Abstract
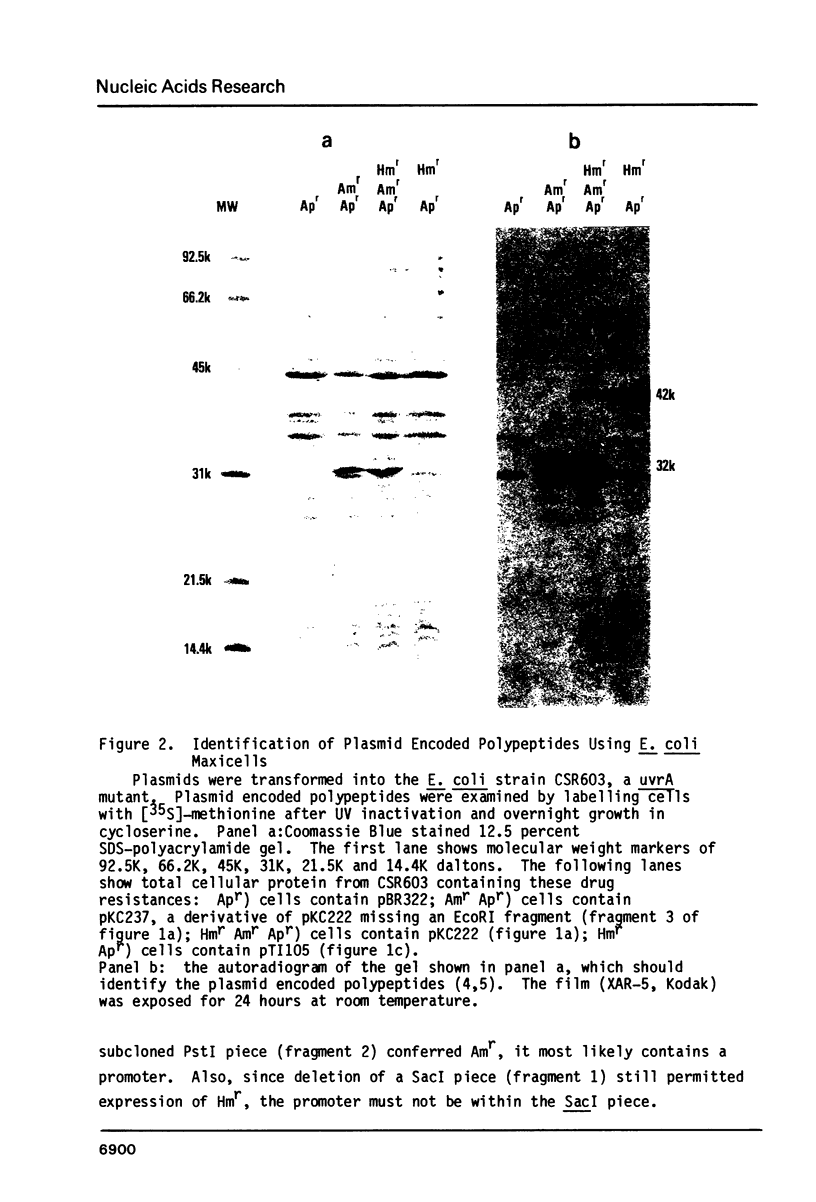
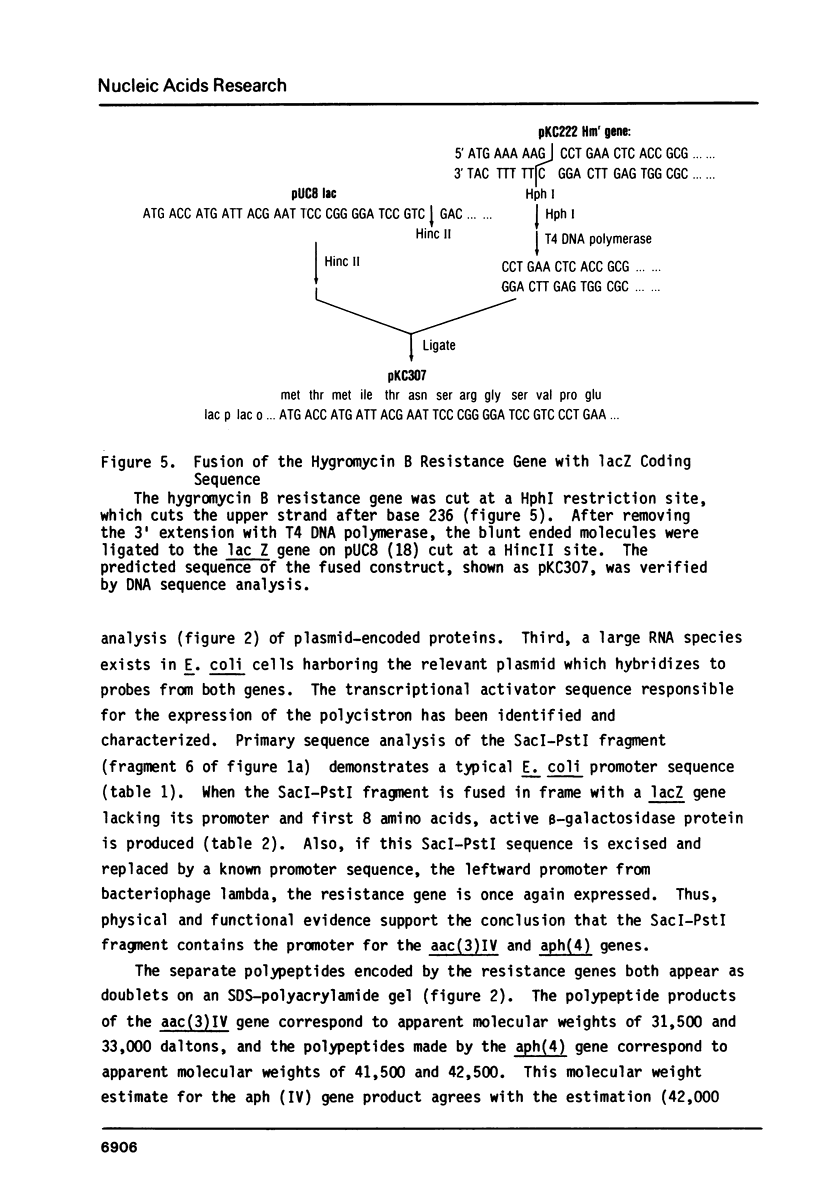
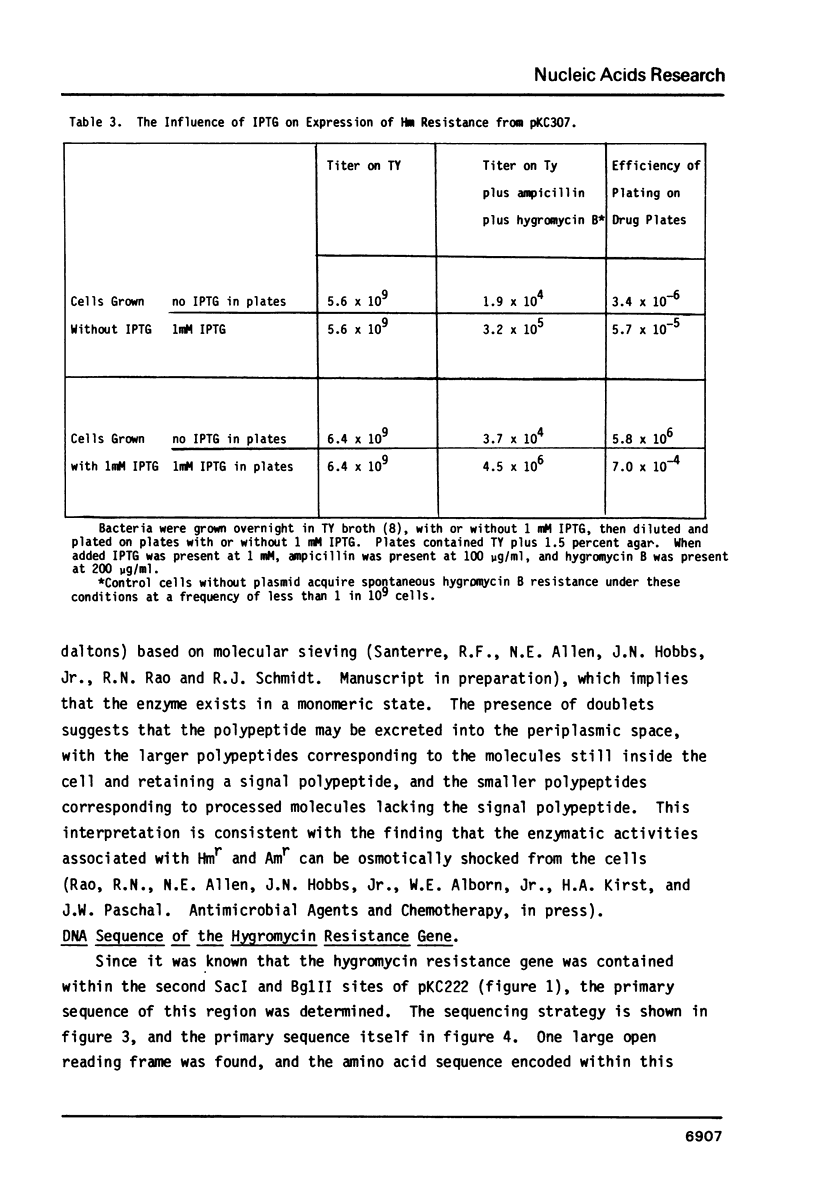
We have characterized hygromycin B and apramycin resistance genes from an E. coli plasmid. We have localized the coding and control regions of these genes by deletion of DNA fragments from plasmids containing the genes. It was found that polypeptides with apparent molecular weights of 33,000 and 31,500 daltons are encoded by the apramycin resistance gene and polypeptides with apparent molecular weights of 42,500 and 41,500 daltons are encoded by the hygromycin B resistance gene. DNA sequence analysis identified a typical promoter sequence upstream of the genes. Deletion of this promoter eliminated both resistance phenotypes, and hygromycin B resistance could be restored by substitution of a promoter from a foreign gene. The region known to be necessary for hygromycin B resistance contained an open reading frame large enough to encode the hygromycin B resistance gene product. This open reading frame was fused with the amino terminus of beta-galactosidase. This hybrid gene conferred hygromycin resistance to E. coli, and expression of resistance was under IPTG control.
Full text
PDF













Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Ames G. F. Resolution of bacterial proteins by polyacrylamide gel electrophoresis on slabs. Membrane, soluble, and periplasmic fractions. J Biol Chem. 1974 Jan 25;249(2):634–644. [PubMed] [Google Scholar]
- Beck E., Ludwig G., Auerswald E. A., Reiss B., Schaller H. Nucleotide sequence and exact localization of the neomycin phosphotransferase gene from transposon Tn5. Gene. 1982 Oct;19(3):327–336. doi: 10.1016/0378-1119(82)90023-3. [DOI] [PubMed] [Google Scholar]
- Casadaban M. J., Chou J., Cohen S. N. In vitro gene fusions that join an enzymatically active beta-galactosidase segment to amino-terminal fragments of exogenous proteins: Escherichia coli plasmid vectors for the detection and cloning of translational initiation signals. J Bacteriol. 1980 Aug;143(2):971–980. doi: 10.1128/jb.143.2.971-980.1980. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Davies J., O'Connor S. Enzymatic modification of aminoglycoside antibiotics: 3-N-acetyltransferase with broad specificity that determines resistance to the novel aminoglycoside apramycin. Antimicrob Agents Chemother. 1978 Jul;14(1):69–72. doi: 10.1128/aac.14.1.69. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Franklin N. C., Bennett G. N. The N protein of bacteriophage lambda, defined by its DNA sequence, is highly basic. Gene. 1979 Dec;8(1):107–119. doi: 10.1016/0378-1119(79)90011-8. [DOI] [PubMed] [Google Scholar]
- González A., Jiménez A., Vázquez D., Davies J. E., Schindler D. Studies on the mode of action of hygromycin B, an inhibitor of translocation in eukaryotes. Biochim Biophys Acta. 1978 Dec 21;521(2):459–469. doi: 10.1016/0005-2787(78)90287-3. [DOI] [PubMed] [Google Scholar]
- Ingolia T. D., Craig E. A. Primary sequence of the 5' flanking regions of the Drosophila heat shock genes in chromosome subdivision 67B. Nucleic Acids Res. 1981 Apr 10;9(7):1627–1642. doi: 10.1093/nar/9.7.1627. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Maxam A. M., Gilbert W. A new method for sequencing DNA. Proc Natl Acad Sci U S A. 1977 Feb;74(2):560–564. doi: 10.1073/pnas.74.2.560. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Oka A., Sugisaki H., Takanami M. Nucleotide sequence of the kanamycin resistance transposon Tn903. J Mol Biol. 1981 Apr 5;147(2):217–226. doi: 10.1016/0022-2836(81)90438-1. [DOI] [PubMed] [Google Scholar]
- Rosenberg M., Court D. Regulatory sequences involved in the promotion and termination of RNA transcription. Annu Rev Genet. 1979;13:319–353. doi: 10.1146/annurev.ge.13.120179.001535. [DOI] [PubMed] [Google Scholar]
- Sancar A., Hack A. M., Rupp W. D. Simple method for identification of plasmid-coded proteins. J Bacteriol. 1979 Jan;137(1):692–693. doi: 10.1128/jb.137.1.692-693.1979. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sancar A., Rupert C. S. Determination of plasmid molecular weights from ultraviolet sensitivities. Nature. 1978 Mar 30;272(5652):471–472. doi: 10.1038/272471a0. [DOI] [PubMed] [Google Scholar]
- Scherer G. F., Walkinshaw M. D., Arnott S., Morré D. J. The ribosome binding sites recognized by E. coli ribosomes have regions with signal character in both the leader and protein coding segments. Nucleic Acids Res. 1980 Sep 11;8(17):3895–3907. doi: 10.1093/nar/8.17.3895. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sutcliffe J. G. Complete nucleotide sequence of the Escherichia coli plasmid pBR322. Cold Spring Harb Symp Quant Biol. 1979;43(Pt 1):77–90. doi: 10.1101/sqb.1979.043.01.013. [DOI] [PubMed] [Google Scholar]
- Thomas P. S. Hybridization of denatured RNA and small DNA fragments transferred to nitrocellulose. Proc Natl Acad Sci U S A. 1980 Sep;77(9):5201–5205. doi: 10.1073/pnas.77.9.5201. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vieira J., Messing J. The pUC plasmids, an M13mp7-derived system for insertion mutagenesis and sequencing with synthetic universal primers. Gene. 1982 Oct;19(3):259–268. doi: 10.1016/0378-1119(82)90015-4. [DOI] [PubMed] [Google Scholar]
- West R. W., Jr, Neve R. L., Rodriguez R. L. Construction and characterization of E. coli promoter-probe plasmid vectors. I. Cloning of promoter-containing DNA fragments. Gene. 1979 Nov;7(3-4):271–288. doi: 10.1016/0378-1119(79)90048-9. [DOI] [PubMed] [Google Scholar]
- Wozney J., Hanahan D., Morimoto R., Boedtker H., Doty P. Fine structural analysis of the chicken pro alpha 2 collagen gene. Proc Natl Acad Sci U S A. 1981 Feb;78(2):712–716. doi: 10.1073/pnas.78.2.712. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zaret K. S., Sherman F. DNA sequence required for efficient transcription termination in yeast. Cell. 1982 Mar;28(3):563–573. doi: 10.1016/0092-8674(82)90211-2. [DOI] [PubMed] [Google Scholar]




