Abstract
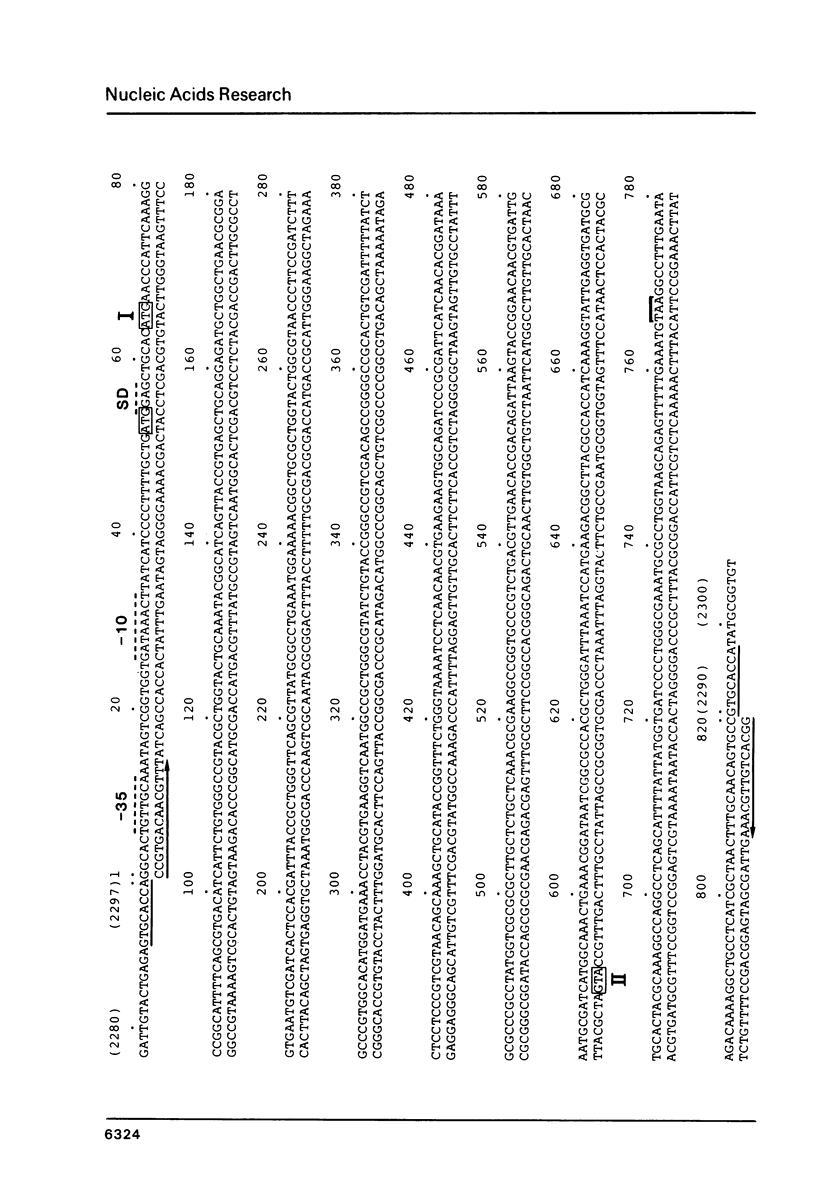
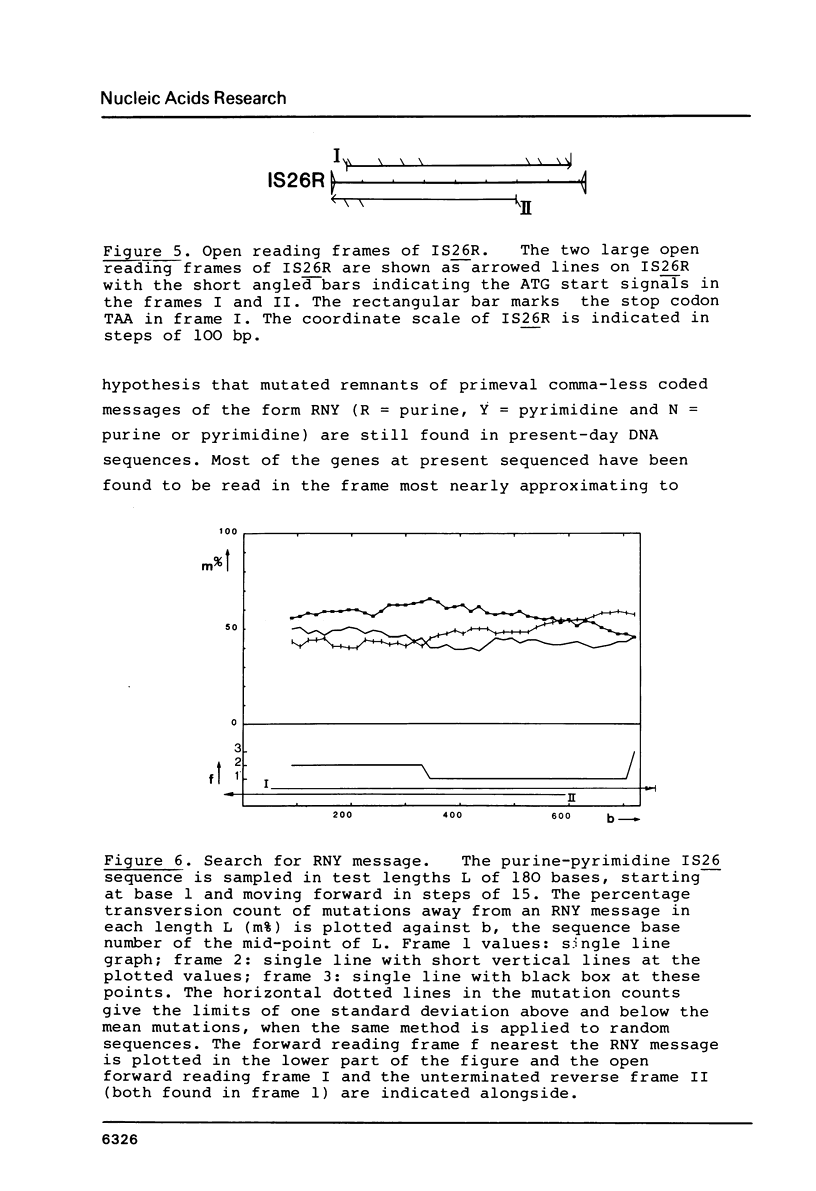
The DNA sequence of a new IS element, the IS26, is 820 bp long and carries 14 bp perfect terminal inverted repeats. Upon integration, IS26 generates an 8 bp duplication of its target sequence. A large open reading frame within IS26 could code for a protein of 234 amino acids. On its reverse strand, IS26 also carries one large open reading frame, 591 bp long, which contains no stop codon within IS26.
Full text
PDF
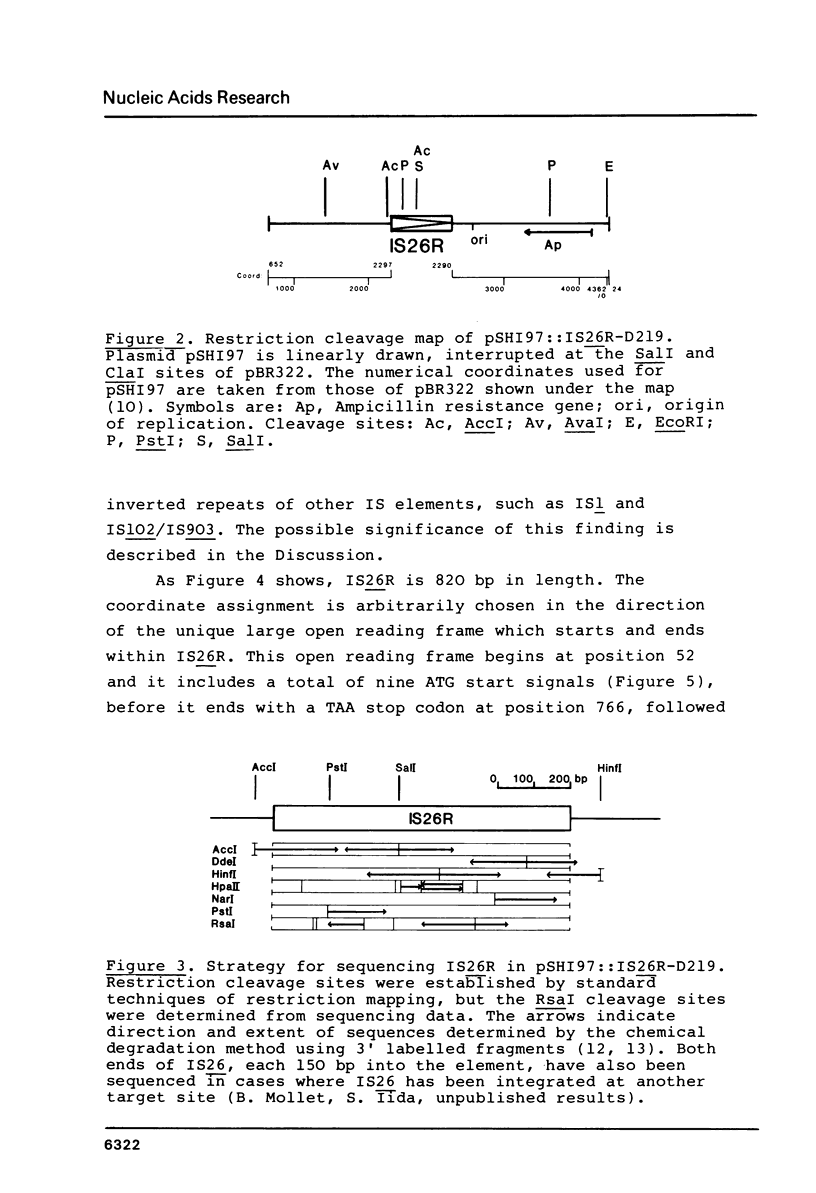









Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Auerswald E. A., Ludwig G., Schaller H. Structural analysis of Tn5. Cold Spring Harb Symp Quant Biol. 1981;45(Pt 1):107–113. doi: 10.1101/sqb.1981.045.01.019. [DOI] [PubMed] [Google Scholar]
- Bernardi A., Bernardi F. Complete sequence of an IS element present in pSC101. Nucleic Acids Res. 1981 Jun 25;9(12):2905–2911. doi: 10.1093/nar/9.12.2905. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bolivar F., Rodriguez R. L., Greene P. J., Betlach M. C., Heyneker H. L., Boyer H. W., Crosa J. H., Falkow S. Construction and characterization of new cloning vehicles. II. A multipurpose cloning system. Gene. 1977;2(2):95–113. [PubMed] [Google Scholar]
- Bräu B., Piepersberg W. Cointegrational transduction and mobilization of gentamicin resistance plasmid pWP14a is mediated by IS140. Mol Gen Genet. 1983;189(2):298–303. doi: 10.1007/BF00337820. [DOI] [PubMed] [Google Scholar]
- Calos M. P., Johnsrud L., Miller J. H. DNA sequence at the integration sites of the insertion element IS1. Cell. 1978 Mar;13(3):411–418. doi: 10.1016/0092-8674(78)90315-x. [DOI] [PubMed] [Google Scholar]
- Calos M. P., Miller J. H. Transposable elements. Cell. 1980 Jul;20(3):579–595. doi: 10.1016/0092-8674(80)90305-0. [DOI] [PubMed] [Google Scholar]
- Campbell A., Berg D. E., Botstein D., Lederberg E. M., Novick R. P., Starlinger P., Szybalski W. Nomenclature of transposable elements in prokaryotes. Gene. 1979 Mar;5(3):197–206. doi: 10.1016/0378-1119(79)90078-7. [DOI] [PubMed] [Google Scholar]
- Campbell A., Starlinger P., Berg D. E., Botstein D., Lederberg E. M., Novick R. P., Szybalski W. Nomenclature of transposable elements in prokaryotes. Plasmid. 1979 Jul;2(3):466–473. doi: 10.1016/0147-619x(79)90030-1. [DOI] [PubMed] [Google Scholar]
- Engler J. A., van Bree M. P. The nucleotide sequence and protein-coding capability of the transposable element IS5. Gene. 1981 Aug;14(3):155–163. doi: 10.1016/0378-1119(81)90111-6. [DOI] [PubMed] [Google Scholar]
- Ghosal D., Sommer H., Saedler H. Nucleotide sequence of the transposable DNA-element IS2. Nucleic Acids Res. 1979 Mar;6(3):1111–1122. doi: 10.1093/nar/6.3.1111. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Grindley N. D. IS1 insertion generates duplication of a nine base pair sequence at its target site. Cell. 1978 Mar;13(3):419–426. doi: 10.1016/0092-8674(78)90316-1. [DOI] [PubMed] [Google Scholar]
- Grindley N. D., Joyce C. M. Genetic and DNA sequence analysis of the kanamycin resistance transposon Tn903. Proc Natl Acad Sci U S A. 1980 Dec;77(12):7176–7180. doi: 10.1073/pnas.77.12.7176. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Grindley N. D., Sherratt D. J. Sequence analysis at IS1 insertion sites: models for transposition. Cold Spring Harb Symp Quant Biol. 1979;43(Pt 2):1257–1261. doi: 10.1101/sqb.1979.043.01.142. [DOI] [PubMed] [Google Scholar]
- Halling S. M., Kleckner N. A symmetrical six-base-pair target site sequence determines Tn10 insertion specificity. Cell. 1982 Jan;28(1):155–163. doi: 10.1016/0092-8674(82)90385-3. [DOI] [PubMed] [Google Scholar]
- Halling S. M., Simons R. W., Way J. C., Walsh R. B., Kleckner N. DNA sequence organization of IS10-right of Tn10 and comparison with IS10-left. Proc Natl Acad Sci U S A. 1982 Apr;79(8):2608–2612. doi: 10.1073/pnas.79.8.2608. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Iida S., Marcoli R., Bickle T. A. Variant insertion element IS1 generates 8-base pair duplications of the target sequence. Nature. 1981 Nov 26;294(5839):374–376. doi: 10.1038/294374a0. [DOI] [PubMed] [Google Scholar]
- Iida S., Meyer J., Arber W. Genesis and natural history of IS-mediated transposons. Cold Spring Harb Symp Quant Biol. 1981;45(Pt 1):27–43. doi: 10.1101/sqb.1981.045.01.006. [DOI] [PubMed] [Google Scholar]
- Iida S., Meyer J., Linder P., Goto N., Nakaya R., Reif H. J., Arber W. The kanamycin resistance transposon Tn2680 derived from the R plasmid Rts1 and carried by phage P1Km has flanking 0.8-kb-long direct repeats. Plasmid. 1982 Sep;8(2):187–198. doi: 10.1016/0147-619x(82)90056-7. [DOI] [PubMed] [Google Scholar]
- Johnsrud L. DNA sequence of the transposable element IS1. Mol Gen Genet. 1979 Jan 31;169(2):213–218. doi: 10.1007/BF00271673. [DOI] [PubMed] [Google Scholar]
- Klaer R., Kühn S., Tillmann E., Fritz H. J., Starlinger P. The sequence of IS4. Mol Gen Genet. 1981;181(2):169–175. doi: 10.1007/BF00268423. [DOI] [PubMed] [Google Scholar]
- Kleckner N. Transposable elements in prokaryotes. Annu Rev Genet. 1981;15:341–404. doi: 10.1146/annurev.ge.15.120181.002013. [DOI] [PubMed] [Google Scholar]
- Kröger M., Hobom G. Structural analysis of insertion sequence IS5. Nature. 1982 May 13;297(5862):159–162. doi: 10.1038/297159a0. [DOI] [PubMed] [Google Scholar]
- Labigne-Roussel A., Briaux-Gerbaud S., Courvalin P. Tn1525, a kanamycin R determinant flanked by two direct copies of IS15. Mol Gen Genet. 1983;189(1):90–101. doi: 10.1007/BF00326060. [DOI] [PubMed] [Google Scholar]
- Maxam A. M., Gilbert W. A new method for sequencing DNA. Proc Natl Acad Sci U S A. 1977 Feb;74(2):560–564. doi: 10.1073/pnas.74.2.560. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Maxam A. M., Gilbert W. Sequencing end-labeled DNA with base-specific chemical cleavages. Methods Enzymol. 1980;65(1):499–560. doi: 10.1016/s0076-6879(80)65059-9. [DOI] [PubMed] [Google Scholar]
- Ohtsubo H., Nyman K., Doroszkiewicz W., Ohtsubo E. Multiple copies of iso-insertion sequences of IS1 in Shigella dysenteriae chromosome. Nature. 1981 Aug 13;292(5824):640–643. doi: 10.1038/292640a0. [DOI] [PubMed] [Google Scholar]
- Ohtsubo H., Ohtsubo E. Nucleotide sequence of an insertion element, IS1. Proc Natl Acad Sci U S A. 1978 Feb;75(2):615–619. doi: 10.1073/pnas.75.2.615. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ohtsubo H., Zenilman M., Ohtsubo E. Insertion element IS102 resides in plasmid pSC101. J Bacteriol. 1980 Oct;144(1):131–140. doi: 10.1128/jb.144.1.131-140.1980. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Oka A., Nomura N., Sugimoto, Sugisaki H., Takanami M. Nucleotide sequence at the insertion sites of a kanamycin transposon. Nature. 1978 Dec 21;276(5690):845–847. doi: 10.1038/276845a0. [DOI] [PubMed] [Google Scholar]
- Oka A., Sugisaki H., Takanami M. Nucleotide sequence of the kanamycin resistance transposon Tn903. J Mol Biol. 1981 Apr 5;147(2):217–226. doi: 10.1016/0022-2836(81)90438-1. [DOI] [PubMed] [Google Scholar]
- Rosenberg M., Court D. Regulatory sequences involved in the promotion and termination of RNA transcription. Annu Rev Genet. 1979;13:319–353. doi: 10.1146/annurev.ge.13.120179.001535. [DOI] [PubMed] [Google Scholar]
- Schoner B., Kahn M. The nucleotide sequence of IS5 from Escherichia coli. Gene. 1981 Aug;14(3):165–174. doi: 10.1016/0378-1119(81)90112-8. [DOI] [PubMed] [Google Scholar]
- Shepherd J. C. From primeval message to present-day gene. Cold Spring Harb Symp Quant Biol. 1983;47(Pt 2):1099–1108. doi: 10.1101/sqb.1983.047.01.124. [DOI] [PubMed] [Google Scholar]
- Shepherd J. C. Method to determine the reading frame of a protein from the purine/pyrimidine genome sequence and its possible evolutionary justification. Proc Natl Acad Sci U S A. 1981 Mar;78(3):1596–1600. doi: 10.1073/pnas.78.3.1596. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shine J., Dalgarno L. The 3'-terminal sequence of Escherichia coli 16S ribosomal RNA: complementarity to nonsense triplets and ribosome binding sites. Proc Natl Acad Sci U S A. 1974 Apr;71(4):1342–1346. doi: 10.1073/pnas.71.4.1342. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Simsek M., DasSarma S., RajBhandary U. L., Khorana H. G. A transposable element from Halobacterium halobium which inactivates the bacteriorhodopsin gene. Proc Natl Acad Sci U S A. 1982 Dec;79(23):7268–7272. doi: 10.1073/pnas.79.23.7268. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sutcliffe J. G. Complete nucleotide sequence of the Escherichia coli plasmid pBR322. Cold Spring Harb Symp Quant Biol. 1979;43(Pt 1):77–90. doi: 10.1101/sqb.1979.043.01.013. [DOI] [PubMed] [Google Scholar]
- Takano T., Ikeda S. Phage P1 carrying kanamycin resistance gene of R factor. Virology. 1976 Mar;70(1):198–200. doi: 10.1016/0042-6822(76)90252-x. [DOI] [PubMed] [Google Scholar]
- Terawaki Y., Takayasu H., Akiba T. Thermosensitive replication of a kanamycin resistance factor. J Bacteriol. 1967 Sep;94(3):687–690. doi: 10.1128/jb.94.3.687-690.1967. [DOI] [PMC free article] [PubMed] [Google Scholar]



