP biomedicals (Solon, OH). Carbobenzoxy-l-leucyl-l-leucyl-leucinal (MG-132) and actinomycin-D (Act-D) were purchased from Calbiochem (San Diego, CA). Chemiluminescence Western blotting detection reagents were obtained from GE Healthcare Life Sciences (Piscataway, NJ). [γ-32P]-ATP was supplied by Perkin Elmer (Boston, MA). 2,3,7,8-Tetrachlorodibenzofuran (TCDF) and [3H]-TCDD (13 Ci/mmole) were obtained from Dr. Stephen Safe (Texas A&M University). Dual-Luciferase Reporter Assay System was obtained from Promega Corporation (Madison, WI). All other chemicals were purchased from Fisher Scientific (Toronto, ON).
2.2. Animals and ethics
All experimental procedures involving animals were approved by the University of Alberta Health Sciences Animal Policy and Welfare Committee. Male Hartley guinea pigs weighing 250–300 g and male C57BL/6 mice weighing 20-25 g were obtained from Charles River Canada (St. Constant, QC, Canada). All animals were exposed to 12 h light/dark cycles and were allowed free access to food and water.
2.3. Cell culture
Human hepatoma HepG2 and murine hepatoma Hepa 1c1c7 cell lines were purchased from American Type Culture Collection (Manassas, VA). Both cells were maintained in Dulbecco’s modified Eagle’s medium supplemented with heat-inactivated fetal bovine serum (10%, v/v), L-glutamine (2 mM), penicillin (100 IU/mL), and streptomycin (100 μg/mL). Cells were grown in 75-cm2 tissue culture flasks at 37°C in a 5% CO2 humidified incubator.
2.4. Chemical treatments
Cells were treated in serum-free medium with TCDD in presence of various concentrations of harmine or harmol as specified under each experiment. Harmine, harmol, TCDD, MG-132, and Act-D were dissolved in dimethyl sulfoxide (DMSO) whereas CHX was dissolved in sterile distilled water. In all treatments, the DMSO concentration did not exceed 0.05% (v/v).
2.5. Determination of cell viability
The effect of harmine and harmol on HepG2 and Hepa 1c1c7 cell viability was determined by measuring the capacity of reducing enzymes present in viable cells to convert MTT to formazan crystals as described previously (Mosmann, 1983).
2.6. RNA isolation and Real-time polymerase chain reaction (Real-time PCR)
HepG2 cells and Hepa 1c1c7 were pre-incubated with increasing concentrations of harmine or harmol (0.5-12.50 μM) for 30 min before the addition of TCDD (1 nM) for 6 h. Thereafter, the total RNA was isolated using TRIzol reagent, according to the manufacturer’s instructions (Invitrogen) as described previously (El Gendy and El-Kadi, 2009). Primers used in the current study were chosen from previous study (Anwar-Mohamed and El-Kadi, 2009); human CYP1A1: forward primer 5′-CGG CCC CGG CTC TCT-3′, reverse primer 5′-CGG AAG GTC TCC AGG ATG AA-3′, and human β-actin: forward primer 5′-CTG GCA CCC AGC ACA ATG-3′, reverse primer 5′-GCC GAT CCA CAC GGA GTA CT-3′, mouse Cyp1a1: forward primer 5′-GGT TAA CCA TGA CCG GGA ACT-3′, reverse primer 5′-TGC CCA AAC CAA AGA GAG TGA-3′, and mouse β-actin: forward primer 5′-TAT TGG CAA CGA GCG GTT CC-3′, reverse primer 5′-GGC ATA GAG GTC TTT ACG GAT GTC-3′. The primers were purchased from Integrated DNA technologies (IDT, Coralville, IA). Real-time PCR reactions were performed on an ABI 7500 instrument (Applied Biosystems), using SYBR® Green PCR Master Mix as described previously (El Gendy and El-Kadi, 2009).
2.7. Protein extraction and Western blot analysis
HepG2 cells and Hepa 1c1c7 were treated with increasing concentrations of harmine or harmol (0.5-12.5 μM) for 30 min before the addition of TCDD (1 nM) for further 24 h. Thereafter, cells were collected and the cell lysate was obtained using lysis buffer. Proteins (50 μg for HepG2, and 25 μg for Hepa 1c1c7) in sample buffer were separated using 10% SDS-PAGE and electrophoretically transferred to a nitrocellulose membrane. The bands were visualized with the enhanced chemiluminescence method according to the manufacturer’s instructions (GE Healthcare, Piscataway, NJ). The intensity of CYP1A1 protein bands was quantified relative to the signals obtained for GAPDH protein, using a densitometer (TBX, Tobias associates, Inc., PA).
2.8. Determination of CYP1A1 enzymatic activity
CYP1A1-dependent 7-ethoxyresorufin O-deethylase (EROD) activity was performed on intact, living cells using 7ER as a substrate as described previously (Sinal and Bend, 1997). The CYP1A1 enzymatic activity was normalized to cellular protein content using a modified fluorescence method (Lorenzen and Kennedy, 1993).
2.9. Transient transfection and luciferase assay
HepG2 cells (3×104 cells per well) were plated onto 12-well cell culture plates. Each well was co-transfected with 1.5 μg of XRE-driven luciferase reporter plasmid pGudLuc 6.1 and 0.1 μg of the renilla luciferase pRL-CMV vector, used for normalization of transfection efficiency. The pRL-CMV vector was obtained from Promega Corporation (Madison,WI). Transfection procedure was carried out using Lipofectamine 2000 reagent according to the manufacturer’s instructions (Invitrogen) and the luciferase assay was performed according to the manufacturer’s instructions (Promega). The luciferase activity was reported as relative light unit (RLU) of firefly luciferase to renilla luciferase (Fluc/Rluc).
2.10. Electrophoretic mobility shift assay (EMSA)
Guinea pig hepatic cytosolic extracts (2 mg) were incubated with harmine or harmol (250 μM) for 30 min before the addition of TCDD (20 nM) for 2 h. The ability of harmine or harmol to affect the transformation and DNA binding of TCDD-activated AhR was tested using a synthetic pair of post-labeled oligonucleotides for XRE binding site as described previously (Denison et al., 2002).
2.11. Competitive ligand binding assay
Hydroxyapatite (HAP) assay was performed to determine the ligand binding ability of harmine and harmol as described previously (Denison et al., 1986). Briefly, untreated guinea pig and C57BL/6 mouse hepatic cytosols were diluted to 2 mg/mL in MEDG buffer (3-(N-morpholino) propanesulfonic acid (25 mM), pH 7.5, ethylenediaminetetraacetic acid (1 mM), dithiotreitol (1 mM), and glycerol (10%, v/v). Several aliquots of guinea pig cytosols (200 μL) were incubated at room temperature for 1 h with [3H]TCDD (2 nM) alone (total binding), [3H]TCDD (2 nM) and TCDF (200 nM, 100-fold excess of competitor, nonspecific binding) or [3H]TCDD (2 nM) in the presence of increasing concentrations of harmine or harmol (1 μM and 25 μM). All chemicals were dissolved in DMSO, in which DMSO content in reactions was adjusted to 2% (v/v) where necessary. Thereafter, hydroxyapatite suspension (250 μL) was added to the different reaction mixtures and incubated for an additional 30 min with gentle vortexing every 10 min. The reactions were washed three times with 1 mL of MEGT buffer (3-(N-morpholino) propanesulfonic acid (25 mM), pH 7.5, ethylenediaminetetraacetic acid (1 mM), glycerol (10%, v/v), and Tween 80 (0.5%, v/v). The HAP pellets were transferred to 4 mL scintillation vials, scintillation cocktail was added, and reactions were counted in a scintillation counter.
2.12. CYP1A1 mRNA stability
The posttranscriptional effects of harmine and harmol were tested using the Act-D chase assay. HepG2 cells were pre-treated with TCDD (1 nM) for 6 h. Thereafter, cells were washed and incubated with Act-D (5 μg/mL), to inhibit further RNA synthesis, immediately before treatment with harmine (2.5 μM) or harmol (2.5 μM). Total RNA was extracted at 0, 1, 3, 6, and 12 h after incubation with harmine or harmol. Thereafter, real-time PCR was carried out as described previously (El Gendy and El-Kadi, 2009). The mRNA half-life values were determined from semilog plots of mRNA amounts, expressed as percentage of treatment at t = 0, versus time.
2.13. CYP1A1 protein stability
The posttranslational effects of harmine and harmol were investigated using the CHX chase assay. HepG2 cells were pre-treated with TCDD (1 nM) for 24 h. Thereafter, cells were washed three times with PBS and incubated with CHX (10 μg/mL), to inhibit further protein synthesis, immediately before treatment with harmine (2.5 μM) or harmol (2.5 μM). Cell homogenates were extracted at 0, 1, 3, 6, and 12 h after incubation with harmine or harmol. CYP1A1 protein was measured by Western blotting as described previously (El Gendy and El-Kadi, 2010). The protein half-life values were determined from semilog plots of integrated densities versus time.
2.14. Direct inhibitory study
To test the direct inhibitory effect of harmine and harmol on CYP1A1 enzyme, a method similar to EROD assay was performed, with slight modifications. Briefly, HepG2 cells were incubated with TCDD (1nM) for 24 h. Thereafter, media were removed from the cells, washed three times with PBS, and increasing concentrations (0.5-12.5 μM) of harmine and harmol in assay buffer (Tris (0.05 M), NaCl (0.1 M), pH 7.8) were added to the cells for 15 min prior to the addition of 7ER (2 μM final concentration) as a substrate for EROD measurement. The remaining CYP1A1 enzymatic activity was normalized to cellular protein content using a modified fluorescence method (Lorenzen and Kennedy, 1993).
2.15. Statistical analysis
All results are presented as mean ± S.E.M., and statistical differences between treatment groups were determined using one way ANOVA followed by Student-Newman-Keuls post hoc test using SigmaStat 3.5 program for Windows, Systat Software Inc. (San Jose, CA).
3. Results
3.1. Effect of harmine and harmol on cell viability
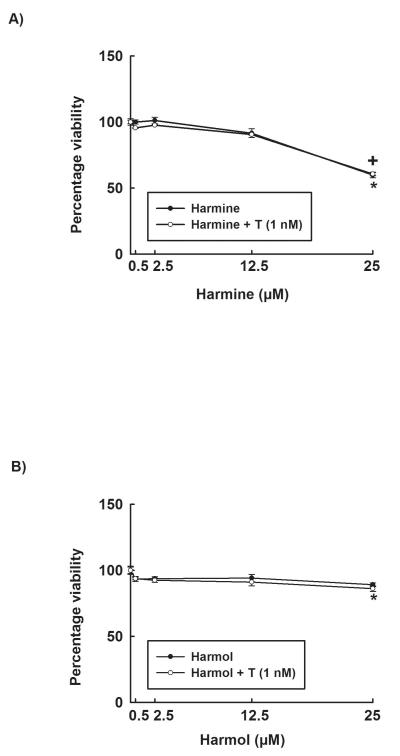
To determine the nontoxic concentrations, harmine and harmol were tested for their potential cytotoxicity in HepG2 and Hepa 1c1c7 cells. Our results showed that incubation of Hepa 1c1c7 cells with increasing concentrations (0.5-25 μM) of harmine or harmol for 24 h did not significantly affect the cell viability up to 12.50 μM (data not shown). Similarly, harmine and harmol did not affect HepG2 cell viability up to 12.50 μM (Fig. 2A&2B). However, the highest concentration (25 μM) tested of harmine, significantly decreased HepG2 cell viability to 60% in presence or absence of TCDD (Fig. 2A). Moreover, harmol (25 μM) significantly decreased HepG2 cell viability in presence of TCDD to 85% (Fig. 2B). Therefore, we have chosen the concentrations between 0.5-12.5 μM for both harmine and harmol as safe concentrations for the following experiments.
Figure 2. Effect of harmine and harmol on HepG2 cell viability.
The effect of increasing concentrations of harmine (A) and harmol (B) on HepG2 cell viability was tested using the MTT assay. Data are expressed as percent of control, which is set at 100%, ± S.E.M. (n = 5). (+) P < 0.05 compared with control(C), (*) P < 0.05 compared with TCDD (T).
3.2. Effect of harmine and harmol on dioxin-mediated induction of CYP1A1 mRNA, protein, and activity levels in HepG2 cells
To investigate whether harmine alters the CYP1A1 mRNA level, HepG2 cells were pre-incubated with increasing concentrations of harmine (0.5-12.50 μM) for 30 min before the addition of TCDD for 6 h. Thereafter, CYP1A1 mRNA was quantified using real-time PCR. Our data showed that harmine significantly decreased the TCDD-mediated induction of CYP1A1 mRNA in a concentration-dependent manner by 27%, 64%, and 88% with harmine concentrations of 0.5, 2.5, and 12.5 μM, respectively (Fig. 3A).
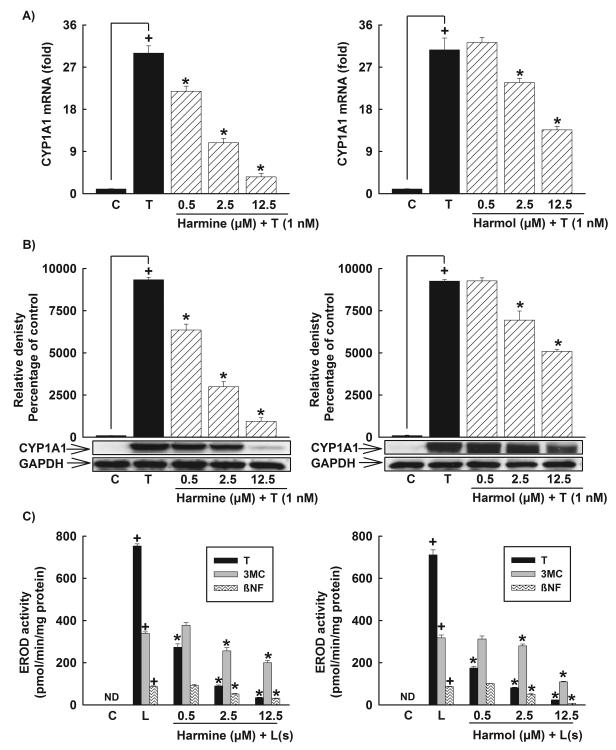
Figure 3. Effect of harmine and harmol on CYP1A1 mRNA, protein, and catalytic activity in HepG2 cells.
Cells were incubated with increasing concentrations of harmine or harmol (0.5-12.5 μM) 30 min before the addition of TCDD (1nM) for an additional 6 h for mRNA or 24 h for protein and catalytic activity. A, The amount of CYP1A1 mRNA was quantified using real-time PCR and normalized to β-actin housekeeping gene. Values represent the mean of fold change ± S.E.M. (n=4). B, Protein was separated on a 10% SDS-PAGE and CYP1A1 protein was determined using the enhanced chemiluminescence method. The intensity of bands was normalized to GAPDH signals, which was used as loading control. One of three representative experiments is shown. C, CYP1A1 activity was determined using CYP1A1-dependent EROD assay. Values represent mean activity ± S.E.M. (n = 8). (+) P < 0.05 compared with Control (C), (*) P < 0.05 compared with TCDD (T), (L); ligand.
Western blot analysis was employed to determine the effect of harmine on the expression of CYP1A1 at the protein level. Consistent with the mRNA results, harmine showed a significant concentration-dependent decrease in TCDD-mediated induction of CYP1A1 protein by 32%, 68%, and 90% with harmine concentrations of 0.5, 2.5, and 12.5 μM, respectively (Fig. 3B). To determine whether harmine has a similar effect on the CYP1A1 catalytic activity, HepG2 cells were incubated with increasing concentrations of harmine (0.5-12.5 μM) 30 min before the addition of TCDD (1 nM) for 24 h. Thereafter, CYP1A1 catalytic activity was determined using EROD assay. Our results showed that harmine significantly decreased the TCDD-mediated induction of the CYP1A1 catalytic activity by 64%, 88%, and 95% with harmine concentrations of 0.5, 2.5, and 12.5 μM, respectively (Fig. 3C). To determine whether the effect of harmine is AhR ligand specific, we tested the effect of harmine on two other AhR ligands, namely, 3MC (0.25 μM) and βNF (10 μM). Our results showed that harmine significantly decreased the induction of CYP1A1 by 41% and 66% in the presence of βNF and 25% and 41% in the presence of 3MC, with harmine concentrations of 2.5 and 12.5 μM, respectively (Fig. 3C). The order of inhibition of harmine against different AhR ligands was TCDD > βNF > 3MC (Fig. 3C).
To investigate whether the effect of harmine is not due to its active metabolite, we examined the effect of its main metabolite, harmol, on the CYP1A1 mRNA level in human HepG2 cells. Our results showed that harmol significantly decreased the TCDD-mediated induction of CYP1A1 mRNA in HepG2 cells in a concentration-dependent manner by 23% and 56% with harmol concentrations of 2.5 and 12.5 μM, respectively (Fig. 3A).
Similarly, harmol significantly decreased the TCDD-mediated induction of CYP1A1 at the protein level by 25% and 45% with harmol concentrations of 2.5 and 12.5 μM, respectively (Fig. 3B). At the CYP1A1 catalytic activity level, harmol decreased all AhR ligand-mediated induction of CYP1A1 catalytic activity in HepG2 cells. Harmol showed a higher activity against TCDD, where it significantly decreased the induction level of CYP1A1 catalytic activity by 75%, 89%, and 97% with harmol concentrations of 0.5, 2.5, and 12.5 μM, respectively (Fig. 3C). Moreover, harmol significantly decreased the induction of CYP1A1 catalytic activity by 41% and 91% in the presence of βNF and by 12% and 66% in the presence of 3MC, with harmol concentrations of 2.5 and 12.5 μM, respectively (Fig. 3C). The order of inhibition of harmol against different AhR ligands was similar to harmine; TCDD > βNF > 3MC (Fig. 3C). Furthermore, neither harmine nor harmol alone significantly affected CYP1A1 at the catalytic activity level in HepG2 cells (data not shown).
3.3. Effect of harmine and harmol on dioxin-mediated induction of Cyp1a1 mRNA, protein, and activity levels in Hepa 1c1c7 cells
In an attempt to explore whether the effect of harmine and harmol is species specific, we examined their effect on TCDD-mediated induction of Cyp1a1 using murine hepatoma cells, Hepa 1c1c7. Our results showed that harmine decreased the TCDD-mediated induction of Cyp1a1 mRNA by 25% and 29% with concentrations of 2.5 and 12.5 μM, respectively (Fig. 4A). Furthermore, harmol showed a more pronounced effect than harmine in Hepa 1c1c7 where it decreased the TCDD-mediated induction of Cyp1a1 mRNA by 53%, 63%, and 71% with harmol concentrations of 0.5, 2.5, and 12.5 μM, respectively (Fig. 4A).
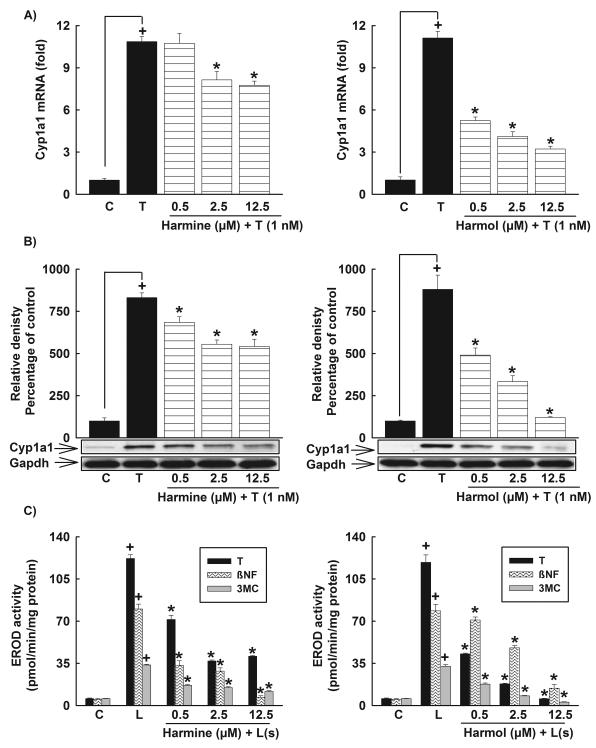
Figure 4. Effect of harmine and harmol on Cyp1a1 mRNA, protein, and catalytic activity in Hepa 1c1c7 cells.
Cells were incubated with increasing concentrations of harmine or harmol (0.5-12.5 μM) 30 min before the addition of TCDD (1nM) for an additional 6 h for mRNA or 24 h for protein and catalytic activity. A, The amount of Cyp1a1 mRNA was quantified using real-time PCR and normalized to β-actin housekeeping gene. Values represent the mean of fold change ± S.E.M. (n=4). B, Protein was separated on a 10% SDS-PAGE and Cyp1a1 protein was determined using the enhanced chemiluminescence method. The intensity of bands was normalized to Gapdh signals, which was used as loading control. One of three representative experiments is shown. C, Cyp1a1 activity was determined using Cyp1a1-dependent EROD assay. Values represent mean activity ± S.E.M. (n = 8). (+) P < 0.05 compared with Control (C), (*) P < 0.05 compared with TCDD (T), (L); ligand.
At the protein level, both compounds significantly inhibited the TCDD-mediated induction of Cyp1a1 in a concentration-dependent manner. Harmine significantly decreased the TCDD-mediated induction of Cyp1a1 protein by 18%, 33%, and 35% with harmine concentrations of 0.5, 2.5, and 12.5 μM, respectively (Fig. 4B). Moreover, harmol showed a more pronounced effect than harmine, where it decreased the TCDD-mediated induction of Cyp1a1 protein by 44%, 62%, and 86% with 0.5, 2.5, and 12.5 μM, respectively (Fig. 4B).
At the catalytic activity level, both harmine and harmol significantly decreased the Cyp1a1 catalytic activity induced by all tested AhR ligands in a concentration-dependent manner. According to the percentage inhibition of the induced Cyp1a1 catalytic activity level, the order of inhibition of harmine against different AhR ligands was βNF > TCDD > 3MC (Fig. 4C). In contrast to harmine, the order of inhibition of harmol against different AhR ligands was TCDD > 3MC > βNF (Fig. 4C). Similar to HepG2 cells, neither harmine nor harmol alone significantly affected Cyp1a1 at the catalytic activity level in Hepa 1c1c7 cells (data not shown).
3.4. Transcriptional effect of harmine and harmol on CYP1A1 gene
In an attempt to explore the effect of harmine and harmol on the AhR-dependent transcriptional activation, HepG2 cells were transiently co-transfected with the XRE-driven luciferase reporter gene and renilla luciferase vector, which is used for normalization of transfection efficiency. Our results showed that TCDD alone significantly induced the luciferase activity by 1300% as compared with the control (Fig. 5A). On the other hand, harmine and harmol significantly decreased the TCDD-induced luciferase activity by 42% and 22%, respectively (Fig. 5A).
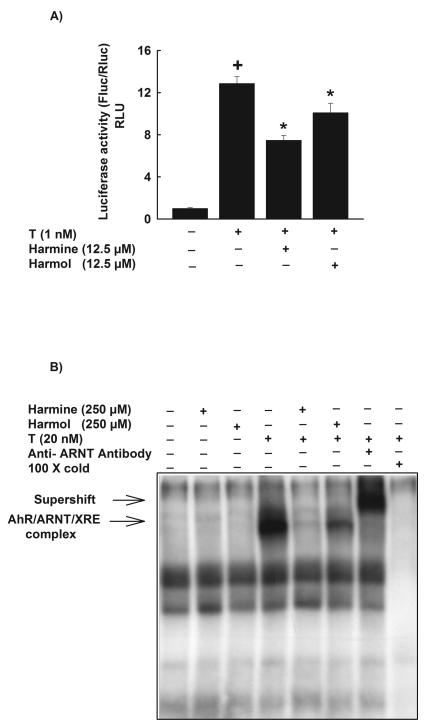
Figure 5. Effect of harmine and harmol on XRE-luciferase activity and AhR activation using electrophoretic mobility shift assay (EMSA).
A, HepG2 cells were transiently co-transfected with XRE-luciferase reporter plasmid pGudLuc 6.1. and renilla luciferase control plasmid pRL-CMV. Cells were treated with DMSO, harmine (12.5 μM) or harmol (12.5 μM) 30 min before the addition of TCDD (1nM) for an additional 24 h. Cells were lysed and luciferase activity is reported as relative light unit (RLU) of firefly luciferase to renilla luciferase (Fluc/Rluc) (mean ± S.E.M., n = 4). (+) P < 0.05 compared with Control (C), (*) P < 0.05 compared with TCDD (T). B, In vitro AhR activity was measured by EMSA using guinea pig hepatic cytosolic extracts. Cytosolic extracts were incubated with DMSO, harmine (250 μM) or harmol (250 μM) for 30 min before the addition of TCDD (20 nM) for 2 h. The mixtures were tested for binding activity to a [γ-32P]-labeled XRE consensus oligonucleotide for an additional 15 min. The products of this binding were separated on a 4% polyacrylamide gel. The specificity of the binding was confirmed by competition assays using anti-ARNT antibody or a 100 fold molar excess of unlabeled XRE. AhR-ARNT-XRE complex formed on the gel was visualized by autoradiography. One representative of three experiments is shown.
In order to test the ability of harmine and harmol to directly interfere with AhR and subsequent DNA binding to XRE, EMSA was performed using untreated guinea pig hepatic cytosol incubated either with vehicle (DMSO), harmine (250 μM) or harmol (250 μM) in the absence and presence of TCDD (20 nM) for 2 h. Figure 5B shows that both harmine and harmol (250 μM) alone did not alter the AhR activity, while TCDD (20 nM) alone induced the AhR activity and the formation of AhR/ARNT/XRE complex. On the other hand, pre-incubation of guinea pig cytosolic extracts with harmine or harmol significantly inhibited the TCDD-mediated activation of AhR and the formation of AhR/ARNT/XRE complex (Fig. 5B). The specificity of the binding was confirmed by the competition assays using anti-ARNT antibody or a 100 fold molar excess of unlabeled XRE (Fig. 5B).
To determine whether harmine or harmol are direct ligands for the AhR, a ligand competition binding assay using hydroxyapatite was performed (Fig. 6). In this assay, we used untreated guinea pig and mouse hepatic cytosols to study the binding ability of harmine and harmol to AhR from two different species. Moreover, the total binding is the overall binding of [3H]-TCDD to cytosolic AhR protein. However, to account for the non-specific binding that happens not through the AhR or not through the ligand-binding center of the AhR, reactions were conducted in the presence of 100-fold excess of the competitor. We have chosen TCDF rather than TCDD because of its higher solubility as TCDD would not be soluble at 200 nM. Thus, the specific binding of [3H]-TCDD to the AhR was calculated by subtracting the non-specific binding from the total binding. Our results demostrated that harmine at concentrations of 1 μM and 25 μM was able to significantly displace [3H]-TCDD (2 nM) by 19% and 74%, and by 20% and 63% using guinea pig and mouse cytosols, respectively (Fig. 6A&6B). To a lower degree, harmol displaced [3H]-TCDD (2 nM) by 6% and 19%, and by 9% and 25% using guinea pig and mouse cytosols, respectively (Fig. 6A&6B). The effect was significant for all tested concentrations of harmol except for the 1 μM concentration in the guinea pig cytosol treatment (Fig. 6A).
Figure 6. AhR ligand binding ability of harmine and harmol.
A, Guinea pig hepatic cytosol (2 mg/mL) or B, C57BL/6 mouse hepatic cytosols (2 mg/mL) were incubated with [3H]-TCDD (2 nM) alone (total binding), [3H]-TCDD (2 nM) and TCDF (200 nM, 100-fold excess of competitor, nonspecific binding), or [3H]-TCDD (2 nM) in the presence of increasing concentrations of harmine or harmol (1 and 25 μM) and the samples analyzed by the hydroxyapatite assay as described under Material and methods. Values were adjusted for nonspecific binding and expressed as % specific binding relative to the absence of a competitor ligand. Values are presented as the mean ± S.E.M. (n = 9). (*) P < 0.05 compared with [3H]-TCDD.
3.5. Posttranscriptional effect of harmine and harmol
The level of gene expression is regulated by the transcription rate and the elimination rate through processing or degradation. Therefore, we tested the effect of harmine and harmol on the stability of CYP1A1 mRNA transcripts in HepG2 cells, using the Act-D chase experiment. If the effect of harmine or harmol on CYP1A1 involves posttranscriptional effects through destabilization of CYP1A1 mRNA, a decrease in CYP1A1 mRNA half-life would be expected. Our results showed that TCDD-induced CYP1A1 mRNA degraded with a half-life of 4.9 ± 0.4 h (Fig. 6A). Moreover, treatment with harmine or harmol did not significantly alter CYP1A1 mRNA half-life which was 5.4 ± 0.26 h and 4.9 ± 0.04 h, respectively (Fig. 7A).
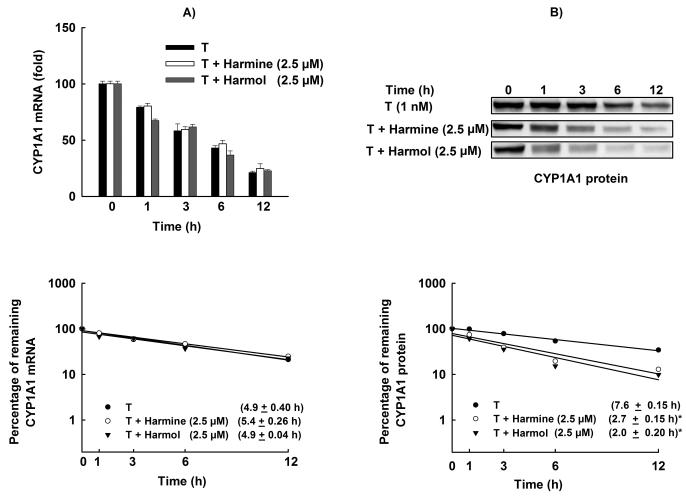
Figure 7. Effect of harmine and harmol on CYP1A1 mRNA and protein stability.
HepG2 cells were treated with TCDD (1 nM) for 6 h for mRNA stability and 24 h for protein stability assays. Thereafter, the cells were washed and incubated with fresh media containing harmine (2.5 μM) or harmol (2.5 μM) plus Act-D (5 μg/mL, the mRNA synthesis inhibitor) or CHX (10 μg/mL, the protein translation inhibitor). A, Total RNA was extracted at 0, 1, 3, 6, and 12 h after incubation with harmine or harmol and subjected to real-time PCR. B, Protein was separated on a 10% SDS-PAGE and CYP1A1 protein was determined using the enhanced chemiluminescence method. The intensities of CYP1A1 protein bands were normalized to GAPDH signals, which were used as loading controls (data not shown). mRNA and protein decay curves were analyzed individually, and the half-life was estimated from the slope of a straight line fitted by linear regression analysis (r2 ≥ 0.85) to a semilog plot, expressed as a percent of treatment at time = 0 h (maximum, 100%) level, versus time. The half-lives obtained from three independent experiments were then used to calculate the mean half-life (mean ± S.E.M., n = 3). (*) P < 0.05 compared with TCDD (T).
3.6. Posttranslational effect of harmine and harmol
The effect of harmine and harmol on the activity level was much higher than that obtained with the protein level which raises the possibility of posttranslational modifications. Therefore, we tested the effect of harmine and harmol on CYP1A1 protein stability in HepG2 cells using CHX chase experiment. Figure 7B shows that CYP1A1 protein induced by TCDD degraded with a half-life of 7.6 ± 0.15 h. Furthermore, harmine and harmol significantly reduced the stability of CYP1A1 protein which degraded with half-lives of 2.7 ± 0.15 h and 2.0 ± 0.20 h, respectively (Fig. 7B).
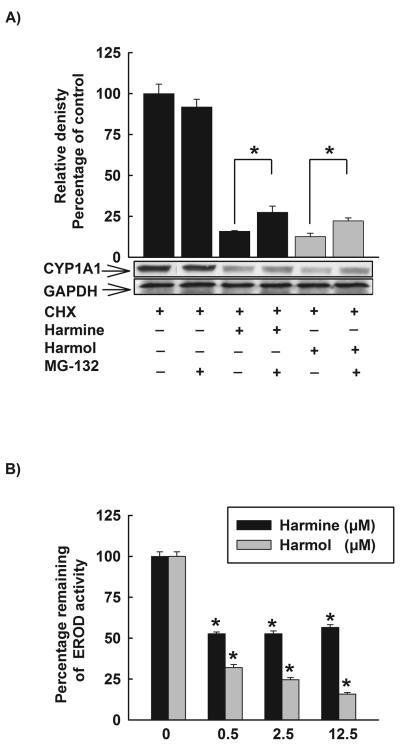
In order to elucidate the underlying mechanisms of the posttranslational modifications of harmine and harmol, we tested the role of ubiquitin-proteasomal pathway. HepG2 cells were treated with TCDD (1 nM) for 24 h, thereafter, cells were washed three times with PBS and incubated with fresh media containing CHX (10 μg/mL) alone, CHX (10 μg/mL) and harmine (2.5 μM) or CHX (10 μg/mL) and harmol (2.5 μM) in the absence and presence of the proteasomal inhibitor, MG-132 (0.5 μM). Total protein was extracted after 6 h and CYP1A1 protein was determined using Western blot analysis. Our data showed that MG-132 alone did not affect the level of CYP1A1 protein (Fig. 8A). On the other hand, MG-132 significantly induced the level of CYP1A1 protein for CHX and harmine or CHX and harmol treated cells (Fig. 8A).
Figure 8. Posttranslational modifications of CYP1A1 by harmine and harmol.
A, Effect of proteasomal inhibitor, MG-132, on the reduced CYP1A1 protein stability by harmine and harmol. HepG2 cells were treated with TCDD (1 nM) for 24 h. Thereafter, cells were washed three times with PBS and incubated with fresh media containing CHX alone (10 μg/mL, the protein translation inhibitor), CHX (10 μg/mL) and harmine (2.5 μM) or CHX (10 μg/mL) and harmol (2.5 μM) in the absence and presence of the proteasomal inhibitor, MG-132 (0.5 μM). After 6 h incubation, total protein was extracted and CYP1A1 protein was determined using Western blot analysis. The intensities of CYP1A1 protein bands were normalized to GAPDH signals, which were used as loading controls. Values represent mean of relative densities and expressed as percentage of control (untreated cells) ± S.E.M. One of three representative experiments is shown. (*) P < 0.05 compared with the relevant treatment. B, The direct inhibitory effects of harmine and harmol on CYP1A1 enzyme. HepG2 cells were pre-treated with TCDD (1nM) for 24 h, thereafter media were removed, washed three times with PBS, and increasing concentrations of harmine and harmol (0.5-12.5 μM) in assay buffer were added for 15 min prior to the addition of 7ER (2 μM final concentration) for the EROD measurement. Results are expressed as percentage of remaining EROD activity (mean ± S.E.M, n = 8). (*) P < 0.05 compared with control (untreated cells).
In addition, we tested the direct inhibitory effect of harmine and harmol on CYP1A1 enzyme. HepG2 cells were treated for 24 h with TCDD (1 nM), thereafter, the cells were washed twice with PBS and increasing concentrations of harmine or harmol in assay buffer were further incubated for 15 min before the addition of the substrate (7ER, 2 μM final concentration). The remaining CYP1A1 activity was measured using EROD assay. Figure 8B shows that harmine and harmol possess direct inhibitory effects on CYP1A1 enzyme, where harmine significantly inhibited the CYP1A1 activity by approximately 50% with all tested concentrations (0.5, 2.5, and 12.5 μM) (Fig. 8B). Moreover, harmol significantly inhibited the CYP1A1 activity in a concentration-dependent manner by 68, 75, and 84% with harmol concentrations of 0.5, 2.5, and 12.5 μM, respectively (Fig. 8B).
4. Discussion
The current study provides the first mechanistic evidence that harmine and its main metabolite, harmol, significantly inhibit the induction of CYP1A1 by dioxin at the transcriptional and posttranslational levels. AhR activation can result in several biological and toxic effects that depend on the type of AhR ligand (Bradshaw and Bell, 2009). The effect of AhR ligands on cell cycle, inflammation, and cancer cell proliferation raised the therapeutic potential for activators/inhibitors of the AhR signaling pathway (Bradshaw and Bell, 2009; Zhao et al., 2010). Dioxins are metabolically stable AhR ligands that produce a spectrum of TCDD-like AhR-dependent toxicity and carcinogenicity (Mandal, 2005). Accordingly, AhR has been used as a target for screening of new chemopreventative agents (Puppala et al., 2008). Numerous AhR antagonists have shown promising results against several carcinogen-activating agents. It has been previously reported that the genotoxicity associated with benzo(a)pyrene in mice was inhibited by AhR antagonists such as 3′-methoxy-4′-nitroflavone and resveratrol (Dertinger et al., 2001; Revel et al., 2003). However, several AhR antagonists lack specificity and can act as partial agonists, therefore, the search for new AhR antagonist is still in progress (Puppala et al., 2008; Signorelli and Ghidoni, 2005; Zhou and Gasiewicz, 2003).
Harmine is metabolized in the liver and extrahepatic tissues to its main metabolite, harmol, by the cytochrome P450s, mainly CYP2D6 and CYP1A2 (Fig.1) (Yu et al., 2003). It was previously reported that harmine and harmol possess antimutagenic and antigenotoxic effects in yeast and mammalian cells, respectively (Moura et al., 2007). These effects have been correlated to the hydroxyl radical-scavenging and the antioxidant properties of harmine and harmol (Moura et al., 2007). In our attempt to search for new chemopreventative agents from natural sources, we have shown that the extract of Peganum harmala fruiting tops decreases the TCDD-mediated induction of Cyp1a1 at mRNA, protein, and enzyme activity levels in murine hepatoma Hepa 1c1c7 cell line (El Gendy et al., 2010). Furthermore, we confirmed that the underlying mechanism of Peganum harmala extract involves modulation of AhR signaling pathway. Additionally, harmine was identified as the main active ingredient of the plant extract and was responsible for the inhibition of the dioxin-mediated induction of Cyp1a1 activity in murine hepatoma Hepa 1c1c7 cells (El Gendy et al., 2010). However, the effect of harmine and harmol on dioxin-mediated induction of CYP1A1 has not been studied before in human hepatoma HepG2 cells. Therefore, we determined the effect of harmine and its metabolite, harmol, in human hepatoma HepG2 cells and compared their effect to that obtained in murine hepatoma Hepa 1c1c7 cells.
In the current study, we tested whether harmine and its main metabolite, harmol, alter the CYP1A1 enzyme in HepG2 cells. Our results showed that harmine significantly reduced the dioxin-mediated induction of CYP1A1 in a concentration-dependent manner at mRNA, protein, and activity levels. In order to test whether the effect of harmine is due to the parent compound or due to its active metabolite, harmol, we examined both compounds in HepG2 cells. Our results demonstrated that similar to harmine, harmol significantly reduced the dioxin-mediated induction of CYP1A1 in a concentration-dependent manner at mRNA, protein, and activity levels in HepG2 cells. However, the effect of harmine was more pronounced.
The fact that the response of AhR agonists/antagonists can vary between different species (Zhang et al., 2003; Zhou et al., 2003), prompted us to test the effect of harmine and harmol on murine hepatoma Hepa 1c1c7 cells. Our results showed that both harmine and harmol significantly reduced the dioxin-mediated induction of Cyp1a1 in a concentration-dependent manner at mRNA, protein, and activity levels in Hepa 1c1c7 cells. In contrast to HepG2 cells, the effect of harmol is higher than the effect of harmine in Hepa 1c1c7 cells. Several postulations have been proposed to explain the species dependence of some of the AhR antagonists across AhR regulated genes. Of these, intrinsic variation in binding affinity for AhR and the differences in the rate of uptake and metabolism of AhR antagonists are different between different species (Murray et al., 2010). In this context, the differences between the effect of harmine and harmol among different species can be attributed to the structural differences between harmine and its metabolite, harmol. Harmine structure contains methoxyl group that is altered by metabolism to hydroxyl group in its metabolite, harmol. The differences in structure of harmine and harmol might affect the uptake and the binding of dioxin to AhR and the subsequent recruitment of different coregulatory proteins among different species (Suzuki and Nohara, 2007).
Furthermore, to examine whether the effect of harmine and harmol is AhR ligand specific, we examined the effect of harmine and harmol on different AhR ligands-mediated induction of CYP1A1 activity, namely, 3MC and βNF. Our results demonstrated that both harmine and harmol significantly inhibited all AhR ligands-mediated induction of CYP1A1 activity in both human and murine hepatoma cell lines, suggesting that the effect of harmine and harmol is not AhR ligand dependent. It has been previously reported that AhR antagonists show variable inhibitory effects against different AhR ligands/agonists. In this context, it was previously reported that 6,2′,4′,-trimethoxyflavone (TMF) antagonizes the TCDD and its related halogenated aromatic hydrocarbons (HAH), as well as non-HAH effects (Murray et al., 2010). On the other hand, CH223191 was found to preferentially inhibit the ability of HAH, but not other non-HAH such as βNF on the AhR signaling pathway (Zhao et al., 2010). Most importantly, both harmine and harmol inhibited the CYP1A1 activity induced by the tested AhR ligands in a similar order in human hepatoma HepG2 cells. However, harmine and harmol showed a different order of inhibition against the tested AhR ligands in murine hepatoma Hepa 1c1c7 cells. The variation in potency of harmine and harmol against different AhR ligands could be attributed to the differences in structure between harmine and its metabolite, harmol. Harmine and harmol could act as selective AhR modulators and alter the AhR protein or the ligand binding domain on the AhR in such a way that they interact differently with the tested AhR ligands (Zhao et al., 2010).
To investigate the underlying mechanisms of harmine and harmol against dioxin-mediated induction of CYP1A1, we tested whether harmine and harmol inhibit CYP1A1 at transcriptional level. For this purpose we examined the effect of harmine and harmol on AhR-dependent luciferase reporter assay and AhR activation and transformation using EMSA. As expected, harmine and harmol significantly inhibited the TCDD-induced AhR-dependent luciferase activity. Moreover, both harmine and harmol inhibited the TCDD-mediated activation and binding of AhR to the XRE using EMSA, confirming the involvement of a transcriptional mechanism. Most importantly, the result of EMSA demonstrates that neither harmine nor harmol alone significantly induced AhR activation or transformation when incubated with guinea pig hepatic cytosols. In the same context, both harmine and harmol alone did not affect CYP1A1 at the activity level in human and murine hepatoma cells (data not shown). This result suggests that both harmine and harmol are not partial agonists for AhR at the used concentrations. To further study whether harmine and harmol are AhR ligands, a ligand competition binding assay using hydroxyapatite was performed. The assay was carried out using untreated guinea pig and mouse hepatic cytosols. Our data demonstrated that both harmine and harmol possess an AhR ligand binding affinity both in guinea pig and mouse cytosols. These data are in agreement with the results of EMSA assay and suggest that both harmine and harmol are AhR antagonists at the tested concentrations.
In addition, we investigated the role of posttranscriptional and posttranslational mechanisms using Act-D- and CHX-chase experiments, respectively. Our results demonstrated that harmine and harmol did not alter the stability of CYP1A1 mRNA in HepG2 cells. However, both compounds significantly decreased the CYP1A1 protein stability in HepG2 cells as indicated by the lower half-life of CYP1A1 protein after incubation either with harmine or harmol. We postulate that, the presence of methoxyl or hydroxyl group in harmine and harmol aromatic structures, respectively, plays a role in the stability of CYP1A1 protein by both compounds. This conclusion is substantiated by the fact that harman, an aromatic β-carboline, which lacks those functioning groups, did not alter the stability of CYP1A1 protein in HepG2 cells (El Gendy and El-Kadi, 2010).
Several mechanisms have been proposed to explain the protein degradation including but not limited to ubiquitin-proteasomal, autophagy-lysosome and calpain pathways (Taguchi et al., 2011). However, ubiquitin-proteasomal pathway possess an important role in CYP1A regulation (Pollenz, 2007; Wiseman and Vijayan, 2007). Therefore, we tested the effect of ubiquitin-proteasomal pathway in the posttranslational modifications of harmine and harmol by using the proteasomal inhibitor, MG-132. Our results showed that inhibition of ubiquitin-proteasomal pathway significantly induced the CYP1A1 protein level of harmine and harmol treated cells, implying the involvement of the ubiquitin-proteasomal pathway in the posttranslational modifications of harmine and harmol. Moreover, the direct inhibitory effect of harmine and harmol on CYP1A1 enzyme was tested. In this assay we incubated HepG2 cells with TCDD for 24 h to induce the level of CYP1A1 enzyme, thereafter, increasing concentrations of harmine and harmol were added for 15 min before the addition of the substrate (7ER). Our data showed that harmine and harmol significantly reduced the level of CYP1A1 enzyme activity as measured by EROD assay, suggesting that both compounds possess a direct inhibitory effect on CYP1A1 enzyme that participates in their posttranslational modifications.
In conclusion we demonstrated that harmine and harmol inhibit the dioxin-mediated induction of CYP1A1 at transcriptional and posttranslational levels. Furthermore, these data may represent novel mechanisms by which harmine and its main metabolite, harmol, inhibit the dioxin-mediated effects.
Acknowledgments
This work was supported by the Natural Sciences and Engineering Council of Canada (NSERC) grant RGPIN 250139 to A.O.S.E., and the National Institutes of Environmental Health Sciences research grant R01ES07685 to M.S.D. M.A.M.E. is the recipient of the Egyptian government scholarship. We are grateful to Dr. Loren Kline (University of Alberta, AB) for providing us with guinea pig livers.
Abbreviations
- Act-D
actinomycin D
- AhR
aryl hydrocarbon receptor
- βNF
β-naphthoflavone
- CHX
cycloheximide
- CYP
cytochrome P450
- EMSA
electrophoretic mobility shift assay
- 7ER
7-ethoxyresorufin
- EROD
7-ethoxyresorufin O-deethylase
- GAPDH
glyceraldehyde-3-phosphate dehydrogenase
- HAH
halogenated aromatic hydrocarbons
- 3MC
3-methylcholanthrene
- MG-132
carbobenzoxy-l-leucyl-l-leucyl-leucinal
- MTT
(3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyl tetrazolium bromide)
- PAH
Polycyclic aromatic hydrocarbon
- TCDD
2,3,7,8-tetrachlorodibenzo-p-dioxin
- TCDF
2,3,7,8-tetrachlorodibenzofuran
- XRE
xenobiotic responsive element
Footnotes
Conflict of Interest Statement The authors have declared no conflict of interests
References
- Anwar-Mohamed A, El-Kadi AO. Sulforaphane induces CYP1A1 mRNA, protein, and catalytic activity levels via an AhR-dependent pathway in murine hepatoma Hepa 1c1c7 and human HepG2 cells. Cancer Lett. 2009;275:93–101. doi: 10.1016/j.canlet.2008.10.003. [DOI] [PubMed] [Google Scholar]
- Arshad N, Zitterl-Eglseer K, Hasnain S, Hess M. Effect of Peganum harmala or its beta-carboline alkaloids on certain antibiotic resistant strains of bacteria and protozoa from poultry. Phytother Res. 2008;22:1533–1538. doi: 10.1002/ptr.2528. [DOI] [PubMed] [Google Scholar]
- Bradshaw TD, Bell DR. Relevance of the aryl hydrocarbon receptor (AhR) for clinical toxicology. Clin Toxicol (Phila) 2009;47:632–642. doi: 10.1080/15563650903140423. [DOI] [PubMed] [Google Scholar]
- Cao R, Chen H, Peng W, Ma Y, Hou X, Guan H, Liu X, Xu A. Design, synthesis and in vitro and in vivo antitumor activities of novel beta-carboline derivatives. Eur J Med Chem. 2005a;40:991–1001. doi: 10.1016/j.ejmech.2005.04.008. [DOI] [PubMed] [Google Scholar]
- Cao R, Peng W, Chen H, Ma Y, Liu X, Hou X, Guan H, Xu A. DNA binding properties of 9-substituted harmine derivatives. Biochem Biophys Res Commun. 2005b;338:1557–1563. doi: 10.1016/j.bbrc.2005.10.121. [DOI] [PubMed] [Google Scholar]
- Cao R, Peng W, Wang Z, Xu A. beta-Carboline alkaloids: biochemical and pharmacological functions. Curr Med Chem. 2007;14:479–500. doi: 10.2174/092986707779940998. [DOI] [PubMed] [Google Scholar]
- Denison MS, Harper PA, Okey AB. Ah receptor for 2,3,7,8-tetrachlorodibenzo-p-dioxin. Codistribution of unoccupied receptor with cytosolic marker enzymes during fractionation of mouse liver, rat liver and cultured Hepa-1c1 cells. Eur J Biochem. 1986;155:223–229. doi: 10.1111/j.1432-1033.1986.tb09480.x. [DOI] [PubMed] [Google Scholar]
- Denison MS, Nagy SR. Activation of the aryl hydrocarbon receptor by structurally diverse exogenous and endogenous chemicals. Annu Rev Pharmacol Toxicol. 2003;43:309–334. doi: 10.1146/annurev.pharmtox.43.100901.135828. [DOI] [PubMed] [Google Scholar]
- Denison MS, Pandini A, Nagy SR, Baldwin EP, Bonati L. Ligand binding and activation of the Ah receptor. Chem Biol Interact. 2002;141:3–24. doi: 10.1016/s0009-2797(02)00063-7. [DOI] [PubMed] [Google Scholar]
- Dertinger SD, Nazarenko DA, Silverstone AE, Gasiewicz TA. Aryl hydrocarbon receptor signaling plays a significant role in mediating benzo[a]pyrene- and cigarette smoke condensate-induced cytogenetic damage in vivo. Carcinogenesis. 2001;22:171–177. doi: 10.1093/carcin/22.1.171. [DOI] [PubMed] [Google Scholar]
- Di Giorgio C, Delmas F, Ollivier E, Elias R, Balansard G, Timon-David P. In vitro activity of the beta-carboline alkaloids harmane, harmine, and harmaline toward parasites of the species Leishmania infantum. Exp Parasitol. 2004;106:67–74. doi: 10.1016/j.exppara.2004.04.002. [DOI] [PubMed] [Google Scholar]
- El Gendy MA, El-Kadi AO. Peganum harmala L. differentially modulates cytochrome P450 gene expression in human hepatoma HepG2 cells. Drug Metab Lett. 2009;3:212–216. doi: 10.2174/187231209790218163. [DOI] [PubMed] [Google Scholar]
- El Gendy MA, El-Kadi AO. Harman induces CYP1A1 enzyme through an aryl hydrocarbon receptor mechanism. Toxicol Appl Pharmacol. 2010;249:55–64. doi: 10.1016/j.taap.2010.08.014. [DOI] [PubMed] [Google Scholar]
- El Gendy MA, Somayaji V, El-Kadi AO. Peganum harmala L. is a candidate herbal plant for preventing dioxin mediated effects. Planta Med. 2010;76:671–677. doi: 10.1055/s-0029-1240633. [DOI] [PubMed] [Google Scholar]
- Herraiz T, Gonzalez D, Ancin-Azpilicueta C, Aran VJ, Guillen H. beta-Carboline alkaloids in Peganum harmala and inhibition of human monoamine oxidase (MAO) Food Chem Toxicol. 2010;48:839–845. doi: 10.1016/j.fct.2009.12.019. [DOI] [PubMed] [Google Scholar]
- Im JH, Jin YR, Lee JJ, Yu JY, Han XH, Im SH, Hong JT, Yoo HS, Pyo MY, Yun YP. Antiplatelet activity of beta-carboline alkaloids from Perganum harmala: a possible mechanism through inhibiting PLCgamma2 phosphorylation. Vascul Pharmacol. 2009;50:147–152. doi: 10.1016/j.vph.2008.11.008. [DOI] [PubMed] [Google Scholar]
- Lorenzen A, Kennedy SW. A fluorescence-based protein assay for use with a microplate reader. Anal Biochem. 1993;214:346–348. doi: 10.1006/abio.1993.1504. [DOI] [PubMed] [Google Scholar]
- Mandal PK. Dioxin: a review of its environmental effects and its aryl hydrocarbon receptor biology. J Comp Physiol B. 2005;175:221–230. doi: 10.1007/s00360-005-0483-3. [DOI] [PubMed] [Google Scholar]
- Mosmann T. Rapid colorimetric assay for cellular growth and survival: application to proliferation and cytotoxicity assays. J Immunol Methods. 1983;65:55–63. doi: 10.1016/0022-1759(83)90303-4. [DOI] [PubMed] [Google Scholar]
- Moura DJ, Richter MF, Boeira JM, Pegas Henriques JA, Saffi J. Antioxidant properties of beta-carboline alkaloids are related to their antimutagenic and antigenotoxic activities. Mutagenesis. 2007;22:293–302. doi: 10.1093/mutage/gem016. [DOI] [PubMed] [Google Scholar]
- Murray IA, Flaveny CA, DiNatale BC, Chairo CR, Schroeder JC, Kusnadi A, Perdew GH. Antagonism of aryl hydrocarbon receptor signaling by 6,2′,4′-trimethoxyflavone. J Pharmacol Exp Ther. 2010;332:135–144. doi: 10.1124/jpet.109.158261. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pollenz RS. Specific blockage of ligand-induced degradation of the Ah receptor by proteasome but not calpain inhibitors in cell culture lines from different species. Biochem Pharmacol. 2007;74:131–143. doi: 10.1016/j.bcp.2007.03.021. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Puppala D, Lee H, Kim KB, Swanson HI. Development of an aryl hydrocarbon receptor antagonist using the proteolysis-targeting chimeric molecules approach: a potential tool for chemoprevention. Mol Pharmacol. 2008;73:1064–1071. doi: 10.1124/mol.107.040840. [DOI] [PubMed] [Google Scholar]
- Revel A, Raanani H, Younglai E, Xu J, Rogers I, Han R, Savouret JF, Casper RF. Resveratrol, a natural aryl hydrocarbon receptor antagonist, protects lung from DNA damage and apoptosis caused by benzo[a]pyrene. J Appl Toxicol. 2003;23:255–261. doi: 10.1002/jat.916. [DOI] [PubMed] [Google Scholar]
- Samoylenko V, Rahman MM, Tekwani BL, Tripathi LM, Wang YH, Khan SI, Khan IA, Miller LS, Joshi VC, Muhammad I. Banisteriopsis caapi, a unique combination of MAO inhibitory and antioxidative constituents for the activities relevant to neurodegenerative disorders and Parkinson’s disease. J Ethnopharmacol. 2010;127:357–367. doi: 10.1016/j.jep.2009.10.030. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shah PP, Saurabh K, Pant MC, Mathur N, Parmar D. Evidence for increased cytochrome P450 1A1 expression in blood lymphocytes of lung cancer patients. Mutat Res. 2009;670:74–78. doi: 10.1016/j.mrfmmm.2009.07.006. [DOI] [PubMed] [Google Scholar]
- Shimada T, Fujii-Kuriyama Y. Metabolic activation of polycyclic aromatic hydrocarbons to carcinogens by cytochromes P450 1A1 and 1B1. Cancer Sci. 2004;95:1–6. doi: 10.1111/j.1349-7006.2004.tb03162.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Signorelli P, Ghidoni R. Resveratrol as an anticancer nutrient: molecular basis, open questions and promises. J Nutr Biochem. 2005;16:449–466. doi: 10.1016/j.jnutbio.2005.01.017. [DOI] [PubMed] [Google Scholar]
- Sinal CJ, Bend JR. Aryl hydrocarbon receptor-dependent induction of cyp1a1 by bilirubin in mouse hepatoma hepa 1c1c7 cells. Mol Pharmacol. 1997;52:590–599. doi: 10.1124/mol.52.4.590. [DOI] [PubMed] [Google Scholar]
- Slattery ML, Samowtiz W, Ma K, Murtaugh M, Sweeney C, Levin TR, Neuhausen S. CYP1A1, cigarette smoking, and colon and rectal cancer. Am J Epidemiol. 2004;160:842–852. doi: 10.1093/aje/kwh298. [DOI] [PubMed] [Google Scholar]
- Song Y, Kesuma D, Wang J, Deng Y, Duan J, Wang JH, Qi RZ. Specific inhibition of cyclin-dependent kinases and cell proliferation by harmine. Biochem Biophys Res Commun. 2004;317:128–132. doi: 10.1016/j.bbrc.2004.03.019. [DOI] [PubMed] [Google Scholar]
- Suzuki T, Nohara K. Regulatory factors involved in species-specific modulation of arylhydrocarbon receptor (AhR)-dependent gene expression in humans and mice. Journal of biochemistry. 2007;142:443–452. doi: 10.1093/jb/mvm152. [DOI] [PubMed] [Google Scholar]
- Taguchi K, Motohashi H, Yamamoto M. Molecular mechanisms of the Keap1-Nrf2 pathway in stress response and cancer evolution. Genes Cells. 2011;16:123–140. doi: 10.1111/j.1365-2443.2010.01473.x. [DOI] [PubMed] [Google Scholar]
- Wiseman SB, Vijayan MM. Aryl hydrocarbon receptor signaling in rainbow trout hepatocytes: role of hsp90 and the proteasome. Comp Biochem Physiol C Toxicol Pharmacol. 2007;146:484–491. doi: 10.1016/j.cbpc.2007.05.006. [DOI] [PubMed] [Google Scholar]
- Yu AM, Idle JR, Krausz KW, Kupfer A, Gonzalez FJ. Contribution of individual cytochrome P450 isozymes to the O-demethylation of the psychotropic beta-carboline alkaloids harmaline and harmine. J Pharmacol Exp Ther. 2003;305:315–322. doi: 10.1124/jpet.102.047050. [DOI] [PubMed] [Google Scholar]
- Zhang S, Qin C, Safe SH. Flavonoids as aryl hydrocarbon receptor agonists/antagonists: effects of structure and cell context. Environ Health Perspect. 2003;111:1877–1882. doi: 10.1289/ehp.6322. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhao B, Degroot DE, Hayashi A, He G, Denison MS. CH223191 is a ligand-selective antagonist of the Ah (Dioxin) receptor. Toxicol Sci. 2010;117:393–403. doi: 10.1093/toxsci/kfq217. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhou J, Gasiewicz TA. 3′-methoxy-4′-nitroflavone, a reported aryl hydrocarbon receptor antagonist, enhances Cyp1a1 transcription by a dioxin responsive element-dependent mechanism. Arch Biochem Biophys. 2003;416:68–80. doi: 10.1016/s0003-9861(03)00274-1. [DOI] [PubMed] [Google Scholar]
- Zhou JG, Henry EC, Palermo CM, Dertinger SD, Gasiewicz TA. Species-specific transcriptional activity of synthetic flavonoids in guinea pig and mouse cells as a result of differential activation of the aryl hydrocarbon receptor to interact with dioxin-responsive elements. Mol Pharmacol. 2003;63:915–924. doi: 10.1124/mol.63.4.915. [DOI] [PubMed] [Google Scholar]