Abstract
The over-all 5-methylcytosine (m5C) content of DNA from normal tissues varies considerably in a tissue-specific manner. By high-performance liquid chromatography, we have examined the m5C contents of enzymatic digests of DNA from 103 human tumors including benign, primary malignant and secondary malignant neoplasms. The diversity and large number of these tumor samples allowed us to compare the range of DNA methylation levels from neoplastic tissues to that of normal tissues from humans. Most of the metastatic neoplasms had significantly lower genomic m5C contents than did most of the benign neoplasms or normal tissues. The percentage of primary malignancies with hypomethylated DNA was intermediate between those of metastases and benign neoplasms. These findings might reflect an involvement of extensive demethylation of DNA in tumor progression. Such demethylation could be a source of the continually generated cellular diversity associated with cancer.
Full text
PDF


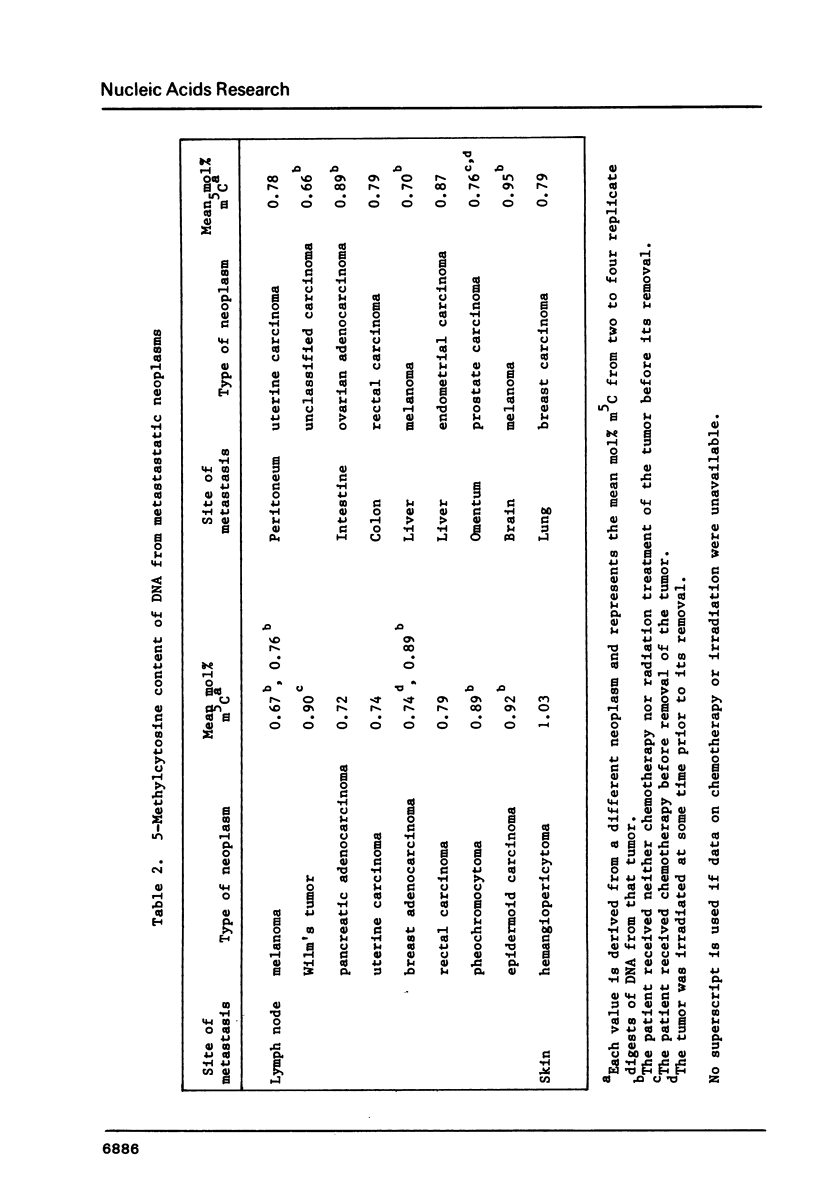
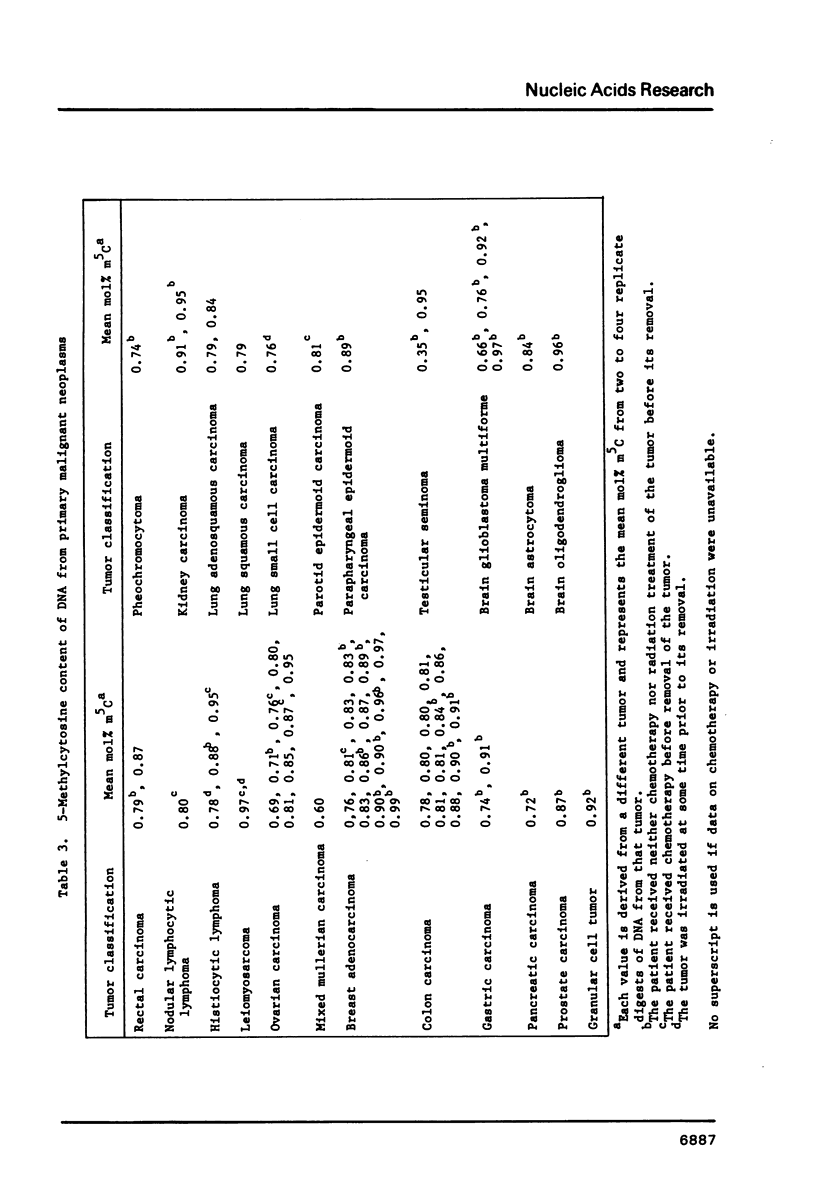







Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Bagshawe K. D. Some facets of trophoblastic neoplasia in man. J Reprod Fertil Suppl. 1982 Nov;31:175–179. [PubMed] [Google Scholar]
- Barlogie B., Drewinko B., Schumann J., Göhde W., Dosik G., Latreille J., Johnston D. A., Freireich E. J. Cellular DNA content as a marker of neoplasia in man. Am J Med. 1980 Aug;69(2):195–203. doi: 10.1016/0002-9343(80)90379-4. [DOI] [PubMed] [Google Scholar]
- Busslinger M., deBoer E., Wright S., Grosveld F. G., Flavell R. A. The sequence GGCmCGG is resistant to MspI cleavage. Nucleic Acids Res. 1983 Jun 11;11(11):3559–3569. doi: 10.1093/nar/11.11.3559. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Christy B., Scangos G. Expression of transferred thymidine kinase genes is controlled by methylation. Proc Natl Acad Sci U S A. 1982 Oct;79(20):6299–6303. doi: 10.1073/pnas.79.20.6299. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Compere S. J., Palmiter R. D. DNA methylation controls the inducibility of the mouse metallothionein-I gene lymphoid cells. Cell. 1981 Jul;25(1):233–240. doi: 10.1016/0092-8674(81)90248-8. [DOI] [PubMed] [Google Scholar]
- Cooper D. N., Taggart M. H., Bird A. P. Unmethylated domains in vertebrate DNA. Nucleic Acids Res. 1983 Feb 11;11(3):647–658. doi: 10.1093/nar/11.3.647. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Creusot F., Acs G., Christman J. K. Inhibition of DNA methyltransferase and induction of Friend erythroleukemia cell differentiation by 5-azacytidine and 5-aza-2'-deoxycytidine. J Biol Chem. 1982 Feb 25;257(4):2041–2048. [PubMed] [Google Scholar]
- Doerfler W. DNA methylation--a regulatory signal in eukaryotic gene expression. J Gen Virol. 1981 Nov;57(Pt 1):1–20. doi: 10.1099/0022-1317-57-1-1. [DOI] [PubMed] [Google Scholar]
- Ehrlich M., Gama-Sosa M. A., Huang L. H., Midgett R. M., Kuo K. C., McCune R. A., Gehrke C. Amount and distribution of 5-methylcytosine in human DNA from different types of tissues of cells. Nucleic Acids Res. 1982 Apr 24;10(8):2709–2721. doi: 10.1093/nar/10.8.2709. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ehrlich M., Wang R. Y. 5-Methylcytosine in eukaryotic DNA. Science. 1981 Jun 19;212(4501):1350–1357. doi: 10.1126/science.6262918. [DOI] [PubMed] [Google Scholar]
- Feinberg A. P., Vogelstein B. Hypomethylation distinguishes genes of some human cancers from their normal counterparts. Nature. 1983 Jan 6;301(5895):89–92. doi: 10.1038/301089a0. [DOI] [PubMed] [Google Scholar]
- Feinberg A. P., Vogelstein B. Hypomethylation of ras oncogenes in primary human cancers. Biochem Biophys Res Commun. 1983 Feb 28;111(1):47–54. doi: 10.1016/s0006-291x(83)80115-6. [DOI] [PubMed] [Google Scholar]
- Fidler I. J., Hart I. R. Biological diversity in metastatic neoplasms: origins and implications. Science. 1982 Sep 10;217(4564):998–1003. doi: 10.1126/science.7112116. [DOI] [PubMed] [Google Scholar]
- Gama-Sosa M. A., Midgett R. M., Slagel V. A., Githens S., Kuo K. C., Gehrke C. W., Ehrlich M. Tissue-specific differences in DNA methylation in various mammals. Biochim Biophys Acta. 1983 Jun 24;740(2):212–219. doi: 10.1016/0167-4781(83)90079-9. [DOI] [PubMed] [Google Scholar]
- Gama-Sosa M. A., Wang R. Y., Kuo K. C., Gehrke C. W., Ehrlich M. The 5-methylcytosine content of highly repeated sequences in human DNA. Nucleic Acids Res. 1983 May 25;11(10):3087–3095. doi: 10.1093/nar/11.10.3087. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hirt B. Selective extraction of polyoma DNA from infected mouse cell cultures. J Mol Biol. 1967 Jun 14;26(2):365–369. doi: 10.1016/0022-2836(67)90307-5. [DOI] [PubMed] [Google Scholar]
- Jones P. A., Taylor S. M. Cellular differentiation, cytidine analogs and DNA methylation. Cell. 1980 May;20(1):85–93. doi: 10.1016/0092-8674(80)90237-8. [DOI] [PubMed] [Google Scholar]
- Jähner D., Stuhlmann H., Stewart C. L., Harbers K., Löhler J., Simon I., Jaenisch R. De novo methylation and expression of retroviral genomes during mouse embryogenesis. Nature. 1982 Aug 12;298(5875):623–628. doi: 10.1038/298623a0. [DOI] [PubMed] [Google Scholar]
- Kappler J. W. The 5-methylcytosine content of DNA: tissue specificity. J Cell Physiol. 1971 Aug;78(1):33–36. doi: 10.1002/jcp.1040780106. [DOI] [PubMed] [Google Scholar]
- Keshet E., Cedar H. Effect of CpG methylation on Msp I. Nucleic Acids Res. 1983 Jun 11;11(11):3571–3580. doi: 10.1093/nar/11.11.3571. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kuo K. C., McCune R. A., Gehrke C. W., Midgett R., Ehrlich M. Quantitative reversed-phase high performance liquid chromatographic determination of major and modified deoxyribonucleosides in DNA. Nucleic Acids Res. 1980 Oct 24;8(20):4763–4776. doi: 10.1093/nar/8.20.4763. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lapeyre J. N., Becker F. F. 5-Methylcytosine content of nuclear DNA during chemical hepatocarcinogenesis and in carcinomas which result. Biochem Biophys Res Commun. 1979 Apr 13;87(3):698–705. doi: 10.1016/0006-291x(79)92015-1. [DOI] [PubMed] [Google Scholar]
- Mermod J. J., Bourgeois S., Defer N., Crépin M. Demethylation and expression of murine mammary tumor proviruses in mouse thymoma cell lines. Proc Natl Acad Sci U S A. 1983 Jan;80(1):110–114. doi: 10.1073/pnas.80.1.110. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Stein R., Razin A., Cedar H. In vitro methylation of the hamster adenine phosphoribosyltransferase gene inhibits its expression in mouse L cells. Proc Natl Acad Sci U S A. 1982 Jun;79(11):3418–3422. doi: 10.1073/pnas.79.11.3418. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vanyushin B. F., Mazin A. L., Vasilyev V. K., Belozersky A. N. The content of 5-methylcytosine in animal DNA: the species and tissue specificity. Biochim Biophys Acta. 1973 Mar 28;299(3):397–403. doi: 10.1016/0005-2787(73)90264-5. [DOI] [PubMed] [Google Scholar]
- Wang R. Y., Shedlarski J. G., Farber M. B., Kuebbing D., Ehrlich M. Two sequence-specific endonucleases from Xanthomonas oryzae. Characterization and unusual properties. Biochim Biophys Acta. 1980 Feb 29;606(2):371–385. doi: 10.1016/0005-2787(80)90047-7. [DOI] [PubMed] [Google Scholar]
- van der Ploeg L. H., Flavell R. A. DNA methylation in the human gamma delta beta-globin locus in erythroid and nonerythroid tissues. Cell. 1980 Apr;19(4):947–958. doi: 10.1016/0092-8674(80)90086-0. [DOI] [PubMed] [Google Scholar]